Abstract
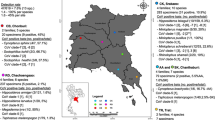
This is the first country-wide surveillance of bat-borne viruses in Kenya spanning from 2012–2015 covering sites perceived to have medium to high level bat-human interaction. The objective of this surveillance study was to apply a non-invasive approach using fresh feces to detect viruses circulating within the diverse species of Kenyan bats. We screened for both DNA and RNA viruses; specifically, astroviruses (AstVs), adenoviruses (ADVs), caliciviruses (CalVs), coronaviruses (CoVs), flaviviruses, filoviruses, paramyxoviruses (PMVs), polyomaviruses (PYVs) and rotaviruses. We used family-specific primers, amplicon sequencing and further characterization by phylogenetic analysis. Except for filoviruses, eight virus families were detected with varying distributions and positive rates across the five regions (former provinces) studied. AstVs (12.83%), CoVs (3.97%), PMV (2.4%), ADV (2.26%), PYV (1.65%), CalVs (0.29%), rotavirus (0.19%) and flavivirus (0.19%). Novel CalVs were detected in Rousettus aegyptiacus and Mops condylurus while novel Rotavirus-A-related viruses were detected in Taphozous bats and R. aegyptiacus. The two Rotavirus A (RVA) strains detected were highly related to human strains with VP6 genotypes I2 and I16. Genotype I16 has previously been assigned to human RVA-strain B10 from Kenya only, which raises public health concern, particularly considering increased human-bat interaction. Additionally, 229E-like bat CoVs were detected in samples originating from Hipposideros bats roosting in sites with high human activity. Our findings confirm the presence of diverse viruses in Kenyan bats while providing extended knowledge on bat virus distribution. The detection of viruses highly related to human strains and hence of public health concern, underscores the importance of continuous surveillance.

Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Aljofan M. 2015. Henipaviruses are not yet history. Future Virol, 10: 507–515.
Anthony SJ, Epstein JH, Murray KA, Navarrete-Macias I, Zambrana-Torrelio CM., Solovyov A. Ojeda-Flores R, Arrigo NC, Islam A, Ali-Khan S, Hosseini P, Bogich TL, Olival KJ, Sanchez-Leon MD, Karesh WB, Goldstein T, Luby SP, Morse SS, Mazet JAK, Daszak P, Lipkin WI 2013. A strategy to estimate unknown viral diversity in mammals. mBio, 4: e00598.
Asano KM, Gregori F, Hora AS, Scheffer KC, Fahl WO, Iamamoto K, Mori E, Silva FDF, Taniwaki SA, Brandão PE. 2016. Group A rotavirus in Brazilian bats: description of novel T15 and H15 genotypes. Arch of Virol, 161: 3225.
Baker KS, Leggett RM, Bexfield NH, Alston M, Daly G, Todd S, Tachedjian M, Holmes CEG, Crameri S, Wang L-F, Heeney JL, Suu-Ire R, Kellam P, Cunningham AA, Wood JLN, Caccamo M, Murcia PR. 2013. Metagenomic study of the viruses of African straw-coloured fruit bats: Detection of a chiropteran poxvirus and isolation of a novel adenovirus. Virol J, 441: 95–106.
Calisher CH, Childs JE, Field HE, Holmes KV, Schountz T. 2006. Bats: Important reservoir hosts of emerging viruses. Clin Microbiol Rev, 19: 531–545.
Chu DKW, Poon LLM, Guan Y, Peiris JSM. 2008. Novel astroviruses in insectivorous bats. J Virol, 82: 9107–9114.
Chua KB, Bellini WJ, Rota PA, Harcourt BH, Tamin A, Lam SK, Ksiazek TG, Rollin PE, Zaki SR, Shieh WJ, Goldsmith CS, Gubler DJ, Roehrig JT, Eaton B, Gould R, Olson J, Field H, Daniels P, Ling AE, Peters CJ, Anderson LJ, Mahy BW. 2000. Nipah virus: a recently emergent deadly paramyxovirus. Science, 288: 1432–1435.
Conrardy C, Tao Y, Kuzmin IV, Niezgoda M, Agwanda B, Breiman RF, Anderson LJ, Rupprecht CE, Tong S. 2014. Molecular detection of adenoviruses, rhabdoviruses, and paramyxoviruses in bats from Kenya. Am J Trop Med. Hyg, 91: 258–266.
Corman VM, Baldwin HJ, Tateno AF, Zerbinati RM, Annan A, Owusu M, Nkrumah EE, Maganga GD, Oppong S, Adu-Sarkodie Y, Vallo P, Ribeiro LV, Filho F, Leroy EM, Thiel V, Hoek L, Poon LLM, Tschapka M, Drexler JF. 2015. Evidence for an ancestral association of human coronavirus 229E with bats. J Virol, 89: 11858–11870.
Crossley BM, Mock RE, Callison SA, & Hietala SK. 2012. Identification and characterization of a novel alpaca respiratory coronavirus most closely related to the human coronavirus 229E. Viruses, 4: 3689–3700.
Dacheux L, Cervantes-Gonzalez M, Guigon G, Thiberge JM, Vandenbogaert M, Maufrais C, Caro V, Bourhy H. 2014. A preliminary study of viral metagenomics of French bat species in contact with humans: Identification of new mammalian viruses. PLoS ONE, 9: e87194.
Drexler JF, Corman VM, Gloza-Rausch F, Seebens A, Annan A, Ipsen A, Krupa T, Muller MA, Kalko EKV, Adu-Sarkodie Y, Oppong S, Drosten C. 2009. Henipavirus RNA in African bats. PloS One, 4: e6367.
Drexler JF, Corman VM, Wegner T, Tateno AF, Zerbinati RM, Gloza-Rausch F, Seebens A, Muller MA, Drosten C. 2011. Amplification of emerging viruses in a bat colony. Emerg Infect Dis, 17: 449–456.
Esona MD, Banyai K, Foytich K, Freeman M, Mijatovic-Rustempasic S, Hull J, Kerin T, Steele AD, Armahd GE, Geyer A, Page N, Agbaya VA, Forbi JC, Aminui M, Gautama R, Seheri LM, Nyangao J, Glass R, Bowena MD, Gentsch JR. 2011. Genomic characterization of human rotavirus G10 strains from the African Rotavirus Network: Relationship to animal rotaviruses. Infect Genet Evol, 11: 237–241.
Esona MD, Mijatovic-Rustempasic S, Conrardy C, Tong S, Kuzmin IV, Agwanda B, Breiman RF, Manyai K, Niezgoda M, Rupprecht E, Gentsch JR, Bowen MD. 2010. Reassortant group a rotavirus from straw-colored fruit bat (Eidolon helvum). Emerg Infect Dis, 16: 1844–1852.
Gatherer D. 2014. The 2014 Ebola virus disease outbreak in West Africa. J Gen Virol, 95: 1619–1624.
Ge X, Li J, Peng C, Wu L, Yang X, Wu Y, Zhang Y, Shi Z. 2011. Genetic diversity of novel circular ssDNA viruses in bats in China. J Gen Virol, 92: 2646–2653.
Ghosh S, Gatheru Z, Nyangao J, Adachi N, Urushibara N, Kobayashi N. 2011. Full genomic analysis of a simian SA11-like G3P. Infect Genet Evol, 11: 57–63.
He B, Yang F, Yang W, Zhang Y, Feng Y, Zhou J, Xie J, Feng Y, Bao X, Guo H, Li Y, Kia L, Li N, Matthijnssens J, Zhang H, Tu C. 2013. Characterization of a Novel G3P. J Virol, 87: 12357–12366.
Hu B, Chmura AA, Li J, Zhu G, Desmond JS, Zhang Y, Zhang W, Epstein JH, Daszak P, Shi Z. 2014. Detection of diverse novel astroviruses from small mammals in China. J Gen Virol, 95: 2442–2449.
Hu B, Ge X, Wang LF, Shi Z. 2015. Bat origin of human coronaviruses. Virol J, 12: 221.
Johnson E, Johnson B, Silverstein D, Tukei P, Geisbert T, Sanchez A, Jahrling P. 1996. Characterization of a new Marburg virus isolated from a 1987 fatal case in Kenya. Arch Virol, 11: 101–114.
Jones KE, Patel NG, Levy M. A, Storeygard A, Balk D, Gittleman JL, Daszak P 2008. Global trends in emerging infectious diseases. Nature, 451: 990–993.
Kading RC, Gilbert AT, Mossel EC, Crabtree MB, Kuzmin IV, Niezgoda M, Agwanda B, Markotter W, Weil MR, Montgomery JM, Rupprecht CE, Miller BR. 2013. Isolation and molecular characterization of Fikirini rhabdovirus, a novel virus from a Kenyan bat. J Gen Virol, 94: 2393–2398.
Kemenesi G, Dallos B, Gorfol T, Boldogh S, Estok P, Kurucz K, Kutas A, Foldes F, Oldal M, Nemeth V, Martella V, Banyai K, Jakab F. 2014. Molecular survey of RNA viruses in Hungarian bats: discovering novel astroviruses, coronaviruses, and caliciviruses. Vector-Borne Zoonot Dis, 14: 846–855.
Kim HK, Yoon SW, Kim DJ, Koo BS, Noh JY, Kim JH, Choi YG, Na W, Chang KT, S Ong D, Jeong DG. 2016. Detection of Severe Acute Respi ratory Syndrome-Like, Middle East Respiratory Syndrome-Like bat coronaviruses and group H Rotavirus in faeces of Korean bats. Transbound Emerg Dis, 63: 365–372.
Kuzmin IV, Mayer AE, Niezgoda M, Markotter W, Agwanda B, Breiman RF, Rupprecht CE. 2010a. Shimoni bat virus, a new representative of the Lyssavirus genus. Virus Res, 149:197–210.
Kuzmin IV, Niezgoda M, Franka R, Agwanda B, Markotter W, Breiman RF, Shieh W-J, Zaki SR, Rupprecht CE. 2010b. Marburg virus in fruit bat, Kenya. Emerg Infect Dis, 16: 352–354.
Leroy EM, Kumulungui B, Pourrut X, Rouquet P, Hassanin A, Yaba P, Delicat A, Paweska JT, Gonzalez JP, Swanepoel R. 2005. Fruit bats as reservoirs of Ebola virus. Nature, 438: 575–576.
Liu WB, Li ZX, Du Y, Wen G. 2015. Ebola virus disease: from epidemiology to prophylaxis. Mil Med Res, 2: 7.
Matthijnssens J, Ciarlet M, Heiman E, Arijs I, Delbeke T, McDonald SM, Palombo EA, Iturriza-Gomara M, Maes P, Patton JT, Rahman M, Van Ranst M. 2008a. Full genome-based classification of rotaviruses reveals a common origin between human Wa-Like and porcine rotavirus strains and human DS-1-like and bovine rotavirus strains. J Virol, 82: 3204–3219.
Matthijnssens J, Ciarlet M, Rahman M, Attoui H, Bányai K, Estes MK, Gentsch JR, Itturiza-Gomara M, Kirkwood C, Martella V, Merens PPC, Nakagomi O, Patton JT, Ruggeri FM, Saif LJ, Santos N, Steyer A, Taniguchi K, Desselberger U, Van Ranst M. 2008b. Recommendations for the classification of group A rotaviruses using all 11 genomic RNA segments. Arch Virol, 153: 1621–1629.
Moratelli R, Calisher CH. 2015. Bats and zoonotic viruses: Can we confidently link bats with emerging deadly viruses? Mem. I. Oswaldo-Cruz, 110: 1–22.
O’Shea TJ, Cryan PM, Cunningham AA, Fooks AR, Hayman DTS, Luis AD, Peel AJ, Plowright RK, Wood JLN. 2014. Bat flight and zoonotic viruses. Emerg Infect Dis, 20: 741–745.
Olival, KJ. 2016. To cull, or not to cull, bat is the question. Eco-Health, 13: 6–8.
Patterson BD, Webala PW. 2012. Keys to the bats (Mammalia: Chiroptera) of East Africa. Fieldiana Life and Earth Sciences, 12: 6.
Plowright RK, Eby P, Hudson PJ, Smith IL, Westcott D, Bryden WL, Martin G, Tabor GM, Skerratt LF, Anderson DL, Crameri G, Quammen D, Jordan D, Freeman P, Wand LF, Epstein JH, Marsh GA, Kung NY, McCallum H. 2015. Ecological dynamics of emerging bat virus spillover. Proc Biol Sci, 282: 2014–2124.
Shi Z. 2013. Emerging infectious diseases associated with bat viruses. Sci China Ser C, 56: 678–682.
Smith DH, Isaacson M, Johnson KM, Bagshawe A, Johnson BK, Swanapoel R, Johnson KM, Killey M, Bagshawe A, Siongok T, Keruga WK. 1982. Marburg-Virus Disease in Kenya. Lancet, 319: 816–820.
Sutherland LJ, Cash AA, Huang YJS, Sang RC, Malhotra I, Moormann AM, King CL, Weaver SC, King CH, La Beaud AD. 2011. Serologic evidence of arboviral infections among humans in Kenya. Am J Trop Med Hyg, 85: 158–161.
Tamura K, Peterson D, Peterson N, Stecher G, Nei M & Kumar S. 2011. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol, 28: 2731–2739.
Tao Y, Shi M, Conrardy C, Kuzmin IV, Recuenco S, Agwanda B, Alvares DA, Ellison JA, Gilbert AT, Moran D, Niezgoda M, Lindblade KA, Holmes EC, Brieman RF, Rupprecht CE, Tong S. 2013. Discovery of diverse polyomaviruses in bats and the evolutionary history of the Polyomaviridae. J Gen Virol, 94: 738–748.
Teeling EC, Springer MS, Madsen O, Bates P, O’Brien SJ, Murphy WJ. 2005. A molecular phylogeny for bats illuminates biogeography and the fossil record. Science, 307: 580–584.
Tong S, Conrardy C, Ruone S, Kuzmin IV, Guo X, Tao Y, Niezgoda M, Haynes L, Agwanda B, Breiman RF, Anderson LJ, Rupprecht CE. 2009. Detection of novel SARS-like and other coronaviruses in bats from Kenya. Emerg Infect Dis, 15: 482–485.
Tse H, Chan WM, Li KSM, Lau SKP, Woo PCY & Yuen KY. 2012. Discovery and genomic characterization of a novel bat sapovirus with unusual genomic features and phylogenetic position. PLoS ONE, 7: e34987.
Van Thiel PPAM, De Bie RMA, Eftimov F, Tepaske R, Zaaijer HL, Van Doornum GJJ, Schutten M, Osterhaus ADM, Majoie CBL, Aronica E, Fehlner-Gardiner C, Wandeler AI, Kager PA. 2009. Fatal human rabies due to duvenhage virus from a bat in Kenya: Failure of treatment with coma-induction, ketamine, and antiviral drugs. PLoS Neglect Trop. D, 3: 1–8.
Wang M, Yan M, Xu H, Liang W, Kan B, Zheng B, Chen H, Zheng H, Xu Y, Zhang E, Wang H, Ye J, Li G, Li M, Cui Z, Liu YF, Guo RT, Liu XN, Zhan LH, Zhou DH, Zhao A, Hai R, Yu D, Guan Y, Xu J. 2005. SARS-CoV infection in a restaurant from palm civet. Emerg Infect Dis, 11: 1860–1865.
Woolhouse MEJ. 2002. Population biology of emerging and reemerging pathogens. Trends Microbiol, 10: 3–7.
Xia L, Fan Q, He B, Xu L, Zhang F, Hu T, Wang Y, Li N, Qiu W, Zheng Y, Matthijnssens J, Tu C. 2014. The complete genome sequence of a G3P[10] Chinese bat rotavirus suggests multiple bat rotavirus inter-host species transmission events. Infect Genet Evol, 28: 1–4.
Yinda CK, Zeller M, Maes P, Deboutte W, Beller L, Heylen E, Ghogomu SM, Ranst MV, Matthijnssens J. 2016. Evidence for reassortment of highly divergent novel rotaviruses from bats in Cameroon, without evidence for human interspecies transmissions. Sci Rep, 6: 34209.
Zaki AM, Van Boheemen S, Bestebroer TM, Osterhaus AD, Fouchier RA. 2012. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. New Eng J Med, 367: 1814–1820.
Zhong NS, Zheng BJ, Li YM, Poon LM, Xie ZH, Chan KH, Li PH, Tan SY, Chang Q, Xie JP, Liu XQ, XU J, Li DX, Yuen KY, Peiris JSM, Guan Y. 2003. Epidemiology and cause of severe acute respiratory syndrome (SARS) in Guangdong, People’s Republic of China, in February, 2003. Lancet, 362: 1353–1358.
Acknowledgments
We are grateful for the Government of Kenya who permitted this study, specifically the Directors of Kenya Wildlife Service and National Museums of Kenya. We also appreciate the field and laboratory teams in Kenya and China who assisted in many different ways towards the success of this work. This work was funded by Sino-Africa Joint Research Center (SAJC201313 and SAJC 201605).
Author information
Authors and Affiliations
Corresponding authors
Additional information
ORCID: 0000-0002-0780-0399
ORCID: 0000-0001-8089-163X
This article is published with open access at Springerlink.com
Electronic supplementary material
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Waruhiu, C., Ommeh, S., Obanda, V. et al. Molecular detection of viruses in Kenyan bats and discovery of novel astroviruses, caliciviruses and rotaviruses. Virol. Sin. 32, 101–114 (2017). https://doi.org/10.1007/s12250-016-3930-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12250-016-3930-2