Abstract
Glial cells in the central nervous system (CNS) are composed of oligodendrocytes, astrocytes and microglia. They contribute more than half of the total cells of the CNS, and are essential for neural development and functioning. Studies on the fate specification, differentiation, and functional diversification of glial cells mainly rely on the proper use of cell- or stage-specific molecular markers. However, as cellular markers often exhibit different specificity and sensitivity, careful consideration must be given prior to their application to avoid possible confusion. Here, we provide an updated overview of a list of well-established immunological markers for the labeling of central glia, and discuss the cell-type specificity and stage dependency of their expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glial cells, or glia, were first described by neuroscientists including Rudolf Virchow, Santiago Ramón y Cajal, and Pío del Río-Hortega over a century ago [1]. With time, it was demonstrated that glial cells are diverse in type and function. It is now clear that these cells constitute more than half of the total number in the mammalian central nervous system (CNS) [2, 3]. Central glia can be classified as macroglia, which refers to both astrocytes and oligodendrocytes, and microglia. They have distinct embryonic origins, developmental trajectories, and functions. During early CNS development, neural progenitor cells or radial glial cells (RGCs) first give rise to neurons, followed by glial progenitor cells. Glial progenitors then produce either oligodendrocyte progenitor cells (OPCs) or astrocyte precursor cells (APCs), which subsequently undergo a series of morphological and molecular changes to become functionally mature oligodendrocytes or astrocytes [4]. In contrast, microglia are CNS-resident macrophages which originate from the yolk sac and fetal liver during early embryogenesis [5, 6].
In the vertebrate CNS, oligodendrocytes form myelin sheathes around axons to facilitate the rapid propagation of action potentials and provide trophic support for the myelinated axons [7]. Deficits of myelin are found in many neurological diseases, such as multiple sclerosis and the leukodystrophies [8]. The development of oligodendrocytes is coordinated by a large cohort of intracellular factors and extracellular signals [9, 10]. In mouse spinal cord, OPCs are initially generated from the OLIG2+ ventral neuroepithelial cells of the motor neuron progenitor (pMN) domain around embryonic day 12.5 (E12.5), and later from the ASCL1+ dp3-dp5 neuroepithelial domains in the dorsal spinal cord at ~ E14.5 [11, 12]. In the embryonic telencephalon, the first group of OPCs originate from the NKX2.1+ medial ganglionic eminence (MGE) and anterior entopeduncular area (AEP) in the ventral forebrain at E12.5–E14.5. The second wave of OPCs are generated from the lateral (LGE) and/or caudal ganglionic eminences at E14.5–E16.5. Starting at ~ E17.5, neural progenitors in the dorsal cortical region give rise to the third wave of OPCs [13]. The generation of OPCs follows a chronological order from ventral to dorsal along the entire anterior-posterior axis. Once generated, OPCs proliferate rapidly and migrate into the surrounding regions. OPCs start to differentiate into newly-formed oligodendrocytes (NFOs) at E18 in the spinal cord and at ~ P2–P3 in the forebrain. NFOs begin to contact and wrap around neuronal axons, express myelin proteins; and further differentiate into mature oligodendrocytes elaborating compact myelin sheaths. Notably, OPCs display significant functional redundancy throughout the CNS both during development and in adulthood, as the OPCs eliminated in one region are quickly replenished by OPCs from surrounding areas [13,14,15]. Studies over the past several decades have successfully identified a large number of stage-specific oligodendrocyte markers, critical tools for investigating the molecular and genetic control of oligodendrocyte development and axonal myelination in health and disease [16].
Astrocytes are the most functionally diverse glial cells and are involved in nearly all aspects of CNS physiology and functioning. The functional roles of astrocytes include but are not limited to maintaining the blood-brain barrier (BBB), providing physical and trophic support to neurons, axon guidance, synapse formation and remodeling, modulating synaptic transmission, regulating osmotic pressure, and ion homeostasis [17]. As noted earlier, APCs also originate from late RGCs in the ventricular zone of the CNS after neurogenesis. At the early stage of neural development, APCs undergo rapid local proliferation to expand their population. However, there are several unique features in the development of astrocytes as compared to that of oligodendrocytes. First, astrocytes arise from nearly all of the ventricular regions in the CNS and migrate to their final destinations in the direction of their radial glial processes [17]. Second, there is no functional redundancy among astrocytes in different regions [18]. Astrocytes are allocated to spatial regions in accordance with their sites of embryonic origin in the ventricular zone (VZ), and do not migrate tangentially to adjacent areas throughout life, even after acute CNS injury [19]. Third, RGCs, APCs, and astrocytes share many of the same molecular markers, such as GFAP and ALDH1L1 [20, 21]. Therefore, unlike the development of oligodendrocytes, there are few definite stage-specific molecular markers for the astrocyte lineage, and transcription factors that control the differentiation and maturation of astrocytes have yet to be found. Based on these and other findings, we propose that astrocytes are dormant neural progenitor cells that are influenced by local environments and are functionally adapted to support the local neuronal populations [22].
Unlike oligodendrocytes and astrocytes that are derived from neural progenitor cells, microglia are CNS-resident macrophages of blood origin. The CNS macrophages comprise microglia and border-associated macrophages (BAMs, also termed CNS-associated macrophages) in the meninges, choroid plexus, and perivascular spaces. Although microglia have been studied for decades, their developmental origin has been under debate for quite a long time. Now it is known that the bulk of microglia originate from hematopoietic progenitors in the yolk sac and enter the CNS tissue during early embryonic development [23, 24]. With the closure of BBB, renewal of the microglial population in the CNS mainly depends on their own proliferation [25]. With the aid of a number of cell-specific markers, microglia have been found to play critical roles in the development and functional maintenance of the CNS, including synaptic elimination, neural circuit wiring, axon tract fasciculation, vasculature tip cell fusion, synapse pruning, and inflammation [26].
In the past decade, a significant progress has been made in the studies of glial development, homeostasis, reactivation, regeneration, and other functions in the CNS. Clearly, these studies are highly dependent on the proper use of specific molecular markers for particular developmental stages or functional states. Theoretically, the most important features for a “perfect” marker gene are its high specificity and affinity. However, marker genes often display varying degrees of specificity and efficiency, and they even vary at different developmental stages or between species. Therefore, the proper use of cellular markers is critical for drawing correct conclusions.
Here, we provide a comprehensive and updated review of a list of well-characterized glial markers and discuss the pros and cons of these markers in their applications (Table 1).
Markers for Cells of Oligodendrocyte Lineage
SOX10
The SRY-box transcription factor SOX10 is one of the earliest identified pan-oligodendroglial markers and is exclusively expressed in cells of the oligodendrocyte lineage in the CNS [27]. Genetic studies revealed that SOX10 plays a central role in controlling oligodendrocyte development, as conventional deletion of the Sox10 gene leads to a dramatic delay of OPC differentiation with little impact on OPC generation and distribution in spinal tissue [28]. However, Sox10-KO enhances the phenotype of Sox9 mutants in OPC generation [29, 30], while the double knock-out of Sox8 and Sox10 genes in differentiated oligodendrocytes results in demyelination in the mouse CNS [31, 32]. These findings suggest that SOX10 regulates nearly all stages of oligodendrocyte development, including their initial generation from neural progenitors, terminal differentiation, and myelin maintenance. As a specific marker for the oligodendroglial lineage, SOX10 is activated in OPCs immediately after their fate specification in the ventricular regions (Fig. 1A, B). SOX10 expression is maintained in OPCs and markedly upregulated at the onset of OPC differentiation. It is worth highlighting that Sox10 is considered to be the most reliable pan-oligodendrocyte marker in both brain and spinal cord, due to its specificity and high efficiency in labeling all cells of oligodendrocyte lineage throughout the life span.
Molecular markers for cells of oligodendrocyte lineage. A Diagram of oligodendrocyte development and stage-specific markers. B OLIG2+ cells include SOX10+ OPCs (white arrowheads), SOX10- astrocytes (arrows), and MIPCs in the SVZ of P0 mouse cortex (blue arrow heads). C Nearly all ALDH1L1+ astrocytes express OLIG2 in P7 mouse cortex (arrows). D, E PDGFRA and NG2 label OPCs in P0 cortex (arrowheads). Note that NG2 is also expressed in vascular pericytes (arrows in E), while PDGFRA is not. F–J CC1 but not MYRF is weakly expressed in astrocytes in P7 cortex (arrows). At P15, all of the CC1+ cells are MYRF+ differentiated oligodendrocytes (arrowheads). K, L Expression patterns of ASPA and CNPase in mouse cortex. M Brain myelin structures visualized by gold-based staining. NPCs, neural progenitor cells; OPCs, oligodendrocyte progenitor cells; Pre-OLs, pre-oligodendrocytes; NFOs, newly-formed oligodendrocytes; OLs, oligodendrocytes; VZ/SVZ, (sub)ventricular zone; IZ, intermediate zone; CP, cortical plate; CC, corpus callosum. Scale bars, 50 μm.
OLIG2
This gene is probably the most frequently-used marker for cells of oligodendrocyte lineage, partly due to its commercial availability. However, OLIG2 is not always a reliable marker for labeling oligodendroglia. In the spinal cord, OLIG2 expression is initially activated in neural progenitors of the pMN domain at ~ E8.5, but is rapidly lost from their motor neuron progeny during neurogenesis [33, 34]. As development proceeds, OLIG2 expression is retained in OPCs but is slightly downregulated in differentiated oligodendrocytes [35, 36]. In the brain, OLIG2 is broadly expressed in the VZ of the ventral telencephalon, including the LGE, MGE, and AEP regions at embryonic stages [37,38,39]. In the dorsal cortical region, the earliest OLIG2-positive cells appear at ~ E17.5, coincident with the onset of local gliogenesis [40, 41]. These OLIG2+ pluripotent precursor cells, termed multipotent intermediate progenitor cells (MIPCs) (also named pri-OPCs or pre-OPCs in other studies [42, 43]), are the common precursors of cortex-derived astrocytes, oligodendrocytes, and olfactory bulb interneurons (OBiNs) [40, 44] (OLIG2+ cells in the VZ and subventricular zone (SVZ), Fig. 1B). Despite its rapid downregulation in OBiNs, OLIG2 expression is sustained in glial cell progeny, including newborn astrocytes (APCs) and OPCs [40, 44, 45] (Fig. 1B, C). In the cortical region of neonatal mice, nearly all astrocytes are immunoreactive to OLIG2 until P7 (Fig. 1C). Within the next week or so, the expression level of OLIG2 is rapidly downregulated as they progress further along the astrocyte lineage. By the age of animal weaning, ~ 99% of the OLIG2+ cells in cortical tissue co-express SOX10, and vice versa, indicating that OLIG2 predominantly labels oligodendrocyte cells at this stage [46] (Fig. 1B). In summary, OLIG2 is an excellent pan-oligodendroglial marker in the spinal cord, since only a small fraction of pMN-derived astrocytes expresses this gene during embryonic stages [47, 48]. However, OLIG2 cannot be used as a specific oligodendrocyte marker in embryonic or early postnatal brain tissue. Consistent with the regional difference in OLIG2 expression, deletion of the Olig2 gene affects the development of cortical astrocytes [45, 49], but not that of spinal astrocytes [50, 51].
PDGFRA and NG2
Platelet-derived growth factor α receptor (PDGFRA) and NG2 proteoglycan (also known as chondroitin sulfate proteoglycan 4) are commonly-used markers for OPCs in the CNS. In rodents, PDGFRA is exclusively expressed in OPCs from their origin from neural progenitor cells in both the embryonic and adult CNS (Fig. 1D) [52, 53]. The function of the PDGFA/PDGFRA signaling pathway is to stimulate OPC proliferation, but meanwhile to inhibit their differentiation. Ablation of either Pdgfa or Pdgfra at embryonic stages causes premature differentiation of OPCs, resulting in severe hypomyelination due to the reduced proliferation and faster depletion of the progenitor pool [54, 55].
NG2 is another widely-used OPC marker, and NG2+ glial cells produce myelinating oligodendrocytes in adult CNS tissue [46, 56,57,58]. However, unlike PDGFRA, NG2 is also expressed in vascular pericytes, especially in the embryonic and newborn brain (Fig. 1E) [59, 60]. In general, PDGFRA is a more reliable OPC marker than NG2 for both in situ hybridization and immunostaining, and the pericytes must be culled when NG2 is used in labeling OPCs.
NKX2.2
Homeodomain transcription factor NKX2.2 is selectively upregulated in differentiating OPCs or pre-oligodendrocytes, but is rapidly down-regulated after oligodendrocyte differentiation [61]. Genetic studies have demonstrated that NKX2.2 controls the timing of OL differentiation, as conditional deletion of Nkx2.2 causes a developmental delay of OL differentiation [62], and overexpression of NKX2.2 in OPCs results in precocious OL differentiation [55]. NKX2.2 promotes oligodendrocyte differentiation by directly inhibiting Pdgfra expression. However, NKX2.2 is also expressed in a subset of neurons in the ventral spinal cord and ventral thalamus [63,64,65]. Thus, identification of pre-myelinating oligodendrocytes by NKX2.2 expression in the gray matter must be combined with other lineage-specific transcription factors such as OLIG2 or SOX10.
CC1
The monoclonal antibody anti-adenomatous polyposis coli (APC) clone CC1 is widely used to mark differentiated/mature oligodendrocytes without labeling myelin. However, previous studies have shown that the CC1 antibody does not bind APC [66]. In fact, the expression of APC is completely different from that of CC1 [66,67,68,69]. It is now clear that the CC1 antibody recognizes a specific isoform of the RNA-binding protein Quaking (QKI, encoded by the Qk gene), namely QKI7 [69]. During early development, Qk is broadly expressed in neural progenitor cells and later in their glial progeny, including OPCs and newborn APCs (Fig. 1F) [70]. With time, Qk expression is gradually reduced in astrocyte lineages, but is strongly upregulated in differentiated oligodendrocytes (Fig. 1G, J) [71]. As an antibody recognizing the cytoplasmic isoform QKI7 of QK proteins, CC1 is a useful and reliable marker for counting the number of mature oligodendrocytes in postnatal animals (older than P15 in the brain and P7 in the spinal cord). When used in the CNS of neonatal mice, astrocytes and OPCs with weak CC1 expression should be excluded from the mature oligodendrocyte population based on their morphology, gene expression level, and tissue distribution.
Myelin Proteins
Myelin is made of lipids (>70% of its dry weight) and proteins. Myelin proteins play important roles in cell adhesion, axon-myelin interactions, and the integrity of compact myelin structure. The most abundant proteins in CNS myelin sheaths include PLP (proteolipid protein 1), MBP (myelin basic protein), CNP (2′,3′-cyclic nucleotide 3′-phosphodiesterase, CNPase), MAG (myelin-associated glycoprotein), and MOG (myelin oligodendrocyte glycoprotein) [72]. These proteins constitute >30% of the total myelin-associated proteins [8]. Detection of their expression at the RNA or protein level directly monitors the differentiation state of oligodendrocytes both in vitro and in vivo. These myelin genes are activated when OPCs start to differentiate, and are sustained in myelinating and myelinated cells. The expression of myelin genes commences at ~E18 in mouse spinal cord or P2–P3 in the cortex by RNA in situ hybridization, but their proteins can only be detected ~ 2 days later. Generally speaking, the chronological order of expression of these markers in vivo is CNP/MBP, PLP/MAG, MOG/CC1, although the temporal difference is quite small. It should be noted that myelin proteins are predominantly localized in the myelin processes of oligodendrocytes. Thus, immunostaining against these proteins in CNS tissue usually reflects the biomass of myelin sheaths, and is not suitable for cell counting (Fig. 1L). Should the quantitative analysis of cell numbers be required, RNA in situ hybridization of these myelin genes (except for Mbp mRNA, which is distributed in cellular processes as well) would be a better choice.
MYRF
The myelin regulatory factor gene (MYRF) is a novel type of membrane-bound transcriptional factor that is evolutionarily conserved from invertebrates to vertebrates. It is initially synthesized as a type-II membrane protein, which subsequently undergoes homo-trimerization and self-cleavage on the endoplasmic reticulum (ER) membrane. Once cleaved on the ER membrane, the N-terminal trimers of MYRF are released and translocate into the nucleus to function as a transcriptional activator [73, 74]. As the downstream target of SOX10, MYRF is a key regulator of myelin gene expression, which is essential for myelin formation and maintenance [75]. In the CNS, MYRF expression is strictly restricted to oligodendrocytes. Its expression is selectively upregulated in oligodendrocytes at the beginning of cell differentiation, and sustained in mature oligodendrocytes, in a pattern similar to that of myelin genes such as Plp and Mbp (Fig. 1H–J) [75, 76]. Conditional ablation of Myrf in OPCs blocks myelinogenesis during development or myelin maintenance/repair in adulthood [76]. Given its higher specificity and nuclear localization, MYRF is considered to be a better marker than CC1 for differentiated oligodendrocytes and cell counting.
Other Oligodendroglial Markers
In addition to the markers described above, a number of others are also frequently used to identify cells of oligodendrocyte lineage in a stage-specific manner. For instance, the monoclonal antibody O4, like NKX2.2, preferentially labels immature differentiating oligodendrocytes (or pre-oligodendrocytes) before the expression of MBP and PLP [64, 77, 78]. This mouse IgM antibody works well in both culture and tissue sections. Besides, Enpp6 and Bmp4 have recently been shown to selectively label newly-formed oligodendrocytes, but their expression is downregulated in more mature myelinating oligodendrocytes [79, 80]. Unfortunately, working antibodies have not been developed for these proteins. As oligodendrocytes undergo terminal maturation, they start to express Opalin (also known as TMEM10) and ASPA (aspartoacylase) [81,82,83,84]. The expression of these two new markers in mature oligodendrocytes occurs several days later than that of MBP and PLP in vivo, i.e. at ~ P5–P7 in spinal tissue or P10–P14 in the corpus callosum. For identification of more mature oligodendrocytes, ASPA seems to be the best choice due to its preponderant localization in the cell body (Fig. 1K). Another marker for mature oligodendrocytes is CAII (carbonic anhydrase 2), and this gene is a specific marker for type I/II oligodendrocytes which are predominantly myelinate small-diameter axons [85, 86]. In addition, a simple myelin staining method, Black-Gold, was developed some years ago, based on the specific affinity of gold phosphate complex with lipidic myelin structures [87]. This chemical staining method is relatively fast and simple compared to immunostaining, and produces much higher resolution than the traditional Luxol Fast Blue staining (Fig. 1M).
Markers for Cells of Astrocyte Lineage
GFAP
Glial fibrillary acidic protein (GFAP) was one of the first identified astrocyte markers [88, 89]. For decades, GFAP has been widely used as the standard astrocyte marker in numerous studies because of its robust staining and the lack of better markers. GFAP labels cultured astrocytes in vitro and reactive astrocytes in injured or pathological CNS tissues. In normal CNS tissue, GFAP is predominantly expressed by astrocytes in the white matter (fibrous astrocytes) and the parenchyma region near the leptomeninges, but very weakly in gray matter protoplasmic astrocytes. However, GFAP also labels the late radial glial cells in embryonic cortical tissue [40]. It is worth mentioning that GFAP has a high degree of homology to other types of neurofilament proteins (e.g. Nestin and Vimentin). Therefore, the specificity of GFAP antibodies must be carefully characterized, due to the potential cross-reactivity issue.
S100B
Besides GFAP, S100B (S100 protein, beta polypeptide, neural; S100β) has also been widely used for decades as an astrocyte biomarker in both the developing and adult CNS [90, 91]. S100B is a Ca2+-binding protein, and has been implicated in several neurological diseases including Alzheimer’s disease, Parkinson’s disease, and neuropathic pain [92]. However, emerging evidence suggests that S100B is also expressed in cells of oligodendrocyte lineage [93, 94]. Our recent study demonstrated that the earliest S100B+ cells are differentiating oligodendrocytes before P4 in the forebrain, rather than astrocytes as previously thought. Its expression in cortical gray matter astrocytes occurs only after P4, a time-point when astrocytes have already migrated from the germline region [95]. Unexpectedly, we found that almost all of the S100B+ cells in the white matter are SOX10+/MYRF+ oligodendrocytes from the neonatal stage to adulthood. In the postnatal CNS, S100B marks protoplasmic astrocytes in the gray matter and maturing oligodendrocytes in both the gray and white matter (Fig. 2B). Nevertheless, S100B still holds advantages as a marker for astrocytes. First, nearly all protoplasmic astrocytes in the gray matter express S100B in post-weaning animals. Second, S100B protein is predominantly localized in the cell body and thus highly suitable for cell counting. Third, unlike other astrocyte markers, S100B is not expressed by radial glial progenitor cells and newborn astrocyte progenitors (Fig. 2A) [95]. Thus, S100B remains an excellent choice for gray matter astrocytes when combined with SOX10 to exclude its expression in oligodendrocytes.
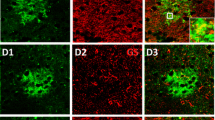
Cellular markers for cells of astrocyte lineage. A Diagram of astrocyte development and the expression of specific markers. B Besides gray matter astrocytes (arrows), many S100B+ cells are co-labeled with the pan-oligodendrocyte marker SOX10 (arrowheads). C ALDH1L1 expression is only detected in astrocytes, but not in SOX10+ oligodendrocytes. D FABP7 is an operational astrocyte marker. E, F Nearly all of SOX10+ OPCs co-express SOX9 in P0 forebrain (arrowheads), and SOX9 primarily marks astrocytes in the adult brain. G All ALDH1L1+ astrocytes co-express SOX9 in the brain. H AQP4 is localized in astrocyte process, and APQ4 immunolabeling reveals the entire network of vessels covered by astrocytic end-feet. RGCs, radial glial cells; APCs, astrocyte progenitor cells; VZ/SVZ, (sub)ventricular zone; IZ, intermediate zone; CP, cortical plate; CC, corpus callosum. Scale bars, 50 μm.
ALDH1L1 and ACSBG1
Recent studies on astroglial gene expression profiling have identified two new astroglial marker genes, Aldh1l1 and Acsbg1. Aldh1l1 (aldehyde dehydrogenase 1 family, member L1) encodes a folate enzyme of tetrahydrofolate synthesis. Several studies have demonstrated the specific expression of ALDH1L1 in astrocytes [96, 97]. We have also confirmed that ALDH1L1+ cells out of the germline region never co-express the pan-oligodendroglial marker SOX10 in both the embryonic and adult brain, indicating their astrocyte identity (Fig. 2C) [95]. ACSBG1, a protein with very long-chain acyl-CoA synthetase activity, has been reported to display an expression pattern similar to that of ALDH1L1 [97]. However, detailed analysis of its spatiotemporal expression pattern is still required to validate its specificity in the astrocyte lineage. In addition, it has been noted that the expression of ALDH1L1 decreases in adulthood [96], calling for better astrocyte markers for the adult CNS.
SOX9 and NFIA
To date, the transcriptional regulation of astrocyte development is still largely unknown. The SRY-box transcriptional factor SOX9 was originally reported to determine glial fate choice in the developing spinal cord. SOX9 is initially expressed in neural progenitor cells, and later in glial cells in the marginal zone, including astrocytes and OPCs (Fig. 2E) [29]. Intriguingly, SOX9 expression is gradually downregulated in the oligodendrocyte lineage as development proceeds and becomes progressively restricted to astrocytes in the adult mouse CNS (Fig. 2F, G) [29, 98]. NFIA (nuclear factor I/A) is a target transcriptional factor of SOX9 and has been implicated in the regulation of astrogliogenesis [99, 100]. Although the expression pattern of NFIA in glial cells is quite similar to that of SOX9, it is also expressed by motor neurons in the spinal cord and deeper layer projection neurons in the cortex [101]. As NFIA down-regulation in oligodendrocytes occurs much slower than SOX9, a higher proportion of SOX10+ OPCs is still more immunoreactive to NFIA than to SOX9 in the adult mouse CNS. Therefore, SOX9 and NFIA could serve as pan-astrocyte markers due to their high immunoreactivity and nuclear localization, but only when OPCs are excluded.
Other Astrocyte Markers
A number of other astrocytic markers have also been described. GLAST and GLT-1, also known as EAAT1/SLC1A3 and EAAT2/SLC1A2, respectively, are the primary astrocytic glutamate transporters in the adult CNS, accounting for > 90% of synaptic glutamate clearance [102, 103]. While the GLAST expression pattern is consistent with a pan-astrocyte marker, GLT-1 is also expressed in neurons in both the brain and the spinal cord [104, 105]. As a transmembrane protein, GLAST immunostaining displays punctate/reticular-like structures and is not suitable for outlining the shape of astrocytes in vivo, especially in adult animals. Nonetheless, we found that GLAST specifically labels cultured astrocyte in vitro. Glutamine synthase (GS, Glul) is another commonly-used astrocyte marker [106]. However, it has recently been reported that GS is activated in mature oligodendrocytes in mouse brain and spinal cord [107, 108]. The onset of GS expression in mature oligodendrocytes is between P21 and P28 in mouse brain, which is later than the appearance of the mature oligodendrocyte marker ASPA. FABP7 (fatty acid binding protein 7, also known as BLBP) has occasionally been used to label astrocytes in some studies [109]. It has also been suggested that FABP7 is expressed in OPCs in both mouse and chicken CNS during development and in adulthood [110]. Nevertheless, FABP7 can still be used as an approximate astrocyte marker, since its expression in OPCs is very weak (Fig. 2D). Other astrocytic markers include Aldolase C (ALDOC), AQP4 (aquaporin 4) (Fig. 2H), and the newly-described transcriptional factor ZBTB20 [111,112,113]. However, more detailed analyses with molecular and genetic approaches are needed to determine the specificity and expression dynamics of these markers.
Markers for Microglia
CNS macrophages consist of two distinct types of cell, microglia in the parenchyma and border-associated macrophages (BAMs) in the meninges, choroid plexus, and perivascular spaces. Both cell types are derived from erythromyeloid progenitors during early embryonic development (Fig. 3A) [23, 114]. Therefore, it is not surprising that microglia share many of the same molecular markers with BAMs. It is well documented that two functional states are associated with microglia: resting microglia with ramified morphology, and reactive microglia with an amoeboid shape in response to a pathological environment. In the previous studies, several molecular markers have been identified that can discern microglia from BAMs, or resting from reactive microglia.
Molecular markers for microglia. A Diagram of microglial development and lineage-specific markers. B CD206 marks BAMs (arrows) but not microglia. C IBA1 expression in microglia in P0 and P15 mouse brain. D P2RY12 expression in microglia of P0 cortex. E CD68 labels reactive microglia after CNS injury. F Expression of Hexb mRNA in microglia revealed by in situ hybridization. BAMs, border-associated macrophages; VZ/SVZ, (sub)ventricular zone; IZ, intermediate zone; CP, cortical plate. Scale bars, 50 μm.
IBA1 and CX3CR1
To date, the most reliable and effective method for detection of microglia relies on the discovery of IBA1 protein by a Japanese group in 1996 and the demonstration of its microglial specificity [115, 116]. IBA1 protein, encoded by the Aif1 gene (allograft inflammatory factor 1), was initially reported to be involved in the membrane ruffling and phagocytosis of macrophages/microglia [117]. Several high-affinity specific antibodies are commercially available for the IBA1 protein. IBA1 is highly specific and capable of labeling both resting and activated microglia [118]. Since IBA1 protein is distributed in the cell body and the tiny processes of microglia, it allows the observation of morphological changes and the calculation of cell numbers (Fig. 3C). One apparent disadvantage of the IBA1 marker is that it not only labels microglia, but also macrophages, including BAMs and blood-derived monocytes [115]. Another pan-microglia marker is CX3CR1, with the same expression pattern as IBA1 [119]. CX3CR1 is a G protein-coupled receptor involved in cell adhesion and migration. However, antibodies for this protein are scarce. Instead, the reporter mouse line Cx3cr1-EGFP has been developed and is widely used for microglia labeling [120].
P2RY12 and CD68
These markers are frequently used to distinguish between resting and reactivated microglia. P2RY12 is a Gi-coupled metabotropic purinergic receptor which mediates microglial responses to extracellular nucleotides [121]. Loss of P2ry12 in microglia switches their morphology from a highly ramified resting state to an amoeboid reactive state. Moreover, P2RY12 is strongly expressed in resting microglia, but drastically reduced after microglial activation (Fig. 3D) [121]. CD68 is a 110-kDa transmembrane glycoprotein present in monocytes and tissue macrophages [122]. It has been reported that CD68 participates in the pathological activation of macrophages by low density lipoproteins, and functions as an inhibitor of immune reactions [123]. The expression of CD68 is not conspicuous in resting microglia, but is strongly upregulated when they are activated (Fig. 3E). As a marker for reactive microglia, one disadvantage of CD68 protein is its “dotted” distribution in the membrane.
CD206 and Hexb
Thanks to the advent of single-cell RNA sequencing technology, scientists are now able to distinguish microglia from BAMs and monocytes at the molecular level. While CD206 detects BAMs and monocytes, Hexb specifically labels microglia. This is an important milestone for the diagnosis of BBB damage under pathological conditions. CD206, encoded by the Mrc1 gene, is a well-defined marker for BAMs and other macrophages without cross-labeling microglial cells (Fig. 3B) [124]. In contrast, the Hexb gene is a stable marker for microglia in both health and pathological conditions without discernable expression in BAMs (Fig. 3F) [125]. Thus, Hexb is an important marker for distinguishing microglia from surrounding macrophages. In addition, other molecular markers for CNS microglia and macrophages have also been suggested in some recent studies. For instance, TMEM119 and Siglec-H appear to be specific to microglia [126, 127], and LYVE1 preferentially labels BAMs and macrophages [128]. Unfortunately, there are no high-quality commercial antibodies for many of the aforementioned microglial markers.
Conclusions
In general, an ideal cellular marker should be specific enough to label only one type of cell or a particular stage of cell development with high affinity. For cells of oligodendrocyte lineage, abundant stage-specific markers have been identified and high-quality antibodies have been developed accordingly. However, for astrocytes, the situation is quite different. To date, there is a lack of definitive staging for astrocyte development, partly due to the lack of stage-specific markers. Many widely-used astrocyte markers are not specific or effective enough to distinguish among RGCs, APCs, and mature astrocytes. It is now becoming clear that astrocytes, radial glial cells, and even cerebellar Bergmann cells share a similar gene expression profile, including Glast, Fabp7, Aldh1l1, GFAP, Hopx, Tnc, Fgfr3, Acsbg1, Dbi, and Qk, as well as the transcriptional factors Sox2, Sox9, Nfia, Zeb1, and Zbtb20 [20, 40]. Although a few markers (e.g. S100B, GS, and NFIA) are capable of detecting most cells of astrocyte lineage, they also label other neuronal cell types such as oligodendrocytes or neurons. Therefore, a combinatory expression of multiple markers should be considered for accurate identification of astrocyte subpopulations or developmental stages.
Intriguingly, mature astrocytes share some of the same molecular marker genes (e.g. S100B and GS) with mature oligodendrocytes in the CNS [95, 107]. When these markers are chosen to label astrocytes, the oligodendrocyte markers SOX10, CC1, and MYRF can be used in combination for more definitive identification. This suggests that these two types of macroglia may share some common metabolic pathways. Thus, part of supporting functions of astrocytes could be taken over by myelin structures. Conceivably, myelinated axons, especially those in the white matter, are remote from their cell bodies and some metabolic support may be provided by myelin.
Another important point is that some of the markers (e.g. Sox2, Fabp7, Sox9, Nfia, Qk, Zeb1, and Zbtb20) expressed in RGCs and astrocytes are also maintained in OPCs, in keeping with the suggestion that RGCs are the common precursors of OPCs and astrocytes. As a matter of fact, several recent studies have identified a special class of intermediate progenitor cells (IPCs) in cortical regions that express EGFR, ASCL1, and OLIG2 during gliogenesis in both mice and primates [40, 44]. These EGFR+/ASCL1+/OLIG2+ multipotent progenitors are described as “pre-OPCs”, “pri-OPCs”, “mGPCs (multipotent glial progenitor cells)”, and “MIPCs (multipotent IPCs)” in different studies [42, 43, 129, 130]. Unexpectedly, these MIPCs are tripotent precursors as they produce not only OPCs, but also astrocytes and olfactory bulb interneurons [40]. Since the expression of OLIG2 does not restrict the fate of cells to OPCs, it seems to be more appropriate to name this type of cell “MPICs”. In light of this new finding, SOX10, rather than OLIG2, would be the best choice for labeling OPCs among glial populations in brain tissue.
Based on the similar gene expression profiling among RGCs, APCs, and astrocytes, we recently hypothesized that astrocytes are the resting neural precursor cells without undergoing terminal differentiation and cell-cycle exit, providing structural and metabolic support for local neurons [22]. Functional adaptions to the local neuronal milieu may explain the heterogeneity of astrocytes in the CNS. In support of this hypothesis, astrocytes retain some degree of stemness and reactive astrocytes are capable of differentiating into neurons or OPCs in vitro or in vivo under certain pathological conditions [131, 132]. Also, astrocytes in different regions retain a radial migration property and fail to migrate into adjacent regions after injury. Thus, astrocytes in different regions cannot be replaced by each other after lesions, contrary to the functional redundancy displayed by OPCs from different embryonic origins [19, 133].
Another issue to be aware of is the species differences in the expression of molecular markers. Although the fundamental principles are roughly the same, primates and rodents do exhibit some species-diversity in their cellular and molecular organization. For instance, cortical regions in human but not mouse develop an enlarged germinal zone called the outer subventricular zone, populated by multipotent outer radial glial cells [134, 135]. It has also been reported that GFAP in human brain and spinal cord is expressed by almost all of the astrocyte populations, while it only labels a subset of astrocytes in the murine CNS [40, 44]. Again, the classical OPC marker PDGFRA is weakly expressed in HOPX+/OLIG2+ MIPCs in the developing human cortex, but not mouse cortex before they differentiate into OPCs [44]. Therefore, it is not recommended to define a cell type with only one single marker, especially during early embryonic stages.
Methodologically, there are usually two major experimental procedures for in situ detection of a particular cell type in a tissue. One is through RNA in situ hybridization (ISH), which detects target mRNAs, and the other is by immunostaining based on specific antibody-antigen biochemical interactions. There are several advantages for ISH detection of molecular markers. First, the RNA probes for ISH are easier to prepare under laboratory conditions, especially when antibodies are not available. Second, mRNAs are usually localized to the cell body, so it is convenient for counting the number of cells of interest. Third, the difference in the nucleic acid sequence of homologous genes is greater than that of the amino-acid sequence, so the specificity of ISH is in general very high. However, its drawbacks are also apparent, as ISH is somewhat time-consuming and unsuitable for multiple-channel labeling and morphological observations. Immunostaining, including immunofluorescence, immunohistochemistry, and immunocytochemistry, is the most commonly used cell labeling technique. Immunostaining relies on specific antibodies. There are several means to validate the specificity of a particular antibody: (1) the expression pattern of a target protein detected by immunostaining should be consistent with that of its mRNA labeled by ISH; (2) knockout verification has now become a standard approach, especially when it is performed in tissue sections; (3) examination of cross-recognition between homologous proteins is recommended, considering that knockout verification of antibodies is mostly done in cell cultures, and homologous proteins that can be recognized non-specifically may not appear in vitro.
To date, the cellular markers for astrocytes and microglia are still relatively lacking, making it difficult to define their developmental stages, cell subtypes, or distinct functional states. At present, with the aid of single-cell RNA sequencing, it is more operable to identify novel cell type-specific markers. However, before being used as a marker, these candidate genes must be verified by both RNA in situ analysis and immunostaining.
References
Somjen GG. Nervenkitt: Notes on the history of the concept of neuroglia. Glia 1988, 1: 2–9.
Herculano-Houzel S. The glia/neuron ratio: How it varies uniformly across brain structures and species and what that means for brain physiology and evolution. Glia 2014, 62: 1377–1391.
von Bartheld CS, Bahney J, Herculano-Houzel S. The search for true numbers of neurons and glial cells in the human brain: A review of 150 years of cell counting. J Comp Neurol 2016, 524: 3865–3895.
Rowitch DH, Kriegstein AR. Developmental genetics of vertebrate glial–cell specification. Nature 2010, 468: 214–222.
Ginhoux F, Guilliams M. Tissue-resident macrophage ontogeny and homeostasis. Immunity 2016, 44: 439–449.
Thion MS, Ginhoux F, Garel S. Microglia and early brain development: An intimate journey. Science 2018, 362: 185–189.
Nave KA, Werner HB. Ensheathment and myelination of axons: Evolution of glial functions. Annu Rev Neurosci 2021, 44: 197–219.
Nave KA, Werner HB. Myelination of the nervous system: Mechanisms and functions. Annu Rev Cell Dev Biol 2014, 30: 503–533.
Huang H, Zhao XF, Zheng K, Qiu M. Regulation of the timing of oligodendrocyte differentiation: Mechanisms and perspectives. Neurosci Bull 2013, 29: 155–164.
He L, Lu QR. Coordinated control of oligodendrocyte development by extrinsic and intrinsic signaling cues. Neurosci Bull 2013, 29: 129–143.
Richardson WD, Kessaris N, Pringle N. Oligodendrocyte wars. Nat Rev Neurosci 2006, 7: 11–18.
Cai J, Qi Y, Hu X, Tan M, Liu Z, Zhang J. Generation of oligodendrocyte precursor cells from mouse dorsal spinal cord independent of Nkx6 regulation and Shh signaling. Neuron 2005, 45: 41–53.
Kessaris N, Fogarty M, Iannarelli P, Grist M, Wegner M, Richardson WD. Competing waves of oligodendrocytes in the forebrain and postnatal elimination of an embryonic lineage. Nat Neurosci 2006, 9: 173–179.
Zhu Q, Whittemore SR, Devries WH, Zhao X, Kuypers NJ, Qiu M. Dorsally-derived oligodendrocytes in the spinal cord contribute to axonal myelination during development and remyelination following focal demyelination. Glia 2011, 59: 1612–1621.
Fang M, Yu Q, Ou B, Huang H, Yi M, Xie B, et al. Genetic evidence that dorsal spinal oligodendrocyte progenitor cells are capable of myelinating ventral axons effectively in mice. Neurosci Bull 2020, 36: 1474–1483.
Elbaz B, Popko B. Molecular control of oligodendrocyte development. Trends Neurosci 2019, 42: 263–277.
Khakh BS, Deneen B. The emerging nature of astrocyte diversity. Annu Rev Neurosci 2019, 42: 187–207.
Bayraktar OA, Fuentealba LC, Alvarez-Buylla A, Rowitch DH. Astrocyte development and heterogeneity. Cold Spring Harb Perspect Biol 2014, 7: a020362.
Tsai HH, Li H, Fuentealba LC, Molofsky AV, Taveira-Marques R, Zhuang H, et al. Regional astrocyte allocation regulates CNS synaptogenesis and repair. Science 2012, 337: 358–362.
Heng X, Guo Q, Leung AW, Li JY. Analogous mechanism regulating formation of neocortical basal radial glia and cerebellar Bergmann glia. eLife 2017, 6: e23253.
Mori T, Buffo A, Götz M. The novel roles of glial cells revisited: The contribution of radial Glia and astrocytes to neurogenesis. Curr Top Dev Biol 2005, 69: 67–99.
Zheng K, Huang H, Yang J, Qiu M. Origin, molecular specification, and stemness of astrocytes. Dev Neurobiol 2022, 82: 149–159.
Ginhoux F, Greter M, Leboeuf M, Nandi S, See P, Gokhan S, et al. Fate mapping analysis reveals that adult microglia derive from primitive macrophages. Science 2010, 330: 841–845.
De S, Van Deren D, Peden E, Hockin M, Boulet A, Titen S, et al. Two distinct ontogenies confer heterogeneity to mouse brain microglia. Development 2018, 145: 152306.
Ajami B, Bennett JL, Krieger C, Tetzlaff W, Rossi FMV. Local self-renewal can sustain CNS microglia maintenance and function throughout adult life. Nat Neurosci 2007, 10: 1538–1543.
Li Q, Barres BA. Microglia and macrophages in brain homeostasis and disease. Nat Rev Immunol 2018, 18: 225–242.
Kuhlbrodt K, Herbarth B, Sock E, Hermans-Borgmeyer I, Wegner M. Sox10, a novel transcriptional modulator in glial cells. J Neurosci 1998, 18: 237–250.
Stolt CC, Rehberg S, Ader M, Lommes P, Riethmacher D, Schachner M, et al. Terminal differentiation of myelin-forming oligodendrocytes depends on the transcription factor Sox10. Genes Dev 2002, 16: 165–170.
Stolt CC, Lommes P, Sock E, Chaboissier MC, Schedl A, Wegner M. The Sox9 transcription factor determines glial fate choice in the developing spinal cord. Genes Dev 2003, 17: 1677–1689.
Finzsch M, Stolt CC, Lommes P, Wegner M. Sox9 and Sox10 influence survival and migration of oligodendrocyte precursors in the spinal cord by regulating PDGF receptor alpha expression. Development 2008, 135: 637–646.
Stolt CC, Lommes P, Friedrich RP, Wegner M. Transcription factors Sox8 and Sox10 perform non-equivalent roles during oligodendrocyte development despite functional redundancy. Development 2004, 131: 2349–2358.
Turnescu T, Arter J, Reiprich S, Tamm ER, Waisman A, Wegner M. Sox8 and Sox10 jointly maintain myelin gene expression in oligodendrocytes. Glia 2018, 66: 279–294.
Wu S, Wu Y, Capecchi MR. Motoneurons and oligodendrocytes are sequentially generated from neural stem cells but do not appear to share common lineage-restricted progenitors in vivo. Development 2006, 133: 581–590.
Richardson WD, Smith HK, Sun T, Pringle NP, Hall A, Woodruff R. Oligodendrocyte lineage and the motor neuron connection. Glia 2000, 29: 136–142.
Lu QR, Yuk DI, Alberta JA, Zhu Z, Pawlitzky I, Chan J, et al. Sonic hedgehog-regulated oligodendrocyte lineage genes encoding bHLH proteins in the mammalian central nervous system. Neuron 2000, 25: 317–329.
Zhou Q, Wang S, Anderson DJ. Identification of a novel family of oligodendrocyte lineage-specific basic Helix-loop-Helix transcription factors. Neuron 2000, 25: 331–343.
Takebayashi H, Yoshida S, Sugimori M, Kosako H, Kominami R, Nakafuku M, et al. Dynamic expression of basic helix-loop-helix Olig family members: Implication of Olig2 in neuron and oligodendrocyte differentiation and identification of a new member, Olig3. Mech Dev 2000, 99: 143–148.
Petryniak MA, Potter GB, Rowitch DH, Rubenstein JLR. Dlx1 and Dlx2 control neuronal versus oligodendroglial cell fate acquisition in the developing forebrain. Neuron 2007, 55: 417–433.
Tekki-Kessaris N, Woodruff R, Hall AC, Gaffield W, Kimura S, Stiles CD, et al. Hedgehog-dependent oligodendrocyte lineage specification in the telencephalon. Development 2001, 128: 2545–2554.
Li X, Liu G, Yang L, Li Z, Zhang Z, Xu Z, et al. Decoding cortical glial cell development. Neurosci Bull 2021, 37: 440–460.
Huang H, Rubenstein JL, Qiu M. Cracking the codes of cortical glial progenitors: Evidence for the common lineage of astrocytes and oligodendrocytes. Neurosci Bull 2021, 37: 437–439.
Huang W, Bhaduri A, Velmeshev D, Wang S, Wang L, Rottkamp CA, et al. Origins and proliferative states of human oligodendrocyte precursor cells. Cell 2020, 182: 594-608.e11.
Weng Q, Wang J, Wang J, He D, Cheng Z, Zhang F, et al. Single-cell transcriptomics uncovers glial progenitor diversity and cell fate determinants during development and gliomagenesis. Cell Stem Cell 2019, 24: 707-723.e8.
Yang L, Li Z, Liu G, Li X, Yang Z. Developmental origins of human cortical oligodendrocytes and astrocytes. Neurosci Bull 2022, 38: 47–68.
Cai J, Chen Y, Cai WH, Hurlock EC, Wu H, Kernie SG, et al. A crucial role for Olig2 in white matter astrocyte development. Development 2007, 134: 1887–1899.
Rivers LE, Young KM, Rizzi M, Jamen F, Psachoulia K, Wade A, et al. PDGFRA/NG2 glia generate myelinating oligodendrocytes and piriform projection neurons in adult mice. Nat Neurosci 2008, 11: 1392–1401.
Masahira N, Takebayashi H, Ono K, Watanabe K, Ding L, Furusho M, et al. Olig2-positive progenitors in the embryonic spinal cord give rise not only to motoneurons and oligodendrocytes, but also to a subset of astrocytes and ependymal cells. Dev Biol 2006, 293: 358–369.
Ohayon D, Escalas N, Cochard P, Glise B, Danesin C, Soula C. Sulfatase 2 promotes generation of a spinal cord astrocyte subtype that stands out through the expression of Olig2. Glia 2019, 67: 1478–1495.
Ono K, Takebayashi H, Ikeda K, Furusho M, Nishizawa T, Watanabe K, et al. Regional- and temporal-dependent changes in the differentiation of Olig2 progenitors in the forebrain, and the impact on astrocyte development in the dorsal pallium. Dev Biol 2008, 320: 456–468.
Lu QR, Sun T, Zhu Z, Ma N, Garcia M, Stiles CD, et al. Common developmental requirement for Olig function indicates a motor neuron/oligodendrocyte connection. Cell 2002, 109: 75–86.
Zhou Q, Anderson DJ. The bHLH transcription factors OLIG2 and OLIG1 couple neuronal and glial subtype specification. Cell 2002, 109: 61–73.
Pringle NP, Mudhar HS, Collarini EJ, Richardson WD. PDGF receptors in the rat CNS: During late neurogenesis, PDGF alpha-receptor expression appears to be restricted to glial cells of the oligodendrocyte lineage. Development 1992, 115: 535–551.
Pringle NP, Richardson WD. A singularity of PDGF alpha-receptor expression in the dorsoventral axis of the neural tube may define the origin of the oligodendrocyte lineage. Development 1993, 117: 525–533.
Fruttiger M, Karlsson L, Hall AC, Abramsson A, Calver AR, Boström H, et al. Defective oligodendrocyte development and severe hypomyelination in PDGF-a knockout mice. Development 1999, 126: 457–467.
Zhu Q, Zhao X, Zheng K, Li H, Huang H, Zhang Z, et al. Genetic evidence that Nkx2.2 and Pdgfra are major determinants of the timing of oligodendrocyte differentiation in the developing CNS. Development 2014, 141: 548–555.
Nishiyama A, Watanabe M, Yang Z, Bu J. Identity, distribution, and development of polydendrocytes: NG2-expressing glial cells. J Neurocytol 2002, 31: 437–455.
Polito A, Reynolds R. NG2-expressing cells as oligodendrocyte progenitors in the normal and demyelinated adult central nervous system. J Anat 2005, 207: 707–716.
Zuo H, Nishiyama A. Polydendrocytes in development and myelin repair. Neurosci Bull 2013, 29: 165–176.
Stallcup WB. The NG2 proteoglycan in pericyte biology. Adv Exp Med Biol 2018, 1109: 5–19.
Stallcup WB, Beasley L. Bipotential glial precursor cells of the optic nerve express the NG2 proteoglycan. J Neurosci 1987, 7: 2737–2744.
Cai J, Zhu Q, Zheng K, Li H, Qi Y, Cao Q, et al. Co-localization of Nkx6.2 and Nkx2.2 homeodomain proteins in differentiated myelinating oligodendrocytes. Glia 2010, 58: 458–468.
Qi Y, Cai J, Wu Y, Wu R, Lee J, Fu H, et al. Control of oligodendrocyte differentiation by the Nkx2.2 homeodomain transcription factor. Development 2001, 128: 2723–2733.
Kurrasch DM, Cheung CC, Lee FY, Tran PV, Hata K, Ingraham HA. The neonatal ventromedial hypothalamus transcriptome reveals novel markers with spatially distinct patterning. J Neurosci 2007, 27: 13624–13634.
Xu X, Cai J, Fu H, Wu R, Qi Y, Modderman G, et al. Selective expression of Nkx-2.2 transcription factor in chicken oligodendrocyte progenitors and implications for the embryonic origin of oligodendrocytes. Mol Cell Neurosci 2000, 16: 740–753.
Fu H, Qi Y, Tan M, Cai J, Takebayashi H, Nakafuku M, et al. Dual origin of spinal oligodendrocyte progenitors and evidence for the cooperative role of Olig2 and Nkx2.2 in the control of oligodendrocyte differentiation. Development 2002, 129: 681–693.
Lang J, Maeda Y, Bannerman P, Xu J, Horiuchi M, Pleasure D, et al. Adenomatous polyposis coli regulates oligodendroglial development. J Neurosci 2013, 33: 3113–3130.
Bhat RV, Baraban JM, Johnson RC, Eipper BA, Mains RE. High levels of expression of the tumor suppressor gene APC during development of the rat central nervous system. J Neurosci 1994, 14: 3059–3071.
Brakeman JSF, Gu SH, Wang XB, Dolin G, Baraban JM. Neuronal localization of the adenomatous polyposis coli tumor suppressor protein. Neuroscience 1999, 91: 661–672.
Bin JM, Harris SN, Kennedy TE. The oligodendrocyte-specific antibody ‘CC1’ binds quaking 7. J Neurochem 2016, 139: 181–186.
Takeuchi A, Takahashi Y, Iida K, Hosokawa M, Irie K, Ito M, et al. Identification of qk as a glial precursor cell marker that governs the fate specification of neural stem cells to a glial cell lineage. Stem Cell Rep 2020, 15: 883–897.
Bhat RV, Axt KJ, Fosnaugh JS, Smith KJ, Johnson KA, Hill DE, et al. Expression of the APC tumor suppressor protein in oligodendroglia. Glia 1996, 17: 169–174.
Jahn O, Tenzer S, Werner HB. Myelin proteomics: Molecular anatomy of an insulating sheath. Mol Neurobiol 2009, 40: 55–72.
Bujalka H, Koenning M, Jackson S, Perreau VM, Pope B, Hay CM, et al. MYRF is a membrane-associated transcription factor that autoproteolytically cleaves to directly activate myelin genes. PLoS Biol 2013, 11: e1001625.
Li Z, Park Y, Marcotte EM. A Bacteriophage tailspike domain promotes self-cleavage of a human membrane-bound transcription factor, the myelin regulatory factor MYRF. PLoS Biol 2013, 11: e1001624.
Hornig J, Fröb F, Vogl MR, Hermans-Borgmeyer I, Tamm ER, Wegner M. The transcription factors Sox10 and Myrf define an essential regulatory network module in differentiating oligodendrocytes. PLoS Genet 2013, 9: e1003907.
Emery B, Agalliu D, Cahoy JD, Watkins TA, Dugas JC, Mulinyawe SB, et al. Myelin gene regulatory factor is a critical transcriptional regulator required for CNS myelination. Cell 2009, 138: 172–185.
Bansal R, Warrington AE, Gard AL, Ranscht B, Pfeiffer SE. Multiple and novel specificities of monoclonal antibodies O1, O4, and R-MAb used in the analysis of oligodendrocyte development. J Neurosci Res 1989, 24: 548–557.
Sommer I, Schachner M. Monoclonal antibodies (O1–O4) to oligodendrocyte cell surfaces: An immunocytological study in the central nervous system. Dev Biol 1981, 83: 311–327.
Xiao L, Ohayon D, McKenzie IA, Sinclair-Wilson A, Wright JL, Fudge AD, et al. Rapid production of new oligodendrocytes is required in the earliest stages of motor-skill learning. Nat Neurosci 2016, 19: 1210–1217.
Zhang S, Wang Y, Zhu X, Song L, Zhan X, Ma E, et al. The Wnt effector TCF7l2 promotes oligodendroglial differentiation by repressing autocrine BMP4-mediated signaling. J Neurosci 2021, 41: 1650–1664.
Golan N, Adamsky K, Kartvelishvily E, Brockschnieder D, Möbius W, Spiegel I, et al. Identification of Tmem10/Opalin as an oligodendrocyte enriched gene using expression profiling combined with genetic cell ablation. Glia 2008, 56: 1176–1186.
Jiang W, Yang W, Yang W, Zhang J, Pang D, Gan L, et al. Identification of Tmem10 as a novel late-stage oligodendrocytes marker for detecting hypomyelination. Int J Biol Sci 2013, 10: 33–42.
Moffett JR, Arun P, Ariyannur PS, Garbern JY, Jacobowitz DM, Namboodiri AMA. Extensive aspartoacylase expression in the rat central nervous system. Glia 2011, 59: 1414–1434.
Madhavarao CN, Moffett JR, Moore RA, Viola RE, Namboodiri MAA, Jacobowitz DM. Immunohistochemical localization of aspartoacylase in the rat central nervous system. J Comp Neurol 2004, 472: 318–329.
Butt AM, Ibrahim M, Ruge FM, Berry M. Biochemical subtypes of oligodendrocyte in the anterior medullary velum of the rat as revealed by the monoclonal antibody Rip. Glia 1995, 14: 185–197.
Butt AM, Ibrahim M, Gregson N, Berry M. Differential expression of the L- and S-isoforms of myelin associated glycoprotein (MAG) in oligodendrocyte unit phenotypes in the adult rat anterior medullary velum. J Neurocytol 1998, 27: 271–280.
Schmued L, Slikker W Jr. Black-Gold: A simple, high-resolution histochemical label for normal and pathological myelin in brain tissue sections. Brain Res 1999, 837: 289–297.
Eng LF, Ghirnikar RS, Lee YL. Glial fibrillary acidic protein: GFAP-thirty-one years (1969–2000). Neurochem Res 2000, 25: 1439–1451.
Eng LF, Vanderhaeghen JJ, Bignami A, Gerstl B. An acidic protein isolated from fibrous astrocytes. Brain Res 1971, 28: 351–354.
Boyes BE, KiM SU, Lee V, Sung SC. Immunohistochemical co-localization of S-100b and the glial fibrillary acidic protein in rat brain. Neuroscience 1986, 17: 857–865.
Ludwin SK, Kosek JC, Eng LF. The topographical distribution of S-100 and GFA proteins in the adult rat brain: An immunohistochemical study using horseradish peroxidase-labelled antibodies. J Comp Neurol 1976, 165: 197–207.
Rothermundt M, Peters M, Prehn JHM, Arolt V. S100B in brain damage and neurodegeneration. Microsc Res Tech 2003, 60: 614–632.
Hachem S, Aguirre A, Vives V, Marks A, Gallo V, Legraverend C. Spatial and temporal expression of S100B in cells of oligodendrocyte lineage. Glia 2005, 51: 81–97.
Vives V, Alonso G, Solal AC, Joubert D, Legraverend C. Visualization of S100B-positive neurons and glia in the central nervous system of EGFP transgenic mice. J Comp Neurol 2003, 457: 404–419.
Du J, Yi M, Zhou F, He W, Yang A, Qiu M, et al. S100B is selectively expressed by gray matter protoplasmic astrocytes and myelinating oligodendrocytes in the developing CNS. Mol Brain 2021, 14: 154.
Yang Y, Vidensky S, Jin L, Jie C, Lorenzini I, Frankl M, et al. Molecular comparison of GLT1+ and ALDH1L1+ astrocytes in vivo in astroglial reporter mice. Glia 2011, 59: 200–207.
Cahoy JD, Emery B, Kaushal A, Foo LC, Zamanian JL, Christopherson KS, et al. A transcriptome database for astrocytes, neurons, and oligodendrocytes: A new resource for understanding brain development and function. J Neurosci 2008, 28: 264–278.
Sun W, Cornwell A, Li J, Peng S, Osorio MJ, Aalling N, et al. SOX9 is an astrocyte-specific nuclear marker in the adult brain outside the neurogenic regions. J Neurosci 2017, 37: 4493–4507.
Deneen B, Ho R, Lukaszewicz A, Hochstim CJ, Gronostajski RM, Anderson DJ. The transcription factor NFIA controls the onset of gliogenesis in the developing spinal cord. Neuron 2006, 52: 953–968.
Kang P, Lee HK, Glasgow SM, Finley M, Donti T, Gaber ZB, et al. Sox9 and NFIA coordinate a transcriptional regulatory cascade during the initiation of gliogenesis. Neuron 2012, 74: 79–94.
Chen KS, Harris L, Lim JWC, Harvey TJ, Piper M, Gronostajski RM, et al. Differential neuronal and glial expression of nuclear factor I proteins in the cerebral cortex of adult mice. J Comp Neurol 2017, 525: 2465–2483.
Pajarillo E, Rizor A, Lee J, Aschner M, Lee E. The role of astrocytic glutamate transporters GLT-1 and GLAST in neurological disorders: Potential targets for neurotherapeutics. Neuropharmacology 2019, 161: 107559.
Rao P, Yallapu MM, Sari Y, Fisher PB, Kumar S. Designing novel nanoformulations targeting glutamate transporter excitatory amino acid transporter 2: Implications in treating drug addiction. J Pers Nanomed 2015, 1: 3–9.
Shibata T, Yamada K, Watanabe M, Ikenaka K, Wada K, Tanaka K, et al. Glutamate transporter GLAST is expressed in the radial glia-astrocyte lineage of developing mouse spinal cord. J Neurosci 1997, 17: 9212–9219.
Takasaki C, Okada R, Mitani A, Fukaya M, Yamasaki M, Fujihara Y, et al. Glutamate transporters regulate lesion-induced plasticity in the developing somatosensory cortex. J Neurosci 2008, 28: 4995–5006.
Anlauf E, Derouiche A. Glutamine synthetase as an astrocytic marker: Its cell type and vesicle localization. Front Endocrinol (Lausanne) 2013, 4: 144.
Xin W, Mironova YA, Shen H, Marino RAM, Waisman A, Lamers WH, et al. Oligodendrocytes support neuronal glutamatergic transmission via expression of glutamine synthetase. Cell Rep 2019, 27: 2262-2271.e5.
Ben Haim L, Schirmer L, Zulji A, Sabeur K, Tiret B, Ribon M, et al. Evidence for glutamine synthetase function in mouse spinal cord oligodendrocytes. Glia 2021, 69: 2812–2827.
Sharifi K, Morihiro Y, Maekawa M, Yasumoto Y, Hoshi H, Adachi Y, et al. FABP7 expression in normal and stab-injured brain cortex and its role in astrocyte proliferation. Histochem Cell Biol 2011, 136: 501.
Sharifi K, Ebrahimi M, Kagawa Y, Islam A, Tuerxun T, Yasumoto Y, et al. Differential expression and regulatory roles of FABP5 and FABP7 in oligodendrocyte lineage cells. Cell Tissue Res 2013, 354: 683–695.
Staugaitis SM, Zerlin M, Hawkes R, Levine JM, Goldman JE. Aldolase C/zebrin II expression in the neonatal rat forebrain reveals cellular heterogeneity within the subventricular zone and early astrocyte differentiation. J Neurosci 2001, 21: 6195–6205.
Hubbard JA, Hsu MS, Seldin MM, Binder DK. Expression of the astrocyte water channel aquaporin-4 in the mouse brain. ASN Neuro 2015, 7: 1759091415605486.
Nagao M, Ogata T, Sawada Y, Gotoh Y. Zbtb20 promotes astrocytogenesis during neocortical development. Nat Commun 2016, 7: 11102.
Utz SG, See P, Mildenberger W, Thion MS, Silvin A, Lutz M, et al. Early fate defines microglia and non-parenchymal brain macrophage development. Cell 2020, 181: 557-573.e18.
Imai Y, Ibata I, Ito D, Ohsawa K, Kohsaka S. A novel Geneiba1in the major histocompatibility complex class III region encoding an EF hand protein expressed in a monocytic lineage. Biochem Biophys Res Commun 1996, 224: 855–862.
Ito D, Imai Y, Ohsawa K, Nakajima K, Fukuuchi Y, Kohsaka S. Microglia-specific localisation of a novel calcium binding protein, Iba1. Mol Brain Res 1998, 57: 1–9.
Ohsawa K, Imai Y, Kanazawa H, Sasaki Y, Kohsaka S. Involvement of Iba1 in membrane ruffling and phagocytosis of macrophages/microglia. J Cell Sci 2000, 113(Pt 17): 3073–3084.
Ito D, Tanaka K, Suzuki S, Dembo T, Fukuuchi Y. Enhanced expression of Iba1, ionized calcium-binding adapter molecule 1, after transient focal cerebral ischemia in rat brain. Stroke 2001, 32: 1208–1215.
Harrison JK, Jiang Y, Chen S, Xia Y, Maciejewski D, McNamara RK, et al. Role for neuronally derived fractalkine in mediating interactions between neurons and CX3CR1-expressing microglia. Proc Natl Acad Sci U S A 1998, 95: 10896–10901.
Jung S, Aliberti J, Graemmel P, Sunshine MJ, Kreutzberg GW, Sher A, et al. Analysis of fractalkine receptor CX(3)CR1 function by targeted deletion and green fluorescent protein reporter gene insertion. Mol Cell Biol 2000, 20: 4106–4114.
Haynes SE, Hollopeter G, Yang G, Kurpius D, Dailey ME, Gan WB, et al. The P2Y12 receptor regulates microglial activation by extracellular nucleotides. Nat Neurosci 2006, 9: 1512–1519.
Holness CL, Simmons DL. Molecular cloning of CD68, a human macrophage marker related to lysosomal glycoproteins. Blood 1993, 81: 1607–1613.
Jiang Z, Shih DM, Xia YR, Lusis AJ, de Beer FC, de Villiers WJS, et al. Structure, organization, and chromosomal mapping of the gene encoding macrosialin, a macrophage-restricted protein. Genomics 1998, 50: 199–205.
Galea I, Palin K, Newman TA, Van Rooijen N, Perry VH, Boche D. Mannose receptor expression specifically reveals perivascular macrophages in normal, injured, and diseased mouse brain. Glia 2005, 49: 375–384.
Masuda T, Amann L, Sankowski R, Staszewski O, Lenz M, D’Errico P, et al. Novel Hexb-based tools for studying microglia in the CNS. Nat Immunol 2020, 21: 802–815.
Konishi H, Kobayashi M, Kunisawa T, Imai K, Sayo A, Malissen B, et al. Siglec-H is a microglia-specific marker that discriminates microglia from CNS-associated macrophages and CNS-infiltrating monocytes. Glia 2017, 65: 1927–1943.
Satoh JI, Kino Y, Asahina N, Takitani M, Miyoshi J, Ishida T, et al. TMEM119 marks a subset of microglia in the human brain. Neuropathology 2016, 36: 39–49.
Mrdjen D, Pavlovic A, Hartmann FJ, Schreiner B, Utz SG, Leung BP, et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, aging, and disease. Immunity 2018, 48: 380-395.e6.
Fu Y, Yang M, Yu H, Wang Y, Wu X, Yong J, et al. Heterogeneity of glial progenitor cells during the neurogenesis-to-gliogenesis switch in the developing human cerebral cortex. Cell Rep 2021, 34: 108788.
Trevino AE, Müller F, Andersen J, Sundaram L, Kathiria A, Shcherbina A, et al. Chromatin and gene-regulatory dynamics of the developing human cerebral cortex at single-cell resolution. Cell 2021, 184: 5053-5069.e23.
Sharif N, Calzolari F, Berninger B. Direct in vitro reprogramming of astrocytes into induced neurons. Methods Mol Biol 2021, 2352: 13–29.
Zamboni M, Llorens-Bobadilla E, Magnusson JP, Frisén J. A widespread neurogenic potential of neocortical astrocytes is induced by injury. Cell Stem Cell 2020, 27: 605-617.e5.
Bergles DE, Richardson WD. Oligodendrocyte development and plasticity. Cold Spring Harb Perspect Biol 2015, 8: a020453.
Smart IHM, Dehay C, Giroud P, Berland M, Kennedy H. Unique morphological features of the proliferative zones and postmitotic compartments of the neural epithelium giving rise to striate and extrastriate cortex in the monkey. Cereb Cortex 2002, 12: 37–53.
Hansen DV, Lui JH, Parker PRL, Kriegstein AR. Neurogenic radial glia in the outer subventricular zone of human neocortex. Nature 2010, 464: 554–561.
Acknowledgments
This review was supported by the National Natural Science Foundation of China (31900703 and 32170969) and a Ministry of Science and Technology China Brain Initiative Grant (2022ZD0204701).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors claim that there are no conflicts of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, H., He, W., Tang, T. et al. Immunological Markers for Central Nervous System Glia. Neurosci. Bull. 39, 379–392 (2023). https://doi.org/10.1007/s12264-022-00938-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12264-022-00938-2