Abstract
In the present study, the arsenic bioremediation ability of Bacillus licheniformis (dubbed as A6) was determined. The strain was isolated from metal polluted wastewater and was identified on the basis of 16S rRNA sequence homology with accession number of KX 785,171. The bacterium showed resistance against multiple toxic heavy metals, and MIC against arsenic was 3000 µg/ml. Resistance of the bacterium against other toxic metal ions was 3000 µg/ml (Cr), 50 µg/ml (Hg), 1000 µg/ml (Mn), 4000 µg/ml (Se), 500 µg/ml (Pb), 100 µg/ml (Co), 70 µg/ml (Cd) and 100 µg/ml (Zn). The optimum growth temperature was 37 °C while pH was 7. The strain also showed resistance against commonly used antibiotics except ceftriaxone 30 µg and amoxicillin with clavulanic acid (2:1) 3 µg. B. licheniformis could oxidize arsenite into arsenate 86 and 98% after 48 and 96 h from the medium at optimum growth conditions. Due to its high oxidation potential, B. licheniformis can be used in the biological treatment of wastewater containing arsenic.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Arsenic (As) falls in the category of metalloid and its toxicity is very high (Xiong et al. 2006). It is a ubiquitous metal (Naureen and Rehman 2016). It is odorless and tasteless. The discovery of arsenic goes to Albertus Magnus in 1250. Its name is derived from “arsenikon” which means yellow color. There are three allotropic forms of arsenic named as yellow arsenic, gray arsenic and black arsenic. Its atomic number is 33, while the atomic mass is 74.992. As exist in two oxidation states i.e., + 3 and + 5. Heavy metals have a very toxic effect on human beings as well as their environment. Human beings are intoxicated by heavy metals through exposures, taking food, drinking water containing metals or metalloids (Tamás and Martinoia 2006). Having the potential role in physiological processes, living organisms require these metals as a cofactor for enzymes in biological pathways (Cummings et al. 1999).
The arsenic concentration is almost increasing in every part of the earth (Islam et al. 2004; Kumari et al. 2018; Sher et al. 2020a, b). Due to huge industries in Pakistan, the heavy metals are increasing in our environment (Sher et al. 2020a, b). As a result, As-concentration is also increasing in ground, and drinking water that is above the World Health Organization (WHO) recommended limit i.e., 10 ppb (Prasad et al. 2013). Apart from anthropogenic activities, there are also natural processes by which As come in our environment i.e., land erosion, leaching, volcanism and weathering of rocks (Naureen and Rehman 2016). It has been reported that arsenic has a lethal effect on human beings due to groundwater contamination (Mandal and Suzuki 2002). The long period of arsenic exposure causes arsenicosis. Arsenic causes skin, bladder and lung cancer. As-toxicity depends upon its form, the mobility of As+3 is higher than other arsenic forms that is why its toxicity is higher than others (Kumar et al. 2004). Arsenic disturbs enzyme activity because of interaction with the sulfonyl group (Patra et al. 2004). Řezanka and Sigler (2008) reported that arsenic can replace phosphorus from the DNA due to its similarity with phosphorus.
Most commonly used conventional treatment methods e.g., soil washing, land filling, flushing, physico-chemical extraction and excavation could be used but, these methods are not eco-friendly and expensive due to usage of chemical substances. Mechanisms of arsenic detoxification via microorganisms are oxidation (conversion of arsenite into arsenate), reduction (conversion of arsenate into arsenite) and methylation (addition of methyl group).
Eradication of metal pollutants from the environment has been a challenge for a long time. It is important to establish an efficient and low cost method for the removal of toxic pollutants including heavy metal ions (Sher et al. 2020a). As conventional cleanup strategies are being used to remove the heavy metal ions from the polluted areas but these are expensive and feasible only in small areas. Researchers have found new cost effective methods that include the use of microorganisms (Naureen and Rehman 2016; Sher et al. 2020b), biomass (Jaafari and Yaghmaeian 2019; Sultana et al. 2020) and live plants (Zargar et al. 2013; Ojuederie and Babalola 2017).
Microorganisms especially bacteria are efficient in converting arsenite into arsenate through oxidation–reduction processes. A huge variety of bacterial strains have the ability to cause arsenic oxidation including Agrobacterium tumefaciens (Kashyap et al. 2006), Pseudomonas arsenitoxidans (Matlakowska et al. 2008), P. lubricans (Rehman et al. 2010), Pantoea sp., Enterobacter sp., Pseudomonas sp. Comamonas sp. (Liao et al. 2011) and Microbacterium oxydans (Sarkar et al. 2013). The arsenate, oxidized form of arsenic, has very low toxicity due to its low mobility as compared to the arsenite, the reduced form of arsenic.
The isolation and characterization of arsenic resistant bacteria from industrial wastewater were the main objective of this study. The bacterium was optimized for its growth conditions and was also checked for its As-oxidation ability under different parameters.
Materials and methods
Wastewater sample collection
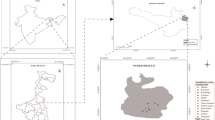
Industrial wastewater was collected from the chemical industry from Sheikhupura (Pakistan) which has a long history for metal usage. Its geographical coordinates are 31.7167° N, 73.9850° E. The samples were taken in sterile bottles and some physical parameters i.e., temperature, pH and color were also noted.
Isolation and selection of bacterial strain
Bacterial strains resistance against arsenic, isolated from wastewater samples, was determined by providing arsenic in Luria–Bertani (LB) medium (Sigma-Aldrich, USA). The industrial wastewater sample was diluted tenfold in a normal saline and 10–3, 10–4 and 10–5 dilutions were used. Then, wastewater sample dilution (100 µl) was spread on LB agar plates, which were already supplemented with 400 µg sodium arsenite/ml for the isolation of arsenite resistant bacterial strains. The autoclaved LB agar plates were prepared for the spreading of samples. The following composition of LB agar was used, 10 g NaCl, 10 g tryptone, 5 g yeast extract, 15 g agar in one liter of distilled water. The colonies of bacteria were observed after incubation at 37 °C for 24 h. The different bacteria having different morphology were selected and streaked again and again until purified bacterial strains were obtained. Quadrant streaking was performed to get pure culture of the isolates.
Morphological characterization
On the basis of shaped and appearance, different bacterial strains from mix culture were selected for further streaking to get pure bacterial strain. After getting a pure bacterial cell culture, its morphology was observed under a light microscope. The morphology of the colony was observed with naked eyes.
Biochemical characterization of isolated bacterium
Different biochemical tests i.e., gram staining, spore staining, acid fast staining, capsular staining, oxidase test, catalase test, citrate utilization test, indole test, methyl red test, Voges–Proskauer test, triple sugar iron test and motility test were performed for bacterial identification (Cappuccino and Sherman 2008).
Arsenic oxidation screening by AgNO3 method
For As3+ oxidizing bacterial screening, AgNO3 method was used (Simeonova et al. 2004). Arsenic containing (100 µg/ml) nutrient-agar plates were inoculated with isolated metal resistant bacteria and then incubated at 37 °C for 72 h. After the appearance of bacterial growth, 0.1 M AgNO3 was flooded on the plates and then placed at room temperature for overnight.
Molecular identification
For molecular identification of bacterial strain, genomic DNA was extracted according to Carozzi et al. (1991). The 16S rRNA gene was amplified by using the universal primer through PCR. The 16S rRNA gene product was sequenced and compared with already gene bank data in NCBI.
Optimum bacterial growth conditions
The bacterial growth conditions were determined on the bases of temperature and pH.
Optimization of temperature
For determination of optimum growth temperature, 5 ml N-broth was taken in three sets of tubes for three temperatures, i.e., 28, 37 and 42 °C. Test tubes containing N-broth were autoclaved, and inoculated with overnight log phase culture and incubated at respective temperature on shaker for 24 h. After incubation, optical density (OD) was taken at 600 nm using a LAMBDA 650 UV/Vis spectrophotometer (PerkinElmer, USA).
Optimization of pH
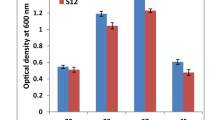
For determination of optimum pH, six sets of nutrient broth in tubes with different pH (5, 6, 7, 8, 9, 10) were prepared. These tubes were inoculated with overnight log phase culture and incubated on shaker at 37 °C for 24 h. After incubation, OD was taken at 600 nm with the help of spectrophotometer.
Bacterial growth curves
For the isolated bacterial strain, the growth curve was formed in the presence and absence of arsenic. Autoclaved N- broth containing 100 µg/ml arsenite in 250 ml flasks was used as a stress medium. The broth was inoculated with I ml overnight bacterial culture and incubated on shaker at 37 °C. Then, 2 ml of aliquots were taken at a regular time interval (4, 8, 12, 16, 20, 24 h) from both stress and non- stress cultures. The OD was measured at 600 nm.
Bioremediation assay
The bioremediation ability of the bacterium was determined at different temperatures, pH and arsenic concentrations. For this purpose, 5 ml of autoclaved N-broth was taken in test tubes and inoculated with overnight bacterial culture. Then, these tubes were placed in a shaking incubator for 96 h. After 2 days, 1 ml broth culture was taken and centrifuged at 5000 rpm for 5 min. Then supernatant was taken and checked for arsenite concentration by Safranin O dye method. After 4 days of incubation, similar process was repeated. Finally, supernatant was used for arsenite determination by the spectrophotometer process. The As+3 oxidation potential of bacterium was determined at 25, 30, 37 and 42 °C after 48 and 96 h. Likewise, As+3 oxidation ability of strain A6 was checked at pH (3, 5, 7 and 9). The As+3 oxidation potential B. licheniformis A6 was determined at 100, 300, 500 and 1000 µg/ml. Each experiment was done in triplicate.
Arsenic estimation
Arsenic was estimated by a spectrophotometric method using the safranin O dye (Pasha and Narayana 2008). In this method, a sample (1 ml) containing As was taken into a 10 ml tube and then added KIO3 (1 ml) solution followed by 1 M HCl. The mixture was gently shaken till the yellow color appeared and then added 0.5 ml of Safranin dye solution. Then the mixture was shaken for 2 min, and added 2 ml of acetate buffer to maintain pH 4, and diluted up to 100 ml by adding distilled water. Then 1 ml of the mixture was taken into the cuvettes, and OD was determined by a spectrophotometer at 532 nm.
Effect of sodium chloride on bacteria
The bacterium was cultured in the presence of As 100 µg/ml and NaCl (0–500 mM). NaCl was added in LB-broth according to above mentioned concentration, and OD was determined by spectrophotometer at 532 nm.
Multiple metal resistances
Multiple metal resistance of the given isolate was checked against different metal ions. The following metals were used Cr, Mn, Pb, Se, Co, Cd, Zn and Hg in the given concentrations 100, 500, 1000 µg/ml, up to 5000 µg/ml. Nutrient-agar plates supplemented with these metals were set, streaked the bacteria and incubated at 37 °C for 24 h.
Antibiotic Resistance
The given bacteria were screened for its resistance against nine regularly used antibiotics disks. Norfloxacin (30 µg), Imipenem (10 µg), amoxicillin/clavulanic acid (2:1), tetracycline (30 µg), ceftriaxone (30 µg), ciprofloxacin (5 µg) and nalidixic acid (30 µg) were used. Muller Hinton-agar plates were prepared with the lawn of bacteria, and then placed the disks of antibiotics onto the MH-agar plates. Results were observed by the growth of bacterial strain on agar plates containing antibiotic disk after 24 h of incubation at \(37 ^\circ \mathrm{C}\).
Statistical analysis
All the experiments were performed in triplicates. Also mean and standard error was calculated for all the experiments.
Results and discussion
Isolation and metal resistance
Physico-chemical characteristics of the wastewater samples determined were color, temperature and pH. Color of the wastewater was dark gray. The temperature and pH ranged between 25 and 32 °C and 6–7.4, respectively. Initially, 37 isolates were isolated from 3 different wastewater samples, but on the basis of minimum inhibitory concentration regarding arsenic 9 strains were selected named as (IT6, A6, S1, S4, S9, S12, P6, A15a and A15b) which were resistant more than 500 µg/ml arsenic. While the other strains showed low MICs in the range of (100–400 µg/ml). While in the nine selected strains, strain A6 was selected because of higher MIC which was 3000 µg/ml and strong arsenic oxidizing ability.
B. licheniformis was found resistant against arsenite upto 3000 µg/ml. The bacterium also showed resistance against multiple metals ion, Cr (3000 µg/ml), Hg (50 µg/ml), Mn (1000 µg/ml), Se (4000 µg/ml), Pb (500 µg/ml), Co (100 µg/ml), Cd (70 µg/ml) and against Zn the resistance was (100 µg/ml). In the present investigation, nine antibiotics disks were used, against the arsenic resistant bacterial strain. The strain A6 showed weak resistance against cefuroxime sodium and nalidixic acid but A6 showed strong resistance against norfloxacin, amikacin, imipenem, tetracycline and ciprofloxacin. The isolated strain was sensitive against amoxicillin/clavulanic acid (2:1) 30 µg and ceftriaxone 30 µg disks.
The diversity of arsenic resistance bacteria is high in such an environment where arsenic level is high or medium (Cai et al. 2009). In one of the studies, 12 strains of bacteria with an arsenite MICs greater than 20 mM were obtained from the high arsenic contaminated soil sample (Cai et al. 2009). Selective pressure due to high arsenic concentration leads to low diversity of metal resistant organisms (Achour-Rokbani et al. 2007; Jackson et al. 2005). Microorganisms take some time to build resistance against metals (Pennanen et al. 1996). Turpeinen also ascertained that a variety of arsenic resistance bacteria is high in complex contaminated soil samples from different metals compared to dry soil (Turpeinen et al. 2004).
In the present study, cross metal resistance of bacterium was checked against heavy metals i.e., chromium, zinc, manganese, and it was found that strain AS6 has resistance against most of the metals. In one of the other study, it was found that the bacteria named as B. safensis MS11 has high resistance regarding arsenite and arsenate (40, 400 mM) and for boron (200 mM) along with the high concentration of salt 15% in LB-medium. B. safensis MS11 also has resistance against Cd, Cr, Cu, Ni, Pb and Zn (Raja and Omine 2012). B. safensis can be used in decontamination of soil having high salt concentration for the removal of arsenic. The arsenic resistant bacteria belong to genera (Actinobacteria, Microbacterium, Pseudomonas and Rhizobium), apart from arsenic other metals resistance mechanisms are also found in these genera (Paul et al. 2014).
Characterization of metal resistant bacterial strain
The bacterial colony shape was circular, medium size and color was light white, margin was entire, elevation was flat and transparency was opaque, and texture was smooth. The organism was gram positive rod, catalase negative and oxidation test was positive (Table 1). The 16S rRNA gene was sequenced partially and submitted to NCBI to check the similarity and the blast query indicated that this gene has highest homology to the Bacillus licheniformis. The nucleotide sequences were deposited to Genbank in FASTA format under the accession number of KX 785,171.
Arsenic presence in the industrial wastewater is lethal for human beings as well as for life in water bodies and terrestrial environments (Mateos et al. 2006). These kinds of heavy metal can easily be detoxified by the way of bioremediation. The biggest problem of arsenic is its carcinogenic effect due to its toxicity (Shakoori et al. 2010). In many studies, mostly the arsenic oxidizing bacteria were recognized from Acinetobacter, Comamonas, Pseudomonas, Stenotrophomonas, Delftia, Agrobacterium and Bacillus as the major genera (Cai et al. 2009).
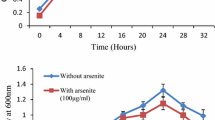
The optimum pH for the isolated B. lichenifromis was 7 while optimum temperature was 37 °C. The growth curves were studied under As-stress and non-stress condition. In the presence of As, the lag phase of the given strain was extended for a bit of time (Fig. 1). Isolated strain was grown in arsenic presence (100 µg/ml) and NaCl (0–500 mM). It was interpreted that NaCl concentration was inversely proportional to the growth of bacteria after a certain limit. Maximum growth was observed at concentration of 100 mM NaCl and minimum when the NaCl was 500 mM.
Arsenite oxidizing potential of the bacterial strain
The brownish precipitate appearance after the use of 0.1 M AgNO3 on the streak growth plates indicates that the isolated strain has arsenic oxidation potential. The AgNO3 reaction depends upon the arsenic ion either arsenite or arsenate. While the reaction among AgNO3 and As (V) generate yellow precipitate (Fig. 2).
Arsenic bioremediation was determined at 25, 30, 37 and 42 °C with the 100 µg/ml. As+3 concentration after 48 and 96 h. Strain A6 showed maximum arsenite oxidation potential 86% after 48 and 98% after 96 h at 37 °C\(.\) At 25\(^\circ \mathrm{C},\) 56% bioremediation activity was observed while at 30\(^\circ \mathrm{C},\) 75% and 42 °C, it was 75 and 17%, respectively. After 96 h incubation, the bacterium showed 64, 83 and 23%. As+3 oxidation potential at 25, 30, 37 and 42 °C, respectively (Fig. 3a).
As+3 oxidation potential was determined at pH 3, 5, 7 and 9 with 100 µg As/ml after 48 and 96 h, Strain A6 had As+3 oxidation potential of 86 and 98% after 48 and 96 h at pH 7. At pH 3 the strain showed 9% oxidation potential, at pH 5, 71%, while at pH 9 the oxidation ability was about 82%. The oxidation potential of the bacterium A6 was 14% (pH3), 85% (pH5), 98% (pH7) and 92% (pH9) after 96 h incubation at 37 °C (Fig. 3b).
As+3 oxidation potential of the bacterium was also checked at various arsenite concentrations i.e., 100, 300, 500 and 1000 µg/ml. After 48 and 96 h. Strain A6 had As+3 oxidation potential of 86% after 48 h and 98% after 96 h at 100 µg/ml. At concentration 300 µg/ml, B. licheniformis showed oxidation ability of 81%, at 500 µg /ml, 56% and at 1000 µg/ ml, the oxidation ability was 19% after 48 h of incubation. The oxidation ability of the bacterium after 96 h of incubation was 98% (100 µg/ml), 86% (300 µg/ml), 56% (500 µg/ml) and 30% (1000 µg/ml) (Fig. 3c).
Bacteria and other organisms play a very important role in different biogeochemical cycles including arsenic cycle, to convert different metal ions into their different oxidation states having different solubility, mobility and toxicity (Silver and Phung 2005). Microorganisms like bacteria, fungi, ciliates, algae, mosses, macrophytes used different ways for detoxification or removal of metals from different environments (Volesky and Holan 1995). Gadd (1990) and Lovley and Coates (1997) reported that microorganisms response to metals present in them by many ways such as Bio sorption of metals, Adsorption, interaction of metal with the molecules in cytosol, entrance into the capsule of cell, precipitation, oxidation of metal (arsenite into arsenate), reduction of metal (arsenate into arsenite) as well as protein-DNA adduct formation (Zhitkovich and Costa 1992) and production of some protein due to metal stress (Ballatori 2002).
The highest arsenic oxidizing bacteria are Pseudomonas, Agrobacterium and Corynebacterium glutamicum, the later one is industrially important for the amino acid manufacturing having MIC (up to 12 mM arsenite and > 400 mM arsenite) (Lovley and Coates 1997). In the present study, the strain A6 showed resistance against As up to 40 mM.
In one of the studies, it was examined that the arsenite oxidation potential was 92% for B. cereus and 88% for A. junii after 6 days of incubation in industrial wastewater (Naureen and Rehman 2016). While the above isolated strain Bacillus licheniformis showed arsenite oxidation potential 86 and 98% after 48 and 96 h, respectively. Due to its strong oxidation potential, this isolated strain can be used to purify wastewater containing As and other heavy metals containing wastewater.
Conclusion
In the present investigation, bacterial strain, isolated from industrial wastewater, showed MIC value upto 40 mM against arsenite. The bacterium was also able to resist other toxic metal ions e.g., Cd, Cr, Mn, Zn, Se, Hg, Pb, Co. The arsenite oxidizing potential of B. licheniformis was 86 and 98% after 48 and 96 h at pH 7. This bacterial strain could be used to treat the industrial waste to ameliorate the toxic metal ions. For practical use, further research work is required to investigate arsenite oxidizing potential of this bacterial strain with experiments using original industrial wastewater in batch and continuous bioreactors.
References
Achour-Rokbani A, Bauda P, Billard P (2007) Diversity of arsenite transporter genes from arsenic-resistant soil bacteria. Res Microbiol 158:128–137
Ballatori N (2002) Transport of toxic metals by molecular mimicry. Environ Health Perspect 110:689–694
Cai L, Liu G, Rensing C, Wang G (2009) Genes involved in arsenic transformation and resistance associated with different levels of arsenic-contaminated soils. BMC Microbiol 9:4
Cappuccino JG, Sherman N (2008) Microbiology: a laboratory manual, vol 9. Pearson/Benjamin Cummings, Boston
Carozzi NB, Kramer VC, Warren GW, Evola S, Koziel MG (1991) Prediction of insecticidal activity of Bacillus thuringiensis strains by polymerase chain reaction product profiles. Appl Environ Microbiol 57:3057–3061
Cummings DE, Caccavo F, Fendorf S, Rosenzweig RF (1999) Arsenic mobilization by the dissimilatory Fe (III)-reducing bacterium Shewanella alga BrY. Environ Sci Technol 33:723–729
Gadd GM (1990) Heavy metal accumulation by bacteria and other microorganisms. Experientia 46:834–840
Islam FS, Gault AG, Boothman C, Polya DA, Charnock JM, Chatterjee D, Lloyd JR (2004) Role of metal-reducing bacteria in arsenic release from Bengal delta sediments. Nature 430(6995):68–84
Jaafari J, Yaghmaeian K (2019) Optimization of heavy metal biosorption onto freshwater algae (Chlorella coloniales) using response surface methodology (RSM). Chemosphere 217:447–455
Jackson C, Harrison K, Dugas S (2005) Enumeration and characterization of culturable arsenate resistant bacteria in a large estuary. Syst Appl Microbiol 28:727–734
Kashyap DR, Botero LM, Lehr C, Hassett DJ, McDermott TR (2006) A Na+:H+ antiporter and a molybdate transporter are essential for arsenite oxidation in Agrobacterium tumefaciens. J Bacteriol 188:1577–1584
Kumar PR, Chaudhari S, Khilar KC, Mahajan S (2004) Removal of arsenic from water by electrocoagulation. Chemosphere 55:1245–1252
Kumari P, Rastogi A, Shukla A, Srivastava S, Yadav S (2018) Prospects of genetic engineering utilizing potential genes for regulating arsenic accumulation in plants. Chemosphere 211(1):397–406
Liao VHC et al (2011) Arsenite-oxidizing and arsenate-reducing bacteria associated with arsenic-rich groundwater in Taiwan. J Contaminant Hydrol 123:20–29
Lovley DR, Coates JD (1997) Bioremediation of metal contamination. Curr Opin Biotechnol 8:285–289
Mandal BK, Suzuki KT (2002) Arsenic round the world: a review. Talanta 58:201–235
Mateos LM, Ordóñez E, Letek M, Gil JA (2006) Corynebacterium glutamicum as a model bacterium for the bioremediation of arsenic. Inter Microbiol 9:207–215
Matlakowska R, Drewniak L, Sklodowska A (2008) Arsenic-hypertolerant pseudomonads isolated from ancient gold and copper-bearing black shale deposits. Geomicrobiol J 25:357–362
Naureen A, Rehman A (2016) Arsenite oxidizing multiple metal resistant bacteria isolated from industrial effluent: their potential use in wastewater treatment. World J Microbiol Biotechnol 32:1–9
Ojuederie OB, Babalola OO (2017) Microbial and plant-assisted bioremediation of heavy metal polluted environments: a review. Int J Environ Res Public Health 14(12):1504
Pasha C, Narayana B (2008) Determination of arsenic in environmental and biological samples using toluidine blue or safranine O by simple spectrophotometric method. Bull Environ Contam Toxicol 81:47–51
Patra M, Bhowmik N, Bandopadhyay B, Sharma A (2004) Comparison of mercury, lead and arsenic with respect to genotoxic effects on plant systems and the development of genetic tolerance. Environ Exp Bot 52:199–223
Paul D, Poddar S, Sar P (2014) Characterization of arsenite-oxidizing bacteria isolated from arsenic-contaminated groundwater of West Bengal. J Environ Sci Hlth, Part A 49:1481–1492
Pennanen T, Frostegard A, Fritze H, Baath E (1996) Phospholipid fatty acid composition and heavy metal tolerance of soil microbial communities along two heavy metal-polluted gradients in coniferous forests. Appl Environ Microbiol 62:420–428
Raja CE, Omine K (2012) Arsenic, boron and salt resistant Bacillus safensis MS11 isolated from Mongolia desert soil. Afr J Biotechnol 11:2267–2275
Rehman A, Butt SA, Hasnain S (2010) Isolation and characterization of arsenite oxidizing Pseudomonas lubricans and its potential use in bioremediation of wastewater. Afr J Biotechnol 9:1493–1498
Řezanka T, Sigler K (2008) Biologically active compounds of semi-metals. Phytochemistry 69:585–606
Prasad KS, Ramanathan A, Paul J, Subramanian V, Prasad R (2013) Biosorption of arsenite (As+3) and arsenate (As+5) from aqueous solution by Arthrobacter sp. Biomass Environ Technol 19(1):2701–2708
Sarkar A, Kazy SK, Sar P (2013) Characterization of arsenic resistant bacteria from arsenic rich groundwater of West Bengal, India. Ecotoxicology 22:363–376
Shakoori FR, Aziz I, Rehman A, Shakoori A (2010) Isolation and characterization of arsenic reducing bacteria from industrial effluents and their potential use in bioremediation of wastewater. Pak J Zool 42:331–338
Sher S, Hussain SZ, Rehman A (2020a) Phenotypic and genomic analysis of multiple heavy metal resistant Micrococcus luteus strain AS2 isolated from industrial waste water and its potential use in arsenic bioremediation. Appl Microbiol Biotechnol 104(5):2243–2254
Sher S, Hussain SZ, Rehman A (2020b) Multiple resistance mechanisms in Staphylococcus sp. strain AS6 under arsenite stress and its potential use in amelioration of wastewater. J King Saud Uni Sci 32:3050–3058
Silver S, Phung LT (2005) A bacterial view of the periodic table: genes and proteins for toxic inorganic ions. J Indus Microbiol Biotechnol 32:587–605
Simeonova DD, Lièvremont D, Lagarde F, Muller DA, Groudeva VI, Lett M-C (2004) Microplate screening assay for the detection of arsenite-oxidizing and arsenate-reducing bacteria. FEMS Microbiol Lett 237:249–253
Sultana N, Hossain SMZ, Mohammed ME, Irfan MF, Haq B, Faruque MO, Razzak SA, Hossain MM (2020) Experimental study and parameters optimization of microalgae based heavy metals removal process using a hybrid response surface methodology-crow search algorithm. Sci Rep 10:15068
Tamás MJ, Martinoia E (2006) Molecular biology of metal homeostasis and detoxification. Springer
Turpeinen R, Kairesalo T, Häggblom MM (2004) Microbial community structure and activity in arsenic-, chromium-and copper-contaminated soils. FEMS Microbiol Ecol 47:39–50
Volesky B, Holan Z (1995) Biosorption of heavy metals. Biotechnol Prog 11:235–250
Xiong J, Wang W, Fan H, Cai L, Wang G (2006) Arsenic resistant bacteria in mining wastes from Shangrao coal mine of China. Environ Sci Technol 1:535–540
Zargar M, Sarrafzadeh MH, Taheri B, Tavakoli O (2013) The surveying of soil and groundwater pollution in a petroleum refinery and the potential of bioremediation for oil decontamination. Petrol Sci Technol 31:2585–2595
Zhitkovich A, Costa M (1992) A simple, sensitive assay to detect DNA-protein cromlinks in intact cells and in vivo. Carcinogenesis 13:1485–1489
Funding
No funding was received for this research work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
The current study does not contain experiments involving animals/Humans.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sher, S., Sultan, S. & Rehman, A. Characterization of multiple metal resistant Bacillus licheniformis and its potential use in arsenic contaminated industrial wastewater. Appl Water Sci 11, 69 (2021). https://doi.org/10.1007/s13201-021-01407-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13201-021-01407-3