Abstract
Background
Monkeypox is endemic to African region and has become of Global concern recently due to its outbreaks in non-endemic countries. Although, the disease was first recorded in 1970, no monkeypox specific drug or vaccine exists as of now.
Methods
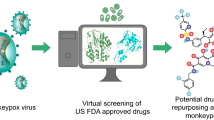
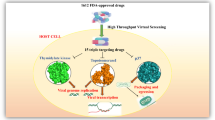
We applied drug repositioning method, testing effectiveness of currently approved drugs against emerging disease, as one of the most affordable approaches for discovering novel treatment measures. Techniques such as virtual ligand-based and structure-based screening were applied to identify potential drug candidates against monkeypox.
Results
We narrowed down our results to 6 antiviral and 20 anti-tumor drugs that exhibit theoretically higher potency than tecovirimat, the currently approved drug for monkeypox disease.
Conclusions
Our results indicated that selected drug compounds displayed strong binding affinity for p37 receptor of monkeypox virus and therefore can potentially be used in future studies to confirm their effectiveness against the disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Monkeypox is a disease caused by monkeypox virus, an enveloped double-stranded DNA virus, which belongs to the family Orthopoxviruses and include vaccinia, cowpox, smallpox and variola (Mpox (Monkeypox), 2023). The first human monkeypox case was identified in 1970 in the Demographic Republic of Congo after a recent eradication of smallpox in the region (Breman et al. 1980). Since then, cases of monkeypox were reported in total of 11 countries in the African continent (Mpox (Monkeypox), 2023). The first monkeypox outbreak outside of Africa occurred in 2003 in the United States of America (CDC 2022). The disease gained global attention in May 2022, after multiple cases of human monkeypox were identified in non-endemic countries (Mpox (Monkeypox), 2023). The natural reservoir of the virus remains unknown. According to Center for Disease Control and Prevention (CDC) monkeypox spread across the World with a total of 31,800 registered cases as of August 9, 2022 (CDC 2022). 31,425 cases occurred in countries with no previous historical reports of the virus outbreaks (CDC 2022). The virus can be transmitted through animal bites, direct contact with bodily fluids, and through skin-to-skin and intimate contacts with an infected person (CDC 2022). The major symptom of monkeypox disease is a rash on the surface of the skin; additional symptoms include fever, chills, swollen lymph nodes, weakness, muscle aches, headache and flu like symptom (CDC 2022).
p37 (encoded by F13L gene), the main extracellular protein, which is encoded by all members of the family Orthopoxviruses, is a 372 amino acid protein expressed during the late phase of infection. It participates in formation of the viral double membrane when the virus exits from the host cell (Blasco and Moss 1991). p37 interacts with elements of trans-Golgi (TGN) that envelope viral particles leading to formation of the triple-wrapped virus prior to being transported to cell surface and release (Blasco and Moss 1991). The protein has no analogs in the host organism making it an ideal drug target (Chen et al. 2009). Tecovirimat, also known as ST-246, was specifically developed to combat orthopoxviruses; its mechanism of action is specifically targeted towards p37, as it has been identified as the drug’s target through genetic mapping of tecovirimat-resistant mutant viruses. (Yang et al. 2005). In vivo tests demonstrated that oral administration of tecovirimat to the infected mice protected them from lethal orthopoxvirus infection (Yang et al. 2005). Study showed that p37 inhibition by tecovirimat prevented its interaction with Rab9 GTPase and TIP47, thus, halting the formation of cell-associated enveloped viruses (CEV) and extracellular enveloped viruses (EEV) (Chen et al. 2009).
Currently, no specific drug targeting monkeypox is available on the market. However, due to its genetic similarity in key regions to smallpox (96.3%), tecovirimat, a drug approved for smallpox, can be prescribed for monkepox treatment as well (Duraffour et al. 2010; Merchlinsky et al. 2019; Desai et al. 2022). Traditional methods of drug development may take decades before a successful drug candidate can be developed and delivered to patients. Drug repositioning or repurposing is a method of determining new applications of existing drugs to treat common and rare diseases (Ashburn and Thor 2004; Hurle et al. 2013; Li et al. 2016). There are several benefits involved when utilizing this method, including lower health risk, since drugs used have already been shown to be sufficiently safe, shorter time period before approving a drug for treatment and significantly less required investment (Ashburn and Thor 2004; Hurle et al. 2013; Li et al. 2016). Due to the importance of p37 protein, we selected it as a target for in silico drug repositioning conducted in this work.
Materials and methods
Data retrieval
The p37 envelope protein sequence was retrieved from National Center for Biotechnology Information (NCBI; accession number: NP_536472.1) (Sayers et al. 2022). Structure of tecovirimat in canonical simplified molecular-input line-entry system (SMILES) was retrieved from ChEMBL database (ChEMBL ID: CHEMBL1257073) (Gaulton et al. 2017). 3D structure of tecovirimat is present in Fig. 1c and d.
(a) A complete docked model of p37 protein and tecovirimat with the binding pocket (red); (b) A close-up view on the binding pocket of p37 with active amino acid residues (red) interacting with the ligand (cyan); (c) A schematic diagram of p37-tecovirimat interaction; (d) 3D representation of tecovirimat structure
3D structures preparation
We utilized Iterative Threading ASSEmbly Refinement (I-TASSER; https://zhanggroup.org/I-TASSER/) web-server to prepare p37 protein using the template structure and determine the binding pocket for ligands (Zhang 2008; Roy et al. 2010). This server identifies suitable template found in Protein Data Bank (PDB) using threading method and then performs template-based fragment assembly of the final model(Berman et al. 2000, 2003). PROCHECK and ProSA-Web servers were utilized to assess the quality of the resulting protein model (Laskowski et al. 1993, 1996; Wiederstein and Sippl, 2007).
Open Babel software was used in this work to prepare 3D structure of ligands with added hydrogens atoms from their corresponding SMILES (O’Boyle et al. 2011).
Virtual screening
To identify potential ligands, we performed ligand-based virtual screening (LBVS) using SWISS-SIMILARITY (SS) server of SwissDrugDesign group (http://www.swisssimilarity.ch/) and structure-based virtual screening (SBVS) using PHARMIT (https://pharmit.csb.pitt.edu/) (Sunseri and Koes 2016; Zoete et al. 2016; Bragina et al. 2022). The screened compounds from PHARMIT were retrieved using ChEMBL web-resource client (https://github.com/chembl/chembl_webresource_client) (Davies et al. 2015). LBVS allows identification of potential ligands based on their similarity with the query structure, while SBVS utilizes information of binding pocket of protein target to determine suitable candidates (Hamza et al. 2012; Li and Shah 2017).
p37 – ligand docking
Chimera 1.15 software was used to visualize and analyze protein and ligand (Pettersen et al. 2004). Hydrogens were added to the structures and DockPrep plugin was utilized to prepare receptor structure for docking analysis (Pettersen et al. 2004; Allen et al. 2015). Protein-ligand docking was performed with AutoDock Vina software (Trott and Olson 2009). It is a molecular modelling simulation software, which docks ligands to a box, defined by the set of x, y and z coordinates (Trott and Olson 2009). AutoDock Vina is a direct successor of Autodock and is significantly faster, being able to utilize multiple cores for the simulations (Trott and Olson 2009). Box size was set to 20 for all coordinates and centered to x = 73.38, y = 69.46 and z = 64.25; the energy range was set to 3 with exhaustiveness level of 8 and 10 modes. Binding energy of the best protein-ligands modes was assessed and LigProt software was used to visualize protein-ligand interactions (Wallace et al. 1995; Laskowski and Swindells 2011). We performed p37 – tecovirimat docking and used the results as a reference for other docking simulations. LigProt software was utilized to visualize protein-ligand interaction (Supplementary Fig. 1).
Results
Receptor structure preparation
Tungstate-inhibited phospholipase D (PDB: 1V0R) was selected as a template for the p37 protein. The resulting C-score predicted by I-TASSER was 0.23 with cluster size of 8, highlighting the adequate quality of this model (Fig. 2a, b). C-score is a confidence score for predicted models, it is based on the significance of threading template alignments and the convergence parameters of the structure assembly simulations (Zhang 2008; Roy et al. 2010). ProSA-Web evaluation revealed that the resulting model had a Z-score of -8.18 (Fig. 2d), which is a good indicator. Z-score is an indicator of general model quality, which represents a comparative analysis of the query model against other proteins available in PDB (Wiederstein and Sippl, 2007). PROCHECK web-server analysis indicated the following results: 233 AAs (68.7%) in the most favored regions, 87 (25.7%) AAs in additional allowed regions, 15 (4.4%) AAs in generously allowed regions and 4 (1.2%) AAs in disallowed regions (Fig. 2c). The ligand-binding site residues were predicted as follows: Phe52, Leu118, Cys120, Ser135, Asn312, Lys314, Asn329 and Asp331.
(a) A 3D model of p37 protein (ribbon) with highlighted active site residues (red); (b) A surface representation of p37 protein model with highlighted binding pocket (red); (c) Ramachandran plot (PROCHECK) of p37 model (233 (68.7%) residues in most favored regions, 87 (25.7%) residues in additional allowed regions, 15 (4.4%) residues in generously allowed regions, 4 (1.2%) residues in disallowed regions); (d) Validation of the final model with ProSA-Web (Z-score = -8.18)
Ligand-based virtual screening: SWISS-SIMILARITY
SWISS-SIMILARITY server allows to perform ligand-based virtual screening of chemical libraries and retrieves compounds most similar to the user’s query (Zoete et al. 2016; Bragina et al. 2022). We used tecovirimat as an input and retrieved a total of 29 approved drug compounds from ChEMBL database using variety of methods (Supplementary Table 1). The similarity of selected compounds to tecovirimat varied between 0.852 (CHEMBL3622821, upadacitinib) and 0.023 (CHEMBL480, Lansoprazole) (Supplementary able 1).
Structure-based virtual screening: PHARMIT
PHARMIT is a web-server designed to perform structure-based virtual screening using pharmacophore, molecular shape and energy minimization (Sunseri and Koes 2016). We utilized exclusive shape with tolerance 1.0 and selected pharmacophore parameters were set with radius of 1. As with LBVS method, we screened ChEMBL30 database and screened a total of 11,612,000 compounds and only phase 4 drugs were retrieved for the final analysis. As a result, a total of 592 phase 4 drugs were selected for the docking analysis (Supplementary Table 2).
Receptor – ligand docking
The p37 – tecovirimat docking demonstrated strong binding with the top model scoring at -8.2 kcal/mol, which was subsequently used as a reference point for further analysis (Fig. 1a, b. The complete list of docking results for ligands retrieved from ligand screening of SWISS-SIMILARITY and PHARMIT can be found in Supplementary Tables 1 and Supplementary Table 2 correspondingly. The top scoring ligand obtained via SWISS-SIMILARITY was durasteride, an antiandrogenic compound used to treat symptomatic benign prostatic hyperplasia, with − 9.3 kcal/mol binding energy and the lowest scoring ligand was sitagliptin, a dipeptidyl peptidase-4 (DPP-4) inhibitor used to manage type 2 diabetes melitus, with − 8.2 kcal/mol binding energy score (Wishart et al. 2018). The docking analysis of 592 phase 4 drugs screened with the help of PHARMIT produced multiple results (56 drug compounds) with top scoring models ranging from − 9.6 - -8.2 kcal/mol binding energy (Table 1). Among selected drugs from PHARMIT screening six compounds were identified as antiviral drugs including glecaprevir (-9.3 kcal/mol), bictegravir (-8.8 kcal/mol), velpatasvir (-8.6 kcal/mol), remdesivir (-8.3 kcal/mol), dolutegravir (-8.2 kcal/mol) and cabotegravir (-8.2 kcal/mol) (Table 1). 19 compounds were identified as various anticancer drugs (Table 1). Screening results of SWISS-SIMILARITY revealed no antiviral drugs and one antitumor agent (pexidartinib, -8.5 kcal/mol) (Table 1) (Wishart et al. 2018).
Discussion
Currently there are no monkeypox specific vaccines and drugs available and smallpox treatment is prescribed to counteract monkeypox disease (Breman et al. 1980; CDC 2022; Mpox (Monkeypox), 2023). Tecovirimat is one of the drugs prescribed to treat smallpox incidence, which demonstrated decent effectiveness (Duraffour et al. 2010; Desai et al. 2022). It was discovered through high-throughput screening in 2002 and was shown to be effective against all orthopoxviruses (Duraffour et al. 2010; Merchlinsky et al. 2019; Desai et al. 2022). In 2018, the Food and Drug Administration (FDA) approved the drug for smallpox treatment (Hoy 2018). Tecovirimat inhibits the formation of p37, a major envelope protein, required to produce virions (Duraffour et al. 2010). Thus, it limits the virus from exiting infected cells preventing spread of the infection (Duraffour et al. 2010; Desai et al. 2022).
In this study we attempted to perform virtual screening based on the tecovirimat mode of binding to p37 envelope protein utilizing both LBVS and SBVS methods. To our knowledge this is the first drug repositioning study dedicated to identify monkeypox drug candidates using both virtual screening methods. We, first, prepared tertiary structure of p37 protein using I-TASSER server and analyzed it using PROCHECK (Ramachandran plot) and PROSA-Web server. Overall, the model demonstrated good ratio of residues in allowed and disallowed regions with strong Z-score (X-ray, NMR) in the range normally found in native proteins of the similar size, indicating a suitability of the predicted tertiary structure for further analysis. The ligand-binding pocket was also predicted by I-TASSER server and included a total of eight residues.
p37–tecovirimat docking was performed to create a reference result with the strongest binding energy of -8.2 kcal/mol. We considered this value as the lowest threshold for the screening drugs to be considered as candidates for monkeypox treatment therapy. SWISS-SIMILARITY server was used to perform LBVS and PHARMIT was utilized for SBVS. The important aspect of any drug discovery process is to assume its safety to the user. After primary screening (LBVS and SBVS) we selected only phase 4 drugs (approved for use in the general population). Since monkeypox is a viral disease, we were primarily interested in antiviral drugs. We also included anti-tumor drugs in our list since previous repurposing studies showed successful repositioning of the said compounds to treat or alleviate symptoms of viral diseases (Ciliberto et al. 2020; Aldea et al. 2021). LBVS produced only a single antitumor drug, while SBVS identified six antiviral agents and 19 antitumor compounds (Table 1). LBVS and SBVS produced significantly different results due to the totally opposite methods utilized in drug screening. Since LBVS was focused on the ligand structure, it identified compounds that closely resembled tecovirimat, whereas SBVS which is based on the binding pocket of the receptor, determined drugs that would most likely fit in it. Thus, the drugs produced by LBVS and SBVS and selected for the final docking did not overlap as we originally theorized. However, due to this disagreement, we were able to obtain higher variety in the compounds to analyze and compare.
The final candidates were selected based on their interaction strength with p37, mimicking possible real receptor-ligand interactions. Our results indicate the possibility of identified candidate drugs as treatments against monkeypox. We believe that results produced in this study will bring us one step closer to identifying a more effective drug in treating and alleviating symptoms of monkeypox. Nevertheless, one should consider that our results are a product of computer simulations and to produce actual viable data, in vitro/in vivo confirmations are required to confirm the effectiveness of the drugs, which will be performed in future studies.
Conclusions
Developing novel drugs against emerging diseases such as monkeypox takes significant amount of time and resources. Alternatively, drug repurposing provides a relatively quick and efficient approach to identify drug candidates from existing drug pool. Thus, this study was conducted to identify those candidates using SBVS and LBVS methods together with 3D protein structure modelling and virtual docking. The results revealed 6 antiviral and 20 anti-tumor drugs with binding affinity stronger than that of tecovirimat, indicating that the selected compounds have high chances to be effective against the disease.
Data availability
The datasets used in this study are available in the ChEMBL database (https://www.ebi.ac.uk/chembl/) and all accession numbers are provided in the article. The additional information about this work is available from the corresponding author upon a reasonable request.
References
Aldea M et al (2021) ‘Repurposing of Anticancer Drugs Expands Possibilities for Antiviral and Anti-Inflammatory Discovery in COVID-19’, Cancer Discovery, 11(6), pp. 1336–1344. Available at: https://doi.org/10.1158/2159-8290.CD-21-0144
Allen WJ et al (2015) ‘DOCK 6: Impact of new features and current docking performance’, Journal of Computational Chemistry, 36(15), pp. 1132–1156. Available at: https://doi.org/10.1002/jcc.23905
Ashburn TT, Thor KB (2004) ‘Drug repositioning: identifying and developing new uses for existing drugs’, Nature Reviews Drug Discovery, 3(8), pp. 673–683. Available at: https://doi.org/10.1038/nrd1468
Berman M, H. et al (2000) ‘The Protein Data Bank’ 28(1):235–242
Berman H, Henrick K, Nakamura H (2003) ‘Announcing the worldwide Protein Data Bank’, Nature Structural & Molecular Biology, 10(12), pp. 980–980. Available at: https://doi.org/10.1038/nsb1203-980
Blasco R, Moss B (1991) ‘Extracellular vaccinia virus formation and cell-to-cell virus transmission are prevented by deletion of the gene encoding the 37,000-Dalton outer envelope protein’, 65(11). Available at: https://doi.org/10.1128/jvi.65.11.5910-5920.1991
Bragina ME et al (2022) ‘The SwissSimilarity 2021 Web Tool: Novel Chemical Libraries and Additional Methods for an Enhanced Ligand-Based Virtual Screening Experience’, International Journal of Molecular Sciences, 23(2), p. 811. Available at: https://doi.org/10.3390/ijms23020811
Breman JG et al (1980) Human monkeypox, 1970-79. Bull World Health Organ 58(2):165–182
CDC (2022) Monkeypox, Centers for Disease Control and Prevention. Available at: https://www.cdc.gov/poxvirus/monkeypox/ (Accessed: 15 September 2022)
Chen Y et al (2009) ‘Vaccinia virus p37 interacts with host proteins associated with LE-derived transport vesicle biogenesis’, Virology Journal, 6, p. 44. Available at: https://doi.org/10.1186/1743-422X-6-44
Ciliberto G, Mancini R, Paggi MG (2020) ‘Drug repurposing against COVID-19: focus on anticancer agents’, Journal of Experimental & Clinical Cancer Research, 39(1), p. 86. Available at: https://doi.org/10.1186/s13046-020-01590-2
Davies M et al (2015) ‘ChEMBL web services: streamlining access to drug discovery data and utilities’, Nucleic Acids Research, 43(Web Server issue), pp. W612–W620. Available at: https://doi.org/10.1093/nar/gkv352
Desai AN et al (2022) ‘Compassionate Use of Tecovirimat for the Treatment of Monkeypox Infection’, JAMA [Preprint]. Available at: https://doi.org/10.1001/jama.2022.15336
Duraffour S, Andrei G, Snoeck R (2010) Tecovirimat, a p37 envelope protein inhibitor for the treatment of smallpox infection. IDrugs 13(3):181–191
Gaulton A et al (2017) ‘The ChEMBL database in 2017’, Nucleic Acids Research, 45(D1), pp. D945–D954. Available at: https://doi.org/10.1093/nar/gkw1074
Hamza A, Wei N-N, Zhan C-G (2012) ‘Ligand-Based Virtual Screening Approach Using a New Scoring Function’, Journal of chemical information and modeling, 52(4), pp. 963–974. Available at: https://doi.org/10.1021/ci200617d
Hoy SM (2018) ‘Tecovirimat: First Global Approval’, Drugs, 78(13), pp. 1377–1382. Available at: https://doi.org/10.1007/s40265-018-0967-6
Hurle MR et al (2013) ‘Computational Drug Repositioning: From Data to Therapeutics’, Clinical Pharmacology & Therapeutics, 93(4), pp. 335–341. Available at: https://doi.org/10.1038/clpt.2013.1
Laskowski RA, Swindells MB (2011) ‘LigPlot+: multiple ligand-protein interaction diagrams for drug discovery’, Journal of Chemical Information and Modeling, 51(10), pp. 2778–2786. Available at: https://doi.org/10.1021/ci200227u
Laskowski RA et al (1993) ‘PROCHECK: a program to check the stereochemical quality of protein structures’, Journal of Applied Crystallography, 26(2), pp. 283–291. Available at: https://doi.org/10.1107/S0021889892009944
Laskowski RA et al (1996) ‘AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR’, Journal of biomolecular NMR, 8(4), pp. 477–486. Available at: https://doi.org/10.1007/BF00228148
Li Q, Shah S (2017) ‘Structure-Based Virtual Screening’, in C.H. Wu, C.N. Arighi, and K.E. Ross (eds) Protein Bioinformatics: From Protein Modifications and Networks to Proteomics. New York, NY: Springer (Methods in Molecular Biology), pp. 111–124. Available at: https://doi.org/10.1007/978-1-4939-6783-4_5
Li J et al (2016) ‘A survey of current trends in computational drug repositioning’, Briefings in Bioinformatics, 17(1), pp. 2–12. Available at: https://doi.org/10.1093/bib/bbv020
Merchlinsky M et al (2019) ‘The development and approval of tecoviromat (TPOXX®), the first antiviral against smallpox’, Antiviral research, 168, pp. 168–174. Available at: https://doi.org/10.1016/j.antiviral.2019.06.005
Mpox (2023) (Monkeypox) WHO. Available at: https://www.who.int/news-room/fact-sheets/detail/monkeypox (Accessed: 15 September 2022)
O’Boyle NM et al (2011) ‘Open Babel: An open chemical toolbox’, Journal of Cheminformatics, 3(1), p. 33. Available at: https://doi.org/10.1186/1758-2946-3-33
Pettersen EF et al (2004) ‘UCSF Chimera–a visualization system for exploratory research and analysis’, Journal of Computational Chemistry, 25(13), pp. 1605–1612. Available at: https://doi.org/10.1002/jcc.20084
Roy A, Kucukural A, Zhang Y (2010) ‘I-TASSER: a unified platform for automated protein structure and function prediction’, Nature Protocols, 5(4), pp. 725–738. Available at: https://doi.org/10.1038/nprot.2010.5
Sayers EW et al (2022) ‘Database resources of the national center for biotechnology information’, Nucleic Acids Research, 50(D1), pp. D20–D26. Available at: https://doi.org/10.1093/nar/gkab1112
Sunseri J, Koes DR (2016) ‘Pharmit: interactive exploration of chemical space’, Nucleic Acids Research, 44(W1), pp. W442–W448. Available at: https://doi.org/10.1093/nar/gkw287
Trott O, Olson AJ (2009) ‘AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading - Trott – 2010 - Journal of Computational Chemistry - Wiley Online Library’, 31, pp. 455–461. Available at: https://doi.org/10.1002/jcc.21334
Wallace AC, Laskowski RA, Thornton JM (1995) ‘LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions’, Protein Engineering, 8(2), pp. 127–134. Available at: https://doi.org/10.1093/protein/8.2.127
Wiederstein M, Sippl MJ (2007) ‘ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins’, Nucleic Acids Research, 35(Web Server), pp. W407–W410. Available at: https://doi.org/10.1093/nar/gkm290
Wishart DS et al (2018) ‘DrugBank 5.0: a major update to the DrugBank database for 2018’, Nucleic Acids Research, 46(D1), pp. D1074–D1082. Available at: https://doi.org/10.1093/nar/gkx1037
Yang G et al (2005) ‘An Orally Bioavailable Antipoxvirus Compound (ST-246) Inhibits Extracellular Virus Formation and Protects Mice from Lethal Orthopoxvirus Challenge’, Journal of Virology, 79(20), pp. 13139–13149. Available at: https://doi.org/10.1128/JVI.79.20.13139-13149.2005
Zhang Y (2008) ‘I-TASSER server for protein 3D structure prediction’, BMC Bioinformatics, 9(1), p. 40. Available at: https://doi.org/10.1186/1471-2105-9-40
Zoete V et al (2016) ‘SwissSimilarity: A Web Tool for Low to Ultra High Throughput Ligand-Based Virtual Screening’, Journal of Chemical Information and Modeling, 56(8), pp. 1399–1404. Available at: https://doi.org/10.1021/acs.jcim.6b00174
Acknowledgements
Not applicable.
Funding
This work was supported by the BK21 FOUR Program of the Department of Agricultural Biotechnology, Seoul National University, Seoul, Korea.
Author information
Authors and Affiliations
Contributions
V.L., Y.L., C.L. and H.K. conceived this study. V.L., C.L. and H.K. designed experimental portion; V.L., Y.L and C.L. conducted the experimental part under supervision of H.K. V.L. and Y.L. collected the data for analysis and performed in silico experiments. V.L., Y.L., C.L. and H.K. wrote the manuscript and edited it.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, V., Lee, Y., Lee, C. et al. Repurposing existing drugs for monkeypox: applications of virtual screening methods. Genes Genom 45, 1347–1355 (2023). https://doi.org/10.1007/s13258-023-01449-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-023-01449-8