Abstract
A Thinopyrum intermedium × Thinopyrum ponticum synthetic hybrid wheatgrass is an excellent source of leaf and stem rust resistance produced by N.V.Tsitsin. Wheat line Mv9kr1 was crossed with this hybrid (Agropyron glael) in Hungary in order to transfer its advantageous agronomic traits into wheat. As the wheat parent was susceptible to leaf rust, the transfer of resistance was easily recognizable in the progenies. Three different partial amphiploid lines with leaf rust resistance were selected from the wheat/Thinopyrum hybrid derivatives by multicolour genomic in situ hybridization. Chromosome counting on the partial amphiploids revealed 58 chromosomes (18 wheatgrass) in line 194, 56 (14 wheatgrass) in line 195 and 54 (12 wheatgrass) in line 196. The wheat chromosomes present in these lines were identified and the wheatgrass chromosomes were characterized by fluorescence in situ hybridization using the repetitive DNA probes Afa-family, pSc119.2 and pTa71. The 3D wheat chromosome was missing from the lines. Molecular marker analysis showed the presence of the Lr24 leaf rust resistance gene in lines 195 and 196. The morphological traits were evaluated in the field during two consecutive seasons in two different locations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The perennial wheatgrasses possess several favourable features for wheat improvement, such as tolerance to biotic and abiotic stresses, leading to better crop safety, yield and quality. Intermediate wheatgrass [Thinopyrum intermedium (Host) Barkworth & D.R. Dewey] and tall wheatgrass [Thinopyrum ponticum (Podp.) Z.-W. Liu & R.-C. Wang] are the two most common introduced species. Because of the sterility of wheat × Thinopyrum F1 hybrids, complete amphiploids or more frequently partial amphiploids are the starting material for successful gene transfer (Jiang et al. 1994). Colchicine treatment on the F1 hybrids leads to the formation of chromosome-doubled amphiploid plants. Partial amphiploids can be selected among the progenies of backcrossed F1 hybrids. The high number of homoeologous chromosomes causes genetic instability in the amphiploids. As a result of substitutions and deletions, partial amphiploid plants carry a stabilized genome. In the case of bread wheat/polyploid Thinopyrum partial amphiploids, genetically stable lines with 56 chromosomes (8×) were reported (Banks et al. 1993; Fedak et al. 2000; Han et al. 2004; Oliver et al. 2006; Sepsi et al. 2008; Bao et al. 2009; Chang et al. 2010; Georgieva et al. 2011; and Zheng et al. 2014), while in durum wheat/polyploid Thinopyrum partial amphiploids 42 chromosomes (6×) were observed (Zeng et al. 2013).
Intermediate and tall wheatgrasses are not only important forage crops but also valuable gene reservoirs for wheat (Triticum aestivum L.) improvement. Almost half of leaf rust resistance genes, 30 % of stem rust resistance genes and 10 % of yellow rust resistance genes have been introduced into bread wheat from closely related and/or wild species (Salina et al. 2015). A significant proportion of them were derived from polyploid Thinopyrum species (Wang 2011). Chromosomal segments of Thinopyrum ponticum (2n = 10× = 70) carrying the leaf rust resistance genes Lr19 (Friebe et al. 1994), Lr24 (McIntosh et al. 1977) and Lr29 (Procunier et al. 1995) and the stem rust resistance genes Sr24 (Sears 1973), Sr25 (McIntosh et al. 1977), Sr26 (Friebe et al. 1994) and Sr43 (Kim et al. 1993) were transferred into wheat. Lr24 is completely linked with Sr24 while Sr25 often shows complete linkage to Lr19. Thinopyrum intermedium was used as a source of the Lr38 (Friebe et al. 1992), Sr44 (Friebe et al. 1996), Bdv2 (Banks et al. 1995), Bdv3 (Sharma et al. 1995), Bdv4 (Lin et al. 2006), Yr50 (Liu et al. 2013) and Wsm1 (Liang et al. 1979) resistance genes via wheat-alien introgressions. These translocations can result from either spontaneous or induced (Friebe et al. 1996) recombination.
Generic relationships within the Triticeae are problematic (Kellogg 2006). Tall wheatgrass was previously classified as Agropyron elongatum and intermediate wheatgrass as Agropyron glaucum in the genus Agropyron (Hitchcock 1951). Dewey (1984) reduced the Agropyron genus based on the presence of the P genome. Tall and intermediate wheatgrass were relocated to the Thinopyrum genus as Thinopyrum ponticum (Podp.) Z.-W. Liu & R.-C. Wang and Th. intermedium (Host) Barkworth & D.R. Dewey, respectively. Polyploid Thinopyrum species contain genomes similar to the J (Jb, Eb) genome of the diploid Th. bessarabicum (Östergren 1940) or the E (Je, Ee) genome of Th. elongatum (Cauderon and Saigne 1961), which are closely related (Ceoloni et al. 2014), and sometimes a third genome from Pseudoroegneria designated as St, previously designated as S (Wang et al. 1995).
The first successful crosses between wheat and wheatgrasses were made in 1930 by NV Tsitsin (Armstrong 1936). Wheat × wheatgrass hybrids were produced to breed wheat with perennial growth habit in the former Soviet Union (Verushkine and Shechurdine 1933). A synthetic hybrid was produced by crossing Th. intermedium (former name Agropyron glaucum) with Th. ponticum (former name Agropyron elongatum) in the 1950s by NV Tsitsin, the aim being to analyse the genome composition of the Agropyron species used in wheat/wheatgrass crosses (Tsitsin 1979). The hybrid plants were named as Agropyron glael by Tsitsin, as an abbreviation of glaucum and elongatum. This name (A. glael) will be used hereafter in this article. The hybrid plants had 56 chromosomes. A number of A. glael plants were maintained in Martonvásár (Hungary) thanks to cooperation between the Hungarian Academy of Sciences and the Moscow Research Institute of Agriculture “Nemchinovka” in the 1960s.
The aim of this study was to describe the chromosome composition of three newly selected wheat/A. glael partial amphiploids by means of multicolour genomic in situ hybridization (mcGISH) and fluorescence in situ hybridization (FISH). A further aim was to characterize artificial powdery mildew inoculation and spontaneous leaf rust and yellow rust infection together with the molecular marker analysis of some Thinopyrum-derived Lr genes present in the lines. The morphological parameters of the partial amphiploid lines were also described.
Materials and methods
Plant material
The A. glael perennial wheatgrass clone was kindly provided by GD Lapchenko from the Moscow Research Institute of Agriculture ’Nemchinovka’. The clone has been maintained in the perennial nursery in Martonvásár since the 1960s by the Hungarian breeder Dezső Szalay. Wheat genotype Mv9kr1, containing both the recessive crossability alleles (kr1kr1kr2kr2) (Molnár-Láng et al. 1996), was crossed with A. glael in 2001. Young inflorescences of F1 plants were used for callus induction and were multiplied in tissue culture as described by Molnár-Láng et al. (1991). Regenerated plants were grown in the phytotron under the conditions described by Tischner et al. (1997). Chinese Spring wheat was the pollinator during backcrossing. The BC1F5-BC1F8 lines, were analysed cytogenetically.
Sequential mcGISH and FISH
Chromosome preparation was carried out as described by Lukaszewski et al. (2004). McGISH was performed in order to simultaneously visualize the different Thinopyrum chromosomes in the BC1 self-pollinated progenies. J (Eb) genomic DNA from Th. bessarabicum labelled with biotin-11-dUTP (Roche Diagnostics, Mannheim, Germany) and St genomic DNA from Ps. spicata labelled with digoxigenin-11-dUTP was produced using the random primed labelling protocol. The hybridization mixture contained 100 ng each of the labelled probes/slide, dissolved in a 15 μl mixture of 100 % formamide, 20 × SSC and 10 % dextran-sulphate at a ratio of 5:1:4, and 3000 ng Triticum aestivum DNA (BBAADD) as a block. Hybridization was performed at 42 °C overnight. Streptavidin-FITC (Roche) and Anti-Digoxigenin-Rhodamine (Roche) dissolved in TNB (Tris-NaCl-blocking buffer) were used in the detection phase. After rinsing off the mcGISH signals, three-colour FISH was performed using three repetitive DNA probes: Afa-family, pSc119.2 and pTa71. Hybridization and detection were carried out as reported by Kruppa et al. (2013). The slides were screened using a Zeiss Axioskop-2 fluorescence microscope equipped with filter sets appropriate for DAPI (Zeiss Filterset 01), and for the simultaneous detection of FITC and Rhodamine (Zeiss filter set 24). Images were captured with a Spot CCD camera (Diagnostic Instruments) and processed with Image Pro Plus software (Media Cybernetics).
Molecular marker analysis
Four primer pairs were used for the detection of the absence or presence of certain Thinopyrum-derived leaf rust and stem rust resistance genes in the partial amphiploid lines. Genomic DNA was extracted from fresh young leaves of wheat cultivars Chinese Spring, Mv9kr1, the wheatgrass species Th. intermedium, Th. ponticum, the synthetic hybrid A. glael, the positive control wheat lines SO91-1027 (Lr19), TC24 (Thatcher*6/Agent, Lr24), TC29 (Thatcher*6//CS7D/Ag#11, Lr29) and Sunelg (Sr26) and the three partial amphiploid lines (lines 194, 195, 196) with a DNeasy Plant Kit (Qiagen, Germany). The STS marker STSLr19 130 with the primer pair GbF-GbR (Lr19, Prins et al. 2001), STS marker J09-STS with the primer pair J09/1-J09/2 (Lr24, Schachermayr et al. 1995) and a SCAR marker with Lr29F18-Lr29R18 primers (Lr29, Procunier, http://maswheat.ucdavis.edu/protocols/Lr29/), were used to reveal the presence of the Lr19, Lr24 and Lr29 leaf rust resistance genes (derived from Thinpyrum sp.) in the partial amphiploid lines. Multiplex PCR with markers Sr26#43 (a dominant STS marker for the presence of Sr26) and BE518379 (6AL-specific, dominant for the absence of Sr26) (Liu et al. 2010) were used to characterize the presence of Sr26. PCR reactions were performed in an Applied Biosystem 9700 PCR (Life Technologies, California, USA) in a final volume of 20 μl containing 200 ng DNA template, 5× Green Go Taq Flexi Buffer (Promega), 2.34 mM MgCl2, 0.9 μM of each dNTP, 10 pmol forward and reverse primers and 1 U GoTaq DNA Polymerase (5 U/μ, Promega). The PCR products were separated using SeaKem 1.5 % agarose gels (Lonza, Rockland, ME, USA) and the fragments were stained using ethidium bromide. A 100-bp DNA ladder (GelPilot 100 bp Plus Ladder, Qiagen, Germany) was used to estimate molecular weight. The patterns were documented and analysed using a Syngene G-BOX documentation system (Syngene, Maryland, USA).
Phenotypic evaluation of the plants
The partial amphiploid lines and the parental wheat genotype (Mv9kr1) were grown in the pesticide-free Tükrös nursery in Martonvásár in two consecutive seasons (2013–2014 and 2014–2015) with 10 seeds in each 1 m row and a row distance of 15 cm. The same genotypes were sown in the breeder’s nursery in Lászlópuszta in the 2014–2015 season in plots of 2 m2. Ten plants were randomly selected from each genotype for analysis. Plant height and tillering (spikes per plant) were measured in the field immediately before harvest. The traits fertility (seeds per spikelet), length of the main spike, number of spikelets per main spike and number of seeds per main spike were measured after harvest. Differences in morphological characteristics between the partial amphiploid line and the control Mv9kr1 genotype were determined by means of the MS Excel Student’s t-test for paired data at the P = 0.05 significance level.
Artificial powdery mildew inoculation and spontaneous leaf rust and stripe rust infection
Powdery mildew resistance was tested under greenhouse conditions. Blumeria graminis f.sp. tritici isolate P07-14 (virulent on differentials with genes Pm1, Pm2, Pm3a, Pm3d, Pm4a, Pm4b, Pm5, Pm6, Pm7, Pm8 or Pm17 or the gene combinations Pm1,2,9, Pm2,4b,8, Pm2,6, Pm2,Mld; avirulent on: Pm3b, Pm3c, Pm3f) was used for inoculation. Ten plants of each genotype (2 partial amphiploids + 2 parents + Carsten V susceptible check) were grown in three randomized replications under an isolator (18 °C, relative air humidity of 80–90 %). The inoculum was shaken on to the leaf surface 9–10 days after sowing. The type of infection was determined ten days after inoculation using the method recommended by Nover (1957). Resistant genotypes gave a score of 0–2, while those with scores of 3–4 were susceptible.
Each year several rows of the leaf rust (Puccinia triticina) -susceptible wheat cultivar Mv9kr1 were planted in the nursery adjacent to the plots of Mv9kr1/A. glael BC1 selfed progenies. Leaf rust and yellow rust (Puccinia striiformis f.sp. tritici) resistance were described using observations on spontaneous infection in the last three years.
Results
Crosses
The hybridization of Mv9kr1 wheat and A. glael resulted in 255 F1 grains. The first successful backcrossing with the wheat genotype Chinese Spring resulted in five BC1 grains in 2004, but only two of them were viable. The first BC1 plant (line 0566) carried 49 chromosomes and was backcrossed with Mv9kr1, but none of the 11 BC2 grains originating from 0566 were viable. The other BC1 plant (No.0567, 62 chromosomes) had four spikes, three of which were self-pollinated resulting in 46 BC1F2 grains, while the fourth was backcrossed with Mv9kr1, resulting in 19 BC2 seeds. Derivatives of these plants have been maintained, self-pollinated and selected for leaf rust resistance in the Tükrös nursery since 2006. Plants of the leaf rust-resistant BC1F5-BC1F8 lines were analysed cytogenetically and grown in the phytotron.
Molecular cytogenetic analysis
Partial amphiploid line 194: 58 chromosomes
The chromosome number and genome composition of the wheat–A. glael partial amphiploids were analysed in somatic metaphase spreads from 5–20 individual plants by sequential mcGISH and FISH.
McGISH allowed nine pairs of A. glael chromosomes to be discriminated (Fig. 1a). Biotin-labelled J genomic DNA hybridized to the entire length of four pairs of submetacentric chromosomes (Ag1-Ag4). Ag5 exhibited a special hybridization pattern: St genomic DNA hybridized to the centromeric and pericentromeric region, while J genomic DNA hybridized to the other parts of the chromosome with the exception of the telomeric region, which remained unlabelled. This chromosome could be identified as JS. The remaining four pairs of chromosomes were labelled by St genomic DNA (Ag6-Ag9) but with faint intensity in the case of Ag8. Chromosomes belonging to the St genome differed greatly in chromosome length and fluorescence intensity. The smallest St chromosome was acro- or telocentric (Ag9), while the others were nearly metacentric. Among the 18 fluorescing chromosomes, two pairs carried a terminal unlabelled region, suggesting that intergenomic rearrangement had taken place. St genomic DNA gave a strong hybridization signal on the satellite region of the wheat chromosomes. J genomic DNA hybridized, though with lower intensity, to six wheat chromosomes, while others were unlabelled.
a Multicolour genomic in situ hybridization (mcGISH) on mitotic chromosomes of the partial amphiploid lines 194 derived from the Mv9kr1 (wheat) × Thinopyrum synthetic hybrid (Agropyron glael, hybrid of Thinopyrum intermedium and Thinopyrum ponticum) cross using J (Thinopyrum bessarabicum, green) and St (Pseudoroegneria spicata, red) genomic DNA probes. Wheat chromosomes are unlabelled (brown). Alien chromosomes are indicated with arrowheads. b The fluorescent in situ hybridization (FISH) pattern on the same cell of lines 194 using Afa-family (red), pSc119.2 (green) and pTa71 (yellow) repetitive DNA probes. A. glael chromosomes are numbered in yellow, not based on homology, while the wheat chromosomes are numbered in white. Scale bar: 10 μm
Twenty pairs of chromosome were blocked by wheat DNA instead of 21, showing that one pair of wheat chromosomes was substituted by a pair of alien chromosomes. FISH with repetitive DNA probes (Afa-family, pTa71, pSc119.2) was used for the identification of the 40 wheat chromosomes and detected the complete absence of the 3D chromosome (3D nullisomy) (Fig. 1b). When the mcGISH and FISH results were compared, the six wheat chromosomes with J hybridization signals were identified as the D-genome. The FISH probes also hybridized to alien chromosomes. All the Thinopyrum chromosomes had an Afa-family hybridization pattern in the telomeric region and three chromosomes had strong pTa71 signals in this region too. The centromeric and pericentromeric regions remained unlabelled, with only two chromosomes having Afa-family signals. A karyogram was constructed for the wheatgrass chromosomes present in this line and the FISH signals were summarized in an idiogram (Fig. 2a).
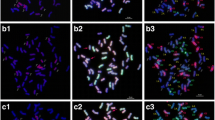
Wheatgrass chromosomes in Mv9kr1 wheat/Thinopyrum synthetic hybrid (Agropyron glael, hybrid of Thinopyrum intermedium and Thinopyrum ponticum) partial amphiploid lines 194 (a), 195 (b) and 196 (c). Multicolour genomic in situ hybridization (mcGISH) karyograms using J (Thinopyrum bessarabicum, green) and St (Pseudoroegneria spicata, red) genomic DNA are presented in the top lanes. Fluorescent in situ hybridization (FISH) karyograms resulting from simultaneous hybridization with Afa-family (red), pSc119.2 (green) and pTa71 (yellow) repetitive DNA probes are presented in the middle lanes. Idiograms of the FISH patterns of the wheatgrass chromosomes are shown in the bottom lanes. Scale bar: 5 μm
According to the mcGISH and FISH results the genome composition of line 194 is 14A+ 14B+ 12D + 8J + 8St + 2JS.
Partial amphiploid line 195: 56 chromosomes
Based on the mcGISH results seven pairs of chromosomes were identified as wheatgrass (Fig. 3a), five pairs of which seem to belong to the J genome as they were mainly green, and two pairs to the St genome, as they fluoresced red, though the hybridization pattern showed some specific features. A very bright red fluorescence signal was observed on the short arm of Ag6, while on the other arm the fluorescence was less intense. As the whole chromosome was red, it was classified as an St chromosome. Ag7 was also identified as an St chromosome, though the fluorescence signal was much fainter than in Ag6. A strong St genomic pattern was observed in the distal part of the short arms of the Ag1, Ag3 and Ag4 chromosomes, while other parts were green, which could be the result of a translocation between the J and St genomes.
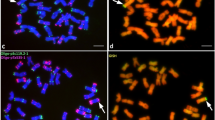
Multicolour genomic in situ hybridization (mcGISH) on mitotic chromosomes of the partial amphiploid lines 195 (a) and 196 (c) derived from the Mv9kr1 (wheat) × Thinopyrum synthetic hybrid (Agropyron glael, hybrid of Thinopyrum intermedium and Thinopyrum ponticum) cross using J (Thinopyrum bessarabicum, green) and St (Pseudoroegneria spicata, red) genomic DNA probes. Wheat chromosomes are unlabelled (brown). Alien chromosomes are indicated with arrowheads. The fluorescent in situ hybridization (FISH) pattern on the same cell of lines 195 (b) and 196 (d) using Afa-family (red), pSc119.2 (green) and pTa71 (yellow) repetitive DNA probes. A. glael chromosomes are numbered in yellow, not based on homology, while the wheat chromosomes are numbered in white. Four 4D chromosomes present in line 195 (b) are marked with blue arrowheads. Translocations between 3BS and an unidentified chromosome arm are marked with blue (d). Scale bar: 10 μm
The wheat chromosomes among which the D chromosomes exhibited slight fluorescence with the J genome probe were characterized using FISH. The chromosome-specific patterns identified two pairs of 4D and no 3D among the 42 wheat chromosomes, so this genotype was identified as a nullitetrasomic line (N3DT4D). During the FISH characterization of the A. glael chromosomes, only the Afa-family probe hybridized to Ag4, Ag5, Ag6 and Ag7, while a strong yellow pTa71 signal on Ag2 and Ag3 marked the NOR region of these chromosomes (Fig. 3b). A faint green pSc119.2 signal was visible in the distal part of the Ag1 short arm. A karyogram was constructed for the wheatgrass chromosomes present in this line and the FISH signals were summarized in an idiogram (Fig. 2b).
The chromosome composition of the progeny of the Mv9kr1/A. glael// Chinese Spring hybrid line 196 is 14A + 14B + 14D (nullitetrasomic line N3DT4D) + 10 J (including J-St translocations) + 4St
Partial amphiploid line 196: 54 chromosomes
McGISH discriminated six pairs of A. glael chromosomes, four pairs of which were hybridized strongly by J genomic DNA (Ag1-Ag4) over their entire length and exhibited great differences in chromosome length (Fig. 3c). The smallest J chromosome (Ag3) was nearly metacentric, while the others were acro- or telocentric. Digoxigenin-labelled St genomic DNA hybridized to the short arm of Ag5, while the long arm remained unlabelled. The last pair (Ag6) showed faint red fluorescence and was identified as St.
Twenty-one pairs of wheat chromosome were unlabelled, though the D chromosomes showed a low level of fluorescence intensity. FISH with repetitive DNA probes (Afa-family, pTa71, pSc119.2) was used for the identification of the 42 wheat chromosomes and showed the complete absence of the 3D chromosome (3D nullisomy) (Fig. 3d). This chromosome was substituted by another, which had 3BS as the longer arm and an unidentifiable small segment as the shorter arm. This small segment was not totally unlabelled by mcGISH, having weak green fluorescence like that observed for D genome-related chromosomes, suggesting the D or J genomic origin of the unknown segment. The FISH probes also hybridized to alien chromosomes. The Ag1, Ag4 and Ag5 chromosomes had Afa-family hybridization patterns in the subtelomeric region and two chromosomes (Ag2 and Ag3) had a strong pTa71 signal in the telomeric region. The centromeric and pericentromeric regions of the alien chromosomes remained unlabelled with the exception of Ag6, which had Afa-family signals. Probe pSc119.2 gave only a weak signal on the telomeric region of Ag1. A karyogram was constructed for the wheatgrass chromosomes present in this line and the FISH signals were summarized in an idiogram (Fig. 2c).
On the basis of the mcGISH and FISH results the genome composition of line 196 is 14A+ 14B + 2 3BS-D/J? translocation + 12D + 8J + 4St.
Molecular marker analysis
The STSLr19 130 marker gave PCR products of the expected 130 bp fragment size in the positive control wheat line S091-1027 and in the wheatgrasses Th. intermedium, Th. ponticum and A. glael. The primer pairs failed to amplify any fragments in the wheat parents Mv9kr1 and Chinese Spring and in the partial amphiploid lines, signalling the absence of Lr19.
The J09-STS marker, which had complete linkage with Lr24, amplified the 310 bp fragment in the positive control wheat line TC24, in the wheatgrasses Th. intermedium, Th. ponticum and A. glael, and in the partial amphiploid lines 195 and 196. Line 194 showed no band intensity (Fig. 4).
Agarose gel electrophoresis patterns of the J09-STS (Lr24) marker. The following DNA templates were used: positive controls (TC24); wheat genotype Mv9kr1, wheat cultivar Chinese Spring (CS), Thinopyrum ponticum, Thinopyrum intermedium, Thinopyrum synthetic hybrid (Agropyron glael, hybrid of Thinopyrum intermedium and Thinopyrum ponticum), Mv9kr1 wheat/Agropyron glael partial amphiploid lines 195 (two samples), 194 and 196. A 100-bp DNA ladder was used to estimate molecular weight
With the help of the Lr29-linked Lr29F18-Lr29R18 primers, PCR products were identified in the TC29 positive control, Th. intermedium and Th. ponticum, while these primers gave no amplification products in A. glael, the wheat parents Mv9kr1 and Chinese Spring or the partial amphiploid lines.
The Sr26#43 marker showed the presence of Sr26 in the positive wheat control line Sunelg, Th. ponticum and A. glael, as PCR products were amplified at the expected 207 bp size. The BE518379 marker showed band intensity at 303 bp size for the absence of Sr26 in the wheat parents Mv9kr1 and Chinese Spring and the partial amphiploid lines.
Phenotypic evaluation of the plants
Phenotypically the partial amphiploids were closer to Triticum aestivum, whereas the adult plants expressed the characteristics of both parents. When the plants were evaluated in the field, the partial amphiploids were found to possess longer spikes (Fig. 5, Table 1) (10.1-13.2 cm) with good fertility (1.7-2.4 seeds/spikelet) and therefore produced no fewer kernels (39-55/spike) than the wheat parent (34-53/spike), except for dwarf line 196, which exhibited significantly lower fertility parameters in all the trials.
The seeds had characteristics intermediate between those of Thinopyrum and T. aestivum, as they were relatively thin and long, with darker brown colour and harder glumes than wheat. The flowering and harvesting times of the partial amphiploids were 10 to 15 days later in the field than for the wheat genotypes in all the years. All the partial amphiploids displayed a vigorous growth habit.
The results obtained for the MS Excel Student’s t-test can be found in Table 1. There were significant differences in morphological characters between the partial amphiploid lines and the control parental genotype Mv9krl. Line 194 had significantly longer spikes with more spikes per plant than the wheat parent in all the experiments, and the plants were significantly taller in both nurseries in 2015. When line 195 was evaluated in the Tükrös breeder’s nursery in 2015, the plant height, length of main spike and number of spikelets per main spike were found to be significantly higher than in wheat, but the fertility (number of seeds per spikelet) was lower. In the case of plant height and the length of the main spike the dwarf line 196 differed significantly from the wheat parental genotype Mv9kr1 in all the trials. The fertility and number of seeds/main spike were significantly lower than in wheat in two of the three experiments.
Reaction to powdery mildew and rusts
Spontaneous leaf rust infection occurred in the pesticide-free Tükrös nursery in the years 2010–2015. During the developmental stage, the wheat–A. glael partial amphiploid lines were highly resistant (type 0) to the leaf rust isolates transmitted from the leaf rust-susceptible spreader rows in the Tükrös prebreeding nursery, while the wheat parents Mv9kr1 (type 4) and Chinese Spring (type 3) were heavily infected by the leaf rust pathogen in all five years (Fig. 6a).
a Symptoms of spontaneous leaf rust infection on the flag-leaves of the susceptible wheat genotypes Mv9kr1 and Chinese Spring (CS) and of the leaf rust-resistant Mv9kr1/Thinopyrum synthetic hybrid (Agropyron glael, hybrid of Thinopyrum intermedium and Thinopyrum ponticum) partial amphiploid lines 194, 195 and 196. b Stripe rust infection on leaf of the susceptible wheat genotypes Mv9kr1 and Chinese Spring and healthy leaves of partial amphiploid lines 194 and 195. Pesticide-free nursery, Martonvásár, Hungary, 2014
Yellow rust infection was observed in 2014 and 2015 in the Tükrös nursery when, the disease occurred spontaneously. The Mv9kr1 and Chinese Spring cultivars were susceptible, while the partial amphiploids showed excellent resistance (Fig. 6b).
The partial amphiploids and their wheat parents were screened using isolates of powdery mildew. The three partial amphiploids and the wheat parents were highly susceptible (type 4) in the seedling stage.
Discussion
In 2001 a crossing programme was begun using the wheat genotpye Mv9kr1 and A. glael (synthetic hybrid of Th. intermedium and Th. ponticum) wheatgrass in order to incorporate the disease resistance of A. glael into wheat (Molnár-Láng et al. 2012). The female wheat parent Mv9kr1 carried the kr1 recessive gene, allowing high crossability in wheat × alien hybridizations (Molnár-Láng et al. 2010). As this wheat genotype is susceptible to leaf rust and yellow rust (Türkösi et al. 2014), the successful transfer of rust resistance from A. glael was easily recognizable in the hybrid progenies. As spontaneous leaf rust disease occurred in the nursery in Martonvásár in 2010–2015 (pesticide-free nursery, weather conditions conducive to fungi) there was no need for artificial inoculation. Partial amphiploid lines were selected from among the BC1 self-pollinated progenies. The aim of this work was to describe the chromosome composition and disease resistance of these unique lines by means of mcGISH and FISH and to compare their phenotypic components with those of the wheat parent Mv9kr1. Wheat/wheatgrass partial amphiploids originating from a cross with A. glael (Th. intermedium × Th. ponticum hybrid) have not previously been reported.
Chromosome counting on the partial amphiploids revealed 58 chromosomes (40 wheat + 18 alien) in line 194, 56 (42 wheat + 14 alien) in line 195 and 54 (42 wheat + 12 alien) in line 196. Other authors observed similar results in the case of wheat/Th. intermedium and wheat/Th. ponticum partial amphiploids. Most of the hexaploid wheat/Thinopyrum sp. partial amphiploids reported contained 56 chromosomes, consisting of 42 wheat and 14 Thinopyrum (Fedak et al. 2000; Han et al. 2004; Oliver et al. 2006; Georgieva et al. 2011; Bao et al. 2014; Zheng et al. 2014), though in some cases fewer wheat (38, 40) and more Thinopyrum chromosomes (16, 18) were observed (Chen et al. 1995; Fedak et al. 2000; Li et al. 2003; Yang et al. 2006; Oliver et al. 2006; Sepsi et al. 2008).
Wheat chromosome 3D was eliminated from the partial amphiploids. These BC1F8 lines were separated from each other in BC1F3, so the elimination of this chromosome probably happened earlier. Among the ABD genomes of hexaploid wheat, the D genome showed the closest homology to the J genome of Thinopyrum (Hsiao et al. 1995; Liu et al. 2007), which was confirmed by the more frequent presence of D-J substititutions and translocations than A-J or B-J (Qi et al. 2007). This close generic relationship could be observed during mcGISH, when J genomic probe DNA hybridized in some cases to D genome-related chromosomes. The hybridization pattern of St genomic DNA also had distinguishing features, as the NOR region of wheat chromosomes 1B and 6B and satellited wheatgrass chromosomes gave fluorescence signals with this probe in all cases. Tang et al. (2000) also described this phenomenon in Th. intermedium.
Decreased fluorescence intensity, J-St translocations in the telomeric region of Js chromosomes, and unlabelled chromosome parts in all types of chromosomes were observed during mcGISH. Chen et al. (2001) reported a high frequency of chromosome pairing between J-JS, J-St and JS-St chromosomes, as the result of which genetic exchange is possible between these genomes. Several minor J-St and JS-St translocations were observed in the partial amphiploids. These translocations may have occurred during the formation of A. glael. As the J-JS-St chromosomes paired at high frequency, it may be that A. glael is not only a hybrid of the two wheatgrass species, that the genetic composition has changed or been enriched with DNA sequences from other species during the long maintenance period (decades), as wheatgrass species are open-pollinating and very polymorphic. As A. glael contains chromosomes from the two most valuable Thinopyrum species, changes in its genome could result in new invaluable genetic material for wheat breeding.
Among the wheatgrass chromosomes in the partial amphiploids, the FISH signals of those belonging to the J genome were compared to the FISH karyotype of the E genome (Je, Ee , Th. elongatum), published by Linc et al. (2012). Ag1 (lines 194, 195 and 196) was very similar to the 3E of Thinopyrum elongatum, while Ag2 (lines 194, 195, 196) was quite similar to 5E. Ag4 (line 194) showed the FISH pattern of 2E. The pTa71 FISH probe, which carries rDNA sequences, gave a strong hybridization signal in Ag2 and Ag3 in lines 194, 195 and 196, whereas Linc et al. (2012) only detected this on 5E. The pSc119.2 probe gave a signal on almost all the E chromosomes of the diploid Th. elongatum (Linc et al. 2012), but in the amphiploid lines only Ag1 (lines 194, 195, 196) and Ag5-Ag6 (line 194) hybridized with this probe. There were thus fewer FISH signals on the J chromosomes of A. glael than on diploid Th. elongatum. During allopolyploidization, rapid genomic events may eliminate non-coding, low-copy DNA sequences from homoeologous chromosomes, while reducing or amplifying high-copy DNA sequences, eliminating rRNA genes or repatterning chromosomes (Feldman and Levy 2005). When the FISH pattern of wheat and its progenitors (T. urartu, Aegilops speltoides, Ae. tauschii) were compared, a reduction in the number of FISH signals was also observed in wheat (Molnár et al. 2014).
Many wheat/Th. intermedium or wheat/Th. ponticum partial amphiploid lines have been reported to carry leaf rust resistance (Li et al. 2003; Han et al. 2004; Sepsi et al. 2008; Chang et al. 2010; Georgieva et al. 2011). The partial amphiploid lines identified in this study had excellent resistance to leaf rust, when observed over several years, but were susceptible to powdery mildew. In addition, the findings suggested that the partial amphiploids might carry different Lr and/or Yr genes, because they contained different types of wheatgrass chromosomes. The Lr24 gene was detected in lines 195 and 196, but line 194 was also resistant to leaf rust. As the Mv9kr1 wheat parent is susceptible to leaf rust, it was concluded that the resistance of the three partial amphiploids originated from A. glael.
Phenotypically the partial amphiploids were similar to T. aestivum, but also expressed the characteristics of the wheatgrass parent and showed good viability. These lines were not just maintained in the nursery, but were used after successful propagation in new crossing programmes with modern, high-yielding wheat varietes in order to decrease the number of wheatgrass chromosomes and to incorporate leaf rust and yellow rust resistance through wheat-A. glael translocations. The selection and identification of resistant progenies is now in progress.
References
Armstrong JM (1936) Hybridization of Triticum and Agropyron, crossing results and description of the first generation hybrids. Can J Res 14:190–202
Banks PM, Xu SJ, Wang RRC, Larkin PJ (1993) Varying chromosome composition of 56 chromosomes wheat-Thinopyrum intermedium partial amphiploids. Genome 36:207–215. doi:10.1139/g93-029
Banks PM, Larkin PJ, Bariana HS, Lagudah ES, Appels R, Waterhouse PM, Brettell RIS, Chen X, Xu HJ, Xin ZY, Qian YT, Zhou XM, Cheng ZM, Zhou GH (1995) The use of cell culture for subchromosomal introgressions of barley yellow dwarf virus resistance from Thinopyrum intermedium to wheat. Genome 38:395–405. doi:10.1139/g95-051
Bao Y, Li X, Liu S, Cui F, Wang H (2009) Molecular cytogenetic characterization of a new wheat-Thinopyrum intermedium partial amphiploid resistant to powdery mildew and stripe rust. Cytogenet Genome Res 126:390–395. doi:10.1159/000266169
Bao Y, Wu X, Zhang C, Li X, He F, Qi X, Wang H (2014) Chromosomal constitutions and reactions to powdery mildew and stripe rust of four novel wheat–Thinopyrum intermedium partial amphiploids. J Genet Genom 41:663–666. doi:10.1016/j.jgg.2014.11.003
Cauderon Y, Saigne B (1961) New interspecific and intergeneric hybrids involving Agropyrum. Wheat Inf Serv 12:13–14
Ceoloni C, Kuzmanovic L, Gennaro A, Forte P, Giorgi D, Grossi MR, Bitti A (2014) Genomes, chromosomes and genes of the wheatgrass genus Thinopyrum: the value of their transfer into wheat for gains in cytogenomic knowledge and sustainable breeding. In: Tuberosa R (ed) Genomics of plant genetic resources, vol 2, Crop productivity, food security and nutritional quality. Springer, Dordrecht, pp 333–358
Chang ZJ, Zhang XJ, Yang ZJ, Zhan HX, Li X, Liu C, Zhang CZ (2010) Characterization of a partial wheat–Thinopyrum intermedium amphiploid and its reaction to fungal diseases of wheat. Hereditas 147:304–312. doi:10.1111/j.1601-5223.2010.02156.x
Chen Q, Conner RL, Laroche A (1995) Identification of the parental chromosomes of the wheat–alien amphiploid Agrotana by genomic in situ hybridization. Genome 38:1163–1169. doi:10.1139/g95-154
Chen Q, Conner RL, Laroche A, Ahmad F (2001) Molecular cytogenetic evidence for a high level of chromosome pairing among different genomes in Triticum aestivum – Thinopyrum intermedium hybrids. Theor Appl Genet 102:847–852. doi:10.1007/s001220000496
Dewey DR (1984) The genomic system of classification as a guide to intergeneric hybridization in the perennial Triticeae. In: Gustafson JP (ed) Gene manipulation in plant improvement. Plenum Press, New York, pp 209–279
Fedak G, Chen Q, Conner RL, Laroche A, Petroski R, Armstrong KW (2000) Characterization of wheat-Thinopyrum partial amphiploids by meiotic analysis and genomic in situ hybridization. Genome 43:712–719. doi:10.1139/gen-43-4-712
Feldman M, Levy AA (2005) Allopolyploidy – a shaping force in the evolution of wheat genomes. Cytogenet Genome Res 109:250–258. doi:10.1159/000082407
Friebe B, Zeller FJ, Mukai Y, Forster BP, Bartos P, Mcintosh RA (1992) Characterization of wheat-Agropyron intermedium derivatives carrying resistance against leaf, stripe and stem rust by C-banding, in situ hybridization and isozyme analysis. Theor Appl Genet 83:775–782. doi:10.1007/BF00226697
Friebe B, Jiang J, Knott DR, Gill BS (1994) Compensation indices of radiation-induced wheat-Agropyron elongatum translocations conferring resistance to leaf rust and stem rust. Crop Sci 34:400–404. doi:10.2135/cropsci1994.0011183X003400020018x
Friebe B, Jiang J, Raupp WJ, Mcintosh RA, Gill BS (1996) Characterization of wheat-alien translocations conferring resistance to diseases and pests: current status. Euphytica 91:59–87. doi:10.1007/BF00035277
Georgieva M, Sepsi A, Tyankova N, Molnár-Láng M (2011) Molecular cytogenetic characterization of two high protein wheat-Thinopyrum intermedium partial amphiploids. J Appl Genet 52:269–277. doi:10.1007/s13353-011-0037-1
Han F, Liu B, Fedak G, Liu Z (2004) Genomic constitution and variation in five partial amphiploids of wheat–Thinopyrum intermedium as revealed by GISH, multicolor GISH and seed storage protein analysis. Theor Appl Genet 109:1070–1076. doi:10.1007/s00122-004-1720-y
Hitchcock AS (1951) Descriptions of genera and species. In: Chase A (ed) Manual of grasses of the United States, 2nd edn. Government Printing Office, Washington, pp 27–980
Hsiao C, Chatterton NJ, Asay KH, Jensen KB (1995) Phylogenetic relationships of the monogenomic species of the wheat tribe, Triticeae (Poaceae), inferred from nuclear rDNA (internal transcribed spacer) sequences. Genome 38:211–223. doi:10.1139/g95-026
Jiang J, Friebe B, Gill BS (1994) Recent advances in alien gene transfer in wheat. Euphytica 73:199–212. doi:10.1007/BF00036700
Kellogg EA (2006) Beyond taxonomy: prospects for understanding morphological diversity in the grasses (Poaceae). Darwiniana 44:7–17
Kim NS, Armstrong K, Knott DR (1993) Molecular detection of Lophopyrum chromatin in wheat-Lophopyrum recombinants and their use in the physical mapping of chromosome 7D. Theor Appl Genet 85:561–567. doi:10.1007/BF00220914
Kruppa K, Türkösi E, Szakács É, Cseh A, Molnár-Láng M (2013) Development and identification of a 4HL.5DL wheat/barley centric fusion using GISH, FISH and SSR markers. Cereal Res Commun 41:221–229. doi:10.1556/CRC.2012.0038
Li HJ, Chen Q, Conner RL, Guo B, Zhang Y, Graf RJ, Laroche A, Jia X, Liu G, Chu C (2003) Molecular characterization of a wheat–Thinopyrum ponticum partial amphiploid and its derivatives for resistance to leaf rust. Genome 46:906–913. doi:10.1139/g03-053
Liang GH, Wang RRC, Niblett CL, Heyne EG (1979) Registration of B-6-37-1 wheat germplasm. Crop Sci 19:421
Lin ZS, Huang DH, Du LP, Ye XG, Xin ZY (2006) Identification of wheat-Thinopyrum intermedium 2Ai-2 ditelosomic addition and substitution lines with resistance to barley yellow dwarf virus. Plant Breed 125:114–119. doi:10.1111/j.1439-0523.2006.01167.x
Linc G, Sepsi A, Molnár-Láng M (2012) A FISH karyotype to study chromosome polymorphisms for the Elytrigia elongata E genome. Cytogenet Genome Res 136:138–144. doi:10.1159/000334835
Liu Z, Li DY, Zhang XY (2007) Genetic relationships among five basic genomes St, E, A, B and D in Triticeae revealed by genomic southern and in situ hybridization. J Integr Plant Biol 49:1080–1086. doi:10.1111/j.1672-9072.2007.00462.x
Liu S, Yu LX, Singh RP, Jin Y, Sorrels ME, Anderson JA (2010) Diagnostic and co-dominant PCR markers for wheat stem rust resistance genes Sr25 and Sr26. Theor Appl Genet 120:691–697. doi:10.1007/s00122-009-1186-z
Liu J, Chang Z, Zhang X, Yang Z, Li X, Jia J, Zhan H, Guo H, Wang J (2013) Putative Thinopyrum intermedium-derived stripe rust resistance gene Yr50 maps on wheat chromosome arm 4BL. Theor Appl Genet 126:265–274. doi:10.1007/s00122-012-1979-3
Lukaszewski AJ, Rybka K, Korzun V, Malyshev SV, Lapinski B, Whitkus R (2004) Genetic and physical mapping of homoeologous recombination points involving wheat chromosome 2B and rye chromosome 2R. Genome 47:36–45. doi:10.1139/g03-089
McIntosh RA, Dyck PL, Green GJ (1977) Inheritance of leaf rust and stem rust resistances in wheat cultivars Agent and Agatha. Aust J Agric Res 28:37–45. doi:10.1071/AR9770037
Molnár I, Kubalakova M, Šimkova H, Farkas A, Cseh A, Megyeri M, Vrána J, Molnár-Láng M, Doležel J (2014) Flow cytometric chromosome sorting from diploid progenitors of bread wheat, T. urartu, Ae. speltoides and Ae. tauschii. Theor Appl Genet 127:1091–1104. doi:10.1007/s00122-014-2282-2
Molnár-Láng M, Galiba G, Kovács G, Sutka J (1991) Changes in the fertility and meiotic behaviour of barley (Hordeum vulgare) × wheat (Triticum aestivum) hybrids regenerated from tissue cultures. Genome 34:261–266. doi:10.1139/g91-041
Molnár-Láng M, Linc G, Sutka J (1996) Transfer of the recessive crossability allele kr1 from Chinese Spring into the winter wheat variety Martonvásári 9. Euphytica 90:301–305. doi:10.1007/BF00027480
Molnár-Láng M, Cseh A, Szakács É, Molnár I (2010) Development of a wheat genotype combining the recessive crossability alleles kr1kr1kr2kr2 and the 1BL.1RS translocation, for the rapid enrichment of 1RS with new allelic variation. Theor Appl Genet 120:1535–1545. doi:10.1007/s00122-010-1274-0
Molnár-Láng M, Linc G, Szakács É, Molnár I, Cseh A, Schneider A, Kruppa K (2012) Wheat-alien introgression programme in Martonvásár. In: III Vavilov International Conference Abstracts, Russian Academy of Sciences, St. Petersburg pp 239–240
Nover I (1957) Sechsjährige Beobachtungen über die physiologische Spezialisierung des echten Mehltaus (Erysiphe graminis DC.) von Weizen und Gerste in Deutschland. Phytopathol Z 31:85–107
Oliver RE, Xu SS, Stack RW, Friesen TL, Jin Y, Cai X (2006) Molecular cytogenetic characterization of four partial wheat–Thinopyrum ponticum amphiploids and their reaction to Fusarium head blight, tan spot, and Stagonospora nodorum blotch. Theor Appl Genet 112:1473–1479. doi:10.1007/s00122-006-0250-1
Östergren G (1940) Cytology of Agropyron junceum, and A. repens and their spontaneous hybrids. Hereditas 26:305–316
Prins R, Groenewald JZ, Marais GF et al (2001) AFLP and STS tagging of Lr19, a gene conferring resistance to leaf rust in wheat. Theor Appl Genet 103:618–624. doi:10.1007/PL00002918
Procunier JD, Townley-Smith TF, Fox S, Prashar S, Gray M, Kim WK, Czarnecki E, Dyck PL (1995) PCR-based RAPD/DGGE markers linked to leaf rust resistance genes Lr29 and Lr25 in wheat (Triticum aestivum L.). J Genet Breed 49:92–97
Qi L, Friebe B, Zhang P, Gill BS (2007) Homoeologous recombination, chromosome engineering and crop improvement. Chromosome Res 15:3–19. doi:10.1007/s10577-006-1108-8
Salina EA, Adonina IG, Badaeva ED, Kroupin PY, Stasyuk AI, Leonova IN, Shishkina AA, Divashuk MG, Starikova EV, Khuat TML, Syukov VV, Karlov GI (2015) A Thinopyrum intermedium chromosome in bread wheat cultivars as a source of genes conferring resistance to fungal diseases. Euphytica 204:91–101. doi:10.1007/s10681-014-1344-5
Schachermayr GM, Messmer MM, Feuillet C, Winzeler H, Winzeler M, Keller B (1995) Identification of molecular markers linked to the Agropyron elongatum-derived leaf rust resistance gene Lr24 in wheat. Theor Appl Genet 90:982–990. doi:10.1007/BF00222911
Sears ER (1973) Agropyron-wheat transfers induced by homoeologous pairing. In: Sears ER, Sears LMS (eds) Proceedings of 4th International Wheat Genetics Symposium. University of Missouri, Columbia, pp 191–199
Sepsi A, Molnár I, Szalay D, Molnár-Láng M (2008) Characterization of a leaf rust-resistant wheat–Thinopyrum ponticum partial amphiploid BE-1, using sequential multicolor GISH and FISH. Theor Appl Genet 116:825–834. doi:10.1007/s00122-008-0716-4
Sharma HC, Ohm H, Goulart L, Lister R, Appels R, Benlhabib O (1995) Introgression and characterization of barley yellow dwarf virus resistance from Thinopyrum intermedium into wheat. Genome 38:406–413. doi:10.1139/g95-052
Tang S, Li Z, Jia X, Larkin PJ (2000) Genomic in situ hybridization (GISH) analyses of Thinopyrum intermedium, its partial amphiploid Zhong 5, and disease-resistant derivatives in wheat. Theor Appl Genet 100:344–352. doi:10.1007/s001220050045
Tischner T, Kőszegi B, Veisz O (1997) Climatic programmes used in the Martonvásár phytotron most frequently in recent years. Acta Agron Hung 45:85–104
Tsitsin NV (1979) Cytogenetic analyses of wheatgrass species and interspecific hybrids (in Russian). In: Tsitsin NV (ed) Difficulties of distant hybridization. Nauka Press, Moscow, pp 48–53
Türkösi E, Farkas A, Aranyi NR, Hoffmann B, Tóth V, Molnár-Láng M (2014) Improvement of the agronomic traits of a wheat–barley centric fusion by introgressing the 3HS.3BL translocation into a modern wheat cultivar. Genome 57:601–607. doi:10.1139/gen-2014-0187
Verushkine S, Shechurdine A (1933) Hybrids between wheat and couch grass. Fertile Triticum-Agropyrum hybrids of great scientific and practical interest. J Hered 24:329–335
Wang RRC (2011) Agropyron and Psathyrostachys. In: Kole C (ed) Wild crop relatives: genomic and breeding resources. Cereals. Springer, Berlin, pp 77–108
Wang RRC, von Bothmer R, Dvorak J, Fedak G, Linde-Laursen I, Muramatsu M (1995) Genome symbols in the Triticeae (Poaceae). In: Wang RRC, Jensen KB, Jaussi C (eds) Proceedings of 2nd International Triticeae Symposium. Utah State University Publications on Design and Production, Logan, pp 29–34
Yang ZJ, Li GR, Chang ZJ, Zhou JP, Ren ZL (2006) Characterization of a partial amphiploid between Triticum aestivum cv. Chinese Spring and Thinopyrum intermedium ssp. trichophorum. Euphytica 149:11–17. doi:10.1007/s10681-005-9010-6
Zeng J, Cao W, Fedak G, Sun S, McCallum B, Fetch T, Xue A, Zhou Y (2013) Molecular cytological characterization of two novel durum–Thinopyrum intermedium partial amphiploids with resistance to leaf rust, stem rust and Fusarium head blight. Hereditas 150:10–16. doi:10.1111/j.1601-5223.2012.02262.x
Zheng Q, Lv Z, Niu Z, Li B, Li H, Xu SS, Han F, Li Z (2014) Molecular cytogenetic characterization and stem rust resistance of five wheat − Thinopyrum ponticum partial amphiploids. J Genet Genomics 41:591–599. doi:10.1016/j.jgg.2014.06.003
Acknowledgments
This work was funded by the Hungarian National Scientific Research Fund (OTKA K 104382 and K 108555). Special thanks to Dezső Szalay, who kindly provided A. glael for the crosses. The authors gratefully acknowledge the excellent technical assistance of F. Tóth, J. Bucsi and I. Könyves-Lakner. Thanks are due to Barbara Hooper for revising the manuscript linguistically.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
This manuscript has no financial or non-financial competing interests.
Additional information
Communicated by: Andrzej Górny
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Kruppa, K., Türkösi, E., Mayer, M. et al. McGISH identification and phenotypic description of leaf rust and yellow rust resistant partial amphiploids originating from a wheat × Thinopyrum synthetic hybrid cross. J Appl Genetics 57, 427–437 (2016). https://doi.org/10.1007/s13353-016-0343-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-016-0343-8