Abstract
Introduction
We aimed to explore the real-world clinical application value and challenges of metagenomic next-generation sequencing (mNGS) for pulmonary infection diagnosis.
Methods
We retrospectively reviewed the results of mNGS and conventional tests from 140 hospitalized patients with suspected pulmonary infections from January 2019 to December 2020. The sample types included bronchoalveolar lavage fluid, lung tissue by transbronchial lung biopsy, pleural effusion, blood, and bronchial sputum. Apart from the mNGS reports that our patients received, an extra comprehensive and thorough literature search was conducted.
Results
Significant differences were noticed in the positive detection rates of pathogens between mNGS and conventional diagnostic testing (115/140, 82.14% vs 50/140, 35.71%, P < 0.05). The percentage of mNGS-positive patients was significantly higher than that of conventional testing-positive patients with regard to bacterial detection (P < 0.01), but no significant differences were found with regard to fungal detection (P = 0.67). Significant statistical differences were found between mixed infection cases (15, 22.70%) and single infection cases (4, 7.84%) in terms of diabetes (P = 0.03). The most frequent pattern of mixed infection was bacteria and fungi mixed infection (40, 40/89 = 44.94%), followed by bacteria mixed infection (29, 29/89 = 32.58%). The sensitivity of mNGS in pulmonary infection diagnosis was much higher than that of conventional test (89.17% vs 50.00%; P < 0.01), but the specificity was the opposite (75.00% vs 81.82%; P > 0.05).
Conclusion
mNGS is a valuable tool for the detection of pulmonary infections, especially mixed pulmonary infections. The most common combinations we found were bacterial–fungal coinfection and bacterial–bacterial coinfection. Still, there are many challenges in the clinical application of mNGS in the diagnosis of pulmonary infections. There is still a lot of work to be done in interpreting the mNGS reports, because both clinical judgment and literature analysis strategy need to be refined.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Why carry out this study? |
Pulmonary infections remain important causes of morbidity and mortality in the world. Improved diagnostic methods with better sensitivity, speed, and spectrum for pathogen detection are urgently needed. |
We aimed to explore the real-world clinical application value and challenges of metagenomic next-generation sequencing (mNGS) for pulmonary infection diagnosis. |
What was learned from the study? |
Significant differences were noticed in the positive detection rates of pathogens between mNGS and conventional diagnostic testing (115/140, 82.14% vs 50/140, 35.71%, P < 0.05) and the sensitivity of mNGS in pulmonary infection diagnosis was much higher than that of conventional test (89.17% vs 50.00%; P < 0.01). |
mNGS is a valuable tool for the detection of pulmonary infections, especially mixed pulmonary infections. There is still a lot of work to be done in interpreting the mNGS reports, because both clinical judgment and literature analysis strategy need to be refined. |
Digital Features
This article is published with digital features, including a summary slide, to facilitate understanding of the article. To view digital features for this article go to https://doi.org/10.6084/m9.figshare.14696046.
Introduction
Pulmonary infections remain important causes of morbidity and mortality in the world [1, 2]. Although various assays exist, rapid and accurate diagnosis of causative pathogens is still difficult. Traditional culture methods, only used for fungal and bacterial detection, are time-consuming and exhibit low rates of positive detection, and therefore cannot meet the clinical needs. Other detection methods like polymerase chain reaction (PCR) and immunology techniques can only identify several specific pathogens. Moreover, infections with multiple pathogens and the appearance of multidrug-resistant pathogens make the identification of the causative agent more difficult. Therefore, improved diagnostic methods with better sensitivity, speed, and spectrum for pathogen detection are urgently needed.
Metagenomic next-generation sequencing (mNGS) is a novel and promising approach which combines high-throughput sequencing with bioinformatics analysis. mNGS requires shorter analysis time, has a wide range of detectable pathogens (all pathogens including bacteria, fungi, viruses, and parasites can be simultaneously identified by a single assay), simple sample-processing, is less affected by prior antibiotic use, and has semiquantitative value in follow-up [3].
Our study summarized the mNGS results for 140 adult patients with suspected pulmonary infection and aimed to evaluate the clinical performance and effect of mNGS in pathogen diagnosis.
Methods
Ethics Statement
The study was conducted in accordance with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. This study was approved by the ethics committee of The Affiliated Wuxi Second People’s Hospital of Nanjing Medical University. Since the data were completely anonymous, informed consent was waived.
Patients and Sample Collection
We retrospectively reviewed 140 hospitalized patients with suspected pulmonary infections in the Department of Respiratory and Critical Care, The Affiliated Wuxi Second People’s Hospital of Nanjing Medical University from January 2019 to December 2020. The attending physician judged whether the patient had suspected pulmonary infection on the basis of clinical manifestations and imaging examination results.
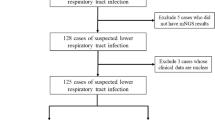
The inclusion criteria were as follows: (1) patients who were suspected of having suspected pulmonary infections; (2) patients who agreed to undergo the mNGS examination; (3) bronchoalveolar lavage fluid (BALF) samples and detection process passed quality control for mNGS; and (4) patients whose medical data were recorded completely.
The exclusion criteria were as follows: (1) patients who refused to undergo the mNGS examination; (2) BALF samples or detection process failed to pass quality control for mNGS; (3) patients with incomplete clinical and laboratory data;
Bronchoscopy was performed by experienced bronchoscopists to obtain specimens including BALF and lung tissue. Transbronchial lung biopsy (TBLB) was performed and BALF was collected by BF-P-260F ultrathin bronchoscope (Olympus, Tokyo, Japan). DirectPath V1.0 virtual bronchoscopic navigation (Olympus, Tokyo, Japan) was used for assistance in navigation.
Body fluid samples, lung biopsy tissues, and plasma samples were sent to BGI Co., Ltd (Shenzhen, China) for sequencing as described in detail below. The remaining specimens were sent to our microbiological laboratory. The conventional tests were performed by culture of the bacteria or fungi in blood agar or in Sabouraud agar at 35 °C for a maximum period of 5 days. Bacteria or fungi culturing were performed by using the VITEK-II Compact automated microbiological system (bioMérieux, France) [4]. Other conventional diagnostic testing included Chlamydia pneumoniae, Mycoplasma pneumoniae, Legionella pneumophila, cytomegalovirus (CMV), Epstein–Barr virus (EBV), adenovirus, and herpes simplex virus (HSV) serological antibody detection. Xpert testing for Mycobacterium tuberculosis and so on.
Metagenomic Next-Generation Sequencing and Analysis
Body Fluid Sample: Sample Processing and DNA Extraction
A body fluid sample (1.5–3 mL of BALF, pleural effusion, or sputum) was collected from each patient according to standard procedures [5]. A 1.5-mL microcentrifuge tube containing 0.6 mL sample, enzyme, and 1 g of 0.5-mm glass beads was agitated by a vortex mixer, and a 0.3-mL sample was separated into a 1.5-mL microcentrifuge tube and DNA was extracted using the TIANamp Micro DNA Kit (DP316, Tiangen Biotech) following the manufacturer’s operational manual. The extracted DNA samples were used for the construction of DNA libraries [6].
Lung Biopsy Tissue: Sample Processing and DNA Extraction
Lung biopsy tissue was minced into small pieces and ground with a tissue homogenate machine. A 10-mg sample was separated into a 1.5-mL microcentrifuge tube and DNA was extracted using the TIANamp Micro DNA Kit (DP316, Tiangen Biotech) following the manufacturer’s operational manual.
Plasma Sample: Sample Processing and DNA Extraction
Blood (3–5 mL) was drawn from patients and centrifuged at 4000 rpm for 10 min at 4 °C. DNA was extracted from 300 µL of plasma using the TIANamp Micro DNA Kit (DP316, Tiangen Biotech, Beijing, China) according to the manufacturer’s operational manual.
Construction of DNA Libraries
DNA libraries were constructed through DNA fragmentation (about 150 bp), end repair, adapter ligation, and PCR amplification. An Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA) was used for quality control of the DNA libraries (200–300 bp). Then, libraries with confirmed quality were sequenced by the BGISEQ-50 platform (BGI Co., Ltd, Shenzhen, China) [7].
Sequencing and Bioinformatic Analysis
High-quality sequencing data were generated by removing low-quality reads, followed by computational subtraction of human host sequences mapped to the human reference genome (hg19) using Burrows–Wheeler alignment [8]. The remaining data by removal of low-complexity reads were classified by simultaneously aligning to four microbial genome databases (bacteria, fungi, viruses, and parasites) which were downloaded from the National Center for Biotechnology Information (NCBI) (ftp://ftp.ncbi.nlm.nih.gov/genomes/). RefSeq contains 4945 whole genome sequence of viral taxa, 6350 bacterial genomes or scaffolds, 1064 fungi related to human infection, and 234 parasites associated with human diseases.
Threshold Criteria for Interpretation of mNGS Result
SMRN: number of sequences strictly mapped at species level.
SMRN of microorganisms in sample is higher than in the negative control group.
For different types of microorganisms, the thresholds were set as follows:
Bacterial/virus/fungus/mycoplasma/chlamydia: SMRN ≥ 3.
Parasite: SMRN ≥ 100.
Mycobacterium tuberculosis: SMRN ≥ 1.
Definite pathogens of pulmonary infection; excluding the normal flora of oral cavity, respiratory tract or the skin (via a large number of literature search).
Meets clinical judgment or targeted treatment response by three experienced senior clinicians.
Microorganisms detected by mNGS which meet all the criteria were identified as suspected pathogens.
Statistical Analyses
Continuous variables are expressed as mean ± standard deviation (SD) and categorical variables are expressed as numbers (percentage). We tested for differences in continuous variables using t test and categorical variables with chi-square test as appropriate. The 2 × 2 contingency tables were established to determine sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV).
The results are presented with 95% confidence intervals (CIs).
Data analyses were performed using SPSS 22.0 software and a two-tailed value of P of 0.05 was considered to represent a significant difference.
Results
Baseline Characteristics of Patients
A total of 140 patients (62 men, 44.28%; 78 women, 55.72%) with suspected pulmonary infection were included in this study (Table 1).
The mean age was 59.16 ± 14.89 years and the majority of patients were above 40 but at most 70 years old (61.43%).
Among them, 75 (75/140 = 53.57%) patients had underlying diseases, including 23 cases of type 2 diabetes (16.43%), 16 (11.43%) cases of connective tissue disease, 14 (10.00%) cases of bronchiectasis, 10 (7.14%) cases of extrapulmonary malignancies, 7 (5.00%) cases of previous history of tuberculosis, 7 (5.00%) cases of previous history of lung cancer, 5 (3.57%) cases of chronic obstructive pulmonary disease, and 1 case of bronchial asthma.
A total of 128 patients underwent bronchoscopy and 9 patients underwent TBLB. The main sample types of mNGS in our patients were BALF (119, 85.00%), while other sample types included lung tissue (9), pleural effusion (6), blood (5), bronchial sputum (1), and so on. The mean time from admission to sampling was 4.3 days.
Concordance Between mNGS and Conventional Diagnostic Testing
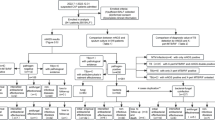
In the present study, mNGS and conventional diagnostic testing (including culture) were both positive for pathogens detection in 48 (48/140 = 34.29%) cases. A total of 67 (67/140 = 48.10%) cases were positive for pathogens detection by mNGS only. There were only 2 cases positive for pathogens detection by conventional test only. mNGS and conventional test were both negative for pathogens detection in 23 (23/140 = 16.43%) cases (Fig. 1).
Concordance between metagenomic next-generation sequencing (mNGS) and conventional testing. Both+, results of mNGS and conventional testing were both positive; Both−, results of mNGS and conventional testing were both negative; mNGS+, only the mNGS result was positive, conventional testing was not; only conventional testing+, only the conventional testing result was positive, mNGS was not
For the 48 both-positive cases, results of mNGS and conventional tests were totally matched in 2 (2/48 = 4.17%) cases and were completely mismatched in 26 (26/48 = 54.17%) cases. A total of 20 (20/48 = 41.67%) cases were found to be partly matched which means at least one detected pathogen was overlapped between conventional diagnostic testing and mNGS (Fig. 1).
Comparison of Pathogenic Detection Between mNGS and Conventional Diagnostic Testing
Significant differences were noticed in the positive detection rates of pathogens between mNGS and conventional diagnostic testing (115/140, 82.14% vs 50/140, 35.71%, P < 0.05).
Among pathogens detected by mNGS (Fig. 2a) and conventional test, Parvimonas micra (n = 8) was the most frequently detected Gram-positive bacteria, followed by Streptococcus pneumoniae (n = 7), Abiotrophia defectiva (n = 7), Enterococcus faecium (n = 6), Streptococcus constellatus (n = 5), and Actinomyces israelii (n = 5).
According to the results of mNGS (Fig. 2b) and conventional test, Haemophilus parainfluenzae (n = 19) was the most frequently detected Gram-negative bacteria, followed by Pseudomonas aeruginosa (n = 13), Acinetobacter baumannii (n = 10), Klebsiella pneumoniae (n = 9), Prevotella melaninogenica (n = 8), and Pyramidobacter piscolens (n = 7).
The most commonly fungi detected by mNGS (Fig. 2c) and conventional test were Candida albicans (n = 23), Pneumocystis jirovecii (n = 4), and Aspergillus fumigatus (n = 3).
The results of mNGS showed that a total of 21 patients were virus-positive (21/140 = 15%) and the most frequently detected viruses were Human gammaherpesvirus 4 (EBV) (n = 16), Human betaherpesvirus 7 (HHV-7) (n = 5), and Human betaherpesvirus 5 (CMV) (n = 4) (Fig. 2d).
In addition, six cases of Mycoplasma, one case of Chlamydia psittaci, and seven cases of M. tuberculosis were also detected by mNGS or conventional test (Fig. 2e).
The percentage of mNGS-positive patients was significantly higher than that of conventional testing-positive patients with regard to bacterial detection (P < 0.01), but no significant differences were found with regard to fungal detection (P = 0.67) (Fig. 3).
The overlap of positivity between mNGS and conventional testing for fungi and bacteria. Both+, results of mNGS and conventional testing were both positive; mNGS+, only the mNGS result was positive, conventional testing was not; conventional testing+, only the conventional testing result was positive, mNGS was not
Streptococcus infantis, Oribacterium parvum, Prevotella pallens, Kingella oralis, Mogibacterium timidum, Treponema maltophilum, Dialister invisus, Streptococcus parasanguinis, Neisseria subflava, Rothia dentocariosa, Campylobacter showae, Actinomyces johnsonii, Johnsonella ignava, Cardiobacterium hominis, Scardovia wiggsiae, Cardiobacterium valvarum, Desulfomicrobium orale, Fretibacterium fastidiosum, Actinomyces gerencseriae, etc. were not interpreted as pathogens, because they were known as normal flora of the oral cavity, respiratory tract, or the skin and were not considered to cause pulmonary infection so far (via literature searches).
Comparison of Clinical Characteristics Between Mixed and Single Pulmonary Infection
Mixed pulmonary infection was defined when two or more infectious pathogens were identified by mNGS or conventional test. As shown in Table 2, there were 66 cases of mixed pulmonary infection and 51 cases of single pulmonary infection.
No difference in age between mixed infection cases (60.15 ± 15.90) and single infection cases (58.45 ± 13.55) was observed (P = 0.54).
There were no significant differences in gender or inflammatory indices between the two groups.
The most common symptoms in both groups were cough, fever, and thoracalgia.
Additionally, there were no significant differences in underlying pulmonary disease, including bronchiectasis, chronic obstructive pulmonary disease (COPD), previous history of tuberculosis, bronchial asthma, and lung cancer, between these two groups.
Significant statistical differences were found between mixed infection cases (15, 22.70%) and single infection cases (4, 7.84%) in terms of diabetes (P = 0.03). However, there were no differences between the two groups in terms of other underlying extrapulmonary diseases like connective tissue disease, and malignant tumor (11, 16.7% vs 5, 9.80%; 4, 6.10% vs 4, 7.84%, respectively).
No difference in time from admission to test between the two groups (3.97 ± 3.00 vs 4.69 ± 3.83) was observed (P = 0.26).
Diagnostic Performance of the Combination of mNGS and Conventional Diagnostic Testing in Mixed Infection
A total of 69 patients (69/140 = 49.29%) were diagnosed with a mixed infection using only the mNGS method. When combined with conventional test results, the diagnostic ratio of the mixed infection was increased to 63.57% (89/140) (Fig. 4).
The most frequent pattern of mixed infection was bacteria and fungi mixed infection (40 patients, 40/89 = 44.94%), followed by bacteria mixed infection (29 patients, 29/89 = 32.58%) (Fig. 4).
Eight patients were diagnosed as bacteria, fungi, viruses, and mycoplasma mixed infection (8/89 = 8.99%); eight patients were diagnosed as bacteria, fungi and viruses mixed infection (8/89 = 8.99%) (Fig. 4).
Comparison of mNGS and Conventional Test in the Diagnosis of Pulmonary Infection
The performance of mNGS and conventional test in diagnosis of pulmonary infection is shown in Table 3.
The sensitivity and specificity of diagnosing pulmonary infection by mNGS were 89.17% (95% CI 83.61–94.73) and 75.00% (95% CI 63.48–86.52), respectively, with NPV and PPV being 95.54% and 53.57%.
As for conventional test, the sensitivity and specificity of diagnosing pulmonary infection were 50.00% (95% CI 40.98–59.02) and 81.82% (95% CI 65.70–97.94), respectively, with NPV and PPV being 93.65% and 23.38%.
The sensitivity of mNGS in pulmonary infection diagnosis was much higher than that of conventional test (89.17% vs 50.00%; P < 0.01), but the specificity was the opposite, with no statistically significant difference (75.00% vs 81.82%; P > 0.05).
Discussion
In our study, mNGS exhibited better performance than conventional test for detecting bacteria (P < 0.01), whereas it was not superior to conventional test with regard to fungal detection (P = 0.67). This was inconsistent with previous findings. According to Miao et al., the sensitivity of mNGS is not better than that of culture for recognizing bacteria, but mNGS has superior feasibility in detecting fungi [9]. Another study found that compared with sequencing, culture is able to identify most bacterium-associated pneumonia [10]. Unlike the aforementioned studies, Xie et al. reported that there was no significant difference between conventional microbiologic test and mNGS with regard to fungal detection; however, mNGS displayed a significantly higher positive rate for detecting pathogenic bacteria [11]. Possible explanations for this divergence are due to different diseases, different sample types, and different test conditions of mNGS and conventional test. Moreover, the positive rate of pathogenic fungi cultivation in our study is relatively higher.
Our results showed that the sensitivity of mNGS in pulmonary infection pathogen detection was much higher than that of the conventional test (89.17% vs 50.00%; P < 0.01), while the specificity of the conventional test was better than mNGS (81.82% vs 75.00%; P > 0.05). This outcome is inconsistent with the study by Cai et al., who found that the sensitivity and specificity of mNGS were both superior to those of culture [12]. However, another study showed that the specificity of mNGS and conventional test are nearly the same (95.2% and 95.2%, respectively; P = 1.0) [13]. Similarly, it has also been reported that the specificity of mNGS was comparable to that of conventional test [9]. On the other hand, other reports have reached similar conclusions to ours [14, 15]. Several explanations might account for these differences.
In general, the overall positive rate of mNGS (115/140, 82.14%) for pathogens in pulmonary infections was significantly higher than that of conventional test (35.71%, P < 0.05). This result is consistent with previous studies. In a multicenter, prospective study, Parize et al. showed that clinically relevant pathogens were identified in a significantly higher proportion of patients with mNGS than conventional microbiological methods (36/101 (36%) vs 11/101 (11%), respectively, P < 0.001) [16]. Langelier et al. reported that mNGS has a greater capacity for detecting microorganisms compared with current microbial diagnostics [17]. These findings, along with our result, confirm that mNGS can raise the detection rate of respiratory pathogens.
In terms of mixed infection diagnosis, the proportion of mixed pulmonary infections in our patients was much higher than we originally thought. We found that a total of 69 (69/140 = 49.29%) cases were positive for mixed infection by mNGS only. However, when combined with conventional test results, the positive ratio of the mixed infection increased to 63.57% (89/140). These results are in agreement with Fang et al.’s findings which showed that, compared with using solely the mNGS method, the diagnostic ratio of the mixed infection was increased from 55.6% to 58.3% (42/72) when combining mNGS and conventional test [4]. Their results and ours both indicate that the combination of mNGS and conventional test contributes to the diagnostic ratio of the mixed pulmonary infection. Prior studies have shown that mNGS was significantly better at the identification of coinfections than conventional tests like serological antibody, PCR, or culture [16, 18, 19].
This study found that the most common combinations were bacterial–fungal coinfection and bacterial–bacterial coinfection. Unlike our result, Wang et al. showed that the most common mixed pulmonary infection was the combination of viruses and fungi [14]. In another study, Legoff et al. observed that bacterial–viral coinfection was the most frequent combinations [20]. In 2020, Wu et al. reported that the most common combinations were viral–bacterial coinfection and bacterial–bacterial coinfection in patients with normal immune function, while bacterial–viral coinfection and bacterial–fungal coinfection were the most frequent combinations in patients with compromised immune function [21]. The inconsistency between our study and other studies might result from the low proportion of immunocompromised patients in this study. Another possible explanation is that our hospital is located in the lower reaches of the Yangtze River, a high humidity area in which fungal infections are common. C. albicans was the major fungal pathogen in this region [22]. Furthermore, we observed triple mixed pulmonary infection (bacteria, fungi, and viruses) and fourfold mixed pulmonary infection (bacteria, fungi, viruses, and mycoplasma) in some patients, the majority of whom were diabetic. Similar results have been reported before [21]. Owing to the advantages mNGS, so many patterns of mixed pulmonary infection can be detected. On the other hand, mixed infections can hardly be identified by traditional culture methods, because various microorganisms interact with, and inhibit, each other. It is known that severe pneumonia is usually not caused by a single pathogen but by coinfections. Thus, mNGS is a more favorable method that may improve clinical guidance in the diagnosis and treatment of patients with pneumonia.
Few studies have discussed the value of mNGS in etiological detection for pulmonary infection in patients with diabetes. In terms of diabetes mellitus, in our study, significant statistical differences were observed between mixed pulmonary infection cases (15, 22.70%) and single infection cases (4, 7.84%) (P = 0.03). This outcome is contrary to that of Wu et al. who found no differences between the patients with and without mixed infections in terms of diabetes [21]. Diabetes is a risk factor for suffering pulmonary infection like community-acquired pneumonia, CAP [23]. People with diabetes are at two times greater risk of developing a pulmonary infection than healthy people [24]. In the present study, mNGS was performed on 24 patients with diabetes and lower respiratory tract infection for pathogen identification. Our data show that patients with diabetes are more susceptible to mixed respiratory tract infections, especially bacterial and fungal infections. Patients with diabetes mellitus were prone to invasive fungal infections [25]. The most common pathogens are A. fumigatus and C. albicans [26, 27]. This is consistent with the results of our current study. In addition, diabetes mellitus increases active pulmonary tuberculosis risk and worsens its outcomes [28]. Two patients with diabetes were diagnosed with active pulmonary tuberculosis by mNGS in this study. These results indicate that for the diabetic population, mNGS are well suited for etiological detection of pulmonary infection.
mNGS is a bias-free assay that can detect all pathogens in the environment; however, this test method itself cannot distinguish pathogenic microorganisms from colonizing microorganisms, background microorganisms, and contaminated microorganisms [29]. For now, no authoritative guide is available to the interpretation of the mNGS report and therefore this remains a great challenge for clinicians [29, 30].
In order to interpret the results of mNGS more accurately and objectively, in our study, we conducted an extra comprehensive and thorough literature search in PubMed in an attempt to identify published cases of pulmonary infection due to pathogens that we detected, without time limits. We found out in mNGS reports that our patients received that some microorganisms which do not cause human pulmonary infections (e.g., Torque teno virus 15, Shigella dysenteriae, Cardiobacterium hominis), some microorganisms which are not pathogenic in humans, or some microorganisms whose pathogenicity is as yet unclear (e.g., Bacteroides vulgatus, Bifidobacterium longum, Parabacteroides merdae, Pyramidobacter piscolens, and Megasphaera micronuciformis) were all included in the pathogenic microorganism list. This problem is relatively easy to solve by reading related literature. The most troublesome issue is opportunistic pathogens. Some microorganisms, for instance, S. constellatus, are part of the normal flora of the oral or respiratory tracts of humans, but pulmonary infections due to them have been widely reported [31, 32]; on the contrary, although reports have shown that some microorganisms, such as A. israelii, can cause respiratory infection [33], the number of articles was quite limited (sometimes only one to two). In fact, the types of pathogens that cause pulmonary infections are constantly evolving. For example, the isolation rate of H. parainfluenzae in patients with pneumonia is increasing year by year; although it remains controversial, this bacterium has been increasingly regarded as an important pathogen of respiratory infections [34]. All of the aforementioned problems pose obvious difficulties and challenges to clinicians when performing a literature search. Therefore, we think that the aforementioned two lists in mNGS reports to some extent can facilitate the interpretation, but it is somewhat arbitrary and can only provide limited reference values to clinicians. An extra comprehensive and elaborate literature search should be conducted.
Moreover, these findings could not be generalized. Various clinical factors including age, symptoms and signs, laboratory tests, imaging performance, immune status, care status, the risk for aspiration pneumonia, ventilator use, tracheal intubation, antibiotic treatment history, treatment response, and so on should all be taken into account [4, 16, 21, 35]. In different patients, even for the same bacteria, we may need to change our judgment.
Certainly, one should also consider the particularity and complexity of mNGS in diagnosing pulmonary infectious. In the context of the misuse and unreasonable administration of antibiotics, numerous opportunistic pulmonary infections have become a tough clinical problem in immunocompromised patients. In cerebrospinal fluid and brain biopsy samples, which are considered sterile, the result of mNGS is easy to interpret [36,37,38]. In contrast, it is much more difficult to evaluate mNGS reports in BALF and sputum [39, 40]. Recently, it has been reported that in patients with severe pneumonia, the utility of mNGS for the simultaneous detection of pathogens from both BALF and blood samples is strongly recommended [41].
The present study has certain limitations. Neither RNA sequencing nor drug resistance testing was performed. Nearly all patients were given empirical antimicrobic treatment before pathogen-detecting test sampling, which might decrease the rate of positive culture. The number of some sample types like blood and bronchial sputum is low. Although a comprehensive and thorough literature search was conducted, how to reasonably interpret the results of the literature search is still worth discussing. Similarly, although clinical judgment was made by three experienced senior clinicians, as a result of the lack of unified standards, subjective bias is unavoidable.
In future, we think that assembling an interdisciplinary team (consisting of medical microbiologists, computational biologists, epidemiologists, infectious disease specialists, and other clinicians) and targeted training are critical to better use of mNGS [28]. With decreasing sequencing costs, drug-resistance gene detection by mNGS may become more feasible.
Conclusions
mNGS exhibited better performance than conventional tests for detecting bacteria, whereas it was not superior to conventional tests with regard to fungal detection. The sensitivity of mNGS in pulmonary infection pathogen detection was much higher than that of conventional tests, while the specificity of conventional tests was better than mNGS. The overall positive rate of mNGS for pathogens in pulmonary infections was significantly higher than that of conventional tests. mNGS is a valuable tool for the detection of mixed pulmonary infections and the most common combinations we found were bacterial–fungal coinfection and bacterial–bacterial coinfection. Significant statistical differences were found between mixed infection cases and single infection cases in terms of diabetes. However, there are still many challenges in the clinical application of mNGS in the diagnosis of pulmonary infections. There is still a lot of work to be done in interpreting the mNGS reports, because both clinical judgment and literature analysis strategy need to be refined.
References
Langelier C, Kalantar KL, Moazed F, et al. Integrating host response and unbiased microbe detection for lower respiratory tract infection diagnosis in critically ill adults. Proc Natl Acad Sci USA. 2018;115(52):E12353–62.
Magill SS, Edwards JR, Bamberg W, et al. Multistate point-prevalence survey of health care-associated infections. N Engl J Med. 2014;370(13):1198–208.
Goldberg B, Sichtig H, Geyer C, Ledeboer N, Weinstock GM. Making the leap from research laboratory to clinic: challenges and opportunities for next-generation sequencing in infectious disease diagnostics. MBio. 2015;6(6):e01888-e1915.
Fang X, Mei Q, Fan X, et al. Diagnostic value of metagenomic next-generation sequencing for the detection of pathogens in bronchoalveolar lavage fluid in ventilator-associated pneumonia patients. Front Microbiol. 2020;11:599756.
Chen X, Ding S, Lei C, et al. Blood and bronchoalveolar lavage fluid metagenomic next-generation sequencing in pneumonia. Can J Infect Dis Med Microbiol. 2020;2020:6839103.
Long Y, Zhang Y, Gong Y, et al. Diagnosis of sepsis with cell-free DNA by next-generation sequencing technology in ICU patients. Arch Med Res. 2016;47(5):365–71.
Jeon YJ, Zhou Y, Li Y, et al. The feasibility study of non-invasive fetal trisomy 18 and 21 detection with semiconductor sequencing platform. PLoS ONE. 2014;9(10):e110240.
Li H, Durbin R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 2009;25(14):1754–60.
Miao Q, Ma Y, Wang Q, et al. Microbiological diagnostic performance of metagenomic next-generation sequencing when applied to clinical practice. Clin Infect Dis. 2018;67(suppl_2):S231-s240.
Toma I, Siegel MO, Keiser J, et al. Single-molecule long-read 16S sequencing to characterize the lung microbiome from mechanically ventilated patients with suspected pneumonia. J Clin Microbiol. 2014;52(11):3913–21.
Xie Y, Du J, Jin W, et al. Next generation sequencing for diagnosis of severe pneumonia: China, 2010–2018. J Infect. 2019;78(2):158–69.
Cai Y, Fang X, Chen Y, et al. Metagenomic next generation sequencing improves diagnosis of prosthetic joint infection by detecting the presence of bacteria in periprosthetic tissues. Int J Infect Dis. 2020;96:573–8.
Huang Z, Li W, Lee GC, et al. Metagenomic next-generation sequencing of synovial fluid demonstrates high accuracy in prosthetic joint infection diagnostics: mNGS for diagnosing PJI. Bone Jt Res. 2020;9(7):440–9.
Wang J, Han Y, Feng J. Metagenomic next-generation sequencing for mixed pulmonary infection diagnosis. BMC Pulm Med. 2019;19(1):252.
Huang J, Jiang E, Yang D, et al. Metagenomic next-generation sequencing versus traditional pathogen detection in the diagnosis of peripheral pulmonary infectious lesions. Infect Drug Resist. 2020;13:567–76.
Parize P, Muth E, Richaud C, et al. Untargeted next-generation sequencing-based first-line diagnosis of infection in immunocompromised adults: a multicentre, blinded, prospective study. Clin Microbiol Infect. 2017;23(8):574 e1-574 e6.
Langelier C, Zinter MS, Kalantar K, et al. Metagenomic sequencing detects respiratory pathogens in hematopoietic cellular transplant patients. Am J Respir Crit Care Med. 2018;197(4):524–8.
Zhang P, Chen Y, Li S, et al. Metagenomic next-generation sequencing for the clinical diagnosis and prognosis of acute respiratory distress syndrome caused by severe pneumonia: a retrospective study. PeerJ. 2020;8:e9623.
Paul L, Comstock J, Edes K, Schlaberg R. Gestational psittacosis resulting in neonatal death identified by next-generation RNA sequencing of postmortem, formalin-fixed lung tissue. Open Forum Infect Dis. 2018;5(8):ofy172.
Legoff J, Zucman N, Lemiale V, et al. Clinical significance of upper airway virus detection in critically ill hematology patients. Am J Respir Crit Care Med. 2019;199(4):518–28.
Wu X, Li Y, Zhang M, et al. Etiology of severe community-acquired pneumonia in adults based on metagenomic next-generation sequencing: a prospective multicenter study. Infect Dis Ther. 2020;9(4):1003–15.
Wang H, Xiao M, Chen SC, et al. In vitro susceptibilities of yeast species to fluconazole and voriconazole as determined by the 2010 National China Hospital Invasive Fungal Surveillance Net (CHIF-NET) study. J Clin Microbiol. 2012;50(12):3952–9.
Arias Fernández L, Pardo Seco J, Cebey-López M, et al. Differences between diabetic and non-diabetic patients with community-acquired pneumonia in primary care in Spain. BMC Infect Dis. 2019;19(1):973.
Kim EJ, Ha KH, Kim DJ, Choi YH. Diabetes and the risk of infection: a national cohort study. Diabetes Metab J. 2019;43(6):804–14.
Lin CY, Wang IT, Chang CC, et al. Comparison of clinical manifestation, diagnosis, and outcomes of invasive pulmonary aspergillosis and pulmonary mucormycosis. Microorganisms. 2019;7(11):531.
Ghanaat F, Tayek JA. Weight loss and diabetes are new risk factors for the development of invasive aspergillosis infection in non-immunocompromized humans. Clin Pract (Lond). 2017;14(5 Spec Iss):296–301.
da Silva RB, Neves RP, Hinrichsen SL, de Lima-Neto RG. Candidemia in a public hospital in Northeastern Brazil: epidemiological features and risk factors in critically ill patients. Rev Iberoam Micol. 2019;36(4):181–5.
Ugarte-Gil C, Alisjahbana B, Ronacher K, et al. Diabetes mellitus among pulmonary tuberculosis patients from 4 tuberculosis-endemic countries: the TANDEM study. Clin Infect Dis. 2020;70(5):780–8.
Simner PJ, Miller S, Carroll KC. Understanding the promises and hurdles of metagenomic next-generation sequencing as a diagnostic tool for infectious diseases. Clin Infect Dis. 2018;66(5):778–88.
Dulanto Chiang A, Dekker JP. From the pipeline to the bedside: advances and challenges in clinical metagenomics. J Infect Dis. 2020;221(Suppl 3):S331-s340.
Hirai J, Sakanashi D, Haranaga S, et al. Case-control study of pneumonia patients with Streptococcus anginosus group bacteria in their sputum. J Infect Chemother. 2016;22(12):794–9.
Noguchi S, Yatera K, Kawanami T, et al. The clinical features of respiratory infections caused by the Streptococcus anginosus group. BMC Pulm Med. 2015;15:133.
Morris JF, Sewell DL. Necrotizing pneumonia caused by mixed infection with Actinobacillus actinomycetemcomitans and Actinomyces israelii: case report and review. Clin Infect Dis. 1994;18(3):450–2.
Doern GV, Brogden-Torres B. Optimum use of selective plated media in primary processing of respiratory tract specimens from patients with cystic fibrosis. J Clin Microbiol. 1992;30(10):2740–2.
Li Y, Sun B, Tang X, et al. Application of metagenomic next-generation sequencing for bronchoalveolar lavage diagnostics in critically ill patients. Eur J Clin Microbiol Infect Dis. 2020;39(2):369–74.
Salzberg SL, Breitwieser FP, Kumar A, et al. Next-generation sequencing in neuropathologic diagnosis of infections of the nervous system. Neurol Neuroimmunol Neuroinflamm. 2016;3(4):e251.
Mongkolrattanothai K, Naccache SN, Bender JM, et al. Neurobrucellosis: unexpected answer from metagenomic next-generation sequencing. J Pediatr Infect Dis Soc. 2017;6(4):393–8.
Wilson MR, Suan D, Duggins A, et al. A novel cause of chronic viral meningoencephalitis: Cache Valley virus. Ann Neurol. 2017;82(1):105–14.
Geng S, Mei Q, Zhu C, et al. Metagenomic next-generation sequencing technology for detection of pathogens in blood of critically ill patients. Int J Infect Dis. 2021;103:81–7.
Pendleton KM, Erb-Downward JR, Bao Y, et al. Rapid pathogen identification in bacterial pneumonia using real-time metagenomics. Am J Respir Crit Care Med. 2017;196(12):1610–2.
Chen J, Zhao Y, Shang Y, et al. The clinical significance of simultaneous detection of pathogens from bronchoalveolar lavage fluid and blood samples by metagenomic next-generation sequencing in patients with severe pneumonia. J Med Microbiol. 2021;70(1). https://doi.org/10.1099/jmm.0.001259.
Acknowledgements
Funding
This work was supported by the Special Project in Clinical Medicine from the Nantong University, Nantong, China No.2019LY045 (the study); Clinical Key Specialty Construction Foundation of Jiangsu Province (the journal’s Rapid Service Fee). The funders had no role in the study design, conduct, analysis, interpretation of study results, decision to publish, or preparation of the manuscript.
Authorship
All named authors meet the International Committee of Medical Journal Editors (ICMJE) criteria for authorship for this article, take responsibility for the integrity of the work as a whole, and have given their approval for this version to be published.
Authors’ Contributions
All authors contributed to the study conception and design. Material preparation was performed by GX, BZ, XR, JJ and TH. Data collection was performed by GX, BZ, XW, LB, YX and HZ. Analysis was performed by GX, BZ, XW and HZ. The first draft of the manuscript was written by GX and BZ. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript. GX and BZ are the co-first authors and contributed equally to this study.
Disclosures
Guijuan Xie, Bo Zhao, Xun Wang, Liang Bao, Yiming Xu, Xian Ren, Jiali Ji, Ting He and Hongqing Zhao declare that they have no conflicts of interest to this work.
Compliance with Ethics Guidelines
The study was conducted in accordance with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. This study was approved by the ethics committee of The Affiliated Wuxi Second People’s Hospital of Nanjing Medical University. Since the data were completely anonymous, informed consent was waived.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Open Access
This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Xie, G., Zhao, B., Wang, X. et al. Exploring the Clinical Utility of Metagenomic Next-Generation Sequencing in the Diagnosis of Pulmonary Infection. Infect Dis Ther 10, 1419–1435 (2021). https://doi.org/10.1007/s40121-021-00476-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40121-021-00476-w