Abstract
Background
We have previously shown that hsa-miR-520d-5p can convert cancer cells into induced pluripotent stem cells (iPSCs) or mesenchymal stem cells (MSCs) via a dedifferentiation by a demethylation mechanism.
Methods
We tested the effect of miR-520d-5p on human fibroblasts to determine whether it could be safely used in normal cells for future clinical therapeutic applications. After we transfected the microRNA into fibroblasts, we analyzed the phenotypic changes, gene expression levels, and stemness induction in vitro, and we evaluated tumor formation in an in vivo xenograft model.
Results
The transfected fibroblasts turned into CD105+ cell populations, survived approximately 24 weeks, and exhibited increases in both the collagen-producing ability and in differentiation. Combinatorial transfection of small interfering RNAs for miR-520d-5p target genes (ELAVL2, GATAD2B, and TEAD1) produced similar results to miR-520d-5p transfection. These molecules converted normal cells into MSCs and not iPSCs.
Conclusions
In vitro data indicate the potent usefulness of this small molecule as a therapeutic biomaterial in normal cells and cancer cells because CD105+ cells never converted to iPSCs despite repeated transfections and all types of transfectants lost their tumorigenicity. This maintenance of a benign state following miR-520d-5p transfection appears to be caused by p53 upregulation. We conclude that miR-520d-5p may be a useful biomaterial at an in vitro level.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Hsa-miR-520d-5p can transform hepatoma cells into non-malignant cells [human induced pluripotent stem cell (hiPSC)-like cells] via the dedifferentiation induced by both DNA and RNA demethylation. |
Because we did not observe malignant transformation or even benign tumorigenicity in normal cells receiving it, we could confirm the safety, lack of toxicity in vitro, and potential applications of hsa-miR-520d-5p and its three target genes in future anti-cancer or anti-aging therapies. |
A fuller understanding of the diverse functions of hsa-miR-520d-5p may help to overcome some of the current struggles in the field of medical science. |
1 Introduction
Mature microRNAs (miRNAs; single-stranded RNA molecules of 18–23 nucleotides) control gene expression in many cellular processes [1]. miRNAs typically reduce the stability of messenger RNAs (mRNAs), including genes that mediate tumorigenic processes, such as apoptosis, cell cycle regulation, differentiation, inflammation, invasion, and stress responses [2]. We previously reported the effect of hsa-miR-520d-5p transfection into hepatoma cells or undifferentiated cancer cells [3].
Interestingly, the regulation of gene expression by miRNAs influences the overall development and life-support function with a certain purpose in normal and cancer cells [4, 5]. Targeting by miRNA is carried out via base-pair interactions between the 5′ end of miRNAs and sites within the coding and/or untranslated regions (UTRs) of transcriptional form of genes targeted by miRNAs; target sites in the 3′UTR lead to more effective translational dysfunction [6, 7]. Because one miRNA generally targets at least hundreds of mRNAs, it is considered to be extremely difficult to elucidate miRNA regulatory pathways [8]. In the case of miR-520d-5p, there are more than 1,000 predicted target genes, according to available bioinformatics.
Because it is common knowledge that conventional anti-cancer agents have a negligible toxic effect on normal cells [9, 10], although molecular targeted anti-cancer agents including antibody drugs are not so toxic to normal cells, but that the anti-cancer effects gradually tend to get weaker, it is expected that normal cells transfected with miR-520d-5p will be less subject to toxic effects. In a previous study, 520d-5p-transfection induced undifferentiated cancer cells or hepatomas with a high or low degree of differentiation into a benign state due to p53 upregulation [3]. Because fibroblasts have recently been shown to be a useful starting material for induced pluripotent stem cell (iPSC) generation in the field of regenerative medicine, in this study we attempt to confirm whether miR-520d-5p transfection can turn fibroblasts into novel stem cells retaining p53 upregulation, unlike iPSC [11, 12]. If miR-520d-5p is a good candidate for use in anti-cancer therapies, it will be important to determine whether local or systemic administration has an effect on normal cells [10]. Because it is also necessary to test whether it has toxicity and tumorigenicity in vivo, in this study we examined the influence and effect of miR-520d-5p on fibroblasts and vascular endothelial cells both in vitro and in vivo, to prove the safety. Additionally, we examined the effects of small interfering RNAs (siRNAs) for the predicted target genes, including ELAVL2 [3].
2 Methods
2.1 Cells
To determine the in vitro and in vivo effects of hsa-miR-520d-5p expression and to explore its safety for future systemic administration, we used three cell lines and lentiviral vectors. Human iPSCs (hiPSCs) (HPS0002) were provided by the RIKEN BioResource Center Cell Bank (Ibaraki, Japan), and both human umbilical vein endothelial cells (HUVECs) and normal human dermal fibroblast (NHDF) cells were provided by TAKARA BIO Inc. (Tokyo, Japan). To examine the effect of miR-520d-5p on normal cells in vitro and in vivo, we used a human fibroblast cell line (NHDF-Ad derived from adults) and HUVECs, cultured in fibroblast basal medium (FBM)-2 medium using the fibroblast growth medium chemically defined (FGM-CD) Bullet Kit and EBM-2 using EBM-2 SingleQuots, respectively (TAKARA BIO Inc.). The hiPSCs were cultured in ReproStem medium (ReproCell, Tokyo, Japan) with 5 ng/mL basic fibroblast growth factor (bFGF)-2. In addition, the human mesangial cell line 293FT (Invitrogen Japan K.K., Tokyo, Japan) was used for producing miR-520d-expressing lentivirus as previously reported [13]. The 293FT cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10 % fetal bovine serum (FBS), 0.1 mM minimum essential medium (MEM) non-essential amino acids solution, 2 mM l-glutamine, and 1 % penicillin/streptomycin.
2.2 Lentiviral Vector Constructs
To examine the effects of miR-520d-5p over-expression on normal cells, we transfected pMIRNA1-miR-520d-5p/green fluorescent protein (GFP) (20 μg; System Biosciences, Mountain View, CA, USA) or the mock vector pCDH/lenti/GFP (20 μg) into 293FT cells. To harvest viral particles, the cells were centrifuged at 170,000 × g (120 min, 4 °C). The viral pellets were collected, and the viral copy numbers were measured using a Lenti-XTM quantitative reverse transcriptase–polymerase chain reaction (qRT-PCR) Titration kit (Clontech, Mountain View, CA, USA). For NHDF-Ad or HUVEC infection, 1.0 × 106 copies of the lentivirus were used per 10 cm culture dish. To confirm the status of GATAD2B and TEAD1 as candidate target genes of miR-520d-5p, short hairpin RNAs (shRNAs) for GATAD2B and TEAD1 were purchased from GeneCopoeia (Rockville, MD, USA). The siRNA sequences for ELAVL2 have been described in a previous report and the siRNA sequences for GATAD2B and TEAD1 are as follows [3]:
GATAD2B-1: 5′-tcttcgcttgaatctgttgtcaagagcaacagattcaagcgaaga-3′
GATAD2B-2: 5′-gcagaatgcagcatctattgtcaagagcaatagatgctgcattctg-3′
GATAD2B-3: 5′-gccttccacacatgttgatgtcaagagcatcaacatgtgtggaagg-3′
GATAD2B-4: 5′-gtgtcagcgtacaacatccttcaagagaggatgttgtacgctgaca-3′
TEAD1-1: 5′-ggactctgcagataagccaatcaagagttggcttatctgcagagtc-3′
TEAD1-2: 5′-gactgccattcataacaagctcaagaggcttgttatgaatggcag-3′
TEAD1-3: 5′-ggcatgccaaccattcttactcaagaggtaagaatggttggcatgc-3′
TEAD1-4: 5′-gaaggtggcttaaaggaacttcaagagagttcctttaagccacctt-3′
The scrambled sequence for miR-520d-5p was 5′-gaguccgcctcuatagacaa-3′, and 5′-auucguuguuauugu-3′ was used as a scramble for GATAD2B and TEAD1. The siRNA lentiviral constructs were transfected into NHDF cells as per the optimized method. siRNAs for three genes (ELAVL2, GATAD2B, and TEAD1) were transfected into NHDF-Ad cells individually and in all possible combinations, for a total of seven different transfections.
2.2.1 Immunodeficient Mice and the In Vivo Study
One week after the lentiviral infection, 5 × 107 NHDF and HUVEC cells were harvested and injected intraperitoneally (n = 8), intramuscularly into the right femoral region (n = 8), or subcutaneously into the right flank (n = 8) (injection volume, 200 µL). Six-week-old immunodeficient mice (KSN/Slc; Shimizu Laboratory Supplies, Kyoto, Japan) were allowed free access to water and food for 12–24 weeks after the injections until they were anesthetized with 100 mg/kg nembutal and euthanized. All animals were housed and fed in the Division of Laboratory Animal Science at Tottori University (Tottori, Japan) under a protocol that was approved by the Japanese Association for Accreditation for Laboratory Animal Care, and the animal research and handling were performed in strict accordance with the federal Institutional Animal Care and Use Committee guidelines. All experiments reported in this study were approved by an institutional committee.
2.3 Gene Expression Analysis by Reverse Transcriptase–Polymerase Chain Reaction (RT-PCR)
Total RNA, including the small RNA fraction, was extracted from cultured cells or homogenized mouse tissues with the mirVana miRNA Isolation Kit (Ambion, Austin, TX, USA). Mature miRNA (miR-520d-5p: 25 ng/μL) was quantified with the Mir-X™ miRNA qRT-PCR SYBR® kit (TAKARA BIO Inc., Tokyo, Japan) according to the manufacturer’s instructions. The gels were run under the same experimental conditions. The polymerase chain reaction (PCR) and data collection analyses were performed with a BioFlux LineGene (Toyobo, Nagoya, Japan). To analyze the reverse transcriptase–PCR (RT-PCR) results, all data except those for human telomerase reverse transcriptase (hTERT) were normalized to the internal control β-actin. The hTERT expression was estimated by the copy number according to a previously developed quantification method [14]. The U6 small nuclear RNA was used as an internal control. The total RNA (50 ng/μL) was reverse transcribed and amplified using the KAPA SYBR FAST One-Step qRT-PCR Kit (NIPPON Genetics Co., Ltd, Tokyo, Japan). The RNA quantification was confirmed by sequencing with high reproducibility. Electronic Supplementary Material (ESM) Table S1 shows the primer sequences that were used for mRNA or miRNA quantification. The data were calculated statistically by a one-way ANOVA or a Mann–Whitney U test, and the significant differences are shown as *P < 0.05 and **P < 0.01.
2.4 Western Blotting
We performed Western blotting with 20 μg/μL of protein and the i-Blot gel transfer system (Invitrogen, Tokyo, Japan). The anti-CD105, -p53, -SIRT1, -caspase antibodies were diluted 1:500, and the anti-β-actin antibody was diluted 1:1,000 according to the manufacturer’s instructions. The chemiluminescent signals were detected within 1 min with an LAS-4000 (Fujifilm, Tokyo, Japan).
2.5 Immunocytochemistry
Immunohistochemical studies were performed with antibodies to detect a multilineage-differentiating stress-enduring (muse) cell marker (anti-CD105), markers of differentiation [anti-Osteocalcin, anti-Aggrecan, or anti-fatty-acid-binding protein (FABP) antibodies] and the Mesenchymal Stem Cell Marker Antibody Panel according to the manufacturer’s instructions (R&D Systems, Minneapolis, MN, USA). The NHDF-Ad and HUVEC cells were infected with lentiviral particles that contained hsa-miR-520d-5p. The transfectants were harvested and transferred to a new culture dish for microscopic examination or to a slide chamber for immunostaining.
2.6 Type I Collagen Quantification
To examine the ability of fibroblasts or 520d-fibroblasts to produce type I collagen, 200 µL of supernatant was collected from cultures consisting of 1 × 106 cells in 25 cm2 dishes, and the quantity of collagen that the 520d-NHDF cells produced was measured using the Human Collagen Type I ELISA kit (ACEL, Inc., Sagamihara, Japan).
2.7 Histological Examination
The lung, the liver, intraperitoneal or post-peritoneal metastases, and the tumor volumes were investigated macroscopically or under a dissecting microscope with bright-field imaging. The tissue samples were fixed in 10 % buffered formalin overnight, washed with phosphate buffered saline (PBS), transferred to 70 % ethanol, embedded in paraffin, sectioned, and stained with hematoxylin–eosin (HE).
2.8 Fluorescence Detection in Cells
To estimate the efficacy of infection by the miR-520d-5p-expressing lentiviral vector, GFP expression was detected with an OLYMPUS IX71 microscope with a TH4-100 power supply (Tokyo, Japan).
2.9 Measurement of 5-Hydroxymethyl-Cytosine (5-hmC) Percentage
The 5-hydroxymethyl-cytosine levels were measured in the NHDF-Ad cells, the mock-NHDF-Ad cells, the 520d-NHDF-Ad cells, and hiPSC cells (200 ng of DNA from each), using the MethylFlash Hydroxymethylated DNA Quantification kit (Colorimetric) according to the manufacturer’s instruction (EPIGENTEK, Farmingdale, NY, USA).
2.10 Telomere Length Measurement
The telomere length in the NHDF-Ad cells and the 520d-NHDF cells was measured using the TeloTAGGG Telomere Length Assay (Roche Applied Science, Basel, Switzerland).
2.11 Prediction of GATAD2B and TEAD1 as Target Genes of miR-520d-5p
The potential target genes for miR-520d-5p (MIMAT0002855: cuacaaagggaagcccuuuc) were predicted using several databases (miRBase: http://www.mirbase.org, DIANA-MICROT: http://diana.cslab.ece.ntua.gr/DianaTools/, miRDB: http://mirdb.org, RNA22-HAS: http://cm.jefferson.edu/rna22v1.0, TargetMiner: http://www.isical.ac.in/~bioinfo_miu, mircoRNA.org: http://www.microrna.org/microrna, and TargetScan-VERT: http://www.targetscan.org/cgi-bin/targetscan/vert_50). After confirming the gene downregulation by RT-PCR, we examined the gene expression in cells transfected with siRNAs against GATAD2B and TEAD1 (siGATAD2B-NHDF and siTEAD1-NHDF; four different siRNAs for each gene; see Sect. 2.2, Lentiviral Vector Constructs) and compared the results with the gene expression levels in the 520d-NHDF cells. We performed RT-PCR, Western blotting, immunocytochemistry, and cell cycle analysis as previously described. To investigate the binding of miR-520d-5p to the 3′UTR of GATAD2B or TEAD1, sequences (or mismatch sequences) that corresponded with predictive miR-520d-5p-binding sites were ligated into the multiple cloning sites (MCS) (PmeI and XhoI) of psiCHECK-2 (Promega KK, Tokyo, Japan). The signals were detected with the Dual-Luciferase Reporter expression assay (Promega KK, Tokyo, Japan). Synthesized miR-520d-5p (MBL, Nagoya, Japan) or the control vector pMIRNA1/GFP (System Biosciences, Mountain View, CA, USA) were co-transfected into NHDF cells together with each prepared luciferase expression vector (firefly or Renilla). Synthesized hsa-miR-520d-3p (MBL, Nagoya, Japan) or pLKO.1 (Addgene, Cambridge, MA, USA) with mismatch sequences were used as the controls. Forty-eight hours after transfection, the luciferin expression (RLU) was measured with an Infinite F500 microplate reader (TECAN, Männedorf, Switzerland). The RLU (Renilla/firefly) was standardized to a control (n = 4). The siGATAD2B-NHDF cells or the siTEAD1-NHDF cells were injected into the right hindquarters of immunodeficient mice, which were then fed routinely for an average of 3 months and observed for tumorigenicity and estimation of tumor quality.
The following six pairs of sequences were inserted into psiCHECK-2:
GATAD2B-3′UTR sense 1, 5′-TCGAGTGTATTCTTTGTAAAATTCGTTT-3′; antisense 1, 5′-AAACGAATTTTACAAAGAATACA-3′;
GATAD2B-3′UTR sense 2, 5′-TCGAGGAACTGCTTTGTAAAGATGG-3′; antisense 2, 5′-AAACCATCTTTACAAAGCAGTTC-3′;
GATAD2B-3′UTR mismatch sense, 5′-TCGAGTGTATTCTTAATAAAATTCGTTT-3′; mismatch antisense, 5′-AAACGAATTTTATTAAGAATACA-3′;
TEAD1-3′UTR sense 1, 5′-TCGAGACTTATCTTTGTAACTAATGTTT-3′; antisense 1, 5′-AAACATTAGTTACAAAGATAAGT-3′;
TEAD1-3′UTR sense 2, 5′-TCGAGAAAGTGCTTTGTAAAAAAAGTTT-3′; antisense 2, 5′-AAACTTTTTTTACAAAGCACTTT-3′;
TEAD1-3′UTR mismatch sense 3, 5′-TCGAGACTTATCTTAATAACTAATGTTT-3′; and mismatch antisense 3, 5′-AAACATTAGTTATTAAGATAAGT-3′.
2.12 Differentiation of 520d-NHDF Cells by Differentiation-Inducing Agents
Following treatment with differentiation agents, the induction of osteogenic, chondrogenic, and adipogenic differentiation in the 520d-NHDF-Ad cells was examined using the Mesenchymal Stem Cell Identification Kits (R&D Systems, Inc., Minneapolis, MN, USA).
2.13 Statistical Analysis
Mann–Whitney U test, t-test or a one-way ANOVA were used for comparisons between the controls, the mock-treated and the miR-520d-5p results with one observed variable. P < 0.05 was considered significant (*P < 0.05, **P < 0.01). In the box plots, the top and bottom of each box represent the 25th and 75th percentiles, respectively, providing the interquartile range. The line through the box indicates the median, and the error bars indicate the 5th and 95th percentiles.
3 Results
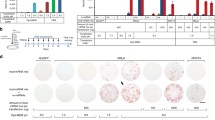
NHDF cells [520d-NHDF (hsa-miR-520d-5p-expressing-NHDF cells)] that were infected with the miR-520d-5p-expressing lentiviral vector were observed until cell death. Cells experienced senescence three times, and each time transfection just after senescence restored cell growth. However, the fourth transfection never restored senescence and led to cell death (at 22–24 weeks) (Fig. 1a). By contrast, NHDF cells without transfection or mock-treated-NHDF (mock-NHDF) cells failed to restore cell growth, and cells experienced cell death at 5–6 weeks or at 8–10 weeks, respectively. Senescent NHDF cells received the first miR-520d-5p transfection at 6–7 weeks after the cultures began growing (Fig. 1b; top left), resulting in two signs of growth restoration (arrows indicate the phenotypes) (Fig. 1b; bottom left and top right). GFP expression in spherical new cell populations indicated the success of the transfections (Fig. 1b; bottom right).
a A time course of the lentiviral transfection of miR-520d-5p into NHDF-Ad cells. The NHDF-Ad cells experienced cell death at 5–6 weeks after the culture began growing. However, transfection of miR-520d-5p into the parental cells after the first senescence (S; indicated by an arrow) resulted in the extension of their lifespan to a total of 18–19 weeks, including two more cycles of senescence and subsequent miR-520d-5p-transfection, until the cells finally underwent apoptosis. b The phenotypic changes of the NHDF-Ad cells are shown. Top the parental cells experienced senescence at approximately 6 weeks. After transfection with miR-520d-5p, giant and spheroid populations emerged, and the new cells generated fibroblasts radially one after another. Bottom the new fibroblast-like cells were slightly longer in shape or more rapidly proliferating compared with the NHDF-Ad cells (left). The giant cells express GFP, indicating the successful transfection of miR-520d-5p into the NHDF-Ad cells; the merged image shows the overlap of the GFP expression with the giant cells [differential interference contrast (DIC) image at 40× magnification] (right). The magnification levels or scale bars are shown in each figure. The two arrows indicate the two types of phenotypes of the transfectants. All mock-transfectants showed the similar phenotype and process till the cell death to parental cells. c A representative transfectant at 22 weeks was shown by DIC (left), GFP expression (middle), and CD105 expression (right by immunocytochemistry). CD105 was expressed mainly in the cytoplasm and the cell membrane. IgG controls of 520d-transfectants in 20 W were shown to the right column. Mock-transfectants and parental cells in senescence showed the similar staining level to IgG controls. d A representative gene expression profile shown by RT-PCR. Nanog and p53 were strongly expressed and ELAVL2 and AID were significantly downregulated to 0.11 and 0.22, respectively. The expression of each mRNA was normalized to β-actin (n = 8, *P < 0.05, **P < 0.01 by Mann–Whitney U test). e The quantity of collagen produced by the fibroblasts was measured from the supernatant (126 µL) from NHDF-Ad cells cultured in 10 cm dishes (n = 3). The data showed that transfection with miR-520d-5p stimulated the collagen-producing ability of fibroblasts (transfectants). S indicates the senescent state of the transfectants. f The DNA methylation level (5-hmC percent) was measured in the 520d-NHDF-Ad cells relative to the NHDF cells, the mock-NHDF cells and the hiPSCs (at 2 weeks) (n = 4). The 520d-NHDF cells were significantly demethylated compared with the parental cells (*P < 0.05 by Mann–Whitney U test). The hiPSCs generally had a lower level of methylation than the NHDF-Ad cells, but significant differences were not observed. AID activation-induced cytidine deaminase, cntl control, DIC differential interference contrast, GFP green fluorescent protein, hiPSC human induced pluripotent stem cell, mRNA messenger RNA, NHDF normal human dermal fibroblast, NHDF-Ad normal human dermal fibroblast derived from adults, RT-PCR reverse transcriptase–polymerase chain reaction
The two types of 520d-NHDF cells, as well as the 520d-NHDF cells at 22 weeks just before senescence, expressed the mesenchymal marker CD105 (Fig. 1c; left and right) as well as GFP (Fig. 1c; middle) and were converted to spherical cell populations of 1–5 cells per 6 cm plate (Fig. 1b; top right). The 520d-NHDF cells transcriptionally upregulated Nanog, p53, and Oct4, but they downregulated activation-induced cytidine deaminase (AID) and embryonic lethal, abnormal vision, Drosophila-like 2 (ELAVL2) (Fig. 1d) [3, 15, 16]. The ability of the NHDF cells to produce type I collagen was restored by the transfection with miR-520d-5p two out of three times. In Fig. 1e, the solid arrow indicates the general reduction of type I collagen produced by the NHDF cells, while the dotted arrow indicates the gradual reduction rescued by miR-520d-5p.
The DNA methylation level was examined in the NHDF cells because miR-520d-5p has been shown to lead to demethylation in cancer cells [3]. The methylation level (5-hmC percent in DNA) was significantly reduced in the 520d-NHDF cells 2 weeks post-transfection [17] compared with NHDF cells, but there was no significant difference between the mock-NHDF, the 520d-NHDF, and the hiPSCs (*P < 0.01; Fig. 1f). In adult fibroblasts, the effect of 520d-5p on DNA demethylation was very weak. However, the 520d-NHDF cells experiencing senescence (at 18–24 weeks) had significantly elevated 5-hmC level relative to NHDF cells, mock-NHDF cells, or hiPSCs (ESM Fig. S1).
Using bioinformatic algorithms, miR-520d-5p was predicted to bind to sequences including TACAAA in the 3′UTRs of TEAD1 and GATAD2B, according to the same method performed regarding ELAVL2 (Fig. 2a) [3]. A luciferase reporter assay revealed that two such sites [TTACAAAG (3′UTR-1) and TACAAAG (3′UTR-2)] included in both the GATAD2B 3′UTR and the TEAD1 3′UTR are actually targeted by miR-520d-5p (Fig. 2b).
a A luciferase reporter expression assay was performed to examine whether miR-520d-5p actually targets TEAD1 and GATAD2B. Out of the more than ten predicted binding sites in the 3′UTRs of TEAD1 and GATAD2B, two sequences [TTACAAG and TACAAAG in the 3′UTR of TEAD1 (top) or GATAD2B (bottom)] were chosen to test for miR-520d-5p binding. These sites were predicted based on the four databases described above. b A luciferase reporter expression assay revealed that miR-520d-5p bound to both the 3′UTR of TEAD1 and the 3′UTR of GATAD2B. In contrast, a mismatch of miR-520d-5p (a control for standardizing data) and a scramble group did not significantly bind to the 3′UTR sites, resulting in a failure to induce luciferase expression. Using a luciferase reporter expression assay, the potent sites of miR-520d-5p binding to the 3′UTR were identified and representatively shown at base pairs 4,569–4,581 and 7,463–7,776 in the 3′UTR of TEAD1 and 3,622–3,639 in the 3′UTR of GATAD2B, respectively. The minus signs (−) in the synthesized miR-520d-5p sequence represent the mismatches in the synthesized miR-520d-3p, and the minus signs in the control vector pMIRNA1/GFP sequence represent the mismatch sequences within miR-520d-5p that was expressed from pMIRNA1-520d-5p/GFP (n = 4). The data were analyzed by t test (*P < 0.05 and **P < 0.01). c The phenotypic changes in fibroblasts induced by siETG (siELAVL2/siTEAD1/siGATAD2B) are shown. siETG induced large cell populations that could generate fibroblasts radially outwards. The fibroblasts were cultured for 4 weeks under routine culture conditions. A bar (lower right) indicates the scale. d Immunocytochemistry was performed on the fibroblasts (NHDF-Ad cells) 4 weeks after the cultures began growing. A representative fibroblast with no siRNA transfection is shown (left), and it expressed CD105 very weakly in the nucleus (right). e Immunocytochemistry was performed on the siETG transfectants at 22 weeks after the cultures began growing. Almost all the fibroblasts expressed CD105 strongly (left), and a representative magnified fibroblast expressed CD105 in the cytoplasm and the cell membrane (right). A DIC image of the fibroblast is shown in the middle part. DIC differential interference contrast, GFP green fluorescent protein, NHDF-Ad normal human dermal fibroblast derived from adults, siRNA small interfering RNA, UTR untranslated region
Because combinatorial transfection with three siRNAs (siETG: siELALVL2/siTEAD1/siGATAD2B) reproduced the CD105 upregulation and resulted in similar phenotypic changes as those seen following miR-520d-5p transfection, we simultaneously transfected the three siRNAs into NHDF cells and examined the effects. Single or pairwise transfections of the siRNAs had a weaker effect than siETG transfection, which could generate major cell growth resulting in radial generation of new fibroblasts (Fig 2c). Compared with scramble-transfected NHDF cells (Fig. 2d), CD105 was continuously upregulated in the cytoplasm of the 520d-NHDF cells (Fig. 2e). Expression of CD105, Nanog, Oct4, p53, and SIRT1 [18] mRNAs was upregulated in response to miR-520d-5p-transfection, but hTERT failed to respond to the last transfection (the arrows indicate the timing of transfection with 520d, and S indicates the senescence of the transfectants) (Fig. 3a). Although insufficient cell numbers prevented cell harvest at 16 weeks post-transfection, CD105, p53, and caspase-8 were translationally upregulated in response to siETG transfection. SIRT1 did not appear to be similarly upregulated (ESM Fig. 2S).
a The relative ratios of transcriptional expression obtained by RT-PCR in siETG-NHDF-Ad cells, which were cultured until cell death following transfection, are depicted. The siETG-NHDF cells expressed CD105, hTERT, Nanog, Oct4, p53, and SIRT1 in response to the first two transfections but the third transfection induced little to no increase in gene expression. Very similar results were obtained from the 520d-NHDF cells. The data were normalized to the expression level in the parental NHDF cells, and the relative averages were shown (n = 4). b Induced differentiation was confirmed by comparing the levels of commercially provided differentiation markers in 520d-NHDF-Ad cells (left), and in mock-NHDF-Ad cells (right). Osteogenic, chondrogenic, and adipogenic differentiation were induced in the 520d-NHDF cells (left part of 520d-NHDF-Ad) as detected by Osteocalcin, Aggrecan, and FABP expression, respectively (right part of 520d-NHDF-Ad), indicating that these transfected cells have mesenchymal stem cell-like capabilities. The GFP expression in these cells (middle part of 520d-NHDF-Ad) indicates successful transfection. The scale bar in each figure is 20 µm. FABP fatty-acid-binding protein, GFP green fluorescent protein, hTERT human telomerase reverse transcriptase, mRNA messenger RNA, NHDF normal human dermal fibroblast, NHDF-Ad normal human dermal fibroblast derived from adults, RT-PCR reverse transcriptase–polymerase chain reaction
As a further test, we transfected miR-520d-5p into HUVECs (ESM Fig. S3A: left). 520d-HUVECs (hsa-miR-520d-5p-overexpressing HUVECs) gave rise to some grouped cells (ESM Fig. S3A: middle and right) but did not show the significant alterations in gene expression (CD34, CD45, and ROBO4) peculiar to their precursors, the endothelial progenitor cells (EPCs) (ESM Fig. S3B) [19–21]. In addition, the telomere length in the transfectants did not show significant differences between the 520d-NHDF cells, the mock-NHDF cells, and the NHDF cells until 22 weeks after transfection.
Osteogenic, chondrogenic, and adipogenic differentiation of the 520d-NHDF-Ad cells was confirmed (Fig. 3b). Osteocalcin, Aggrecan, and FABP expression were observed (top, middle, and bottom, respectively: Fig. 3b; left). Control cells transfected with a mock-vector showed very weak expression of each gene (Fig. 3b; right). GFP expression was used to confirm the successful transfection of miR-520d-5p into the NHDF cells, although GFP was expressed very weakly in the chondrogenic NHDF cells.
To confirm the in vitro results in vivo, 1.0 × 106 viral copies of miR-520d-5p were transfected into 1.0 × 106 NHDF-Ad cells or HUVECs, which were then inoculated into athymic KSN/Slc mice. The NHDF or HUVEC cells that received 1.0 × 106 copies of miR-520d-5p (n = 24) did not generate any malignant or benign tumors in mice (intraperitoneally, subcutaneously, or intramuscularly; n = 8 mice for each). The inoculated mice did not show any significant body weight loss (mean ± standard error of the mean and range: 27.7 ± 0.5 and 25.5–29.6 g for 520d-NHDF and 27.5 ± 0.4 and 25.6–29.8 g for 520d-HUVEC), compared with the control mice (27.8 ± 0.4 and 25.7–29.6 g). Furthermore, they did not show any toxicity or exhibit any disorders. In summary, the in vivo study demonstrated that mice inoculated with 520d-NHDF or 520d-HUVEC cells at a sufficient and effective viral titer (in viral copy number/cell) never gave rise to any tumorous lesions in any tissues over a 24-week period.
4 Discussion
The accumulated data have clarified that miRNAs are involved in numerous biological processes, and miRNA dysregulation frequently occurs in various carcinomas [22–24]. Many recent studies have attempted to better understand the role of miRNAs and reveal the importance of miRNA-mediated regulation in cancer and in normal cells [25–27]. The potential application of miRNA families in iPS generation has been reported previously [28]. The results we have reported on the effects of miR-520d-5p in carcinogenesis and normal stem cell differentiation strongly suggest the actual possibility of transforming cancer stem cells (CSCs) into normal stem cells [iPSCs or mesenchymal stem cells (MSCs)] [3, 29].
Muse cells are distinct stem cells in the mesenchymal cell population with the capacity to self-renew, to differentiate into cells representative of all three germ layers from a single cell, and to repair damaged tissues by spontaneous differentiation into tissue-specific cells without forming teratomas [30]. We describe some procedures for isolating and evaluating these cells. Muse cells are also a practical cell source for hiPSCs with markedly high generation efficiency [31]. They can be sorted from commercially available mesenchymal cells (such as adult human bone marrow stromal cells and dermal fibroblasts, or from fresh adult human bone marrow samples) as cells that are double positive for stage-specific embryonic antigen-3 (SSEA-3) and CD105 [32].
This study was performed to confirm the safety of miR-520d-5p in normal cells. The miR-520d-5p-transfected fibroblasts expressed CD105/Endoglin (Fig. 1c), significantly elevated p53 and Nanog expression (Fig. 1d) [33], increased their collagen-producing ability (Fig. 1e) [34], and extended their longevity to approximately 24 weeks, although unlike hTERT, miR-520d-5p is not sufficient for immortalization [35]. Because the elongation or maintenance of telomere length was not observed despite a temporary increase in hTERT expression in response to miR-520d-5p [36, 37], the transfectants followed a natural aging process.
Evidence suggests that transient telomerase expression and only a very small average telomere elongation by hTERT resulted in a 50 % increase in the lifespan of human fibroblasts [36]. The DNA methylation level was rapidly and significantly reduced in response to the first three miR-520d-5p transfections (Fig. 1f), but the fourth transfection failed to reduce DNA methylation and the cells experienced senescence and eventually cell death (ESM Fig. 1S). This study has shown that it is safe to transfect miR-520d-5p even into HUVECs (ESM Fig. 3S), suggesting that this molecule may be a candidate for in vivo trials.
TEAD1 and GATAD2B may be target genes of miR-520d-5p. It is known that TEAD1 is a transcription factor that is a key component of the Hippo signaling pathway [37]. Signaling plays a role in organ size control and tumor suppression by restricting proliferation and promoting apoptosis, and it also acts by mediating gene expression of YAP1 and WWTR1/TAZ, thereby regulating cell proliferation, migration and induction of epithelial to mesenchymal transitions (EMT) [38]. GATAD2B functions as a transcriptional repressor and also as an enhancer of MBD2-mediated repression [39, 40].
TEAD1 and GATAD2B are each predicted to have more than ten binding sites for miR-520d-5p in their 3′UTR sequences [representative sites starting at base pairs 4,569 and 7,463 (TTACAAAG), and 506 and 7,256 (TACAAAG) for TEAD1 and base pairs 930, 3,300, and 5,309 (TTACAAAG), and 3,622 (TACAAAG) for GATAD2B] (Fig. 2a), which suggests that miR-520d-5p possibly binds to at least four sites in each 3′UTR (Fig. 2b). Suppression of target genes by each siRNA for ELAVL2, TEAD1, and GATAD2B was confirmed using RT-PCR (ESM Fig. 4S). Because combinatorial lentiviral transfection of three siRNAs (siETG) including siELAVL2 showed similar phenotypes to miR-520d-5p, including increased lifespan and CD105 upregulation (Figs. 2c–e, 3a, b), siETG and miR-520d-5p may activate senescent cells by changing their metabolic and/or methylation status. This may allow normal cells to extend their life beyond their natural lifespan, although the upregulation of CD105 gradually decreased until cell death, as did the miR-520d-5p-transfection-induced upregulation of Nanog, Oct4, p53, SIRT1, and hTERT (Fig. 3a). Transfection of siETG into hepatoma cells (HLF or Huh7) induced a reduction in DNA demethylation almost to the levels seen in hiPSCs (ESM Fig. 5S).
Interestingly, in vivo injection of siETG-NHDF-Ad and siETG-HUVEC never gave rise to any tumors, any ectopic tissues formation, or any scars, suggesting that these molecules are safe and non-toxic enough to administer systemically with an optimized drug delivery system, and they have met the minimum requirements for advancement to preclinical trials for development as novel anti-cancer or rejuvenation agents. Combinatorial transfection of siRNAs for these three predicted target genes may have similar effects to those from miR-520d-5p alone.
We previously reported that miR-520d-5p can induce CD105 expression in undifferentiated cancer cells [3]. CD105-expressing fibroblasts can differentiate into cartilage, bone, or adipocytes, indicating that they have characteristics of MSCs. They are similar to muse cells but may not be identical (ESM Fig. 6S). Even fibroblasts transfected with miR-520d-5p more than 20 times during their 20–24 weeks were not induced into iPSCs, indicating that this miRNA might be safe, as normal cells or mock-transfected normal cells had different responses to this miRNA transfection than did cancer cells, as we reported previously [3]. This appears to be due to differences in p53 expression. p53 is upregulated in normal cells throughout the development or the differentiation, whereas it is upregulated in cancer cells following their miR-520d-5p-transfection-induced transformation to benignancy or normalcy. This may be compatible with the finding that miR-520d-5p-transfection cannot maintain the spheroid form of iPSCs because of the promotion of an epithelial-mesenchymal transition (EMT) [41]. miR-520d-5p may thus be useful for quality control following the induction of iPSC differentiation.
Furthermore, the reason we did not examine the expression of SSEA-3 with sugar chains in this study, although it is known as another biomarker of muse cells, is because we regard it as a non-essential marker in normal cells because it also strongly expresses in cancer cells as well as colorectal cancer [42]. The function and regulation of miRNAs is different between cancer cells and normal cells. Understanding the induction of demethylation or dedifferentiation may be crucial in interpreting these findings [43].
5 Conclusions
Taken together, hsa-miR-520d-5p transfection can induce fibroblasts to form MSCs with CD105 positivity (muse-like cells), as confirmed in in vitro studies and in an in vivo xenografted model. The essence of this mechanism is the induction of dedifferentiation via demethylation. However, as the effects on normal cells, the safety, and the effectiveness of miR-520d-5p were confirmed, this study suggests that this molecule may be a candidate for use in nucleic acid-based therapies. This study may provide the first evidence of juvenescence of fibroblasts with a single miRNA. The limitations of this study are that it comprised in vitro data and that no effects on normal blood cells have been examined yet.
In conclusion, in this in vitro study, we demonstrate the potential applications of miR-520d-5p and its three target genes in future anti-cancer or anti-aging therapies.
References
Lee Y, Jeon K, Lee JT, Kim S, Kim VN. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 2002;21:4663–70.
Zeng Y, Cullen BR. Sequence requirements for micro RNA processing and function in human cells. RNA. 2003;9:112–23.
Tsuno S, Wang X, Shomori K, Hasegawa J, Miura N. Hsa-miR-520d induces hepatoma cells to form normal liver tissues via a stemness-mediated process. Sci Rep. 2014;4:3852.
Farazi TA, Spitzer JI, Morozov P, Tuschl TX. miRNA in human cancer. J Pathol. 2011;223:102–15.
Miura N, Shimizu M, Shinoda W, Tsuno S, Sato R, Wang X, et al. Human RGM249-derived small RNAs may regulate tumor malignancy. Nucleic Acid Ther. 2013;23:332–43.
Guo H, Ingolia NT, Weissman JS, Bartel DP. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature. 2010;466:835–40.
Lai EC. Micro RNAs are complementary to 3′ UTR sequence motifs that mediate negative post-transcriptional regulation. Nat Genet. 2002;30:363–4.
Saetrom O, Snøve O Jr, Saetrom P. Weighted sequence motifs as an improved seeding step in microRNA target prediction algorithms. RNA. 2005;11:995–1003.
Middleton G, Brown S, Lowe C, Maughan T, Gwyther S, Oliver A, et al. A randomised phase III trial of the pharmacokinetic biomodulation of irinotecan using oral ciclosporin in advanced colorectal cancer: results of the Panitumumab, Irinotecan & Ciclosporin in COLOrectal cancer therapy trial (PICCOLO). Eur J Cancer. 2013;49:3507–16.
Hornberg JJ, Laursen M, Brenden N, Persson M, Thougaard AV, Toft DB, et al. Exploratory toxicology as an integrated part of drug discovery. Part II: screening strategies. Drug Discov Today. 2014;19(8):1137–44.
Okita K, Matsumura Y, Sato Y, Okada A, Morizane A, Okamoto S, et al. A more efficient method to generate integration-free human iPS cells. Nat Methods. 2011;8:409–12.
Hirami Y, Osakada F, Takahashi K, Okita K, Yamanaka S, Ikeda H, et al. Generation of retinal cells from mouse and human induced pluripotent stem cells. Neurosci Lett. 2009;24:126–31.
Antic D, Keene JD. Embryonic lethal abnormal visual RNA-binding proteins involved in growth, differentiation, and posttranscriptional gene expression. Am J Hum Genet. 1997;61:273–8.
Wiszniak SE, Dredge BK, Jensen KB. HuB (elavl2) mRNA is restricted to the germ cells by post-transcriptional mechanisms including stabilisation of the message by DAZL. PLoS One. 2011;6:e20773.
Piccolo FM, Fisher AG. Getting rid of DNA methylation. Trends Cell Biol. 2014;24(2):136–43.
Chen H, Liu X, Chen H, Cao J, Zhang L, Hu X, et al. Role of SIRT1 and AMPK in mesenchymal stem cells differentiation. Ageing Res Rev. 2014;13:55–64.
Liu T, Zou G, Gao Y, Zhao X, Wang H, Huang Q, et al. High efficiency of reprogramming CD34+ cells derived from human amniotic fluid into induced pluripotent stem cells with Oct4. Stem Cells Dev. 2012;21:2322–32.
Hynes K, Menicanin D, Mrozik KM, Gronthos S, Bartold PM. Generation of functional mesenchymal stem cells from different induced pluripotent stem cell lines. Stem Cells Dev. 2014;15:1084–96.
Pircher A, Wellbrock J, Fiedler W, Heidegger I, Gunsilius E. New antiangiogenic strategies beyond inhibition of vascular endothelial growth factor with special focus on axon guidance molecules. Oncology. 2014;3:46–52.
Ghildiyal M, Zamore PD. Small silencing RNAs: an expanding universe. Nat Rev Genet. 2009;10:94–108.
Ladeiro Y, Couchy G, Balabaud C, Bioulac-Sage P, Pelletier L, Rebouissou S, et al. MicroRNA profiling in hepatocellular tumors is associated with clinical features and oncogene/tumor suppressor gene mutations. Hepatology. 2008;47:1955–63.
Meltzer PS. Cancer genomics: small RNAs with big impacts. Nature. 2005;435:745–6.
Farazi TA, Hoell JI, Morozov P, Tuschl T. MicroRNAs in human cancer. Adv Exp Med Biol. 2013;774:1–20.
Li Z, Yang CS, Nakashima K, Rana TM. Small RNA-mediated regulation of iPS cell generation. EMBO J. 2011;30:823–34.
Miura N, Sato R, Tsukamoto T, Shimizu M, Kabashima H, Takeda M, et al. A noncoding RNA gene on chromosome 10p15.3 may function upstream of hTERT. BMC Mol Biol. 2009;2(10):5.
Lin SL, Chang DC, Chang-Lin S, Lin CH, Wu DT, Chen DT, et al. Mir-302 reprograms human skin cancer cells into a pluripotent ES-cell-like state. RNA. 2008;14:2115–24.
Lin SL, Chang DC, Lin CH, Ying SY, Leu D, Wu DT. Regulation of somatic cell reprogramming through inducible mir-302 expression. Nucleic Acids Res. 2010;39:1054–65.
Kuroda Y, Wakao S, Kitada M, Murakami T, Nojima M, Dezawa M. Isolation, culture and evaluation of multilineage-differentiating stress-enduring (Muse) cells. Nat Protoc. 2013;8:1391–415.
Ogura F, Wakao S, Kuroda Y, Tsuchiyama K, Bagheri M, Heneidi S, et al. Human adipose tissue possesses a unique population of pluripotent stem cells with non-tumorigenic and low telomerase activities: potential implications in regenerative medicine. Stem Cells Dev. 2014;1:717–28.
Vega Crespo A, Awe JP, Reijo Pera R, Byrne JA. Human skin cells that express stage-specific embryonic antigen 3 associate with dermal tissue regeneration. Biores Open Access. 2012;1:25–33.
Abdelalim EM, Tooyama I. Knockdown of p53 suppresses Nanog expression in embryonic stem cells. Biochem Biophys Res Commun. 2014;443:652–7.
Longo C, Galimberti M, De Pace B, Pellacani G, Bencini PL. Laser skin rejuvenation: epidermal changes and collagen remodeling evaluated by in vivo confocal microscopy. Lasers Med Sci. 2013;28:769–76.
Morales CP, Holt SE, Ouellette M, Kaur KJ, Yan Y, Wilson KS, et al. Absence of cancer-associated changes in human fibroblasts immortalized with telomerase. Nat Genet. 1999;21:115–8.
Steinert S, Shay JW, Wright WE. Transient expression of human telomerase extends the life span of normal human fibroblasts. Biochem Biophys Res Commun. 2000;14:1095–8.
Sun S, Stoflet ES, Cogan JG, Strauch AR, Getz MJ. Negative regulation of the vascular smooth muscle alpha-actin gene in fibroblasts and myoblasts: disruption of enhancer function by sequence-specific single-stranded-DNA-binding proteins. Mol Cell Biol. 1995;15:2429–36.
Zhang H, Liu CY, Zha ZY, Zhao B, Yao J, Zhao S, et al. TEAD transcription factors mediate the function of TAZ in cell growth and epithelial-mesenchymal transition. J Biol Chem. 2009;5:13355–62.
Brackertz M, Boeke J, Zhang R, Renkawitz R. Two highly related p66 proteins comprise a new family of potent transcriptional repressors interacting with MBD2 and MBD3. J Biol Chem. 2002;25:40958–66.
Joshi P, Greco TM, Guise AJ, Luo Y, Yu F, Nesvizhskii AI, et al. The functional interactome landscape of the human histone deacetylase family. Mol Syst Biol. 2013;9:672.
Termén S, Tan EJ, Heldin CH, Moustakas A. p53 regulates epithelial-mesenchymal transition induced by transforming growth factor β. J Cell Physiol. 2013;228:801–13.
Suzuki Y, Haraguchi N, Takahashi H, Uemura M, Nishimura J, Hata T, et al. SSEA-3 as a novel amplifying cancer cell surface marker in colorectal cancers. Int J Oncol. 2013;42:161–7.
De Carvalho DD, You JS, Jones PA. DNA methylation and cellular reprogramming. Trends Cell Biol. 2010;20:609–17.
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Yamamoto M, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131:861–72.
Miura N, Nakamura H, Sato R, Tsukamoto T, Harada T, Takahashi S, et al. Clinical usefulness of serum telomerase reverse transcriptase (hTERT) mRNA and epidermal growth factor receptor (EGFR) mRNA as a novel tumor marker for lung cancer. Cancer Sci. 2006;97:1366–73.
Author contributions
The author(s) have made the following declarations about their contributions: Norimasa Miura conceived and designed the experiments; Yoshitaka Ishihara, Satoshi Tsuno, Taro Yamashita, and Yusuke Endo performed the experiments; Satoshi Tsuno and Norimasa Miura analyzed the data; Satoshi Tsuno contributed reagents/materials/analysis tools; Satoshi Kuwamoto contributed to the pathological diagnosis; and Satoshi Tsuno and Norimasa Miura wrote the manuscript. Junichi Hasegawa provided an environment for this study.
Conflict of interest
The authors declare no conflict of interest.
Funding and support
This work was supported by a Grant-in-Aid for Research for Promoting Technological Seeds B (development type), the Takeda Science Foundation, the Princess Takamatsu Cancer Research Fund, the Adaptable and Seamless Technology Transfer Program through Target-driven R&D (Exploratory Research) of Japan Science and Technology Agency (JST) and the JSPS KAKENHI grant (Grant-in-Aid for Challenging Exploratory Research) number 23659285, which all have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Figure 1 The alteration of the relative 5-hmC percent induced by miR-520d-5p was examined to understand the general DNA methylation levels during the entire culture period. The 520d-NHDF cells at 18–24 weeks were significantly hypermethylated, compared with the parental cells, the mock-NHDF cells or the hiPSCs, indicating that the demethylation that followed the earlier miR-520d-5p transfections seemed to have failed, and cell death approached. **: P < 0.01 by Mann-Whitney U-test.
Supplementary Figure 2 Western blotting for representative proteins was carried out using siETG-NHDF-Ad cells. CD105 and caspase-8 were upregulated in response to siETG transfection, p53 protein levels were elevated a little later after the transfection and SIRT1 tended to be upregulated during senescence. These findings were compatible with the transcriptional expression profiling except for p53 and SIRT1. S: senescence, T: transfection by siETG. The protein level of β-actin is a used as a control.
Supplementary Figure 3 A. HUVECs (left) were transfected with miR-520d-5p, and the two types (accumulated cell groups or spheroid-like cells) of phenotypic changes seen in the transfectants (520d-HUVECs) are shown (the former; 40× magnification in the middle, the latter; 100× on the right). The accumulated cell groups lay scattered around the culture dish. B. Cell markers of vascular endothelial cell precursors (CD34, CD45, and ROBO4) were examined by RT-PCR, and no changes in gene expression were observed in the 520d-HUVECs. The data were normalized to β-actin mRNA expression levels. NS not significant.
Supplementary Figure 4 Relative mRNA expression of target genes (ELAVL2, TEAD1, and GATAD2B) to β-actin by each siRNA transfection into NHDF-Ad. Average level was shown representatively (n = 3). Each siRNA was suppressed the expression of target genes to some extent.
Supplementary Figure 5 The relative 5-hmC levels were measured to understand the general DNA methylation levels in two hepatoma cell lines (left: HLF, right: Huh7) transfected with siETG. The results indicate that siETG induced a similar effect to transfection with miR-520d-5p alone. The 5-hmC levels in the siETG-HLF cells and the siETG-Huh7 cells were similar to the 5-hmC levels in the hiPSCs. siGATAD2B next to siETG in the hepatoma cells was effective for demethylating them, indicating that this transcriptional factor may be implicated in methylation process in well-differentiated hepatoma, unlike in undifferentiated hepatoma. The data were normalized to average 5-hmC levels in the HLF cells (n = 4 or 6).
Supplementary Figure 6 Time-lapse video observations of siETG-NHDF-Ad cells (at 22 weeks) for 2 days showed cell morphology and proliferation (100× magnification). The morphology was very similar to that of the 520d-NHDF-Ad cells.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Ishihara, Y., Tsuno, S., Kuwamoto, S. et al. Hsa-miR-520d Converts Fibroblasts into CD105+ Populations. Drugs R D 14, 253–264 (2014). https://doi.org/10.1007/s40268-014-0064-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40268-014-0064-6