Abstract
Purpose of Review
Microbial infections in chronic wounds can often lead to lower-limb amputation, decrease in quality of life, and increase in mortality rate, and there is an unmet need to distinguish between pathogens and colonisers in these chronic wounds. Hence, identifying the composition of healthy skin microbiota, microbes associated with chronic wound and healing processes, and microbial interactions and host response in healing wounds vs. non-healing wounds can help us in formulating innovative individual-centric treatment protocols.
Recent Findings
This review highlights various metabolites and biomarkers produced by microbes that have been identified to modulate these interactions, particularly those involved in host–microbe and microbe–microbe communication. Further, considering that many skin commensals demonstrate contextual pathogenicity, we provide insights into promising initiatives in the wound microbiome research.
Summary
The skin microbiome is highly diverse and variable, and considering its importance remains to be a hotspot of medical investigations and research to enable us to prevent and treat skin disorders and chronic wound infections. This is especially relevant now considering that non-healing and chronic wounds are highly prevalent, generally affecting lower extremities as seen in diabetic foot ulcers, venous leg ulcers, and pressure ulcers. Pathogenic bacteria are purported to have a key role in deferring healing of wounds. However, the role of skin microflora in wound progression has been a subject of debate. In this review, we discuss biomarkers associated with chronic wound microenvironment along with the relevance of skin microflora and their metabolites in determining the chronicity of wounds.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Human skin protects us from physical, chemical, and immunological risks from the outside world and also hosts a major ecosystem that harbours skin’s indigenous microbiota or the skin microbiome [1]. In all likelihood, the most significant role of this multifunctional organ is to safeguard against infections from invading microbes. Although the skin serves to prevent pathogenic microbes from gaining entry into the host, the presence of hair follicles and other appendages provides the human skin with a surface area of approximately 30 m2 which allows interaction with a diverse array of microbes [2]. The moist, sebaceous, and dry microenvironments of the skin harbour a variety of microbial populations, including bacteria, fungi, viruses, and microeukaryotes [1].
For well over a century, scientists have debated whether microbes on the skin play a definitive role in causing disease or are merely colonisers [3]. The microbial populations of skin protect against foreign microbes via both direct and indirect processes, like the production of antimicrobial compounds (direct) [4] and competitive exclusion (indirect) [5]. However, the role of skin-dwelling commensals varies depending on the species, with certain species alternating between healthy and diseased states. An altered microbiome, together with a break in skin’s protective barrier, increases the risk of infection, progressing to skin-related disorders. Infection has been identified as one of the most important contributors to the development and maintenance of chronic wounds [6].
Chronic wounds are a serious healthcare issue, with diabetic foot ulcers (DFU), pressure ulcers (decubitus ulcers (DU)), venous leg ulcers (VLU), and non-healing surgical wounds being the most prevalent. Chronic wounds are more common in elderly people who have latent medical problems like diabetes, vascular disease, pulmonary disorders, kidney malfunctions, and obesity [7]. Impaired wound healing has also been linked to a weakened immune system, poor nutrition, and prolonged mechanical stress [8]. Chronic wounds are attributed to disturbingly high mortality rates: the 5-year mortality rate of diabetic foot ulcers is 30.5%, which is comparable to the mortality rate linked with cancer (31%) [7]. Chronic non-healing wounds are also accompanied by significant treatment expenses reaching $95 billion in annual healthcare expenses [9]. Unfortunately, effective therapies are still missing, despite the rising frequency and high costs of care [7].
A characteristic aspect of these non-healing wounds is the prevalence of skin commensals in them, associated with varied bacterial communities nestled in an altered microenvironment. Recent investigations have taken recourse to evaluation of markers and indicators such as enzyme activity, volatile molecules, or other metabolites along with sensors for changes in pH, temperature, and odour, in addition to microbiological investigations, to better understand the complex dynamics of chronic wound healing better [10,11,12,13]. This review examines the importance of the human skin microbiome and how these important natural communities are altered in chronic wounds. We also reflect upon the gamut of microbe–host and microbe–microbe interactions that affect the skin and its robustness, including wound healing modulation along with a discussion on a variety of microbiome-derived metabolites that have been identified as important intermediaries in skin microbial populations.
Skin Microbiome
The indigenous microflora of skin as a whole has been linked to the proper establishment of an intact boundary over a lifetime. The interactions between the diverse skin commensals that inhabit this vast plane are either neutral or mutually profitable. For instance, during brief developmental phase in early life, skin commensals and hair follicles work in accordance to facilitate tolerance to skin commensal microbes and maintain skin immune cell homeostasis [14]. The dynamic environment of the skin and its associated microbiome is the most reliable predictors of temporal age, outperforming hallmarks of the gut and oral microbiome [15,16,17]. These commensal communities are not only just considered hitchhikers, but contribute actively to keeping the skin barrier intact. For instance, skin microbiota has been linked to crucial barrier function processes such as regulation of the skin inflammatory response, epidermal differentiation, and augmentation of wound healing [18, 19].
The healthy skin microbiome is heavily influenced by several host-related factors including age, anatomical site, and others (Fig. 1). Studies have shown that the skin microflora of an individual is developed intra-partum, where the mode of maternal delivery has a significant impact on microbial makeup [20, 21]. The colonisation of the skin by microbes is also heavily influenced by anatomical site, as evidenced by diverse microbial populations dwelling in various epidermal topographical niches [22]. A 16S rRNA gene-based study of a healthy human skin microbiome revealed the presence of minimum 19 phyla along with more than 1000 species of bacteria from 20 different skin sites [23•]. Phyla Actinobacteria (52%), Firmicutes (24%), Proteobacteria (17%), and Bacteroidetes (7%) made up the most of epidermal microflora, while Corynebacteria (Actinobacteria), Propionibacteria (Actinobacteria), and Staphylococci (Firmicutes) were the most prevalent genera [23•, 24]. Moisture content and anatomical site both play a role in determining microbial composition. For instance, sebaceous sites like occiput, glabella, alar crease, external auditory canal, retro-auricular crease, and back have the highest microbial burden [23•] and are favoured by Propionibacterium and Staphylococcus, while sites with high moisture content are dominated by Corynebacterium and Staphylococcus. Even though dry regions (like volar forearm, hypothenar palm, and buttock) comprised a higher proportion of beta-proteobacteria, Flavobacteriales, and other Gram negative organisms, it showed high microbial richness and variance overall [24]. Corresponding to epidermal colonisation, recent research shows that the cutaneous microbiome transcends into the sub-epidermal layers of the skin, accompanied by greater abundance but a lower proportion of Proteobacteria (Burkholderiales and Pseudomonadales) and Actinobacteria [25]
While the relevance of anatomical locations and moisture content in determining the human skin microbiome has been clearly established, studies have also shown that genetics and environmental variables including climate play a role in defining the ‘normal’ microbiome [31, 32]. For instance, skin commensals obtained from forearms of Venezuelan subjects (dominance of Staphylococcus and Proteobacteria) varied significantly from those of Americans (primarily Propionibacterium) [33]. Furthermore, besides gender and age, the cutaneous microbiome can also be reliable predictors of whether residents reside in urban or rural location amidst the same metropolitan region [26]. In a similar study, Hospodsky et al. [34] found that hands of women in Tanzania contained more soil-associated microorganisms including members of Rhodobacteraceae and Nocardioidaceae than women in the USA. In another study by Wang et al. [35], the pan-microbiome concept suggests that the microbial assemblage of healthy skin differed between nations and displayed considerable variation in Chinese subjects as opposed to another ethnic group, Pakistanis. The ethnic and environmental variances highlight the need to broaden our existing knowledge on skin microbiome diversity to include a wider range of geographic and cultural communities since variations in skin commensals might significantly contribute to wound healing and therapy.
Skin and Microbial Interactions
The stratum corneum, the outermost layer of the skin, is made up of densely packed, keratinocytes supported by cornified envelopes providing a powerful shield to the host from external influences. Moreover, stratum corneum’s low water content along with hydrolysis of epidermal phospholipids into free fatty acids (FFA) decreases the pH of the skin surface, making it unsuitable for pathogen colonisation [36]. The human skin is also known to release several antimicrobial peptides (AMP) which are effective against a wide range of pathogens, like bacteria, viruses, and various parasites. Primary skin-derived AMPs are cathelicidins and beta-defensins, while several other proteins such as ribonucleases and peptidoglycan recognition proteins with potent antimicrobial activity have also been reported from skin tissue [37].
Rather than living off the host, skin microbial community plays an active part in antimicrobial defence via both indirect and direct methods. Skin microflora, indirectly, works as a competitive barrier against potential infections by colonising the epidermal niches and harnessing the available resources [16]. Furthermore, microbial metabolism products may indirectly boost the skin’s antibacterial potential. Lipase activity, for example, enables the production of FFAs by hydrolysis of sebum lipids in a variety of skin commensals, including Corynebacterium acnes and Staphylococcus epidermidis [38]. Similarly, the nasal commensal Corynebacterium accolens was reported to hinder pathogenic Streptococcus pneumoniae via conversion of cutaneous triacylglycerols to FFAs [39], suggesting that FFAs, synthesised by skin commensals, may have a direct antibacterial effect in addition to lowering skin pH. Through the synthesis of antimicrobial compounds, the local commensal microbial communities also contribute to colonisation resistance. Coagulase negative Staphylococci have shown to suppress the growth of closely-related bacterial species by producing bacteriocins — extremely potent compounds which are heat-resistant and synthesised in ribosomes [40,41,42]. Bacteriocins are reported to be synthesised by various skin commensals. For instance, S. epidermidis are reported to generate a variety of bacteriocins and phenol-soluble modulins that selectively eradicate pathogenic bacteria however inactive towards S. epidermidis [43]. The antibacterial arsenal of S. epidermidis includes the serine protease Esp that have been reported to deter Staphylococcus aureus in human nasal carriage [43]. Likewise, Staphylococcus lugdunensis produces lugdunin, a non-ribosomal peptide, which was shown to be efficacious against a variety of skin infections and particularly effective in decreasing the nasal carriage of S. aureus [44] (Fig. 2).
The production of antibiotics by skin commensals may also be considered as one of the major defence strategies adopted against infectious bacterial pathogens. Mupirocin, a topical antibiotic that is used to treat staphylococcal or streptococcal skin infections [48], is produced by Pseudomonas fluorescens, a bacterium which is occasionally discovered among cutaneous commensal populations [49]. Interestingly, mupirocin resistance is inherent in certain skin microbes, like Micrococcus spp. and Corynebacterium spp. [50]. This beneficial property of mupirocin enables the targeted treatment of harmful pathogens while conserving the local microbiota. The commensal microbes that live in the human body not only strive for niche possession, but also defend against invasive pathogens and hence have developed multiple mechanisms to overcome microbial competition, most of which remain largely unexplored.
Chronic Wounds
Venous, diabetic, and pressure ulcers are some of the most frequent chronic wounds [7]. Venous leg ulcers (VLUs) develop when there is venous dysfunction. VLUs are most common among chronic non-healing wounds contributing 60–80% of cases and are caused by a variety of factors. Inept veins or valves, as well as reduced muscle performance, can result in inadequate calf muscle pump activity, which can cause ambulatory venous hypertension leading to local venous dilation and pooling [51]. This results in the entrapment of leucocytes which may release proteolytic enzymes damaging tissues. Venous pooling also causes inter-endothelial pore expansion and the deposition of fibrin and other macromolecules, which trap growth factors and render them useless for wound healing. Bed-bound, wheelchair-bound, and neurologically disabled individuals are more likely to develop arterial, diabetic, and pressure ulcers, which is often followed by necrosis of the affected tissue. Lower-extremity ulcers, which affect roughly 23% of people with autoimmune illness, can be a late debilitating consequence of connective tissue disorders [52]. The pathogenic processes that create arterial, diabetic, and pressure ulcers are mostly ischemic in nature, resulting from several factors such as lymphatic obstruction, reperfusion, and cell deformation. Tissue inflammation, in all its manifestations, is a critical process implicated in the aetiology of ulcer development.
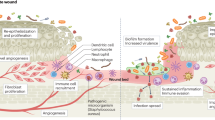
Wound healing and repair are a complicated and systematic process that is closely regulated by various types of cells which are, in turn, regulated by a number of growth factors, cytokines, and chemokines. Wounds become chronic if this process is hampered in any way and does not allow skin barrier to reseal the open wound. Chronic wounds of all forms have been reported to show fibrosis along with increased rate of keratinocyte proliferation and lack of migration. In contrast to the typical wound repair, chronic wounds have been reported to show hampered angiogenesis, recruitment and impeded activation of stem cells as well as extracellular matrix remodelling, with persistent inflammation [45•, 53,54,55]. Wounds allow bacteria from the skin microflora and those from the environment, to get access to the underlying cells and tissues to colonise and develop in optimum circumstances [56]. During the normal cutaneous wound healing process, commensal bacteria interact with skin cells, which help in modulating the innate immune response [57]. Chronic, hard-to-heal wounds generally harbour polymicrobial biofilms that encourage pathogenic expansion in the wound bed and impeded wound healing [58•]. Biofilm formation has been reported in 60% of chronic wounds as opposed to just 6% of acute wounds in a clinical examination of wound specimens (n = 50) from adult patients [58•]. Furthermore, Wolcott et al. [58•] determined that Pseudomonas was the most dominant genus in chronic wound biofilms as well as the most prevalent bacterial genus observed in monomicrobial biofilms. Microbial clustering on wound surface results in biofilms, which are enclosed in an exopolymeric material made up of polysaccharides, lipids, and protein [59]. Under these conditions, microbes tend to suitably alter their reproductive rate and metabolic activities and utilise quorum sensing to relay changes in dynamics of their population density via organic signalling molecules synthesised by them. Biofilms trigger the host immune responses on a constant basis, thereby delaying the wound from entering the proliferative phase, hindering the healing of the wound [59].
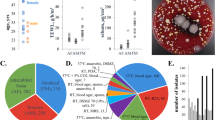
The composition of the human skin microbiome changes over time, while the presence and number of bacteria in wounded skin vary according to wound type. The three major phyla, namely, Firmicutes, Proteobacteria, and Actinobacteria, which are found in healthy skin are also found in pressure ulcers [60]. In their clinical observation study of chronic wounds from 2963 subjects, Wolcott et al. [58•] found that Staphylococcus was the most common and these chronic wounds (DFU, DU, VLU, and non-healing surgical wounds) were predominated by S. aureus and S. epidermidis. Furthermore, despite the fact that microbial heterogeneity was unaffected by wound type, the most prevalent genera were noted to be S. epidermidis, whereas Pseudomonas aeruginosa was found to be relatively abundant in the examined wounds, indicating biofilm development. This investigation corroborated the previous findings of James et al. [61] who using culture and molecular techniques established Gram-positive cocci to be the most predominant bacterial population in chronic wound specimens, whereas Gram-negatives were shown to be involved in the production of biofilms in chronic wounds with the predominance of Staphylococcus and Pseudomonas in chronic wounds. Despite the fact that chronic wounds are normally exposed to high amounts of oxygenation, anaerobic bacteria have been found to be more prevalent in chronic wounds than acute wounds, with Finegoldia, Prevotella, Peptoniphilus, Peptostreptococcus, and Anaerococcus as regular components of chronic wound microbiome (Table 1).
With respect to species diversity, DFUs were found to be considerably less diverse than unwounded skin samples from diabetic subjects as well as control skin samples, in terms of three different alpha diversity estimates, namely, observed species richness, Chao 1 estimator, and Shannon index [70]. Furthermore, permutational multivariate analysis of variance demonstrated that the beta diversity of bacterial populations in control skin was considerably different from that of DFUs. While Corynebacterium was reported to be the most frequent genus in diabetic foot associated ulcers and osteomyelitis (n = 20) by Johani et al. [71], Gardiner et al. [70] reported Staphylococcus as the most dominant genus in chronic DFUs, followed by Acinetobacter and Corynebacterium. However, using NGS approach, Kalan et al. [72] reported that certain S. aureus strains were exclusively present in unhealed wounds, while few other generalist strains were more broadly distributed across wound types suggesting an association between strain type and adverse clinical outcome of the wound in DFU subjects. Further, a similar result was obtained in the type 2 diabetic mouse model, with wounds inoculated with S. aureus strains showing poor outcomes with slow healing rate and poor re-epithelialisation [72]. Moreover, microbial community transition dynamics can significantly affect wound healing rates. For example, Loesche et al. [62] classified the microbial communities associated with DFUs into four major community types, and the temporal transitions between these types dictated the wound healing rates. Further, treatment with systemic antibiotics could only destabilise the microbiota but did not significantly alter the relative abundance of specific bacterial taxa.
Chronic wounds are polymicrobial in nature, and hence, understanding the dynamic and complex interplay between pathogens and skin commensals, rather than just the presence or absence of particular bacteria, is probably more helpful in terms of understanding the wound dynamics and progression [73]. The pathogenic impact of anaerobes, for instance, might be exacerbated by the presence of aerobes because they utilise oxygen, causing tissue hypoxia and favouring anaerobe proliferation [74]. This may be considered a symbiotic connection, where two or more species cooperate to enhance virulence and hinder healing [75]. It is now well established that the two of the most prevalent pathogens in infections of chronic wounds are P. aeruginosa and S. aureus. They are usually detected together, and combined infections are more harmful than their monomicrobial infections [76, 77]. In addition, P. aeruginosa and S. aureus have been demonstrated to have higher antibiotic resistance when cultured together in a wound model [76]. Also, Bacteroides fragilis has been identified as the most common anaerobic bacterium isolated in DFUs in various investigations [78], and it plays an essential role in the composition of microbial populations and biological interactions. In an interaction study, Mastropaolo et al. [79] compared the synergistic interactions among partners in polymicrobial wound infections such as B. fragilis, C. perfringens, and E. coli in an obese diabetic mouse model and showed strong synergism between B. fragilis and E. coli but not C. perfringens.
Overall, there is mounting evidence that polymicrobial interactions may promote the pathogenic capacity of other microbes or reduce their virulence and hence have a considerable influence on the degree of severity as well as the progression of wound infection. Hence, it is critical to investigate beyond the mere presence or absence of microbes in such non-healing wounds but a further attempt to understand possible microbial interconnections.
Microbial Interactions in Chronic Wounds
Chronic wounds are usually polymicrobial in nature. Such infections are accompanied by persistent biofilms and show higher resistance to antibiotic therapy in comparison to monomicrobial infections [80]. In one of the earliest attempts to demonstrate polymicrobial infections, guinea pigs were co-infected with E. coli and B. fragilis, and after 7-day post-infection, each species showed an increase in bacterial CFU by more than 100 times in comparison to monomicrobial infection, as well as increase in inflammation and purulence, which was reported suggestive of poor healing and indicated pathogenic synergy [81]. Likewise, co-infection of ulcers with P. aeruginosa and S. aureus has also been linked to the persistent non-healing condition [77]. However, certain investigations on the co-infection of P. aeruginosa and S. aureus have also revealed that P. aeruginosa inhibits the proliferation of S. aureus [46, 82]. A study involving mouse wound excisional model revealed that detection of peptidoglycan by P. aeruginosa is critical for its competitive edge in the vicinity of other Gram-positive taxa. The presence of N-acetylglucosamine or peptidoglycan fragments induces secretion of elastase and pyocyanin by P. aeruginosa through the proposed two-component response regulator PA0601 [46]. When P. aeruginosa and S. aureus were co-infected at the same time in the wound, P. aeruginosa exceeded S. aureus by more than 100-fold at 4 dpi. The results showed that sensing of S. aureus peptidoglycan by P. aeruginosa triggered the release of lytic virulence factors allowing P. aeruginosa to overtake S. aureus in co-infected wounds, although P. aeruginosa deletion mutants of PA0601 (involved in peptidoglycan sensing) were unable to outnumber S. aureus in the same way [46]. During co-infection with P. aeruginosa in porcine wounds, virulence factor protein A of S. aureus was drastically under-expressed by more than threefold at 2 and 4 dpi, whereas alpha-hemolysin and Panton-Valentine leukocidin (PVL) that cause necrosis in wounds were significantly upregulated at 4 dpi. Furthermore, compared to their monomicrobial infection, P. aeruginosa and S. aureus co-infection caused downregulation of pro-inflammatory cytokines, including IL-1 alpha, IL-1 beta, IL-6, and IL-8 [45•]. These findings reveal that negative interactions between microbes can lead to changes in the expression and activity of multiple virulence factors along with host immune response regulators, giving a competitive advantage to a few select species over others in a polymicrobial infection (Fig. 2).
Polymicrobial infections compromise the integrity of skin’s surface and can exacerbate the chronicity of wounds with impaired healing. In a porcine burn wound model, co-infected with P. aeruginosa and Acinetobacter baumannii, the mammalian tight junction proteins zona-occludens-1 and zona-occludens-2 (ZO-1, ZO-2) were significantly downregulated as opposed to non-infected controls, leading to a functionally compromised epidermis [83]. In another study of porcine wound infection model, upon P. aeruginosa and S. aureus coinfections, there was a marked decrease in wound re-epithelisation due to inhibition of keratinocyte growth factor 1, compared to monomicrobial infections [45•]. These findings also show that polymicrobial infections with biofilms hinder wound closure and may make the host more susceptible to other opportunistic infections. The microorganisms in chronic wounds and the neighbouring healthy skin are often not quite easily distinguishable [70, 84].
Commensals have traditionally been overlooked in acute wounds; however, there is increasing support that they may play a bigger part in the pathogenesis and prognosis of chronic wounds. Corynebacterium spp., for instance, releases a secretory component that suppresses the agr regulatory system in the pathogen S. aureus and its virulence factors [85], thereby preventing colonisation and infection on healthy skin. However, C. striatum has been identified as an emerging multidrug-resistant wound pathogen [86], causing a high proliferation rate in the epidermal layer in diabetic wound model in mice [72], where competition between C. striatum and S. aureus may result in additional damage to wound tissue. S. epidermidis, an identified skin commensal, is also known to play a role in wound worsening; yet, certain strains of S. epidermidis enhance the ability of immune system to speed up the healing of the wound [87]. In a comparative examination of diabetic mouse models, db/db diabetic mice wounds had a larger proportion of Staphylococci than wounds of db/ + diabetic mice, suggesting a negative correlation to wound healing ability [88]. Furthermore, in diabetic mice wounds with the predominance of Staphylococci, genes involved with an acute inflammatory response were significantly upregulated, and the persistent inflammation was attributed to the chronicity of the wounds [89].
Our present knowledge of how the local skin microbiome impacts the predisposition for wound infections is limited. Future research aimed at studying the potential of local microbiota to manipulate the host response and how their interactions with other opportunistic pathogen(s) during infection would affect wound healing efficacy can provide significant clues on managing chronic wounds. Simply put, commensal activity is crucial in a wound environment, but its ecological importance is complicated and should not be overlooked. Identifying the secreted components generated by these organisms would be a significant step forward in understanding how they build up microbial populations, communicate with other microbial communities, and regulate dialogue with the host cell.
Conclusion
The skin is a unique and important part of the human body that shields and protects us from our surroundings. It also offers a variety of habitats with unique microenvironments that contribute to a diverse array of microbial communities taking residing there. While tremendous strides have been made in recognising the significance of the skin microbiome in the establishment of host immune responses and microbial resistance, there is still much more to learn. The majority of current research has been concentrated on prominent skin microbes belonging to the genera Staphylococcus and Cutibacterium, which include both helpful and harmful species. However, other major genera such as Corynebacterium, Kocuria, Micrococcus, and Brevibacterium (belonging to the phylum Actinobacteria which is also a predominant group associated with human skin) have not been extensively researched. All of the mentioned genera are known to cause infection, but they are mainly understudied, and their effects on the skin and other skin microbiota are unclear. Even well-studied species like S. epidermidis are less explored in the context of molecular processes that drive microbe–microbe interactions and immune responses. Even though it has been well established that fungal colonisers contribute heavily to skin microbial populations and in regulating skin health, their interactions and immunomodulatory roles are a little-known component of the vast skin microbiome. The potential of microbes to adapt to changes in the wound microenvironment leads to virulence, exacerbation of wound, and delay in wound healing. To provide effective therapies for chronic wounds, it is critical to examine the function of microbes at the cellular and molecular levels, not only focusing on bacteria but also on other natural flora such as viruses and fungus. A better knowledge of the interactions between skin cells, normal flora, and their environment can help us in distinguishing between normal and pathological healing and aid researchers in developing better therapies or strategies for effectively eliminating the pathogenic microorganisms found in chronic wounds. Understanding the molecular markers and defence mechanisms that regulate the fine line between the commensal and pathogenic nature of microbes will help develop innovative therapies and even direct towards engineering a ‘healthy’ skin microbiome. Determining critical microbial elements and metabolomic patterns that could be employed as diagnostic biomarkers to determine clinical populations at risk of wound infection can lead to non-invasive diagnostic and therapeutic procedures that are more precise.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance
Callewaert C, Ravard Helffer K, Lebaron P. Skin microbiome and its interplay with the environment. Am J Clin Dermatol. 2020;21:4–11. https://doi.org/10.1007/s40257-020-00551-x.
Gallo RL. Human skin is the largest epithelial surface for interaction with microbes. J Invest Dermatol. 2017;137:1213–4. https://doi.org/10.1016/j.jid.2016.11.045.
Chen YE, Fischbach MA, Belkaid Y. Skin microbiota–host interactions. Nature. 2018;553:427–36. https://doi.org/10.1038/nature25177.
Nakatsuji T, Chen TH, Narala S, et al. Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Sci Transl Med. 2017;9:eaah4680. https://doi.org/10.1126/scitranslmed.aah4680.
SanMiguel A, Grice EA. Interactions between host factors and the skin microbiome. Cell Mol Life Sci. 2015;72(1499–515):10. https://doi.org/10.1007/s00018-014-1812-z.
Tipton CD, Sanford NE, Everett JA, et al. Chronic wound microbiome colonization on mouse model following cryogenic preservation. PLoS One. 2019;14:e0221565. https://doi.org/10.1371/journal.pone.0221565.
Sen CK. Human wound and its burden: updated 2020 compendium of estimates. Adv Wound Care. 2021;10:281–92. https://doi.org/10.1089/wound.2021.0026.
Fayne RA, Borda LJ, Egger AN, Tomic-Canic M. The potential impact of social genomics on wound healing. Adv Wound Care. 2020;9:325–31. https://doi.org/10.1089/wound.2019.1095.
Nussbaum SR, Carter MJ, Fife CE, et al. An economic evaluation of the impact, cost, and Medicare policy implications of chronic nonhealing wounds. Value Health. 2018;21:27–32. https://doi.org/10.1016/j.jval.2017.07.007.
Sharifuzzaman M, Chhetry A, Zahed MA, et al. Smart bandage with integrated multifunctional sensors based on MXene-functionalized porous graphene scaffold for chronic wound care management. Biosens Bioelectron. 2020;169: 112637. https://doi.org/10.1016/j.bios.2020.112637.
Simoska O, Duay J, Stevenson KJ. Electrochemical detection of multianalyte biomarkers in wound healing efficacy. ACS Sens. 2020;5:3547–57. https://doi.org/10.1021/acssensors.0c01697.
Sun T, He J, Qian S, et al. Collaborative detection for wound infections using electronic nose and FAIMS technology based on a rat wound model. Sens Actuators B: Chem. 2020;320: 128595. https://doi.org/10.1016/j.snb.2020.128595.
Iglesias-Mayor A, Amor-Gutiérrez O, Toyos-Rodríguez C, Bassegoda A, Tzanov T, de la Escosura-Muñiz A. Electrical monitoring of infection biomarkers in chronic wounds using nanochannels. Biosens Bioelectron. 2022;209: 114243. https://doi.org/10.1016/j.bios.2022.114243.
Scharschmidt TC, Vasquez KS, Pauli ML, et al. Commensal microbes and hair follicle morphogenesis coordinately drive Treg migration into neonatal skin. Cell Host Microbe. 2017;21:467–77. https://doi.org/10.1016/j.chom.2017.03.001.
Kim HJ, Kim JJ, Myeong NR, et al. Segregation of age-related skin microbiome characteristics by functionality. Sci Rep. 2019;9:1–11. https://doi.org/10.1038/s41598-019-53266-3.
Dimitriu PA, Iker B, Malik K, Leung H, Mohn WW, Hillebrand GG. New insights into the intrinsic and extrinsic factors that shape the human skin microbiome. MBio. 2019;10:e00839-e919. https://doi.org/10.1128/mBio.00839-19.
Huang S, Haiminen N, Carrieri AP, et al. Human skin, oral, and gut microbiomes predict chronological age. Msystems. 2020;5:e00630-e719. https://doi.org/10.1128/mSystems.00630-19.
Linehan JL, Harrison OJ, Han SJ, et al. Non-classical immunity controls microbiota impact on skin immunity and tissue repair. Cell. 2018;172:784–96. https://doi.org/10.1016/j.cell.2017.12.033.
Constantinides MG, Link VM, Tamoutounour S, et al. MAIT cells are imprinted by the microbiota in early life and promote tissue repair. Science. 2019;366:peaax6624. https://doi.org/10.1126/science.aax6624.
Dominguez-Bello MG, Costello EK, Contreras M, et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci USA. 2010;107:11971–5. https://doi.org/10.1073/pnas.1002601107.
Zhu T, Liu X, Kong FQ, et al. Age and mothers: potent influences of children’s skin microbiota. J Invest Dermatol. 2019;139:2497-5.e6. https://doi.org/10.1016/j.jid.2019.05.018.
Oh J, Byrd AL, Deming C, Conlan S, Kong HH, Segre JA. Biogeography and individuality shape function in the human skin metagenome. Nature. 2014;514:59–64. https://doi.org/10.1038/nature13786.
Grice EA, Kong HH, Conlan S, et al. Topographical and temporal diversity of the human skin microbiome. Science. 2009;324:1190–2. https://doi.org/10.1126/science.1171700 (16S rRNA sequence analysis of 20 distinct skin sites of healthy humans. 16S rRNA sequence analysis of 20 distinct skin sites of healthy humans (n=20) demonstrating that physiologically related regions have similar bacterial populations.
Oh J, Byrd AL, Park M, Kong HH, Segre JA. NISC Comparative Sequencing Program. Temporal stability of the human skin microbiome. Cell. 2016;165:854–66. https://doi.org/10.1016/j.cell.2016.04.008.
Nakatsuji T, Chiang HI, Jiang SB, Nagarajan H, Zengler K, Gallo RL. The microbiome extends to subepidermal compartments of normal skin. Nat Commun. 2013;4:1–8. https://doi.org/10.1038/ncomms2441.
Ying S, Zeng DN, Chi L, et al. The influence of age and gender on skin-associated microbial communities in urban and rural human populations. PLoS One. 2015;10: e0141842. https://doi.org/10.1371/journal.pone.0141842.
Chu DM, Ma J, Prince AL, Antony KM, Seferovic MD, Aagaard KM. Maturation of the infant microbiome community structure and function across multiple body sites and in relation to mode of delivery. Nat Med. 2017;23:314–26. https://doi.org/10.1038/nm.4272.
Oh J, Conlan S, Polley EC, Segre JA, Kong HH. Shifts in human skin and nares microbiota of healthy children and adults. Genome Med. 2012;4:1–11. https://doi.org/10.1186/gm378.
Capone KA, Dowd SE, Stamatas GN, Nikolovski J. Diversity of the human skin microbiome early in life. J Invest Dermatol. 2011;131:2026–32. https://doi.org/10.1038/jid.2011.168.
Luna PC. Skin microbiome as years go by. Am J Clin Dermatol. 2020;21:12–7. https://doi.org/10.1007/s40257-020-00549-5.
Gupta VK, Paul S, Dutta C. Geography, ethnicity or subsistence-specific variations in human microbiome composition and diversity. Front Microbiol. 2017;1162. https://doi.org/10.3389/fmicb.2017.01162
Wang Y, Yu Q, Zhou R, Feng T, Hilal MG, Li H. Nationality and body location alter human skin microbiome. Appl Microbiol Biotechnol. 2021;105:5241–56. https://doi.org/10.1007/s00253-021-11387-8.
Blaser MJ, Dominguez-Bello MG, Contreras M, et al. Distinct cutaneous bacterial assemblages in a sampling of South American Amerindians and US residents. ISME J. 2013;7:85–95. https://doi.org/10.1038/ismej.2012.81.
Hospodsky D, Pickering AJ, Julian TR. Hand bacterial communities vary across two different human populations. Microbiology. 2014;160:1144–52. https://doi.org/10.1099/mic.0.075390-0.
Wang X, Zhou H, Chen D, et al. Whole-genome sequencing reveals a prolonged and persistent intrahospital transmission of Corynebacterium striatum, an emerging multidrug-resistant pathogen. J Clin Microbiol. 2019;57:e00683-e719. https://doi.org/10.1128/JCM.00683-19.
Swaney MH, Kalan LR. Living in your skin: microbes, molecules, and mechanisms. Infect Immun. 2021;89:e00695-e720. https://doi.org/10.1128/IAI.00695-20.
Nguyen AV, Soulika AM. The dynamics of the skin’s immune system. Int J Mol Sci. 2019;20:1811. https://doi.org/10.3390/ijms20081811.
Jia Y, Gan Y, He C, Chen Z, Zhou C. The mechanism of skin lipids influencing skin status. J Dermatol Sci. 2018;89:112–9. https://doi.org/10.1016/j.jdermsci.2017.11.006.
Bomar L, Brugger SD, Yost BH, Davies SS, Lemon KP. Corynebacterium accolens releases antipneumococcal free fatty acids from human nostril and skin surface triacylglycerols. MBio. 2016;7:e01725-e1815. https://doi.org/10.1128/mBio.01725-15.
Lynch D, O’Connor PM, Cotter PD, Hill C, Field D, Begley M. Identification and characterisation of capidermicin, a novel bacteriocin produced by Staphylococcus capitis. PLoS One. 2019;14: e0223541. https://doi.org/10.1371/journal.pone.0223541.
O’Sullivan JN, Rea MC, O’Connor PM, Hill C, Ross RP. Human skin microbiota is a rich source of bacteriocin-producing staphylococci that kill human pathogens. FEMS Microbiol Ecol. 2019;95:fiy241. https://doi.org/10.1093/femsec/fiy241.
O’Sullivan JN, O’Connor PM, Rea MC. Nisin J, a novel natural nisin variant, is produced by Staphylococcus capitis sourced from the human skin microbiota. J Bacteriol. 2020;202:e00639-e719. https://doi.org/10.1128/JB.00639-19.
Newstead LL, Varjonen K, Nuttall T, Paterson GK. Staphylococcal-produced bacteriocins and antimicrobial peptides: their potential as alternative treatments for Staphylococcus aureus infections. Antibiotics. 2020;9:40. https://doi.org/10.3390/antibiotics9020040.
Zipperer A, Konnerth MC, Laux C, et al. Human commensals producing a novel antibiotic impair pathogen colonization. Nature. 2016;535:511–6. https://doi.org/10.1038/nature18634.
Pastar I, Nusbaum AG, Gil J, et al. Interactions of methicillin resistant Staphylococcus aureus USA300 and Pseudomonas aeruginosa in polymicrobial wound infection. PLoS One. 2013;8: e56846. https://doi.org/10.1371/journal.pone.0056846. This study highlights the importance of bacterial interactions in multi-species wound infections in a porcine wound model.
Korgaonkar A, Trivedi U, Rumbaugh KP, Whiteley M. Community surveillance enhances Pseudomonas aeruginosa virulence during polymicrobial infection. Proc Natl Acad Sci USA. 2013;110:1059–64. https://doi.org/10.1073/pnas.1214550110.
Byrd AL, Belkaid Y, Segre JA. The human skin microbiome. Nat Rev Microbiol. 2018;16:143–55. https://doi.org/10.1038/nrmicro.2017.157.
Khoshnood S, Heidary M, Asadi A, et al. A review on mechanism of action, resistance, synergism, and clinical implications of mupirocin against Staphylococcus aureus. Biomed Pharmacother. 2019;109:1809–18. https://doi.org/10.1016/j.biopha.2018.10.131.
Huang P, Yue SJ, Cai YY, et al. rpeA, a global regulator involved in mupirocin biosynthesis in Pseudomonas fluorescens NCIMB 10586. Appl Microbiol Biotechnol. 2021;105:9309–19. https://doi.org/10.1007/s00253-021-11683-3.
Thomas CM, Hothersall J, Willis CL, Simpson TJ. Resistance to and synthesis of the antibiotic mupirocin. Nat Rev Microbiol. 2010;8:281–9. https://doi.org/10.1038/nrmicro2278.
Ortega MA, Fraile-Martínez O, García-Montero C, et al. Understanding chronic venous disease: a critical overview of its pathophysiology and medical management. J Clin Med. 2021;10:3239. https://doi.org/10.3390/jcm10153239.
Shanmugam VK, Angra D, Rahimi H, McNish S. Vasculitic and autoimmune wounds. J Vasc Surg Venous Lymphat Disord. 2017;5:280–92. https://doi.org/10.1016/j.jvsv.2016.09.006.
Ramirez HA, Pastar I, Jozic I, et al. Staphylococcus aureus triggers induction of miR-15B-5P to diminish DNA repair and deregulate inflammatory response in diabetic foot ulcers. J Invest Dermatol. 2018;138:1187–96. https://doi.org/10.1016/j.jid.2017.11.038.
Stone RC, Stojadinovic O, Rosa AM, et al. A bioengineered living cell construct activates an acute wound healing response in venous leg ulcers. Sci Transl Med. 2017;9:eaaf8611. https://doi.org/10.1126/scitranslmed.aaf8611.
Stone RC, Stojadinovic O, Sawaya AP, et al. A bioengineered living cell construct activates metallothionein/zinc/MMP8 and inhibits TGFβ to stimulate remodeling of fibrotic venous leg ulcers. Wound Repair Regen. 2020;28:164–76. https://doi.org/10.1111/wrr.12778.
Tomic-Canic M, Burgess JL, O’Neill KE, Strbo N, Pastar I. Skin microbiota and its interplay with wound healing. Am J Clin Dermatol. 2020;21:36–43. https://doi.org/10.1007/s40257-020-00536-w.
Harrison OJ, Linehan JL, Shih HY, et al. Commensal-specific T cell plasticity promotes rapid tissue adaptation to injury. Science. 2019;363:eaat6280. https://doi.org/10.1126/science.aat6280.
Wolcott RD, Hanson JD, Rees EJ, et al. Analysis of the chronic wound microbiota of 2,963 patients by 16S rDNA pyrosequencing. Wound Repair Regen. 2016;24:163–74. https://doi.org/10.1111/wrr.12370. One of the largest cohort study with respect to chronic wounds where bacterial communities in 2963 various chronic wounds were analysed using 16S rDNA pyrosequencing.
Wu YK, Cheng NC, Cheng CM. Biofilms in chronic wounds: pathogenesis and diagnosis. Trends Biotechnol. 2019;37:505–17. https://doi.org/10.1016/j.tibtech.2018.10.011.
Ammons MCB, Morrissey K, Tripet BP, et al. Biochemical association of metabolic profile and microbiome in chronic pressure ulcer wounds. PLoS One. 2015;10: e0126735. https://doi.org/10.1371/journal.pone.0126735.
James GA, Swogger E, Wolcott R, et al. Biofilms in chronic wounds. Wound Repair Regen. 2008;16:37–44. https://doi.org/10.1111/j.1524-475X.2007.00321.x.
Loesche M, Gardner SE, Kalan L, et al. Temporal stability in chronic wound microbiota is associated with poor healing. J Invest Dermatol. 2017;137:237–44. https://doi.org/10.1016/j.jid.2016.08.009.
Smith DM, Snow DE, Rees E, et al. Evaluation of the bacterial diversity of pressure ulcers using bTEFAP pyrosequencing. BMC Med Genomics. 2010;3:1–2. https://doi.org/10.1186/1755-8794-3-41.
Dunyach-Remy C, Salipante F, Lavigne JP, et al. Pressure ulcers microbiota dynamics and wound evolution. Sci Rep. 2021;11:1–13. https://doi.org/10.1038/s41598-021-98073-x.
Sloan TJ, Turton JC, Tyson J, et al. Examining diabetic heel ulcers through an ecological lens: microbial community dynamics associated with healing and infection. J Med Microbiol. 2019;68:230–40. https://doi.org/10.1099/jmm.0.000907.
Travis J, Malone M, Hu H, et al. The microbiome of diabetic foot ulcers: a comparison of swab and tissue biopsy wound sampling techniques using 16S rRNA gene sequencing. BMC Microbiol. 2020;20:1–14. https://doi.org/10.1186/s12866-020-01843-2.
Davies CE, Hill KE, Newcombe RG, et al. A prospective study of the microbiology of chronic venous leg ulcers to reevaluate the clinical predictive value of tissue biopsies and swabs. Wound Repair Regen. 2007;15:17–22.
Wolcott RD, Gontcharova V, Sun Y, Dowd SE. Evaluation of the bacterial diversity among and within individual venous leg ulcers using bacterial tag-encoded FLX and titanium amplicon pyrosequencing and metagenomic approaches. BMC Microbiol. 2009;9:1–11. https://doi.org/10.1186/1471-2180-9-226.
Moore K, Hall V, Paull A, et al. Surface bacteriology of venous leg ulcers and healing outcome. J Clin Pathol. 2010;63:830–4. https://doi.org/10.1136/jcp.2010.077032.
Gardiner M, Vicaretti M, Sparks J, et al. A longitudinal study of the diabetic skin and wound microbiome. PeerJ. 2017;5: e3543. https://doi.org/10.7717/peerj.3543.
Johani K, Fritz BG, Bjarnsholt T, et al. Understanding the microbiome of diabetic foot osteomyelitis: insights from molecular and microscopic approaches. Clin Microbiol Infect. 2019;25:332–9. https://doi.org/10.1016/j.cmi.2018.04.036.
Kalan LR, Meisel JS, Loesche MA, et al. Strain-and species-level variation in the microbiome of diabetic wounds is associated with clinical outcomes and therapeutic efficacy. Cell Host Microbe. 2019;25:641–55. https://doi.org/10.1016/j.chom.2019.03.006.
Suryaletha K, John J, Radhakrishnan MP, George S, Thomas S. Metataxonomic approach to decipher the polymicrobial burden in diabetic foot ulcer and its biofilm mode of infection. Int Wound J. 2018;15:473–81. https://doi.org/10.1111/iwj.12888.
Park JU, Oh B, Lee JP, Choi MH, Lee MJ, Kim BS. Influence of microbiota on diabetic foot wound in comparison with adjacent normal skin based on the clinical features. Biomed Res Int. 2019;2019:7459236. https://doi.org/10.1155/2019/7459236.
Ibberson CB, Whiteley M. The social life of microbes in chronic infection. Curr Opin Microbiol. 2020;53:44–50. https://doi.org/10.1016/j.mib.2020.02.003.
DeLeon S, Clinton A, Fowler H, Everett J, Horswill AR, Rumbaugh KP. Synergistic interactions of Pseudomonas aeruginosa and Staphylococcus aureus in an in vitro wound model. Infect Immun. 2014;82:4718–28. https://doi.org/10.1128/IAI.02198-14.
Alves PM, Al-Badi E, Withycombe C, Jones PM, Purdy KJ, Maddocks SE. Interaction between Staphylococcus aureus and Pseudomonas aeruginosa is beneficial for colonisation and pathogenicity in a mixed biofilm. Pathog Dis. 2018;76:fty003. https://doi.org/10.1093/femspd/fty003.
Sadeghpour-Heravi F, Zakrzewski M, Vickery K, Armstrong DG, Hu H. Bacterial diversity of diabetic foot ulcers: current status and future prospectives. J Clin Med. 2019;8:1935. https://doi.org/10.3390/jcm8111935.
Mastropaolo MD, Evans NP, Byrnes MK, Stevens AM, Robertson JL, Melville SB. Synergy in polymicrobial infections in a mouse model of type 2 diabetes. Infect Immun. 2005;73:6055–63. https://doi.org/10.1128/IAI.73.9.6055-6063.2005.
Orazi G, O’Toole GA. “It takes a village”: mechanisms underlying antimicrobial recalcitrance of polymicrobial biofilms. J Bacteriol. 2019;202:e00530-e619. https://doi.org/10.1128/JB.00530-19.
Kelly MJ. The quantitative and histological demonstration of pathogenic synergy between Escherichia coli and Bacteroides fragilis in guinea-pig wounds. J Med Microbiol. 1978;11:513–23. https://doi.org/10.1099/00222615-11-4-513.
Orazi G, Ruoff KL, O’Toole GA. Pseudomonas aeruginosa increases the sensitivity of biofilm-grown Staphylococcus aureus to membrane-targeting antiseptics and antibiotics. MBio. 2019;10:01501–19. https://doi.org/10.1128/mBio.01501-19.
Roy S, Elgharably H, Sinha M, et al. Mixed-species biofilm compromises wound healing by disrupting epidermal barrier function. J Pathol. 2014;233:331–43. https://doi.org/10.1002/path.4360.
Verbanic S, Shen Y, Lee J, Deacon JM, Chen IA. Microbial predictors of healing and short-term effect of debridement on the microbiome of chronic wounds. NPJ Biofilms Microbiomes. 2020;6:1–11. https://doi.org/10.1038/s41522-020-0130-5.
Hardy BL, Dickey SW, Plaut RD, et al. Corynebacterium pseudodiphtheriticum exploits Staphylococcus aureus virulence components in a novel polymicrobial defense strategy. MBio. 2019;10(1):e02491-e2518. https://doi.org/10.1128/mBio.02491-18.
Wang X, Zhou H, Chen D, et al. Whole-genome sequencing reveals a prolonged and persistent intrahospital transmission of Corynebacterium striatum, an emerging multidrug-resistant pathogen. J Clin Microbiol. 2019;57:e00683-e719. https://doi.org/10.1128/JCM.00683-19.
Brown MM, Horswill AR. Staphylococcus epidermidis—Skin friend or foe? PLoS Pathog. 2020;16: e1009026. https://doi.org/10.1371/journal.ppat.1009026.
Grice EA, Snitkin ES, Yockey LJ. Longitudinal shift in diabetic wound microbiota correlates with prolonged skin defense response. Proc Natl Acad Sci USA. 2010;107:14799–804. https://doi.org/10.1073/pnas.1004204107.
Cohen TS, Takahashi V, Bonnell J, et al. Staphylococcus aureus drives expansion of low-density neutrophils in diabetic mice. J Clin Invest. 2019;129:2133–44. https://doi.org/10.1172/JCI126938.
Acknowledgements
We thank the Science and Engineering Board (SERB), Department of Science and Technology (DST), DBT BUILDER – Interdisciplinary Life Science Programme for Advance Research and Education (DB-ILSPARE), TIFAC-CORE, India, Indo-German Science and Technology Centre (IGSTC), and Manipal Academy of Higher Education (MAHE) for the support. CS thanks MAHE for Dr. TMA Pai PhD Scholarship and TSM thanks MAHE for intramural funding.
Funding
Open access funding provided by Manipal Academy of Higher Education, Manipal
Author information
Authors and Affiliations
Contributions
All the authors contributed to the idea of the article. Literature search and compilation of data were performed by Chandni. The first draft was written by Chandni and critically revised by Murali and Satyamoorthy. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Human and Animal Rights and Informed Consent
This review article does not include any studies with human or animal subjects performed by any of the authors.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Bacteriology
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sachdeva, C., Satyamoorthy, K. & Murali, T.S. Microbial Interplay in Skin and Chronic Wounds. Curr Clin Micro Rpt 9, 21–31 (2022). https://doi.org/10.1007/s40588-022-00180-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40588-022-00180-4