Abstract
The SWI/SNF chromatin remodeling complex utilizes the energy of ATP hydrolysis to facilitate chromatin access and plays essential roles in DNA-based events. Studies in animals, plants and fungi have uncovered sophisticated regulatory mechanisms of this complex that govern development and various stress responses. In this review, we summarize the composition of SWI/SNF complex in eukaryotes and discuss multiple functions of the SWI/SNF complex in regulating gene transcription, mRNA splicing, and DNA damage response. Our review further highlights the importance of SWI/SNF complex in regulating plant immunity responses and fungal pathogenesis. Finally, the potentials in exploiting chromatin remodeling for management of crop disease are presented.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Eukaryotic genome are organized and compacted by 146 base pairs of DNA wrapped around a histone octamer, forming structures named nucleosomes, which enable long DNA strands to precisely fit into the nucleus (Mittal & Roberts, 2020; Ojolo et al. 2018). Within the nucleus, DNA strands are highly folded, constrained, and compacted into higher order chromatin structures, and the dynamic regulation of chromatin can ensure the appropriate timing, location and sequence of cellular DNA-based events (Luger et al. 2012; Roberts & Orkin, 2004). Therefore, mechanisms that govern chromatin dynamics are integral components of eukaryotic gene regulation, that include covalent histones, DNA modifications and ATP-dependent chromatin remodeling complexes (Centore et al. 2020; Roberts & Orkin, 2004).
Chromatin remodelers utilize the energy derived from ATP (adenosine triphosphate) hydrolysis to move, destabilize, eject, or restructure nucleosomes, thus modulating the access of transcription machinery to DNA (Becker & Hörz, 2002; Hohmann & Vakoc, 2014). The ATP-dependent chromatin remodeling complex can be divided into four distinct subfamilies: SWI/SNF, ISWI, CHD and INO80. These subfamilies share a conserved ATPase domain but functions in a largely non-redundant manner to govern discrete biological processes, such as transcriptional regulation, DNA replication, DNA repair, homologous recombination and chromosomal segregation (Clapier & Cairns, 2009; Hohmann & Vakoc, 2014; Masliah-Planchon et al. 2015). Among the four chromatin remodeling complex subfamilies, the SWI/SNF complex is evolutionarily conserved and was first discovered through genetic screens and biochemical purification in budding yeast Saccharomyces cerevisiae (Mittal & Roberts, 2020; Neigeborn & Carlson, 1984). Notably, the SWI/SNF complex is the most strongly associated regulator of chromatin access (Clapier et al. 2017; Euskirchen et al. 2012). Many studies have shown that the SWI/SNF complex regulates various stress responses and developmental pathways in budding yeast, Drosophila, and human through precise control of gene expression (Kasten et al. 2011; Kwon & Wagner, 2007). In this review, we highlight key functions and regulatory mechanisms of this complex in regulating plant-pathogen interactions.
Compositions of the SWI/SNF complex
The conserved components of SWI/SNF complex were characterized in several eukaryotic organisms including S. cerevisiae, Arabidopsis and human (Table 1). All core and actin-related subunits, as well as the transcription associated component Swp73/Snf12, are well conserved in various eukaryotes. However, each organism uniquely constructs distinct SWI/SNF complex using both the conserved components and unique subunits. Even though the complex normally contains 9–12 subunits (Roberts & Orkin, 2004; Smith et al. 2003), only five of these, i.e. Swi2/Snf2 (the ATPase subunit), Snf5, Swi3, Arp7 and Arp9, are comparable or almost comparable to the entire complex when we take budding yeast as an example (Phelan et al. 1999; Roberts & Orkin, 2004; Zhang et al. 2018). Studies on the subunit architecture in S. cerevisiae revealed that the loss of Arp7 or Arp9 disrupts the catalytic core of SWI/SNF (Zhang et al. 2018). While in the absence of Snf5, the catalytic ternary complex Snf2-Arp7-Arp9 could be fully detached, suggesting its role in coordinating the distinct modules in SWI/SNF (Dutta et al. 2017; Zhang et al. 2018). In addition, the locations of Snf5, Swp82 and Swi1 indicate that these subunits are associated with the binding of the complex with transcription factors (Prochasson et al. 2003; Zhang et al. 2018).
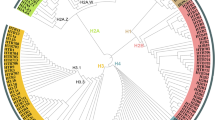
The number of conserved components also varies from species to species, especially the most vital components associated with ATPase activity (Table 1). This finding prompted us to construct a phylogenetic tree to characterize the homologs of Snf2 and their functional domains in animals, plants and fungi. As shown in Fig. 1, the predicted ATPases were clustered into six groups: Group 1, SNF2 and BRG1; Group 2, CHR12; Group 3, BRM and SYD; Group 4, DDM1; Group 5, CHD1; and Group 6, ISWI. All group members contain two highly conserved domains, SNF2_N and Helicase_C, while different groups having unique domains. For instance, only group 1 members contain HSA and bromo domain, and the bromo domain is capable of binding to acetylated histones, which suggests that members of group 1 may function as reader proteins of acetylation (Jarończyk et al. 2021). SnAC domain is distributed in groups 1 and 2, and the SYD domain in members of group 3. The QLQ domain is present only in groups 1 and 3. Group 4 members have the shortest protein length, and they only contain the two conserved domains SNF2_N and Helicase_C. Interestingly, almost all members of group 4 originate from plants, and most of them are DDM1 or DDM1-like proteins. These proteins play a role in maintaining DNA methylation even though they have no methyltransferase activity (Ramirez-Prado et al. 2018; Vongs et al. 1993). Another noticeable finding is that the chromo domain is strictly distributed within the group 5. This domain is implicated in the binding of the proteins that are found to methylate histone tails and RNAs (Lu et al. 2020; Pray-Grant et al. 2005). It is noteworthy that the CHD3 members of group 5 own three extra domains, i.e., PHD, DUF1086, and DUF1087. Among them, the PHD is considered as a histone code reader (Wang et al. 2021). In addition, the components of group 6 also contain two distinct domains SLIDE and HAND that are associated with extra-nucleosomal DNA and the entry site of nucleosomes (Dang & Bartholomew, 2007).
Phylogenetic tree of ATPases in animals, plants, and fungi. The sequence of ScSnf2 (YOR290C) was used as a bait to elicit all sequences with high homology and annotation, and the threshold was set to e-value = 1e− 10. A maximum-likelihood phylogenetic tree was constructed with sequences of ATPases or their homologs. All predicted ATPases were attributed to six groups (Group 1–6) based on the composition of domains. Different groups and conserved domains of ATPases were indicated by different colors. Am: Apis mellifera; At: Arabidopsis thaliana; Af: Aspergillus fumigatus; Bm: Bombyx mori; Bc: Botrytis cinerea; Bd: Brachypodium distachyon; Bn: Brassica napus; Ce: Caenorhabditis elegans; Ca: Candida albicans; Candida g: Candida glabrata; Cp: Candida parapsilosis; Cg: Colletotrichum graminicola; Cn: Cryptococcus neoformans; Cs: Cucumis sativus; Dc: Daucus carota; Dm: Drosophila melanogaster; Ff: Fusarium fujikuroi; Fg: Fusarium graminearum; Fo: Fusarium oxysporum; Fv: Fusarium verticillioides; Hc: Histoplasma capsulatum; Hs: Homo sapiens; Nc: Neurospora crassa; Nt: Nicotiana tabacum; Os: Oryza sativa; Pb: Paracoccidioides brasiliensis; Pa: Podospora anserine; Po: Pyricularia oryzae; Sc: Saccharomyces cerevisiae; Sp: Schizosaccharomyces pombe; Ss: Sclerotinia sclerotiorum; Sl: Solanum lycopersicum; St: Solanum tuberosum; Tt: Thermothelomyces thermophiles; Tr: Trichoderma reesei; Yl: Yarrowia lipolytica; Zm: Zea mays
Functional mechanisms of the SWI/SNF complex
Regulatory roles of the SWI/SNF complex in gene transcription
The SWI/SNF remodelers are key regulators of nucleosome positioning, which typically controls chromatin accessibility and binding sites for transcriptional machinery at the gene promoters or enhancers, thus leading to either gene activation or repression (Clapier et al. 2017; Euskirchen et al. 2012). Genome-wide analysis showed that approximately 5% of genes are regulated by the SWI/SNF at the transcription level in the budding yeast (Sudarsanam et al. 2000), flies (Zraly et al. 2006), as well as mice (Gresh et al. 2005). However, due to the low intracellular level and the lack of intrinsic DNA binding specificity of the SWI/SNF complexes, they need to be guided by gene-specific transcriptional regulators, covalent histone modifiers or long noncoding RNAs (lncRNAs) to facilitate specific gene loci targeting in various organisms (Peterson & Workman, 2000; Sanz et al. 2012).
In A. thaliana, the physical interaction of SWI/SNF subunits with different proteins governs a wide range of developmental processes, such as embryo, leaf and flower organ development, response to plant hormones, and abiotic stresses (Ramirez-Prado & Benhamed, 2021; Reyes, 2014). A plant-unique H3K27 demethylase REF6 targets the CTCTGYTY motif-containing genomic loci through its zinc-finger (ZnF) domains and further facilitates recruitment of the BRM complex (Li et al. 2016). The Arabidopsis SYD (a homolog of the yeast Snf2p ATPase) can act as a transcriptional repressor of the meristem identity switch in the floral transition via interacting with and altering activity of the plant-specific transcriptional activator LEAFY (Wagner & Meyerowitz, 2002). Upon auxin sensing, the MONOPTEROS transcription factor recruits the BRM complex to increase DNA accessibility for induction of key regulators of flower primordium initiation (Wu et al. 2015). In Arabidopsis, some lncRNAs are produced by a specialized RNA Polymerase V (Pol V). The Pol V-produced lncRNAs can be associated with IDN2, a lncRNA-binding protein. Interestingly, SWI3B, an essential subunit of the BRM complex physically interacts with DN2, and subsequently contributes to lncRNAs-mediated transcriptional silencing (Zhu et al. 2013).
The fungal SWI/SNF complexes work in concert with various transcription factors and covalent histone modifiers to facilitate maintaining proper chromatin accessibility landscapes during fungal development and stress responsive processes (Table 2), which is consistent with those in plants. When we look at yeast as an example, the SWI/SNF complex functions together with regulatory protein Yap8 to mediate transcriptional activation of ACR2 and ACR3 in response to arsenic stress in S. cerevisiae (Menezes et al. 2017). While transcription factor Cha4 recruits SWI/SNF to initiate SRG1 transcription by remodeling the two nucleosomes located at the SRG1 transcription start site when serine is available to the cells (Hainer et al. 2011). When yeast encounters cell wall stress, the downstream transcription factor of cell wall integrity pathway Rlm1 physically interacts with SWI/SNF to direct its association to target promoters (Sanz et al. 2012). Moreover, several reports showed that the SWI/SNF complex is recruited by multiple transcription factors. For instance, Adr1 and Cat8 recruits SWI/SNF complex upon glucose repression (Biddick et al. 2008), whereas two classes of transcriptional activators HSF and Msn2/4 associate with this complex to modulate chromatin disassembly at heat shock gene promoters (Erkina et al. 2008; Shivaswamy & Iyer, 2008). One interesting report demonstrated that the yeast repressor Sko1 recruits Cyc8(Ssn6)-Tup1 corepressor complex to regulate transcription of genes that are induced upon hyperosmotic stress. During this process, the MAP kinase Hog1 associates with target promoters, phosphorylates Sko1, and converts Sko1 into a transcriptional activator. Subsequently, the formation of Sko1/Hog1/Tup1 ternary transcription activator complex is important for SWI/SNF recruitment during the transcriptional induction process (Proft & Struhl, 2002). Additionally, histone modifiers can also mediate the recruitment of SWI/SNF complex. As yeast cells progress through the cell cycle, the activator Swi5 enters into nuclei at the end of anaphase, which recruits both SWI/SNF and Spt-Ada-Gcn5-Acetyltransferase (SAGA) complexes to the HO endonuclease promoter (Cosma et al. 2016). Nucleosome arrays provide a functional link between histone acetylation and the SWI/SNF complex, and the retention of SWI/SNF is mediated by histone acetylation (Hassan et al. 2001). Further studies suggest that SWI/SNF preferentially displaces acetylated histones from the array relative to total histones, and the acetyl-lysine binding domain, Swi2/Snf2 bromodomain, plays a vital role in this process (Chandy et al. 2006; Mitra et al. 2006). Upon phosphate depletion, chaperone anti-silencing function 1 (Asf1) recruits SWI/SNF complex to promote chromatin disassembly at the yeast PHO5 promoter (Adkins et al. 2007).
Involvement of the SWI/SNF complex in mRNA splicing
In higher eukaryotes, mRNA splicing is important for controlling both qualitative and quantitative aspects of gene expression. Moreover, the alternative splicing from a single gene can generate multiple functionally distinct protein isoforms, which greatly expands the genetic plasticity of an organism and has emerged as a vital layer of gene regulation in response to diverse stresses (Black, 2000; Blencowe, 2006; Pleiss et al. 2007). Changes in chromatin structure have been shown to affect mRNA splicing (Allemand et al. 2016; Kornblihtt, 2006). The genome-wide mapping of nucleosome positioning from different organisms shows that nucleosomes are particularly enriched at intron-exon junctions (Luco et al. 2011), which is evolutionarily conserved from plants to mammals, suggesting an essential role of nucleosome positioning in exon definition (Luco et al. 2011; Schwartz et al. 2009).
The SWI/SNF has been reported to interact with Pol II (Kornblihtt, 2006), thus influencing the efficiency of splicing (Allemand et al. 2016). For example, the accumulation of phosphorylated RNA Pol II in a central block of alternative exon of the CD44 gene in human cells can be caused by the overexpression of Brm, the core ATPase of SWI/SNF complex, which leads to the increased accumulation of mature mRNA (Batsché et al. 2006). Furthermore, the SWI/SNF complex also regulates mRNA splicing by modulating the transcription elongation rate of RNA Pol II through its subunit SNF5, therefore affecting the transcription of genes involved in hormone signaling (Zraly & Dingwall, 2012). The maize SWI3D protein, ZmCHB101, impacts alternative splicing by influencing transcriptional elongation rate mediated by RNA Pol II in response to osmotic stress (Yu et al. 2019). Schwabish and Struhl (2007) reported that SWI/SNF affects RNA Pol II elongation thereby influencing splicing through two possible mechanisms, including the association with Pol II or factors that travel with elongating Pol II, and recognition of distorted chromatin that occur during transcriptional elongation process to permit passage of Pol II.
A widely reported mechanism of SWI/SNF controlling splicing is by recruitment of splicing machinery, e.g., spliceosomal components, splicing factors (SFs) and RNA binding proteins (RBPs). In fission yeast, SWI/SNF contributes to splicing catalysis by promoting the recruitment of Prp2 ATPase, which functions to destabilize SF3 immediately before the first step of catalysis (Patrick et al. 2015). Similarly, SWI/SNF is considered as a key regulator in S. cerevisiae meiotic splicing, because it can lead to the redistribution of spliceosomes from ribosomal protein genes (RPGs), where the splicing of RPGs can be finely tuned to the environmental conditions and nutrient availability encountered by cells (Pleiss et al. 2007; Venkataramanan et al. 2017). Human and Drosophila SWI/SNF family influences splicing when adapting to environmental stimuli via physical interaction and snRNPs U1/U5 recruitment (Batsché et al. 2006; Tyagi et al. 2009). Studies in HeLa and Schizosaccharomyces pombe cells both revealed that interactions between U2 snRNP and SWI/SNF subunits influence splicing process (Cavellán et al. 2006; Fair & Pleiss, 2017; Makarov et al. 2012; Patrick et al. 2015). Specifically, human SWI/SNF complex may serve as an adaptor for U2 snRNP association with chromatin. SWI/SNF proteins were identified as components of spliceosomal complex E (Makarov et al. 2012), and the overexpression of U2 snRNP components in ΔSWI/SNF cells led to inefficient splicing of many introns in fission yeast (Patrick et al. 2015). Moreover, studies in Drosophila indicated that SWI/SNF plays a role in pre-mRNA processing, possibly by modulating the recruitment and/or assembly of splicing factors (Waldholm et al. 2011). BAF57/SMARCE1 interacts with splicing factor SRSF1 to regulate mechanical stress-induced alternative splicing of cyclin D1 in osteoblast cells (Feng et al. 2021). In addition to splicing factors, recruitment of RNA binding proteins is also an important way to modulate splicing by SWI/SNF complex. For instance, in human cells, the ATPase Brm in concert with the mRNA-binding protein p54 regulate the splicing of telomerase reverse transcriptase (TERT) by accelerating exon-inclusion (Ito et al. 2008). Expression of the ATPase BRG1 in cervical cancer C33A cells promotes the local recruitment of splicing-RNA binding factors to chromatin and RNA, and further alters their binding to the nascent pre-mRNA, subsequently affecting alternative splicing (Zapater et al. 2019).
Taken together, SWI/SNF complex influences splicing by two distinct but conserved mechanisms, including modulation of RNA Pol II accumulation/elongation and recruitment of splicing machinery (Allemand et al. 2008; Batsché et al. 2006; Kornblihtt, 2006) (Fig. 2). Moreover, the function of splicing in response to external stimuli is an ubiquitously accepted way that allows eukaryotes to quickly adjust the abundance of functional transcripts to environmental perturbations (Allemand et al. 2008; Capovilla et al. 2015; Pleiss et al. 2007). While studies in fungi remain limited, especially those in pathogenic fungi, summarizing SWI/SNF interactors affecting splicing events upon stress stimuli in plants and animals is important and provides significant reference for research in pathogenic fungi.
Two mechanisms by which SWI/SNF complex may affect splicing to adapt environmental stimuli. A The overexpression of Brm causes accumulation of phosphorylated RNA Pol II, thereby leading to increased mature mRNA. The core subunits of SWI/SNF complex SNF5 and Swi3, can modulate the transcription elongation rate of RNA Pol II at the target pre-mRNA, thus affecting the transcripts of hormone signaling and osmotic response, respectively. B SWI/SNF complex regulates pre-mRNA splicing by recruiting components of spliceosome, SFs (Splicing factors), as well as RBPs (RNA binding proteins) to target pre-mRNA. Both of the mechanisms promote splicing progress
Involvement of the SWI/SNF complex in DNA damage repair (DDR)
DNA damage can be corrected by DNA repair pathways (Ataian & Krebs, 2006; Lindahl, 2000). However, the genomic DNA packaged inside chromatin hinders DNA accessibility, and therefore the DNA repair and signaling machineries have to overcome this chromatin barrier to access the lesion (Ataian & Krebs, 2006; Smeenk & van Attikum, 2013; Wang et al. 2020). An increase in chromatin mobility has also been observed at lesions of budding yeast and human cells exposed to DNA damage (Hauer et al. 2017; Miné-Hattab & Rothstein, 2012; Roukos et al. 2013), which suggests a tight relationship between DNA damage repair (DDR) and chromatin structure.
ATP-dependent chromatin remodeling complex SWI/SNF members function directly in nucleotide-excision repair (NER), double-strand break (DSB) repair and other DDR pathways by modifying chromatin structure around DNA damage sites and further recruiting DDR proteins (Bao & Shen, 2007; Mittal & Roberts, 2020; Ribeiro-Silva et al. 2019; Smeenk & van Attikum, 2013) (Fig. 3). NER is a versatile DNA repair pathway that can remove a variety of structurally unrelated lesions including UV-induced bulky DNA adducts (Spivak, 2015). In S. cerevisiae, damage-recognition heterodimer Rad4-Rad23 associates with Snf6 and Snf5, two subunits of the SWI/SNF complex to increase DNA accessibility for NER in chromatin, and their association is stimulated by UV irradiation (Gong et al. 2006). Moreover, BRG1 facilitates NER at different stages by modulating chromatin relaxation and stabilizing xeroderma pigmentosum group C protein (XPC) at the damage sites, which subsequently promotes xeroderma pigmentosum group G protein (XPG), and proliferates cell nuclear antigen (PCNA) recruitment to complete the repair in mammalian cells (Zhao et al. 2009). Inactivation of ATPase subunits downregulates GTF2H1, a core subunit of the transcription factor IIH (TFIIH) complex, thus compromising TFIIH stability and NER pathway (Ribeiro-Silva et al. 2018).
Regulatory mechanisms of the SWI/SNF complex in various DNA damage repair pathways. A Various models of how different subunits of SWI/SNF complex can recruit or be recruited by DNA damage associated factors to facilitate DNA damage repair (DDR) pathways, such as NER, HR, NHEJ and other DDR pathways. B Phosphorylation of ATPase or acetylation/phosphorylation of histone are implicated to promote the localization of SWI/SNF complex and further recruit DNA damage associated factors at the damage loci
In addition to NER, the SWI/SNF complex also functions in homologous recombination (HR) and non-homologous end-joining (NHEJ), which are two major conserved DSB repair pathways (Bao & Shen, 2007; Smeenk & van Attikum, 2013). For example, the chromatin remodeling activity of SWI/SNF contributes to Mre11-Rad50-Xrs2 (MRX) recruitment and resection initiation during HR, as nucleosome eviction at a DSB site is observed to be delayed in a SWI/SNF mutant (Wiest et al. 2017). Promoting the Rad51- and Rad54-dependent strand invasion during recombinational repair of the mating-type loci also requires SWI/SNF complex to alleviate heterochromatic constraints (Hauer & Gasser, 2017; Sinha et al. 2009). Consistently, downregulation of BRG1 and BRM in human cells reduced HR efficiency by 40–50% and 15%, respectively. BRG1 is important for activating ATM- and Rad3-related (ATR) kinase and reducing nucleosome density at DSBs, which further changes the chromatin structure and promotes CtIP nuclease recruitment, thus stimulating DNA end resection and HR (Hays et al. 2020). In both yeast and mammalian cells, BRM stimulates recruitment of NHEJ factors KU70/KU80, while BRG1 promotes HR-associated DNA end resection and RPA and RAD51 loading (Ribeiro-Silva et al. 2019). SWI/SNF complex also participates in other DDR pathways. For example, inactivation of ATPase subunits compromise H2AX phosphorylation, thereby affecting their function in DSB repair (Park et al. 2006). Meanwhile, the depletion BRG1 increases R-loops and R-loop-dependent DNA breaks as well as transcription-replication conflicts (Bayona-Feliu et al. 2021). In plant cells, the presence of an appropriate level of the SWI/SNF subunit SWI3B enhances the dissociation of structural maintenance complex 5 (SMC5) from chromosomes for its further recruitment at DSBs during DNA damage (Jiang et al. 2019).
Depletion, mutation or loss of SWI/SNF subunits has been shown to cause defects in DNA damage repair, suggesting that SWI/SNF complex is rapidly recruited to DSBs (Harrod et al. 2020). But how does SWI/SNF complex know when and where to find targets? Post-transcriptional modifications have been implicated to promote the localization of SWI/SNF to DNA damage sites. For instance, A-T mutated (ATM)-mediated phosphorylation of BRG1 and BAF170 increases the association of SWI/SNF complex with the early DDR protein BRIT1/MCPH1 (Kwon et al. 2015; Peng et al. 2009). The Ser-721 phosphorylation of BRG1 by ATM facilitates DSB repair by stimulating the association with γ-H2AX nucleosomes via enhancing the affinity to acetylated H3 (Kwon et al. 2015). The histone H2B kinase AMPK2, a major substrate of the tumor suppressor LKB1, is recruited to DSBs by LKB1. Disruption of the AMPK2 phosphorylation site impairs BRM recruitment to DSB sites, which further fails to activate NHEJ pathway (Ui et al. 2014). In addition, the acetylation of histone H3 and H4 has been shown to facilitate SWI/SNF recruitment. CBP and p300 acetylate H4 K5/K8/K12/K16 and H3 K18 to facilitate SWI/SNF chromatin remodeling at DSBs, which may provide access to the damage site for the NHEJ factors Ku70/Ku80 (Ogiwara et al. 2011). The SWI/SNF subunit Taf14 functions as a selective reader of histone H3 Lys9 acetylation (H3K9ac), and disruption of this binding in cells impairs the transcription of DNA damage response genes (Shanle et al. 2015). γH2AX recruits histone acetyltransferase GCN5 to trigger acetylation of H3, which subsequently attracts SWI/SNF binding (Lee et al. 2010; Park et al. 2006). Other mechanisms can also contribute to SWI/SNF recruitment at DSBs, for example, DDR protein BRIT1/MCPH1 associates with core subunits of SWI/SNF and promotes their recruitment and retention (Lukas et al. 2011; Peng et al. 2009). During methyl methanesulfonate (MMS) induction in S. cerevisiae, the activator Crt1 facilitates the recruitment of TFIID and SWI/SNF, which in turn promotes chromatin remodeling and preinitiation complex assembly (Ghosh & Pugh, 2011).
Roles of the SWI/SNF complex in plant-pathogen interaction
As sessile organisms, plants must properly modulate their gene expression to survive in the environment, which is greatly influenced by the dynamic chromatin structure (Chang et al. 2020; Song et al. 2021). Mounting evidence has shown that SWI/SNF complex plays important roles in plant abiotic stress responses, including temperatures, drought, salt, osmotic stress and hormone signaling pathways as well as biotic stresses (Bhadouriya et al. 2020; Chang et al. 2020; Song et al. 2021; Thouly et al. 2020). In this review, we focus on deciphering the roles of this chromatin remodeling complex in response to plant biotic stress, which will advance our understanding in plant-pathogen interactions.
The SWI/SNF complex regulates plant immune response
Unlike animals, plants lack an adaptive immune system that produces antibodies and lack mobile cells to detect, prevent or reduce infections once perceiving microbial pathogens. Instead, plants have evolved an innate immunity system to defend against microbial attack (Ding & Wang, 2015). Phytohormones including salicylic acid (SA), jasmonic acid (JA), ethylene (ET), abscisic acid (ABA), auxins, gibberellins (GA), cytokinins, and brassinosteroids, emerged as important players in plant immune response (Pieterse et al. 2009). Accumulating evidence has shown that chromatin structures play essential roles in regulating plant defense responses via physical association with the components or regulators of phytohormone signaling pathways (Chen et al. 2017; Ding & Wang, 2015; Ojolo et al. 2018; Ramirez-Prado et al. 2018; Sarnowska et al. 2016).
In Arabidopsis, BRM binds to abscisic acid (ABA)-induced basic domain/leucine zipper transcription factor ABA INSENSITIVE5 (ABI5) locus to stabilize the local nucleosome, thus negatively regulating its expression level in the absence of stress stimuli (Han et al. 2012). A two-hybrid analysis revealed that the core subunit of SWI/SNF complex SWI3B interacts with HYPERSENSITIVE to ABA1 (HAB1), a protein phosphatase type 2C, to modulate ABA signaling (Saez et al. 2008). In addition, the dysregulation of pathogenesis-related (PR) genes in the BRM101 mutant indicates a role for this protein in salicylic acid (SA)-mediated resistance (Bezhani et al. 2007; Chen et al. 2017; Han et al. 2012). BRM also directly associates with the promoters of GA biosynthesis genes to positively regulate its expression, whereas the loss of BRM leads to remarkable decrease level of endogenous bioactive GA (Archacki et al. 2013; Reyes, 2014). Moreover, several subunits of SWI/SNF complex (SWI3B, SWI3D, and BSH) interact with the miRNA-binding protein ARGONAUTE1 (AGO1) to facilitate its binding to chromatin upon JA, auxin, and SA stimuli in Arabidopsis (Liu et al. 2018a; Maury et al. 2019).
In addition to regulating phytohormone signaling pathways, the SWI/SNF complex plays important roles in modulating transcription and splicing of resistant genes. In Arabidopsis, SWP73A directly binds to the promoters of NLR (NOD-like receptor) genes to suppress their expression level. In addition, it may also function as a H3K9me2 reader to enhance this transcription suppression (Huang et al. 2021), as H3K9me2 is a transcriptional repression marker in plants (Pfluger & Wagner, 2007). Moreover, SWP73A affects the alternative splicing of some NLRs through indirectly suppressing the key regulator of RNA splicing CDC5, thus inhibiting the defense responses in Arabidopsis. In turn, bacteria-induced small RNAs silence SWP73A to activate a group of NLRs and trigger robust immune responses upon infection (Huang et al. 2021).
Other than the above mentioned subunits of SWI/SNF complex, another Arabidopsis ATPase, SYD, also plays important roles in plant defense against specific biotic stresses. Genetic, biochemical and biological evidences have shown that SYD is recruited to the promoters of genes controlling jasmonate (JA)- and ethylene (ET)-dependent defense responses to positively regulate their expression levels upon Botrytis cinerea infection (Walley et al. 2008). Moreover, SYD is required for resistance against the necrotrophic pathogen B. cinerea rather than the biotrophic pathogen Pseudomonas syringae, which indicates that chromatin remodeling participates in resistance to pathogens selectively (Walley et al. 2008). SNC1 (Suppressor of NPR1, Constitutive 1) is an intracellular Arabidopsis NLR protein that can be activated to induce defense responses (Zhang et al. 2003). Meanwhile, SYD functions antagonistically with MOS1 and MOS9 at the chromatin level to regulate SNC1 transcription and SNC1-mediated immunity (Johnson et al. 2015). Strikingly, the expression levels of SYD were found to be dramatically decreased upon indoleacetic acid (IAA), ABA, and benzothiadiazole (BTH) treatments and increased upon flg22 treatment, which suggests that SYD responds to pathogen attack at the transcription level (Shu et al. 2021).
In rice (Oryza sativa L.), the putative SWI/SNF2 class ATPase BRHIS1 is downregulated by the rice blast fungus Magnaporthe oryzae infection, suggesting that BRHIS1 plays a negative regulatory function in plant immunity (Li et al. 2015). RNA-seq and ChIP-seq data show that BRHIS1 suppresses the expression of a few defense-related genes (OsPBZc and OsSIRK1) rather than SA pathway genes, therefore regulating plant immunity in an SA-independent manner (Li et al. 2015). Further co-IP assays suggest that BRHIS1 specifically interacts with monoubiquitinated histone variants, H2A.Xa/H2A.Xb/H2A.3 and H2B.7 (Li et al. 2015). However, the enrichment of these histone variants at the promoter regions of OsPBZc and OsSIRK1 is correlated with their increased expression, whereas the BRHIS1 expression is suppressed (Li et al. 2015). These findings indicate that the BRHIS1 can relax chromatin state for defense genes by monoubiquitination local histone variants under normal growth conditions. Upon pathogen attack, BRHIS1 is inhibited, which makes chromatin accessible for defense gene expression. Together, the association between BRHIS1 and monoubiquitinated histone variant allows plants to establish an expression-ready chromatin state for defense genes to facilitate rapid activation of induced plant immune responses (Chen et al. 2017; Li et al. 2015).
The signaling pathways regulates the SWI/SNF complex in plant
Although the pivotal role of SWI/SNF complex in plant immunity has been documented, how this complex is regulated in stress response or pathogen attack remains to be elucidated. One interesting study established a model suggesting that effector-SWI/SNF association plays vital roles during ectomycorrhizal-plant interactions. During infection, the effector-like protein PaMiSSP9.7 encoded by mycorrhizal fungus Pisolithus albus enters host root cells and localizes into the nucleus. Furthermore, it interacts with the SWI3D, a subunit of the SWI/SNF complex in host plant Eucalyptus grandis, that is activated and required to alter the outcome of mycorrhization (Aguirre, 2017).
More recently, Song and colleagues demonstrated the stability and activity of SWI/SNF subunits are also controlled by post-translational modifications (Song et al. 2021). Vicente and colleagues found that treatment with NaCl or ABA resulted in a decline in BRM protein (Vicente et al. 2017), which may be a result of the phosphorylation or dephosphorylation of BRM (Peirats-Llobet et al. 2016). Specifically, phosphorylation of BRM by the kinase SnRK2 contributes to its inhibition, whereas PP2CA-mediated dephosphorylation recovers the ability of BRM to negatively control ABA response (Peirats-Llobet et al. 2016). Another core subunit of SWI/SNF complex, SWI3B, physically interacts with the clade A PP2C phosphatase HAB1 (a core component in ABA signal pathway) and is directly dephosphorylated in an ABA-dependent manner (Saez et al. 2008). In addition to phosphorylation or dephosphorylation, the ubiquitin-proteasome system has also been shown to modulate BRM protein stability (Song et al. 2021). For instance, deletion mutants of brm-3 and the SUMO ligase mms21–1 both show resembling defects in root development, and the protein level of BRM is dramatically decreased in mms21–1 mutant. Further biochemical evidence indicated that BRM is modified by SUMO3, and AtMMS21 enhances this SUMOylation to elevate BRM stability (Zhang et al. 2017). BRM is degraded by 26S proteasome in response to high-boron induced DSBs (Sakamoto et al. 2018), suggesting BRM is a target of 26S proteasome and is required for DSBs tolerance. During this process, BRM associates with histone acetylation to make chromatin more accessible, which could be inhibited by 26S proteasome (Sakamoto et al. 2018).
The SWI/SNF complex plays important roles in fungal pathogenesis
During the infection process, pathogens encounter diverse stresses. To successfully invade hosts, fungi have evolved sophisticated strategies to adjust their developmental processes to adapt to the changes in various stimuli, including both the environment and the host (Łaźniewska et al. 2012). Several reports have demonstrated that the SWI/SNF complex is involved in stress responses and fungal pathogenesis (Table 2). The best characterized model is the most prevalent human fungal pathogen C. albicans. Knock-out mutation of SWI1 or SNF2, two core subunits of SWI/SNF complex, leads to a complete loss of pathogenicity in murine systemic candidiasis by inhibiting the expression of hyphae (filamentation)-specific genes (Mao et al. 2006). When cells encounter serum or nutrient starvation, the ATPase SNF2 is recruited by a synergistic action of NuA4 HAT complex and the hyphae-specific transcription factor Efg1 to the promoters of hyphae-specific genes, indicating that the SWI/SNF complex harnesses histone modifying enzyme to govern Candidia pathogenicity (Lu et al. 2008). Consistently, deletion of SNF5 leads to an altered metabolome and a loss of virulence, which implies its essential roles in maintaining metabolic homeostasis and pathogenicity in C. albicans (Burgain & Pic, 2019). The SWI/SNF complex is also involved in fluconazole resistance by cooperating with the inactivated form of transcription factor Mrr1 to promote nucleosome displacement from MDR1 (a multiple drug resistance gene 1) promoter, which further permits Mrr1 binding (Liu & Myers, 2017). Importantly, the fungal-specific subunit of SWI/SNF complex SNF6 is critical for C. albicans to colonize its host and to cause disease, suggesting SNF6 is a potential antifungal target (Tebbji et al. 2017). Similarly, in Cryptococcus neoformans, SWI/SNF assists transcription factor Znf2 to control yeast-to-hypha differentiation, which opens the promoter regions of hyphal specific genes, including the ZNF2 gene itself, therefore facilitating fungal pathogenesis (Lin & Zhao, 2019).
In Neurospora crassa, white Collar-1 (WC-1) recruits SWI/SNF complex to remodel and loop chromatin at FRQ, thereby activating FRQ expression to initiate the circadian cycle (Wang et al. 2014). In Trichoderma reesei, the transcriptional activator of cellulase/hemicellulase genes, XYR1, interacts with the SNF12 subunit of SWI/SNF complex to remodel chromatin at cellulase gene promoters, thus activating their expression to initiate the cellulolytic response (Cao et al. 2019). In our previous studies, we found two transcription factors FgAreB and FgSR that recruit SWI/SNF complex via direct interaction with Swp73 to orchestrate genes responding to nitrosative and phytoalexin stresses, respectively, during F. graminearum infection (Jian et al. 2021; Liu et al. 2019). Importantly, Swp73 is essential in F. graminearum (Jian et al. 2021). These results are consistent with the cases observed in human, which also showed that BAF60a can serve as a bridge for the interactions between transcription factors and SWI/SNF complex (Iba et al. 2003; Oh et al. 2008). In addition, BAF60 (a Swp73 homolog) RNA interference mutant lines showed severe growth defects in Arabidopsis (Jégu et al. 2014), which further stressed the decisive role of this subunit across eukaryotes. Based on these published reports, we deduced that the disruption of interactions between BAF60 and transcription factors can cause transcriptional repression of a number of genes that participate in stress response.
It is widely accepted that reactive oxygen species (ROS) production by NADPH oxidases is one of the earliest responses of pathogen recognition in both plants and animals (Kadota et al. 2015). ROS acts as antimicrobials to prevent pathogen entry, and ROS-mediated DNA oxidative damage can be corrected by DNA repair pathways (Wang et al. 2020). Studies have established that the SWI/SNF complex is required for DNA damage repair, which is conserved from fungi to humans (Bao & Shen, 2007; Bohm et al. 2021; Harrod et al. 2020; Jiang et al. 2019; Ribeiro-Silva et al. 2019; Wiest et al. 2017). Therefore, we propose that the SWI/SNF complex plays a critical function in responses to ROS during plant-pathogen interactions. Although studies about the canonical SWI/SNF participates in this process have not been reported yet, another subtype of SWI/SNF family RSC has been characterized to modulate ROS response in phytopathogen-plant interactions. For example, silencing SFH1 (Snf5 homolog) in Sclerotinia sclerotiorum causes defects in hyphal growth and decreases ROS accumulation, suggesting its function in ROS production (Liu et al. 2018b). Also in the soil-borne pathogenic fungus Verticillium dahlia, VdDpb4 and VdIsw2 of ISWI chromatin remodeling complex play roles in maintaining chromatin structure for positioning nucleosomes and transcription regulation of DDR genes in response to ROS stress during plant infection (Wang et al. 2020).
In addition to ROS, host perception of pathogens can also provoke reactive nitrogen species (RNS), leading to nitrosative stress (NS) (Arasimowicz-Jelonek & Floryszak-Wieczorek, 2016). It is worth mentioning that our recent work suggests SWI/SNF complex can be recruited by transcription factor FgAreB to the nitrosative stress response (NSR) gene promoters, subsequently promoting NSR gene expression in F. graminearum (Jian et al. 2021). Furthermore, a transcriptional repressor, FgIxr1 was found to compete with the SWI/SNF complex to bind FgAreB, which negatively regulates NS response. In turn, NS promotes FgIxr1 degradation, therefore, enhancing the recruitment of the SWI/SNF complex by FgAreB (Jian et al. 2021). Taken together, SWI/SNF complex participates in plant-pathogen interaction via different routes (Fig. 4).
The SWI/SNF complex is involved in plant-pathogen interaction. A During Eucalyptus grandis-Pisolithus albus interaction, the effector-like protein PaMiSSP9.7 produced by the mycorrhizal fungus enters plant root nucleus to interact with Swi3D of SWI/SNF complex. B SWI/SNF complex mediates fungal development and pathogenesis by reprogramming the expression of stress responsive genes to phytohormone, ROS and RNS during various pathogen-plant interactions
Future perspectives
Nucleosomal structure is a barrier to transcription, DNA replication, and genome-wide DNA repair (Jansen & Verstrepen, 2011). SWI/SNF chromatin remodeling complex can utilize the energy derived from ATP hydrolysis to maintain proper nucleosome organization, thereby controlling major intracellular DNA-based biological processes (Becker & Hörz, 2002; Hohmann & Vakoc, 2014). Phylogenetic analyses and literatures suggest that the components of SWI/SNF complex and their mechanisms of operation are evolutionarily conserved across eukaryotes. However, studies of fungal SWI/SNF complex are limited, especially in phytopathogenic fungi, when compared to those in human and plants. Thus, summarizing and discussing the composition and function of SWI/SNF complex in higher eukaryotes will help advance our understanding of this complex in pathogenic fungi.
Genetic and biochemical experiments have revealed that the SWI/SNF complex is an essential regulator of numerous chromosomal processes, and its dysregulation leads to severe defects in development and stress response across eukaryotes (Clapier et al. 2017; Euskirchen et al. 2012; Jégu et al. 2014; Kwon & Wagner, 2007). This remodeler is built in a modular mode, with specific subunits that interact with or being regulated by specific activators/repressors/covalent histone modifiers or other functional proteins governing diverse stress responses. But how do these specific interactions provide such varied targeting repertoire, and how do they enable particular remodeler outcomes at specific locations? It will be of great interest to discover the SWI/SNF subunit interacting partners, combined with genetic, proteomics, transcriptomic and other high-throughput sequencing techniques, which will open new avenues to characterize a divergent set of fungal SWI/SNFs and their specific biological roles in pathogen-plant interaction. Further characterization of the SWI/SNF complex, including its organization, assembly, 3D structure, interaction partners, molecular regulatory mechanisms, and their roles in pathogen-plant interactions are expected to generate novel strategies that will help develop prevention measurements of plant diseases.
Availability of data and materials
Not applicable.
References
Adkins MW, Williams SK, Linger J, Tyler JK (2007) Chromatin disassembly from the PHO5 promoter is essential for the recruitment of the general transcription machinery and coactivators. Mol Cell Biol 27(18):6372–6382. https://doi.org/10.1128/MCB.00981-07
Aguirre CB (2017) Characterisation of Eucalyptus grandis SWEET and SWI/SNF proteins during symbiosis with Pisolithus microcarpus. Western Sydney University, Australia
Allemand E, Batsché E, Muchardt C (2008) Splicing, transcription, and chromatin: a ménage à trois. Curr Opin Genet Dev 18(2):145–151. https://doi.org/10.1016/j.gde.2008.01.006
Allemand E, Myers MP, Garcia-Bernardo J, Harel-Bellan A, Krainer AR, Muchardt C (2016) A broad set of chromatin factors influences splicing. PLoS Genet 12(9):e1006318. https://doi.org/10.1371/journal.pgen.1006318
Arasimowicz-Jelonek M, Floryszak-Wieczorek J (2016) Nitric oxide in the offensive strategy of fungal and oomycete plant pathogens. Front Plant Sci 7:252. https://doi.org/10.3389/fpls.2016.00252
Archacki R, Buszewicz D, Sarnowski TJ, Sarnowska E, Rolicka AT, Tohge T, Fernie AR, Jikumaru Y, Kotlinski M, Iwanicka-Nowicka R, Kalisiak K, Patryn J, Halibart-Puzio J, Kamiya Y, Davis SJ, Koblowska MK, Jerzmanowski A (2013) BRAHMA ATPase of the SWI/SNF chromatin remodeling complex acts as a positive regulator of gibberellin-mediated responses in Arabidopsis. PLoS One 8(3):e58588. https://doi.org/10.1371/journal.pone.0058588
Ataian Y, Krebs JE (2006) Five repair pathways in one context: chromatin modification during DNA repair. Biochem Cell Biol 84(4):490–504. https://doi.org/10.1139/o06-075
Bao Y, Shen X (2007) Chromatin remodeling in DNA double-strand break repair. Curr Opin Genet Dev 17(2):126–131. https://doi.org/10.1016/j.gde.2007.02.010
Batsché E, Yaniv M, Muchardt C (2006) The human SWI/SNF subunit Brm is a regulator of alternative splicing. Nat Struct Mol Biol 13(1):22–29. https://doi.org/10.1038/nsmb1030
Bayona-Feliu A, Barroso S, Muñoz S (2021) The SWI/SNF chromatin remodeling complex helps resolve R-loop-mediated transcription-replication conflicts 53(7): 1050–1063 doi: https://doi.org/10.1038/s41588-021-00867-2
Becker PB, Hörz W (2002) ATP-dependent nucleosome remodeling. Annu Rev Biochem 71(1):247–273. https://doi.org/10.1146/annurev.biochem.71.110601.135400
Bezhani S, Winter C, Hershman S, Wagner JD, Kennedy JF, Kwon CS, Pfluger J, Su Y, Wagner D (2007) Unique, shared, and redundant roles for the Arabidopsis SWI/SNF chromatin remodeling ATPases BRAHMA and SPLAYED. Plant Cell 19(2):403–416. https://doi.org/10.1105/tpc.106.048272
Bhadouriya SL, Mehrotra S, Basantani MK, Loake GJ, Mehrotra R (2020) Role of chromatin architecture in plant stress responses: an update. Front Plant Sci 11:603380. https://doi.org/10.3389/fpls.2020.603380
Biddick RK, Law GL, Young ET (2008) Adr1 and Cat8 mediate coactivator recruitment and chromatin remodeling at glucose-regulated genes. PLoS One 3(1):e1436. https://doi.org/10.1371/journal.pone.0001436
Black DL (2000) Protein diversity from alternative splicing: a challenge for bioinformatics and post-genome biology. Cell 103(3):367–370. https://doi.org/10.1016/s0092-8674(00)00128-8
Blencowe BJ (2006) Alternative splicing: new insights from global analyses. Cell 126(1):37–47. https://doi.org/10.1016/j.cell.2006.06.023
Bohm KA, Hodges AJ, Czaja W, Selvam K, Smerdon MJ, Mao P, Wyrick JJ (2021) Distinct roles for RSC and SWI/SNF chromatin remodelers in genomic excision repair. Genome Res 31(6):1047–1059. https://doi.org/10.1101/gr.274373.120
Cao Y, Zheng F, Zhang W, Meng X, Liu W (2019) Trichoderma reesei XYR1 recruits SWI/SNF to facilitate cellulase gene expression. Mol Microbiol 112(4):1145–1162. https://doi.org/10.1111/mmi.14352
Capovilla G, Pajoro A, Immink RG, Schmid M (2015) Role of alternative pre-mRNA splicing in temperature signaling. Curr Opin Plant Biol 27:97–103. https://doi.org/10.1016/j.pbi.2015.06.016
Cavellán E, Asp P, Percipalle P, Farrants AK (2006) The WSTF-SNF2h chromatin remodeling complex interacts with several nuclear proteins in transcription. J Biol Chem 281(24):16264–16271. https://doi.org/10.1074/jbc.M600233200
Centore RC, Sandoval GJ, Soares LMM, Kadoch C, Chan HM (2020) Mammalian SWI/SNF chromatin remodeling complexes: emerging mechanisms and therapeutic strategies. Trends Genet 36(12):936–950. https://doi.org/10.1016/j.tig.2020.07.011
Chang YN, Zhu C, Jiang J, Zhang H, Zhu JK, Duan CG (2020) Epigenetic regulation in plant abiotic stress responses. J Integr Plant Biol 62(5):563–580. https://doi.org/10.1111/jipb.12901
Chen W, Zhu Q, Liu Y, Zhang Q (2017) Chromatin remodeling and plant immunity. Adv Protein Chem Struct Biol 106:243–260. https://doi.org/10.1016/bs.apcsb.2016.08.006
Clapier CR, Cairns BR (2009) The biology of chromatin remodeling complexes. Annu Rev Biochem 78(1):273–304. https://doi.org/10.1146/annurev.biochem.77.062706.153223
Clapier CR, Iwasa J, Cairns BR, Peterson CL (2017) Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat Rev Mol Cell Biol 18(7):407–422. https://doi.org/10.1038/nrm.2017.26
Cosma MP, Tanaka T, Nasmyth K (2016) Ordered recruitment of transcription and chromatin remodeling factors to a cell cycle- and developmentally regulated promoter. Cell 166(3):781. https://doi.org/10.1016/j.cell.2016.07.016
Dang W, Bartholomew B (2007) Domain architecture of the catalytic subunit in the ISW2-nucleosome complex. Mol Cell Biol 27(23):8306–8317. https://doi.org/10.1128/MCB.01351-07
Ding B, Wang GL (2015) Chromatin versus pathogens: the function of epigenetics in plant immunity. Front Plant Sci 6:675. https://doi.org/10.3389/fpls.2015.00675
Dutta A, Sardiu M, Gogol M, Gilmore J, Zhang D, Florens L, Abmayr SM, Washburn MP, Workman JL (2017) Composition and function of mutant SWI/SNF complexes. Cell Rep 18(9):2124–2134. https://doi.org/10.1016/j.celrep.2017.01.058
Erkina TY, Tschetter PA, Erkine AM (2008) Different requirements of the SWI/SNF complex for robust nucleosome displacement at promoters of heat shock factor and Msn2- and Msn4-regulated heat shock genes. Mol Cell Biol 28(4):1207–1217. https://doi.org/10.1128/MCB.01069-07
Euskirchen G, Auerbach RK, Snyder M (2012) SWI/SNF chromatin-remodeling factors: multiscale analyses and diverse functions. J Biol Chem 287(37):30897–30905. https://doi.org/10.1074/jbc.R111.309302
Fair BJ, Pleiss JA (2017) The power of fission: yeast as a tool for understanding complex splicing. Curr Genet 63(3):375–380. https://doi.org/10.1007/s00294-016-0647-6
Feng J, Xu X, Fan X, Yi Q, Tang L (2021) BAF57/SMARCE1 interacting with splicing factor srsf1 regulates mechanical stress-induced alternative splicing of cyclin D1. Genes (Basel) 12(2):306. https://doi.org/10.3390/genes12020306
Ghosh S, Pugh BF (2011) Sequential recruitment of SAGA and TFIID in a genomic response to DNA damage in Saccharomyces cerevisiae. Mol Cell Biol 31(1):190–202. https://doi.org/10.1128/MCB.00317-10
Gong F, Fahy D, Smerdon MJ (2006) Rad4-Rad23 interaction with SWI/SNF links ATP-dependent chromatin remodeling with nucleotide excision repair. Nat Struct Mol Biol 13(1):902–907. https://doi.org/10.1038/nsmb1152
Gresh L, Bourachot B, Reimann A, Guigas B, Fiette L, Garbay Set al (2005) The SWI/SNF chromatin-remodeling complex subunit SNF5 is essential for hepatocyte differentiation. EMBO J 24(18): 3313–3324. doi: https://doi.org/10.1038/sj.emboj.7600802
Hainer SJ, Pruneski JA, Mitchell RD, Monteverde RM, Martens JA (2011) Intergenic transcription causes repression by directing nucleosome assembly. Genes Dev 25(1):29–40. https://doi.org/10.1101/gad.1975011
Han SK, Sang Y, Rodrigues A, Wu MF, Rodriguez PL, Wagner D (2012) The SWI2/SNF2 chromatin remodeling ATPase BRAHMA represses abscisic acid responses in the absence of the stress stimulus in Arabidopsis. Plant Cell 24(12):4892–4906. https://doi.org/10.1105/tpc.112.105114
Harrod A, Lane KA, Downs JA (2020) The role of the SWI/SNF chromatin remodelling complex in the response to DNA double strand breaks. DNA Repair (Amst) 93:102919. https://doi.org/10.1016/j.dnarep.2020.102919
Hassan AH, Neely KE, Workman JL (2001) Histone acetyltransferase complexes stabilize SWI/SNF binding to promoter nucleosomes. Cell 104(6):817–827. https://doi.org/10.1016/s0092-8674(01)00279-3
Hauer MH, Gasser SM (2017) Chromatin and nucleosome dynamics in DNA damage and repair. Genes Dev 31(22):2204–2221. https://doi.org/10.1101/gad.307702.117
Hauer MH, Seeber A, Singh V, Thierry R, Sack R, Amitai A, Kryzhanovska M, Eglinger J, Holcman D, Owen-Hughes T, Gasser SM (2017) Histone degradation in response to DNA damage enhances chromatin dynamics and recombination rates. Nat Struct Mol Biol 24(2):99–107. https://doi.org/10.1038/nsmb.3347
Hays E, Nettleton E, Carter C, Morales M, Vo L, Passo M, Vélez-Cruz R (2020) The SWI/SNF ATPase BRG1 stimulates DNA end resection and homologous recombination by reducing nucleosome density at DNA double strand breaks and by promoting the recruitment of the CtIP nuclease. Cell Cycle 19(22):3096–3114. https://doi.org/10.1080/15384101.2020.1831256
Hohmann AF, Vakoc CR (2014) A rationale to target the SWI/SNF complex for cancer therapy. Trends Genet 30(8):356–363. https://doi.org/10.1016/j.tig.2014.05.001
Huang CY, Rangel DS, Qin X, Bui C, Li R, Jia Z, Cui X, Jin H (2021) The chromatin-remodeling protein BAF60/SWP73A regulates the plant immune receptor NLRs. Cell Host Microbe 29(3):425–434.e424. https://doi.org/10.1016/j.chom.2021.01.005
Iba H, Mizutani T, Ito T (2003) SWI/SNF chromatin remodelling complex and retroviral gene silencing. Rev Med Virol 13(2):99–110. https://doi.org/10.1002/rmv.378
Ito T, Watanabe H, Yamamichi N, Kondo S, Tando T, Haraguchi T, Mizutani T, Sakurai K, Fujita S, Izumi T, Isobe T, Iba H (2008) Brm transactivates the telomerase reverse transcriptase (TERT) gene and modulates the splicing patterns of its transcripts in concert with p54(nrb). Biochem J 411(1):201–209. https://doi.org/10.1042/BJ20071075
Jansen A, Verstrepen KJ (2011) Nucleosome positioning in Saccharomyces cerevisiae. Microbiol Mol Biol Rev 75(2):301–320. https://doi.org/10.1128/MMBR.00046-10
Jarończyk K, Sosnowska K, Zaborowski A, Pupel P, Bucholc M, Małecka E, Siwirykow N, Stachula P, Iwanicka-Nowicka R, Koblowska M, Jerzmanowski A, Archacki R (2021) Bromodomain-containing subunits BRD1, BRD2, and BRD13 are required for proper functioning of SWI/SNF complexes in Arabidopsis. Plant Commun 2(4):100174. https://doi.org/10.1016/j.xplc.2021.100174.
Jégu T, Latrasse D, Delarue M, Hirt H, Domenichini S, Ariel F, Crespi M, Bergounioux C, Raynaud C, Benhamed M (2014) The BAF60 subunit of the SWI/SNF chromatin-remodeling complex directly controls the formation of a gene loop at FLOWERING LOCUS C in Arabidopsis. Plant Cell 26(2):538–551. https://doi.org/10.1105/tpc.113.114454
Jian Y, Liu Z, Wang H, Chen Y, Yin Y, Zhao Y (2021) Interplay of two transcription factors for recruitment of the chromatin remodeling complex modulates fungal nitrosative stress response. Nat Commun 12(1):2576. https://doi.org/10.1038/s41467-021-22831-8
Jiang J, Mao N, Hu H, Tang J, Han D, Liu S, Wu Q, Liu Y, Peng C, Lai J, Yang C (2019) A SWI/SNF subunit regulates chromosomal dissociation of structural maintenance complex 5 during DNA repair in plant cells. Proc Natl Acad Sci 116(30):15288–15296. https://doi.org/10.1073/pnas.1900308116
Johnson KC, Xia S, Feng X, Li X (2015) The chromatin remodeler SPLAYED negatively regulates SNC1-mediated immunity. Plant Cell Physiol 56(8):1616–1623. https://doi.org/10.1093/pcp/pcv087
Kadota Y, Shirasu K, Zipfel C (2015) Regulation of the NADPH oxidase RBOHD during plant immunity. Plant Cell Physiol 56(8):1472–1480. https://doi.org/10.1093/pcp/pcv063
Kasten MM, Clapier CR, Cairns BR (2011) SnapShot: chromatin remodeling: SWI/SNF. Cell 144(2):310.e311. https://doi.org/10.1016/j.cell.2011.01.007
Kornblihtt AR (2006) Chromatin, transcript elongation and alternative splicing. Nat Struct Mol Biol 13(1):5–7. https://doi.org/10.1038/nsmb0106-5
Kwon CS, Wagner D (2007) Unwinding chromatin for development and growth: a few genes at a time. Trends Genet 23(8):403–412. https://doi.org/10.1016/j.tig.2007.05.010
Kwon SJ, Park JH, Park EJ, Lee SA, Lee HS, Kang SW, Kwon J (2015) ATM-mediated phosphorylation of the chromatin remodeling enzyme BRG1 modulates DNA double-strand break repair. Oncogene 34(3):303–313. https://doi.org/10.1038/onc.2013.556
Łaźniewska J, Macioszek VK, Kononowicz AK (2012) Plant-fungus interface: the role of surface structures in plant resistance and susceptibility to pathogenic fungi. Physiol Mol Plant P 78:24–30. https://doi.org/10.1016/j.pmpp.2012.01.004
Lee HS, Park JH, Kim SJ, Kwon SJ, Kwon J (2010) A cooperative activation loop among SWI/SNF, gamma-H2AX and H3 acetylation for DNA double-strand break repair. EMBO J 29(8):1434–1445. https://doi.org/10.1038/emboj.2010.27
Li C, Gu L, Gao L, Chen C, Wei CQ, Qiu Q, Chien CW, Wang S, Jiang L, Ai LF, Chen CY, Yang S, Nguyen V, Qi Y, Snyder MP, Burlingame AL, Kohalmi SE, Huang S, Cao X, Wang ZY, Wu K, Chen X, Cui Y (2016) Concerted genomic targeting of H3K27 demethylase REF6 and chromatin-remodeling ATPase BRM in Arabidopsis. Nat Genet 48(6):687–693. https://doi.org/10.1038/ng.3555
Li X, Jiang Y, Ji Z, Liu Y, Zhang Q (2015) BRHIS1 suppresses rice innate immunity through binding to monoubiquitinated H2A and H2B variants. EMBO Rep 16(9):1192–1202. https://doi.org/10.15252/embr.201440000
Lin J, Zhao Y, Ferraro AR, Yang E, Lewis ZA, Lin X (2019) Transcription factor Znf2 coordinates with the chromatin remodeling SWI/SNF complex to regulate Cryptococcal cellular differentiation. Commun Biol 2(1):412. https://doi.org/10.1038/s42003-019-0665-2
Lindahl T (2000) Suppression of spontaneous mutagenesis in human cells by DNA base excision-repair. Mutat Res 462(2–3):129–135. https://doi.org/10.1016/s1383-5742(00)00024-7
Liu C, Xin Y, Xu L, Cai Z, Xue Y, Liu Y, Xie D, Liu Y, Qi Y (2018a) Arabidopsis ARGONAUTE 1 binds chromatin to promote gene transcription in response to hormones and stresses. Dev Cell 44(3):348–361.e347. https://doi.org/10.1016/j.devcel.2017.12.002
Liu L, Wang Q, Sun Y, Zhang Y, Zhang X, Liu J, Yu G, Pan H (2018b) Sssfh1, a gene encoding a putative component of the RSC chromatin remodeling complex, is involved in hyphal growth, reactive oxygen species accumulation, and pathogenicity in Sclerotinia sclerotiorum. Front Microbiol 9:1828. https://doi.org/10.3389/fmicb.2018.01828
Liu Z, Jian Y, Chen Y, Kistler HC, He P, Ma Z (2019) A phosphorylated transcription factor regulates sterol biosynthesis in fusarium graminearum. Nat Commun 10(1):1228. https://doi.org/10.1038/s41467-019-09145-6
Liu Z, Myers LC (2017) Candida albicans SWI/SNF and mediator complexes differentially regulate Mrr1-induced MDR1 expression and fluconazole resistance. Antimicrob Agents Chemother 61(11):e01344–e01317. https://doi.org/10.1128/AAC.01344-17
Lu Y, Su C, Mao X, Raniga PP, Liu H, Chen J (2008) Efg1-mediated recruitment of NuA4 to promoters is required for hypha-specific SWI/SNF binding and activation in Candida albicans. Mol Biol Cell 19(10):4260–4272. https://doi.org/10.1091/mbc.e08-02-0173
Lu Y, Tan F, Zhao Y, Zhou SL, Chen XS, Hu YF, Zhou DX (2020) A chromodomain-helicase-DNA-binding factor functions in chromatin modification and gene regulation. Plant Physiol 183(3):1035–1046. https://doi.org/10.1104/pp.20.00453
Luco RF, Allo M, Schor IE, Kornblihtt AR, Misteli T (2011) Epigenetics in alternative pre-mRNA splicing. Cell 144(1):16–26. https://doi.org/10.1016/j.cell.2010.11.056
Luger K, Dechassa ML, Tremethick DJ (2012) New insights into nucleosome and chromatin structure: an ordered state or a disordered affair? Nat Rev Mol Cell Biol 13(7):436–447. https://doi.org/10.1038/nrm3382
Lukas J, Lukas C, Bartek J (2011) More than just a focus: the chromatin response to DNA damage and its role in genome integrity maintenance. Nat Cell Biol 13(10):1161–1169. https://doi.org/10.1038/ncb2344
Makarov EM, Owen N, Bottrill A, Makarova OV (2012) Functional mammalian spliceosomal complex E contains SMN complex proteins in addition to U1 and U2 snRNPs. Nucleic Acids Res 40(6):2639–2652. https://doi.org/10.1093/nar/gkr1056
Mao X, Cao F, Nie X, Liu H, Chen J (2006) The SWI/SNF chromatin remodeling complex is essential for hyphal development in Candida albicans. FEBS Lett 580(11):2615–2622. https://doi.org/10.1016/j.febslet.2006.04.009
Masliah-Planchon J, Bièche I, Guinebretière JM, Bourdeaut F, Delattre O (2015) SWI/SNF chromatin remodeling and human malignancies. Annu Rev Pathol 10(1):145–171. https://doi.org/10.1146/annurev-pathol-012414-040445
Maury S, Sow MD, Le Gac AL, Genitoni J, Lafon-Placette C, Mozgova I (2019) Phytohormone and chromatin crosstalk: the missing link for developmental plasticity? Front Plant Sci 10:395. https://doi.org/10.3389/fpls.2019.00395
Menezes RA, Pimentel C, Silva AR, Amaral C, Merhej J, Devaux F et al (2017) Mediator, SWI/SNF and SAGA complexes regulate Yap8-dependent transcriptional activation of ACR2 in response to arsenate. Biochim Biophys Acta Gene Regul Mech 1860(4):472–481. https://doi.org/10.1016/j.bbagrm.2017.02.001
Miné-Hattab J, Rothstein R (2012) Increased chromosome mobility facilitates homology search during recombination. Nat Cell Biol 14(5):510–517. https://doi.org/10.1038/ncb2472
Mittal P, Roberts CWM (2020) The SWI/SNF complex in cancer - biology, biomarkers and therapy. Nat Rev Clin Oncol 17(7):435–448. https://doi.org/10.1038/s41571-020-0357-3
Neigeborn L, Carlson M (1984) Genes affecting the regulation of SUC2 gene expression by glucose repression in Saccharomyces cerevisiae. Genetics 108(4):845–858. https://doi.org/10.1093/genetics/108.4.845
Ogiwara H, Ui A, Otsuka A, Satoh H, Yokomi I, Nakajima S, Yasui A, Yokota J, Kohno T (2011) Histone acetylation by CBP and p300 at double-strand break sites facilitates SWI/SNF chromatin remodeling and the recruitment of non-homologous end joining factors. Oncogene 30(18):2135–2146. https://doi.org/10.1038/onc.2010.592
Oh J, Sohn DH, Ko M, Chung H, Jeon SH, Seong RH (2008) BAF60a interacts with p53 to recruit the SWI/SNF complex. J Biol Chem 283(18):11924–11934. https://doi.org/10.1074/jbc.M705401200
Ojolo SP, Cao S, Priyadarshani S, Li W, Yan M, Aslam M et al (2018) Regulation of plant growth and development: a review from a chromatin remodeling perspective. Front Plant Sci 9:1232. https://doi.org/10.3389/fpls.2018.01232
Park JH, Park EJ, Lee HS, Kim SJ, Hur SK, Imbalzano AN, Kwon J (2006) Mammalian SWI/SNF complexes facilitate DNA double-strand break repair by promoting gamma-H2AX induction. EMBO J 25(17):3986–3997. https://doi.org/10.1038/sj.emboj.7601291
Patrick KL, Ryan CJ, Xu J, Lipp JJ, Nissen KE, Roguev A, Shales M, Krogan NJ, Guthrie C (2015) Genetic interaction mapping reveals a role for the SWI/SNF nucleosome remodeler in spliceosome activation in fission yeast. PLoS Genet 11(3):e1005074. https://doi.org/10.1371/journal.pgen.1005074
Peirats-Llobet M, Han SK, Gonzalez-Guzman M, Jeong CW, Rodriguez L, Belda-Palazon B, Wagner D, Rodriguez PL (2016) A direct link between abscisic acid sensing and the chromatin-remodeling ATPase BRAHMA via core ABA signaling pathway components. Mol Plant 9(1):136–147. https://doi.org/10.1016/j.molp.2015.10.003
Peng G, Yim EK, Dai H, Jackson AP, Burgt I, Pan MR et al (2009) BRIT1/MCPH1 links chromatin remodelling to DNA damage response. Nat Cell Biol 11(7):865–872. https://doi.org/10.1038/ncb1895
Peterson CL, Workman JL (2000) Promoter targeting and chromatin remodeling by the SWI/SNF complex. Curr Opin Genet Dev 10(2):187–192. https://doi.org/10.1016/s0959-437x(00)00068-x
Pfluger J, Wagner D (2007) Histone modifications and dynamic regulation of genome accessibility in plants. Curr Opin Plant Biol 10(6):645–652. https://doi.org/10.1016/j.pbi.2007.07.013
Phelan ML, Sif S, Narlikar GJ, Kingston RE (1999) Reconstitution of a core chromatin remodeling complex from SWI/SNF subunits. Mol Cell 3(2):247–253. https://doi.org/10.1016/s1097-2765(00)80315-9
Pieterse CM, Leon-Reyes A, Van der Ent S, Van Wees SC (2009) Networking by small-molecule hormones in plant immunity. Nat Chem Biol 5(5):308–316. https://doi.org/10.1038/nchembio.164
Pleiss JA, Whitworth GB, Bergkessel M, Guthrie C (2007) Rapid, transcript-specific changes in splicing in response to environmental stress. Mol Cell 27(6):928–937. https://doi.org/10.1016/j.molcel.2007.07.018
Pray-Grant MG, Daniel JA, Schieltz D, Yates JR 3rd, Grant PA (2005) Chd1 chromodomain links histone H3 methylation with SAGA- and SLIK-dependent acetylation. Nature 433(7024):434–438. https://doi.org/10.1038/nature03242
Prochasson P, Neely KE, Hassan AH, Li B, Workman JL (2003) Targeting activity is required for SWI/SNF function in vivo and is accomplished through two partially redundant activator-interaction domains. Mol Cell 12(4):983–990. https://doi.org/10.1016/s1097-2765(03)00366-6
Proft M, Struhl K (2002) Hog1 kinase converts the Sko1-Cyc8-Tup1 repressor complex into an activator that recruits SAGA and SWI/SNF in response to osmotic stress. Mol Cell 9(6):1307–1317. https://doi.org/10.1016/s1097-2765(02)00557-9
Ramirez-Prado JS, Benhamed M (2021) New partners for old friends: plant SWI/SNF complexes. Mol Plant 14(6):870–872. https://doi.org/10.1016/j.molp.2021.05.017
Ramirez-Prado JS, Piquerez SJM, Bendahmane A, Hirt H, Raynaud C, Benhamed M (2018) Modify the histone to win the battle: chromatin dynamics in plant-pathogen interactions. Front Plant Sci 9:355. https://doi.org/10.3389/fpls.2018.00355
Reyes JC (2014) The many faces of plant SWI/SNF complex. Mol Plant 7(3):454–458. https://doi.org/10.1093/mp/sst147
Ribeiro-Silva C, Aydin ÖZ, Mesquita-Ribeiro R, Slyskova J, Helfricht A, Marteijn JA, Hoeijmakers JHJ, Lans H, Vermeulen W (2018) DNA damage sensitivity of SWI/SNF-deficient cells depends on TFIIH subunit p62/GTF2H1. Nat Commun 9(1):4067. https://doi.org/10.1038/s41467-018-06402-y
Ribeiro-Silva C, Vermeulen W, Lans H (2019) SWI/SNF: complex complexes in genome stability and cancer. DNA Repair (Amst) 77:87–95. https://doi.org/10.1016/j.dnarep.2019.03.007
Roberts CW, Orkin SH (2004) The SWI/SNF complex-chromatin and cancer. Nat Rev Cancer 4(2):133–142. https://doi.org/10.1038/nrc1273
Roukos V, Voss TC, Schmidt CK, Lee S, Wangsa D, Misteli T (2013) Spatial dynamics of chromosome translocations in living cells. Science 341(6146):660–664. https://doi.org/10.1126/science.1237150
Saez A, Rodrigues A, Santiago J, Rubio S, Rodriguez PL (2008) HAB1-SWI3B interaction reveals a link between abscisic acid signaling and putative SWI/SNF chromatin-remodeling complexes in Arabidopsis. Plant Cell 20(11):2972–2988. https://doi.org/10.1105/tpc.107.056705
Sakamoto T, Tsujimoto-Inui Y, Sotta N (2018) Proteasomal degradation of BRAHMA promotes boron tolerance in Arabidopsis. Nat Commun 9(1):5285. https://doi.org/10.1038/s41467-018-07393-6
Sanz AB, García R, Rodríguez-Peña JM, Díez-Muñiz S, Nombela C, Peterson CL, Arroyo J (2012) Chromatin remodeling by the SWI/SNF complex is essential for transcription mediated by the yeast cell wall integrity MAPK pathway. Mol Biol Cell 23(14):2805–2817. https://doi.org/10.1091/mbc.E12-04-0278
Sarnowska E, Gratkowska DM, Sacharowski SP, Cwiek P, Tohge T, Fernie AR, Siedlecki JA, Koncz C, Sarnowski TJ (2016) The role of SWI/SNF chromatin remodeling complexes in hormone crosstalk. Trends Plant Sci 21(7):594–608. https://doi.org/10.1016/j.tplants.2016.01.017
Schwabish MA, Struhl K (2007) The SWI/SNF complex is important for histone eviction during transcriptional activation and RNA polymerase II elongation in vivo. Mol Cell Biol 27(20):6987–6995. https://doi.org/10.1128/MCB.00717-07
Schwartz S, Meshorer E, Ast G (2009) Chromatin organization marks exon-intron structure. Nat Struct Mol Biol 16(9):990–995. https://doi.org/10.1038/nsmb.1659
Shanle EK, Andrews FH, Meriesh H, McDaniel SL, Dronamraju R, DiFiore JV et al (2015) Association of Taf14 with acetylated histone H3 directs gene transcription and the DNA damage response. Genes Dev 29(17):1795–1800. https://doi.org/10.1101/gad.269977.115
Shivaswamy S, Iyer VR (2008) Stress-dependent dynamics of global chromatin remodeling in yeast: dual role for SWI/SNF in the heat shock stress response. Mol Cell Biol 28(7):2221–2234. https://doi.org/10.1128/MCB.01659-07
Shu J, Chen C, Li C, Thapa RK, Song J, Xie X, Nguyen V, Bian S, Liu J, Kohalmi SE, Cui Y (2021) Genome-wide occupancy of Arabidopsis SWI/SNF chromatin remodeler SPLAYED provides insights into its interplay with its close homolog BRAHMA and polycomb proteins. Plant J 106(1):200–213. https://doi.org/10.1111/tpj.15159
Sinha M, Watanabe S, Johnson A, Moazed D, Peterson CL (2009) Recombinational repair within heterochromatin requires ATP-dependent chromatin remodeling. Cell 138(6):1109–1121. https://doi.org/10.1016/j.cell.2009.07.013
Smeenk G, van Attikum H (2013) The chromatin response to DNA breaks: leaving a mark on genome integrity. Annu Rev Biochem 82(1):55–80. https://doi.org/10.1146/annurev-biochem-061809-174504
Smith CL, Horowitz-Scherer R, Flanagan JF, Woodcock CL, Peterson CL (2003) Structural analysis of the yeast SWI/SNF chromatin remodeling complex. Nat Struct Biol 10(2):141–145. https://doi.org/10.1038/nsb888
Song ZT, Liu JX, Han JJ (2021) Chromatin remodeling factors regulate environmental stress responses in plants. J Integr Plant Biol 63(3):438–450. https://doi.org/10.1111/jipb.13064
Spivak G (2015) Nucleotide excision repair in humans. DNA Repair (Amst) 36:13–18. https://doi.org/10.1016/j.dnarep.2015.09.003
Sudarsanam P, Iyer VR, Brown PO, Winston F (2000) Whole-genome expression analysis of snf/swi mutants of Saccharomyces cerevisiae. Proc Natl Acad Sci 97(7):3364–3369. https://doi.org/10.1073/pnas.050407197
Tebbji F, Chen Y, Sellam A, Whiteway M (2017) The genomic landscape of the fungus-specific SWI/SNF complex subunit, Snf6, in Candida albicans. mSphere 2(6):e00497-00517. 2(6). https://doi.org/10.1128/mSphere.00497-17
Thouly C, Le Masson M, Lai X, Carles CC (2020) Unwinding BRAHMA functions in plants. Genes 11(1):90. https://doi.org/10.3390/genes11010090
Tyagi A, Ryme J, Brodin D, Ostlund Farrants AK, Visa N (2009) SWI/SNF associates with nascent pre-mRNPs and regulates alternative pre-mRNA processing. PLoS Genet 5(5):e1000470. https://doi.org/10.1371/journal.pgen.1000470
Ui A, Ogiwara H, Nakajima S, Kanno S, Watanabe R, Harata M, Okayama H, Harris CC, Yokota J, Yasui A, Kohno T (2014) Possible involvement of LKB1-AMPK signaling in non-homologous end joining. Oncogene 33(13):1640–1648. https://doi.org/10.1038/onc.2013.125
Venkataramanan S, Douglass S, Galivanche AR, Johnson TL (2017) The chromatin remodeling complex SWI/SNF regulates splicing of meiotic transcripts in Saccharomyces cerevisiae. Nucleic Acids Res 45(13):7708–7721. https://doi.org/10.1093/nar/gkx373
Vicente J, Mendiondo GM, Movahedi M, Peirats-Llobet M, Juan YT, Shen YY, Dambire C, Smart K, Rodriguez PL, Charng YY, Gray JE, Holdsworth MJ (2017) The Cys-Arg/N-end rule pathway is a general sensor of abiotic stress in flowering plants. Curr Biol 27(20):3183–3190. https://doi.org/10.1016/j.cub.2017.09.006
Vongs A, Kakutani T, Martienssen RA, Richards EJ (1993) Arabidopsis thaliana DNA methylation mutants. Science 260(5116):1926–1928. https://doi.org/10.1126/science.8316832
Wagner D, Meyerowitz EM (2002) SPLAYED, a novel SWI/SNF ATPase homolog, controls reproductive development in Arabidopsis. Curr Biol 12(2):85–94. https://doi.org/10.1016/s0960-9822(01)00651-0
Waldholm J, Wang Z, Brodin D, Tyagi A, Yu S, Theopold U, Farrants AKÖ, Visa N (2011) SWI/SNF regulates the alternative processing of a specific subset of pre-mRNAs in Drosophila melanogaster. BMC Mol Biol 12(1):46. https://doi.org/10.1186/1471-2199-12-46
Walley JW, Rowe HC, Xiao Y, Chehab EW, Kliebenstein DJ, Wagner D, Dehesh K (2008) The chromatin remodeler SPLAYED regulates specific stress signaling pathways. PLoS Pathog 4(12):e1000237. https://doi.org/10.1371/journal.ppat.1000237
Wang B, Kettenbach AN, Gerber SA, Loros JJ, Dunlap JC (2014) Neurospora WC-1 recruits SWI/SNF to remodel frequency and initiate a circadian cycle. PLoS Genet 10(9):e1004599. https://doi.org/10.1371/journal.pgen.1004599
Wang S, Wu XM, Liu CH, Shang JY, Gao F, Guo HS (2020) Verticillium dahliae chromatin remodeling facilitates the DNA damage repair in response to plant ROS stress. PLoS Pathog 16(4):e1008481. https://doi.org/10.1371/journal.ppat.1008481
Wang TY, Wang YX, You CJ (2021) Structural and functional characteristics of plant PHD domain-containing proteins. Yi Chuan 43(4):323–339. https://doi.org/10.16288/j.yczz.20-412
Wiest NE, Houghtaling S, Sanchez JC, Tomkinson AE, Osley MA (2017) The SWI/SNF ATP-dependent nucleosome remodeler promotes resection initiation at a DNA double-strand break in yeast. Nucleic Acids Res 45(10):5887–5900. https://doi.org/10.1093/nar/gkx221
Wu MF, Yamaguchi N, Xiao J, Bargmann B, Estelle M, Sang Y et al (2015) Auxin-regulated chromatin switch directs acquisition of flower primordium founder fate. eLife 4:e09269. https://doi.org/10.7554/eLife.09269
Yu X, Meng X, Liu Y, Wang X, Wang TJ, Zhang A, Li N, Qi X, Liu B, Xu ZY (2019) The chromatin remodeler ZmCHB101 impacts alternative splicing contexts in response to osmotic stress. Plant Cell Rep 38(2):131–145. https://doi.org/10.1007/s00299-018-2354-x
Yudkovsky N, Logie C, Hahn S, Peterson CL (1999) Recruitment of the SWI/SNF chromatin remodeling complex by transcriptional activators. Genes Dev 13(18):2369–2374. https://doi.org/10.1101/gad.13.18.2369
Zapater AG, Mackowiak S, Guo Y, Jordan-Pla A, Friedländer M, Visa N et al (2019) The SWI/SNF subunits BRG1 affects alternative splicing by changing RNA binding factor interactions with RNA. bioRxiv. https://doi.org/10.1101/858852
Zhang J, Lai J, Wang F, Yang S, He Z, Jiang J (2017) A SUMO ligase AtMMS21 regulates the stability of the chromatin remodeler BRAHMA in root development. Plant Physiol 173(3):1574–1582. https://doi.org/10.1104/pp.17.00014
Zhang Y, Goritschnig S, Dong X, Li X (2003) A gain-of-function mutation in a plant disease resistance gene leads to constitutive activation of downstream signal transduction pathways in suppressor of npr1-1, constitutive 1. Plant Cell 15(11):2636–2646. https://doi.org/10.1105/tpc.015842
Zhang Z, Wang X, Xin J, Ding Z, Liu S, Fang Q, Yang N, Xu RM, Cai G (2018) Architecture of SWI/SNF chromatin remodeling complex. Protein cell 9(12):1045–1049. https://doi.org/10.1007/s13238-018-0524-9
Zhao Q, Wang QE, Ray A, Wani G, Han C, Milum K, Wani AA (2009) Modulation of nucleotide excision repair by mammalian SWI/SNF chromatin-remodeling complex. J Biol Chem 284(44):30424–30432. https://doi.org/10.1074/jbc.M109.044982
Zhu Y, Rowley MJ, Böhmdorfer G, Wierzbicki AT (2013) A SWI/SNF chromatin-remodeling complex acts in noncoding RNA-mediated transcriptional silencing. Mol Cell 49(2):298–309. https://doi.org/10.1016/j.molcel.2012.11.011
Zraly CB, Dingwall AK (2012) The chromatin remodeling and mRNA splicing functions of the Brahma (SWI/SNF) complex are mediated by the SNR1/SNF5 regulatory subunit. Nucleic Acids Res 40(13):5975–5987. https://doi.org/10.1093/nar/gks288
Zraly CB, Middleton FA, Dingwall AK (2006) Hormone-response genes are direct in vivo regulatory targets of Brahma (SWI/SNF) complex function. J Biol Chem 281(46):35305–35315. https://doi.org/10.1074/jbc.M607806200
Acknowledgements
We thank Chao Liu and Dr. Zunyong Liu for their help on constructing the phylogenetic tree.
Author details
1 State Key Laboratory of Rice Biology, and Key Laboratory of Molecular Biology of Crop Pathogens and Insects, Institute of Biotechnology, Zhejiang University, Hangzhou 310,058, China. 2 Department of Plant Pathology and Microbiology, Texas A&M University, College Station, TX, USA.
Funding
This work was supported by Science and Technology Project of Zhejiang Province (2018C02G2011110), China Postdoctoral Science Foundation (2021 M692849), National Natural Science Foundation of China (31930088) and China Agriculture Research System of MOF and MARAC (CARS-3-1-29).
Author information
Authors and Affiliations
Contributions
The manuscript was prepared by Y. J, W. S, and Z. M.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jian, Y., Shim, WB. & Ma, Z. Multiple functions of SWI/SNF chromatin remodeling complex in plant-pathogen interactions. Stress Biology 1, 18 (2021). https://doi.org/10.1007/s44154-021-00019-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44154-021-00019-w