Abstract
Background
Recent advances in proteomics technologies such as two-hybrid, phage display and mass spectrometry have enabled us to create a detailed map of biomolecular interaction networks. Initial mapping efforts have already produced a wealth of data. As the size of the interaction set increases, databases and computational methods will be required to store, visualize and analyze the information in order to effectively aid in knowledge discovery.
Results
This paper describes a novel graph theoretic clustering algorithm, "Molecular Complex Detection" (MCODE), that detects densely connected regions in large protein-protein interaction networks that may represent molecular complexes. The method is based on vertex weighting by local neighborhood density and outward traversal from a locally dense seed protein to isolate the dense regions according to given parameters. The algorithm has the advantage over other graph clustering methods of having a directed mode that allows fine-tuning of clusters of interest without considering the rest of the network and allows examination of cluster interconnectivity, which is relevant for protein networks. Protein interaction and complex information from the yeast Saccharomyces cerevisiae was used for evaluation.
Conclusion
Dense regions of protein interaction networks can be found, based solely on connectivity data, many of which correspond to known protein complexes. The algorithm is not affected by a known high rate of false positives in data from high-throughput interaction techniques. The program is available from ftp://ftp.mshri.on.ca/pub/BIND/Tools/MCODE.
Similar content being viewed by others
Background
Recent papers published in Science and Nature among others describe large-scale proteomics experiments that have generated large data sets of protein-protein interactions and molecular complexes [1–7]. Protein structure [8] and gene expression data [9] is also accumulating at a rapid rate. Bioinformatics systems for storage, management, visualization and analysis of this new wealth of data must keep pace. We previously published a simple graph theory method that identified a functional protein complex around the yeast protein Las17 that is involved in actin cytoskeleton rearrangement [10]. Here we extend the method to better apply it to the accumulating information in protein networks.
Currently, most proteomics data is available for the model organism Saccharomyces cerevisiae, by virtue of the availability of a defined and relatively stable proteome, full genome clone libraries [11], established molecular biology experimental techniques and an assortment of well designed genomics databases [12–14]. Using the Biomolecular Interaction Network Database (BIND – http://www.bind.ca) [15] as an integration platform, we have collected 15,143 yeast protein-protein interactions among 4,825 proteins (about 75% of the yeast proteome). Much larger data sets than this will eventually be available for other well studied model organisms as well as for the human proteome. These complex data sets present a formidable challenge for computational biology to develop automated data mining analyses for knowledge discovery.
Here we present the first report that uses a clustering algorithm to identify molecular complexes in a large protein interaction network derived from heterogeneous experimental sources. Based on our previous observation that highly interconnected, or dense, regions of the network may represent complexes [10], the "Molecular Complex Detection" (MCODE) algorithm has been implemented and evaluated on our yeast protein interaction compilation using known molecular complex data from a recent systematic mass spectrometry study of the proteome [7] and from the MIPS database [13].
Predicting molecular complexes from protein interaction data is important because it provides another level of functional annotation above other guilt-by-association methods. Since sub-units of a molecular complex generally function towards the same biological goal, prediction of an unknown protein as part of a complex also allows increased confidence in the annotation of that protein.
MCODE also makes the visualization of large networks manageable by extracting the dense regions around a protein of interest. This is important, as it is now obvious that the current visualization tools present on many interaction databases [15], originally based on the Sun Microsystems embedded spring graph layout Java applet do not scale well to large networks (http://java.sun.com/applets/jdk/1.1/demo/GraphLayout/example1.html).
Algorithm
The MCODE algorithm operates in three stages, vertex weighting, complex prediction and optionally post-processing to filter or add proteins in the resulting complexes by certain connectivity criteria.
A network of interacting molecules can be intuitively modeled as a graph, where vertices are molecules and edges are molecular interactions. If temporal pathway or cell signalling information is known, it is possible to create a directed graph with arcs representing direction of chemical action or direction of information flow, otherwise an undirected graph is used. Using this graph representation of a biological system allows graph theoretic methods to be applied to aid in analysis and solve biological problems. This graph theory approach has been used by other biomolecular interaction database projects such as DIP [16], CSNDB [17], TRANSPATH [18], EcoCyc [19] and WIT [20] and is discussed by Wagner and Fell [21].
Algorithms for finding clusters, or locally dense regions, of a graph are an ongoing research topic in computer science and are often based on network flow/minimum cut theory [22, 23] or more recently, spectral clustering [24]. To find locally dense regions of a graph, MCODE instead uses a vertex-weighting scheme based on the clustering coefficient, Ci, which measures 'cliquishness' of the neighborhood of a vertex [25]. Ci = 2n/k i (k i -1) where k i is the vertex size of the neighborhood of vertex i and n is the number of edges in the neighborhood (the immediate neighborhood density of v not including v). A clique is defined as a maximally connected graph. There is no standard graph theory definition of density, but definitions are normally based on the connectivity level of a graph. Density of a graph, G = (V,E), with number of vertices, |V|, and number of edges, |E|, is defined here as |E|; divided by the theoretical maximum number of edges possible for the graph, |E|max. For a graph with loops (an edge connecting back to its originating vertex), |E|max = |V| (|V|+1)/2 and for a graph with no loops, |E|max = |V| (|V|-1)/2. So, density of G, DG = |E|/|E|max and is thus a real number ranging from 0.0 to 1.0.
The first stage of MCODE, vertex weighting, weights all vertices based on their local network density using the highest k-core of the vertex neighborhood. A k-core is a graph of minimal degree k (graph G, for all v in G, deg(v) >= k). The highest k-core of a graph is the central most densely connected subgraph. We define here the term core-clustering coefficient of a vertex, v, to be the density of the highest k-core of the immediate neighborhood of v (vertices connected directly to v) including v (note that Ci does not include v). The core-clustering coefficient is used here instead of the clustering coefficient because it amplifies the weighting of heavily interconnected graph regions while removing the many less connected vertices that are usually part of a biomolecular interaction network, known to be scale-free [6, 21, 26–29]. A scale-free network has a vertex connectivity distribution that follows a power law, with relatively few highly connected vertices (high degree) and many vertices having a low degree. A given highly connected vertex, v, in a dense region of a graph may be connected to many vertices of degree one (singly linked vertex). These low degree vertices do not interconnect within the neighborhood of v and thus would reduce the clustering coefficient, but not the core-clustering coefficient. The final weight given to a vertex is the product of the vertex core-clustering coefficient and the highest k-core level, k max, of the immediate neighborhood of the vertex. This weighting scheme further boosts the weight of densely connected vertices. This specific weighting function is based on local network density. Many other functions are possible and some may have better performance for this algorithm but these are not evaluated here.
The second stage, molecular complex prediction, takes as input the vertex weighted graph, seeds a complex with the highest weighted vertex and recursively moves outward from the seed vertex, including vertices in the complex whose weight is above a given threshold, which is a given percentage away from the weight of the seed vertex. This is the vertex weight percentage (VWP) parameter. If a vertex is included, its neighbours are recursively checked in the same manner to see if they are part of the complex. A vertex is not checked more than once, since complexes cannot overlap in this stage of the algorithm (see below for a possible overlap condition). This process stops once no more vertices can be added to the complex based on the given threshold and is repeated for the next highest unseen weighted vertex in the network. In this way, the densest regions of the network are identified. The vertex weight threshold parameter defines the density of the resulting complex. A threshold that is closer to the weight of the seed vertex identifies a smaller, denser network region around the seed vertex.
The third stage is post-processing. Complexes are filtered if they do not contain at least a 2-core (graph of minimum degree 2). The algorithm may be run with the 'fluff' option, which increases the size of the complex according to a given 'fluff' parameter between 0.0 and 1.0. For every vertex in the complex, v, its neighbors are added to the complex if they have not yet been seen and if the neighborhood density (including v) is higher than the given fluff parameter. Vertices that are added by the fluff parameter are not marked as seen, so there can be overlap among predicted complexes with the fluff parameter set. If the algorithm is run using the 'haircut' option, the resulting complexes are 2-cored, thereby removing the vertices that are singly connected to the core complex. If both options are specified, fluff is run first, then haircut.
Resulting complexes from the algorithm are scored and ranked. The complex score is defined as the product of the complex subgraph, C = (V,E), density and the number of vertices in the complex subgraph (DC × |V|). This ranks larger more dense complexes higher in the results. Other scoring schemes are possible, but are not evaluated here.
MCODE may also be run in a directed mode where a seed vertex is specified as a parameter. In this mode, MCODE only runs once to predict the single complex that the specified seed is a part of. Typically, when analyzing complexes in a given network, one would find all complexes present (undirected mode) and then switch to the directed mode for the complexes of interest. The directed mode allows one to experiment with MCODE parameters to fine tune the size of the resulting complex according to existing biological knowledge of the system. In directed mode, MCODE will first pre-process the input network to ignore all vertices with higher vertex weight than the seed vertex. If this were not done, MCODE would preferentially branch out to denser regions of the graph, if they exist, which could belong to separate, but denser complexes. Thus, a seed vertex for directed mode should always be the highest density vertex among the suspected complex. There is an option to turn this pre-processing step off, which will allow seeded complexes to branch out into denser regions of the graph, if desired.
The time complexity of the entire algorithm is polynomial O(nmh 3) where n is the number of vertices, m is the number of edges and h is the vertex size of the average vertex neighbourhood in the input graph, G. This comes from the vertex-weighting step. Finding a k-core in a graph proceeds by progressively removing vertices of degree < k until all remaining vertices are connected to each other by degree k or more, and is thus O(n 2). The highest k-core is found by trying to find k-cores from one up until all vertices have been found and cannot go beyond a number of steps equal to the highest degree in the graph. Thus, the highest k-core step is O(n 3). Since this k-core step operates only on the neighbourhood of a vertex, the n in this case is the number of vertices in the average neighbourhood of a vertex, h. The inner loop of the algorithm only operates twice for every edge in the input graph, thus is O(2mh 3). The outer loop operates once on all vertices in the input graph, thus the entire time complexity of the weighting stage is O(n 2mh 3) = O(nmh 3). The complex prediction stage is O(n) and the optional post-processing step can be up to O(cs 2), where c is the number of complexes that were found in the previous step and s is the number of vertices in the largest complex - O(cs 2) to find the 2-core once for each complex.
Even though the fastest min-cut graph clustering algorithms are faster, at O(n 2logn) [30], MCODE has a number of advantages. Since weighting is done once and comprises most of the time complexity, many algorithm parameters can be tried, in O(n), once weighting is complete. This is useful when evaluating many different parameters. MCODE is relatively easy to implement and since it is local density based, has the advantage of a directed mode and a complex connectivity mode. These two modes are generally not useful in typical clustering applications, but are useful for examining molecular interaction networks. Additionally, only those proteins above a given local density threshold are assigned to complexes. This is in contrast to many clustering applications that force all data points to be part of clusters, whether they truly should be part of a cluster or not.
Pseudocode
Stage 1: Vertex Weighting
procedure MCODE-VERTEX-WEIGHTING
input: graph: G = (V,E)
for all v in G do
N = find neighbors of v to depth 1
K = Get highest k-core graph from N
k = Get highest k-core number from N
d = Get density of K
Set weight of v = k × d
end for
end procedure
Stage 2: Molecular Complex Prediction
procedure MCODE-FIND-COMPLEX
input: graph: G = (V,E); vertex weights: W;
vertex weight percentage: d; seed vertex: s
if s already seen then return
for all v neighbors of s do
if weight of v > (weight of s)(1 - d) then add v to complex C
call: MCODE-FIND-COMPLEX (G, W, d, v)
end for
end procedure
procedure MCODE-FIND-COMPLEXES
input: graph: G = (V,E); vertex weights: W;
vertex weight percentage: d
for all v in G do
if not already seen v then call: MCODE-FIND-COMPLEX(G, W, d, v)
end for
end procedure
Stage 3: Post-Processing (optional)
procedure MCODE-FLUFF-COMPLEX
input: graph: G = (V,E); vertex weights: W;
fluff density threshold: d; complex graph: C = (U,F)
for all u in C do
if weight of u >d then add u to complex C
end for
end procedure
procedure MCODE-POST-PROCESS
input: graph: G = (V,E); vertex weights: W; haircut flag: h; fluff flag: f;
fluff density threshold: t; set of predicted complex graphs: C
for all c in C do
if c not 2-core then filter
if h is TRUE then 2-core complex
if f is TRUE then call: MCODE-FLUFF-COMPLEX(G, W, t, c)
end for
end procedure
Overall Process
procedure MCODE
input: graph: G = (V,E); vertex weight percentage: d;
haircut flag: h; fluff flag: f; fluff density threshold: t;
set of predicted complex graphs: C
call: W = MCODE-VERTEX-WEIGHTING (G)
call: C = MCODE-FIND-COMPLEXES (G, W, d)
call: MCODE-POST-PROCESS (G, W, h, f, t, C)
end procedure
Implementation
MCODE has been implemented in ANSI C using the cross-platform NCBI Toolkit; http://www.ncbi.nlm.nih.gov/IEB and the BIND graph library in the SLRI Toolkit; http://sourceforge.net/projects/slritools. Both of these source code libraries are freely available. The actual MCODE source code is not yet freely available. The MCODE program has been compiled and tested on UNIX, Mac OS X and Windows. Because a yeast gene name dictionary is used to recognize input and generate output, the MCODE executable currently only works for yeast proteins in a user friendly manner. The algorithm, however is completely general, via the graph theory abstraction, to any graph and thus to any biomolecular interaction network. MCODE binaries are available from ftp://ftp.mshri.on.ca/pub/BIND/Tools/MCODE.
Results
Evaluation of MCODE
The evaluation of MCODE requires a set of experimentally determined biomolecular interactions and a set of associated experimentally determined molecular complexes. Currently, the largest source for such data is for proteins from the budding yeast, Saccharomyces cerevisiae. Recently, a large-scale mass spectrometry study by Gavin et al [7] provided a large data set of protein interactions with manually annotated molecular complexes. Also available are the protein interaction and complex tables of MIPS [13] and YPD [14]. MCODE was used to automatically predict protein complexes in our collected protein-protein interaction data sets. Resulting complexes were then matched to known molecular complexes from Gavin et al. (the Gavin benchmark) and the MIPS benchmark using an overlap score. Parameter optimization was then used to maximize the biological relevance of predicted complexes according to the given benchmarks. YPD was not used as a current version could not be acquired.
To ensure that MCODE is not unduly affected by the expected high false-positive rate in large-scale interaction data sets, large-scale and literature derived MCODE predictions were compared. MCODE was then used to predict complexes in the entire set of machine readable protein-protein interactions that we could collect for yeast. Complexes of interest were then further examined using the directed mode and complex connectivity mode of MCODE.
Evaluation of MCODE using the Gavin data set of protein interactions and complexes
In this study, we wanted to use all forms of protein interaction data available, which requires mixing of different types of experiments, such as yeast two-hybrid and co-immunoprecipitation. Two-hybrid results are inherently pairwise, whereas copurification results are sets of one or more identified proteins. For a copurification result, only a set of size 2 can be directly considered a pairwise interaction, otherwise it must be modeled as a set of hypothetical interactions. Biochemical copurifications can be thought of as populations of complexes with some underlying pairwise protein interaction topology that is unknown from the experiment. In the general case of the purification used by Gavin et al., one affinity tagged protein was used as bait to pull associated proteins out of a yeast cell lysate. The two extreme cases for the topology underlying the population of complexes from a single purification experiment are a minimally connected 'spoke' model, where the data are modeled as direct bait-associated protein pairwise interactions, and a maximally connected 'matrix' model, where the data are modeled as all proteins connected to all others in the set. The real topology of the set of proteins must lie somewhere between these two extremes.
Population of complexes: C = {b, c, d, e} (b = bait)
Spoke model hypothetical interactions: i S = {b-c, b-d, b-e}
Matrix model hypothetical interactions; i M = {b-b, b-c, b-d, b-e, c-c, c-d, c-e, d-d, d-e, e-e}
Advantages of the spoke model are that it is biologically intuitive, biologists often represent their copurification results in this manner, and is about 3 times more accurate than the matrix model [31]. Disadvantages are that it could misrepresent interactions. The matrix model, alternatively, cannot misrepresent interactions, as all possible interactions are generated, but this is at the cost of generating a large number of false interactions. Matrix topologies are also physically implausible for larger complexes because of increased possibility of steric clash if all subunits are interacting with all others. Ultimately, the spoke model should be reasonable for use in evaluating MCODE.
Gavin et al. raw data from 588 biochemical purifications were represented using the spoke model, described above, to get 3,225 hypothetical protein-protein interactions among 1,363 proteins for input to MCODE. A list of 232 manually annotated protein complexes based on the original purification data reported by Gavin et al. was filtered to remove five reported 'complexes' each composed of a single protein and six complexes of two or three proteins that were already in the data set as part of a larger complex. This yielded a filtered set of 221 complexes that were used to evaluate MCODE, although some of these complexes have significant overlap to other complexes in the set.
To evaluate which parameter choice would allow automatic prediction of protein complexes from the spoke modeled Gavin et al. interaction set that best matched the manually annotated complexes, MCODE was run using all four possible combinations of the two Boolean parameters (haircut: true/false, fluff: true/false) over a full range of 20 vertex weight percentage (VWP) and fluff parameters (0 to 0.95 in 0.05 increments). During this parameter optimization process, MCODE was limited to find complexes of size two or higher.
A scoring scheme was developed to determine how effectively an MCODE predicted complex matched a complex from the benchmark set of complexes. In this case, the benchmark complex set was the Gavin et al. hand-annotated complex set. The overlap score was defined as ω = i 2/a*b, where i is the size of the intersection set of a predicted complex with a known complex, a is the size of the predicted complex and b is the size of the known complex. A protein is part of the intersection set only if it is present in both predicted and known complexes. Thus, a predicted complex that has no proteins in a known complex has ω = 0 and a predicted complex that perfectly matches a known complex has ω = 1. Also, predicted complexes that fully overlap, but are much larger or much smaller than any known complexes will get a low ω. The overlap score of a predicted complex vs. a benchmark complex is then a measure of biological significance of the prediction, assuming that the benchmark set of complexes is biologically relevant. The best parameter choice for MCODE on this protein interaction data set is one that predicts the largest set of complexes that match the largest number of benchmark complexes above a threshold ω. Since there is overlap in the Gavin benchmark complex database, a predicted complex may match more than one known complex with a high ω.
To choose an overlap score that maximizes biological relevance of the predicted complexes without filtering away too many predictions, each of the 840 parameter combinations tested during the parameter optimization stage. The number of MCODE predicted complexes was plotted against the number of matched known complexes over a range of ω thresholds from 'no threshold' to 0.1 to 0.9 (in 0.1 increments). If no ω threshold is used, a predicted complex only needs at least one protein in common with a known complex to be considered a match. If predicted and known complexes are only counted as a match when their ω is above a specific threshold, the number of matched complexes declines with increasing ω threshold, as shown in Figure 1. Interestingly, the average and maximum number of matched known complexes drops more quickly from zero until a ω threshold of 0.2 than from 0.2 to 0.9 indicating that many predicted complexes only have one or a few proteins that overlap with known complexes. A ω threshold of 0.2 to 0.3 thus seems to filter out most predicted complexes that have insignificant overlap with known complexes.
Effect of Overlap Score Threshold on Number of Predicted and Matched Known Complexes for the Gavin Evaluation Figure legend: Average and maximum number of predicted and matched known complexes seen during MCODE parameter optimization (840 parameter combinations) plotted as a function of overlap score threshold. As the stringency for the closeness that a predicted complex must match a known complex is increased (increase in overlap score), fewer predicted complexes match known complexes. Note that these curves do not correspond to the best parameter set, but rather are an average of results from all tried parameter combinations.
Figure 2 shows the range of number of complexes predicted and number of known complexes matched for the 0.2 ω threshold over all tried MCODE parameters. A y = x line is also plotted to show that data points tend to be skewed towards a higher number of matched known complexes than predicted complexes because of the redundancy in the Gavin complex benchmark. Data points closest to the upper right portion of the graph maximize both number of matched known complexes and number of predicted complexes. MCODE parameter combinations that result in these data points therefore optimize MCODE on this data set (according to the overlap score threshold). This result shows that the number of predicted complexes should be similar to the number of matched known complexes for a parameter choice to be reasonable, although the number of matched known complexes may be larger, again, because of some commonality among complexes in the benchmark set. The parameter combination corresponding to the best data point (63,88) at an overlap score threshold of 0.2 is haircut = FALSE, fluff = TRUE, VWP = 0.05 and a fluff density threshold between 0 and 0.1. These parameter optimization results for MCODE over this data set were stable over a range of ω thresholds up to 0.5. Above 0.5, the result was not stable as there were generally too few predicted complexes with high overlap scores (Figure 1).
A specificity versus sensitivity analysis [32] was also performed. Defining the number of true positives (TP) as the number of MCODE predicted complexes with ω over a threshold value and the number of false positives (FP) as the total number of predicted MCODE complexes minus TP. The number of false negatives (FN) equals the number of known benchmark complexes not matched by predicted complexes. Sensitivity was defined as [TP/(TP+FN)] and specificity was defined as [TP/(TP+FP)]. The MCODE parameter choice that optimizes both specificity and sensitivity is the same as from the above analysis. The optimal sensitivity of this analysis was ~0.31 and the corresponding specificity was ~0.79.
The 63 MCODE predicted complexes only matched 88 of the 221 complexes in the known data set indicating that MCODE could not recapitulate the majority of the Gavin complex benchmark solely using protein connectivity information. As mentioned above, there are more matched complexes than predicted because of some redundancy in the benchmark. This low sensitivity is not surprising, since many of the hand-annotated complexes were created directly from single co-immunoprecipitation results, which are not highly interconnected in the spoke model. For example, Cdc3 was used as a bait to co-immunoprecipitate Cdc10, Cdc11, Cdc12 and Ydl225w. A complex was annotated as containing these five proteins, but only Cdc3 was used as bait. If more elements of a complex are used as baits, the proteins become more interconnected and more readily predicted by MCODE. A good example of this is the Arp2/3 complex, which is highly conserved in eukaryotes and is involved in actin cytoskeleton rearrangement. The structure of this complex is known by X-ray crystallography [33] thus actual protein-protein interactions from the structure can be matched up to the co-immunoprecipitation results. MCODE predicted all seven components of the Arp2/3 complex crystal structure and five extra proteins using the optimized parameters. Six out of the seven Arp2/3 subunits were used as baits by Gavin et al. and the resulting benchmark complex included the five extra proteins that MCODE also predicted (Nog2, Pfk1, Prt1, Cct8 and Cct5) that are not in the crystal structure. Cct5 and Cct8 are known to be involved in actin assembly, but Nog2, Pfk1 and Prt1 are not. These extra proteins likely represent non-specific binding in the experimental approach. These two cases are shown diagrammatically in Figure 3. Interestingly, using the haircut parameter would remove all five extra proteins that are not in the crystal structure, leaving only the seven that are present. This shows that while the parameter optimization allows maximum matching of the hand-annotated known complexes, these complexes may not all be physiologically relevant and thus another parameter set may better predict 'real' complexes.
Examples of Gavin Benchmark Complexes Missed and Hit by MCODE Figure legend: Protein complexes are represented as graphs using the spoke model. Vertices represent proteins and edges represent experimentally determined interactions. Blue vertices are baits in the Gavin et al. study. A) A Cdc3 complex hand-annotated by Gavin et al. that was missed by MCODE because of a lack of connectivity information among sub-components. This complex annotation was the result of a single co-immunoprecipitation experiment. B) The Arp2/3 complex as annotated by Gavin et al. and as found by MCODE with parameters optimized to the data set. Note the five extra proteins that have minimal connectivity to main cluster. C) The protein connection map seen from the crystal structure of the Arp2/3 complex. The crystal structure is from Bos taurus (cow), but is assumed to be very similar to yeast based on very high similarity between cow and yeast Arp2/3 subunits.
To explore the effect of certain MCODE parameters on resulting predicted complexes, various features of these complexes were examined while changing specific parameters and keeping all else constant. Linearly increasing the VWP parameter increased the size of the predicted complexes exponentially while reducing the number of complexes predicted in a linear fashion. Figure 4 shows this effect with both fluff and haircut parameters turned off. At high VWP values, very large complexes were predicted and these encompassed most of the data set, thus were not very useful.
Effect of Vertex Weight Percentage Parameter on Predicted Complex Size Figure legend: As the vertex weight percentage (VWP) parameter of MCODE is increased, the number of predicted complexes steadily decreases and the average and largest size of predicted complexes increases exponentially. The y-axis follows a logarithmic scale. For reference, the average and maximum size of the MIPS benchmark complexes are 6 and 81, respectively and of the Gavin benchmark complexes are 11.8 and 88, respectively.
Because using haircut = TRUE would have led MCODE to predict the Arp2/3 complex perfectly (according to the crystal structure as discussed above), we examined if the haircut parameter has any general effect on the number of matched predicted complexes. Setting haircut = TRUE had no significant effect on the number of complexes predicted at high ω thresholds, but generally reduced the number of matched known complexes at low ω thresholds (0 to 0.1) compared to haircut = FALSE. Since the haircut = TRUE option removes less-connected proteins on the fringe of a predicted complex and this reduces the number of predicted complexes with low overlap scores, these fringe proteins likely contribute to low-level overlap (<0.2 ω) of the known complexes.
We also investigated the effect of changing the fluff density threshold when setting fluff = TRUE on the number of matched benchmark complexes. Linearly increasing the fluff density threshold in the MCODE post-processing step linearly decreased the number of matched complexes above an overlap score of 0.2.
Evaluation of MCODE using MIPS data set of protein interactions and complexes
Since the Gavin et al. data set was developed by only one group using a single experimental method, it may not accurately represent protein complex knowledge for yeast. The MIPS protein complex catalogue http://mips.gsf.de/proj/yeast/catalogues/complexes/ is a curated set of 260 protein complexes for yeast that was compiled from the literature and is thus a more realistic data set comprised of varied experiments from many labs using different techniques. After filtering away 50 'complexes' each composed of a single protein and 2 highly similar complexes, we were left with 208 complexes for the MIPS known set. This set did not include information from the recent large-scale mass spectrometry studies [6, 7]. While the MIPS complex catalogue may be incomplete, it is currently the best available public resource for yeast protein complexes that we are aware of.
MCODE was run again with a full combination of parameters, this time over a set of 9088 protein-protein interactions among 4379 proteins which did not include the recent large-scale mass spectrometry studies but included all interactions from the MIPS, YPD and PreBIND databases as well as from the majority of large-scale yeast two-hybrid experiments to date [2–4, 10, 34]. This interaction set is termed 'Pre HTMS'. All of the interactions in this set were published before the last update specified on the MIPS protein complex catalogue and many are included in the MIPS protein interaction table, thus we assumed that the MIPS complex catalogue took into account the information in the known interaction table. Protein complexes found by MCODE in this set were compared to the MIPS protein complex catalogue to evaluate how well MCODE performed at locating protein complexes ab initio.
The same evaluation of MCODE that was done using the Gavin et al. data set was performed with the MIPS data set. From this analysis, including specificity versus sensitivity plots (optimized sensitivity = ~0.27 and specificity = ~0.31), the MIPS complex benchmark optimized parameters were haircut = TRUE, fluff = TRUE, VWP = 0.1 and a fluff density threshold of 0.2. This result was stable up to a ω threshold of 0.6 after which it was difficult to evaluate the results, as there were generally too few predicted complexes above the high ω thresholds. This parameter combination led MCODE to predict 166 complexes of which 52 matched 64 MIPS complexes with a ω of at least 0.2. Examining the ω distribution for this parameter set reveals that, even though this prediction is optimized, most of the predicted complexes don't show overlap to those in the known MIPS set (Figure 5). The complexes predicted here are also different from those predicted from the Gavin interaction data. Nine complexes have an overlap score above 0.2 between these two sets, with the highest overlap score being 0.43 and all the rest being below 0.27. This might signify that either the MIPS complex catalogue is not complete, that there is not enough data in the dataset that MCODE was run on, or a human annotated definition of a complex does not perfectly match with a graph density based definition.
Overlap Score Distributions of Pre HTMS and AllYeast interaction sets with MIPS Complex Benchmark Optimized MCODE Parameter Sets Figure legend: The number of MCODE predicted complexes in the pre-large scale mass spectrometry (Pre HTMS) and AllYeast protein-protein interaction sets with a given overlap score threshold compared to the MIPS benchmark complex set is shown. The majority of predicted complexes have an overlap score of zero meaning that they had no overlap with the catalogue of known MIPS protein complexes.
The effect of the VWP parameter on complex size and of the haircut and fluff parameters on number of matched complexes was very similar to that seen when evaluating MCODE on the Gavin complex benchmark.
Effect of data set properties on MCODE
Since many large-scale protein interaction data sets from yeast are known to contain a high level of false positives [35], we examined the effect these might have on MCODE predictions. Sensitivity vs. specificity was plotted for MCODE predictions, with parameters chosen to maximize these values at ω threshold of 0.2 against the MIPS and Gavin complex benchmarks for the various data sets (Figure 6).
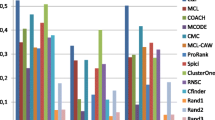
Sensitivity vs. Specificity Plots of MCODE Results Among Various Data Sets Figure legend: Specificity is plotted versus sensitivity of the best MCODE results at an overlap score above 0.2 against both the MIPS (Panel A) and Gavin (Panel B) complex benchmarks. Panel A shows that there are no large inherent differences among interaction data sets resulting from significantly different experimental methods (data set: sensitivity, specificity; Y2H:0.10,0.27; Benchmark:0.29,0.36; HTP Only:0.14;0.24; Pre HTMS:0.27,0.31; AllYeast:0.27,0.26; Gavin Spoke:0.10,0.38). Panel B shows that the Gavin benchmark is expectedly biased towards the Gavin interaction data set and thus should not be used as a general benchmark (data set: sensitivity, specificity; Y2H:0.03,0.10; Benchmark:0.11,0.16; HTP Only:0.24;0.33; Pre HTMS:0.10,0.13; AllYeast:0.27,0.26; Gavin Spoke:0.31,0.79).
MCODE predictions on the high-throughput data sets, termed 'Gavin Spoke', 'Y2H' and 'HTP only' (see Methods), are about as specific as the literature derived interaction data set, but not as sensitive (Figure 6A). MCODE predictions on interaction data sets containing the literature derived benchmark, labelled 'Benchmark', 'Pre HTMS' and 'AllYeast', are generally more sensitive and specific than those containing just the large-scale interaction sets. Since the specificity drops from Benchmark to Pre HTMS to AllYeast, with increasing amounts of large-scale data, it could be argued that addition of this data negatively affects MCODE. However, large-scale data is known to contain a high number of false positives, so it should be expected that these false-positives would not randomly contribute to the formation of dense regions, which are highly unlikely to occur by chance (see below). More complexes should be predicted with the addition of the large-scale data, assuming this data explores previously unseen regions of the interactome, but the high number of false-positives should limit the amount of new complexes compared to the amount of added interactions. The MIPS complex benchmark used here is not expected to contain complexes newly found in large-scale studies, explaining the decrease in specificity. This is exactly what occurs in our analysis. In an effort to further test the effect of large-scale data on MCODE prediction performance, the Benchmark interaction data set was augmented with the addition of interactions from large-scale experiments that only connect proteins in the Benchmark set with each other. Over 3100 interactions were added to the Benchmark data set to create a set of over 6400 interactions. MIPS complex benchmark optimised MCODE predicted 52 complexes matching 66 MIPS benchmark complexes, almost exactly the same number of complexes found using the Benchmark set by itself (Table 1). These analyses strongly suggest the addition of large-scale experimentally derived interactions does not unduly affect the prediction of complexes by MCODE.
It can be seen from Figure 6B that the Gavin complex benchmark set is biased towards the Gavin et al. spoke modeled interaction data. This is expected and is the main reason why the less biased MIPS complex set is used throughout this work as a benchmark instead of the Gavin set.
Since the result of a co-immunoprecipitation experiment is a set of proteins, which we model as binary interactions using the spoke method, we wished to evaluate whether this affects complex prediction compared to an experimental system that generates purely binary interaction results, such as yeast two-hybrid. As can be seen in Table 1, MCODE does find known complexes in the 'Y2H' set of only yeast two-hybrid results, thus this set does contain dense regions that are known protein complexes. This being said, the Y2H set is the least dense of all data sets examined here so is expected to have less dense regions of the network and thus less MCODE predictable complexes per protein present in the set. MCODE predicts a similar amount of complexes as well as finding a similar amount of known complexes in the Y2H and Gavin Spoke data sets indicating that these data sets are not significantly different from each other in the amount of dense network regions that they contain, even though they are different sizes. Taken together, the latter results and those in Figure 6B show that the spoke model is a reasonable representation of the Gavin et al. tandem affinity purification data.
Predicting complexes in the Yeast interactome
Given that MCODE performed reasonably well on test data, we decided to predict complexes in a much larger network. All machine-readable protein-protein interaction data from various data sets [2–7, 10, 13, 14]. were collected and integrated to form a non-redundant set of 15,143 experimentally determined yeast protein interactions encompassing 4,825 proteins, or approximately three quarters of the proteome. This set was termed 'AllYeast'. MCODE was parameter optimized, as above, using the MIPS benchmark. The best resulting parameter set was haircut = TRUE, fluff = TRUE, VWP = 0 and a fluff density threshold of 0.1. With these parameters, MCODE predicted 209 complexes, of which 54 matched 63 MIPS benchmark complexes above an overlap score of 0.2 (see Additional file 1). Complexes found in this manner should be further studied using MCODE in directed mode by specifying a seed vertex and trying different parameters to examine how large a complex can get before seemingly biologically irrelevant proteins are added (see below).
Figure 5 shows that even when a large set of interactions is used as input to MCODE, most of the MCODE predicted complexes do not match well with known complexes in MIPS. The complex size distribution of MCODE predicted complexes matches the shape of the MIPS set, but the MCODE complexes are on average larger (Average MIPS size = 6.0, Average MCODE Predicted size = 9.7). The average number of YPD and GO functional annotation terms per protein in an MCODE predicted complex is similar to that of MIPS complexes (Table 2). This seems to indicate that MCODE is predicting complexes that are functionally relevant. Also, closer examination of the top, middle and bottom five scoring MCODE complexes shows that MCODE can predict biologically relevant complexes (Table 3).
Many of the 209 predicted complexes are of size 2 (9 predicted complexes) or 3 (54 predicted complexes). Complexes of this size may not be significant since it is easy to create high density subgraphs of size 2 or 3, but becomes combinatorially more difficult to randomly create high density subgraphs as the size of the subgraph increases. To examine the relevance of these small predicted complexes of size 2 or 3, we calculated the sensitivity and specificity of the optimized MCODE predictions against the MIPS complex benchmark while disregarding the small complexes. First, complexes of size 2, then of size 3, were removed from the optimized MCODE predicted complex set. Removing each of these sets independently resulted in only small sensitivity and specificity changes. Because both sets overlap the MIPS benchmark, small complexes have been reported as predictions. Also, because MCODE found these small complexes in regions of high local density, they may be good cores for further examination with MCODE in directed mode, especially since the haircut option was turned on here to produce them.
Complexes that are larger and denser are ranked higher by MCODE and these generally correspond to known complexes (see below). Interestingly, some MCODE complexes contain unknown proteins that are highly connected to known complex subunits. For example, the second highest ranked MCODE complex is involved in RNA processing/modification and contains the known polyadenylation factor I complex (Cft1, Cft2, Fip1, Pap1, Pfs2, Pta1, Ysh1, Yth1 and Ykl059c). Seven other proteins involved in mainly RNA processing/modification (Fir1, Hca4, Pcf11, Pti1, Ref2, Rna14, Ssu72) and protein degradation (Uba2 and Ufd1) are highly connected within this predicted complex. Two unknown proteins Pti1 and Yor179c are highly connected to RNA processing/modification proteins and are therefore likely involved in the same process (Figure 7). Pti1 may be an unknown component of the polyadenylation factor I complex. The 23rd highest ranked predicted complex is interesting in that it is involved in cell polarity and cytokinesis and contains two proteins of unknown function, Yhr033w and Yal027w. Yal027w interacts with two kinases, Gin4 and Kcc4, which in turn interact with the components of the Septin complex (Cdc3, Cdc10, Cdc11 and Cdc12) (Figure 8).
The Second Highest Ranked MCODE Predicted Complex is Involved in RNA Processing and Modification . Figure legend: This complex incorporates the known polyadenylation factor I complex (Cft1, Cft2, Fip1, Pap1, Pfs2, Pta1, Ysh1, Yth1 and Ykl059c) and contains other proteins highly connected to this complex, some of unknown function. The fact that the unknown proteins (Yor179c and Pti1) connect more to known RNA processing/modification proteins than to other proteins in the larger data set likely indicates that these proteins function in RNA processing/modification. This complex was ranked second by MCODE from the predicted complexes in the AllYeast interaction set.
An MCODE Predicted Complex Involved in Cytokinesis Figure legend: This predicted complex incorporates the known Septin complex (Cdc3, Cdc10, Cdc11 and Cdc12) involved in cytokinesis and other cytokinesis related proteins. The Yal027w protein is of unknown function, but likely functions in cell cycle control according to this figure, possibly in cytokinesis. This complex was ranked 23rd by MCODE from the predicted complexes in the AllYeast interaction set.
Significance of MCODE predictions
Naïvely, the chance of randomly picking a known protein complex from a protein interaction network depends on the size of the complex and the network. It is easier to pick out a smaller known complex by chance from a smaller network. For instance, in our network of 15,143 interactions among 4,825 proteins, the chance of picking a specific known complex of size three is about one in 1.9 × 1010 (4,825 choose 3). A more realistic model would assume that the proteins are connected and thus would only consider complex choices of size three where all three proteins are connected. The number of choices now depends on the topology of the network. In our large network, there are 6,799 fully connected subnetworks of size three and 313,057 subnetworks of size three with only two interactions (from the triadic census feature of Pajek). Thus now our chance of picking a more realistic complex is one out of 319,856 (1/(6,799 + 313,057) = 3.1 × 10-6). As the size of the complex increases, the number of possible complex topologies increases exponentially and, in a connected network of some reasonable density, so does the number of possible subgraphs that could represent a complex. The density of our large protein interaction network is 0.0013 and is mostly connected (4,689 proteins are in one connected component). Thus, it is expected that if a complex is found in a network with MCODE that matches a known complex, that the result would be highly significant. To understand the significance of complex prediction further, the topology of the protein interaction network would have to be understood in general, in order to build a null model to compare against.
Recent research on modeling complex systems [21, 25, 27] has found that networks such as the world wide web, metabolic networks [26] and protein-protein interaction networks [36] are scale-free. That is, the connectivity distribution of the vertices of the graph follows a power law, with many vertices of low degree and few vertices of high degree. Scale-free networks are known to have large clustering coefficients, or clustered regions of the graph. In biological networks, at least in yeast, these clustered regions seem to correspond to molecular complexes and these subgraphs are what MCODE is designed to find.
To test the significance of clustered regions in biological networks, 100 random permutations of the large set of all 15,143 yeast interactions were made. If the graph to be randomised is considered as a set of edges between two vertices (v 1, v 2), a network permutation is made by randomly permuting the set of all v 2 vertices. The random networks have the same number of edges and vertices as the original network and follow a power-law connectivity distribution, as do the original data sets [37]. Running MCODE with the same parameters as the original network (haircut = TRUE, fluff = TRUE, VWP = 0 and a fluff density threshold of 0.1) on the 100 random networks resulted in an average of 27.4 (SD = 4.4) complexes per network. The size distribution of complexes found by MCODE did not match that of the complexes found in the original network, as some complexes found in the random networks were composed of >1500 proteins. One random network that had an approximately average number of predicted complexes (27) was parameter optimized using the MIPS benchmark to see how parameter choice affects the size distribution and number of predicted complexes. Parameters of haircut = TRUE, fluff = TRUE, VWP = 0.1 and a fluff density threshold of zero produced the maximal number of 81 complexes for this network, but these complexes were composed of on average 27 proteins (without counting an outlier complex of size 1961), which is much larger than normal (e.g. larger than the MIPS set average of 6.0). None of these predicted complexes matched any MIPS complexes above an overlap score of 0.1. Also, the random network complexes had a much higher average number of YPD and GO annotation terms per protein per complex than for MIPS or MCODE on the original network (Table 2). This indicates, as expected, that the random network complexes are composed of a higher level of unrelated proteins than complexes in the original network. Thus, the number, size and functional composition of complexes that MCODE predicts in the large set of all yeast interactions are highly unlikely to occur by chance.
To evaluate the effectiveness of our scoring scheme, which scores larger, more dense complexes higher than smaller, more sparse complexes, we examined the accuracy of MCODE predictions at various score thresholds. As the score threshold for inclusion of complexes is increased, less complexes are included, but a higher percentage of the included complexes match complexes in the benchmark. This is at the expense of sensitivity as many benchmark matching complexes are not included at higher score thresholds (Figure 9). For example, of the ten predicted complexes with MCODE score greater or equal to six, nine match a known complex in either the MIPS or Gavin benchmark above a 0.2 threshold overlap score, yielding an accuracy of 90%. 100% of the five complexes that had an MCODE score better or equal to seven matched known complexes. Thus, complexes that score highly on our simple density based scoring scheme are very likely to be real.
Effect of Complex Score Threshold on MCODE Prediction Accuracy Figure legend: MCODE complexes equal to or greater than a specific score were compared to a benchmark comprising the combined MIPS and Gavin benchmarks. Accuracy was calculated as the number of known complexes better or equal to the threshold score divided by the total number of predicted complexes (matching and non-matching) at that threshold. A complex was deemed to match a known complex if it had an overlap score above 0.2. The number of predicted complexes that matched known complexes at each score threshold is shown as labels on the plot.
Directed mode of MCODE
To simulate an obvious example where the directed mode of MCODE would be useful, MCODE was run with relaxed parameters (haircut = TRUE, fluff = TRUE, VWP = 0.05 and a fluff density threshold of 0.2) compared to the best parameters on the AllYeast network. The resulting fourth highest ranked complex, when visualized, shows two clustered components and represents two protein complexes, the proteasome and an RNA processing complex, both found in the nucleus (Figure 10). This is an example of where a lower VWP parameter would have been superior since it would have divided this large complex into two more functionally related complexes. The highest weighted vertices in the center of each of the two dense regions in Figure 10 are the Rpt1 and Lsm4 proteins. MCODE was run in directed mode starting with these two proteins over a range of VWP parameters from 0 to 0.2, at 0.05 increments. For Lsm4, the parameter set of haircut = TRUE, fluff = FALSE, VWP = 0 was used to find a core complex, which contained 9 proteins fully connected to each other (Dcp1, Kem1, Lsm2, Lsm3, Lsm4, Lsm5, Lsm6, Lsm7 and Pat1). Above this VWP parameter, the core complex branched out into proteasome subunit proteins, which are not part of the Lsm complex (see Figure 11A). Using this VWP parameter, combinations of haircut and fluff parameters were used to further expand the core complex. This process was stopped when the predicted complexes began to include proteins of sufficiently different known biological function to the seed vertex. Proteins, such as Vam6 and Yor320c were included in the complex at moderate fluff parameters (0.4–0.6), but not at higher fluff parameters, and these are known to be localized in membranes outside of the nucleus, thus are likely not functionally related to the Lsm complex proteins. Therefore, the 9 proteins listed above were decided to be the final complex (Figure 11B). This is intuitive because of their maximal density (a 9-clique).
An MCODE Predicted Complex That is Too Large (Relaxed Parameters) Figure legend: An example of a predicted complex that incorporates two complexes, proteasome (left) and an RNA processing complex (right). These should probably be predicted as separate complexes as can be seen by the clear distinction of biological role annotation on one side of this layout compared to the other (purple versus blue). This figure, however, shows the large amount of overall connectivity between these two complexes. This complex was ranked fourth by MCODE from the predicted complexes in the AllYeast interaction set with slightly relaxed parameters compared to the optimized prediction.
MCODE in Directed Mode Figure legend: MCODE was used in directed mode to further study the complex in Figure 10 by using seed vertices from high density regions of the two parts of this complex. A) The result of examining the Lsm complex using MCODE parameters that are too relaxed (haircut = TRUE, fluff = FALSE, VWP = 0.05). B) The final Lsm complex using MCODE parameters of haircut = TRUE, fluff = FALSE and VWP = 0 seeded with Lsm4. C) The final 26S proteasome complex seeded with Rpt1 using MCODE parameters haircut = TRUE, fluff = TRUE and VWP = 0.2. Visible here are two regions of density in this complex corresponding to the 20S proteolytic subunit (left side – mainly Pre proteins) and the 19S regulatory subunit (right side – mainly Rpt and Rpn proteins).
Using this same method of known biological role "titration" on Rpt1 found a complex of 34 proteins (Gal4, Gcn4, Hsm3, Lhs1, Nas6, Pre1, Pre2, Pre3, Pre4, Pre5, Pre6, Pre7, Pre9, Pup3, Rpn10, Rpn11, Rpn13, Rpn3, Rpn5, Rpn6, Rpn7, Rpn8, Rpn9, Rpt1, Rpt2, Rpt3, Rpt4, Rpt6, Rri1, Scl1, Sts1, Ubp6, Ydr179c, Ygl004c) and 160 interactions using the parameter set haircut = TRUE, fluff = TRUE, VWP = 0.2 and a fluff density threshold of 0.3. Two regions of density can be seen here corresponding to the two known subunits of the 26S proteasome. The 20S proteolytic subunit of the proteasome is comprised of 15 proteins (Pre1 to Pre10, Pup1, Pup2, Pup3, Scl1 and Ump1) of which Pre7, Pre8, Pre10, Pup1, Pup2 and Ump1 are not found with MCODE. The 19S regulatory subunit of the proteasome is known to have 21 subunits (Nas6, Rpn1 to Rpn13, Rpt1 to Rpt6 and Ubp6) of which Rpn1, Rpn2, Rpn4, Rpn12 and Rpt5 are not found with MCODE. Known complex components not found by MCODE are not present at a high enough local density regions of the interaction network, possibly because not enough experiments involving these proteins are present in our data set. Figure 11C shows the final Rpt1 seeded complex. Of note, Ygl004c is unknown and binds to almost every Rpt and Rpn protein in the complex although all of these interactions were from a single immunoprecipitation experiment [6]. As well, Rri1 and Ydr179c have unknown function and both bind to each other and to Rpn5. Thus one would predict that these three unknown proteins function with or as part of the 26S proteasome. The protein Hsm3 binds to eight other 19S subunits and is involved in DNA mismatch repair pathways, but is not known to be part of the proteasome, although all of these Hsm3 interactions are from a particular large-scale experiment [7]. Interestingly, Gal4, a transcription factor involved in galactose metabolism, is found to be part of the proteasome complex. While this metabolic functionality seems unrelated to protein degradation, it has recently been shown that the binding is physiologically relevant [38]. These cases illustrate the possible unreliability of both functional annotation and interaction data, but also that seemingly unrelated proteins should not be immediately discounted if found to be part of a complex by MCODE.
Of note, the known topology of the 26S proteasome [39] compares favourably with the complex visualization of Figure 11C without considering stoichiometry. Thus, if enough interactions are known, visualizing complexes may reveal the rough structural outline of large complexes. This should be expected when dealing with actual physical protein-protein interactions since there are few allowed topologies for large complexes considering the specific set of defining interactions and steric clashes between protein subunits.
Complex connectivity
MCODE may also be used to examine the connectivity and relationships between molecular complexes. Once a complex is known using the directed mode, the MCODE parameters can be relaxed to allow branching out into other complexes. The MCODE directed mode preprocessing step must also be turned off to allow MCODE to branch into other connected complexes, which may reside in denser regions of the graph than the seed vertex. As an example, this was done with the Lsm4 seeded complex (Figure 12). MCODE parameters were relaxed to haircut = TRUE, fluff = FALSE, VWP = 0.2 although they could be further relaxed for greater extension out into the network.
Examining Complex Connectivity with MCODE Figure legend: The complexes shown here are known to be nuclear localized and are involved in protein degradation (19S proteasome subunit), mRNA processing (Lsm complex and mRNA Cleavage/Polyadenylation complex), cell cycle (anaphase promoting complex) and transcription (SAGA transcriptional activation complex).
Discussion
This method represents an initial step in taking advantage of the protein function data being generated by many large-scale protein interaction studies. As the experimental methods are further developed, an increasing amount of data will be produced which will require computational methods for efficient interpretation. The algorithm described here allows the automated prediction of protein complexes from qualitative protein-protein interaction data and is thus able to help predict the function of unknown proteins and aid in the understanding of the functional connectivity of molecular complexes in the cell. The general nature of this method may allow complex prediction for molecules other than proteins as well, for example metabolic complexes that include small molecules.
MCODE cannot stand alone in this task; it must be combined with a graph visualization system to ease the understanding of the relationships among molecules in the data set. We use the Pajek program for large network analysis [40] with the Kamada-Kawai graph layout algorithm [41]. Kamada-Kawai models the edges in the graph as springs, randomly places the vertices in a high energy state and then attempts to minimize the energy of the system over a number of time steps. The result is that the Euclidean distance, here in a plane, is close to the graph-theoretic or path distance between the vertices. The vertices are visually clustered based on connectivity. Biologically, this visualization can allow one to see the rough structural outline of large complexes, if enough interactions are known, as evidenced in the proteasome complex analysis above (Figure 11C).
It is important to note and understand the limitations of the current experimental methods (e.g. yeast two-hybrid and co-immunoprecipitation) and the protein interaction networks that these techniques generate when analyzing the resulting data. One common class of false-positive interactions arising from many different kinds of experimental methods is that of indirect interactions. For instance, an interaction may be seen between two proteins using a specific experimental method, but in reality, those proteins do not physically bind each other, and one or more other molecules that are generally part of the same complex mediate the observed interaction. As can be seen for the Arp2/3 complex shown in Figure 3, when pairwise interactions between all combinations of proteins in a complex are studied, this creates a very dense graph. Interestingly, this false-positive effect is normally considered a disadvantage, but is an advantage with MCODE as it increases the density in the region of the graph containing a complex, which can then be more easily predicted.
Apart from the experimental factors that lead to false-positive and false-negative interactions, representational limitations also exist computationally. Temporal and spatial information is not currently described in interaction networks. A complex found by the MCODE approach may not actually exist even though all of the component proteins bind each other in vitro. Those proteins may never be present at the same time and place. For example, molecular complexes that perform different functions sometimes have common subunits as with the three types of eukaryotic RNA polymerases.
Complex stoichiometry, another important aspect of biological data, is not represented either. While it is possible to include full stoichiometry in a graph representation of a biomolecular interaction network, many experimental methods do not provide this information, so a homo-multimeric complex is normally represented as a simple homodimer. When an experiment does provide stoichiometry information, it is not stored in most current databases, such as MIPS and YPD. Thus, one is forced to return to the primary literature to extract the data, an extremely time-consuming task for large data sets.
Some quantitative and statistical information is present when integrating results of large-scale approaches and this is not used in our current graph model. For instance, the number of different types of experiments that find the same interaction, the quality of the experiment, the date the experiment was conducted (newer methods may be superior in certain aspects) and other factors that pertain to the reliability of the interaction could all be considered to determine a reliability index or p-value on edges in the graph. For instance, one may wish to rank results published in high-impact journals above other journals (or vice versa) and rank classical purification methods above high-throughput yeast two-hybrid techniques when determining the quality of the interaction data. It may also be possible to weight vertices on the graph by other quality criteria, such as whether a protein is hypothetical from a gene prediction or not or whether a protein is expressed at a particular time and place in the cell. For example, if one were interested in a certain stage of the cell cycle, proteins that are known to be absent at that stage could be reduced in weight (VWP in the case of MCODE) compared to proteins that are present. It should be noted that any weighting scheme that tries to assess the quality of an interaction might make false assumptions that would prevent the discovery of new and interesting data.
This paper shows that the structure of a biological network can define complexes, which can be seen as dense regions. This may be attributed to indirect interactions accumulating in the literature. Thus, interaction data taken out of context may be erroneous. For instance, if one has a collection of protein interactions from various different experiments done at different times in different labs from a specific complex that form a clique, and if one chooses an interaction from this clique, then how can one verify if it is indirect or not. We would only begin to know if we had a very detailed description of the experiment from the original papers where we could tell the amount of work and quality of work that went into measuring each interaction. Thus with only a qualitative view of interactions, in reference to Dobzhansky [42], nothing in the biomolecular interaction network would make sense except in light of molecular complexes and the functional connections between them. If one had a highly detailed representation of each interaction including time, place, experimental condition, number of experiments, binding sites, chemical actions and chemical state information, one would be able to computationally delve into molecular complexes to resolve topology, structure, function and mechanism down to the atomic level. This information would also help to judge the biological relevance of an interaction. Thus, we require databases like BIND [15] to store this information. The integration of known qualitative and quantitative molecular interaction data in a machine-readable format should allow increasingly accurate protein interaction, molecular complex and pathway prediction, including actual binding site and mechanism information in a sequence and structural context.
Based on our scale-free network analysis, it would seem that real biological networks are organized differently than random models of scale-free networks in that they have higher clustering coefficients around specific regions (complexes) and the vertices in these regions are related to each other, by biological function. Thus, attempts to model biological networks and their evolution in a global way solely using the statistics of scale-free networks may not work, rather modeling should take into account as much extant biological knowledge as possible.
Future work on MCODE could include researching different, possibly adaptive, vertex scoring functions to take into account, for example, the local density of the network past the immediate neighborhood of a vertex and the inclusion of functional annotation and p-values on edges. Time, space and stoichiometry should also be represented on networks and in visualization systems. The process of 'functional annotation titration' in the directed mode of MCODE could be automated.
Conclusions
MCODE effectively finds densely connected regions of a molecular interaction network, many of which correspond to known molecular complexes, based solely on connectivity data. Given that this approach to analyzing protein interaction networks performs well using minimal qualitative information implies that large amounts of available knowledge is buried in large protein interaction networks. More accurate data mining algorithms and systems models could be constructed to understand and predict interactions, complexes and pathways by taking into account more existing biological knowledge. Structured molecular interaction data resources such as BIND will be vital in creating these resources.
Methods
Data sources
All protein interaction data sets from MIPS [13], Gene Ontology [43] and PreBIND http://bioinfo.mshri.on.ca/prebind/ were collected as described previously [6]. The YPD protein interaction data are from March 2001 and were originally requested from Proteome, Inc. http://www.proteome.com. Other interaction data sets are from BIND http://www.bind.ca. A BIND yeast import utility was developed to integrate data from SGD [12], RefSeq [44], Gene Registry http://genome-www.stanford.edu/Saccharomyces/registry.html, the list of essential genes from the yeast deletion consortium [11] and GO terms [43]. This database ensures proper matching of yeast gene names among the multiple data sets that may use different names for the same genes. The yeast proteome used here is defined by SGD and RefSeq and contains 6,334 ORFs including the mitochondrial chromosome. Before performing comparisons, the various interaction data sets were entered into a local instance of BIND as pairwise protein interaction records. The MIPS complex catalogue was downloaded in February 2002.
The protein interaction data sets used here were composed as follows. 'Gavin Spoke' is the spoke model of the raw purifications from Gavin et al [7]. 'Y2H' is all known large-scale [2–5, 10] combined with normal yeast two-hybrid results from MIPS. 'HTP Only' is only high-throughput or large-scale data [2–7, 10] The 'Benchmark' set was constructed from MIPS, YPD and PreBIND as previously described [6]. 'Pre HTMS' was composed of all yeast sets except the recent large-scale mass spectrometry data sets [6, 7]. 'AllYeast' was the combination of all above data sets. All data sets are non-redundant.
Network visualization
Visualization of networks was performed using the Pajek program for large network analysis [40]http://vlado.fmf.uni-lj.si/pub/networks/pajek/ as described previously [6, 10]. using the Kamada-Kawai graph layout algorithm followed by manual vertex adjustments and was formatted using CorelDraw 10. Power law analysis was also accomplished as previously described [6].
References
Fields S: Proteomics. Proteomics in genomeland. Science 2001, 291: 1221–1224. 10.1126/science.291.5507.1221
Uetz P, Giot L, Cagney G, Mansfield TA, Judson RS, Knight JR, et al.: A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature 2000, 403: 623–627. 10.1038/35001009
Ito T, Chiba T, Ozawa R, Yoshida M, Hattori M, Sakaki Y: A comprehensive two-hybrid analysis to explore the yeast protein interactome. Proc Natl Acad Sci U S A 2001, 98: 4569–4574. 10.1073/pnas.061034498
Drees BL, Sundin B, Brazeau E, Caviston JP, Chen GC, Guo W, et al.: A protein interaction map for cell polarity development. J Cell Biol 2001, 154: 549–571. 10.1083/jcb.200104057
Fromont-Racine M, Mayes AE, Brunet-Simon A, Rain JC, Colley A, Dix I, et al.: Genome-wide protein interaction screens reveal functional networks involving Sm-like proteins. Yeast 2000, 17: 95–110. 10.1002/1097-0061(20000630)17:2<95::AID-YEA16>3.0.CO;2-H
Ho Y, Gruhler A, Heilbut A, Bader GD, Moore L, Adams SL, et al.: Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature 2002, 415: 180–183. 10.1038/415180a
Gavin AC, Bosche M, Krause R, Grandi P, Marzioch M, Bauer A, et al.: Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 2002, 415: 141–147. 10.1038/415141a
Christendat D, Yee A, Dharamsi A, Kluger Y, Savchenko A, Cort JR, et al.: Structural proteomics of an archaeon. Nat Struct Biol 2000, 7: 903–909. 10.1038/82823
Kim SK, Lund J, Kiraly M, Duke K, Jiang M, Stuart JM, et al.: A gene expression map for Caenorhabditis elegans. Science 2001, 293: 2087–2092. 10.1126/science.1061603
Tong AH, Drees B, Nardelli G, Bader GD, Brannetti B, Castagnoli L, et al.: A combined experimental and computational strategy to define protein interaction networks for peptide recognition modules. Science 2002, 295: 321–324. 10.1126/science.1064987
Winzeler EA, Shoemaker DD, Astromoff A, Liang H, Anderson K, Andre B, et al.: Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 1999, 285: 901–906. 10.1126/science.285.5429.901
Chervitz SA, Hester ET, Ball CA, Dolinski K, Dwight SS, Harris MA, et al.: Using the Saccharomyces Genome Database (SGD) for analysis of protein similarities and structure. Nucleic Acids Res 1999, 27: 74–78. 10.1093/nar/27.1.74
Mewes HW, Frishman D, Gruber C, Geier B, Haase D, Kaps A, et al.: MIPS: a database for genomes and protein sequences. Nucleic Acids Res 2000, 28: 37–40. 10.1093/nar/28.1.37
Costanzo MC, Crawford ME, Hirschman JE, Kranz JE, Olsen P, Robertson LS, et al.: YPD, PombePD and WormPD: model organism volumes of the BioKnowledge library, an integrated resource for protein information. Nucleic Acids Res 2001, 29: 75–79. 10.1093/nar/29.1.75
Bader GD, Donaldson I, Wolting C, Ouellette BF, Pawson T, Hogue CW: BIND-The biomolecular interaction network database. Nucleic Acids Res 2001, 29: 242–245. 10.1093/nar/29.1.242
Xenarios I, Salwinski L, Duan XJ, Higney P, Kim SM, Eisenberg D: DIP, the Database of Interacting Proteins: a research tool for studying cellular networks of protein interactions. Nucleic Acids Res 2002, 30: 303–305. 10.1093/nar/30.1.303
Takai-Igarashi T, Nadaoka Y, Kaminuma T: A database for cell signaling networks. J Comput Biol 1998, 5: 747–754.
Wingender E, Chen X, Hehl R, Karas H, Liebich I, Matys V, et al.: TRANSFAC: an integrated system for gene expression regulation. Nucleic Acids Res 2000, 28: 316–319. 10.1093/nar/28.1.316
Karp PD, Riley M, Saier M, Paulsen IT, Paley SM, Pellegrini-Toole A: The EcoCyc and MetaCyc databases. Nucleic Acids Res 2000, 28: 56–59. 10.1093/nar/28.1.56
Overbeek R, Larsen N, Pusch GD, D'Souza M, Selkov EJ, Kyrpides N, et al.: WIT: integrated system for high-throughput genome sequence analysis and metabolic reconstruction. Nucleic Acids Res 2000, 28: 123–125. 10.1093/nar/28.1.123
Wagner A, Fell DA: The small world inside large metabolic networks. Proc R Soc Lond B Biol Sci 2001, 268: 1803–1810. 10.1098/rspb.2001.1711
Flake GW, Lawrence S, Giles CL, Coetzee FM: Self-Organization of the Web and Identification of Communities. IEEE Computer 2002, 35: 66–71. 10.1109/2.989932
Goldberg AV: Finding a Maximum Density Subgraph. Technical Report UCB/CSD University of California, Berkeley, CA 1984., 84:
Ng A, Jordan M, Weiss Y: On spectral clustering: Analysis and an algorithm. Advances in Neural Information Processing Systems 14: Proceedings of the 2001 2001.
Watts DJ, Strogatz SH: Collective dynamics of 'small-world' networks. Nature 1998, 393: 440–442. 10.1038/30918
Jeong H, Tombor B, Albert R, Oltvai ZN, Barabasi AL: The large-scale organization of metabolic networks. Nature 2000, 407: 651–654. 10.1038/35036627
Albert R, Jeong H, Barabasi AL: Error and attack tolerance of complex networks. Nature 2000, 406: 378–382. 10.1038/35019019
Barabasi AL, Albert R: Emergence of scaling in random networks. Science 1999, 286: 509–512. 10.1126/science.286.5439.509
Fell DA, Wagner A: The small world of metabolism. Nat Biotechnol 2000, 18: 1121–1122. 10.1038/81025
Hartuv E, Shamir R: A clustering algorithm based on graph connectivity. Information processing letters 1999, 76: 175–181. 10.1016/S0020-0190(00)00142-3
Bader GD, Hogue CW: Analyzing yeast protein-protein interaction data obtained from different sources. Nat Biotechnol 2002, 20: 991–997. 10.1038/nbt1002-991
Baldi P, Brunak S, Chauvin Y, Andersen CA, Nielsen H: Assessing the accuracy of prediction algorithms for classification: an overview. Bioinformatics 2000, 16: 412–424. 10.1093/bioinformatics/16.5.412
Robinson RC, Turbedsky K, Kaiser DA, Marchand JB, Higgs HN, Choe S, et al.: Crystal structure of Arp2/3 complex. Science 2001, 294: 1679–1684. 10.1126/science.1066333
Mayes AE, Verdone L, Legrain P, Beggs JD: Characterization of Sm-like proteins in yeast and their association with U6 snRNA. EMBO J 1999, 18: 4321–4331. 10.1093/emboj/18.15.4321
von Mering C, Krause R, Snel B, Cornell M, Oliver SG, Fields S, et al.: Comparative assessment of large-scale data sets of protein-protein interactions. Nature 2002, 417: 399–403. 10.1038/nature750
Jeong H, Mason SP, Barabasi AL, Oltvai ZN: Lethality and centrality in protein networks. Nature 2001, 411: 41–42. 10.1038/35075138
Maslov S, Sneppen K: Specificity and stability in topology of protein networks. Science 2002, 296: 910–913. 10.1126/science.1065103
Gonzalez F, Delahodde A, Kodadek T, Johnston SA: Recruitment of a 19S proteasome subcomplex to an activated promoter. Science 2002, 296: 548–550. 10.1126/science.1069490
Bochtler M, Ditzel L, Groll M, Hartmann C, Huber R: The proteasome. Annu Rev Biophys Biomol Struct 1999, 28: 295–317. 10.1146/annurev.biophys.28.1.295
Batagelj V, Mrvar A: Pajek – Program for Large Network Analysis. Connections 1998, 2: 47–57.
Kamada T, Kawai S: An algorithm for drawing general indirect graphs. Information processing letters 1989, 31: 7–15. 10.1016/0020-0190(89)90102-6
Dobzhansky T: Nothing in Biology Makes Sense Except in the Light of Evolution. American Biology Teacher 1973, 35: 125–129.
The Gene Ontology Consortium: Gene ontology: tool for the unification of biology. Nat Genet 2000, 25: 25–29. 10.1038/75556
Pruitt KD, Maglott DR: RefSeq and LocusLink: NCBI gene-centered resources. Nucleic Acids Res 2001, 29: 137–140. 10.1093/nar/29.1.137
Acknowledgements
Michel Dumontier, Sherrie Kelly, Katerina Michalickova, Tony Pawson and Mike Tyers provided helpful discussion. This work was supported in part from grants from the Canadian Institutes of Health Research (CIHR), the Ontario Research and Development Challenge Fund and MDS-Sciex to C.H. G.D.B. is supported by an Ontario Graduate Scholarship (OGS).
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
GB conceived of the study and carried out all programming and analyses as a Ph.D. student in the lab of CH. CH supervised the study and provided valuable input for the evaluation analyses.
Electronic supplementary material
12859_2002_52_MOESM1_ESM.zip
Additional File 1: AllYeastPredictedComplexes.zip Zip file containing Pajek .net and annotation files for all 209 complexes found using MCODE on the set of all yeast interactions reported here. Various report files from MCODE are also included as well as basic instructions for using Pajek. (ZIP 510 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Bader, G.D., Hogue, C.W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics 4, 2 (2003). https://doi.org/10.1186/1471-2105-4-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2105-4-2