Abstract
Background
Antibody genes are diversified by somatic hypermutation (SHM), gene conversion and class-switch recombination. All three processes are initiated by the activation-induced deaminase (AID). According to a DNA deamination model of SHM, AID converts cytosine to uracil in DNA sequences. The initial deamination of cytosine leads to mutation and recombination in pathways involving replication, DNA mismatch repair and possibly base excision repair. The DNA sequence context of mutation hotspots at G-C pairs during SHM is DGYW/WRCH (G-C is a hotspot position, R = A/G, Y = T/C, W = A/T, D = A/G/T).
Results
To investigate the mechanisms of AID-induced mutagenesis in a model system, we studied the genetic consequences of AID expression in yeast. We constructed a yeast vector with an artificially synthesized human AID gene insert using codons common to highly expressed yeast genes. We found that expression of the artificial hAIDSc gene was moderately mutagenic in a wild-type strain and highly mutagenic in an ung1 uracil-DNA glycosylase-deficient strain. A majority of mutations were at G-C pairs. In the ung1 strain, C-G to T-A transitions were found almost exclusively, while a mixture of transitions with 12% transversions was characteristic in the wild-type strain. In the ung1 strain mutations that could have originated from deamination of the transcribed stand were found more frequently. In the wild-type strain, the strand bias was reversed. DGYW/WRCH motifs were preferential sites of mutations.
Conclusion
The results are consistent with the hypothesis that AID-mediated deamination of DNA is a major cause of mutations at G-C base pairs in immunoglobulin genes during SHM. The sequence contexts of mutations in yeast induced by AID and those of somatic mutations at G-C pairs in immunoglobulin genes are significantly similar. This indicates that the intrinsic substrate specificity of AID itself is a primary determinant of mutational hotspots at G-C base pairs during SHM.
Similar content being viewed by others
Background
The immune system uses several strategies to modify genetic material to generate various types of high affinity antibodies [1]. These strategies enable production of multiple antibody variants to a wide range of different antigens [2]. Initially, antigen receptors are generated by a site-specific recombination process called V(D)J recombination occurring in the bone marrow [3]. However, this is not sufficient to assure an adequate immune response. Mature B-lymphocytes migrate to the secondary lymphoid organs where they encounter antigens. Upon activation by antigens, mature B-lymphocytes begin to proliferate and form germinal centers, where immunoglobulin genes undergo additional modifications: class switch recombination (CSR), immunoglobulin gene conversion (IGC) and somatic hypermutation (SHM) [4]. SHM, IGC and CSR, all require active transcription [5] and generate diversity of antibodies, that is followed by selection leading to the production of high affinity antibodies [6]. The frequency of mutations during this process is up to six orders of magnitude higher than in other genes [6]. Most of the mutations are base pair substitutions, occurring with a similar frequency at G-C and A-T base pairs. Statistically preferred hotspots for mutations at G-C pairs are RG YW/WRC Y motifs (mutating G-C are underlined, R stands for purine base, Y stands for pyrimidine base and W stands for A or T) [7], or recently refined DG YW/WRC H motifs (D stands for G, T or A) [8]. Hotspots of mutations at A-T pairs are in WA /TW motifs (mutating A-T are underlined) [9].
A major breakthrough in understanding the mechanisms of CSR, IGC and SHM was the discovery that they all depend on activation-induced cytidine deaminase, AID [10–16]. Patients with defective AID have giant germinal centers and elevated levels of only one type of low-affinity antibodies, IgM. They suffer from recurrent bacterial infections in the respiratory tract [17] due to the lack of efficient antibody responses that depend on several crucial steps of B cell terminal differentiation including CSR and SHM. SHM is targeted to specific DNA regions in specialized tissues. Defects in this targeting may result in genome-wide mutagenesis and cancer. B-cell lymphomas possess translocations that bring proto-oncogenes into immunoglobulin loci (see [18]). Constitutive expression of AID in mice leads to an increase of tumor incidence [19].
When discovered, AID was thought to act in mutagenesis and recombination in immunity by RNA editing [10, 11, 20]. It was proposed that AID edits pre-mRNA encoding a nicking endonuclease that initiates SHM, IGC and CSR [5]. This model is called "RNA-editing" [20]. The AID is homologous to the known RNA-editing enzyme APOBEC1, which deaminates cytosine at position 6666 in ApoB100 mRNA and seemingly has no role in immunity. AID possesses the ability to deaminate cytidine, and shuttles between the nucleus and cytoplasm similar to APOBEC1 [4, 5, 21, 22]. A different hypothesis, called "DNA deamination", suggests AID deaminates cytosine directly and that uracil generated in this reaction triggers downstream reactions leading to genetic instability [23–26] (see [27–33] for reviews).
Experimental evidence is accumulating in favor of the DNA deamination hypothesis of AID function [29, 31–34]. AID is able to induce SHM and CSR in hybridomas and in fibroblasts, suggesting that it is the only B-cell specific component required for induction of both genetic events [13, 14, 35]. AID can also induce mutations when expressed in E. coli [24]. These mutations occur in the same DNA sequence motifs as mutations during SHM [8, 36]. Therefore, eukaryotic cell-specific components are not necessary for mutagenesis. This mutator effect is enhanced in uracil-DNA glycosylase-deficient ung1 strains, which are unable to repair uracil in DNA [37], suggesting that the deamination of cytosine to uracil in DNA is the cause of these mutations [24]. It was found that the expression of two homologous deaminases, APOBEC1 and APOBEC3G, is highly mutagenic in bacteria [38]. Almost all mutations arising upon expression of these deaminases were G-A to A-T transitions, consistent with the DNA deamination model. AID deaminates single-stranded and supercoiled double-stranded DNA [39–44] (see also review [31]). AID exhibits clear DNA sequence context specificity, which resembles the specificity of G-C to the A-T component of SHM mutagenesis (G YW/WRC motifs, see[8, 40, 44–46]). The specificity of induced mutations in bacteria is consistent with predominant deamination of the non-transcribed DNA strand [36, 45], which is thought to be single-stranded during transcription (reviewed in [31]). During SHM, however, both DNA strands are targeted for mutagenesis [7], [47]. This discrepancy between the parameters of SHM in vertebrates and deaminase-induced mutagenesis in prokaryotes still needs to be resolved.
To characterize the initial steps of AID-induced mutations, we examined the specificity of the mutator effect of human AID expressed in yeast. We constructed a yeast vector with an artificially synthesized human AID gene insert using codons common to highly expressed yeast genes. We found that expression of the artificial hAIDSc gene was moderately mutagenic in the wild-type strain and highly mutagenic in the ung1 strain, similar to expression of unmodified human AID [48]. This is consistent with the uracil DNA deamination model of mutagenesis. We identified a spectrum of mutations in the CAN1 gene occurring in wild-type and ung1 strains expressing hAIDSc. We compared the sequence context of AID-induced mutations in yeast at G-C bases with somatic mutations in immunoglobulin genes. These comparisons revealed significantly similar properties and further support the hypothesis that AID is a primary cause of mutations at G-C pairs in immunoglobulin genes during SHM.
Results
hAIDSc expression and its mutator effect
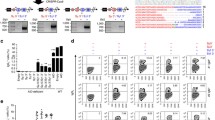
Codon usage is different in yeast and humans. To improve our system of expression of human AID over work published earlier [48], we constructed a new yeast expression vector with the human AID gene recoded to use the same codons utilized by highly expressed yeast genes and with a galactose-inducible promoter. Appropriate transformants were grown in galactose-containing medium and the AID protein was readily detected in yeast extracts by Western blot (Fig. 1, lane 3).
Western blot analysis of hAIDSc expression in yeast. Yeast strain CG379-3-29RL transformed by expression vector pESC-LEU2 or pESC-LEU-hAIDSc were grown to logarithmic phase in a complete minimal medium without leucine. Then cells were washed and transferred into similar medium but containing galactose instead of glucose. Yeast protein extracts were prepared from approximately 200 mg of cells by the glass beads cell disruption method as described in [85]. Proteins were separated using 4–12% gradient PAA NuPage gel (Invitrogen). Transfer to PVDF membrane and reaction with primary antibodies (mouse anti c-myc) and then secondary antibodies (goat antimouse) was accomplished as suggested by the vendor (Western Breeze kit, Invitrogen). Lane 1 – Molecular weight markers (Benchmark, His-Tagged) were detected with antiHisx6 antibodies. Lane 2 – extract of yeast strain containing vector pESC-LEU. Lane 3 – extract of yeast strain containing pESC-LEU-hAIDSc.
The expression of the hAIDSc did not result in any profound growth inhibition; the cell titer usually reached 5 × 107, which is typical for galactose-containing minimal medium (data not shown). Mutation rates were analyzed by fluctuation analysis (Table 1). Our strain permits the detection of various classes of genetic events (see Materials and Methods, and also [48, 49]). Using this strain we can obtain the express information about the specificity of the mutagenic effect.
The expression of the hAIDSc did not induce frameshift mutations to His+ (last column of Table 1). In Ung1+ strains, hAIDSc expression leads to a 7.6 fold increase in Canr forward mutations and a 3 – 6 fold increase in nonsense mutation reversion (Ade+, Trp+). The ung1 mutation per se led to a 5 – 10 fold increase of mutation rates as shown in rows one and four. When the hAIDSc was expressed in the ung1 strain, the mutator effect was multiplicative for Canr forward mutations (82 fold increase over the wild-type strain) and synergistic for nonsense mutation reversion (a 404 – 1290 fold increase over wild-type). TAG and TAA nonsense mutations cannot revert by true back-mutations via G-C to A-T transitions. We have previously shown by genetic analysis and sequencing of revertants that reversion is caused by dominant suppressors and most likely represent mutations in the anticodon of tRNA genes, which could be G-C to A-Ts [48]. The high response of ade5-1 and trp1-289 markers to hAIDSc may reflect the role of transcription in AID-induced hypermutation in yeast, since tRNA genes are transcribed differently from metabolic genes. The ura3-29 allele reversion was stimulated only weakly. It is known, that the allele reverts via various changes at G-C pair in "TCT" DNA sequence context [50], which is different from hotspots of AID deaminations. The results suggest that uracil DNA deamination is the primary source of mutation induced by the hAIDSc in yeast and are consistent with our previous studies [48]. Optimized codon usage did not lead to increased mutagenesis under conditions of constant induction of galactose promoter since the mutagenic potential of expression of the hAIDSc was comparable with expression of native human AID [48].
Mutagenic specificity of hAIDSc
We studied the specificity of mutations in the CAN1 gene induced by the expression of the hAIDSc. Independent Canr mutants were obtained under conditions of hAIDSc expression in the wild-type and the ung1 strain. Results of sequencing of mutants are summarized in the Tables 2, 3 and [see Additional file 1]. Most mutations (64 out of 70 in the wild-type and 62 out of 66 in the ung1 strain) were at G-C base pairs. Transversions comprised 12% of the mutations at the G-C pairs in wild-type and 1.6% in the ung1 strain (Table 4). The decreased proportion of transversions in the ung1 strain is consistent with the data obtained earlier in chicken and mice [26, 51]. We compared these spectra with the spectra of spontaneous mutations in CAN1 in the wild-type strains obtained by Rattrey and coauthors [52], Table 5. The major property of these mutation spectra was a high frequency of frameshift mutations (>20%) [52]. Another feature of the spontaneous mutations is a high frequency of mutations in A-T bases (>50%) and a higher frequency of transversions compared to transitions (>50%) (see also the breakdown of the types of spontaneous mutations obtained previously by other groups [53–55]). These features of CAN1 spontaneous mutations are similar to the properties of mutations observed in the yeast SUP4-o gene [56]. Thus, the spectra of mutations induced by the expression of the hAIDSc are different from spontaneous mutations in yeast genes. This result indicates that spontaneous mutations constitute a minor fraction (if any) of the mutations induced by the expression of the hAIDSc.
G-C mutations may arise by putative deamination on the transcribed or non-transcribed DNA strand. Mutations in the ung1 strain, representing deamination proclivity without of uracil repair, occur at a higher rate on the transcribed strand (Table 4). This is different from the effect of AID expression observed in the most E. coli selective systems [31]. In the wild-type strain, there is some prevalence of mutations due to putative non-transcribed strand deaminations (Table 4) suggesting the possibility that in our system the repair of uracil in the transcribed DNA strand is more efficient than in non-transcribed strand. Clearly, hAIDSc is targeted to both DNA strands in yeast, similar to somatic mutations in G-C bases during SHM [6, 7, 9, 57–59]. It is important to mention that, under normal circumstances, there are no differences in DNA strand preferences between mutation spectra from the wild-type and ung1 strains [60].
Next, we examined whether the DNA context of mutations induced by AID in yeast is similar to SHM mutations in mammals or in E. coli expressing hAID (Table 5). DG YW and G YW motifs [7, 8, 40] were under-represented in mutations occurring spontaneous in wild-type or ung1 strains (Table 5, row 1–2) and were 2 – 5 fold over-represented in mutations induced by AID in yeast (Table 5, rows 3–4). Lists of mutation hotspots are shown in Table 6. Distributions of AID-induced mutation hotspots in the wild-type and ung1 strains are significantly different (Table 6, P = 0.003). The specificity of mutations in yeast correlates better with the hotspot motifs for SHM in mice than does the specificity of AID induced mutations in E. coli. Indeed, out of four comparisons, the indices of preference for mutation hotspot motifs in yeast were higher than in E. coli (compare rows 3–4 with rows 5–8; rows 10–11 with rows 5–8, Table 5). Some properties of mutations in yeast resemble the in vitro AID induced mutation spectrum [40]. For example, one CG YW/WRC G sequence which is not mutable in SHM [8] had a high mutability in the ung1 mutation spectrum (Table 6). It has been suggested that a mammalian DNA repair enzyme, perhaps the uracil-DNA glycosylase, efficiently repairs the lesion of CpG dinucleotides and thus eliminates mutations from CG YW/WRC G motifs in vivo [8]. We have found 5 GG YW and 3 TG YW hotspots (Table 6). Interestingly, no hotspots were found in AG YW motifs (Table 6), which are the most frequent hotspot motifs in mammalian immunoglobulin genes [47]. The lack of mutation hotspots at AG YW could not be attributed to its lesser prevalence because the number of AGYW, GG YW, and TG YW motifs in CAN1 was similar (results not shown). However, a general pattern of mutations in yeast is similar to the targeting of somatic mutation in DG YW/WRC H mutable motifs, which are highly specific for SHM in mammals. In control, no significant targeting of mutations to DG YW/WRC H mutable motifs was found for spontaneous mutations in wild type or ung1 yeast (PW≤Wrandom > 0.05, Table 5, rows 1–2).
We did not find a substantial number of A-T mutations, which typically comprise one-half of all SHM [47, 58, 61]. This result corresponds with earlier published results on the expression of AID in E. coli [24, 36], in yeast [48], in murine fibroblasts [13], and in human hybridomas [35]. Apparently, additional components are required to model the full spectrum of SHM under conditions of AID expression in heterologous systems or in non-B cell tissues.
Discussion
Mutator efficiency and specificity of expression of hAIDSc
Yeast is a well-studied model eukaryotic organism used for various genetic studies. Yeast was used in this study to characterize the mutator effects of ectopic expression of human AID. The CAN1 reporter gene has been chosen because of numerous mutational studies [52, 54] and a well-characterized transcription pattern [62–64]. The results are different from studies of AID effects in prokaryotic models and in vitro experiments. We observed mutations arising due to deamination occurring in both DNA strands. In E.coli, transcription enhances deamination of the non-transcribed DNA strand, which is exposed as single-stranded DNA during the elongation reaction, but not mutation of the transcribed DNA strand, which is likely to be protected by E. coli RNA polymerase [42, 43, 65]. The observed DNA strand targeting of mutations in ung1 yeast closely resembles targeting of somatic mutations in vertebrate immunoglobulin genes (Table 5). Interestingly, there was a significant strand bias of mutations in wild-type yeast toward the non-transcribed strand (Tables 4 and 5). A more efficient repair of the transcribed DNA strand is one possible explanation of this asymmetry. Preferential nucleotide excision repair of the transcribed strand is a well-known phenomenon [66]. The possibility of transcription-coupled repair of uracil bases in DNA has not yet been thoroughly studied.
Interestingly, a strand bias toward the non-transcribed DNA strand was found in Ung-/- Msh2-/- mouse (Table 5, row 8) [67]. The difference between the number of mutations in DG YW/WRC H sites versus all other G:C sites in wild-type and Ung-/-Msh2-/- strains was statistically significant (Fisher exact test, P = 0.04). This may indicate that AID has a preference to the non-transcribed DNA strand as suggested earlier (see review [32]). An excessive DNA deamination of the non-transcribed DNA strand may be compensated by more efficient repair of this strand during the SHM phase 2 [67] causing approximately equal frequencies of mutations in both DNA strands (Table 5). More efficient repair of the non-transcribed strand is consistent with the idea of preferential targeting of the DNA polymerase η to the non-transcribed strand during SHM [9, 25]. In general, the strand specificity of SHM in Ung-/-Msh2-/- mouse is similar to AID-induced mutations in wild-type strains of yeast and E. coli (Table 5). Substantial differences between the observed targeting of AID to the mutable motifs in Ung-/- Msh2-/- and wild-type mouse (Table 5) are not consistent with a hypothesis that mutagenesis during the A:T-focused phase is nearly exclusively targeted to A:T bases [67, 68]. It is possible that mutagenesis during this phase is targeted to both A:T and G:C bases with a preference to A:T bases and no preference to DG YW/WRC H mutable motifs, this is consistent with the observed mutational and context specificity of the DNA polymerase η in vitro [9, 25, 69], DG YW/WRC H-independent mutagenesis of G:C bases will cause erosion of a high initial DG YW/WRC H motif specificity observed in Ung-/-Msh2-/- mouse (Table 5). There are also differences between strand specificity of Ung-/-Msh2-/- mouse and AID-induced mutations in human fibroblasts (Table 5), this might be explained by some differences in AID targeting or transcription-associated repair of uracil between B-lymphocytes and fibroblasts. All these results suggested that a weak strand bias is an intrinsic property of SHM.
A significant difference between in vitro systems and our experiments was observed. AID catalyses multiple deaminations in vitro [40]. We detected 11 clones with multiple mutations (10 clones with two mutations and one clone with three mutations) and checked the number of mutations in the first and second halves of CAN1. If multiple mutations emerge as a result of independent events, half of the clones are expected to have mutations in different halves of CAN1. In six out of 11 clones mutations were located in different parts of CAN1, thus independent mutation events is the most likely explanation. In general, the specificity and distribution of mutations in yeast did not exhibit a pattern of multiple mutations that would have been consistent with postulated processive action of AID [40]. These results are consistent with a high frequency of rearranged immunoglobulin V genes with one somatic mutation (for example, [70]). Apparent non-processive action of AID in vivo may be explained by a competition for binding to the CAN1 DNA sequence between AID and other proteins participating in transcription, replication and/or repair. For example, it is known that replication factor A stimulates AID [71], while the specificity of AID in vitro was studied on DNA without any additional factors. Clearly, this requires additional investigation.
Mechanisms of mutagenesis by AID
The mechanism of SHM initiated by AID may be as follows (see [23, 24, 27, 28, 67, 72]). Deamination of cytosine in DNA leads to the formation of a mismatched U-G base pair. If left unrepaired, further rounds of replication of uracil-containing DNA will generate only transition type mutations, G-C to AT. Uracil removal by uracil-DNA glycosylase leads to an apyrimidinic (AP) site. The AP site may be bypassed by a specialized DNA polymerase and, being a non-coding lesion, could lead to a transition or transversion mutation. The AP site may also be incised by AP-endonuclease and then repaired by the short patch base excision repair (BER) with involvement of error prone DNA polymerases with generation of transitions and transversions (e.g., see [72]). This mechanism generates mutations at G-C pairs. In order to explain the high frequency of mutations during the short patch BER reaction, it should be postulated that the relatively accurate DNA polymerase β is substituted in B-cells by an error-prone polymerase. The candidate is DNA polymerase ι, which is expressed in Burkitt's lymphoma cell line BL-2 [73] and whose inactivation suppresses SHM in this line [74]. However, 129-derived strains of mice, lacking active polymerase η, are fully proficient in SHM [75]. The reason for this discrepancy in not established yet.
Another type of mutation, which comprise about 50% of all mutations during SHM, is a change at the A-T base pairs [47, 58, 61]. The explanation of the mutation origins in A-T base pair is based on several observations. It is known that mutations at A-T base pairs depend on mismatch repair components MSH2, MSH6, EXO1 and error-prone DNA polymerase η [9, 29, 67, 68, 70, 72]. They are thought to be the result of error-prone bypass or repair of abasic site by error-prone polymerases, in particular, DNA polymerases η, ι and ζ [9, 25, 70, 73, 74, 76, 77] (reviewed in [28, 72, 78, 79]). It is possible that they are generated in the following way. Initiation of mismatch repair of a G-U base pair leads to a gap. Gaps may also be generated by long patch BER. Repair of gaps with the involvement of error-prone DNA polymerases may lead to mutations distal to initial G-U pair [25, 68]. Again, it should be postulated that gap repair is unusual in B-cell being inaccurate, since normally it is performed by highly accurate replicative DNA polymerases. The final feature of the current model of AID-initiated genetic modifications is that nicks and gaps, arising during DNA repair, stimulate recombination [16, 48]. SHM in the Ung1-/- mice is greatly biased in favour of transitions, since the pathway via apyrimidinic sites is blocked [26].
Mutations at A-T base pairs are absent in Msh2-/-Ung1-/- mice [67]. Is it important to notice that Ung1 is not a major enzyme involved in the overall repair of G:U mismatches in mice, as suggested by small mutator phenotypes in the Ung1-/- mice and the existence of the robust Smug1 glycosylase [80]. In B-cells, however, the Ung1 alone appears to be crucial for all genetic diversification processes [67, 81]. Mutations at A-T base are not observed when AID is expressed in prokaryotes or in yeast [31, 36, 48], and this work. Therefore, current model systems only partially reconstruct SHM. Delicate balance of mismatch repair and activity of error-prone polymerases, specific for B cells, might be required for the full spectrum of SHM mutations [68]. Changes in the chromatin structure are necessary for SHM [82] and this additional level of regulation should be taken into account when considering different SHM models.
Conclusion
In the present study, we have shown that expression of human AID is mutagenic in yeast and the mutagenic effect is one-two orders of magnitude higher in the ung1 strain. This observation suggests that the cause of the mutator effects is AID-driven DNA deamination. DNA sequence contexts of mutation hotspots coincide with DG YW/WRC H mutable motifs of somatic hypermutation, which is consistent with the DNA deamination model of SHM, suggesting that the intrinsic substrate specificity of AID itself is a primary determinant of mutational hotspots at G-C base pairs during SHM.
Methods
Construction of the expression vector
A new hAID gene was constructed using codons characteristic to highly expressed yeast genes. The DNA Builder program http://cbi.swmed.edu/computation/DNABuilder/dnabuilder.html and yeast codon usage data [83, 84] was used to construct a DNA sequence encoding human AID, with the preferable yeast codons. The DNA corresponding to this sequence and encoding for the c-myc Tag at the C-terminus (hAIDSc) was custom-synthesized and cloned into Bam HI-Sal I cut pESC-LEU (Stratagene) expression vector by the McLab Company (San Francisco). In this construct, the deaminase genes were placed downstream of the strong, galactose-inducible GAL1 promoter. DNA sequencing analysis confirmed the complete sequence of the insert. Protein production was demonstrated by Western blot as described earlier [85], with one modification – the Western Breeze Kit (Invitrogen) was used for detection of the protein in yeast extracts.
Yeast strains
For our experiments with the yeast vector expressing the deaminase genes we used yeast strain CG379-3-29RL (MAT α ura3Δleu2-3,112 trp1-289 bik1::ura3-29RL his7-2 ade5-1 lys2-Tn5-13) [48, 86, 87]. This strain allows concomitant measurement of mutation rates at several loci. These include a) the forward mutation rate at the CAN1 locus, where mutations reflect a variety of substitution, frameshift and more complex events; b) the rate of reversion of nonsense mutations: the trp1-289 (TAG [88]) and ade5-1 (TAA, [89]), where mutations reflect base substitutions in the nonsense codon as well as in suppressor genes encoding tRNAs; c) reversion of the ura3-29 missense mutation TCT which occurs via C-G to T-A transitions, C-G to C-A and C-G to G-C transversions [50]; d) reversion of the his7-2 mutant allele which occurs mainly via + 1 frameshifts in a homopolymeric AT run [49, 90].
Measurement of mutations rates
Mutation rates were analysed by fluctuation analysis [49, 90]. Independent transformants of the wild-type and ung1 derivatives of our basic strain were grown in a complete minimal medium lacking leucine to select for the plasmid, and containing galactose instead of glucose, to induce the hAIDSc expression.
Isolation and sequencing of can1 mutants
Yeast transformants patches originating from single colonies (64 per one plate) were replica-plated onto galactose-containing medium without leucine. After two days, they were replica-plated onto canavanine-containing medium to select for can1 mutants. After five days of incubation, one Canr colony was picked from each streak and streaked onto canavanine-containing medium. Chromosomal DNA from cells originating from one colony of these can1 mutants was isolated using a Yeast DNA Extraction Kit (Epicentre). Subsequent PCR amplification and sequencing was performed as described earlier [91].
In vivo and in vitro mutation spectra
Five in vivo and one in vitro mutation spectra, which have been described before [13, 26, 36, 40, 47] were used in this study. We consider that these large mutation spectra reflect intrinsic bias in mutation process. The compilation of somatic mutations in the VkOx transgene includes data derived from transgenic light chains with multiple copies of the transgene and from cells selected in gut Peyer's Patches (PP). The multiple copies are targeted in the same cell even when the light chain they encode is not part of the antigen binding antibody molecule. This implies that the majority of the mutations accumulated are unselected. In the case of PP derived cells, the selective pressure is multiple, therefore again, the common denominator of the biases observed would reflect the intrinsic biases [92, 93].
Statistical analyses
The Fisher exact test was used to compare frequencies of transitions and transversions. This test was also used to compare the number of mutations in DG YW/WRC H sites versus all other G-C sites in wild-type and Ung-/-Msh2-/- strains of mouse. A Monte Carlo modification of the Pearson χ2 test of spectra homogeneity [94] was used to compare mutation distributions along hotspot positions of the CAN1 sequence. Calculations were done using the COLLAPSE program ftp://ftp.bionet.nsc.ru/pub/biology/dbms/[95]. Mutations in the CAN1 gene were detected using the phenotypic assay described above, however the full list of detectable positions in this gene is not known. We predicted these positions using the SIFT program with default parameters [96]. Mutation hotspots were defined using a threshold for the number of mutations at a site. The threshold is established by analyzing the frequency distribution derived from a mutation spectrum using the CLUSTERM program http://www.itb.cnr.it/webmutation/[97]. Briefly, this program decomposes a mutation spectrum into several homogeneous classes of sites, with each class approximated by a binomial distribution. Variations in mutation frequencies among sites of the same class are random by definition (mutation probability is the same for all sites within a class), but differences between classes are statistically significant. Each site has a probability P(C) to be assigned to class C. A class with the highest mutation frequency is called hotspot class. Sites with of P(Chotspot) ≥ 0.95 of being assigned to the hotspot class Chotspot are defined as hotspot sites. This approach ensures that the assignment is statistically significant and robust (see Rogozin et al. [98] for detailed discussion of this approach and problems associated with its application).
Nucleotide sequence features can be correlated with a mutation spectrum and the correlation can be tested for statistical significance. The significance of correlations between the distribution of mutable motifs and mutations along a target sequence was measured by a Monte Carlo procedure (the CONSEN program) [7]. This approach takes into account frequencies of substitutions for each nucleotide, the possibility of multiple mutations in a site, and context of the mutating sites. The Monte Carlo simulation was run with weighted sites, with the weight of a site defined as:

where Mj is the number of mutations in site j. Wj weights were summed for all sites in the analyzed sequence resulting in the total weight W. A distribution of total weights Wrandom was calculated for 10,000 target sequences with randomly shuffled mutation spectra. Each of the resulting random mutation spectra contained the same number of mutations as the observed spectrum with the same distribution of mutations over randomly chosen sites. The distribution of Wrandom was used to calculate probability PW≤Wrandom. This probability is equal to the fraction of random spectra in which Wrandom is the same or greater than W. Small probability values (PW≤Wrandom ≤ 0.05) indicate a significant correlation between a mutable motif and the mutation frequency [7, 99].
References
Tonegawa S: Somatic generation of antibody diversity. Nature. 1983, 302: 575-581. 10.1038/302575a0.
Honjo T, Nakai S, Nishida Y, Kataoka T, Yamawaki-Kataoka Y, Takahashi N, Obata M, Shimizu A, Yaoita Y, Nikaido T, Ishida N: Rearrangements of immunoglobulin genes during differentiation and evolution. Immunol Rev. 1981, 59: 33-67.
Alt FW, Blackwell TK, Yancopoulos GD: Development of the primary antibody repertoire. Science. 1987, 238: 1079-1087.
Honjo T, Kinoshita K, Muramatsu M: Molecular mechanism of class switch recombination: linkage with somatic hypermutation. Annu Rev Immunol. 2002, 20: 165-196. 10.1146/annurev.immunol.20.090501.112049.
Kinoshita K, Honjo T: Linking class-switch recombination with somatic hypermutation. Nat Rev Mol Cell Biol. 2001, 2: 493-503. 10.1038/35080033.
Milstein C, Rada C: The maturation of immune responce. 1995, London: Academic Press
Rogozin IB, Kolchanov NA: Somatic hypermutagenesis in immunoglobulin genes. II. Influence of neighbouring base sequences on mutagenesis. Biochim Biophys Acta. 1992, 1171: 11-18.
Rogozin IB, Diaz M: DGYW/WRCH is a better predictor of mutability at G:C bases in Ig hypermutation than the widely accepted RGYW/WRCY motif and probably reflects a two-step activation-induced cytidine deaminase-triggered process. J Immunol. 2004, 172: 3382-3384.
Rogozin IB, Pavlov YI, Bebenek K, Matsuda T, Kunkel TA: Somatic mutation hotspots correlate with DNA polymerase eta error spectrum. Nat Immunol. 2001, 2: 530-536. 10.1038/88732.
Muramatsu M, Kinoshita K, Fagarasan S, Yamada S, Shinkai Y, Honjo T: Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potential RNA editing enzyme. Cell. 2000, 102: 553-563. 10.1016/S0092-8674(00)00078-7.
Revy P, Muto T, Levy Y, Geissmann F, Plebani A, Sanal O, Catalan N, Forveille M, Dufourcq-Labelouse R, Gennery A, Tezcan I, Ersoy F, Kayserili H, Ugazio AG, Brousse N, Muramatsu M, Notarangelo LD, Kinoshita K, Honjo T, Fischer A, Durandy A: Activation-induced cytidine deaminase (AID) deficiency causes the autosomal recessive form of the Hyper-IgM syndrome (HIGM2). Cell. 2000, 102: 565-575. 10.1016/S0092-8674(00)00079-9.
Durandy A, Honjo T: Human genetic defects in class-switch recombination (hyper-IgM syndromes). Curr Opin Immunol. 2001, 13: 543-548. 10.1016/S0952-7915(00)00256-9.
Yoshikawa K, Okazaki IM, Eto T, Kinoshita K, Muramatsu M, Nagaoka H, Honjo T: AID enzyme-induced hypermutation in an actively transcribed gene in fibroblasts. Science. 2002, 296: 2033-2036. 10.1126/science.1071556.
Okazaki IM, Kinoshita K, Muramatsu M, Yoshikawa K, Honjo T: The AID enzyme induces class switch recombination in fibroblasts. Nature. 2002, 416: 340-345. 10.1038/nature727.
Arakawa H, Hauschild J, Buerstedde JM: Requirement of the activation-induced deaminase (AID) gene for immunoglobulin gene conversion. Science. 2002, 295: 1301-1306. 10.1126/science.1067308.
Di Noia JM, Neuberger MS: Immunoglobulin gene conversion in chicken DT40 cells largely proceeds through an abasic site intermediate generated by excision of the uracil produced by AID-mediated deoxycytidine deamination. Eur J Immunol. 2004, 34: 504-508. 10.1002/eji.200324631.
Durandy A, Hivroz C, Mazerolles F, Schiff C, Bernard F, Jouanguy E, Revy P, DiSanto JP, Gauchat JF, Bonnefoy JY, Casanova JL, Fischer A: Abnormal CD40-mediated activation pathway in B lymphocytes from patients with hyper-IgM syndrome and normal CD40 ligand expression. J Immunol. 1997, 158: 2576-2584.
Kuppers R: Somatic hypermutation and B cell receptor selection in normal and transformed human B cells. Ann N Y Acad Sci. 2003, 987: 173-179.
Okazaki IM, Hiai H, Kakazu N, Yamada S, Muramatsu M, Kinoshita K, Honjo T: Constitutive expression of AID leads to tumorigenesis. J Exp Med. 2003, 197: 1173-1181. 10.1084/jem.20030275.
Honjo T, Muramatsu M, Fagarasan S: AID: how does it aid antibody diversity?. Immunity. 2004, 20: 659-668. 10.1016/j.immuni.2004.05.011.
Doi T, Kinoshita K, Ikegawa M, Muramatsu M, Honjo T: De novo protein synthesis is required for the activation-induced cytidine deaminase function in class-switch recombination. Proc Natl Acad Sci U S A. 2003, 100: 2634-2638. 10.1073/pnas.0437710100.
Ito S, Nagaoka H, Shinkura R, Begum N, Muramatsu M, Nakata M, Honjo T: Activation-induced cytidine deaminase shuttles between nucleus and cytoplasm like apolipoprotein B mRNA editing catalytic polypeptide 1. Proc Natl Acad Sci U S A. 2004, 101: 1975-1980. 10.1073/pnas.0307335101.
Poltoratsky V, Goodman MF, Scharff MD: Error-prone candidates vie for somatic mutation. J Exp Med. 2000, 192: F27-30. 10.1084/jem.192.10.F27.
Petersen-Mahrt SK, Harris RS, Neuberger MS: AID mutates E. coli suggesting a DNA deamination mechanism for antibody diversification. Nature. 2002, 418: 99-103. 10.1038/nature00862.
Pavlov YI, Rogozin IB, Galkin AP, Aksenova AY, Hanaoka F, Rada C, Kunkel TA: Correlation of somatic hypermutation specificity and A-T base pair substitution errors by DNA polymerase eta during copying of a mouse immunoglobulin kappa light chain transgene. Proc Natl Acad Sci U S A. 2002, 99: 9954-9959. 10.1073/pnas.152126799.
Rada C, Williams GT, Nilsen H, Barnes DE, Lindahl T, Neuberger MS: Immunoglobulin isotype switching is inhibited and somatic hypermutation perturbed in UNG-deficient mice. Curr Biol. 2002, 12: 1748-1755. 10.1016/S0960-9822(02)01215-0.
Storb U, Stavnezer J: Immunoglobulin genes: generating diversity with AID and UNG. Curr Biol. 2002, 12: R725-727. 10.1016/S0960-9822(02)01247-2.
Diaz M, Storb U: A novel cytidine deaminase AIDs in the delivery of error-prone polymerases to immunoglobulin genes. DNA Repair (Amst). 2003, 2: 623-627. 10.1016/S1568-7864(02)00240-9.
Neuberger MS, Harris RS, Di Noia J, Petersen-Mahrt SK: Immunity through DNA deamination. Trends Biochem Sci. 2003, 28: 305-312. 10.1016/S0968-0004(03)00111-7.
Li Z, Woo CJ, Iglesias-Ussel MD, Ronai D, Scharff MD: The generation of antibody diversity through somatic hypermutation and class switch recombination. Genes Dev. 2004, 18: 1-11. 10.1101/gad.1161904.
Bhagwat AS: DNA-cytosine deaminases: from antibody maturation to antiviral defense. DNA Repair (Amst). 2004, 3: 85-89. 10.1016/j.dnarep.2003.09.008.
Barreto VM, Ramiro AR, Nussenzweig MC: Activation-induced deaminase: controversies and open questions. Trends Immunol. 2005, 26: 90-96. 10.1016/j.it.2004.12.004.
Pham P, Bransteitter R, Goodman MF: Reward versus Risk: DNA cytidine deaminases triggering immunity and disease. Biochemistry. 2005, 44: 2703-2715. 10.1021/bi047481+.
Luo Z, Ronai D, Scharff MD: The role of activation-induced cytidine deaminase in antibody diversification, immunodeficiency, and B-cell malignancies. J Allergy Clin Immunol. 2004, 114: 726-735. 10.1016/j.jaci.2004.07.049. quiz 736.
Martin A, Bardwell PD, Woo CJ, Fan M, Shulman MJ, Scharff MD: Activation-induced cytidine deaminase turns on somatic hypermutation in hybridomas. Nature. 2002, 415: 802-806.
Beale RC, Petersen-Mahrt SK, Watt IN, Harris RS, Rada C, Neuberger MS: Comparison of the differential context-dependence of DNA deamination by APOBEC enzymes: correlation with mutation spectra in vivo. J Mol Biol. 2004, 337: 585-596. 10.1016/j.jmb.2004.01.046.
Lindahl T: DNA glycosylases, endonucleases for apurinic/apyrimidinic sites, and base excision-repair. Prog Nucleic Acid Res Mol Biol. 1979, 22: 135-192.
Harris RS, Petersen-Mahrt SK, Neuberger MS: RNA editing enzyme APOBEC1 and some of its homologs can act as DNA mutators. Mol Cell. 2002, 10: 1247-1253. 10.1016/S1097-2765(02)00742-6.
Bransteitter R, Pham P, Scharff MD, Goodman MF: Activation-induced cytidine deaminase deaminates deoxycytidine on single-stranded DNA but requires the action of RNase. Proc Natl Acad Sci U S A. 2003, 100: 4102-4107. 10.1073/pnas.0730835100.
Pham P, Bransteitter R, Petruska J, Goodman MF: Processive AID-catalysed cytosine deamination on single-stranded DNA simulates somatic hypermutation. Nature. 2003, 424: 103-107. 10.1038/nature01760.
Dickerson SK, Market E, Besmer E, Papavasiliou FN: AID mediates hypermutation by deaminating single stranded DNA. J Exp Med. 2003, 197: 1291-1296. 10.1084/jem.20030481.
Ramiro AR, Stavropoulos P, Jankovic M, Nussenzweig MC: Transcription enhances AID-mediated cytidine deamination by exposing single-stranded DNA on the nontemplate strand. Nat Immunol. 2003, 4: 452-456. 10.1038/ni920.
Sohail A, Klapacz J, Samaranayake M, Ullah A, Bhagwat AS: Human activation-induced cytidine deaminase causes transcription-dependent, strand-biased C to U deaminations. Nucleic Acids Res. 2003, 31: 2990-2994. 10.1093/nar/gkg464.
Shen HM, Storb U: Activation-induced cytidine deaminase (AID) can target both DNA strands when the DNA is supercoiled. Proc Natl Acad Sci U S A. 2004, 101: 12997-13002. 10.1073/pnas.0404974101.
Bransteitter R, Pham P, Calabrese P, Goodman MF: Biochemical analysis of hyper-mutational targeting by wild type and mutant AID. J Biol Chem. 2004, 279: 51612-51621. 10.1074/jbc.M408135200.
Yu K, Huang FT, Lieber MR: DNA substrate length and surrounding sequence affect the activation-induced deaminase activity at cytidine. J Biol Chem. 2004, 279: 6496-6500. 10.1074/jbc.M311616200.
Milstein C, Neuberger MS, Staden R: Both DNA strands of antibody genes are hypermutation targets. Proc Natl Acad Sci U S A. 1998, 95: 8791-8794. 10.1073/pnas.95.15.8791.
Poltoratsky VP, Wilson SH, Kunkel TA, Pavlov YI: Recombinogenic phenotype of human activation-induced cytosine deaminase. J Immunol. 2004, 172: 4308-4313.
Pavlov YI, Shcherbakova PV, Kunkel TA: In vivo consequences of putative active site mutations in yeast DNA polymerases alpha, epsilon, delta, and zeta. Genetics. 2001, 159: 47-64.
Pavlov YI, Newlon CS, Kunkel TA: Yeast origins establish a strand bias for replicational mutagenesis. Mol Cell. 2002, 10: 207-213. 10.1016/S1097-2765(02)00567-1.
Di Noia J, Neuberger MS: Altering the pathway of immunoglobulin hypermutation by inhibiting uracil-DNA glycosylase. Nature. 2002, 419: 43-48. 10.1038/nature00981.
Rattray AJ, Shafer BK, McGill CB, Strathern JN: The roles of REV3 and RAD57 in double-strand-break-repair-induced mutagenesis of Saccharomyces cerevisiae. Genetics. 2002, 162: 1063-1077.
Kunz BA, Ramachandran K, Vonarx EJ: DNA sequence analysis of spontaneous mutagenesis in Saccharomyces cerevisiae. Genetics. 1998, 148: 1491-1505.
Tishkoff DX, Filosi N, Gaida GM, Kolodner RD: A novel mutation avoidance mechanism dependent on S. cerevisiae RAD27 is distinct from DNA mismatch repair. Cell. 1997, 88: 253-263. 10.1016/S0092-8674(00)81846-2.
Niimi A, Limsirichaikul S, Yoshida S, Iwai S, Masutani C, Hanaoka F, Kool ET, Nishiyama Y, Suzuki M: Palm mutants in DNA polymerases alpha and eta alter DNA replication fidelity and translesion activity. Mol Cell Biol. 2004, 24: 2734-2746. 10.1128/MCB.24.7.2734-2746.2004.
Kang XL, Yadao F, Gietz RD, Kunz BA: Elimination of the yeast RAD6 ubiquitin conjugase enhances base-pair transitions and G.C->T.A transversions as well as transposition of the Ty element: implications for the control of spontaneous mutation. Genetics. 1992, 130: 285-294.
Foster SJ, Dorner T, Lipsky PE: Somatic hypermutation of VkappaJkappa rearrangements: targeting of RGYW motifs on both DNA strands and preferential selection of mutated codons within RGYW motifs. Eur J Immunol. 1999, 29: 4011-4021. 10.1002/(SICI)1521-4141(199912)29:12<4011::AID-IMMU4011>3.0.CO;2-W.
Spencer J, Dunn M, Dunn-Walters DK: Characteristics of sequences around individual nucleotide substitutions in IgVH genes suggest different GC and AT mutators. J Immunol. 1999, 162: 6596-6601.
Boursier L, Su W, Spencer J: Analysis of strand biased 'G'.C hypermutation in human immunoglobulin V(lambda) gene segments suggests that both DNA strands are targets for deamination by activation-induced cytidine deaminase. Mol Immunol. 2004, 40: 1273-1278. 10.1016/j.molimm.2003.11.026.
Impellizzeri KJ, Anderson B, Burgers PM: The spectrum of spontaneous mutations in a Saccharomyces cerevisiae uracil-DNA-glycosylase mutant limits the function of this enzyme to cytosine deamination repair. J Bacteriol. 1991, 173: 6807-6810.
Rogozin IB, Pavlov YI, Kunkel TA: Response 1 to 'Smaller role for pol ?. Nature Immunology. 2001, 2: 983-984. 10.1038/ni1101-983.
Coffman JA, Rai R, Cooper TG: Genetic evidence for Gln3p-independent, nitrogen catabolite repression-sensitive gene expression in Saccharomyces cerevisiae. J Bacteriol. 1995, 177: 6910-6918.
Rinckel LA, Garfinkel DJ: Influences of histone stoichiometry on the target site preference of retrotransposons Ty1 and Ty2 in Saccharomyces cerevisiae. Genetics. 1996, 142: 761-776.
van der Merwe GK, Cooper TG, van Vuuren HJ: Ammonia regulates VID30 expression and Vid30p function shifts nitrogen metabolism toward glutamate formation especially when Saccharomyces cerevisiae is grown in low concentrations of ammonia. J Biol Chem. 2001, 276: 28659-28666. 10.1074/jbc.M102280200.
Chaudhuri J, Tian M, Khuong C, Chua K, Pinaud E, Alt FW: Transcription-targeted DNA deamination by the AID antibody diversification enzyme. Nature. 2003, 422: 726-730. 10.1038/nature01574.
Hanawalt PC: Subpathways of nucleotide excision repair and their regulation. Oncogene. 2002, 21: 8949-8956. 10.1038/sj.onc.1206096.
Rada C, Di Noia JM, Neuberger MS: Mismatch recognition and uracil excision provide complementary paths to both Ig switching and the A/T-focused phase of somatic mutation. Mol Cell. 2004, 16: 163-171. 10.1016/j.molcel.2004.10.011.
Wilson TM, Vaisman A, Martomo SA, Sullivan P, Lan L, Hanaoka F, Yasui A, Woodgate R, Gearhart PJ: MSH2-MSH6 stimulates DNA polymerase eta, suggesting a role for A:T mutations in antibody genes. J Exp Med. 2005, 201: 637-645. 10.1084/jem.20042066.
Matsuda T, Bebenek K, Masutani C, Rogozin IB, Hanaoka F, Kunkel TA: Error rate and specificity of human and murine DNA polymerase eta. J Mol Biol. 2001, 312: 335-346. 10.1006/jmbi.2001.4937.
Zeng X, Winter DB, Kasmer C, Kraemer KH, Lehmann AR, Gearhart PJ: DNA polymerase eta is an A-T mutator in somatic hypermutation of immunoglobulin variable genes. Nat Immunol. 2001, 2: 537-541. 10.1038/88740.
Chaudhuri J, Khuong C, Alt FW: Replication protein A interacts with AID to promote deamination of somatic hypermutation targets. Nature. 2004, 430: 992-998. 10.1038/nature02821.
Kunkel TA, Pavlov YI, Bebenek K: Functions of human DNA polymerases eta, kappa and iota suggested by their properties, including fidelity with undamaged DNA templates. DNA Repair (Amst). 2003, 2: 135-149. 10.1016/S1568-7864(02)00224-0.
Poltoratsky V, Woo CJ, Tippin B, Martin A, Goodman MF, Scharff MD: Expression of error-prone polymerases in BL2 cells activated for Ig somatic hypermutation. Proc Natl Acad Sci U S A. 2001, 98: 7976-7981. 10.1073/pnas.141222198.
Faili A, Aoufouchi S, Flatter E, Gueranger Q, Reynaud CA, Weill JC: Induction of somatic hypermutation in immunoglobulin genes is dependent on DNA polymerase iota. Nature. 2002, 419: 944-947. 10.1038/nature01117.
McDonald JP, Frank EG, Plosky BS, Rogozin IB, Masutani C, Hanaoka F, Woodgate R, Gearhart PJ: 129-derived strains of mice are deficient in DNA polymerase iota and have normal immunoglobulin hypermutation. J Exp Med. 2003, 198: 635-643. 10.1084/jem.20030767.
Tissier A, Frank EG, McDonald JP, Vaisman A, Fernandez de Henestrosa AR, Boudsocq F, McLenigan MP, Woodgate R: Biochemical characterization of human DNA polymerase iota provides clues to its biological function. Biochem Soc Trans. 2001, 29: 183-187. 10.1042/BST0290183.
Zan H, Komori A, Li Z, Cerutti A, Schaffer A, Flajnik MF, Diaz M, Casali P: The translesion DNA polymerase zeta plays a major role in Ig and bcl-6 somatic hypermutation. Immunity. 2001, 14: 643-653. 10.1016/S1074-7613(01)00142-X.
Goodman MF: Error-prone repair DNA polymerases in prokaryotes and eukaryotes. Annu Rev Biochem. 2002, 71: 17-50. 10.1146/annurev.biochem.71.083101.124707.
Neuberger MS, Di Noia JM, Beale RC, Williams GT, Yang Z, Rada C: Somatic hypermutation at A.T pairs: polymerase error versus dUTP incorporation. Nat Rev Immunol. 2005, 5: 171-178. 10.1038/nri1553.
Nilsen H, Rosewell I, Robins P, Skjelbred CF, Andersen S, Slupphaug G, Daly G, Krokan HE, Lindahl T, Barnes DE: Uracil-DNA glycosylase (UNG)-deficient mice reveal a primary role of the enzyme during DNA replication. Mol Cell. 2000, 5: 1059-1065. 10.1016/S1097-2765(00)80271-3.
Barnes DE, Lindahl T: Repair and Genetic Consequences of Endogenous DNA Base Damage in Mammalian Cells. Annu Rev Genet. 2004, 38: 445-476. 10.1146/annurev.genet.38.072902.092448.
Woo CJ, Martin A, Scharff MD: Induction of somatic hypermutation is associated with modifications in immunoglobulin variable region chromatin. Immunity. 2003, 19: 479-489. 10.1016/S1074-7613(03)00261-9.
Bennetzen JL, Hall BD: Codon selection in yeast. J Biol Chem. 1982, 257: 3026-3031.
Jansen R, Bussemaker HJ, Gerstein M: Revisiting the codon adaptation index from a whole-genome perspective: analyzing the relationship between gene expression and codon occurrence in yeast using a variety of models. Nucleic Acids Res. 2003, 31: 2242-2251. 10.1093/nar/gkg306.
Pavlov YI, Nguyen D, Kunkel TA: Mutator effects of overproducing DNA polymerase eta (Rad30) and its catalytically inactive variant in yeast. Mutat Res. 2001, 478: 129-139.
Shcherbakova PV, Noskov VN, Pshenichnov MR, Pavlov YI: Base analog 6-N-hydroxylaminopurine mutagenesis in the yeast Saccharomyces cerevisiae is controlled by replicative DNA polymerases. Mutat Res. 1996, 369: 33-44.
Shcherbakova PV, Pavlov YI: 3'–>5' exonucleases of DNA polymerases epsilon and delta correct base analog induced DNA replication errors on opposite DNA strands in Saccharomyces cerevisiae. Genetics. 1996, 142: 717-726.
Calderon IL, Contopoulou CR, Mortimer RK: Isolation of a DNA fragment that is expressed as an amber suppressor when present in high copy number in yeast. Gene. 1984, 29: 69-76. 10.1016/0378-1119(84)90167-7.
Achilli A, Matmati N, Casalone E, Morpurgo G, Lucaccioni A, Pavlov YI, Babudri N: The exceptionally high rate of spontaneous mutations in the polymerase delta proofreading exonuclease-deficient Saccharomyces cerevisiae strain starved for adenine. BMC Genet. 2004, 5: 34-10.1186/1471-2156-5-34.
Shcherbakova PV, Kunkel TA: Mutator phenotypes conferred by MLH1 overexpression and by heterozygosity for mlh1 mutations. Mol Cell Biol. 1999, 19: 3177-3183.
Kozmin SG, Pavlov YI, Kunkel TA, Sage E: Roles of Saccharomyces cerevisiae DNA polymerases Poleta and Polzeta in response to irradiation by simulated sunlight. Nucleic Acids Res. 2003, 31: 4541-4552. 10.1093/nar/gkg489.
Betz AG, Rada C, Pannell R, Milstein C, Neuberger MS: Passenger transgenes reveal intrinsic specificity of the antibody hypermutation mechanism: clustering, polarity, and specific hot spots. Proc Natl Acad Sci U S A. 1993, 90: 2385-2388.
Gonzalez-Fernandez A, Milstein C: Analysis of somatic hypermutation in mouse Peyer's patches using immunoglobulin kappa light-chain transgenes. Proc Natl Acad Sci U S A. 1993, 90: 9862-9866.
Adams WT, Skopek TR: Statistical test for the comparison of samples from mutational spectra. J Mol Biol. 1987, 194: 391-396. 10.1016/0022-2836(87)90669-3.
Khromov-Borisov NN, Rogozin IB, Pegas Henriques JA, de Serres FJ: Similarity pattern analysis in mutational distributions. Mutat Res. 1999, 430: 55-74.
Ng PC, Henikoff S: SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res. 2003, 31: 3812-3814. 10.1093/nar/gkg509.
Glazko GV, Milanesi L, Rogozin IB: The subclass approach for mutational spectrum analysis: application of the SEM algorithm. J Theor Biol. 1998, 192: 475-487. 10.1006/jtbi.1998.0668.
Rogozin IB, Kondrashov FA, Glazko GV: Use of mutation spectra analysis software. Hum Mutation. 2001, 17: 83-102. 10.1002/1098-1004(200102)17:2<83::AID-HUMU1>3.0.CO;2-E.
Rogozin IB, Pavlov YI: Theoretical analysis of mutation hotspots and their DNA sequence context specificity. Mutat Res. 2003, 544: 65-85. 10.1016/S1383-5742(03)00032-2.
Acknowledgements
This work was supported in part by MedCen Foundation and RFBR. We are grateful to Barry Gold, Eugene Koonin, Elena Vasunina, Robert Lahue and Tadayoshi Bessho for fruitful discussions. We thank Dr. Polina Shcherbakova for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
This work was designed by YIP, TAK and IBR. YIP and CF were involved in vector construction, Western blot analysis of proteins, mutation rates measurements and isolation of can1 mutants. VIM and LA performed PCR and DNA sequencing and were involved in analysis of mutations. IBR did statistical analysis of mutation spectra.
Electronic supplementary material
12865_2005_71_MOESM1_ESM.pdf
Additional File 1: Spectra of can1 mutations in yeast. Types of nucleotides changes found are shown above the DNA sequence of the non-transcribed strand. A. ung1 strain, B. wild-type strain. Most of the mutations were single base pair substitutions. The multiple mutations found in the CAN1 are listed below. wild-type strain 452,C->T; 1728,T->G 123,A->G; 1392,G->A 123,A->G; 412,C->T 589,T->A; 1151,C->G 1485,G->T; 1486,C->A ung1 strain 553,C->A; 1633,C->T 1018,G->A; 1728,T->G 299,G->A; 1475,C->T 424,G->A; 980,G->A 572,T->C; 573,C->T 509,G->A; 516,G->A; 806,G->A (PDF 48 KB)
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
Open Access This article is published under license to BioMed Central Ltd. This is an Open Access article is distributed under the terms of the Creative Commons Attribution License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Mayorov, V.I., Rogozin, I.B., Adkison, L.R. et al. Expression of human AID in yeast induces mutations in context similar to the context of somatic hypermutation at G-C pairs in immunoglobulin genes. BMC Immunol 6, 10 (2005). https://doi.org/10.1186/1471-2172-6-10
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2172-6-10