Abstract
Background
Obesity associated type 2 diabetes mellitus is a metabolic disorder ; however, the etiology of obesity associated type 2 diabetes mellitus remains largely unknown. There is an urgent need to further broaden the understanding of the molecular mechanism associated in obesity associated type 2 diabetes mellitus.
Methods
To screen the differentially expressed genes (DEGs) that might play essential roles in obesity associated type 2 diabetes mellitus, the publicly available expression profiling by high throughput sequencing data (GSE143319) was downloaded and screened for DEGs. Then, Gene Ontology (GO) and REACTOME pathway enrichment analysis were performed. The protein - protein interaction network, miRNA - target genes regulatory network and TF-target gene regulatory network were constructed and analyzed for identification of hub and target genes. The hub genes were validated by receiver operating characteristic (ROC) curve analysis and RT- PCR analysis. Finally, a molecular docking study was performed on over expressed proteins to predict the target small drug molecules.
Results
A total of 820 DEGs were identified between healthy obese and metabolically unhealthy obese, among 409 up regulated and 411 down regulated genes. The GO enrichment analysis results showed that these DEGs were significantly enriched in ion transmembrane transport, intrinsic component of plasma membrane, transferase activity, transferring phosphorus-containing groups, cell adhesion, integral component of plasma membrane and signaling receptor binding, whereas, the REACTOME pathway enrichment analysis results showed that these DEGs were significantly enriched in integration of energy metabolism and extracellular matrix organization. The hub genes CEBPD, TP73, ESR2, TAB1, MAP 3K5, FN1, UBD, RUNX1, PIK3R2 and TNF, which might play an essential role in obesity associated type 2 diabetes mellitus was further screened.
Conclusions
The present study could deepen the understanding of the molecular mechanism of obesity associated type 2 diabetes mellitus, which could be useful in developing therapeutic targets for obesity associated type 2 diabetes mellitus.
Similar content being viewed by others
Introduction
Obesity associated type 2 diabetes is one of the most common metabolic disorder worldwide [1]. Type 2 diabetes mellitus is characterized by insulin deficiency due to pancreatic β-cell inactivation and insulin resistance [2]. Genetic factors, hyperinsulinemia, atherogenic dyslipidemia, glucose intolerance, hypertension, prothrombic state, hyperuricemia and polycystic ovary syndrome are the key risk factors for the occurrence and progression of type 2 diabetes mellitus [3]. Obesity associated type 2 diabetes mellitus affects the vital organs such as heart [4], brain [5], kidney [6] and eye [7]. Etiology and advancement of obesity associated type 2 diabetes mellitus is more complex and still understandable. Therefore, it is essential to understand the precise molecular mechanisms associated in the progression of obesity associated type 2 diabetes mellitus and thus to establish valid diagnostic and therapeutic strategies.
Current evidence has shown that genetic predisposition plays a key role in the advancement of obesity associated type 2 diabetes mellitus [8]. Recently, several genes and pathways have been found to participate in the occurrence and advancement of obesity associated type 2 diabetes mellitus [9], including FGF21 [10], pro-opiomelanocortin (POMC) [11], PI3K/AKT pathway [12] and JAK/STAT pathway [13]. However, the current knowledge is insufficient to explain and understand how these crucial genes and signaling pathways are associated with advancement of obesity associated type 2 diabetes mellitus. Therefore, there is a great need to find new prognostic and diagnostics biomarkers, and to advance novel techniques to enlighten the molecular mechanisms controlling the progression of obesity associated type 2 diabetes mellitus.
Bioinformatics analysis of expression profiling by high throughput sequencing data has shown great promise to discover potential key genes and signaling pathways with significant roles in metabolic disorder [14], to identify new prognostic and diagnostics biomarkers, and biological processes implicated in obesity associated type 2 diabetes mellitus. In this investigation, using bioinformatics analysis, we aimed to investigate expression profiling by high throughput sequencing data to determine differentially expressed genes (DEGs) and significant pathways in obesity associated type 2 diabetes mellitus. After searching the Gene Expression Omnibus (GEO) database [15], we selected RNA sequencing dataset GSE143319 for identifying DEGs for obesity associated type 2 diabetes mellitus. This dataset gives more information about obesity associated type 2 diabetes mellitus elevates patient’s risk of nonalcoholic steatohepatitis (NASH), cardiovascular disease and cancer. Gene Ontology (GO) and pathway enrichment analysis were performed. A hub and target genes were identified from protein-protein interaction (PPI) network, modules, miRNA-target genes regulatory network and TF-target gene regulatory network. Subsequently, hub genes were validated by using receiver operating characteristic (ROC) curve and RT- PCR analysis. Finally, molecular docking studies performed for prediction of small drug molecules.
Materials and Methods
RNA sequencing data
The expression profiling by high throughput sequencing dataset GSE143319 deposited by Ding et al [16] into the GEO database were obtained on the GPL20301 platform (Illumina HiSeq 4000 (Homo sapiens)). This dataset is provided for 30 samples, including 15 samples of metabolically healthy obese and 15 samples of a metabolically unhealthy obese.
Identification of DEGs
The limma [17] in R bioconductor package was utilized to screen DEGs between metabolically healthy obese and metabolically unhealthy obese. These DEGs were identified as important genes that might play an important role in the development of obesity associated type 2 diabetes mellitus. The cutoff criterion were ∣log fold change (FC)∣ > 0.2587 for up regulated genes, ∣log fold change (FC)∣ < -0.2825 for down regulated genes and adjusted P value < 0.05.
GO and pathway enrichment analyses
ToppGene (ToppFun) (https://toppgene.cchmc.org/enrichment.jsp) [18], which is a useful online database that integrates biologic data and provides a comprehensive set of functional annotation information of genes as well as proteins for users to analyze the functions or signaling pathways. GO (https://geneontology.org/) [19] enrichment analysis (biologic processes [BP], cellular components [CC], and molecular functions [MF]) is a strong bioinformatics tool to analyze and annotate genes. The REACTOME (https://reactome.org/) [20] is a pathway database resource for understanding high-level gene functions and linking genomic information from large scale molecular datasets. To analyze the function of the DEGs, biologic analyses were performed using GO and REACTOME pathway enrichment analysis via ToppGene online database.
PPI network construction and module analysis
IMEX interactome (https://www.imexconsortium.org/) [21] online PPI database was using to identify the hub gene information in PPI network. Analyzing the interactions and functions of DEGs might give information about the controlling the progression of obesity associated type 2 diabetes mellitus. Cytoscape (version 3.8.2) (www.cytoscape.org) is a bioinformatics platform for constructing and visualizing PPI network [22]. Therefore, the topological properties includes node degree [23], betweenness centrality [24], stress centrality [25], closeness centrality [26] are analyzed in using Java plug-in Network Analyzer to obtain hub genes in the PPI network. The plug-in PEWCC1 of Cytoscape was applied to detect densely connected regions in PPI network. The significant modules in the PPI network was selected using PEWCC1 (https://apps.cytoscape.org/apps/PEWCC1) [27]. The criteria for selection were set as follows: Max depth = 100, degree cut-off = 2, node score cut-off = 0.2, PEWCC1 scores >5, and K-score = 2.
Target gene – miRNA regulatory network construction and analysis
Obesity associated type 2 diabetes mellitus relating miRNAs and experimentally validated target genes were identified from miRNet database (https://www.mirnet.ca/) [28]. Obesity associated type 2 diabetes mellitus relating miRNAs and target genes were identified through target genes - miRNA regulatory network. Then the target genes - miRNA regulatory network was constructed and visualized by using Cytoscape software.
Target gene – TF network regulatory construction and analysis
Obesity associated type 2 diabetes mellitus relating TFs and experimentally validated target genes were identified from TFs database NetworkAnalyst database (https://www.networkanalyst.ca/) [29]. Obesity associated type 2 diabetes mellitus relating TFs and target genes were identified through target genes - TF regulatory network. Then the target genes -TF regulatory network was constructed and visualized by using Cytoscape software.
Receiver operating characteristic (ROC) curve analysis
The ROC curve was used to calculate classifiers in bioinformatics applications. To further assess the predictive accuracy of the hub genes, ROC analysis was performed to discriminate metabolically healthy obese from metabolically unhealthy obese. ROC curves for hub genes were generated using pROC in R [30] based on the obtained DEGs and their expression profiling by high throughput sequencing dataset. The area under the curve (AUC) was evaluated and used to compare the diagnostic value of hub genes.
Validation of the expression levels of candidate genes by RT-PCR
Quantitative RT-PCR was conducted to validate the expressions of these hub genes in obesity associated type 2 diabetes mellitus. Total RNAs were extracted from Primary Subcutaneous Pre adipocytes; Normal Human cell line (ATCC® PCS-210-010™) and 3T3-L1 cells (ATCC® CL-173) using TRI Reagent® (Sigma, USA) according to instruction, followed by reverse transcription with Reverse transcription cDNA kit (Thermo Fisher Scientific, Waltham, MA, USA) and cDNA amplification through 7 Flex real-time PCR system (Thermo Fisher Scientific, Waltham, MA, USA). The expressions of hub genes were normalized to against beta actin expression. The data were calculated by the 2−ΔΔCt method [31]. A primer used in the current investigation was listed in Table 1.
Molecular docking studies
The Surflex-Docking docking studies for the designed molecules were performed using module SYBYL-X 2.0 perpetual software. Using ChemDraw Tools, the molecules were sketched and imported and saved into sdf format using open free software from Babel. The co-crystallised protein structures of CEBPD, TP73, ESR2, TAB1 and MAP 3K5 of its PDB code 3L4W, 2XWC, 1U3Q, 5NZZ & 2CLQwas extracted from Protein Data Bank [32,33,34,35,36]. Together with the TRIPOS force field, GasteigerHuckel (GH) charges were added to all designed derivatives for the structure optimization process. Furthermore, energy minimization was carried out using MMFF94s and MMFF94 algorithm process. The processing of protein was accomplished after the incorporation of protein. The co-crystallized ligand and all water molecules were expelled from the crystal structure; more hydrogen was added and the side chain was optimized. TRIPOS force field was used to minimize complexity of structure. The interaction efficiency of the compounds with the receptor was expressed in kcal / mol units by the Surflex-Dock score. The best spot between the protein and the ligand was inserted into the molecular region. The visualization of ligand interaction with receptor is done by using discovery studio visualizer.
Results
Identification of DEGs
As presented in the cluster heat map of Fig. 1, total 820 DEGs, comprising 409 up regulated and 411 down regulated genes, were identified between metabolically healthy obese samples and metabolically unhealthy obese samples. DEGs were illustrated by volcano plot (Fig.2), and the top up regulated and down regulated DEGs are listed in Table 2.
Gene ontology and pathway enrichment analyses
DEGs were divided into up regulated genes and down regulated genes. GO and REACTOME pathway enrichment analysis were conducted for DEGs. Results of GO categories were presented by functional groups, which include BP, CC, and MF, and are listed in Table 3. In group BP, up regulated genes enriched in regulation of ion transmembrane transport and oxoacid metabolic process, while the down regulated genes enriched in cell adhesion and response to endogenous stimulus. For group CC, up regulated genes enriched in intrinsic component of plasma membrane and mitochondrion, while down regulated genes enriched in integral component of plasma membrane and supra molecular fiber. In addition, GO results of group MF showed that up regulated genes enriched in transferase activity, transferring phosphorus-containing groups and transporter activity and down regulated genes enriched in signaling receptor binding and molecular transducer activity. Several significant enriched pathways were acquired through REACTOME pathway analysis (Table 4). The enriched pathways for up regulated genes included integration of energy metabolism and neuronal system, while, down regulated genes enriched in extracellular matrix organization and GPCR ligand binding.
PPI network construction and module analysis
PPI network complex consisted of 3648 nodes and 6305 edges, wherein node and edge represented gene and interaction between genes (Fig.3a). Moreover, CEBPD, TP73, ESR2, TAB1, MAP 3K5, FN1, UBD, RUNX1, PIK3R2 and TNF were identified as hub genes and are listed in Table 5. In addition, module analysis was conducted to detect the highly connected regions of PPI network, and two significant modules were identified (Fig.3b and Fig.3c). Further GO and pathway enrichment analysis revealed that genes in these modules were mostly implicated in regulation of ion transmembrane transport, oxoacid metabolic process, intrinsic component of plasma membrane, extracellular matrix organization and supra molecular fiber.
PPI network and the most significant modules of DEGs. a The PPI network of DEGs was constructed using Cytoscape b The most significant module was obtained from PPI network with 4 nodes and 6 edges for up regulated genes c The most significant module was obtained from PPI network with 6 nodes and 10 edges for down regulated genes. Up regulated genes are marked in green; down regulated genes are marked in red
Target gene – miRNA regulatory network construction and analysis
The target genes - miRNA regulatory network was constructed, including 1982 miRNAs and 245 target genes. As shown in the integrated target genes - miRNA regulatory network (Fig.4), FASN targeted 147 miRNAs (ex, hsa-mir-4314), SREBF1 targeted 81 miRNAs (ex, hsa-mir-5688), CKB targeted 72 miRNAs (ex, hsa-mir-583), CACNA1A targeted 69 miRNAs (ex, hsa-mir-632), ESR2 targeted 61 miRNAs (ex, hsa-mir-3176), MAP 1B targeted 249 miRNAs (ex, hsa-mir-1299), RUNX1 targeted 125 miRNAs (ex, hsa-mir-4530), PRNP targeted 106 miRNAs (ex, hsa-mir-4477a), FN1 targeted 105 miRNAs (ex, hsa-mir-606) and DAB2 targeted 75 miRNAs (ex, hsa-mir-1343-3p6) and are listed in Table 6.
Target gene-TF s regulatory network construction and analysis
The target genes -TF regulatory network was constructed, including 333 TFs and 204 target genes. As shown in the integrated target genes -TF regulatory network (Fig. 5), SREBF1 targeted 94 TFs (ex, ATF4), FASN targeted 71 TFs (ex, CUX1), SLC9A3R1 targeted 63 TFs (ex, MBD2), CKB targeted 50 TFs (ex, IRF4), TGM2 targeted 50 TFs (ex, SIN3A), PIK3R2 targeted 73 TFs (ex, ZNF143), FLNC targeted 53 TFs (ex, SMARCE1), RUNX1 targeted 53 TFs (ex, ZBTB7A), FN1 targeted 45 TFs (ex, CREB1) and TRIM63 targeted 22 TFs (ex, RELA) and are listed in Table 6.
Receiver operating characteristic (ROC) curve analysis
The ROC curve analysis was used to assess the predictive accuracy of hub genes. AUC was determined and used to prefer the most appropriate cut-off gene expression levels. ROC curves and AUC values are presented in Fig. 6. All AUC values exceeded 0.72, while the up regulated genes CEBPD, TP73, ESR2, TAB1 and MAP 3K5, and down regulated genes FN1, UBD, RUNX1, PIK3R2 and TNF had AUC values > 0.75.
Validation of the expression levels of candidate genes by RT-PCR
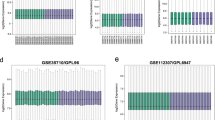
To further verify the expression level of hub genes in obese samples, RT-PCR was performed to calculate the mRNA levels of the ten hub genes identified in the present study (CEBPD, TP73, ESR2, TAB1, MAP 3K5, FN1, UBD, RUNX1, PIK3R2 and TNF) in obese samples. As illustrated in Fig. 7, the expression of CEBPD, TP73, ESR2, TAB1, MAP 3K5 were significantly up regulated in obese samples compared with normal control tissues, while FN1, UBD, RUNX1, PIK3R2 and TNF were significantly down regulated in obese samples compared with normal control tissues. The present RT-PCR results were in line with the prior bioinformatics analysis, suggesting that these hub genes might be associated with progression of obesity associated type 2 diabetes mellitus.
Molecular docking studies
In the current research, the docking simulation was conducted to recognize the active site conformation and major interactions responsible for complex stability with the binding sites receptor. Drug design software Sybyl X 2.1 was used to perform docking experiments on novel molecules containing thiazolidindioneheterocyclic ring. Molecules containing the heterocyclic ring of thiazolidinedione are constructed based on the pioglitazone structure and are most widely used alone or in conjunction with other anti-diabetic drugs. Obesity associated type 2 diabetes mellitus is a chronic disorder that prevents insulin from being used by the body the way it should. It's said that people with obesity associated type 2 diabetes mellitus have insulin resistance, oral hypoglycaemic agents are used either alone or in combination of two or more drugs. Pioglitazone (Glitazones) are commonly used either alone or in combination in obesity associated type 2 diabetes mellitus. The one protein in each over expressed genes in obesity associated type 2 diabetes mellitus are selected for docking studies. The X-RAY crystallographic structure of one protein from each over-expressed genes of CEBPD, TP73, ESR2, TAB1 and MAP 3K5, and their co-crystallized PDB code of 4LY9, 2XWC, 2IOG, 5NZZ and 5UP3 respectively were selected for docking. The examinations of the designed molecules were performed to recognize the potential molecule. The foremost of the designed molecules obtained C-score greater than 6 and are said to be active. A total of 24 designed molecules few molecules have excellent good binding energy (C-score) greater than 8 respectively. Few of the designed molecules obtained good binding scores such as molecule TZP20, TZPS8, TZP22, TZPS10 (Fig.8) obtained binding core of 12.212, 11.489, 11.013 and 10.851 with 5UP3 and molecule TZP22, TZPS8, TZPS10 obtained binding score of 9.482, 9.329 and 9.252 with 2XWC and molecule TZP20, TZPS10 obtained binding score 7.359 and 6.848 with 5NZZ and molecule TZP22, TZP21, TZPS9 obtained binding score 11.053, 10.716 and 10.669 with 2IOG respectively. The molecule TZP23, TZPS5, TZPS2 obtained bind score 4.336 to 4.319 with 5NZZ and molecule TZPS10 of binding core 4.633 with 2IOG respectively. The binding score of the predicted molecules are compared with that of the standard pioglitaone obtained bind score of 10.1314, 9.834, 9.8244, 9.8284 and 7.4321 with 2IOG, 2XWC, 4LY9, 5UP3 and 5NZZ, the values are depicted in Table 7. The molecule TZP22 obtained good binding score with all proteins and hydrogen bonding and other bonding interactions with amino acids with protein code 2IOG are depicted by 3D (Fig.9) and 2D (Fig.10) figures.
Discussion
Obesity associated type 2 diabetes mellitus is the most common aggressive metabolic disorder [37]. However, the most key challenge in treating obesity associated type 2 diabetes mellitus is the presence of complexity [38]. Although previous investigations have reported various potential molecular markers linked with the advancement of obesity associated type 2 diabetes mellitus, the molecular mechanism underlying its pathogenesis has not been generally studied [39]. In the present investigation, a total of 820 DEGs were identified, containing 409 up regulated genes and 411 down regulated genes. SULT1C2 [40] and UBD (ubiquitin D) [41] were responsible for progression of kidney diseases, but these genes might be liable for advancement of obesity associated type 2 diabetes mellitus. HLA-DQA1 was associated with progression of T2DM [42]. SPX (spexin hormone) [43] and APOB (apolipoprotein B) [44] are a critical proteins plays an important role in obesity associated type 2 diabetes mellitus.
The GO and pathway enrichment analysis of DEG are closely related to obesity associated type 2 diabetes mellitus. Genes such as KCNE5 [45], SHANK3 [46], CASQ2 [47], EDNRA (endothelin receptor type A) [48], EPHB4 [49], ALPK3 [50], WNT11 [51], IRAK2 [52], FBN1 [53], SFRP2 [54], CLCA2 [55], NEXN (nexilin F-actin binding protein) [56], PALLD (palladin, cytoskeletal associated protein) [57], DAB2 [58], NRP2 [59], THBS2 [60], CSF1R [61], KCNA2 [62], CACNA1C [63], F2R [64], UCHL1 [65], CCL18 [66], ITGB1BP2 [67] and FMOD (fibromodulin) [68] were reportedly involved in cardio vascular diseases, but these genes might be key for progression of obesity associated type 2 diabetes mellitus. Hu et al. [69], Liu et al. [70], Eltokhi et al. [71], Cai et al. [72], Pfeiffer et al. [73], Lin et al. [74], Royer-Zemmour et al. [75], Pastor et al. [76], Goodspeed et al. [77], Zhang et al. [78], Rogers et al. [79], Su et al. [80] and Foale et al. [81] reported that NRXN1, CRHR1, SHANK2, PSEN2, CKB (creatine kinase B), CD200R1, SRPX2, PTPRZ1, SLC6A1, GABRB2, KCNA1, ASAH1 and LINGO1 were the genes expressed in progression of neuropsychiatric disorders, but these genes might be involved in advancement of obesity associated type 2 diabetes mellitus. Reports indicate that genes include SPHK2 [82], NPC1L1 [83], CNTFR (ciliaryneurotrophic factor receptor) [84], SLC2A4 [85], EDA (ectodysplasin A) [86], TGM2 [87], GCK (glucokinase) [88], FASN (fatty acid synthase) [89], FAP (fibroblast activation protein alpha) [90], PRNP (prion protein) [91], LYVE1 [92], SERPINE1 [93], TNF (tumor necrosis factor) [94], FASLG (Fas ligand) [95], HGF (hepatocyte growth factor) [96], FNDC5 [97], LBP (lipopolysaccharide binding protein) [98] and LOX (lysyl oxidase) [99] were the genes expressed in obesity associated type 2 diabetes mellitus. Hirai et al [100], Vuori et al [101], Porta et al [102], Nomoto et al [103] and Blindbæk et al [104] demonstrates that VAMP2, CACNB2, SLC19A3, PFKFB3 and MFAP4 were the genes essential for progression of type 1 diabetes, but these genes might be key for advancement of obesity associated type 2 diabetes mellitus. Genes such as CACNA1A [105], ALK (ALK receptor tyrosine kinase) [106], SLC4A4 [107], STOX1 [108], COL3A1 [109], VNN1 [110], SLC4A7 [111], BDKRB2 [112], DRD1 [113] and LPAR1 [114] have reported significantly linked with hypertension, but these genes might be crucial for progression of obesity associated type 2 diabetes mellitus. Genes such as KCNE2 [115], DLL1 [116], ACVR1C [117], RGS3 [118], MLXIPL (MLX interacting protein like) [119], PAG1 [120], SLC2A10 [121] and GRB14 [122] play important role in type 2 diabetes mellitus progression. A recent investigation has indicated that genes such as GPIHBP1 [123], FGFRL1 [124], DAPK2 [125], MAP 3K5 [126], ANKK1 [127], GK (glycerol kinase) [128], SPHK1 [129], GNG3 [130], FSTL3 [131], SLIT2 [132], CCDC80 [133], RND3 [134], PTGER4 [135], RUNX1 [136], ADAM12 [137], OLR1 [138], THBS1 [139], CD28 [140], TRPV4 [141], ATRN (attractin) [142], MRC1 [143], SEMA3C [144], HTR2B [145], NOX4 [146], TACR1 [147], BAMBI [148], PDGFD (platelet derived growth factor D) [149], APLN (apelin) [150], MFAP5 [151] and LUM (lumican) [152] are associated with a development of obesity. A previous investigation found that genes such asDDR1 [153], TAB1 [154], NEK8 [155], SERPINE2 [156], FCGR2B [157], ANGPT2 [158], FN1 [159], SOCS5 [158], SMOC2 [160], CD2 [161] and SCN9A [162] expression were associated with a kidney diseases, but these genes might be responsible for advancement of obesity associated type 2 diabetes mellitus.
In addition, an investigation reported that hub genes serve an essential role in maintaining the entire PPI network and its modules are indispensable. 10 hub genes, including CEBPD, TP73, ESR2, TAB1, MAP 3K5, FN1, UBD, RUNX1, PIK3R2 and TNF, were identified as the key genes responsible for progression of obesity associated type 2 diabetes mellitus. Investigation has demonstrated that CEBPD (CCAAT enhancer binding protein delta) is involved in obesity [163]. An investigation by Domingues-Montanari et al. [164] demonstrated that key gene ESR2 was involved in the progression of cardio vascular disease, but this gene might be responsible for progression of obesity associated type 2 diabetes mellitus. TP73, PIK3R2, SLC9A3R1, KRT5, KRT14 and TFAP2C are novel biomarkers for pathogenesis of obesity associated type 2 diabetes mellitus.
The miRNA-target gene regulatory network and TF-target gene regulatory network highlighted in the current investigation provides new theoretical guidance for further exploring the molecular mechanism of obesity associated type 2 diabetes mellitus and provides a new perspective for understanding the underlying biological processes of this diseases, and miRNA and TF targeted therapy. Eberlé et al [165], Cheng et al [166], Cavallari et al [167], Qi et al [168] and Yan et al [169] indicated that SREBF1, MBD2, IRF4, CREB1 and RELA (Nuclear factor-kB) were the genes responsible for advancement of obesity associated type 2 diabetes mellitus. Matsha et al [170] and Ding et al [171] demonstrated that hsa-mir-1299 and hsa-mir-4530 were the miRNAs liable for progression of type 2 diabetes mellitus. Hall et al [172] and Salazar-Mendiguchía et al [173] reported that FLNC (filamin C) and TRIM63 were the genes involved in progression of cardio vascular disease, but these genes might be essential for development of obesity associated type 2 diabetes mellitus. Xiao et al [174], Stratigopoulos et al [175] and Zhou et al [176] noted that ATF4, CUX1 and ZBTB7A were the genes responsible for advancement of obesity. MAP 1B, hsa-mir-4314, hsa-mir-5688, hsa-mir-583, hsa-mir-632, hsa-mir-3176, hsa-mir-4477a, hsa-mir-606, hsa-mir-1343-3p6, SIN3A, ZNF143 and SMARCE1 are the novel biomarkers for pathogenesis of obesity associated type 2 diabetes mellitus.
However, this investigation had distinct limitations. First, the mechanisms of several hub genes in the pathological process of obesity associated type 2 diabetes mellitus remain unclear, permit further investigation. Moreover, the potency of our small molecule drug screening in diminishing side effects remains to be assessed.
In conclusion, with the integrated bioinformatics analysis for expression profiling by high throughput sequencing in obesity associated type 2 diabetes mellitus, ten hub genes associated with the pathogenesis and prognosis of obesity associated type 2 diabetes, including CEBPD, TP73, ESR2, TAB1, MAP 3K5, FN1, UBD, RUNX1, PIK3R2 and TNF. These hub genes were associated with progression of obesity associated type 2 diabetes mellitus and first five (CEBPD, TP73, ESR2, TAB1 and MAP 3K5) of them might be linked with targeted therapy. These hub genes might be regarded as new diagnostic and prognostic biomarkers for obesity associated type 2 diabetes mellitus. However, further in-depth investigation (in vivo and in vitro experiment) is necessary to elucidate the biological function of these genes in obesity associated type 2 diabetes mellitus.
Availability of data and materials
The datasets supporting the conclusions of this article are available in the GEO (Gene Expression Omnibus) (https://www.ncbi.nlm.nih.gov/geo/) repository. [(GSE143319) (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE143319]
References
Pulgaron ER, Delamater AM. Obesity and type 2 diabetes in children: epidemiology and treatment. Curr Diab Rep. 2014;14(8):508. https://doi.org/10.1007/s11892-014-0508-y.
Taylor R. Type 2 diabetes: etiology and reversibility. Diab Care. 2013;36(4):1047–55. https://doi.org/10.2337/dc12-1805.
Catalán V, Gómez-Ambrosi J, Rodríguez A, Ramírez B, Rotellar F, Valentí V, Silva C, Gil MJ, Salvador J, Frühbeck G. Increased circulating and visceral adipose tissue expression levels of YKL-40 in obesity-associated type 2 diabetes are related to inflammation: impact of conventional weight loss and gastric bypass. J Clin Endocrinol Metab. 2011;96(1):200–9. https://doi.org/10.1210/jc.2010-0994.
Fukushima A, Lopaschuk GD. Acetylation control of cardiac fatty acid β-oxidation and energy metabolism in obesity, diabetes, and heart failure. Biochim Biophys Acta. 2016;1862(12):2211–20. https://doi.org/10.1016/j.bbadis.2016.07.020.
Pugazhenthi S, Qin L, Reddy PH. Common neurodegenerative pathways in obesity, diabetes, and Alzheimer's disease. Biochim Biophys Acta Mol Basis Dis. 2017;1863(5):1037–45. https://doi.org/10.1016/j.bbadis.2016.04.017.
Chagnac A, Zingerman B, Rozen-Zvi B, Herman-Edelstein M. Consequences of Glomerular Hyperfiltration: The Role of Physical Forces in the Pathogenesis of Chronic Kidney Disease in Diabetes and Obesity. Nephron. 2019;143(1):38–42. https://doi.org/10.1159/000499486.
Cheung N, Wong TY. Obesity and eye diseases. Surv Ophthalmol. 2007;52(2):180–95. https://doi.org/10.1016/j.survophthal.2006.12.003.
Temelkova-Kurktschiev T, Stefanov T. Lifestyle and genetics in obesity and type 2 diabetes. Exp Clin Endocrinol Diabetes. 2012;120(1):1–6. https://doi.org/10.1055/s-0031-1285832.
Wen X, Qian C, Zhang Y, Wu R, Lu L, Zhu C, Cheng X, Cui R, You H, Mei F, et al. Key pathway and gene alterations in the gastric mucosa associated with obesity and obesity-related diabetes. J Cell Biochem. 2019;120(4):6763–71. https://doi.org/10.1002/jcb.27976.
Kruse R, Vienberg SG, Vind BF, Andersen B, Højlund K. Effects of insulin and exercise training on FGF21, its receptors and target genes in obesity and type 2 diabetes. Diabetologia. 2017;60(10):2042–51. https://doi.org/10.1007/s00125-017-4373-5.
Davison LJ, Holder A, Catchpole B, O'Callaghan CA. The Canine POMC Gene, Obesity in Labrador Retrievers and Susceptibility to Diabetes Mellitus. J Vet Intern Med. 2017;31(2):343–8. https://doi.org/10.1111/jvim.14636.
Huang X, Liu G, Guo J, Su Z. The PI3K/AKT pathway in obesity and type 2 diabetes. Int J Biol Sci. 2018;14(11):1483–96. https://doi.org/10.7150/ijbs.27173.
Gurzov EN, Stanley WJ, Pappas EG, Thomas HE, Gough DJ. The JAK/STAT pathway in obesity and diabetes. FEBS J. 2016;283(16):3002–15. https://doi.org/10.1111/febs.13709.
Xin Y, Kim J, Okamoto H, Ni M, Wei Y, Adler C, Murphy AJ, Yancopoulos GD, Lin C, Gromada J. RNA Sequencing of Single Human Islet Cells Reveals Type 2 Diabetes Genes. Cell Metab. 2016;24(4):608–15. https://doi.org/10.1016/j.cmet.2016.08.018.
Clough E, Barrett T. The Gene Expression Omnibus Database. Methods Mol Biol. 2016;1418:93–110. https://doi.org/10.1007/978-1-4939-3578-9_5.
Ding X, Iyer R, Novotny C, Metzger D, Zhou HH, Smith GI, Yoshino M, Yoshino J, Klein S, Swaminath G, et al. Inhibition of Grb14, a negative modulator of insulin signaling, improves glucose homeostasis without causing cardiac dysfunction. Sci Rep. 2020;10(1):3417. https://doi.org/10.1038/s41598-020-60290-1.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7):e47. https://doi.org/10.1093/nar/gkv007.
Chen J, Bardes EE, Aronow BJ, Jegga AG. ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res. 2009; 37(Web Server issue):W305-W311. doi:https://doi.org/10.1093/nar/gkp427
Thomas PD. The Gene Ontology and the Meaning of Biological Function. Methods Mol Biol. 2017;1446:15–24. https://doi.org/10.1007/978-1-4939-3743-1_2.
Fabregat A, Jupe S, Matthews L, Sidiropoulos K, Gillespie M, Garapati P, Haw R, Jassal B, Korninger F, May B, et al. The Reactome Pathway Knowledgebase. Nucleic Acids Res. 2018;46(D1):D649–55. https://doi.org/10.1093/nar/gkx1132.
Orchard S, Kerrien S, Abbani S, Aranda B, Bhate J, Bidwell S, Bridge A, Briganti L, Brinkman FS, Cesareni G, et al. Protein interaction data curation: the International Molecular Exchange (IMEx) consortium. Nat Methods. 2012;9(4):345–50. https://doi.org/10.1038/nmeth.1931.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–504. https://doi.org/10.1101/gr.1239303.
Przulj N, Wigle DA, Jurisica I. Functional topology in a network of protein interactions. Bioinformatics. 2004;20(3):340–8. https://doi.org/10.1093/bioinformatics/btg415.
Nguyen TP, Liu WC, Jordán F. Inferring pleiotropy by network analysis: linked diseases in the human PPI network. BMC Syst Biol. 2011; 5:179. Published 2011 Oct 31. doi: https://doi.org/10.1186/1752-0509-5-179
Shi Z, Zhang B. Fast network centrality analysis using GPUs. BMC Bioinformatics. 2011;12:149. https://doi.org/10.1186/1471-2105-12-149.
Fadhal E, Gamieldien J, Mwambene EC. Protein interaction networks as metric spaces: a novel perspective on distribution of hubs. BMC Syst Biol. 2014;8:6. https://doi.org/10.1186/1752-0509-8-6.
Zaki N, Efimov D, Berengueres J. Protein complex detection using interaction reliability assessment and weighted clustering coefficient. BMC Bioinform. 2013;14:163. https://doi.org/10.1186/1471-2105-14.
Fan Y, Xia J. miRNet-Functional Analysis and Visual Exploration of miRNA-Target Interactions in a Network Context. Methods Mol Biol. 2018;1819:215–33. https://doi.org/10.1007/978-1-4939-8618-7_10.
Zhou G, Soufan O, Ewald J, Hancock REW, Basu N, Xia J. NetworkAnalyst 3.0: a visual analytics platform for comprehensive gene expression profiling and meta-analysis. Nucleic Acids Res. 2019;47:W234–41. https://doi.org/10.1093/nar/gkz240.
Robin X, Turck N, Hainard A, Tiberti N, Lisacek F, Sanchez JC, Müller M. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinform. 2011;12:77. https://doi.org/10.1186/1471-2105-12-77.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25(4):402–8. https://doi.org/10.1006/meth.2001.
Naim MJ, Alam O, Alam MJ, Hassan MQ, Siddiqui N, Naidu VGM, Alam MI. Design, synthesis and molecular docking of thiazolidinedione based benzene sulphonamide derivatives containing pyrazole core as potential anti-diabetic agents. Bioorg Chem. 2018;76:98–112. https://doi.org/10.1016/j.bioorg.2017.11.010.
Mohammadi-Khanaposhtani M, Rezaei S, Khalifeh R, Imanparast S, Faramarzi MA, Bahadorikhalili S, Safavi M, Bandarian F, Nasli Esfahani E, Mahdavi M, et al. Design, synthesis, docking study, α-glucosidase inhibition, and cytotoxic activities of acridine linked to thioacetamides as novel agents in treatment of type 2 diabetes. Bioorg Chem. 2018;80:288–95. https://doi.org/10.1016/j.bioorg.2018.06.035.
Rojas LB, Gomes MB. Metformin: an old but still the best treatment for type 2 diabetes. Diabetol Metab Syndr. 2013;5(1):6. https://doi.org/10.1186/1758-5996-5-6.
Thakral S, Narang R, Kumar M, Singh V. Synthesis, molecular docking and molecular dynamic simulation studies of 2-chloro-5-[(4-chlorophenyl) sulfamoyl]-N-(alkyl/aryl)-4-nitrobenzamide derivatives as antidiabetic agents. BMC Chem. 2020;14(1):1–6. https://doi.org/10.1186/s13065-020-00703-4.
Srikanth Kumar K, Lakshmana Rao A, Basaveswara Rao MV. Design, synthesis, biological evaluation and molecular docking studies of novel 3-substituted-5-[(indol-3-yl)methylene]-thiazolidine-2,4-dione derivatives. Heliyon. 2018;4(9):e00807. https://doi.org/10.1016/j.heliyon.2018.e00807.
Mohammad S, Ahmad J. Management of obesity in patients with type 2 diabetes mellitus in primary care. Diab Metab Syndr. 2016;10(3):171–81. https://doi.org/10.1016/j.dsx.2016.01.017.
Rao M, Gao C, Xu L, Jiang L, Zhu J, Chen G, Law BYK, Xu Y. Effect of Inulin-Type Carbohydrates on Insulin Resistance in Patients with Type 2 Diabetes and Obesity: A Systematic Review and Meta-Analysis. J Diab Res. 2019:5101423. https://doi.org/10.1155/2019/5101423.
Al-Sulaiti H, Diboun I, Agha MV, et al. Metabolic signature of obesity-associated insulin resistance and type 2 diabetes. J Transl Med. 2019;17(1):348. https://doi.org/10.1186/s12967-019-2096-8.
Sugimura K, Tanaka T, Tanaka Y, Takano H, Kanagawa K, Sakamoto N, Ikemoto S, Kawashima H, Nakatani T. Decreased sulfotransferase SULT1C2 gene expression in DPT-induced polycystic kidney. Kidney Int. 2002;62(3):757–62. https://doi.org/10.1046/j.1523-1755.2002.00512.x.
Zhang JY, Wang M, Tian L, Genovese G, Yan P, Wilson JG, Thadhani R, Mottl AK, Appel GB, Bick AG, et al. UBD modifies APOL1-induced kidney disease risk. Proc Natl Acad Sci U S A. 2018;115(13):3446–51. https://doi.org/10.1073/pnas.1716113115.
Ma ZJ, Sun P, Guo G, Zhang R, Chen LM. Association of the HLA-DQA1 and HLA-DQB1 Alleles in Type 2 Diabetes Mellitus and Diabetic Nephropathy in the Han Ethnicity of China. J Diab Res. 2013;2013:452537. https://doi.org/10.1155/2013/452537.
Kołodziejski PA, Pruszyńska-Oszmałek E, Korek E, Sassek M, Szczepankiewicz D, Kaczmarek P, Nogowski L, Maćkowiak P, Nowak KW, Krauss H, et al. Serum levels of spexin and kisspeptin negatively correlate with obesity and insulin resistance in women. Physiol Res. 2018;67(1):45–56. https://doi.org/10.33549/physiolres.933467.
Onat A, Can G, Hergenç G, Yazici M, Karabulut A, Albayrak S. Serum apolipoprotein B predicts dyslipidemia, metabolic syndrome and, in women, hypertension and diabetes, independent of markers of central obesity and inflammation. Int J Obes (Lond). 2007;31(7):1119–25. https://doi.org/10.1038/sj.ijo.0803552.
Ravn LS, Hofman-Bang J, Dixen U, Larsen SO, Jensen G, Haunsø S, Svendsen JH, Christiansen M. Relation of 97T polymorphism in KCNE5 to risk of atrial fibrillation. Am J Cardiol. 2005;96(3):405–7. https://doi.org/10.1016/j.amjcard.2005.03.086.
Man W, Gu J, Wang B, Zhang M, Hu J, Lin J, Sun D, Xiong Z, Gu X, Hao K, et al. SHANK3 Co-ordinately Regulates Autophagy and Apoptosis in Myocardial Infarction. Front Physiol. 2020;11:1082. https://doi.org/10.3389/fphys.2020.01082.
Refaat MM, Aouizerat BE, Pullinger CR, Malloy M, Kane J, Tseng ZH. Association of CASQ2 polymorphisms with sudden cardiac arrest and heart failure in patients with coronary artery disease. Heart Rhythm. 2014;11(4):646–52. https://doi.org/10.1016/j.hrthm.2014.01.015.
Pritchard AB, Kanai SM, Krock B, Schindewolf E, Oliver-Krasinski J, Khalek N, Okashah N, Lambert NA, Tavares ALP, Zackai E, et al. Loss-of-function of Endothelin receptor type A results in Oro-Oto-Cardiac syndrome. Am J Med Genet A. 2020;182(5):1104–16. https://doi.org/10.1002/ajmg.a.61531.
Yang D, Jin C, Ma H, Huang M, Shi GP, Wang J, Xiang M. EphrinB2/EphB4 pathway in postnatal angiogenesis: a potential therapeutic target for ischemic cardiovascular disease. Angiogenesis. 2016;19(3):297–309. https://doi.org/10.1007/s10456-016-9514-9.
Jorholt J, Formicheva Y, Vershinina T, Kiselev A, Muravyev A, Demchenko E, Fedotov P, Zlotina A, Rygkov A, Vasichkina E, et al. Cardiomyopathy and Skeletal Muscle Features Associated with ALPK3 Homozygous and Compound Heterozygous Variants. Genes (Basel). 2020;11(10):1201. https://doi.org/10.3390/genes11101201.
Touma M, Kang X, Gao F, Zhao Y, Cass AA, Biniwale R, Xiao X, Eghbali M, Coppola G, Reemtsen B, et al. Wnt11 regulates cardiac chamber development and disease during perinatal maturation. JCI Insight. 2017;2(17):e94904. https://doi.org/10.1172/jci.insight.94904.
Liu Z, Zhao N, Zhu H, Zhu S, Pan S, Xu J, Zhang X, Zhang Y, Wang J. Circulating interleukin-1β promotes endoplasmic reticulum stress-induced myocytes apoptosis in diabetic cardiomyopathy via interleukin-1 receptor-associated kinase-2. Cardiovasc Diabetol. 2015;14:125. https://doi.org/10.1186/s12933-015-0288-y.
Barrett PM, Topol EJ. The fibrillin-1 gene: unlocking new therapeutic pathways in cardiovascular disease. Heart. 2013;99(2):83–90. https://doi.org/10.1136/heartjnl-2012-301840.
Wu Y, Liu X, Zheng H, Zhu H, Mai W, Huang X, Huang Y. Multiple Roles of sFRP2 in Cardiac Development and Cardiovascular Disease. Int J Biol Sci. 2020;16(5):730–8. https://doi.org/10.7150/ijbs.40923.
Mao Z, Wang Y, Peng H, He F, Zhu L, Huang H, Huang X, Lu X, Tan X. A newly identified missense mutation in CLCA2 is associated with autosomal dominant cardiac conduction block. Gene. 2019;714:143990. https://doi.org/10.1016/j.gene.2019.143990.
Wu C, Yan H, Sun J, Yang F, Song C, Jiang F, Li Y, Dong J, Zheng GY, Tian XL, et al. NEXN is a novel susceptibility gene for coronary artery disease in Han Chinese. PLoS One. 2013;8(12):e82135. https://doi.org/10.1371/journal.pone.0082135.
Hoke M, Schillinger M, Dick P, Exner M, Koppensteiner R, Minar E, Mlekusch W, Schlager O, Wagner O, Mannhalter C. Polymorphism of the palladin gene and cardiovascular outcome in patients with atherosclerosis. Eur J Clin Invest. 2011;41(4):365–71. https://doi.org/10.1111/j.1365-2362.2010.02416.x.
Wang Y, Wang Y, Adi D, He X, Liu F, Abudesimu A, Fu Z, Ma Y. Dab2 gene variant is associated with increased coronary artery disease risk in Chinese Han population. Medicine (Baltimore). 2020;99(27):e20924. https://doi.org/10.1097/MD.0000000000020924.
Harman JL, Sayers J, Chapman C, Pellet-Many C. Emerging Roles for Neuropilin-2 in Cardiovascular Disease. Int J Mol Sci. 2020;21(14):5154. https://doi.org/10.3390/ijms21145154.
Wang Y, Fu W, Xie F, Wang Y, Chu X, Wang H, Shen M, Wang Y, Wang Y, Sun WL, et al. Common polymorphisms in ITGA2, PON1 and THBS2 are associated with coronary atherosclerosis in a candidate gene association study of the Chinese Han population. J Hum Genet. 2010;55(8):490–4. https://doi.org/10.1038/jhg.2010.53.
Wei Y, Zhu M, Corbalán-Campos J, Heyll K, Weber C, Schober A. Regulation of Csf1r and Bcl6 in macrophages mediates the stage-specific effects of microRNA-155 on atherosclerosis. Arterioscler Thromb Vasc Biol. 2015;35(4):796–803. https://doi.org/10.1161/ATVBAHA.114.304723.
Long QQ, Wang H, Gao W, Fan Y, Li YF, Ma Y, Yang Y, Shi HJ, Chen BR, Meng HY, et al. Long Noncoding RNA Kcna2 Antisense RNA Contributes to Ventricular Arrhythmias via Silencing Kcna2 in Rats With Congestive Heart Failure. J Am Heart Assoc. 2017;6(12):e005965. https://doi.org/10.1161/JAHA.117.005965.
Beitelshees AL, Navare H, Wang D, Gong Y, Wessel J, Moss JI, Langaee TY, Cooper-DeHoff RM, Sadee W, Pepine CJ, et al. CACNA1C gene polymorphisms, cardiovascular disease outcomes, and treatment response. Circ Cardiovasc Genet. 2009;2(4):362–70. https://doi.org/10.1161/CIRCGENETICS.109.857839.
Gigante B, Vikström M, Meuzelaar LS, Chernogubova E, Silveira A, Hooft FV, Hamsten A, de Faire U. Variants in the coagulation factor 2 receptor (F2R) gene influence the risk of myocardial infarction in men through an interaction with interleukin 6 serum levels. Thromb Haemost. 2009;101(5):943–53.
Lei Q, Yi T, Li H, Yan Z, Lv Z, Li G, Wang Y. Ubiquitin C-terminal hydrolase L1 (UCHL1) regulates post-myocardial infarction cardiac fibrosis through glucose-regulated protein of 78 kDa (GRP78). Sci Rep. 2020;10(1):10604. https://doi.org/10.1038/s41598-020-67746-4.
de Jager SC, Bongaerts BW, Weber M, Kraaijeveld AO, Rousch M, Dimmeler S, van Dieijen-Visser MP, Cleutjens KB, Nelemans PJ, van Berkel TJ, et al. Chemokines CCL3/MIP1α, CCL5/RANTES and CCL18/PARC are independent risk predictors of short-term mortality in patients with acute coronary syndromes. PLoS One. 2012;7(9):e45804. https://doi.org/10.1371/journal.pone.0045804.
Ruppert V, Meyer T, Richter A, Maisch B, Pankuweit S. German Competence Network of Heart Failure. Identification of a missense mutation in the melusin-encoding ITGB1BP2 gene in a patient with dilated cardiomyopathy. Gene. 2013;512(2):206–10. https://doi.org/10.1016/j.gene.2012.10.055.
Andenæs K, Lunde IG, Mohammadzadeh N, Dahl CP, Aronsen JM, Strand ME, Palmero S, Sjaastad I, Christensen G, Engebretsen KVT, et al. The extracellular matrix proteoglycan fibromodulin is upregulated in clinical and experimental heart failure and affects cardiac remodeling. PLoS One. 2018;13(7):e0201422. https://doi.org/10.1371/journal.pone.0201422.
Hu Z, Xiao X, Zhang Z, Li M. Genetic insights and neurobiological implications from NRXN1 in neuropsychiatric disorders. Mol Psychiatr. 2019;24(10):1400–14. https://doi.org/10.1038/s41380-019-0438-9.
Liu Z, Liu W, Yao L, Yang C, Xiao L, Wan Q, Gao K, Wang H, Zhu F, Wang G, et al. Negative life events and corticotropin-releasing-hormone receptor1 gene in recurrent major depressive disorder. Sci Rep. 2013;3:1548. https://doi.org/10.1038/srep01548.
Eltokhi A, Rappold G, Sprengel R. Distinct Phenotypes of Shank2 Mouse Models Reflect Neuropsychiatric Spectrum Disorders of Human Patients With SHANK2 Variants. Front Mol Neurosci. 2018;11:240. https://doi.org/10.3389/fnmol.2018.00240.
Cai Y, An SS, Kim S. Mutations in presenilin 2 and its implications in Alzheimer's disease and other dementia-associated disorders. Clin Interv Aging. 2015;10:1163–72. https://doi.org/10.2147/CIA.S85808.
Pfeiffer FE, Homburger HA, Houser OW, Baker HL Jr, Yanagihara T. Elevation of serum creatine kinase B-subunit levels by radiographic contrast agents in patients with neurologic disorders. Mayo Clin Proc. 1987;62(5):351–7. https://doi.org/10.1016/s0025-6196(12)65438-x.
Lin S, He L, Shen R, Fang F, Pan H, Zhu X, Wang M, Zhou Z, Liu Z, Wang X, et al. Identification of the CD200R1 promoter and the association of its polymorphisms with the risk of Parkinson's disease. Eur J Neurol. 2020;27(7):1224–30. https://doi.org/10.1111/ene.14224.
Royer-Zemmour B, Ponsole-Lenfant M, Gara H, Roll P, Lévêque C, Massacrier A, Ferracci G, Cillario J, Robaglia-Schlupp A, Vincentelli R, et al. Epileptic and developmental disorders of the speech cortex: ligand/receptor interaction of wild-type and mutant SRPX2 with the plasminogen activator receptor uPAR. Hum Mol Genet. 2008;17(23):3617–30. https://doi.org/10.1093/hmg/ddn256.
Pastor M, Fernández-Calle R, Di Geronimo B, Vicente-Rodríguez M, Zapico JM, Gramage E, Coderch C, Pérez-García C, Lasek AW, Puchades-Carrasco L, et al. Development of inhibitors of receptor protein tyrosine phosphatase β/ζ (PTPRZ1) as candidates for CNS disorders. Eur J Med Chem. 2018;144:318–29. https://doi.org/10.1016/j.ejmech.2017.11.080.
Goodspeed K, Pérez-Palma E, Iqbal S, Cooper D, Scimemi A, Johannesen KM, Stefanski A, Demarest S, Helbig KL, Kang J, et al. Current knowledge of SLC6A1-related neurodevelopmental disorders. Brain Commun. 2020;2:fcaa170. https://doi.org/10.1093/braincomms/fcaa170.
Zhang T, Li J, Yu H, Shi Y, Li Z, Wang L, Wang Z, Lu T, Wang L, Yue W, et al. Meta-analysis of GABRB2 polymorphisms and the risk of schizophrenia combined with GWAS data of the Han Chinese population and psychiatric genomics consortium. PLoS One. 2018;13(6):e0198690. https://doi.org/10.1371/journal.pone.0198690.
Rogers A, Golumbek P, Cellini E, Doccini V, Guerrini R, Wallgren-Pettersson C, Thuresson AC, Gurnett CA. De novo KCNA1 variants in the PVP motif cause infantile epileptic encephalopathy and cognitive impairment similar to recurrent KCNA2 variants. Am J Med Genet A. 2018;176(8):1748–52. https://doi.org/10.1002/ajmg.a.38840.
Su Y, Yang L, Li Z, Wang W, Xing M, Fang Y, Cheng Y, Lin GN, Cui D. The interaction of ASAH1 and NGF gene involving in neurotrophin signaling pathway contributes to schizophrenia susceptibility and psychopathology. Prog Neuropsychopharmacol Biol Psychiatry. 2021;104:110015. https://doi.org/10.1016/j.pnpbp.2020.110015.
Foale S, Berry M, Logan A, Fulton D, Ahmed Z. LINGO-1 and AMIGO3, potential therapeutic targets for neurological and dysmyelinating disorders? Neural Regen Res. 2017;12(8):1247–51. https://doi.org/10.4103/1673-5374.213538.
Ravichandran S, Finlin BS, Kern PA, Özcan S. Sphk2-/- mice are protected from obesity and insulin resistance. Biochim Biophys Acta Mol Basis Dis. 2019;1865(3):570–6. https://doi.org/10.1016/j.bbadis.2018.12.012.
Labonté ED, Camarota LM, Rojas JC, Jandacek RJ, Gilham DE, Davies JP, Ioannou YA, Tso P, Hui DY, Howles PN. Reduced absorption of saturated fatty acids and resistance to diet-induced obesity and diabetes by ezetimibe-treated and Npc1l1-/- mice. Am J Physiol Gastrointest Liver Physiol. 2008;295(4):G776–83. https://doi.org/10.1152/ajpgi.90275.2008.
Watt MJ, Dzamko N, Thomas WG, Rose-John S, Ernst M, Carling D, Kemp BE, Febbraio MA, Steinberg GR. CNTF reverses obesity-induced insulin resistance by activating skeletal muscle AMPK. Nat Med. 2006;12(5):541–8. https://doi.org/10.1038/nm1383.
Yang Q, Graham TE, Mody N, Preitner F, Peroni OD, Zabolotny JM, Kotani K, Quadro L, Kahn BB. Serum retinol binding protein 4 contributes to insulin resistance in obesity and type 2 diabetes. Nature. 2005;436(7049):356–62. https://doi.org/10.1038/nature03711.
Awazawa M, Gabel P, Tsaousidou E, Nolte H, Krüger M, Schmitz J, Ackermann PJ, Brandt C, Altmüller J, Motameny S, et al. A microRNA screen reveals that elevated hepatic ectodysplasin A expression contributes to obesity-induced insulin resistance in skeletal muscle. Nat Med. 2017;23(12):1466–73. https://doi.org/10.1038/nm.4420.
Ludvigsen TP, Olsen LH, Pedersen HD, Christoffersen BØ, Jensen LJ. Hyperglycemia-induced transcriptional regulation of ROCK1 and TGM2 expression is involved in small artery remodeling in obese diabetic Göttingen Minipigs. Clin Sci (Lond). 2019;133(24):2499–516. https://doi.org/10.1042/CS20191066.
Cockburn BN, Ostrega DM, Sturis J, Kubstrup C, Polonsky KS, Bell GI. Changes in pancreatic islet glucokinase and hexokinase activities with increasing age, obesity, and the onset of diabetes. Diabetes. 1997;46(9):1434–9. https://doi.org/10.2337/diab.46.9.1434.
Berndt J, Kovacs P, Ruschke K, Klöting N, Fasshauer M, Schön MR, Körner A, Stumvoll M, Blüher M. Fatty acid synthase gene expression in human adipose tissue: association with obesity and type 2 diabetes. Diabetologia. 2007;50(7):1472–80. https://doi.org/10.1007/s00125-007-0689-x.
Williams KH, Viera de Ribeiro AJ, Prakoso E, Veillard AS, Shackel NA, Bu Y, Brooks B, Cavanagh E, Raleigh J, et al. Lower serum fibroblast activation protein shows promise in the exclusion of clinically significant liver fibrosis due to non-alcoholic fatty liver disease in diabetes and obesity. Diab Res Clin Pract. 2015;108(3):466–72. https://doi.org/10.1016/j.diabres.2015.02.024.
de Brito G, Lupinacci FC, Beraldo FH, Santos TG, Roffé M, Lopes MH, de Lima VC, Martins VR, Hajj GN. Loss of prion protein is associated with the development of insulin resistance and obesity. Biochem J. 2017;474(17):2981–91. https://doi.org/10.1042/BCJ20170137.
Michurina SV, Ishchenko IY, Arkhipov SA, Klimontov VV, Rachkovskaya LN, Konenkov VI, Zavyalov EL. Effects of Melatonin, Aluminum Oxide, and Polymethylsiloxane Complex on the Expression of LYVE-1 in the Liver of Mice with Obesity and Type 2 Diabetes Mellitus. Bull Exp Biol Med. 2016;162(2):269–72. https://doi.org/10.1007/s10517-016-3592-y.
Kaur P, Reis MD, Couchman GR, Forjuoh SN, Greene JF, Asea A. SERPINE 1 Links Obesity and Diabetes: A Pilot Study. J Proteomics Bioinform. 2010;3(6):191–9. https://doi.org/10.4172/jpb.1000139.
Serino M, Menghini R, Fiorentino L, Amoruso R, Mauriello A, Lauro D, Sbraccia P, Hribal ML, Lauro R, Federici M. Mice heterozygous for tumor necrosis factor-alpha converting enzyme are protected from obesity-induced insulin resistance and diabetes. Diabetes. 2007;56(10):2541–6. https://doi.org/10.2337/db07-0360.
Blüher M, Klöting N, Wueest S, Schoenle EJ, Schön MR, Dietrich A, Fasshauer M, Stumvoll M, Konrad D. Fas and FasL expression in human adipose tissue is related to obesity, insulin resistance, and type 2 diabetes. J Clin Endocrinol Metab. 2014;99(1):E36–44. https://doi.org/10.1210/jc.2013-2488.
Muratsu J, Iwabayashi M, Sanada F, Taniyama Y, Otsu R, Rakugi H, Morishita R. Hepatocyte Growth Factor Prevented High-Fat Diet-Induced Obesity and Improved Insulin Resistance in Mice. Sci Rep. 2017;7(1):130. https://doi.org/10.1038/s41598-017-00199-4.
Xiong XQ, Geng Z, Zhou B, Zhang F, Han Y, Zhou YB, Wang JJ, Gao XY, Chen Q, Li YH, et al. FNDC5 attenuates adipose tissue inflammation and insulin resistance via AMPK-mediated macrophage polarization in obesity. Metabolism. 2018;83:31–41. https://doi.org/10.1016/j.metabol.2018.01.013.
Kim KE, Cho YS, Baek KS, Li L, Baek KH, Kim JH, Kim HS, Sheen YH. Lipopolysaccharide-binding protein plasma levels as a biomarker of obesity-related insulin resistance in adolescents. Korean J Pediatr. 2016;59(5):231–8. https://doi.org/10.3345/kjp.2016.59.5.231.
Daley EJ, Pajevic PD, Roy S, Trackman PC. Impaired Gastric Hormone Regulation of Osteoblasts and Lysyl Oxidase Drives Bone Disease in Diabetes Mellitus. JBMR Plus. 2019;3(10):e10212. https://doi.org/10.1002/jbm4.10212.
Hirai H, Miura J, Hu Y, Larsson H, Larsson K, Lernmark A, Ivarsson SA, Wu T, Kingman A, Tzioufas AG, et al. Selective screening of secretory vesicle-associated proteins for autoantigens in type 1 diabetes: VAMP2 and NPY are new minor autoantigens. Clin Immunol. 2008;127(3):366–74. https://doi.org/10.1016/j.clim.2008.01.018.
Vuori N, Sandholm N, Kumar A, Hietala K, Syreeni A, Forsblom C, Juuti-Uusitalo K, Skottman H, Imamura M, Maeda S, et al. CACNB2 Is a Novel Susceptibility Gene for Diabetic Retinopathy in Type 1 Diabetes. Diabetes. 2019;68(11):2165–74. https://doi.org/10.2337/db19-0130.
Porta M, Toppila I, Sandholm N, Hosseini SM, Forsblom C, Hietala K, Borio L, Harjutsalo V, Klein BE, Klein R, et al. Variation in SLC19A3 and Protection From Microvascular Damage in Type 1 Diabetes. Diabetes. 2016;65(4):1022–30. https://doi.org/10.2337/db15-1247.
Nomoto H, Pei L, Montemurro C, Rosenberger M, Furterer A, Coppola G, Nadel B, Pellegrini M, Gurlo T, Butler PC, et al. Activation of the HIF1α/PFKFB3 stress response pathway in beta cells in type 1 diabetes. Diabetologia. 2020;63(1):149–61. https://doi.org/10.1007/s00125-019-05030-5.
Blindbæk SL, Schlosser A, Green A, Holmskov U, Sorensen GL, Grauslund J. Association between microfibrillar-associated protein 4 (MFAP4) and micro- and macrovascular complications in long-term type 1 diabetes mellitus. Acta Diabetol. 2017;54(4):367–72. https://doi.org/10.1007/s00592-016-0953-y.
Hu Z, Liu F, Li M, He J, Huang J, Rao DC, Hixson JE, Gu C, Kelly TN, Chen S, et al. Associations of Variants in the CACNA1A and CACNA1C Genes With Longitudinal Blood Pressure Changes and Hypertension Incidence: The GenSalt Study. Am J Hypertens. 2016;29(11):1301–6. https://doi.org/10.1093/ajh/hpw070.
Tabbò F, D'Aveni A, Tota D, Pignataro D, Bironzo P, Carnio S, Cappia S, Cortese G, Righi L, Novello S. Pulmonary Arterial Hypertension in ALK Receptor Tyrosine Kinase-Positive Lung Cancer Patient: Adverse Event or Disease Spread? J Thorac Oncol. 2019;14(2):e38–40. https://doi.org/10.1016/j.jtho.2018.10.154.
Yang HC, Liang YJ, Chen JW, Chiang KM, Chung CM, Ho HY, Ting CT, Lin TH, Sheu SH, Tsai WC, et al. Identification of IGF1, SLC4A4, WWOX, and SFMBT1 as hypertension susceptibility genes in Han Chinese with a genome-wide gene-based association study. PLoS One. 2012;7(3):e32907. https://doi.org/10.1371/journal.pone.0032907.
Erlandsson L, Ducat A, Castille J, Zia I, Kalapotharakos G, Hedström E, Vilotte JL, Vaiman D, Hansson SR. lpha-1 microglobulin as a potential therapeutic candidate for treatment of hypertension and oxidative stress in the STOX1 preeclampsia mouse model. Sci Rep. 2019;9(1):8561. https://doi.org/10.1038/s41598-019-44639-9.
Samokhin AO, Stephens T, Wertheim BM, Wang RS, Vargas SO, Yung LM, Cao M, Brown M, Arons E, Dieffenbach PB, et al. NEDD9 targets COL3A1 to promote endothelial fibrosis and pulmonary arterial hypertension. Sci Transl Med. 2018;10(445):eaap7294. https://doi.org/10.1126/scitranslmed.aap7294.
Fava C, Montagnana M, Danese E, Sjögren M, Almgren P, Engström G, Hedblad B, Guidi GC, Minuz P, Melander O. Vanin-1 T26I polymorphism, hypertension and cardiovascular events in two large urban-based prospective studies in Swedes. Nutr Metab Cardiovasc Dis. 2013;23(1):53–60. https://doi.org/10.1016/j.numecd.2011.01.012.
Wang L, Li H, Yang B, Guo L, Han X, Li L, Li M, Huang J, Gu D. The Hypertension Risk Variant Rs820430 Functions as an Enhancer of SLC4A7. Am J Hypertens. 2017;30(2):202–8. https://doi.org/10.1093/ajh/hpw127.
Bhupatiraju C, Patkar S, Pandharpurkar D, Joshi S, Tirunilai P. Association and interaction of -58C>T and ±9 bp polymorphisms of BDKRB2 gene causing susceptibility to essential hypertension. Clin Exp Hypertens. 2012;34(3):230–5. https://doi.org/10.3109/10641963.2011.631653.
Zhang H, Sun ZQ, Liu SS, Yang LN. Association between GRK4 and DRD1 gene polymorphisms and hypertension: a meta-analysis. Clin Interv Aging. 2015;11:17–27. https://doi.org/10.2147/CIA.S94510.
Xu K, Ma L, Li Y, Wang F, Zheng GY, Sun Z, Jiang F, Chen Y, Liu H, Dang A, et al. Genetic and Functional Evidence Supports LPAR1 as a Susceptibility Gene for Hypertension. Hypertension. 2015;66(3):641–6. https://doi.org/10.1161/HYPERTENSIONAHA.115.05515.
Lee SM, Baik J, Nguyen D, Nguyen V, Liu S, Hu Z, Abbott GW. Kcne2 deletion impairs insulin secretion and causes type 2 diabetes mellitus. FASEB J. 2017;31(6):2674–85. https://doi.org/10.1096/fj.201601347.
Deng Z, Shen J, Ye J, Shu Q, Zhao J, Fang M, Zhang T. Association between single nucleotide polymorphisms of delta/notch-like epidermal growth factor (EGF)-related receptor (DNER) and Delta-like 1 Ligand (DLL 1) with the risk of type 2 diabetes mellitus in a Chinese Han population. Cell Biochem Biophys. 2015;71(1):331–5. https://doi.org/10.1007/s12013-014-0202-3.
Emdin CA, Khera AV, Aragam K, Haas M, Chaffin M, Klarin D, Natarajan P, Bick A, Zekavat SM, Nomura A, et al. DNA Sequence Variation in ACVR1C Encoding the Activin Receptor-Like Kinase 7 Influences Body Fat Distribution and Protects Against Type 2 Diabetes. Diabetes. 2019;68(1):226–34. https://doi.org/10.2337/db18-0857.
Kim KS, Jung Yang H, Lee IS, Kim KH, Park J, Jeong HS, Kim Y, Seok Ahn K, et al. The aglycone of ginsenoside Rg3 enables glucagon-like peptide-1 secretion in enteroendocrine cells and alleviates hyperglycemia in type 2 diabetic mice. Sci Rep. 2015;5:18325. https://doi.org/10.1038/srep18325.
Mtiraoui N, Turki A, Nemr R, Echtay A, Izzidi I, Al-Zaben GS, Irani-Hakime N, Keleshian SH, Mahjoub T, Almawi WY. Contribution of common variants of ENPP1, IGF2BP2, KCNJ11, MLXIPL, PPARγ, SLC30A8 and TCF7L2 to the risk of type 2 diabetes in Lebanese and Tunisian Arabs. Diabetes Metab. 2012;38(5):444–9. https://doi.org/10.1016/j.diabet.2012.05.002.
Ishii H, Niiya T, Ono Y, Inaba N, Jinnouchi H, Watada H. Improvement of quality of life through glycemic control by liraglutide, a GLP-1 analog, in insulin-naive patients with type 2 diabetes mellitus: the PAGE1 study. Diabetol Metab Syndr. 2017;9:3. https://doi.org/10.1186/s13098-016-0202-0.
Jiang YD, Chang YC, Chiu YF, Chang TJ, Li HY, Lin WH, Yuan HY, Chen YT, Chuang LM. SLC2A10 genetic polymorphism predicts development of peripheral arterial disease in patients with type 2 diabetes. SLC2A10 and PAD in type 2 diabetes. BMC Med Genet. 2010;11:126. https://doi.org/10.1186/1471-2350-11-126.
Harder MN, Ribel-Madsen R, Justesen JM, Sparsø T, Andersson EA, Grarup N, Jørgensen T, Linneberg A, Hansen T, Pedersen O. Type 2 diabetes risk alleles near BCAR1 and in ANK1 associate with decreased β-cell function whereas risk alleles near ANKRD55 and GRB14 associate with decreased insulin sensitivity in the Danish Inter99 cohort. J Clin Endocrinol Metab. 2013;98(4):E801–6. https://doi.org/10.1210/jc.2012-4169.
Aruga M, Tokita Y, Nakajima K, Kamachi K, Tanaka A. The effect of combined diet and exercise intervention on body weight and the serum GPIHBP1 concentration in overweight/obese middle-aged women. Clin Chim Acta. 2017;475:109–15. https://doi.org/10.1016/j.cca.2017.10.017.
Baruch A, Wong C, Chinn LW, Vaze A, Sonoda J, Gelzleichter T, Chen S, Lewin-Koh N, Morrow L, Dheerendra S, et al. Antibody-mediated activation of the FGFR1/Klothoβ complex corrects metabolic dysfunction and alters food preference in obese humans. Proc Natl Acad Sci U S A. 2020;117(46):28992–9000. https://doi.org/10.1073/pnas.2012073117.
Soussi H, Reggio S, Alili R, Prado C, Mutel S, Pini M, Rouault C, Clément K, Dugail I. DAPK2 Downregulation Associates With Attenuated Adipocyte Autophagic Clearance in Human Obesity. Diabetes. 2015;64(10):3452–63. https://doi.org/10.2337/db14-1933.
Haim Y, Blüher M, Konrad D, Goldstein N, Klöting N, Harman-Boehm I, Kirshtein B, Ginsberg D, Tarnovscki T, Gepner Y, et al. ASK1 (MAP 3K5) is transcriptionally upregulated by E2F1 in adipose tissue in obesity, molecularly defining a human dys-metabolic obese phenotype. Mol Metab. 2017;6(7):725–36. https://doi.org/10.1016/j.molmet.2017.05.003.
Aliasghari F, Nazm SA, Yasari S, Mahdavi R, Bonyadi M. Associations of the ANKK1 and DRD2 gene polymorphisms with overweight, obesity and hedonic hunger among women from the Northwest of Iran [published online ahead of print, 2020 Feb 4]. Eat Weight Disord. 2020;10.1007/s40519-020-00851-5. doi:https://doi.org/10.1007/s40519-020-00851-5
Koschinsky T, Gries FA, Herberg L. Regulation of glycerol kinase by insulin in isolated fat cells and liver of Bar Harbor obese mice. Diabetologia. 1971;7(5):316–22. https://doi.org/10.1007/BF01219464.
Xie J, Shao Y, Liu J, Cui M, Xiao X, Gong J, Xue B, Zhang Q, Hu X, Duan H. K27Q/K29Q mutations in sphingosine kinase 1 attenuate high-fat diet induced obesity and altered glucose homeostasis in mice. Sci Rep. 2020;10(1):20038. https://doi.org/10.1038/s41598-020-77096-w.
Schwindinger WF, Borrell BM, Waldman LC, Robishaw JD. Mice lacking the G protein gamma3-subunit show resistance to opioids and diet induced obesity. Am J Physiol Regul Integr Comp Physiol. 2009;297(5):R1494–502. https://doi.org/10.1152/ajpregu.00308.2009.
Founds SA, Ren D, Roberts JM, Jeyabalan A, Powers RW. Follistatin-like 3 across gestation in preeclampsia and uncomplicated pregnancies among lean and obese women. Reprod Sci. 2015;22(4):402–9. https://doi.org/10.1177/1933719114529372.
Liu KY, Sengillo JD, Velez G, Jauregui R, Sakai LY, Maumenee IH, Bassuk AG, Mahajan VB, Tsang SH. Missense mutation in SLIT2 associated with congenital myopia, anisometropia, connective tissue abnormalities, and obesity. Orphanet J Rare Dis. 2018;13(1):138. https://doi.org/10.1186/s13023-018-0885-4.
Osorio-Conles O, Guitart M, Moreno-Navarrete JM, Escoté X, Duran X, Fernandez-Real JM, Gomez-Foix AM, Fernández-Veledo S, Vendrell J. Adipose tissue and serum CCDC80 in obesity and its association with related metabolic disease. Mol Med. 2017;23:225–34. https://doi.org/10.2119/molmed.2017.00067.
Dankel SN, Røst TH, Kulyté A, Fandalyuk Z, Skurk T, Hauner H, Sagen JV, Rydén M, Arner P, Mellgren G. The Rho GTPase RND3 regulates adipocyte lipolysis. Metabolism. 2019;101:153999. https://doi.org/10.1016/j.metabol.2019.153999.
Yasui M, Tamura Y, Minami M, Higuchi S, Fujikawa R, Ikedo T, Nagata M, Arai H, Murayama T, Yokode M. The Prostaglandin E2 Receptor EP4 Regulates Obesity-Related Inflammation and Insulin Sensitivity. PLoS One. 2015;10(8):e0136304. https://doi.org/10.1371/journal.pone.0136304.
Lu F, Liu Q. Validation of RUNX1 as a potential target for treating circadian clock-induced obesity through preventing migration of group 3 innate lymphoid cells into intestine. Med Hypotheses. 2018;113:98–101. https://doi.org/10.1016/j.mehy.2018.02.015.
Masaki M, Kurisaki T, Shirakawa K, Sehara-Fujisawa A. Role of meltrin {alpha} (ADAM12) in obesity induced by high- fat diet. Endocrinology. 2005;146(4):1752–63. https://doi.org/10.1210/en.2004-1082.
Khaidakov M, Mitra S, Kang BY, Wang X, Kadlubar S, Novelli G, Raj V, Winters M, Carter WC, Mehta JL. Oxidized LDL receptor 1 (OLR1) as a possible link between obesity, dyslipidemia and cancer. PLoS One. 2011;6(5):e20277. https://doi.org/10.1371/journal.pone.0020277.
Matsuo Y, Tanaka M, Yamakage H, Sasaki Y, Muranaka K, Hata H, Ikai I, Shimatsu A, Inoue M, Chun TH, et al. Thrombospondin 1 as a novel biological marker of obesity and metabolic syndrome. Metabolism. 2015;64(11):1490–9. https://doi.org/10.1016/j.metabol.2015.07.016.
Poggi M, Morin SO, Bastelica D, Govers R, Canault M, Bernot D, Georgelin O, Verdier M, Burcelin R, Olive D, et al. CD28 deletion improves obesity-induced liver steatosis but increases adiposity in mice. Int J Obes (Lond). 2015;39(6):977–85. https://doi.org/10.1038/ijo.2015.26.
Zhu Y, Wen L, Wang S, Zhang K, Cui Y, Zhang C, Feng L, Yu F, Chen Y, Wang R, et al. Omega-3 fatty acids improve flow-induced vasodilation by enhancing TRPV4 in arteries from diet-induced obese mice. Cardiovasc Res. 2020:cvaa296. https://doi.org/10.1093/cvr/cvaa296.
He L, Gunn TM, Bouley DM, Lu XY, Watson SJ, Schlossman SF, Duke-Cohan JS, Barsh GS. A biochemical function for attractin in agouti-induced pigmentation and obesity. Nat Genet. 2001;27(1):40–7. https://doi.org/10.1038/83741.
Moreno-Navarrete JM, Ortega F, Gómez-Serrano M, García-Santos E, Ricart W, Tinahones F, Mingrone G, Peral B, Fernández-Real JM. The MRC1/CD68 ratio is positively associated with adipose tissue lipogenesis and with muscle mitochondrial gene expression in humans. PLoS One. 2013;8(8):e70810. https://doi.org/10.1371/journal.pone.0070810.
Nam JS, Ahn CW, Park HJ, Kim YS. Semaphorin 3 C is a Novel Adipokine Representing Exercise-Induced Improvements of Metabolism in Metabolically Healthy Obese Young Males. Sci Rep. 2020;10(1):10005. https://doi.org/10.1038/s41598-020-67004-7.
Kwak SH, Park BL, Kim H, German MS, Go MJ, Jung HS, Koo BK, Cho YM, Choi SH, Cho YS, et al. Association of variations in TPH1 and HTR2B with gestational weight gain and measures of obesity. Obesity (Silver Spring). 2012;20(1):233–8. https://doi.org/10.1038/oby.2011.253.
Muñoz M, López-Oliva ME, Rodríguez C, Martínez MP, Sáenz-Medina J, Sánchez A, Climent B, Benedito S, García-Sacristán A, Rivera L, et al. Differential contribution of Nox1, Nox2 and Nox4 to kidney vascular oxidative stress and endothelial dysfunction in obesity. Redox Biol. 2020;28:101330. https://doi.org/10.1016/j.redox.2019.101330.
Klöting N, Wilke B, Klöting I. Alleles on rat chromosome 4 (D4Got41-Fabp1/Tacr1) regulate subphenotypes of obesity. Obes Res. 2005;13(3):589–95. https://doi.org/10.1038/oby.2005.63.
Van Camp JK, De Freitas F, Zegers D, Beckers S, Verhulst SL, Van Hoorenbeeck K, Massa G, Verrijken A, Desager KN, Van Gaal LF, et al. Investigation of common and rare genetic variation in the BAMBI genomic region in light of human obesity. Endocrine. 2016;52(2):277–86. https://doi.org/10.1007/s12020-015-0778-4.
Zhang ZB, Ruan CC, Lin JR, Xu L, Chen XH, Du YN, Fu MX, Kong LR, Zhu DL, Gao PJ. Perivascular Adipose Tissue-Derived PDGF-D Contributes to Aortic Aneurysm Formation During Obesity. Diabetes. 2018;67(8):1549–60. https://doi.org/10.2337/db18-0098.
Suriyaprom K, Pheungruang B, Tungtrongchitr R, Sroijit OY. Relationships of apelin concentration and APLN T-1860C polymorphism with obesity in Thai children. BMC Pediatr. 2020;20(1):455. https://doi.org/10.1186/s12887-020-02350-z.
Vaittinen M, Kolehmainen M, Rydén M, Eskelinen M, Wabitsch M, Pihlajamäki J, Uusitupa M, Pulkkinen L. MFAP5 is related to obesity-associated adipose tissue and extracellular matrix remodeling and inflammation. Obesity (Silver Spring). 2015;23(7):1371-1378. doi:https://doi.org/10.1002/oby.21103
Wolff G, Taranko AE, Meln I, Weinmann J, Sijmonsma T, Lerch S, Heide D, Billeter AT, Tews D, Krunic D, et al. Diet-dependent function of the extracellular matrix proteoglycan Lumican in obesity and glucose homeostasis. Mol Metab. 2019;19:97–106. https://doi.org/10.1016/j.molmet.2018.10.007.
Soomro I, Hong A, Li Z, Duncan JS, Skolnik EY. Discoidin Domain Receptor 1 (DDR1) tyrosine kinase is upregulated in PKD kidneys but does not play a role in the pathogenesis of polycystic kidney disease. PLoS One. 2019;14(7):e0211670. https://doi.org/10.1371/journal.pone.0211670.
Zeng H, Qi X, Xu X, Wu Y. TAB1 regulates glycolysis and activation of macrophages in diabetic nephropathy. Inflamm Res. 2020;69(12):1215–34. https://doi.org/10.1007/s00011-020-01411-4.
Zalli D, Bayliss R, Fry AM. The Nek8 protein kinase, mutated in the human cystic kidney disease nephronophthisis, is both activated and degraded during ciliogenesis. Hum Mol Genet. 2012;21(5):1155–71. https://doi.org/10.1093/hmg/ddr544.
Li YB, Wu Q, Liu J, Fan YZ, Yu KF, Cai Y. miR-199a-3p is involved in the pathogenesis and progression of diabetic neuropathy through downregulation of SerpinE2. Mol Med Rep. 2017;16(3):2417–24. https://doi.org/10.3892/mmr.2017.6874.
Zhou XJ, Cheng FJ, Qi YY, Zhao YF, Hou P, Zhu L, Lv JC, Zhang H. FCGR2B and FCRLB gene polymorphisms associated with IgA nephropathy. PLoS One. 2013;8(4):e61208. https://doi.org/10.1371/journal.pone.0061208.
Tsai YC, Kuo PL, Hung WW, Wu LY, Wu PH, Chang WA, Kuo MC, Hsu YL. Angpt2 Induces Mesangial Cell Apoptosis through the MicroRNA-33-5p-SOCS5 Loop in Diabetic Nephropathy. Mol Ther Nucleic Acids. 2018;13:543–55. https://doi.org/10.1016/j.omtn.2018.10.003.
Ohtsubo H, Okada T, Nozu K, Takaoka Y, Shono A, Asanuma K, Zhang L, Nakanishi K, Taniguchi-Ikeda M, Kaito H, et al. Identification of mutations in FN1 leading to glomerulopathy with fibronectin deposits. Pediatr Nephrol. 2016;31(9):1459–67. https://doi.org/10.1007/s00467-016-3368-7.
Gerarduzzi C, Kumar RK, Trivedi P, Ajay AK, Iyer A, Boswell S, Hutchinson JN, Waikar SS, Vaidya VS. Silencing SMOC2 ameliorates kidney fibrosis by inhibiting fibroblast to myofibroblast transformation. JCI Insight. 2017;2(8):e90299. https://doi.org/10.1172/jci.insight.90299.
Kim JM, Wu H, Green G, Winkler CA, Kopp JB, Miner JH, Unanue ER, Shaw AS. CD2-associated protein haploinsufficiency is linked to glomerular disease susceptibility. Science. 2003;300(5623):1298–300. https://doi.org/10.1126/science.1081068.
Li QS, Cheng P, Favis R, Wickenden A, Romano G, Wang H. SCN9A Variants May be Implicated in Neuropathic Pain Associated With Diabetic Peripheral Neuropathy and Pain Severity. Clin J Pain. 2015;31(11):976–82. https://doi.org/10.1097/AJP.0000000000000205.
Bennett CE, Nsengimana J, Bostock JA, Cymbalista C, Futers TS, Knight BL, McCormack LJ, Prasad UK, Riches K, Rolton D, et al. CCAAT/enhancer binding protein alpha, beta and delta gene variants: associations with obesity related phenotypes in the Leeds Family Study. Diab Vasc Dis Res. 2010;7(3):195–203. https://doi.org/10.1177/1479164110366274.
Domingues-Montanari S, Subirana I, Tomás M, Marrugat J, Sentí M. Association between ESR2 genetic variants and risk of myocardial infarction. Clin Chem. 2008;54(7):1183–9. https://doi.org/10.1373/clinchem.2007.102400.
Eberlé D, Clément K, Meyre D, Sahbatou M, Vaxillaire M, Le Gall A, Ferré P, Basdevant A, Froguel P, Foufelle F. SREBF-1 gene polymorphisms are associated with obesity and type 2 diabetes in French obese and diabetic cohorts. Diabetes. 2004;53(8):2153–7. https://doi.org/10.2337/diabetes.53.8.2153.
Cheng J, Song J, He X, Zhang M, Hu S, Zhang S, Yu Q, Yang P, Xiong F, Wang DW, et al. Loss of Mbd2 Protects Mice Against High-Fat Diet-Induced Obesity and Insulin Resistance by Regulating the Homeostasis of Energy Storage and Expenditure. Diabetes. 2016;65(11):3384–95. https://doi.org/10.2337/db16-0151.
Cavallari JF, Fullerton MD, Duggan BM, Foley KP, Denou E, Smith BK, Desjardins EM, Henriksbo BD, Kim KJ, Tuinema BR, et al. Muramyl Dipeptide-Based Postbiotics Mitigate Obesity-Induced Insulin Resistance via IRF4. Cell Metab. 2017; 25(5):1063-1074.e3. doi:https://doi.org/10.1016/j.cmet.2017.03.021
Qi L, Saberi M, Zmuda E, Wang Y, Altarejos J, Zhang X, Dentin R, Hedrick S, Bandyopadhyay G, Hai T, et al. Adipocyte CREB promotes insulin resistance in obesity. Cell Metab. 2009;9(3):277–86. https://doi.org/10.1016/j.cmet.2009.01.006.
Yan X, Zhu MJ, Xu W, Tong JF, Ford SP, Nathanielsz PW, Du M. Up-regulation of Toll-like receptor 4/nuclear factor-kappaB signaling is associated with enhanced adipogenesis and insulin resistance in fetal skeletal muscle of obese sheep at late gestation. Endocrinology. 2010;151(1):380–7. https://doi.org/10.1210/en.2009-0849.
Matsha TE, Kengne AP, Hector S, Mbu DL, Yako YY, Erasmus RT. MicroRNA profiling and their pathways in South African individuals with prediabetes and newly diagnosed type 2 diabetes mellitus. Oncotarget. 2018;9(55):30485–98. https://doi.org/10.18632/oncotarget.25271.
Ding L, Ai D, Wu R, Zhang T, Jing L, Lu J, Zhong L. Identification of the differential expression of serum microRNA in type 2 diabetes. Biosci Biotechnol Biochem. 2016;80(3):461–5. https://doi.org/10.1080/09168451.2015.1107460.
Hall CL, Akhtar MM, Sabater-Molina M, Futema M, Asimaki A, Protonotarios A, Dalageorgou C, Pittman AM, Suarez MP, Aguilera B, et al. Filamin C variants are associated with a distinctive clinical and immunohistochemical arrhythmogenic cardiomyopathy phenotype. Int J Cardiol. 2020;307:101–8. https://doi.org/10.1016/j.Eijcard.2019.09.048.
Salazar-Mendiguchía J, Ochoa JP, Palomino-Doza J, Domínguez F, Díez-López C, Akhtar M, Ramiro-León S, Clemente MM, Pérez-Cejas A, Robledo M, et al. Mutations in TRIM63 cause an autosomal-recessive form of hypertrophic cardiomyopathy. Heart. 2020;106(17):1342–8. https://doi.org/10.1136/heartjnl-2020-316913.
Xiao Y, Deng Y, Yuan F, Xia T, Liu H, Li Z, Chen S, Liu Z, Ying H, Liu Y, et al. An ATF4-ATG5 signaling in hypothalamic POMC neurons regulates obesity. Autophagy. 2017;13(6):1088–9. https://doi.org/10.1080/15548627.2017.1307488.
Stratigopoulos G, LeDuc CA, Cremona ML, Chung WK, Leibel RL. Cut-like homeobox 1 (CUX1) regulates expression of the fat mass and obesity-associated and retinitis pigmentosa GTPase regulator-interacting protein-1-like (RPGRIP1L) genes and coordinates leptin receptor signaling. J Biol Chem. 2011;286(3):2155–70. https://doi.org/10.1074/jbc.M110.188482.
Zhou JP, Ren YD, Xu QY, Song Y, Zhou F, Chen MY, Liu JJ, Chen LG, Pan JS. Obesity-Induced Upregulation of ZBTB7A Promotes Lipid Accumulation through SREBP1. Biomed Res Int. 2020:4087928. https://doi.org/10.1155/2020/4087928.
Acknowledgements
I thank Jun Yoshino, Washington University School of Medicine, Medicine, St. Louis, USA, very much, the author who deposited their profiling by high throughput sequencing dataset, GSE143319, into the public GEO database.
Conflict of interest
The authors declare that they have no conflict of interest.
Informed consent
No informed consent because this study does not contain human or animals participants.
Author information
Authors and Affiliations
Contributions
P. G - Methodology and validation. B. V - Writing original draft, and review and editing. A. T - Formal analysis and validation. C. V - Software and investigation. I. K - Supervision and resources. The authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Prashanth, G., Vastrad, B., Tengli, A. et al. Investigation of candidate genes and mechanisms underlying obesity associated type 2 diabetes mellitus using bioinformatics analysis and screening of small drug molecules. BMC Endocr Disord 21, 80 (2021). https://doi.org/10.1186/s12902-021-00718-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12902-021-00718-5