Abstract
The innate immune system is the first line of host defense, which responds rapidly to viral infection. Innate recognition of viruses is mediated by a set of pattern recognition receptors (PRRs) that sense viral genomic nucleic acids and/or replication intermediates. PRRs are mainly localized either to the endosomes, the plasma membrane or the cytoplasm. Recent evidence suggested that several proteins located in the nucleus could also act as viral sensors. In turn, these important elements are becoming the target for most viruses to evade host immune surveillance. In this review, we focus on the recent progress in the study of viral recognition and evasion.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Viruses, including influenza virus, Ebola virus, Zika virus, pose significant global public health threats. Immunization is essential for controlling and eliminating viral infections. The innate immune system provides nonspecific defense mechanisms against pathogens and comes into play immediately. Profound understanding of the relationship between viral recognition and viral evasion may provide new insights for the transmission and treatment of viral infectious diseases. In this review, we summarize recent discoveries about the cytosolic and nuclear viral nucleic acids sensing pathway. Besides, elaborate escape strategies for viruses at different steps of the signaling transduction are also included.
Recognition of viral nucleic acids
Many kinds of pathogens experience intra-cellular stage during infection. After invasion, these pathogens utilize and hijack the host cellular environment and materials to facilitate their replication and proliferation. During this period, components of pathogens, such as nucleic acids and polysaccharides can be exposed. These components called the pathogen-associated molecular patterns (PAMPs) [1]. Host have evolved sophisticated mechanism to identify these foreign substances, which is known as the pattern recognition receptors (PRRs). PRRs can recognize PAMPs and initiate downstream signal activation, including the innate immune signaling cascades or programmed cell death processes [2]. Cytosolic sensors play an important role in recognizing intra-cellular PAMPs [3]. According to their structural and functional characteristics, cytosolic PRRs are mainly divided into retinoic acid inducible gene-I-like receptors (RLRs) [4], nucleotide binding oligomerization domain-like receptors (NOD-like receptors, NLRs) [5], AIM2-like receptors (ALRs) [6], cyclic GMP-AMP synthase (cGAS) [7], nucleases and other DExD/H-box family helicases besides RLRs. In the recent years, several proteins present in the nucleus also showed sensor activity, which could sense viral nucleic acid generated in the nucleus. These sensors shared distinct recognition diversities and together made up the cytosolic and nuclear innate immune sensing network.
Cytosolic viral RNA sensing
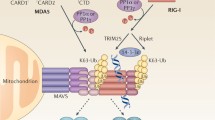
Various kinds of pathogens, such as virus, parasites like Plasmodium and bacteria, can release RNA into the cytoplasm of infected cells [8]. RIG-I like receptors (RLRs) are a class of RNA helicases that can recognize and bind RNA from different origin, including RNA from pathogens, long non-coding RNAs, mitochondrial RNA and small interfering RNAs of cells [9]. Canonical members of RLRs include retinoic acid inducible gene I (RIG-I, also called DDX58), melanoma differentiation associated gene 5 (MDA5) and DExH-box helicase 58 (LGP2, also called DHX58) [4]. These three molecules can recognize double-stranded RNA (dsRNA) in the cytoplasm, but recognition specificity and functional characteristics are different. MDA5 and RIG-I have a similar structure, both containing the N-terminal caspase-associated recruitment domain (CARD), the helicase domain in the middle, and their respective C-terminal region [10]. N-terminal domain mediates the oligomerization of RIG-I and MDA5 and also mediates the interaction of RIG-I and MDA5 with the downstream protein mitochondrial antiviral signaling protein (MAVS). dsRNA recognition is mediated by RIG-I and MDA5 helicase domains and the C-terminal domains [11,12,13,14]. Regarding the specificity of RNA recognition, RIG-I is primarily responsible for recognizing short dsRNA with a 5′-triphosphate (5′-ppp) in the cytoplasm, while MDA5 is primarily responsible for recognizing dsRNA larger than 2000 bp such as genomic RNA of EMCV [15, 16]. Circular RNA synthesized in vitro and derived from virus can also bind and activate RIG-I [17]. Because of the constitutive expression of RLRs, the activity of RLRs needs to be suppressed at resting state to prevent excessive inflammation and the occurrence of autoimmune diseases. Structural and functional studies have revealed that RIG-I exhibits a self-inhibitory effect when not binding to dsRNA [10]. On the other hand, the combination of dsRNA can eliminate this inhibitory effect and release the CARD of RIG-I [18]. RIG-I thus oligomerizes to form the activation status. When combined with dsRNA, every four RIG-I molecules form tetramerization via CTD. RIG-I tetramerization along the dsRNA forms fiber-like aggregates, which ultimately activates downstream MAVS-mediated signaling pathways [12, 13]. Ubiquitination of RIG-I, mediated by the E3 ligases tripartite motif-containing 25 (TRIM25) and Riplet, is critical for its activation. Recent studies have revealed the relative importance of these two enzymes in the activation RIG-I. Both TRIM25 and Riplet are reported to be involved in the activation of RIG-I, but Riplet is considered to be the most important E3 ubiquitin ligase in this process [19]. The consequence of ubiquitination is: on one hand, the K63 ubiquitination of RIG-I mediated by Riplet/TRIM25 promotes the formation of fiber-like aggregation when RIG-I binds to short dsRNA. On the other hand, fiber-like aggregation formed by Riplet/TRIM25 mediated bridging of RIG-I tetramers binding with long dsRNA promotes RIG-I activation [20].
MDA5 also undergoes K63 polyubiquitination mediated activation. In cell free systems, K63 polyubiquitination stabilizes the 2CARD oligomerization of MDA5, promoting the activation of MDA5 [21]. It has been reported that TRIM65 mediates the K63 polyubiquitination of MDA5. TRIM65 catalyzes K63 polyubiquitination of MDA5 on lysine 743, which is important for the antiviral ability of MDA5 [22]. Although having a helicase domain and CTD, unlike RIG-I or MDA5, DHX58 lacks N-terminal CARDs, which prevents it from activating MAVS and downstream signals despite its dsRNA binding function [23]. Nevertheless, DHX58 can still participate in the RLR signaling pathway via regulating RIG-I and MDA5 activation. DHX58 can competitively bind dsRNA, inhibiting RIG-I activation. Meanwhile, such competition effect can protect dsRNA from Dicer mediated cleavage and promote the binding of MDA5 with dsRNA, which is important for the downstream signaling transduction [3, 24, 25].
After binding with RNA in the cytosol, MDA5 or RIG-I is recruited to outer mitochondrial membrane, where it binds MAVS via CARD to promote MAVS oligomerization. The oligomerized MAVS further recruits TRAF2, TRAF3, TRAF5 and TRAF6 to form signalosome. MAVS signalosome then recruits TBK1, providing a platform for phosphorylation of IRF3. Phosphorylated IRF3 dimerizes and translocates into the nucleus to transcriptionally activate the production of antiviral type I interferons and pro-inflammatory cytokines. MAVS can also recruit IKKα/β/γ and activate NFκB to promote various pro-inflammatory cytokines transcription [26,27,28].
RLRs generally exhibit background expression. However, after viral or other pathogen infection, the expression levels of MDA5 and RIG-I will be significantly increased, which indicates that there is a positive feedback mechanism for natural immune signals in the RNA receptor pathway [29, 30].
Various DExD/H box RNA helicases are reported to act as RNA sensors. DDX1-DDX21-DHX36 complex recognizes viral dsRNA in the cytosol and activates TRIF pathway [31]. DDX19A senses viral RNA and activates NLRP3 inflammasome [32]. DHX33 senses cytosolic viral dsRNA and interacts with MAVS to activate innate immune signaling in dendritic cells [33]. Moreover, DHX33 also activates the NLRP3 inflammasome after binding to dsRNA to trigger cell pyroptosis [34]. DDX60 can bind viral DNA and RNA and is essential for RIG-I activation [35, 36].
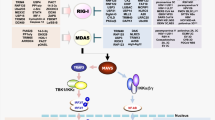
Apart from the sensors raised above, several interferon-inducible sensors participate in viral RNA sensing. DsRNA sensor 2′,5′-oligoadenylate (2-5A) synthetase (OAS) catalyzes ATP to 2′,5′-oligoadenylates, which further activates ribonuclease RNase L to mediate the cleavage of viral dsRNA [37, 38]. Interferon-induced protein with tetratricopeptide repeats 5 (IFIT5) and IFIT1 recognize ssRNA carrying a 5′-triphosphate (5′-ppp) with TPR domain, protecting cells from viral infection [39, 40]. Protein kinase regulated by RNA (PKR) links cellular stress to viral sensing. PKR recognizes viral dsRNA, followed by mediating translation arrest by phosphorylating eIF2α, changing the cell translation pattern and inhibiting viral replication [41]. Moreover, PKR remains inactivated by endogenous circRNA and is activated via RNase L mediated circRNA degradation after viral infection [42]. Adenosine deaminase acting on dsRNA 1 (ADAR1) shares structural similarity with PKR for both having multiple dsRNA binding domains. Full length ADAR1 (p150) is an interferon-stimulated gene (ISG) and mainly localizes in the cytosol [43]. ADAR1 recognizes viral dsRNA and catalyzes the adenosine (A) to inosine (I), resulting in A-to-I editing. Such editing of viral dsRNA mainly suppresses dsRNA-induced signaling, including the blockage of PKR induced translation arrest and inhibition of innate immune signaling activation induced by RIG-I [44, 45]. Leucine-rich repeat (LRR) protein LRRFIP1 recognizes dsRNA and responds to VSV infection. Meanwhile, LRRFIP1 also senses dsDNA from L. monocytogenes. The binding of LRRFIP1 to dsRNA or dsDNA is essential for β-catenin phosphorylation. β-catenin is an important transcriptional co-activator of Ifnb1 promoter [46] (Fig. 1).
Cytosolic viral RNA sensing. The RLR family members MDA5 and RIG-I preferentially recognize long dsRNA and 5′- triphosphate short dsRNA, respectively. RIG-I can also recognize circular RNA synthesized in vitro and derived from virus. After RNA ligand binding, RIG-I and MDA5 undergo conformational changes that expose N-terminal CARDs to mediate downstream signaling. Riplet/TRIM25 or TRIM65-mediated K63-linked ubiquitination induces RIG-I and MDA5 activation, respectively. Both RIG-I and MDA5 can interact with MAVS on the outer membrane of mitochondria and activate MAVS to form signal platform, which further activates NFκB via the IKK complex or IRF3 via TBK1. DHX58 is reported to facilitate MDA5 activation, whereas it drives the inactivation of RIG-I. Besides, DDX1-DDX21-DHX36 complex recognizes viral dsRNA and activates TRIF pathway. DDX19A senses viral RNA and activates NLRP3 inflammasome. DHX33 senses cytosolic viral dsRNA and interacts with MAVS to activate innate immune signaling. DDX60 is needed for the activation of RIG-I by binding to viral RNA. ADAR1 recognizes viral dsRNA suppresses dsRNA-induced signaling. LRRFIP1 recognizes dsRNA and L. monocytogenes dsDNA. Moreover, IFIT1 and IFIT5 can recognize 5′- triphosphate ssRNA
Cytosolic viral DNA sensing
In recent years, various DNA binding proteins have been reported to act as DNA recognition receptors. One of the most important DNA recognition receptors is cyclic GMP-AMP synthase (cGAS). cGAS is an interferon-inducible gene, which means that DNA-mediated innate immune signaling pathways also have positive feedback mechanism as RNA-mediated pathways [47]. After recognizing the double-stranded DNA (dsDNA) located in the cytosol, cGAS uses GTP and ATP to synthesize the second messenger cGAMP. Cytosolic cGAMP activates downstream STING-TBK1-IRF3 signaling. cGAS has different binding modes when recognizing DNA of different lengths. When recognizing dsDNA less than 20 bp, dimerized cGAS each binds one dsDNA nucleotide, forming a 4-molecule complex. When binding the dsDNA longer that 40 bp, the dimerized cGAS lines along the dsDNA and forms a phase-separated structure to efficiently synthesize cGAMP [48]. cGAS activated by viral DNA can induce the activation of innate immunity and up-regulate the expression of type I interferon and various pro-inflammatory cytokines, subsequently activating adaptive immune system to promote the elimination of pathogens [49]. Recently, it has been reported that via N-terminal phosphoinositol binding domain, cGAS can localize to phagocytes plasmalemma. N-terminal domain of cGAS selectively interacts with PI (4,5) P2, mutation of this domain results in cGAS lipid binding defects and mislocates to the cytoplasm and nucleus. The mis-localized cGAS causes strong interferon in response to genotoxic stress, but the cGAS response induced by viral infection is weakened [50].
Several members of DExD/H helicases are reported to respond to cytosolic dsDNA. It has been reported that both DHX9 and DHX36 can sense cytosolic CpG-DNA, with DHX9 recognizing CpG-B DNA with DUF1605 domain and DHX36 recognizing CpG-A DNA with DEAH domain [51]. DDX41 is the first reported cytoplasmic DNA sensor. DDX41 uses its DEAD domain to recognize dsDNA and interact with STING to activate the innate immune signaling [52]. RNA Pol III is the sensor for AT-rich dsDNA in the cytoplasm and is widely expressed. RNA Pol III recognizes viral dsDNA and transcribes it into dsRNA, which in turn recognized by RIG-I to activate innate immune response [53]. The DNA-dependent protein kinase (DNA-PK), a DNA damage response mediator, was reported to act as a dsDNA sensor. DNA-PK is a signaling complex consisting of three members, namely the catalytic subunit DNA-PKcs and DNA binding subunits Ku70 and Ku80 [54]. DNA-PK recognizes cytoplasmic dsDNA of vaccinia virus, which is important for IRF3-mediated innate immune responses in fibroblasts [55]. Moreover, DNA-PK relies on STING, TBK1, and IRF3 to induce the production of cytokines during VACV infection [56]. It is well known that three prime repair exonuclease 1 (TREX1) can protect host from excessive inflammation via efficiently degrading cytosolic endogenous DNA [56, 57]. Moreover, in the case of HIV infection, inhibiting the function of TREX1 can augment the production of type I IFN and suppress HIV replication and spreading, indicating that TREX1 negatively regulates HIV-induced antiviral innate immunity [58].
Another family of cytosolic DNA sensor are PYHIN family proteins. Members of this family contain an N-terminal Pyrin domain, which mediates the interaction with other Pyrin domain containing proteins. PYHIN family members also contain one or two C-terminal HIN domains, which are involved in DNA binding [59, 60]. Human genome contains 4 PYHIN family members, whereas murine genome contains 14 members [61]. Of the PYHIN members, AIM2-like receptors (ALRs), including AIM2, IFI16 and IFI204 (ortholog of human IFI16) are identified as PRRs [6]. Unlike IFI16 or IFI204, AIM2 exists in both human and murine genome. AIM2 exhibits a self-inhibitory state without binding with DNA, which is first thought to be mediated by the self-Pyrin-HIN domain interaction [62]. However, a recent study suggested that Pyrin domain was not self-inhibitory, but provided a platform for ligand binding [63]. AIM2 senses dsDNA from virus such as vaccinia virus and forms AIM2 inflammasomes along with caspase-1 and apoptosis-associated speck-like protein containing CARD (ASC) [64]. IFI16 was reported to bind viral DNA of herpes simplex virus 1 (HSV-1) in the cytoplasm, then mediates the activation of STING-TBK1-IRF3 axis [65]. However, as an ortholog of human IFI16, IFI204 plays controversial roles in innate immune response. IFI204 is important for TLR4 signaling and may facilitate host to fight against Staphylococcus aureus and Francisella infection, but negatively regulates antiviral innate immunity via inhibiting the target promoter binding of IRF7 in the nucleus [66,67,68,69]. Exploring the function of other PYHIN family members, especially their roles in antiviral innate immunity, will help understand the compilated regulation mechanism of innate immune response. Moreover, NLRC3 can act as a DNA sensor. NLRC3 recognizes dsDNA from HSV-1, releasing STING from binding to NLRC3, which is an inactive form of STING, to activate type I interferon induction [70] (Fig. 2). In order to link the DNA/RNA sensors with recognition of viruses, we list the viruses that are sensed by the individual PRRs in Table 1.
Cytosolic viral DNA sensing. DNA is a PAMP that can be delivered to the cytoplasm of host cells during microbial infection. Several DNA sensors have been reported to promote the activation of innate immune signaling by activating a STING-dependent signaling pathway. Among them, cGAS can catalyze ATP and GTP to generate the second messenger cyclic GMP-AMP (cGAMP) and then activate STING. However, the molecular basis by which other DNA sensors activate STING is not well understood. RNA polymerase III can activate the immune response in a STING-independent way, RNA polymerase III can transcribe poly (dA-dT) into dsRNA, which is then recognized by RIG-I. Several members of DExD/H helicases are also respond to cytosolic dsDNA, such as DHX9, DHX36 and DDX41. In IRF3-mediated innate immune responses of fibroblasts, DNA-PK recognizes cytoplasmic dsDNA of vaccinia virus. TREX1 is well known to protect host from excessive inflammation via efficiently degrading cytosolic endogenous DNA. PYHIN family proteins: AIM2 and IFI16. AIM2 senses dsDNA from virus and oligomerizes to form AIM2 inflammasome. IFI16 binds to cytoplasmic viral DNA and mediates the activation of STING-TBK1-IRF3 axis. Moreover, NLRC3 recognizes dsDNA and releases STING, which activates type I interferon induction
Nuclear viral DNA/RNA sensing
Virus, including DNA and some RNA virus, replication generates nucleic acid in the nucleus. Generally, most DNA virus replicate within the host cell nucleus, which leads to their genomic DNA ejection or viral DNA/RNA generation during their replication in host cell nucleus [85]. Most RNA viruses replicate in the cytosol. However, retroviruses and some single-stranded RNA virus, such as influenza virus, can replicate in the nucleus and their genomic RNA can be detected in the nucleus of the infected cells. Most cytoplasmic replicated RNA viruses, such as VSV or Rabies virus, can generate the leader RNA transcript which can present in host cell nucleus and mediate viral replication and transcription.
Most PRRs are localized on the plasma membrane, endosomes or cytoplasm. Recent evidence suggested that several proteins located in the nucleus could also act as viral sensors. For example, IFI16 can recognize HSV-1 virus DNA within the nucleus and leads to the expression of IFNβ via activation of IRF3 signaling [86]. Nuclear IFI16 can also recognize viral DNA from HIV, causing the pyroptosis of infected CD4+ T cells [87]. When human endothelial cells are infected with KSHV, IFI16 can interact with procaspase-1 and ASC to form functional inflammasome. This inflammasome is involved in viral DNA recognition, caspase-1 activation and IL-1β maturation [88].
cGAS is widely accepted as a cytoplasmic dsDNA sensor. Recently several researches suggested that cGAS is primarily a nuclear protein and the tethering prevents genomic DNA from activating cGAS. Nuclear localized cGAS is tightly tethered through salt-resistant interaction. cGAS tethering requires intact nuclear chromatin to keep cGAS in resting state and prevent its own reactivity [89]. In addition, the nucleosome-binding interface occupies the strong dsDNA binding surface on cGAS, thus effectively prevents cGAS from oligomerizing into a functional active state [90]. Lately, cGAS is reported to bind to nuclear IFI16 and can promote the stability of IFI16 in human fibroblasts and keratinocytes during HSV-1 infection [91].
Normally, hnRNPA2B1 is an RNA-binding protein localized in the nucleus. Cao et al. found that hnRNPA2B1 could bind viral DNA in various mouse and human cells nuclei during HSV-1 infection [92,93,94]. hnRNPA2B1 can form a complex with viral DNA and undergo homodimerization and demethylation, resulting in the translocation of the complex into the cytoplasm and the activation of immune response mediated by type I interferon signaling.
Our group confirmed that scaffold attachment factor A (SAFA) localized in the nucleus can act as a trans-activator of antiviral genes and a viral dsRNA sensor. Upon viral infection, SAFA senses dsRNA generated from virus (such as HSV-1 or VSV) replication, oligomerizes, and facilitates antiviral immunity inducing IFNβ production through interacting with DNA topoisomerase 1 (TOP1) and SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 5 (SMARCA5) in human and mouse primary cells. Besides, oligomerized SAFA mediates IFNB1 transcription and maintains antiviral status of infected cells through interacting with enhancers and super-enhancers [95].
NONO (non-POU domain containing octamer binding) is an important sensor for HIV capsid recognition in the nuclei of dendritic cells and macrophages. The directly binding affinity of NONO to weakly pathogenic HIV-2 capsid is stronger than that of highly pathogenic HIV-1. NONO is necessary for cGAS to associate with nuclear HIV DNA and thus activates cGAS. Moreover, NONO can recognize conserved regions in the capsid of HIV and has limited escape tolerance to mutations. The promotion of DNA sensing by cGAS followed by detection of nuclear viral capsid by NONO reveals the basic strategy of differentiating non-self from self in the nucleus [96].
It is well known that hexamethylene bis-acetamide-inducible protein 1 (HEXIM1) inhibits the positive transcription elongation factor b (P-TEFb) and can bind to RNA in the nucleus [97]. P-TEFb controls transcriptional elongation through RNA polymerase II. HEXIM1 is the crucial component of the 7SK RNP complex and plays a significant role in inhibiting RNA polymerase II phosphorylation and succeeding transcriptional elongation [98,99,100,101]. HEXIM1 plays an critical role in regulating the innate immune response mediated by DNA infection, it can bind long non-coding RNA such as NEAT1 to promote the formation of the HDP-RNP complex, which acts as a platform for activation through the cGAS-STING pathway and subsequent IRF3 phosphorylation [102].
More and more proteins have been found to recognize nuclear viral nucleic acids, which raises question of how they distinguish non-self from self-nucleic acids substances. It may be partially explained by the highly ordered structure of the nucleus and far more research is needed.
The strategies of viral innate immune evasion
Viruses have evolved a series of effective tactics to evade the surveillance of host’s innate immunity in order to replicate and spread efficiently. We thus describe the molecular strategies that may be used by viruses to evade host innate immunity, including viral evasion of PRRs detection and block the activation of signaling molecule at different levels.
PRRs function as sensors to initially response to invading pathogens. Nevertheless, viruses have developed different strategies to evade PRRs surveillance. Viruses can utilize cellular membrane to form a confined space or replicate in organelles (endoplasmic reticulum, Golgi complex or mitochondrion) to avoid being detected by RLRs. For example, Dengue virus (DENV) can replicate in endoplasmic reticulum to efficiently hide viral dsRNA from the cytoplasm [103]. Several viruses can modify their genomes to avoid being detected by RLRs. In order to escape the surveillance of RIG-I, Borna disease virus (BDV) encodes phosphatases that convert 5′-triphosphate (5′-ppp) on its genome to 5′-monophosphate (5′-p) [104]. In addition, many viruses can block PRRs recognition of viral nucleic acids by using host-encoded or viral proteins. For example, EBOV’s viral protein 35 (VP35) competitively binds to viral dsRNA to prevent dsRNA from being detected by RIG-I [105].
In addition to developing mechanisms to evade host detection, viruses also target the PRRs level by segregating and modifying the receptors. HSV-1 capsid protein US11 can interact with RIG-I and prevent RIG-I from forming RIG-I/MAVS complex [106]. Lys63 linked polyubiquitylation is important for the activation of RIG-I. The NS1 protein of Influenza A virus (IAV) can inhibit the activity of the E3 ubiquitin ligases TRIM25 or Riplet and thus prevent Lys63 linked ubiquitination of RIG-I [107, 108]. MDA5 is an intracellular sensor that can recognize viral long dsRNA [10, 109]. The V protein of Parainfluenza virus 5 (PIV5-V) can recognize and bind to MDA5 structural motif, disrupting the ATP hydrolysis activity and filament formation of MDA5 [110]. In the absence of viral infection, phosphorylation of threonine or serine leaves MDA5 and RIG-I in an inactive state. In the case of virus infection, PP1α or PP1γ mediated dephosphorylation is crucial for RIG-I and MDA5 activation. The V proteins of Nipah virus (NiV-V) can act as an antagonistic to PP1γ and PP1α, reducing the dephosphorylation of MDA5 and RIG-I, impairing downstream innate immune responses [111].
Degradation of the sensor or the key component is an effective way to inhibit RLR signaling. Intriguingly, many viruses can directly degrade RLRs by expressing proteases. For example, 3C protease of Enterovirus 71 (EV71), Poliovirus, EMCV, Echovirus, Rhinovirus type 1A and Rhinovirus type 16 can directly cleave RIG-I [112], whereas the 2A protease of EV71 can cleave MDA5 [113].
The signal platform formed by MAVS is critical for activating the downstream molecules TBK1 (TANK binding kinase 1) and IRF3 (interferon regulatory factor 3). Degradation of the adaptor MAVS is also a common event during virus infection. 3C protease from hepatitis A virus (HAV), 2A protease and 3C protease from rhinovirus can directly target and cleave MAVS [114,115,116,117]. Interestingly, IAV can block MAVS signaling activation in a proteasome-independent manner. The IAV protein PB1-F2 can bind to MAVS transmembrane region to block type I interferon signaling by decreasing the mitochondrial localized MAVS protein level [118]. In the innate immune response, TBK1 is an important kinase. When the activity of TBK1 is blocked, IRF3 cannot be phosphorylated and cannot be dimerized or translocate into nucleus. The NS3 protein of hepatitis C virus (HCV) and the N1L76 protein of vaccinia virus (VACV) can bind to TBK1 and inhibit the downstream signaling activation [119, 120] (Fig. 3).
Innate immune evasion strategies of targeting RLR signaling pathway. Viruses can escape from host immune clearance. For example, DENV can replicate in endoplasmic reticulum to efficiently hide dsRNA from the cytoplasm. BDV encodes phosphatases to process the 5′-ppp on its genome to 5′-p. The protein viral protein 35 (VP35) of EBOV can interact with dsRNA to prevent dsRNA from being detected by RLRs. Moreover, RLR signaling can be inhibited by viral proteins that either directly bind MDA5, RIG-I, MAVS, TBK1 to inhibit their function or induce their degradation
cGAS acts as a crucial DNA sensor for the detection of viral dsDNA and mediates the production of proinflammatory cytokines and IFNs. Viruses are able to inhibit cGAS dependent innate immune signaling via distinct approaches. Viral proteins can bind to cGAS and affect its DNA-binding activity or enzymatic activity. For example, ORF52 of Kaposi’s sarcoma-associated herpesvirus (KSHV) can interact with cGAS, blocking enzymatic activity of cGAS and impairing downstream innate immune responses, ultimately facilitating the evasion of KSHV [121]. Viral proteins can also mediate the degradation of cGAS. After DENV infection, cGAS can detect the released DNA from the mitochondria. Nevertheless, NS2B protein of DENV can degrade cGAS by lysosomal degradation [122]. IFI16 is the nuclear DNA sensor. The viral E3 ubiquitin ligase ICP0 of HSV-1 can medicate proteasomal degradation of IFI16 [123]. pUL83 protein of the human cytomegalovirus (HCMV) binds to PYD domain of IFI16 and inhibits IFI16 oligomerization, dampening downstream immune signaling activation. Besides, VACV proteins C16 and C4 can antagonize DNA-PK. C4 and C16 can bind to Ku and prevent Ku from binding to DNA [124, 125], thus reducing the production of chemokines and cytokines, decreasing the recruitment of inflammatory cells, inhibiting the IRF3 activation. Moreover, E1A oncoprotein of human adenovirus 5 and the ICP0 protein of HSV-1 can also block DNA-PK induced robust and broad antiviral response [56].
It has been reported that viruses can disturb STING’s function. For example, viral protein vIRF1 from KSHV can bind to STING as an antagonist [121]. In addition, the protease complex NS2B–NS3 of DENV can cleave STING to reduce the IFNs signaling transduction [126] (Fig. 4).
Conclusions
Research over the past two decades have extensively investigated the role of innate immune system and related PRR in virus recognition and initiation of antiviral defenses. Recent studies found viral RNA and DNA sensing in the nucleus of viruses infected cells. The detail regulatory mechanism of nuclear viral recognition and subsequent epigenetic alteration await to be investigated. Besides, viruses have developed diverse strategies to suppress immune responses for evasion. However, The research of immune escape lags far behind immune recognition. Far more research is needed to broadly clarify the mechanisms and identify the potential targets for viral immune evasion. Given the initial promising results, a better understanding of host-viral biology will bring great benefits to the novel clinical applications.
Availability of data and materials
Not applicable.
References
Barton GM, Kagan JC. A cell biological view of toll-like receptor function: regulation through compartmentalization. Nat Rev Immunol. 2009;9(8):535–42. https://doi.org/10.1038/nri2587.
Cao X. Self-regulation and cross-regulation of pattern-recognition receptor signalling in health and disease. Nat Rev Immunol. 2016;16(1):35–50. https://doi.org/10.1038/nri.2015.8.
Bruns AM, Leser GP, Lamb RA, Horvath CM. The innate immune sensor LGP2 activates antiviral signaling by regulating MDA5-RNA interaction and filament assembly. Mol Cell. 2014;55(5):771–81. https://doi.org/10.1016/j.molcel.2014.07.003.
Loo YM, Gale M Jr. Immune signaling by RIG-I-like receptors. Immunity. 2011;34(5):680–92. https://doi.org/10.1016/j.immuni.2011.05.003.
Motta V, Soares F, Sun T, Philpott DJ. NOD-like receptors: versatile cytosolic sentinels. Phys Rev. 2015;95(1):149–78. https://doi.org/10.1152/physrev.00009.2014.
Man SM, Kanneganti TD. Regulation of inflammasome activation. Immunol Rev. 2015;265(1):6–21. https://doi.org/10.1111/imr.12296.
Sun L, Wu J, Du F, Chen X, Chen ZJ. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science. 2013;339(6121):786–91. https://doi.org/10.1126/science.1232458.
Tan X, Sun L, Chen J, Chen ZJ. Detection of microbial infections through innate immune sensing of nucleic acids. Annu Rev Microbiol. 2015;72(1):447–78. https://doi.org/10.1146/annurev-micro-102215-095605.
Hur S. Double-stranded RNA sensors and modulators in innate immunity. Annu Rev Immunol. 2019;37(1):349–75. https://doi.org/10.1146/annurev-immunol-042718-041356.
Chow KT, Gale M Jr, Loo YM. RIG-I and other RNA sensors in antiviral immunity. Annu Rev Immunol. 2018;36(1):667–94. https://doi.org/10.1146/annurev-immunol-042617-053309.
Yoneyama M, Kikuchi M, Natsukawa T, Shinobu N, Imaizumi T, Miyagishi M, et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nat Immunol. 2004;5(7):730–7. https://doi.org/10.1038/ni1087.
Kowalinski E, Lunardi T, McCarthy AA, Louber J, Brunel J, Grigorov B, et al. Structural basis for the activation of innate immune pattern-recognition receptor RIG-I by viral RNA. Cell. 2011;147(2):423–35. https://doi.org/10.1016/j.cell.2011.09.039.
Jiang F, Ramanathan A, Miller MT, Tang GQ, Gale M, Patel SS, et al. Structural basis of RNA recognition and activation by innate immune receptor RIG-I. Nature. 2011;479(7373):423–7. https://doi.org/10.1038/nature10537.
Wu B, Peisley A, Richards C, Yao H, Zeng X, Lin C, et al. Structural basis for dsRNA recognition, filament formation, and antiviral signal activation by MDA5. Cell. 2013;152(1–2):276–89. https://doi.org/10.1016/j.cell.2012.11.048.
Schlee M, Roth A, Hornung V, Hagmann CA, Wimmenauer V, Barchet W, et al. Recognition of 5′ triphosphate by RIG-I helicase requires short blunt double-stranded RNA as contained in panhandle of negative-strand virus. Immunity. 2009;31(1):25–34. https://doi.org/10.1016/j.immuni.2009.05.008.
Kato H, Takeuchi O, Mikamo-Satoh E, Hirai R, Kawai T, Matsushita K, et al. Length-dependent recognition of double-stranded ribonucleic acids by retinoic acid-inducible gene-I and melanoma differentiation-associated gene 5. J Exp Med. 2008;205(7):1601–10. https://doi.org/10.1084/jem.20080091.
Chen YG, Kim MV, Chen X, Batista PJ, Aoyama S, Wilusz JE, et al. Sensing self and foreign circular RNAs by intron identity. Mol Cell. 2017;67(2):228–38e5. https://doi.org/10.1016/j.molcel.2017.05.022.
Rawling DC, Fitzgerald ME, Pyle AM. Establishing the role of ATP for the function of the RIG-I innate immune sensor. eLife. 2015;4:e09391. https://doi.org/10.7554/eLife.09391.
Hayman TJ, Hsu AC, Kolesnik TB, Dagley LF, Willemsen J, Tate MD, et al. RIPLET, and not TRIM25, is required for endogenous RIG-I-dependent antiviral responses. Immunol Cell Biol. 2019;97(9):840–52. https://doi.org/10.1111/imcb.12284.
Cadena C, Ahmad S, Xavier A, Willemsen J, Park S, Park JW, et al. Ubiquitin-dependent and -independent roles of E3 ligase RIPLET in innate immunity. Cell. 2019;177(5):1187–200e16. https://doi.org/10.1016/j.cell.2019.03.017.
Jiang X, Kinch LN, Brautigam Chad A, Chen X, Du F, Grishin NV, et al. Ubiquitin-induced oligomerization of the RNA sensors RIG-I and MDA5 activates antiviral innate immune response. Immunity. 2012;36(6):959–73. https://doi.org/10.1016/j.immuni.2012.03.022.
Lang X, Tang T, Jin T, Ding C, Zhou R, Jiang W. TRIM65-catalized ubiquitination is essential for MDA5-mediated antiviral innate immunity. J Exp Med. 2017;214(2):459–73. https://doi.org/10.1084/jem.20160592.
Yoneyama M, Kikuchi M, Matsumoto K, Imaizumi T, Fujita T. Shared and unique functions of the DExD/H-box helicases RIG-I, MDA5, and LGP2 in antiviral innate immunity. J Immunol. 2005;175(5):2851–8. https://doi.org/10.4049/jimmunol.175.5.2851.
Rothenfusser S, Goutagny N, Diperna G, Gong M, Monks BG, Schoenemeyer A, et al. The RNA helicase Lgp2 inhibits TLR-independent sensing of viral replication by retinoic acid-inducible gene-I. J Immunol. 2005;175(8):5260–8. https://doi.org/10.4049/jimmunol.175.5.2851.
Van der Veen AG, Maillard PV, Schmidt JM, Lee SA, Deddouche-Grass S, Borg A, et al. The RIG-I-like receptor LGP2 inhibits dicer-dependent processing of long double-stranded RNA and blocks RNA interference in mammalian cells. EMBO J. 2018;37(4):e97479. https://doi.org/10.15252/embj.201797479.
Wu B, Peisley A, Tetrault D, Li Z, Egelman EH, Magor KE, et al. Molecular imprinting as a signal-activation mechanism of the viral RNA sensor RIG-I. Mol Cell. 2014;55(4):511–23. https://doi.org/10.1016/j.molcel.2014.06.010.
Fang R, Jiang Q, Zhou X, Wang C, Jiang Z. MAVS activates TBK1 and IKKε through TRAFs in NEMO dependent and independent manner. PLoS Pathog. 2017;13(11):e1006720. https://doi.org/10.1371/journal.ppat.1006720.
Liu S, Chen J, Cai X, Wu J, Chen X, Wu YT, et al. MAVS recruits multiple ubiquitin E3 ligases to activate antiviral signaling cascades. Elife. 2013;2:e00785. https://doi.org/10.7554/eLife.00785.
Dixit E, Kagan JC. Intracellular pathogen detection by RIG-I-like receptors. Adv Immunol. 2013;117:99–125. https://doi.org/10.1016/B978-0-12-410524-9.00004-9.
Ivashkiv LB, Donlin LT. Regulation of type I interferon responses. Nat Rev Immunol. 2014;14(1):36–49. https://doi.org/10.1038/nri3581.
Zhang Z, Kim T, Bao M, Facchinetti V, Jung SY, Ghaffari AA, et al. DDX1, DDX21, and DHX36 helicases form a complex with the adaptor molecule TRIF to sense dsRNA in dendritic cells. Immunity. 2011;34(6):866–78. https://doi.org/10.1016/j.immuni.2011.03.027.
Li J, Hu L, Liu Y, Huang L, Mu Y, Cai X, et al. DDX19A senses viral RNA and mediates NLRP3-dependent inflammasome activation. J Immunol. 2015;195(12):5732–49. https://doi.org/10.4049/jimmunol.1501606.
Liu Y, Lu N, Yuan B, Weng L, Wang F, Liu Y-J, et al. The interaction between the helicase DHX33 and IPS-1 as a novel pathway to sense double-stranded RNA and RNA viruses in myeloid dendritic cells. Cell Mol Immunol. 2014;11(1):49–57. https://doi.org/10.1038/cmi.2013.40.
Mitoma H, Hanabuchi S, Kim T, Bao M, Zhang Z, Sugimoto N, et al. The DHX33 RNA helicase senses cytosolic RNA and activates the NLRP3 inflammasome. Immunity. 2013;39(1):123–35. https://doi.org/10.1016/j.immuni.2013.07.001.
Miyashita M, Oshiumi H, Matsumoto M, Seya T. DDX60, a DEXD/H box helicase, is a novel antiviral factor promoting RIG-I-like receptor-mediated signaling. Mol Cell Biol. 2011;31(18):3802–19. https://doi.org/10.1128/MCB.01368-10.
Oshiumi H, Miyashita M, Okamoto M, Morioka Y, Okabe M, Matsumoto M, et al. DDX60 is involved in RIG-I-dependent and independent antiviral responses, and its function is attenuated by virus-induced EGFR activation. Cell Rep. 2015;11(8):1193–207. https://doi.org/10.1016/j.celrep.2015.04.047.
Hornung V, Hartmann R, Ablasser A, Hopfner K-P. OAS proteins and cGAS: unifying concepts in sensing and responding to cytosolic nucleic acids. Nat Rev Immunol. 2014;14(8):521–8. https://doi.org/10.1038/nri3719.
Nogimori T, Nishiura K, Kawashima S, Nagai T, Oishi Y, Hosoda N, et al. Dom34 mediates targeting of exogenous RNA in the antiviral OAS/RNase L pathway. Nucleic Acids Res. 2019;47(1):432–49. https://doi.org/10.1093/nar/gky1087.
Pichlmair A, Lassnig C, Eberle CA, Górna MW, Baumann CL, Burkard TR, et al. IFIT1 is an antiviral protein that recognizes 5′-triphosphate RNA. Nat Immunol. 2011;12(7):624–30. https://doi.org/10.1038/ni.2048.
Abbas YM, Pichlmair A, Górna MW, Furga GS, Nagar B. Structural basis for viral 5′-PPP-RNA recognition by human IFIT proteins. Nature. 2013;494(7435):60–4. https://doi.org/10.1038/nature11783.
Balachandran S, Roberts PC, Brown LE, Truong H, Pattnaik AK, Archer DR, et al. Essential role for the dsRNA-dependent protein kinase PKR in innate immunity to viral infection. Immunity. 2000;13(1):129–41. https://doi.org/10.1016/s1074-7613(00)00014-5.
Liu CX, Li X, Nan F, Jiang S, Gao X, Guo SK, et al. Structure and degradation of circular RNAs regulate PKR activation in innate immunity. Cell. 2019;177(4):865–80e21. https://doi.org/10.1016/j.cell.2019.03.046.
George CX, Samuel CE. Human RNA-specific adenosine deaminase ADAR1 transcripts possess alternative exon 1 structures that initiate from different promoters, one constitutively active and the other interferon inducible. Comparative Study. 1999;96(8):4621–6. https://doi.org/10.1073/pnas.96.8.4621.
Yang S, Deng P, Zhu Z, Zhu J, Wang G, Zhang L, et al. Adenosine deaminase acting on RNA 1 limits RIG-I RNA detection and suppresses IFN production responding to viral and endogenous RNAs. J Immunol. 2014;193(7):3436–45. https://doi.org/10.4049/jimmunol.1401136.
Toth AM, Li Z, Cattaneo R, Samuel CE. RNA-specific adenosine deaminase ADAR1 suppresses measles virus-induced apoptosis and activation of protein kinase PKR. J Biolumin Chemilumin. 2009;284(43):29350–6. https://doi.org/10.1074/jbc.M109.045146.
Yang P, An H, Liu X, Wen M, Zheng Y, Rui Y, et al. The cytosolic nucleic acid sensor LRRFIP1 mediates the production of type I interferon via a β-catenin-dependent pathway. Nat Immunol. 2010;11(6):487–94. https://doi.org/10.1038/ni.1876.
Schoggins JW, Wilson SJ, Panis M, Murphy MY, Jones CT, Bieniasz P, et al. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature. 2010;472(7344):481–5. https://doi.org/10.1038/nature09907.
Zhang X, Wu J, Du F, Xu H, Sun L, Chen Z, et al. The cytosolic DNA sensor cGAS forms an oligomeric complex with DNA and undergoes switch-like conformational changes in the activation loop. Cell Rep. 2014;6(3):421–30. https://doi.org/10.1016/j.celrep.2014.01.003.
Ablasser A, Chen ZJ. cGAS in action: expanding roles in immunity and inflammation. Science. 2019;363(6431):eaat8657. https://doi.org/10.1126/science.aat8657.
Barnett KC, Coronas-Serna JM, Zhou W, Ernandes MJ, Cao A, Kranzusch PJ, et al. Phosphoinositide interactions position cGAS at the plasma membrane to ensure efficient distinction between self- and viral DNA. Cell. 2019;176(6):1432–46e11. https://doi.org/10.1016/j.cell.2019.01.049.
Kim T, Pazhoor S, Bao M, Zhang Z, Hanabuchi S, Facchinetti V, et al. Aspartate-glutamate-alanine-histidine box motif (DEAH)/RNA helicase a helicases sense microbial DNA in human plasmacytoid dendritic cells. Proc Natl Acad Sci U S A. 2010;107(34):15181–6. https://doi.org/10.1073/pnas.1006539107.
Zhang Z, Yuan B, Bao M, Lu N, Kim T, Liu Y-J. The helicase DDX41 senses intracellular DNA mediated by the adaptor STING in dendritic cells. Nat Immunol. 2011;12(10):959–65. https://doi.org/10.1038/ni.2091.
Chiu YH, Macmillan JB, Chen ZJ. RNA polymerase III detects cytosolic DNA and induces type I interferons through the RIG-I pathway. Cell. 2009;138(3):576–91. https://doi.org/10.1016/j.cell.2009.06.015.
Zhang X, Brann TW, Zhou M, Yang J, Oguariri RM, Lidie KB, et al. Cutting edge: Ku70 is a novel cytosolic DNA sensor that induces type III rather than type I IFN. J Immunol. 2011;186(8):4541–5. https://doi.org/10.4049/jimmunol.1003389.
Ferguson BJ, Mansur DS, Peters NE, Ren H, Smith GL. DNA-PK is a DNA sensor for IRF-3-dependent innate immunity. Elife. 2012;1:e00047. https://doi.org/10.7554/eLife.00047.
Burleigh K, Maltbaek JH, Cambier S, Green R, Gale M Jr, James RC, et al. Human DNA-PK activates a STING-independent DNA sensing pathway. Sci Immunol. 2020;5(43):eaba4219. https://doi.org/10.1126/sciimmunol.aba4219.
Stetson DB, Ko JS, Heidmann T, Medzhitov R. Trex1 prevents cell-intrinsic initiation of autoimmunity. Cell. 2008;134(4):587–98. https://doi.org/10.1016/j.cell.2008.06.032.
Nan Y, Regaladomagdos AD, Stiggelbout B, Min LK, Lieberman J. The cytosolic exonuclease TREX1 inhibits the innate immune response to human immunodeficiency virus type 1. Nat Immunol. 2010;11(11):1005–13. https://doi.org/10.1038/ni.1941.
Martinon F, Hofmann K, Tschopp J. The pyrin domain: a possible member of the death domain-fold family implicated in apoptosis and inflammation. Curr Biol. 2001;11(4):R118–20. https://doi.org/10.1016/s0960-9822(01)00056-2.
Shaw N, Liu ZJ. Role of the HIN domain in regulation of innate immune responses. Mol Cell Biol. 2014;34(1):2–15. https://doi.org/10.1128/MCB.00857-13.
Cridland JA, Curley EZ, Wykes MN, Schroder K, Sweet MJ, Roberts TL, et al. The mammalian PYHIN gene family: phylogeny evolution and expression. BMC Evol Biol. 2012;12(1):140. https://doi.org/10.1186/1471-2148-12-140.
Jin T, Perry A, Jiang J, Smith P, Curry JA, Unterholzner L, et al. Structures of the HIN domain: DNA complexes reveal ligand binding and activation mechanisms of the AIM2 inflammasome and IFI16 receptor. Immunity. 2012;36(4):561–71. https://doi.org/10.1016/j.immuni.2012.02.014.
Morrone SR, Matyszewski M, Yu X, Delannoy M, Egelman EH, Sohn J. Assembly-driven activation of the AIM2 foreign-dsDNA sensor provides a polymerization template for downstream ASC. Nat Commun. 2015;6(1):7827. https://doi.org/10.1038/ncomms8827.
Hornung V, Ablasser A, Charrel-Dennis M, Bauernfeind F, Horvath G, Caffrey DR, et al. AIM2 recognizes cytosolic dsDNA and forms a caspase-1-activating inflammasome with ASC. Nature. 2009;458(7237):514–8. https://doi.org/10.1038/nature07725.
Unterholzner L, Keating SE, Baran M, Horan KA, Jensen SB, Sharma S, et al. IFI16 is an innate immune sensor for intracellular DNA. Nat Immunol. 2010;11(11):997–1004. https://doi.org/10.1038/ni.1932.
Yi Y-S, Jian J, Gonzalez-Gugel E, Shi Y-X, Tian Q, Fu W, et al. p204 is required for canonical lipopolysaccharide-induced TLR4 signaling in mice. EBioMedicine. 2018;29:78–91. https://doi.org/10.1016/j.ebiom.2018.02.012.
Chen W, Yu SX, Zhou FH, Zhang XJ, Gao WY, Li KY, et al. DNA sensor IFI204 contributes to host defense against staphylococcus aureus infection in mice. Front Immunol. 2019;10:474. https://doi.org/10.3389/fimmu.2019.00474.
Storek KM, Gertsvolf NA, Ohlson MB, Monack DM. cGAS and Ifi204 cooperate to produce type I IFNs in response to Francisella infection. J Immunol. 2015;194(7):3236–45. https://doi.org/10.4049/jimmunol.1402764.
Cao L, Ji Y, Zeng L, Liu Q, Zhang Z, Guo S, et al. P200 family protein IFI204 negatively regulates type I interferon responses by targeting IRF7 in nucleus. PLoS Pathog. 2019;15(10):e1008079. https://doi.org/10.1371/journal.ppat.1008079.
Li X, Deng M, Petrucelli AS, Zhu C, Mo J, Zhang L, et al. Viral DNA binding to NLRC3, an inhibitory nucleic acid sensor, unleashes STING, a cyclic dinucleotide receptor that activates type I interferon. Immunity. 2019;50(3):591–9e6. https://doi.org/10.1016/j.immuni.2019.02.009.
Kato H, Sato S, Yoneyama M, Yamamoto M, Uematsu S, Matsui K, et al. Cell type-specific involvement of RIG-I in antiviral response. Immunity. 2005;23(1):19–28. https://doi.org/10.1016/j.immuni.2005.04.010.
Loo YM, Fornek J, Crochet N, Bajwa G, Perwitasari O, Martinez-Sobrido L, et al. Distinct RIG-I and MDA5 signaling by RNA viruses in innate immunity. J Virol. 2008;82(1):335–45. https://doi.org/10.1128/JVI.01080-07.
Plumet S, Herschke F, Bourhis JM, Valentin H, Longhi S, Gerlier D. Cytosolic 5′-triphosphate ended viral leader transcript of measles virus as activator of the RIG I-mediated interferon response. PLoS One. 2007;2(3):e279. https://doi.org/10.1371/journal.pone.0000279.
Habjan M, Andersson I, Klingstrom J, Schumann M, Martin A, Zimmermann P, et al. Processing of genome 5′ termini as a strategy of negative-strand RNA viruses to avoid RIG-I-dependent interferon induction. PLoS One. 2008;3(4):e2032. https://doi.org/10.1371/journal.pone.0002032.
Hornung V, Ellegast J, Kim S, Brzozka K, Jung A, Kato H, et al. 5′-triphosphate RNA is the ligand for RIG-I. Science. 2006;314(5801):994–7. https://doi.org/10.1126/science.1132505.
Kato H, Takeuchi O, Sato S, Yoneyama M, Yamamoto M, Matsui K, et al. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature. 2006;441(7089):101–5. https://doi.org/10.1038/nature04734.
Sumpter R Jr, Loo YM, Foy E, Li K, Yoneyama M, Fujita T, et al. Regulating intracellular antiviral defense and permissiveness to hepatitis C virus RNA replication through a cellular RNA helicase. RIG-I J Virol. 2005;79(5):2689–99. https://doi.org/10.1128/JVI.79.5.2689-2699.2005.
Roth-Cross JK, Bender SJ, Weiss SR. Murine coronavirus mouse hepatitis virus is recognized by MDA5 and induces type I interferon in brain macrophages/microglia. J Virol. 2008;82(20):9829–38. https://doi.org/10.1128/JVI.01199-08.
Gitlin L, Barchet W, Gilfillan S, Cella M, Beutler B, Flavell RA, et al. Essential role of mda-5 in type I IFN responses to polyriboinosinic:polyribocytidylic acid and encephalomyocarditis picornavirus. Proc Natl Acad Sci U S A. 2006;103(22):8459–64. https://doi.org/10.1073/pnas.0603082103.
Pichlmair A, Schulz O, Tan CP, Rehwinkel J, Kato H, Takeuchi O, et al. Activation of MDA5 requires higher-order RNA structures generated during virus infection. J Virol. 2009;83(20):10761–9. https://doi.org/10.1128/JVI.00770-09.
Fredericksen BL, Keller BC, Fornek J, Katze MG, Gale M Jr. Establishment and maintenance of the innate antiviral response to West Nile virus involves both RIG-I and MDA5 signaling through IPS-1. J Virol. 2008;82(2):609–16. https://doi.org/10.1128/JVI.01305-07.
Parvatiyar K, Zhang Z, Teles RM, Ouyang S, Jiang Y, Iyer SS, et al. The helicase DDX41 recognizes the bacterial secondary messengers cyclic di-GMP and cyclic di-AMP to activate a type I interferon immune response. Nat Immunol. 2012;13(12):1155–61. https://doi.org/10.1038/ni.2460.
Kondo T, Kobayashi J, Saitoh T, Maruyama K, Ishii KJ, Barber GN, et al. DNA damage sensor MRE11 recognizes cytosolic double-stranded DNA and induces type I interferon by regulating STING trafficking. Proc Natl Acad Sci U S A. 2013;110(8):2969–74. https://doi.org/10.1073/pnas.1222694110.
Yang P, An H, Liu X, Wen M, Zheng Y, Rui Y, et al. The cytosolic nucleic acid sensor LRRFIP1 mediates the production of type I interferon via a beta-catenin-dependent pathway. Nat Immunol. 2010;11(6):487–94. https://doi.org/10.1038/ni.1876.
Marsh M, Helenius A. Virus entry: open sesame. Cell. 2006;124(4):729–40. https://doi.org/10.1016/j.cell.2006.02.007.
Roy A, Ghosh A, Kumar B, Chandran B. IFI16, a nuclear innate immune DNA sensor, mediates epigenetic silencing of herpesvirus genomes by its association with H3K9 methyltransferases SUV39H1 and GLP. Elife. 2019;8:e49500. https://doi.org/10.7554/eLife.49500.
Booiman T, Kootstra NA. Polymorphism in IFI16 affects CD4(+) T-cell counts in HIV-1 infection. Int J Immunogenet. 2014;41(6):518–20. https://doi.org/10.1111/iji.12157.
Ansari MA, Dutta S, Veettil MV, Dutta D, Iqbal J, Kumar B, et al. Herpesvirus genome recognition induced acetylation of nuclear IFI16 is essential for its cytoplasmic translocation, inflammasome and IFN-beta responses. PLoS Pathog. 2015;11(7):e1005019. https://doi.org/10.1371/journal.ppat.1005019.
Volkman HE, Cambier S, Gray EE, Stetson DB. Tight nuclear tethering of cGAS is essential for preventing autoreactivity. Elife. 2019;8:e47491. https://doi.org/10.7554/eLife.47491.
Boyer JA, Spangler CJ, Strauss JD, Cesmat AP, Liu P, McGinty RK, et al. Structural basis of nucleosome-dependent cGAS inhibition. Science. 2020;370(6515):450–4. https://doi.org/10.1126/science.abd0609.
Orzalli MH, Broekema NM, Diner BA, Hancks DC, Elde NC, Cristea IM, et al. cGAS-mediated stabilization of IFI16 promotes innate signaling during herpes simplex virus infection. Proc Natl Acad Sci U S A. 2015;112(14):E1773–81. https://doi.org/10.1073/pnas.1424637112.
Orzalli MH, Deluca NA, Knipe DM. Nuclear IFI16 induction of IRF-3 signaling during herpesviral infection and degradation of IFI16 by the viral ICP0 protein. Proc Natl Acad Sci U S A. 2012;109(44):E3008–17. https://doi.org/10.1073/pnas.1211302109.
Kerur N, Veettil MV, Sharma-Walia N, Bottero V, Sadagopan S, Otageri P, et al. IFI16 acts as a nuclear pathogen sensor to induce the inflammasome in response to Kaposi sarcoma-associated herpesvirus infection. Cell Host Microbe. 2011;9(5):363–75. https://doi.org/10.1016/j.chom.2011.04.008.
Wang L, Wen M, Cao X. Nuclear hnRNPA2B1 initiates and amplifies the innate immune response to DNA viruses. Science. 2019;365(6454):eaav0758. https://doi.org/10.1126/science.aav0758.
Cao L, Liu S, Li Y, Yang G, Luo Y, Li S, et al. The nuclear matrix protein SAFA surveils viral RNA and facilitates immunity by activating antiviral enhancers and super-enhancers. Cell Host Microbe. 2019;26(3):369–84e8. https://doi.org/10.1016/j.chom.2019.08.010.
Lahaye X, Gentili M, Silvin A, Conrad C, Picard L, Jouve M, et al. NONO detects the nuclear HIV capsid to promote cGAS-mediated innate immune activation. Cell. 2018;175(2):488–501. e22. https://doi.org/10.1016/j.cell.2018.08.062.
Diribarne G, Bensaude O. 7SK RNA, a non-coding RNA regulating P-TEFb, a general transcription factor. RNA Biol. 2009;6(2):122–8. https://doi.org/10.4161/rna.6.2.8115.
Michels AA, Nguyen VT, Fraldi A, Labas V, Edwards M, Bonnet F, et al. MAQ1 and 7SK RNA interact with CDK9/cyclin T complexes in a transcription-dependent manner. Mol Cell Biol. 2003;23(14):4859–69. https://doi.org/10.1128/mcb.23.14.4859-4869.2003.
Yik JH, Chen R, Nishimura R, Jennings JL, Link AJ, Zhou Q. Inhibition of P-TEFb (CDK9/cyclin T) kinase and RNA polymerase II transcription by the coordinated actions of HEXIM1 and 7SK snRNA. Mol Cell. 2003;12(4):971–82. https://doi.org/10.1016/s1097-2765(03)00388-5.
Yik JH, Chen R, Pezda AC, Zhou Q. Compensatory contributions of HEXIM1 and HEXIM2 in maintaining the balance of active and inactive positive transcription elongation factor b complexes for control of transcription. J Biolumin Chemilumin. 2005;280(16):16368–76. https://doi.org/10.1074/jbc.M500912200.
Michels AA, Fraldi A, Li Q, Adamson TE, Bonnet F, Nguyen VT, et al. Binding of the 7SK snRNA turns the HEXIM1 protein into a P-TEFb (CDK9/cyclin T) inhibitor. EMBO J. 2004;23(13):2608–19. https://doi.org/10.1038/sj.emboj.7600275.
Morchikh M, Cribier A, Raffel R, Amraoui S, Cau J, Severac D, et al. HEXIM1 and NEAT1 long non-coding RNA form a multi-subunit complex that regulates DNA-mediated innate immune response. Mol Cell. 2017;67(3):387–99e5. https://doi.org/10.1016/j.molcel.2017.06.020.
Uchida L, Espada-Murao LA, Takamatsu Y, Okamoto K, Hayasaka D, Yu F, et al. The dengue virus conceals double-stranded RNA in the intracellular membrane to escape from an interferon response. Sci Rep. 2014;4(1):7395. https://doi.org/10.1038/srep07395.
Wang H, Vaheri A, Weber F, Plyusnin A. Old World hantaviruses do not produce detectable amounts of dsRNA in infected cells and the 5′ termini of their genomic RNAs are monophosphorylated. J Gen Virol. 2011;92(5):1199–204. https://doi.org/10.1099/vir.0.029405-0.
Cardenas WB, Loo YM, Gale M, Hartman AL, Kimberlin CR, Martinez-Sobrido L, et al. Ebola virus VP35 protein binds double-stranded RNA and inhibits alpha/beta interferon production induced by RIG-I signaling. J Virol. 2006;80(11):5168–78. https://doi.org/10.1128/JVI.02199-05.
Xing J, Wang S, Lin R, Mossman KL, Zheng C. Herpes simplex virus 1 tegument protein US11 downmodulates the RLR signaling pathway via direct interaction with RIG-I and MDA-5. J Virol. 2012;86(7):3528–40. https://doi.org/10.1128/JVI.06713-11.
Gack MU, Albrecht RA, Urano T, Inn KS, Huang IC, Carnero E, et al. Influenza a virus NS1 targets the ubiquitin ligase TRIM25 to evade recognition by the host viral RNA sensor RIG-I. Cell Host Microbe. 2009;5(5):439–49. https://doi.org/10.1016/j.chom.2009.04.006.
Rajsbaum R, Albrecht RA, Wang MK, Maharaj NP, Versteeg GA, Nistal-Villán E, et al. Species-specific inhibition of RIG-I ubiquitination and IFN induction by the influenza a virus NS1 protein. PLoS Pathog. 2012;8(11):e1003059. https://doi.org/10.1371/journal.ppat.1003059.
Said EA, Tremblay N, Al-Balushi MS, Al-Jabri AA, Lamarre D. Viruses seen by our cells: the role of viral RNA sensors. J Immunol Res. 2018;2018:9480497–14. https://doi.org/10.1155/2018/9480497.
Childs K, Stock N, Ross C, Andrejeva J, Hilton L, Skinner M, et al. Mda-5, but not RIG-I, is a common target for paramyxovirus V proteins. Virology. 2007;359(1):190–200. https://doi.org/10.1016/j.virol.2006.09.023.
Davis ME, Wang MK, Rennick LJ, Full F, Gableske S, Mesman AW, et al. Antagonism of the phosphatase PP1 by the measles virus V protein is required for innate immune escape of MDA5. Cell Host Microbe. 2014;16(1):19–30. https://doi.org/10.1016/j.chom.2014.06.007.
Barral PM, Sarkar D, Fisher PB, Racaniello VR. RIG-I is cleaved during picornavirus infection. Virology. 2009;391(2):171–6. https://doi.org/10.1016/j.virol.2009.06.045.
Feng Q, Langereis MA, Lork M, Nguyen M, Hato SV, Lanke K, et al. Enterovirus 2Apro targets MDA5 and MAVS in infected cells. J Virol. 2014;88(6):3369–78. https://doi.org/10.1128/JVI.02712-13.
Wang B, Xi X, Lei X, Zhang X, Cui S, Wang J, et al. Enterovirus 71 protease 2Apro targets MAVS to inhibit anti-viral type I interferon responses. PLoS Pathog. 2013;9(3):e1003231. https://doi.org/10.1371/journal.ppat.1003231.
Mukherjee A, Morosky SA, Delorme-Axford E, Dybdahl-Sissoko N, Oberste MS, Wang T, et al. The coxsackievirus B 3Cpro protease cleaves MAVS and TRIF to attenuate host type I interferon and apoptotic signaling. PLoS Pathog. 2011;7(3):e1001311. https://doi.org/10.1371/journal.ppat.1001311.
Yang Y, Liang Y, Qu L, Chen Z, Yi M, Li K, et al. Disruption of innate immunity due to mitochondrial targeting of a picornaviral protease precursor. Proc Natl Acad U S A. 2007;104(17):7253–8. https://doi.org/10.1073/pnas.0611506104.
Drahos J, Racaniello VR. Cleavage of IPS-1 in cells infected with human rhinovirus. J Virol. 2009;83(22):11581–7. https://doi.org/10.1128/JVI.01490-09.
Varga ZT, Grant A, Manicassamy B, Palese P. Influenza virus protein PB1-F2 inhibits the induction of type I interferon by binding to MAVS and decreasing mitochondrial membrane potential. J Virol. 2012;86(16):8359–66. https://doi.org/10.1128/JVI.01122-12.
Otsuka M, Kato N, Moriyama M, Taniguchi H, Omata M. Interaction between the HCV NS3 protein and the host TBK1 protein leads to inhibition of cellular antiviral response. Hepatology. 2005;41(5):1004–12. https://doi.org/10.1002/hep.20666.
Diperna G, Stack J, Bowie AG, Boyd A, Kotwal G, Zhang Z, et al. Poxvirus protein N1L targets the I-κB kinase complex, inhibits signaling to NF-κB by the tumor necrosis factor superfamily of receptors, and inhibits NF-κB and IRF3 signaling by toll-like receptors. J Biolumin Chemilumin. 2004;279(35):36570–8. https://doi.org/10.1074/jbc.M400567200.
Wu JJ, Li W, Shao Y, Avey D, Fu B, Gillen J, et al. Inhibition of cGAS DNA sensing by a herpesvirus virion protein. Cell Host Microbe. 2015;18(3):333–44. https://doi.org/10.1016/j.chom.2015.07.015.
Aguirre S, Luthra P, Sanchez-Aparicio MT, Maestre AM, Patel J, Lamothe F, et al. Dengue virus NS2B protein targets cGAS for degradation and prevents mitochondrial DNA sensing during infection. Nat Microbiol. 2017;2(5):17037. https://doi.org/10.1038/nmicrobiol.2017.37.
Cuchet-Lourenco D, Anderson G, Sloan E, Orr A, Everett RD. The viral ubiquitin ligase ICP0 is neither sufficient nor necessary for degradation of the cellular DNA sensor IFI16 during herpes simplex virus 1 infection. J Virol. 2013;87(24):13422–32. https://doi.org/10.1128/JVI.02474-13.
Peters NE, Ferguson BJ, Mazzon M, Fahy AS, Krysztofinska E, Arribas-Bosacoma R, et al. A mechanism for the inhibition of DNA-PK-mediated DNA sensing by a virus. PLoS Pathog. 2013;9(10):e1003649. https://doi.org/10.1371/journal.ppat.1003649.
Scutts SR, Ember SW, Ren H, Ye C, Lovejoy CA, Mazzon M, et al. DNA-PK is targeted by multiple vaccinia virus proteins to inhibit DNA sensing. Cell Rep. 2018;25(7):1953–65e4. https://doi.org/10.1016/j.celrep.2018.10.034.
Aguirre S, Maestre AM, Pagni S, Patel JR, Savage T, Gutman D, et al. DENV inhibits type I IFN production in infected cells by cleaving human STING. PLoS Pathog. 2012;8(10):e1002934. https://doi.org/10.1371/journal.ppat.1002934.
Acknowledgments
The authors apologize to all colleagues whose important contributions could not be discussed and cited owing to space constraints.
Code availability in declaration
Not applicable
Funding
Current research in the You laboratory is supported by National Natural Science Foundation of China (31570891 and 31872736), the National Key Research and Development Program of China (Grant #2016YFA0500302) and the Zhuhai Science and Technology Innovation Bureau (ZH22036302200063PWC to Z. Yin).
Author information
Authors and Affiliations
Contributions
writing—original draft preparation, S.L. Z.Z. M.K. Y.Z.; writing—review and editing, F.Y. S.L. L.C. L.C. Z.Y.; supervision, F.Y. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no conflicts of interest with the contents of this article.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, S., Cao, L., Zhang, Z. et al. Cytosolic and nuclear recognition of virus and viral evasion. Mol Biomed 2, 30 (2021). https://doi.org/10.1186/s43556-021-00046-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s43556-021-00046-z