Abstract
The development of all-optical techniques and analytical tools to visualize and manipulate the activity of identified neuronal ensembles enables the characterization of causal relations between neuronal activity and behavioral states. In this chapter, we review the implementation of simultaneous two-photon imaging and holographic optogenetics in conjunction with population analytical tools to identify and reactivate neuronal ensembles to control a visual-guided behavior.
You have full access to this open access chapter, Download protocol PDF
Similar content being viewed by others
Key words
- Two-photon imaging
- Two-photon optogenetics
- Holographic microscope
- Neuronal ensembles
- Pattern completion
- Population vectors
- Control of behavior
1 Introduction
One of the main questions in modern neuroscience is how the activity of identified neuronal populations generates behavioral and mental states [1]. Recent advances in optical techniques that allow the simultaneous recording and manipulation of neurons with single-cell resolution [2, 3] combined with population analytical tools [4,5,6,7,8] suggest that neuronal ensembles are units of brain computation [9, 10]. Neuronal ensembles are groups of neurons with coordinated activity that may underlie sensations, perceptions, emotions, and memories. In order to prove causality between the activity of specific neuronal ensembles and learned behaviors, it is becoming clear that the ability to manipulate and record hundreds of neurons simultaneously needs to be guided by analytical tools that allow the identification of neurons that could have a deterministic impact in brain states. Holographic two-photon imaging [11], two-photon optogenetics [12], and analytical tools [13] offer the possibility to target neuronal ensembles to control behavioral states. The demonstration of the causal relation between neuronal ensemble activity and behavior has been achieved recently in different brain areas [4, 14,15,16,17,18], paving the pathway to design more sophisticated experiments to understand complex mental states in health and disease [19].
The optimal design of all-optical experiments to control learned behaviors with single-cell resolution requires the simultaneous reading and writing of neuronal activity. This could be achieved in many ways [20], but in this chapter we will focus mainly on scanning and parallel optical techniques guided by analytical tools. We describe the implementation of scanning two-photon imaging and parallel two-photon optogenetics using a spatial light modulator (SLM). We also describe the main concepts necessary to identify and target neurons with pattern completion capability [6, 21] that can recall neuronal ensembles related to a visually guided behavior [4].
2 Implementation of Simultaneous Two-Photon Imaging and Two-Photon Optogenetics
2.1 Background
2.1.1 Light-Sensitive Sensors and Actuators of Neuronal Activity
Optical methods are powerful to record and manipulate neuronal activity at single-cell resolution across a large population—a key requirement to investigate neuronal ensembles. Comparing with electrophysiology, optical methods can simultaneously sample and target a large group of neurons with high spatial specificity in a noninvasive manner and can be conveniently implemented for in vivo studies. The cornerstone of optical methods is the optical microscope, which uses light to record and manipulate the neurons that are illuminated. As the neurons in the brain do not typically respond to light, they can be loaded or transfected with light-sensitive sensors and actuators that can respectively report and manipulate their neuronal activity upon light illumination. Using these neuronal activity sensors and actuators, an optical microscope can simultaneously read and write neuronal activity across a large field of view with high spatial specificity.
Calcium [22,23,24,25,26] and voltage indicators [27, 28] are two commonly used sensors for neuronal activity. These indicators are embedded with fluorophores, which can absorb the illumination or excitation light, and emit fluorescence. The efficiency of this light-absorption, fluorescence-emission process is modulated by the intracellular calcium concentration and membrane potential respectively in calcium indicators and voltage indicators (See also Chaps. 1 and 2). Thus, the neuronal activity can be deduced from the time-lapse recording of the fluorescence in individual cells. The commonly used calcium indicators include GCaMP6 [26], jGCaMP7 [29], jRGECO [30], etc. Compared with calcium indicators, voltage indicators are less mature with smaller signal-to-noise ratio and prone to photo-bleach though much progress has been made in the past years [31] (Chap. 2). The counterparts of these activity sensors are actuators, which could be chemical-based neurotransmitter cage-compounds [32, 33] or opsins [34, 35] serving as light-sensitive ion channels. When absorbing the light, the cage-compounds could release the neurotransmitters (uncaging) and the opsins could open or close the ion channels (optogenetics) and thus control the membrane potential. This mechanism thus allows light control of neuronal activity in individual cells.
2.1.2 One-Photon and Multiphoton Excitation
Depending on how light is interacting with the light-sensitive sensors and actuators, optical methods can be classified into two categories: one-photon and multiphoton (i.e., two-photon [36, 37] or three-photon [38, 39]). In one-photon case, the interaction is linear. The fluorescence emission rate of the sensors and the actuation strength of the actuators are proportional to the intensity of the excitation light before saturation. While one-photon excitation is straightforward, it lacks spatial specificity in 3D. As the excitation would form a double cone-shape pattern along the axial direction (Fig. 1), all neurons with the sensors or actuators within the double cone may absorb the light. It is thus very challenging to just excite a single neuron in a 3D volume. Furthermore, in terms of imaging, the out-of-focus fluorescence emission typically becomes background on the image captured at the focal plane and thus reduces the image signal-to-noise ratio. As the light-absorption and fluorescence-emission cycles create phototoxicity, one-photon excitation pays a high price to image the focal plane by creating phototoxicity within the entire double cone volume. Multiphoton excitation can greatly overcome these limitations, as the interaction between the light and the sensors or actuators is nonlinear. In the two-photon case, the excitation effect is proportional to the intensity square of the excitation light before saturation, leading to a strong gradient of the excitation effect along the optical axis in the double cone. Thus, it is feasible to control the incident light so only the light intensity at the focal point (i.e., the tip of the double cone) is strong enough to excite the neuronal sensors or actuators (Fig. 1). This greatly improves the spatial specificity or resolution and reduces the phototoxicity in out-of-focus region. Another advantage of two-photon excitation is that the excitation light has a longer wavelength, which reduces the light scattering effect in scattering brain tissues. Thus two-photon excitation can penetrate much deeper in the brain compared with one-photon excitation. The drawback of two-photon excitation is that much more laser power is required due to a lower efficiency of multiphoton absorption. As the excitation light eventually turns into heat, more heat will be generated inside the brain. This is typically not a concern as in a typical imaging experiment, the laser power used is less than the brain damage threshold [40], even at very deep layers. In the past two decades, two-photon microscopes have become one of the workhorses in neuroscience. Recently, three-photon excitation has been successfully demonstrated for both calcium imaging [39] and optogenetics [41]. Comparing with two-photon excitation, three-photon excitation has an even higher nonlinearity and could further increase spatial specificity and penetration depth, though the laser power should be managed so it will not exceed the brain damage threshold.
Comparison between one-photon and two-photon excitation. In one-photon excitation (a), a double cone is formed by focusing the visible excitation light (indicated by blue color); fluorophores (green) in the entire double cone could be excited. In two-photon excitation (b), while a double cone is still formed by the focus of infrared excitation light (indicated by red color), the fluorescence generation (green) is localized at the vicinity of the focal region. Reprinted with permission from [101], Springer Nature. Experimental illustration of the one-photon and two-photon excitation (0.16 NA for both cases) is shown in the bottom panel. (Reprinted with permission from Ref. [102], Springer Nature)
2.1.3 Basic Setup of Multiphoton Microscope
Since the multiphoton absorption rate is generally low, and it has a nonlinear relationship with the light intensity, a femtosecond laser is required for multiphoton excitation. The femtosecond laser delivers a periodic pulse train, typically with a repetition rate of 1~80 MHz. The temporal width of a pulse is below 300 fs, yielding a very high instantaneous peak power and thus a high multiphoton absorption rate. Unlike one-photon excitation where a large area or volume of sample could be simultaneously illuminated and their fluorescence could be detected through a camera in the case of imaging, the illumination of multiphoton light is typically through a rapid scanning of the laser beam on the sample. In multiphoton imaging, a single pixel detector (versus pixel array in a camera) is used to record the emitted fluorescence (see also Chap. 2). By correlating the temporal signal from the detector and the scanning trajectory, the image can be built. The microscope setup (Fig. 2a) typically includes a raster lateral (xy) scanning system composed of a galvanometer scanner and a resonant scanner (30~60 Hz frame rate), or two galvanometer scanners (4~10 Hz frame rate). For volumetric imaging, an axial (z) scanning system is implemented by adding a piezo-electric controller on the objective lens or inserting axial focusing devices, such as electrically tunable lens [42], spatial light modulator [43], or remote focusing unit [44,45,46] (~ ms focus switching time). Different axial planes are sequentially scanned (Fig. 2b). While this configuration of lateral and axial scanning is straightforward, it may not be the most efficient as it blindly samples the brain tissue. There could be a large volume of “empty” extracellular space without neurons, and among all the neurons, typically only a subset of them are labeled with the calcium indicators. Random access scanning techniques [47,48,49,50,51] can overcome this issue. The laser focus spot can jump rapidly between different regions in the sample in 3D (<20 μs transition time between different regions) (Fig. 2c). This is enabled by acousto-optic deflectors (AODs). The challenges of this technique are to overcome the spatial and temporal distortion of the ultrashort laser pulses, caused by the angular dispersion from the AODs’ phase grating and the group delay dispersion of the AOD crystals, respectively. These limit the imaging field of view. While these distortions can be compensated [51], the optical setup is complex. AODs also have a limitation of high insertion loss. Recently, many advanced scanning mechanisms including beam multiplexing [43, 52,53,54,55] are proposed and demonstrated to increase the imaging throughput. For extensive reviews, see [56,57,58,59].
Basic setup of two-photon laser scanning microscope. (a) Schematics of a typical two-photon laser scanning microscope. The Pockels cell is used to modulate the laser light intensity. The xy-scan system and z-scan system are used to scan the laser focal spot laterally and axially in the sample, respectively. The dichroic mirror transmits the infrared excitation light and reflects the visible fluorescent light to the photomultiplier tube (PMT). (b) Schematics of the typical scanning trajectory in volumetric imaging. (c) Schematics of an exemplary scanning trajectory using random access scanners. The green regions indicate neurons of interest. (Reprinted with permission from Ref. [56], Springer Nature)
The setup of multiphoton photostimulation is similar as multiphoton imaging, but without the detection module. The laser spot on the sample is typically controlled by a pair of galvanometer mirrors, and raster or spirally scanned across each target neuron (1~100 ms) before jumping to another neuron. Alternatively, the combination of beam shaping and temporal focusing techniques [60,61,62,63,64] can be implemented so that a disk pattern with tight axial confinement can be projected to the entire neuron. The scanners no longer perform raster or spiral scanning, but direct the disk excitation pattern to different neurons sequentially [3]. Using the holographic approach, illumination patterns could be simultaneously projected to multiple targeted neurons, so the scanners are no longer required [62,63,64,65,66]. We will discuss this topic in depth, as well as holographic illumination technique that can simultaneously photostimulate multiple neurons, in the next section.
In the following, we focus on our implementation of a microscope that can simultaneously perform calcium imaging to record neuronal activity and optogenetics to manipulate neuronal activity.
2.2 Simultaneous Two-Photon Imaging and Two-Photon Optogenetics
2.2.1 Overall Consideration
An all-optical method [20] refers to simultaneously recording and manipulating the neuronal activity through light. Here, we describe the combination of two-photon calcium imaging and two-photon optogenetics in a microscope. To flexibly image and manipulate neuronal ensembles, it is desired to have the following features in the microscope:
-
(i)
Calcium imaging and optogenetics should be independently controlled and performed.
-
(ii)
Two-photon imaging could image neuronal activity over a large neuronal population with cellular resolution and high temporal resolution (4~30 Hz for calcium imaging).
-
(iii)
Two-photon optogenetics could simultaneously photostimulate multiple neurons with cellular resolution and high temporal precision (sub-millisecond to milliseconds).
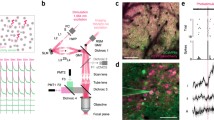
To simultaneously, while independently, perform calcium imaging and optogenetics, the microscope should have two independent beam paths, respectively, for calcium imaging and optogenetics (Fig. 3a) [2, 3, 66,67,68,69]. These two beam paths then merge together before the light is directed into the brain tissue. Each beam path should be implemented with an individual femtosecond laser with a different wavelength, which corresponds to the central excitation spectrum of the calcium indicators and opsins. It is important that they have a different excitation spectrum, so as to prevent crosstalk between imaging and photostimulation (see also Chaps. 1, 2, and 5). In other words, when performing calcium imaging, the imaging laser should not excite the opsin; when performing optogenetics, the photostimulation laser should not excite the fluorescence of the calcium indicator. There are two common options to choose the calcium indicator and opsin pair: green calcium indicators such as GCaMP [26, 29] with red-shifted opsins such as C1V1 [70], or red-shifted calcium indicators such as jRGECO [30] with blue-activated opsins such as ChR2 [34]. While both of these pairs have minimal overlap in the excitation spectrum, it is critical to keep the imaging laser power low and perform control experiments to ensure that the imaging laser does not increase the spiking rate of the neurons (see Note 1). In case the photostimulation laser creates fluorescence artifacts, the imaging data should be processed to eliminate the artifacts. For a successful experiment, it is also important to have a high co-expression rate of the calcium indicators and opsins on a large neuronal population. The cooperation of both constructs into a single virus could be a promising approach [14, 71]. An example of co-expression of GCaMP6s and C1V1 using two different viruses is shown in Fig. 3c.
Simultaneous two-photon calcium imaging and two-photon holographic optogenetics. (a) Two-photon microscope setup with two independent beam paths for calcium imaging and photostimulation. An electrically tunable lens is equipped in the imaging path so the focal plane can be rapidly switched for high-speed volumetric imaging. The dichroic mirror 3 is used to spectrally separate the fluorescence into green and red channel. In the photostimulation path, a half-wave plate is placed before the spatial light modulator to align the polarization of the laser to the active axis of the spatial light modulator. The spatial light modulator then creates a hologram in the sample to photostimulate the regions of interest (e.g., a group of neurons). A pair of relay lens (4f system) is used to transfer the light field to a set of galvanometer mirrors which can spirally scan the holographic pattern. At the intermediate plane of this pair of relay lens, a zeroth-order beam block is used to block the residue light that is not modulated by the spatial light modulator. The imaging laser and photostimulation laser have different wavelengths, and they are combined through the dichroic mirror 1 before being delivered to the sample through the scan lens, tube lens, and objective lens. HWP half-wave plate, PMT photomultiplier tube. (b) Schematics for simultaneous volumetric calcium imaging and 3D holographic patterned photostimulation in mouse cortex. (c) An exemplary field of view showing neurons co-expressing GCaMP6s (green) and C1V1-mCherry (magenta). (Reprinted and adapted from Ref. [69])
To characterize neuronal ensemble activity, it is necessary to image a large population of neurons, typically on the order of hundreds. For visual cortex, a minimal field of view of 200 × 200 μm2 is necessary, though 400 × 400 ~ 600 × 600 μm2 would be more desirable. Using low magnification objective lens, the field of view can go above 1 mm2. Furthermore, volumetric imaging can be performed to image neuronal ensembles in 3D and across different functional layers (Fig. 3a, b). Due to the sequential scanning nature of two-photon microscopy, there is a tradeoff between the field of view and spatiotemporal resolution. Thus, while it is desirable to increase the field of view so as to image more neurons, one should maintain cellular resolution with a good signal-to-noise ratio (i.e., able to distinguish the calcium transients), as well as a frame rate of at least 4–10 Hz. In the setup shown in Fig. 3a, we use a galvanometer and a resonant scanner pair for lateral scanning, and an electrically tunable lens for fast focal plane switching. Volumetric imaging of 3 planes could be achieved at a volume rate >6 Hz, with a field of view ~500 × 500 μm2 per plane [69].
A typical experiment starts with two-photon calcium imaging on the brain tissue. The animal is head-fixed and stabilized on the microscope stage and could perform behavior tasks. The spatial footprint as well as the neuronal activity of individual neurons within the image field of view can then be extracted from the recording. The neuronal ensemble can be identified from the activity pattern (see Subheading 3). Depending on the applications, the users may choose a specific population of neurons to perform optogenetics experiments. It is thus desirable to perform photostimulation on multiple neurons simultaneously. Two-photon holographic illumination enables photostimulating a group of user-selected neurons [2, 12, 14, 66, 68, 69, 72, 73]. We discuss the topic in the following section.
2.2.2 Holographic Illumination
Holographic illumination refers to projecting to the brain tissue a computer-generated holographic pattern. Neurons falling within this pattern can then be photostimulated. Computer-generated hologram is a light field of an arbitrary shape in 3D in an imaging volume, and this shape can be dynamically controlled and changed through a spatial light modulator (Fig. 4a). In the simplest imaging system, which contains a single lens, to generate a hologram in the imaging space (i.e., around the front focal plane of the lens), one can spatially modulate the light field at the back focal plane of the lens. This light field then propagates through the lens, and coherently interferes at the front focal plane and forms the holographic pattern (Fig. 4a). As the light field at the front focal plane and back focal plane of a lens forms a Fourier transform relationship, one can calculate how the light field should be spatially modulated at the back focal plane based on the desired pattern at the front focal plane (Fig. 4b, c). The typical spatial light modulator is based on liquid crystals and is configured to only modulate the phase of the light field. A Gerchberg-Saxton algorithm (Fig. 5) [74], which is an iterative approach, can be used to calculate the phase pattern on the spatial light modulator, given the amplitude of the desired pattern in the imaging space and the amplitude of the light field incident on the spatial light modulator. If the holographic pattern only contains a group of points with different weights, a superposition algorithm [55, 75], which is essentially a single iteration of the Gerchberg-Saxton algorithm, can be used to calculate the phase pattern (Table 1).
Computer-generated hologram. (a) Principle of computer-generated hologram. The collimated laser beam is incident onto the spatial light modulator (SLM), which spatially modulates the wavefront of the light. The light field then propagates through a lens and forms the desired 3D image in the imaging domain through interference. f, focal length of the lens. (b) By modulating the spatial phase profile at the back focal plane of the object lens, different focal spot patterns can be formed in the imaging domain. (Reprinted with permission from Ref. [103], Elsevier). (c) Example of a two-photon SLM hologram. The four panels illustrate the binary target image, the spatial light modulator phase hologram generated by Gerchberg-Saxton algorithm, the squared image (to mimic two-photon excitation) of the projected pattern back calculated from the phase hologram, and the experimentally measured two-photon fluorescence image generated by the SLM. A stylized picture of Cajal is used as a target image. (Reprinted from Ref. [11], Frontiers)
In two-photon microscopes, we could not directly modulate the phase at the back focal plane of the objective lens, as the back focal plane is typically inside the objective lens housing. Relay lens pairs (4f system) can create the conjugate planes of the back focal plane of the objective lens, and the spatial light modulator can be placed in one of those conjugate planes (Fig. 3a). Furthermore, as there is residue light that is not modulated by the spatial light modulator (termed zeroth order beam), it will create a strong focus at the imaging space (see Note 2). The relay lens pairs help to resolve this issue as a small beam block can be placed at the intermediate plane (which is conjugate to the focal plane at the imaging space) to block the zeroth order beam (Fig. 3a). Using the holographic illumination, various patterns can be generated in the imaging space (Fig. 4b, c).
Holographic illumination essentially spatially multiplexes the excitation beams, and allows multiple neurons being photostimulated simultaneously. However, this comes with an increase of laser power on the brain tissue. To alleviate this issue, a laser with a lower repetition rate can be used. When the pulse repetition rate is reduced, the energy in each laser pulse is increased, while keeping the overall average power the same. As the two-photon excitation efficiency is proportional to the square of the laser peak power, the increase of excitation efficiency due to the increase in pulse energy outweighs its reduction due to the reduced number of pulses per unit time. Thus, using a laser with lower repetition rate, a higher overall excitation efficiency can be achieved while keeping the same average power. In other words, to achieve the same excitation efficiency, the average laser power can be reduced. In the holographic photostimulation system described, we used a laser with a repetition rate of 1 MHz [69]. This reduces the overall laser power by 80 times compared with a commonly used 80 MHz femtosecond laser.
Due to the chromatic dispersion and spatial discretization of the SLM pixels, the SLM has a spatially varying diffraction efficiency [75]. The diffraction efficiency drops as the deflection angle increases, limiting the addressable 3D field of view of SLM. To alleviate this effect, the diffraction efficiency can be first measured, and then compensated in the Gerchberg-Saxton algorithm or through the weight factor Ai (Table 1) in the superposition algorithm [55, 75]. The laser intensity across the field of view can then be made uniform. By using an XY galvanometer set to provide a lateral offset to the centroid of the SLM’s addressable field of view, the effective lateral field of view can be further extended. While neurons across this enlarged field of view cannot be targeted simultaneously, groups of neurons located at different sub-fields can be targeted by switching the offset of the XY galvanometer and the phase pattern on the SLM [76].
2.2.3 Spiral Scan Versus Scanless Approach in Holographic Photostimulation
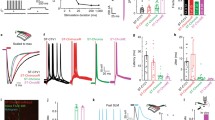
As mentioned in Subheading 2.1, photostimulation can be performed by raster or spirally scanning the laser focal spot across the neuron, or by projecting a disk pattern which matches the morphology of the neuron so the entire neuron can be stimulated at once. The same applies in holographic photostimulation. In the first approach, a group of focal spots are created in the hologram, and each focal spot lies on the centroid of the individual targeted neuron [2, 12, 69]. A set of galvanometer scanners then spirally scan the entire group of focal points (with certain repetitions), so each focal spot spirally scans across the corresponding targeted neuron. In the second approach, a group of disk patterns are created in the hologram and projected to the brain tissue [72, 77, 78]. Each disk spatially overlaps with a targeted neuron. Since the hologram already generates multiple disks for all the target neurons, this method can work without scanners, and thus this is a scanless approach. In two-photon excitation, these two approaches have their own advantages and limitations. The spiral scan approach spatially concentrates the laser intensity into focal spots; as two-photon excitation effect is proportional to light intensity square, this approach has a high excitation efficiency. As the entire neuron is not photostimulated at the same time, to reach the threshold of evoking action potentials, it relies on the accumulation of the excitation effect along the spiral trajectory. The decay constant of the opsin kinetics (tau-off) should thus be longer than the duration of each spiral (i.e., the opsin channels can stay open during the one single spiral scan, which could be less than 1 ms); otherwise the excitation effect cannot be accumulated. The scanless approach, on the other hand, disperses the light intensity across the entire neuron, and thus it requires a higher average power to reach the same excitation strength as the spiral scan approach. However, as the entire neuron is being stimulated at once, it does not pose a limitation on the kinetics of the opsin. It is also expected that the jitter of the delay between the onset of photostimulation and onset of action potential is smaller. As the opsin decay constant (tau-off) is typically larger than the duration of a single spiral scan (which can be <1 ms), the scanning approach is in favor as it takes lower laser power to photoactivate the neurons. In our experiments, we used C1V1 as the opsin. Using spiral scan, it takes about 1.8× less power to photoactivate the neurons than the scanless approach, with the same photostimulation duration (Fig. 6) [69]. For opsins with faster kinetics, a faster spiral scan or the scanless approach may be preferred.
Comparison between spiral scan and scanless holographic approaches for photostimulation. In the scanning approach, the laser spot is spirally scanned over the cell body; in the scanless approach, a disk pattern is generated by the SLM, covering the entire cell body at once. (a) Photostimulation triggered calcium response of a targeted neuron in vivo at mouse layer 2/3 of V1, for different stimulation modalities. For each modality, the multiplication of stimulation duration and the square of the laser power was kept constant over four different stimulation durations. The average response traces are plotted over those from the individual trials. (b) ΔF/F response of individual neurons on different photostimulation conditions (layer 2/3 of V1, over a depth of 100 ~ 270 μm from pial surface; one-way ANOVA test). For each neuron and each stimulation duration, the laser power used in the scanless disk modality is 1 and 1.8 times relative to that in the spiral scan. For each neuron and each modality, the multiplication of the stimulation duration and the square of the laser power was kept constant over four different stimulation durations. (c) Boxplot summarizing the statistics in (b). The central mark indicates the median, and the bottom and top edges of the box indicate the 25th and 75th percentiles, respectively. The whiskers extend to the most extreme data points (99.3% coverage if the data are normal distributed) not considered outliers, and the outliers are plotted individually using the “+” symbol. In this experiment, the mice were transfected with GCaMP6f and C1V1-mCherry. Repetition rate of the photostimulation laser is 1 MHz. The spiral scan consists of 50 rotations to cover the neuronal cell body, and the scanning speed is adjusted to make different stimulation durations. (Reprinted with permission from Ref. [69])
2.2.4 Detailed Implementation of the Microscope for Simultaneous Two-Photon Imaging and Two-Photon Holographic Photostimulation
In this section, we explain the details of how the microscope can be constructed with two beam paths for two-photon imaging and two-photon holographic photostimulation (Fig. 3a). We aim to help the readers understand the inside of a commercial microscope system and meanwhile provide basic guidelines for those who want to home-build the microscope. Here, we use GCaMP6 as calcium indicators, and C1V1-mCherry as the red-shifted opsin. The mCherry can be used to indicate if the opsin is expressed in each neuron. We combine the holographic illumination with spiral scan approach to minimize the laser power onto the brain.
Setup of the Two-Photon Imaging Path
-
(1.1) The imaging laser is typically a wavelength tunable Ti:Sapphire laser with a repetition rate of 80 MHz, and a pulse width of 70~140 fs. For GCaMP imaging, the wavelength can be set to 920~940 nm. A Pockels cell is set up after the laser to modulate the laser power. An optional pulse compressor could be set up after the Pockels cell to optimize the pulse width at the sample.
-
(1.2) An optional electrically tunable lens could be set up as an axial focal control device. A set of XY galvanometer mirrors, or a resonant scanner paired with a galvanometer mirror, can be set up to raster scan the laser focal spot on the sample. The electrically tunable lens and the scanning mirrors are located at the conjugate plane of the back focal plane of the objective lens. 4f systems are used to relay these planes and to magnify the beam size so that the beam can illuminate a large portion of the aperture of the electrically tunable lens and scanning mirrors.
-
(1.3) A dichroic mirror is set up to combine the imaging path and photostimulation path and direct the beam to scan lens and tube lens, which forms a 4f system to magnify the beam at the back focal plane of the objective lens. This beam size should be optimized for specific objective lens so as to achieve a good excitation numerical aperture (NA) for cellular resolution. For population imaging, the objective lens is typically chosen to be 16~40×, and the excitation NA is typically 0.45~0.6. The collection NA should be as large as possible so as much as the emitted fluorescence could be collected.
-
(1.4) A dichroic mirror is set up before the objective lens to transmit the infrared excitation light and reflect the fluorescence emission to photomultiplier tubes (PMTs). A dichroic mirror can further separate the fluorescence spectrum into different channels (typically green and red).
Setup of the Two-Photon Photostimulation Path
-
(2.1) The photostimulation laser can be a high repetition rate fiber laser or a low repetition rate pulse-amplified laser (including optical parametric amplified laser). The latter laser is preferred for holographic photostimulation as its higher pulse energy allows lowering the average photostimulation power per cell, thus enabling photostimulating a large number of neurons with the overall available power budget. For red-shifted opsin such as C1V1, the preferred laser wavelength is 1040~1080 nm. Similar as the imaging path, a Pockels cell and an optional pulse compressor are set up after the laser.
-
(2.2) A spatial light modulator is set up at a conjugate plane to the back focal plane of the objective lens. Following the Pockels cell, a half-wave plate is set up to rotate the polarization of the laser so the light polarization aligns with the active axis of the spatial light modulator. A 4f system is used to magnify the beam size so it fills up the active region of the spatial light modulator. A 4f system is placed after the spatial light modulator with a zeroth-order beam block set up at the intermediate imaging plane. After this 4f system, a set of XY galvanometer mirrors are set up, which is also at the conjugate plane of the back focal plane of the objective lens. Here, the XY galvanometer can not only perform spiral scanning but also provide a lateral offset to the centroid of the SLM’s addressable field of view, thus effectively enlarging the overall field of view, as discussed in earlier sessions. Thus, the XY galvanometer is preferred over galvo/resonant scanner pair, as the latter cannot provide a static beam shift in the resonant scanner’s scanning direction.
-
(2.3) The photostimulation path then combines with the imaging path at the dichroic mirror and is then directed to the sample through the scan lens, tube lens, and objective lens.
Control Electronics and Software
-
(3.1) The electronic control system typically includes analog inputs/outputs to control the Pockels cells, electrically tunable lens, spatial light modulator, and scanners in both beam paths, as well as high-speed digitizers to record the imaging data from the PMT. For home-built microscopes, ScanImage [79] is one commonly used control software.
-
(3.2) The phase pattern on the spatial light modulator can be calculated through a superposition algorithm or a Gerchberg-Saxton algorithm as detailed in Subheading 2.2.2 so a user-defined excitation pattern can be projected to the sample. An open source control software can be found in [80].
Coordinate Calibration
-
(4.1) It is critical to register the photostimulation beam’s target coordinate with the imaging laser’s image coordinate. An autofluorescent plastic slide can serve as the sample, and a 2D holographic pattern can be projected to burn spots on the surface of the autofluorescent plastic slide. The imaging laser can then visualize the burned spots. This can also calibrate the axial focus offset between the two beam paths. An affine transformation can be extracted to map the coordinates between the hologram generation algorithm and the actual imaging system. This can be repeated for a few defocusing depths set in the spatial light modulator, and a linear interpolation of the mapping can be applied for the depths in between.
-
(4.2) Due to the chromatic dispersion and finite pixel size of SLM, the SLM’s beam steering efficiency, also called diffraction efficiency, drops with larger angle, leading to a lower beam power for targets further away from the center field of view (in xy), and nominal focus (in z). This efficiency drop can be analytically calculated [55, 75] or experimentally measured by raster scanning the photostimulation beam on the autofluorescence slide and detecting the fluorescence strength. A linear compensation can be applied in the weighting coefficient among different points in the target pattern to counteract this non-uniformity (see Note 3).
Optogenetics Experiment
-
(5.1) Before each set of experiments on animals, it is a good practice to verify the system (laser power, targeting accuracy, power uniformity among different beamlets from the hologram) by generating groups of random spots through holograms, burning the spots on an autofluorescent plastic slide, and comparing the resultant image with the desired target.
-
(5.2) The calcium imaging is first performed followed by the extraction of the spatial footprint and temporal dynamics of each recorded neuron. To confirm the co-expression of calcium indicators and opsins in an individual neuron, one can image the fluorophore linked to the opsin. A desired group of neurons can then be selected for optogenetics experiments. Laser power and stimulation time should be adjusted to produce responses observed physiologically (see Note 4).
-
(5.3) While the excitation wavelengths between the calcium indicators and opsins are different, the calcium indicator may still generate fluorescence background during photostimulation, particularly when a large number of neurons are photostimulated simultaneously. Imaging processing should be implemented to remove or suppress this background.
3 Identification and Targeting of Neuronal Ensembles Related to Behavior
3.1 Background
The implementation of simultaneous two-photon imaging [11] and two-photon optogenetics [12] allowed the recording and manipulation of tens of cells simultaneously [14, 66, 69]. However, the identification of the neurons that could have a robust impact on the overall population activity also represents a critical step in the design of experiments aiming to control learned behaviors [4, 21, 81].
3.1.1 Multidimensional Reduction Techniques Applied to Population Recordings
It has been demonstrated that the activity of neuronal ensembles could be defined as an array of multidimensional population vectors, where the dimensionality of the array is given by all recorded cells in the field of view [5, 8, 82]. Similar population vectors could be visualized by diverse computational techniques that define clusters in a reduced dimensional space, where each cluster depicts a neuronal ensemble [5, 83]. A neuronal ensemble represents a group of neurons with coordinated activity that repeats at different points in time [10]. The characterization of brain states using multidimensional population vectors is independent of the recording length and could be implemented in chronic experiments where the activity of identified neuronal ensembles could be compared at different days.
3.1.2 Targeting Visualized Neuronal Populations with Two-Photon Optogenetics
Two experimental designs are possible to control behavior targeting visualized neuronal populations (see Note 5). One solution is to target all available neurons that could respond to a given behavioral cue with single-cell precision [14, 16, 18]. The other solution is to target neurons with pattern completion capabilities that could recall physiologically relevant neuronal ensembles related to behavior [4, 6, 10, 19, 21].
In cortical circuits, the synchronous activation of several neurons could simultaneously result in two unwanted scenarios: (i) the generation of epileptiform activity or (ii) the forced engagement of GABAergic circuits. In both scenarios, the physiological recalling of a specialized neuronal ensemble is compromised (see Note 5).
On the other hand, the identification and targeting of neurons with pattern completion capabilities could keep the balance between excitation and inhibition inherent to the microcircuit under study, allowing the recalling of physiological neuronal ensembles that work as attractors [84] evoking a given behavior [84, 85].
3.2 Implementation of Analytical Methods to Recall Neuronal Ensembles Relevant to a Learned Behavior
In this part of the chapter, we describe the procedures and concepts to analyze population activity extracted from calcium imaging recordings. We will focus on the steps after individual neurons have been identified from imaging recordings and their activity has been inferred from calcium transients.
3.2.1 Motion Correction, Identification of Neurons, and Spike Extraction
Simultaneous population recordings allow the characterization of population activity with single-cell resolution but generate large datasets representing an analytical challenge. A common practice used for analysis of simultaneous population recordings is to create a binary representation of neuronal activity where 1’s indicate firing and 0’s indicate silent periods [82, 86]. Recently, several independent research groups have released open-source code to preprocess calcium imaging recordings to extract activity information from a series of images [87,88,89,90].
There are four main steps that need to be considered before performing population analysis on binary arrays (Fig. 7):
-
(i)
Motion correction: Lateral displacements are commonly corrected using TurboReg [91], fast Fourier transform [92] or field approaches [93]. Translations in depth should be avoided or the data should be discarded.
-
(ii)
Identification of neurons: Automatic identification of regions of interest (ROI’s) can be performed using different approaches [87, 88].
-
(iii)
Determination of changes in fluorescence: After the identification of ROI’s, fluorescence traces for each neuron should be extracted.
-
(iv)
Activity inference from calcium transients: The first time derivative, deconvolution, or fitting could be used to infer spiking activity from calcium transients [82, 87, 88]. Note that inferring co-activity between pairs of neurons from raw calcium transients introduces an artifact from the decaying phase of the changes in fluorescence. Such common mistake generates spurious correlations between neurons as demonstrated before [7].
Spike inference from holographic calcium imaging recordings. (a) Representative experiment where two focal planes were recorded simultaneously using holographic two-photon microscopy. (b) Regions of interest (ROIs) detected from the recordings in a. Each ROI corresponds to an identified neuron. (c) Representative changes in fluorescence obtained from ROIs shown in b. (d) Binary arrays representing the activity of one neuron obtained from inferred spikes. (Modified from Ref. [5, 10])
3.2.2 Binary Arrays from Inferred Activity
To construct binary arrays, inferred activity should be threshold usually 3 S.D. above noise level (Fig. 7d). This procedure has been shown to match bursting activity of neurons above 2 action potentials [82]. The onsets from inferred activity can be used to represent the overall population activity. A binary matrix [N × T] is constructed, where N denotes the total number of neurons in the field of view and T represents the total number of time points recorded. Each row in the binary matrix depicts the activity of one neuron and each column in the binary matrix represents the activity profile of a neuronal population (Fig. 8a). To visualize the overall network activity, the binary matrix can be plotted as a raster plot where 1’s are dots. The population synchronicity can be extracted from the time histogram of the raster plot [5, 8].
Neuronal ensembles represented as multidimensional population vectors. (a) Schematic representation of different frames where active neurons are shown in black-filled circles (left). Binary matrix illustrating the overall population activity, where each row is a neuron and each column denotes a population vector. The total number of recorded neurons gives the dimensionality of the array. (b) Neuronal ensembles representing recurrent groups of neurons that are active at different times can be understood as clusters of similar population vectors in a multidimensional space. The cosine similarity could be used as a metric to calculate the angle between population vectors. If the angle is close to 0°, the population vectors are similar, therefore almost the same neurons fired at different times. If the angle is close to 90°, the population vectors are different. (Reprinted from Ref. [10])
3.2.3 Multidimensional Population Vectors Defining Neuronal Ensembles
To identify groups of neurons with coordinated activity, population vectors from a physiologically meaningful time window are constructed; time windows from 100–500 ms have been shown to reflect physiological functions and generate similar results. Population vectors capture the coordinated activation of specific groups of neurons. Once the population vectors are defined, comparing the distribution of coordinated activity that could appear randomly against the observed coordinated activity can show population vectors above chance levels [5, 7]. Only population vectors with more active cells than the ones expected by chance should be considered for further analysis. Significant population vectors capture network activity defining a multidimensional space in which the number of dimensions is dictated by the total number of identified neurons with coordinated activity above chance levels [4,5,6, 8, 10]. It is important to highlight that to characterize population activity, each dot in a multidimensional space should be a population vector instead of an individual neuron (Fig. 8b).
3.2.4 Similarity Measurements on Population Vectors
It has been shown that the use of significant population vectors can be used to discriminate similar patterns of activity repeated at different times [4,5,6, 8]. The representation of network activity as population vectors allows an exhaustive comparison of all population vectors to visualize different experimental conditions. Similarity maps of population vectors represent a valuable tool to visualize recurrent activity of neuronal ensembles. To construct similarity maps, all possible combinations of vector pairs need to be computed. Since population vectors representing the activity of a given group of neurons point to a similar place in a multidimensional space, the angle between population vectors could be used to create a similarity map [8, 82]. The cosine of the angle between a pair of population vectors is defined by their normalized dot product: cos(θ)=V1 • V2 / llV1ll llV2ll. Thus, if the angle is close to zero, the vectors are similar whereas if the vectors are different, they tend to be orthogonal (Fig. 8b).
3.2.5 Identification of Neuronal Ensembles from Population Vectors
Similarity maps constructed from all possible vector pairs could be understood as a low-dimensional representation of the original network activity, where the angles between all population vectors as a function of time are highlighted. Thus, high similarity values in the same row denote recurrent groups of neurons firing together. A cluster of population vectors pointing in a similar direction gives the definition of a neuronal ensemble using the population vector approach. Similarity maps can be transformed to a binary matrix S of size [T × T] (Fig. 9a, b). The factorization of S using singular value decomposition (SVD) factorizes S as the multiplication of three factors: V, ∑, and VT, where V and VT are orthonormal basis and ∑ contains the singular values [5, 8]. To detect neuronal ensembles from recurrent patterns of activity observed in the matrix S, the rate of singular values decay is determined (Fig. 9c). Taking the singular values above chance levels can reproduce the original matrix S with high accuracy (Fig. 9d). Then each factor of the SVD decomposition represents a linearly orthogonal component and each factor defines a neuronal ensemble (Fig. 9e). In the case of primary visual cortex, each neuronal ensemble represents a different orientation of drifting-gratings, a different natural scene or the Go signal from a visually guided Go/No-Go task [4, 5, 7]. Another approach to identify neuronal ensembles is the projection of the multidimensional population vectors into a low dimensional space, in such reduced dimensional space clusters of population vectors depicted with cluster analysis define neuronal ensembles or network states (Fig. 8b) [82]. The advantage of using SVD factorization is that the total number of neuronal ensembles could be systematically detected from the magnitude of the singular values. Thus, recurrent population vectors can be assigned to a given neuronal ensemble and population vectors with low repetition rate are excluded.
Neuronal ensembles defined by similarity maps of population vectors using singular value decomposition (SVD). (a) Similarity map illustrating the angles between all possible pair combinations of population vectors. Note that the recurrent activation of a given group of neurons is visualized as increased similarity index in the same row. An angle between two vectors close to 0° represents a similarity value close to 1, whereas an angle between two vectors close to 90° represents a similarity value close to 0. (b) Binary matrix computed from the similarity map in a, representing significant patterns of activity factorized by SVD. Black patterns in the same row indicate recurrent coactive neurons at different times. (c) Magnitude of singular values used to define the number of neuronal ensembles repeated above chance levels. Red line indicates the double of singular values from shuffled data. The cutoff indicates the number of neuronal ensembles that account for >90% of the data. In general microcircuits of ~100 neurons stimulated with four drifting-gratings can be defined by ~6 neuronal ensembles. (d) Binary matrix of the first 6 factors obtained from SVD of b. (e) Factors from SVD that reproduce the overall response of the imaged focal plane to visual stimuli. Bars on top indicate orientations of drifting-gratings. Empty squares represent spontaneously active neuronal ensembles that appeared in the absence of visual stimuli. (Modified from Ref. [5])
3.2.6 Pattern Completion Properties of Neuronal Ensembles
In the neuronal ensemble framework, pattern completion refers to the ability to recall a whole neuronal ensemble by the activation of few neurons with strong functional connectivity [6]. It has been recently shown that the stimulation of the same neuronal population for several times (~100) was able to imprint an artificial neuronal ensemble in layer 2/3 of primary visual cortex of awake head fixed mice [6]. Imprinted ensembles were composed by neurons that had low probability to fire together, but after the imprinting protocol, the stimulated neurons fired spontaneously in the absence of stimuli (Fig. 10a). The mechanism governing the creation of artificial neuronal ensembles could be explained by Hebbian synaptic plasticity [94], where the connectivity between neurons firing together was strengthened (Fig. 10b). Indeed, neurons responding to a given drifting-grating or belonging to a neuronal ensemble have increased probability to be connected [8, 95]. Imprinted ensembles could also be recalled by the stimulation of individual neurons with pattern completion capabilities (Fig. 10), suggesting that pattern completion could be a general property of different brain areas [21].
Imprinting and recalling of artificial neuronal ensembles in layer 2/3 of primary visual cortex. (a) Spatial map of neurons activated with two-photon optogenetics several times (~100). Red neurons represent repeatedly responding neurons (left). Scale bar: 50 μm. Imprinted ensemble (red neurons) shows spontaneous activity in the absence of any stimuli (middle). The activation of one neuron with pattern completion capability (blue arrow) was able to recall the imprinted ensemble (red neurons). (b) Cartoon illustrating how imprinted and recalled ensembles could be explained by the strengthening of connections of pre-existing ensembles and photostimulated neurons. (Modified from Ref. [10])
3.2.7 Recalling Neuronal Ensembles Related to Behavior
To prove the causal relation between neuronal activity and a learned behavior, it is necessary to identify representative members of neuronal ensembles that could trigger the behavior [81]. During the behavioral training phase photostimulation should be avoided (see Note 6). It has been recently shown, in a visually guided Go/No-Go task, that neuronal ensembles responding to drifting-gratings in layer 2/3 of primary visual cortex increase their reliability for the Go signal whereas reduce their reliability for the No-Go signal [4]. Increased reliability is reflected as strengthened functional connectivity of the Go ensemble allowing neurons with pattern completion capabilities [21] to recall the whole Go ensemble and trigger the perception of the Go signal evoking the learned behavior (Fig. 11).
Controlling behavior using pattern completion properties of Go ensembles. (a) In mice trained in a visually guided Go/No-Go task, the Go signal activates a neuronal ensemble related to the correct execution of the task (green neurons). The activation of the Go ensemble produced licking. (b) Activation of as few as two pattern completion neurons (red neurons) belonging to the Go ensemble is able to recall the whole Go ensemble and produce licking even in the absence of visual stimuli or behavioral cues. (Reprinted from Ref. [4])
4 Considerations for the Implementation of a Visually Guided Go/No-Go Task
-
Training and experiments should be performed at the same hour of the day.
-
That same person should handle the animals.
-
Training should be performed every day, avoiding skipping training sessions on weekends or holidays.
-
The weight of the animals should be systematically documented and maintained at the same level overnight. In general, 1 mL of water should increase weight by 1 g.
-
Changes in weight ±2% of the total mass could significantly alter behavioral performance.
-
To avoid continuous licking, plain water should be preferred over sweetened water.
-
The lick port should be placed 1.5 mm from the mouth. A lick port too close to the mouth could cause increased non-task related licking.
5 Notes
-
1.
A proper calcium indicator and opsin pair should be chosen to minimize crosstalk. It is also critical to control imaging laser power so that neurons are not depolarized by the imaging laser during imaging [Subheading 2.2.1].
-
2.
The zeroth-order beam block in the photostimulation path should be positioned at the intermediate plane of the 4f system, right after the spatial light modulator. While it can block the zeroth-order beam, it also makes the neurons in the center field of view inaccessible to the photostimulation beam. Thus, the zeroth-order beam block should have an intermediate size: if too small, the zeroth-order beam cannot be totally blocked, but if too large, there could be a large number of neurons in the center field of view not accessible by the photostimulation laser [Subheading 2.2.2, Fig. 3a].
-
3.
As the opsin expression level may differ between different neurons, the weight factors for different target neurons should be adjusted in the hologram calculation [55, 75] so that their calcium transients evoked by two-photon optogenetics have a similar response in amplitude [see Subheading 2.2.4. (4.2)].
-
4.
The amplitude of calcium transients in individual neurons evoked by two-photon optogenetic stimulation should be similar to the amplitude of calcium transients evoked by physiological stimuli, otherwise the effects described could be due to over stimulation of neurons [see Subheading 2.2.4. (5.2)].
-
5.
The responses induced by the activation of large numbers of neurons in a given optical field could be attenuated by the engagement of GABAergic circuits. The total number of simultaneously photostimulated neurons in a given optical field should be maintained in the same range as the number of active neurons evoked by physiological conditions (sensory stimuli) [see Subheading 3.1.2].
-
6.
During the training phase of the behavioral task, pairing of sensory stimuli and optogenetic stimuli should be avoided. Otherwise, the effects attributed to optogenetic stimuli could be due to mice learning to identify the optogenetic activation of neurons [see Subheading 3.2.7].
6 Outlook
A general development direction of all-optical methods is to increase the throughput (i.e., field of view and spatiotemporal resolution) of both imaging and photostimulation, while reducing the laser power incident onto the brain. Beam multiplexing techniques are promising approaches to increase the imaging throughput [56,57,58,59]. Larger spatial light modulators with higher pixel count could increase the beam diffraction efficiency and thus the field of view [14]. Faster spatial light modulators could allow rapid switching of the hologram and thus could enable faster photostimulation of neuronal ensembles in specific spatiotemporal patterns. As the simultaneously photostimulated neurons across a 3D volume increase [14, 66, 69, 72], there could be off-target effects through photostimulating the dendritic arbors crossing the cell bodies of target neurons. Somatic restricted viruses where the opsin only expresses in the cell body could alleviate this issue [14, 66, 72, 96].
To access deeper brain regions noninvasively, adaptive optics [97, 98] and three-photon excitation [38, 39, 41, 99, 100] could be used. A spatial light modulator in the photostimulation path could implement adaptive optics, and a spatial light modulator in the imaging path could allow for similar corrections. Three-photon calcium imaging has also been successfully demonstrated [39, 41, 99, 100]. One current challenge is that it needs high power for three-photon optogenetics in deep brain regions. The development of high-efficiency opsins optimized for three-photon excitation could overcome this challenge.
The development of all-optical techniques to simultaneously read and write patterned activity in neuronal populations could help the understanding of perception, memory formation, behavioral states, and pathological conditions with single-cell resolution in the next decades. The systematic documentation of the causality between neuronal ensemble activity and brain states requires further development of analytical tools to visualize and target specific neurons that can control the overall network activity [81]. The adaptation of artificial intelligence algorithms used in big data to understand population activity could be used to design and guide in vivo experiments [13].
Scaling of these techniques to several brain areas to measure and control thousands of neurons in different parts of the brain will require the identification and targeting of neurons with pattern completion capability [21, 81]. This approach could control thousands of neurons by the activation of a small percentage of them, reducing sample heating and deterioration of spatial resolution that comes with increased number of parallel stimulations.
Finally, the development of transgenic mice with genetically encoded calcium or voltage indicators and opsins that use different wavelengths in molecularly identified subpopulations of neurons [20] will allow chronic recordings and stimulation of several brain areas with single cell and molecular identity precision, which represent the next challenge for neuronal ensemble research.
References
Alivisatos AP, Chun M, Church GM, Greenspan RJ, Roukes ML, Yuste R (2012) The brain activity map project and the challenge of functional connectomics. Neuron 74:970–974
Packer AM, Russell LE, Dalgleish HW, Hausser M (2015) Simultaneous all-optical manipulation and recording of neural circuit activity with cellular resolution in vivo. Nat Methods 12:140–146
Rickgauer JP, Deisseroth K, Tank DW (2014) Simultaneous cellular-resolution optical perturbation and imaging of place cell firing fields. Nat Neurosci 17:1816–1824
Carrillo-Reid L, Han S, Yang W, Akrouh A, Yuste R (2019) Controlling visually guided behavior by holographic recalling of cortical ensembles. Cell 178(447–457):e445
Carrillo-Reid L, Miller JE, Hamm JP, Jackson J, Yuste R (2015) Endogenous sequential cortical activity evoked by visual stimuli. J Neurosci 35:8813–8828
Carrillo-Reid L, Yang W, Bando Y, Peterka DS, Yuste R (2016) Imprinting and recalling cortical ensembles. Science 353:691–694
Miller JE, Ayzenshtat I, Carrillo-Reid L, Yuste R (2014) Visual stimuli recruit intrinsically generated cortical ensembles. Proc Natl Acad Sci U S A 111:E4053–E4061
Carrillo-Reid L, Lopez-Huerta VG, Garcia-Munoz M, Theiss S, Arbuthnott GW (2015) Cell assembly signatures defined by short-term synaptic plasticity in cortical networks. Int J Neural Syst 25:1550026
Yuste R (2015) From the neuron doctrine to neural networks. Nat Rev Neurosci 16:487–497
Carrillo-Reid L, Yang W, Kang Miller JE, Peterka DS, Yuste R (2017) Imaging and optically manipulating neuronal ensembles. Annu Rev Biophys 46:271–293
Nikolenko V, Watson BO, Araya R, Woodruff A, Peterka DS, Yuste R (2008) SLM microscopy: scanless two-photon imaging and photostimulation with spatial light modulators. Front Neural Circuit 2:1–14
Packer AM, Peterka DS, Hirtz JJ, Prakash R, Deisseroth K, Yuste R (2012) Two-photon optogenetics of dendritic spines and neural circuits. Nat Methods 9:1202–1205
Pandarinath C, O'Shea DJ, Collins J, Jozefowicz R, Stavisky SD, Kao JC, Trautmann EM, Kaufman MT, Ryu SI, Hochberg LR et al (2018) Inferring single-trial neural population dynamics using sequential auto-encoders. Nat Methods 15:805–815
Marshel JH, Kim YS, Machado TA, Quirin S, Benson B, Kadmon J, Raja C, Chibukhchyan A, Ramakrishnan C, Inoue M et al (2019) Cortical layer-specific critical dynamics triggering perception. Science 365:eaaw5202
Daie K, Svoboda K, Druckmann S (2021) Targeted photostimulation uncovers circuit motifs supporting short-term memory. Nat Neurosci 24:259–265
Dalgleish HW, Russell LE, Packer AM, Roth A, Gauld OM, Greenstreet F, Thompson EJ, Hausser M (2020) How many neurons are sufficient for perception of cortical activity? elife 9:e58889
Gill JV, Lerman GM, Zhao H, Stetler BJ, Rinberg D, Shoham S (2020) Precise holographic manipulation of olfactory circuits reveals coding features determining perceptual detection. Neuron 108(382–393):e385
Robinson NTM, Descamps LAL, Russell LE, Buchholz MO, Bicknell BA, Antonov GK, Lau JYN, Nutbrown R, Schmidt-Hieber C, Hausser M (2020) Targeted activation of hippocampal place cells drives memory-guided spatial behavior. Cell 183:2041–2042
Carrillo-Reid L (2021) Neuronal ensembles in memory processes. Semin Cell Dev Biol 125:136–143
Emiliani V, Cohen AE, Deisseroth K, Hausser M (2015) All-optical interrogation of neural circuits. J Neurosci 35:13917–13926
Carrillo-Reid L, Han S, O'Neil D, Taralova E, Jebara T, Yuste R (2021) Identification of pattern completion neurons in neuronal ensembles using probabilistic graphical models. J Neurosci 41:8577–8588
Tsien RY (1980) New calcium indicators and buffers with high selectivity against magnesium and protons: design, synthesis, and properties of prototype structures. Biochemistry 19:2396–2404
Yuste R, Katz LC (1991) Control of postsynaptic Ca2+ influx in developing neocortex by excitatory and inhibitory neurotransmitters. Neuron 6:333–344
Miyawaki A, Llopis J, Heim R, McCaffery JM, Adams JA, Ikura M, Tsien RY (1997) Fluorescent indicators for Ca2+ based on green fluorescent proteins and calmodulin. Nature 388:882–887
Stosiek C, Garaschuk O, Holthoff K, Konnerth A (2003) In vivo two-photon calcium imaging of neuronal networks. Proc Natl Acad Sci U S A 100:7319–7324
Chen TW, Wardill TJ, Sun Y, Pulver SR, Renninger SL, Baohan A, Schreiter ER, Kerr RA, Orger MB, Jayaraman V et al (2013) Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499:295–300
Gross D, Loew LM (1989) Fluorescent indicators of membrane potential: microspectrofluorometry and imaging. Methods Cell Biol 30:193–218
Peterka DS, Takahashi H, Yuste R (2011) Imaging voltage in neurons. Neuron 69:9–21
Dana H, Sun Y, Mohar B, Hulse BK, Kerlin AM, Hasseman JP, Tsegaye G, Tsang A, Wong A, Patel R et al (2019) High-performance calcium sensors for imaging activity in neuronal populations and microcompartments. Nat Methods 16:649–657
Dana H, Mohar B, Sun Y, Narayan S, Gordus A, Hasseman JP, Tsegaye G, Holt GT, Hu A, Walpita D et al (2016) Sensitive red protein calcium indicators for imaging neural activity. elife 5:e12727
Bando Y, Grimm C, Cornejo VH, Yuste R (2019) Genetic voltage indicators. BMC Biol 17:71
Adams SR, Tsien RY (1993) Controlling cell chemistry with caged compounds. Annu Rev Physiol 55:755–784
Ellis-Davies GC (2007) Caged compounds: photorelease technology for control of cellular chemistry and physiology. Nat Methods 4:619–628
Nagel G, Szellas T, Huhn W, Kateriya S, Adeishvili N, Berthold P, Ollig D, Hegemann P, Bamberg E (2003) Channelrhodopsin-2, a directly light-gated cation-selective membrane channel. Proc Natl Acad Sci U S A 100:13940–13945
Boyden ES, Zhang F, Bamberg E, Nagel G, Deisseroth K (2005) Millisecond-timescale, genetically targeted optical control of neural activity. Nat Neurosci 8:1263–1268
Denk W, Strickler JH, Webb WW (1990) Two-photon laser scanning fluorescence microscopy. Science 248:73–76
Yuste R, Denk W (1995) Dendritic spines as basic functional units of neuronal integration. Nature 375:682–684
Horton NG, Wang K, Kobat D, Clark CG, Wise FW, Schaffer CB, Xu C (2013) three-photon microscopy of subcortical structures within an intact mouse brain. Nat Photonics 7:205–209
Ouzounov DG, Wang T, Wang M, Feng DD, Horton NG, Cruz-Hernandez JC, Cheng YT, Reimer J, Tolias AS, Nishimura N et al (2017) In vivo three-photon imaging of activity of GCaMP6-labeled neurons deep in intact mouse brain. Nat Methods 14:388–390
Podgorski K, Ranganathan GN (2016) Brain heating induced by near infrared lasers during multi-photon microscopy. J Neurophysiol 116(3):1012–1023:jn 00275 02016
Rowlands CJ, Park D, Bruns OT, Piatkevich KD, Fukumura D, Jain RK, Bawendi MG, Boyden ES, So PTC (2017) Wide-field three-photon excitation in biological samples. Light Sci Appl 6:e16255
Grewe BF, Voigt FF, van‘t Hoff M, Helmchen F (2011) Fast two-layer two-photon imaging of neuronal cell populations using an electrically tunable lens. Biomed Opt Express 2:2035–2046
Han S, Yang W, Yuste R (2019) Two-color volumetric imaging of neuronal activity of cortical columns. Cell Rep 27(2229–2240):e2224
Anselmi F, Ventalon C, Begue A, Ogden D, Emiliani V (2011) Three-dimensional imaging and photostimulation by remote-focusing and holographic light patterning. Proc Natl Acad Sci U S A 108:19504–19509
Botcherby EJ, Juskaitis R, Booth MJ, Wilson T (2008) An optical technique for remote focusing in microscopy. Opt Commun 281:880–887
Botcherby EJ, Smith CW, Kohl MM, Debarre D, Booth MJ, Juskaitis R, Paulsen O, Wilson T (2012) Aberration-free three-dimensional multiphoton imaging of neuronal activity at kHz rates. Proc Natl Acad Sci U S A 109:2919–2924
Kaplan A, Friedman N, Davidson N (2001) Acousto-optic lens with very fast focus scanning. Opt Lett 26:1078–1080
Duemani Reddy G, Kelleher K, Fink R, Saggau P (2008) Three-dimensional random access multiphoton microscopy for functional imaging of neuronal activity. Nat Neurosci 11:713–720
Grewe BF, Langer D, Kasper H, Kampa BM, Helmchen F (2010) High-speed in vivo calcium imaging reveals neuronal network activity with near-millisecond precision. Nat Methods 7:399–405
Kirkby PA, Nadella KMNS, Silver RA (2010) A compact acousto-optic lens for 2D and 3D femtosecond based 2-photon microscopy. Opt Express 18:13720–13744
Katona G, Szalay G, Maak P, Kaszas A, Veress M, Hillier D, Chiovini B, Vizi ES, Roska B, Rozsa B (2012) Fast two-photon in vivo imaging with three-dimensional random-access scanning in large tissue volumes. Nat Methods 9:201–208
Cheng A, Goncalves JT, Golshani P, Arisaka K, Portera-Cailliau C (2011) Simultaneous two-photon calcium imaging at different depths with spatiotemporal multiplexing. Nat Methods 8:139–142
Ducros M, Goulam Houssen Y, Bradley J, de Sars V, Charpak S (2013) Encoded multisite two-photon microscopy. Proc Natl Acad Sci U S A 110:13138–13143
Wu J, Liang Y, Hsu C-L, Chavarha M, Evans SW, Shi D, Lin MZ, Tsia KK, Ji N (2019) Kilohertz in vivo imaging of neural activity. bioArxiv:543058
Yang W, Miller JE, Carrillo-Reid L, Pnevmatikakis E, Paninski L, Yuste R, Peterka DS (2016) Simultaneous multi-plane imaging of neural circuits. Neuron 89:269–284
Yang W, Yuste R (2017) In vivo imaging of neural activity. Nat Methods 14:349–359
Ji N, Freeman J, Smith SL (2016) Technologies for imaging neural activity in large volumes. Nat Neurosci 19:1154–1164
Weisenburger S, Vaziri A (2018) A guide to emerging technologies for large-scale and whole-brain optical imaging of neuronal activity. Annu Rev Neurosci 41:431–452
Mertz J (2019) Strategies for volumetric imaging with a fluorescence microscope. Optica 6:1261–1268
Oron D, Tal E, Silberberg Y (2005) Scanningless depth-resolved microscopy. Opt Express 13:1468–1476
Zhu G, van Howe J, Durst M, Zipfel W, Xu C (2005) Simultaneous spatial and temporal focusing of femtosecond pulses. Opt Express 13:2153–2159
Papagiakoumou E, de Sars V, Oron D, Emiliani V (2008) Patterned two-photon illumination by spatiotemporal shaping of ultrashort pulses. Opt Express 16:22039–22047
Papagiakoumou E, de Sars V, Emiliani V, Oron D (2009) Temporal focusing with spatially modulated excitation. Opt Express 17:5391–5401
Papagiakoumou E, Ronzitti E, Emiliani V (2020) Scanless two-photon excitation with temporal focusing. Nat Methods 17:571–581
Papagiakoumou E, Anselmi F, Begue A, de Sars V, Gluckstad J, Isacoff EY, Emiliani V (2010) Scanless two-photon excitation of channelrhodopsin-2. Nat Methods 7:848–854
Mardinly AR, Oldenburg IA, Pegard NC, Sridharan S, Lyall EH, Chesnov K, Brohawn SG, Waller L, Adesnik H (2018) Precise multimodal optical control of neural ensemble activity. Nat Neurosci 21:881–893
Dal Maschio M, Donovan JC, Helmbrecht TO, Baier H (2017) Linking neurons to network function and behavior by two-photon holographic optogenetics and volumetric imaging. Neuron 94(774–789):e775
Forli A, Vecchia D, Binini N, Succol F, Bovetti S, Moretti C, Nespoli F, Mahn M, Baker CA, Bolton MM et al (2018) Two-photon bidirectional control and imaging of neuronal excitability with high spatial resolution in vivo. Cell Rep 22:3087–3098
Yang W, Carrillo-Reid L, Bando Y, Peterka DS, Yuste R (2018) Simultaneous two-photon imaging and two-photon optogenetics of cortical circuits in three dimensions. elife 7:e32671
Yizhar O, Fenno LE, Prigge M, Schneider F, Davidson TJ, O'Shea DJ, Sohal VS, Goshen I, Finkelstein J, Paz JT et al (2011) Neocortical excitation/inhibition balance in information processing and social dysfunction. Nature 477:171–178
Akerboom J, Carreras Calderon N, Tian L, Wabnig S, Prigge M, Tolo J, Gordus A, Orger MB, Severi KE, Macklin JJ et al (2013) Genetically encoded calcium indicators for multi-color neural activity imaging and combination with optogenetics. Front Mol Neurosci 6:2
Pegard NC, Mardinly AR, Oldenburg IA, Sridharan S, Waller L, Adesnik H (2017) Three-dimensional scanless holographic optogenetics with temporal focusing (3D-SHOT). Nat Commun 8:1228
Chen IW, Ronzitti E, Lee BR, Daigle TL, Dalkara D, Zeng H, Emiliani V, Papagiakoumou E (2019) In vivo submillisecond two-photon optogenetics with temporally focused patterned light. J Neurosci 39:3484–3497
Gerchberg RW, Saxton WO (1972) A practical algorithm for the determination of the phase from image and diffraction pictures. Optik 35:237–246
Golan L, Reutsky I, Farah N, Shoham S (2009) Design and characteristics of holographic neural photo-stimulation systems. J Neural Eng 6:066004
Yang SJ, Allen WE, Kauvar I, Andalman AS, Young NP, Kim CK, Marshel JH, Wetzstein G, Deisseroth K (2015) Extended field-of-view and increased-signal 3D holographic illumination with time-division multiplexing. Opt Express 23:32573–32581
Hernandez O, Papagiakoumou E, Tanese D, Fidelin K, Wyart C, Emiliani V (2016) Three-dimensional spatiotemporal focusing of holographic patterns. Nat Commun 7:11928
Mardinly A, Pegard N, Oldenburg I, Sridharan S, Hakim R, Waller L, Adesnik H (2017) 3D all-optical control of functionally defined neurons with cellular resolultion and sub-millisecond precision. In: Optics in the Life Sciences. The Optical Society of America, BrM3B.4
Pologruto TA, Sabatini BL, Svoboda K (2003) ScanImage: flexible software for operating laser scanning microscopes. Biomed Eng Online 2:13
Yang W (2018). https://github.com/wjyangGithub/Holographic-Photostimulation-System
Carrillo-Reid L, Yuste R (2020) Playing the piano with the cortex: role of neuronal ensembles and pattern completion in perception and behavior. Curr Opin Neurobiol 64:89–95
Carrillo-Reid L, Tecuapetla F, Tapia D, Hernandez-Cruz A, Galarraga E, Drucker-Colin R, Bargas J (2008) Encoding network states by striatal cell assemblies. J Neurophysiol 99:1435–1450
Cunningham JP, Yu BM (2014) Dimensionality reduction for large-scale neural recordings. Nat Neurosci 17:1500–1509
Cossart R, Aronov D, Yuste R (2003) Attractor dynamics of network UP states in neocortex. Nature 423:283–289
Hopfield JJ (1982) Neural networks and physical systems with emergent collective computational abilities. Proc Natl Acad Sci U S A 79:2554–2558
Sasaki T, Matsuki N, Ikegaya Y (2007) Metastability of active CA3 networks. J Neurosci 27:517–528
Pnevmatikakis EA, Soudry D, Gao Y, Machado TA, Merel J, Pfau D, Reardon T, Mu Y, Lacefield C, Yang W et al (2016) Simultaneous denoising, deconvolution, and demixing of calcium imaging data. Neuron 89:285–299
Mukamel EA, Nimmerjahn A, Schnitzer MJ (2009) Automated analysis of cellular signals from large-scale calcium imaging data. Neuron 63:747–760
Giovannucci A, Friedrich J, Gunn P, Kalfon J, Brown BL, Koay SA, Taxidis J, Najafi F, Gauthier JL, Zhou P et al (2019) CaImAn an open source tool for scalable calcium imaging data analysis. elife 8:e38173
Pnevmatikakis EA (2019) Analysis pipelines for calcium imaging data. Curr Opin Neurobiol 55:15–21
Thevenaz P, Ruttimann UE, Unser M (1998) A pyramid approach to subpixel registration based on intensity. IEEE Trans Image Process 7:27–41
Guizar-Sicairos M, Thurman ST, Fienup JR (2008) Efficient subpixel image registration algorithms. Opt Lett 33:156–158
Pnevmatikakis EA, Giovannucci A (2017) NoRMCorre: An online algorithm for piecewise rigid motion correction of calcium imaging data. J Neurosci Methods 291:83–94
Hebb DO (1949) The organization of behavior; a neuropsychological theory. Wiley, New York
Ko H, Hofer SB, Pichler B, Buchanan KA, Sjostrom PJ, Mrsic-Flogel TD (2011) Functional specificity of local synaptic connections in neocortical networks. Nature 473:87–91
Shemesh OA, Tanese D, Zampini V, Linghu C, Piatkevich K, Ronzitti E, Papagiakoumou E, Boyden ES, Emiliani V (2017) Temporally precise single-cell-resolution optogenetics. Nat Neurosci 20:1796–1806
Ji N (2017) Adaptive optical fluorescence microscopy. Nat Methods 14:374–380
Booth MJ (2014) Adaptive optical microscopy: the ongoing quest for a perfect image. Light Sci Appl 3:e165
Li B, Wu C, Wang M, Charan K, Xu C (2020) An adaptive excitation source for high-speed multiphoton microscopy. Nat Methods 17:163–166
Weisenburger S, Tejera F, Demas J, Chen B, Manley J, Sparks FT, Traub FM, Daigle T, Zeng HK, Losonczy A et al (2019) Volumetric Ca2+ imaging in the mouse brain using hybrid multiplexed sculpted light microscopy. Cell 177:1050
Helmchen F, Denk W (2005) Deep tissue two-photon microscopy. Nat Methods 2:932–940
Zipfel WR, Williams RM, Webb WW (2003) Nonlinear magic: multiphoton microscopy in the biosciences. Nat Biotechnol 21:1369–1377
Yang W, Yuste R (2018) Holographic imaging and photostimulation of neural activity. Curr Opin Neurobiol 50:211–221
Acknowledgments
CONACYT (INFR-2018-294756 [LCR], CF6653 [LCR]), Burroughs Wellcome Fund (Career Award at the Scientific Interface #1015761) [WY], National Eye Institute (R01EY011787 [RY]), National Institute of Mental Health (R01MH115900) [RY], National Institute of Neurological Disease and Stroke (R01NS110422 [RY], R34NS116740 [RY], R01NS118289 [WY]), the US Army Research Office (W911NF-12-1-0594 MURI) [RY], and the National Science Foundation (CAREER 1847141 [WY], CRCNS 1822550 [RY]). Rafael Yuste is listed as an inventor of the patent: “Devices, apparatus and method for providing photostimulation and imaging of structures” (United States Patent 9846313). Weijian Yang and Rafael Yuste are listed as inventors of the patent: “System, method and computer-accessible medium for multi-plane imaging of neural circuits” (United States Patent 10520712).
Author information
Authors and Affiliations
Corresponding authors
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2023 The Author(s)
About this protocol
Cite this protocol
Carrillo-Reid, L., Yang, W., Yuste, R. (2023). Optical and Analytical Methods to Visualize and Manipulate Cortical Ensembles and Behavior. In: Papagiakoumou, E. (eds) All-Optical Methods to Study Neuronal Function. Neuromethods, vol 191. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2764-8_11
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2764-8_11
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2763-1
Online ISBN: 978-1-0716-2764-8
eBook Packages: Springer Protocols