Abstract
Purpose
Gene mutations drive tumor immune microenvironment (TIME) heterogeneity, in turn affecting prognosis and immunotherapy efficacy. PIK3CA is the most frequently mutated gene in breast cancer (BC), yet its relevance to BC prognosis remains controversial. Herein, we sought to determine the impact of PIK3CA mutation-driven immune genes (PDIGs) on BC prognosis in relation to TIME heterogeneity.
Methods
PIK3CA mutation characteristics were compared and verified between the TCGA-BRCA dataset and a patient cohort from our hospital. PIK3CA mutation-driven differentially expressed genes were identified for consensus clustering and weighted gene co-expression network analysis to select the modules most relevant to the immune subtype. Thereafter, the two were intersected to obtain PDIGs. Univariate Cox, LASSO, and multivariate Cox regression analyses were sequentially performed on PDIGs to obtain a PIK3CA mutation-driven immune signature (PDIS), which was then validated using the Gene Expression Omnibus (GEO) database. Differences in functional enrichment, mutation landscape, immune infiltration, checkpoint gene expression, and drug response were compared between different risk groups.
Results
PIK3CA mutation frequencies in the TCGA and validation cohorts were 34.49% and 40.83%, respectively. PIK3CA mutants were significantly associated with ER, PR, and molecular BC subtypes in our hospital cohort. The PDIS allowed for effective risk stratification and exhibited prognostic power in TCGA and GEO sets. The low-risk patients exhibited greater immune infiltration, higher expression of common immune checkpoint factors, and lower scores for tumor immune dysfunction and exclusion.
Conclusion
The PDIS can be used as an effective prognostic model for predicting immunotherapy response to guide clinical decision-making.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In 2020, female breast cancer (BC) was the most common malignant tumor worldwide and the leading cause of cancer-related death in women (Sung et al. 2021). It is estimated that 287,850 new cases of female BC and 43,250 deaths will occur in 2022 in the US alone (Siegel and Miller 2022). In recent years, surgery, radiotherapy, chemotherapy, immunotherapy, endocrine, and targeted therapy have been widely applied for the treatment of this heterogeneous malignancy, leading to significant progress. However, the global burden of BC remains considerable, necessitating the early identification of high-risk patients and precision medicine-inspired solutions for improved prognosis.
Phosphatidylinositol 3-kinases (PI3Ks) phosphorylate the third hydroxyl of the phosphatidylinositol ring, and they can be classified into three types based on structure (I, II, and III) (Mosele et al. 2020). Type I are the most widely studied, acting in various important biological processes, such as cell growth, proliferation, metabolism, and migration through the PI3K/AKT/mTOR pathway (Samuels et al. 2005). The PIK3CA gene encodes the catalytic subunit of class IA PI3Ks (p110α) (Zardavas et al. 2014). Mutation of PIK3CA can lead to abnormal catalytic activity of PI3Ks and, consequently, promote carcinogenesis in various tissues (Herberts et al. 2020; Hu et al. 2021; Jin et al. 2021; Ugai et al. 2021). PIK3CA somatic mutations occur in approximately 30% of BC patients, being more common in hormone receptor (HR)-positive tumors (Lv et al. 2020). Approximately 80% of PIK3CA mutations occur in the helical and kinase domains, with E542K and E545K in exon 9 as well as H1047R and H1047L in exon 20 as the most common variants (Dirican et al. 2016). In the past few years, PIK3CA has emerged as a promising target for BC treatment, with alpelisib and fulvestrant receiving approval for the treatment of PIK3CA-mutated, endocrine-resistant HR+/HER2− locally-advanced or metastatic BC (De Mattos-Arruda 2020). Moreover, several large clinical trials have reported an association of PIK3CA mutations with favorable prognosis and clinicopathological BC features (Kalinsky et al. 2009; Loi et al. 2013). Therefore, further study of the differences in gene expression between PIK3CA-mutated and wild-type BC should provide valuable insight for predicting prognosis and guiding clinical decision-making.
In light of the impact that the tumor immune microenvironment (TIME) has on tumorigenesis and metastasis, there has been increasing interest in immunotherapy for BC. Immune checkpoint inhibitors (ICIs) targeting cytotoxic T lymphocyte-associated protein 4 (CTLA4), programmed cell death-1 (PD-1), and programmed cell death ligand-1 (PD-L1) have shown promising efficacy in certain tumor types, leading to their entry into the clinic (Gaynor et al. 2022). The immunoregulatory effects of CTLA4 antagonists tremelimumab and ipilimumab have been confirmed in small-scale BC cohorts (Zhu et al. 2021). Clinical trials of CTLA4 antagonist monotherapy or in combination with other immunomodulators are ongoing, with the clinical benefits of CTLA4 inhibition in BC expected to be confirmed in the future. Anti-PD-1/PD-L1 immunotherapy confers a survival benefit to some metastatic triple-negative BC patients (Adams et al. 2019; Schmid et al. 2017). The TIME, which is composed of infiltrating immune cells and various other cell types, is strongly correlated with tumor progression and ICI response (Miller et al. 2021; Oliver et al. 2019; Taylor et al. 2017). Tumor-infiltrating lymphocytes (TILs) are major indicators of immune infiltration in the TIME and can inhibit tumor growth (Bagbudar et al. 2022). Enhancing cytotoxic T cell responses may suppress tumor growth and improve patients survival (Rupp et al. 2022). Studies have shown that BC tumors with a higher proportion of TILs are more sensitive to chemotherapy, radiotherapy, as well as immunotherapy, leading to improved prognosis (Adams et al. 2014; Denkert et al. 2018). Some studies have reported the effect of different mutations on ICI response (Chen et al. 2021a; Collins et al. 2022). Currently, the main challenge of immunotherapy remains the exploration of reliable indicators to predict its potential efficacy.
Hence, we explored the mutation characteristics of PIK3CA in BC samples from the Cancer Genome Atlas Database (TCGA, https://portal.gdc.cancer.gov/) and our institution. Subsequently, a PIK3CA mutation-driven immune signature (PDIS) was developed based on TCGA data, and we validated its capacity for BC patient risk stratification using the Gene Expression Omnibus database (GEO, https://www.ncbi.nlm.nih.gov/geo/). The signature presented herein may be employed to evaluate the TIME, optimize clinical benefit, and predict patient prognosis.
Materials and methods
Data collection and processing
Somatic mutation profile, RNA-seq data of BC and normal samples (FPKM-normalized format), as well as corresponding clinical features (including age, survival time, survival status, TNM stage, pathological stage, and ER/PR/HER2 receptor status) were collected from TCGA. Patients without survival information were excluded from subsequent analyses. We implemented the following inclusion criteria for cohorts from the GEO database: (1) samples were derived from human BC; (2) RNA-seq expression data was available; (3) clinical and prognostic information of patients was available; (4) the number of samples was greater than 50.
PIK3CA mutation analysis of BC samples from local hospital
To validate the results of mutation analysis in TCGA, we obtained surgically resected female BC samples from the Department of Oncology at Second Affiliated Hospital of Xi’an Jiaotong University obtained from January 2019 to July 2022. Screening criteria included: (1) no preoperative neoadjuvant therapy, (2) postoperative pathologically confirmed BC, and (3) pathological tissues eligible for molecular typing. This study was approved by the Ethics Committee of the Second Affiliated Hospital of Xi’an Jiaotong University. The surgical procedure was performed in accordance with the Declaration of Helsinki. All patients signed an informed consent form. We employed the allele-specific polymerase chain reaction (AS-PCR) method for PIK3CA mutation detection in patient samples. DNA was extracted from formalin-fixed, paraffin-embedded (FFPE) samples using the QIAamp DNA FFPE Tissue Kit (Qiagen, Cat No./ID: 56,404). Five hotspot mutations of PIK3CA (H1047R, H1047L, E542K, E545K, E545D) were then examined as per the manufacturer’s instructions using the Human PI3K Gene Mutation Fluorescence PCR diagnostic kit (Amoy Diagnostics, Xiamen, China) on an SLAN-96S fluorescent quantitative PCR instrument. Based on the obtained Ct values, we divided samples into negative, weakly positive, and strongly positive, with the latter two considered to harbor mutations.
Somatic mutation analysis and clinical validation
BC mutation data, including mutated genes, mutation types, and mutation sites, were downloaded from TCGA “Masked Somatic Mutation” database. The waterfall function in the “maftools” package was applied to obtain the mutation landscape of the TCGA-BRCA cohort. To understand the distribution of PIK3CA mutations in BC and their correlation with clinicopathological factors, PIK3CA mutation samples and related information were extracted from the downloaded mutation data, with clinical samples obtained from our local hospital used for comparison and validation. Correlations between PIK3CA mutations and clinical variables were analyzed using Pearson’s Chi-square test or Fisher’s exact test (Shao et al. 2021).
Identification of differentially expressed genes (DEGs) driven by PIK3CA mutations
RNA-seq data of PIK3CAMUT and PIK3CAWT patient tumor samples were extracted. After normalization, the “edgeR” package was further used to identify DEGs driven by PIK3CA mutations in BC patients. The screening criteria were set as the adjusted P < 0.05, log2|fold change (FC)|> 0.
Consensus clustering
Single-sample gene set enrichment analysis (ssGSEA) was used to analyze the infiltration of 22 immune cell types and the activity of seven immune-related pathways in tumor tissue samples (Hänzelmann et al. 2013). Based on the extent of immune infiltration, we applied the “ConsensusClusterPlus” package for consensus clustering to divide BC patients into two groups of high- and low-immune activity (Seiler et al. 2010).
Weighted gene co-expression network analysis
The “WGCNA” package was used to construct a gene co-expression network to aggregate genes with highly correlated expression and identify gene modules closely related to immune subtypes (Liu et al. 2022). Firstly, a similarity matrix was constructed based on expression data, and the Pearson correlation coefficient was calculated to evaluate the similarity between genes (Langfelder and Horvath 2008). An adjacency matrix was then constructed based on the above matrix. The formula was as follows: aij = power (sij, β) =|sij|β, where ajj represents the correlation strength between gene i and gene j, sij is the Pearson correlation coefficient between the two genes, and β is the soft threshold (power) (Zhang and Horvath 2005). By constructing a topological overlap matrix (TOM), hierarchical clustering analysis of genes was performed, and genes with similar expression patterns were divided into gene modules using dynamic branch cut methods (Tian et al. 2020). Each module contained at least 200 genes, and modules with correlations greater than or equal to 0.75 were merged. Finally, the Pearson correlation coefficient between the MEs of each module and the clinical traits was calculated, and the module most related to the immune subtypes were screened for subsequent analysis. Module eigengene (ME, The first principal component in each module) described the overall level of gene expression in the Module. Module membership (MM, Correlation of all gene expression profiles with eigengene of this module) and gene significance (GS, The absolute value of the correlation between genes and phenotypic traits) were used to evaluate the correlation between genes in the module and the module itself, along with the correlation with the corresponding traits of the module (Tian, et al. 2020).
Construction and validation of PDIS
A Venn diagram was used to intersect genes driven by the PIK3CA mutation and the module most relevant to immunity. The obtained genes were termed PIK3CA mutation-driven immune genes (PDIGs). Univariate Cox regression analysis was performed on the obtained PDIGs to screen those related to overall survival (OS) of BC patients. Then, using the “glmnet” and “survival” packages, least absolute shrinkage and selection operator (LASSO) was conducted on the above genes. By controlling the penalty coefficient λ, the coefficients of some genes less related to prognosis were compressed to 0, and the coefficients of genes significantly related to prognosis were retained greater than 0. Then a multivariate model was constructed to further confirm the genes independently associated with prognosis. Risk score = h(t, X) = h0(t) × eƩ (coefi*Expri), where h0(t), coefi, and Expri are the constant, regression coefficient, and gene expression level, respectively (Ren et al. 2022). Patients in TCGA training and GEO validation cohorts were divided into high- and low-risk groups according to the corresponding median risk score, of which the high-risk patients are those with PDIS score above the median and the low-risk patients are those with PDIS score below the median. Kaplan–Meier (KM) survival analysis was performed using the “survival” and “survminer” packages to explore the capacity of PDIS and PIK3CA mutation status or regions to differentiate prognosis between risk groups. Other recently published prognostic models of BC were retrieved from PubMed (https://pubmed.ncbi.nlm.nih.gov/). The “survivalROC” package was used to plot the receiver operating characteristic curves (ROC), whereafter the area under the ROC curve (AUC) and C-index were used to evaluate and compare the predictive validity of multiple models.
Establishment and verification of predictive nomogram
Combining with risk score and other clinicopathological features, R packages “rms” and “regplot” were used to construct a nomogram to quantitatively predict the survival rate of BC patients. Then, using the calibration function and “survivalROC” package, calibration and ROC curves of 1-, 3-, 5-year survival were drawn to verify the predictive ability of the nomogram. Decision curve analysis (DCA) was performed on different prognostic factors using the “ggDCA” package.
Functional enrichment analysis
DEGs between the high- and low-risk groups were screened with log2|FC|> 1 and an adjusted P < 0.05 as the standard. The “clusterProfiler” package was then used to perform Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis to explore the potential function of DEGs (Yu et al. 2012). Next, gene set enrichment analysis (GSEA) was used to explore the signaling pathways associated with different risk subgroups.
Mutation landscape and TMB analysis
The “maftools” package was used to visualize mutations in the high- and low-risk groups as well as to compare the two groups in this regard. The total number of mutations per megabase in each sample was calculated to obtain tumor mutational burden (TMB) (Wan et al. 2020). We compared TMB between the two subgroups and determined the association between TMB and survival.
Evaluation of differences in immune infiltration and checkpoint factor expression
We used the “GSVA” and “GSEABase” packages to conduct ssGSEA, on the basis of which we employed the ESTIMATE algorithm to evaluate immune score, stromal score, estimate score, and tumor purity (Yoshihara et al. 2013). Using the LM22 gene signature matrix downloaded from the CIBERSORT website as a reference (https://cibersortx.stanford.edu/), we determined the relative proportions of 22 immune cell types in each tumor sample (Ren et al. 2021). Finally, the relationship between risk scores, immune scores, immune cell infiltration abundance, and immune checkpoint expression level was analyzed using the “ggplot2” package. Immune function-related genes were collected from TISIDB (an integrated repository portal for tumor-immune system interactions, http://cis.hku.hk/TISIDB/), whereafter the correlation between risk scores and the above genes was analyzed and visualized using the “limma”, “reshape2”, and “RColorBrewer” R packages, to explore the potential mechanism of immune cell infiltration.
Drug response analysis
IC50 refers to the half-inhibitory concentration of the detected drug and is negatively correlated with the effect of chemotherapeutic drugs. We used the “pRRophetic” package and its functions “car”, “ridge”, “preprocessCore”, “genefilter”, and “sva” to calculate IC50 (Pang et al. 2021). Tumor Immune Dysfunction and Exclusion (TIDE) is a newly developed computational method that identifies the potential for tumor immune escape based on transcriptomic data (Jiang et al. 2018). FPKM gene expression data were Z-score normalized, and TIDE scores were then calculated for BC patients, with high TIDE scores predicting poor ICI efficacy. Accordingly, we analyzed the differences in drug response between patients in high- and low-risk groups, visualizing them via the “ggplot2” package.
Statistical analysis
All statistical analyses and charts were generated using R software (version 4.2.0), SPSS software (version 18.0), and Excel (Microsoft Corporation, California). The R package in this article is from CRAN (Comprehensive R Archive Network) or Bioconductor (https://www.bioconductor.org/) for download. KM survival analysis and log-rank tests were used to compare survival between high- and low-risk groups. A two-tailed P-value less than 0.05 was used as the threshold for statistical significance.
Results
BC cases included in the study
The study design is presented in Fig. S1. Mutation analysis included 920 cases, of which 318 were PIK3CAMUT, and 602 were PIK3CAWILD. Then, 1018 samples with RNA-seq expression data, survival, and clinical information from TCGA BC dataset (n = 1057) and normal (n = 111) (Table S1) were further screened for subsequent analysis (individual clinicopathological factors was not available for some patients, which were included after evaluation). Furthermore, 185 BC samples from GSE48390 (n = 81) and GSE42568 (n = 104) were selected as external cohorts from the GEO database, as per screening criteria, to validate gene signature. The included samples and the corresponding survival information are listed in Table S2.
Distribution of PIK3CA in BC and its relationship with clinicopathological factors in TCGA and clinical samples
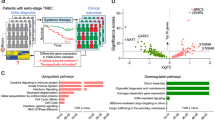
The waterfall diagram of TCGA cohorts showed that PIK3CA was the most frequently mutated gene in BC, with missense mutations being most common (Fig. 1A). A total of 14,599 DEGs were identified between the PIK3CAWT and PIK3CAMUT groups, of which 5316 and 9283 genes were significantly up- or downregulated in the former, respectively (Fig. 1B). We found that 34.49% (318/ 922) of BC samples harbored PIK3CA mutations (28.30% were exon 9 mutations, 33.65% were exon 20 mutations, and 38.05% were other types of mutations) (Fig. 1C). Of the 218 BC cases tested at the local hospital, 89 patients had PIK3CA mutations (Fig. 1D), accounting for 40.83% (39.33% for exon 9 mutation, 52.81% for exon 20 mutation, 7.86% for other types of mutations). H1047R was the most common mutation type in TCGA and samples from the local hospital, accounting for 30.19% (96/ 318) and 41.57% (37/ 89) of PIK3CA mutations, correspondingly.
We then explored the relationship between PIK3CA mutations and various clinicopathological features. In TCGA database, there was no correlation between PIK3CA mutation status or regions and age, T, N, M, pathological stage, ER, PR, nor HER2 status (Table 1). An association between BC subtypes and PIK3CA mutations was not determined due to insufficient information on molecular subtypes. The number of patients included from our hospital was insufficient to explore the relationship of PIK3CA mutations with metastasis and their impact on prognosis. In our clinical validation cohort, PIK3CA mutation regions were significantly associated with ER (P = 0.026) and PR status (P < 0.001), but not with other factors (Table 2). It is worth noting that there was a significant correlation between PIK3CA mutation regions and BC subtype (P = 0.023), with PIK3CA mutations being more frequent in HR + /HER2- subtype tumors.
Immune subtypes distinguished via consensus clustering
Based on the proportion of infiltrating immune cells and pathways, 1057 BC patients were divided into two subgroups (cluster 1 and cluster 2) via consensus clustering (Fig. 2A). PCA analysis revealed that cluster 1 and cluster 2 exhibited obvious separation in spatial distribution (Fig. 2B). The level of immune infiltration was higher in cluster 1 compared to cluster 2, and the former was therefore defined as the high-immunity group (Fig. 2C).
Screening of co-expression module most relevant to immune subtypes based on WGCNA
WGCNA was used to explore the module and hub genes most related to immune subtypes in BC, and β = 5 was selected as the best soft threshold with scale-free R2 = 0.9 (Fig. 3A). Eight modules were finally identified when the dynamic shear tree was used to obtain co-expression modules (Fig. 3B). In addition to immune subtype, we included other clinical characteristics (including mutation status, OS time, OS status, age, stage, T, N, M) from TCGA data when performing modular trait correlation analysis. The heatmap indicated that the green module was most associated with immune subtype (ME = − 0.55, P < 0.001) and was significantly correlated with OS status (ME = − 0.07, P = 0.02) and N stage (ME = − 0.11, P < 0.001), including 1635 genes (Fig. 3C). The scatter plot revealed a strong relationship between Gene significance (GS) and Module Membership (MM) (cor = 0.92, P < 0.001), indicating that these genes were highly correlated not only with the green module but also with immune subtypes, which warranted further investigation (Fig. 3D).
Screening of immune-related genes based on WGCNA. A Analysis of the scale-free fit index (left) and the mean connectivity (right) for various soft-thresholding power. B The hierarchical clustering dendrogram and the corresponding assigned module colors. C Heatmap of correlation between modules obtained by WGCNA and clinical traits. D Scatter plot of relevance between genes within the green module and immune subtypes
Construction of prognostic PDIS in TCGA-BRCA and validation of its predictive ability in GEO cohorts
Five hundred eighty PDIGs were obtained from the intersection of 14,599 PIK3CA mutation-driven genes and 1635 immune genes in the Venn diagram (Fig. 4A). Univariate Cox regression analysis was performed on the above genes, and 136 genes related to the prognosis of BC patients were obtained (Table S3). LASSO regression analysis was then performed, and the coefficients of 26 genes were retained greater than 0 (Fig. 4B, C). Then multivariate Cox regression analysis was performed on these 26 genes to select independent prognostic genes related to survival, and 16 PDIGs were obtained to construct PDIS, including 11 protective genes and 5 risk genes (Figure S2). The risk score was calculated as: Risk score = h0(t) × e (0.4662 * ADORA3) + (0.5468 * C2CD2) + (−0.2922 * CHST10) + (−0.5043 * DTX1) + (−0.1808 * FKBP5) + (−0.2313 * H1−0) + (−0.1762 * HSPA2) + (−0.2945 * JAK2) + (0.2275 * LINC00992) + (−0.2190 * LYSMD2) + (−0.4286 * NFKBIA) + (−0.3825 * PARP12) + (−0.2276 * PDCD4−AS1) + (0.3791 * PPA2) + (0.2722 * PROM2) + (−0.1232 * TCN1). The patients were divided into high- and low-risk groups based on the median risk score. The KM survival curve indicated that OS in the high-risk group was obviously lower than that in the low-risk group (P < 0.001, Fig. 4D). The predicted AUCs of PDIS for 1-, 3-, and 5-year OS were 0.807, 0.800, and 0.790, respectively (Fig. 4E).
Construction and validation of prognostic PDIS. A PDIGs were obtained by Venn diagram. B Ten-fold cross-validation for the coefficients. C Parameter selection of the 26 selected PDIGs in LASSO regression. D KM survival analysis of OS in high- and low-risk patients in TCGA-BRCA cohort. E ROC curves of PDIS predicting OS at 1-, 3-, and 5-year in TCGA-BRCA cohort. F KM survival analysis in the GEO validation cohorts. G ROC curves of PDIS predicting OS in GEO validation cohorts. ROC curves of H Zhang Yi signature, I Peng signature, J Chen signature, K Liu signature, and L Zhang Dai signature predicting OS at 1-, 3-, and 5-year. M Comparison of C-index of predictive power in multiple models. N Univariate and O multivariate analysis of various factors for OS
GSE48390 and GSE42568 cohorts were used to further validate the predictive ability of PDIS. The risk score of each patient was calculated and the cohorts divided into two groups with respect to the median value. KM survival analysis showed that patients in the high-risk group had a significantly worse OS (P < 0.001, Fig. 4F). The AUCs of the PDIS for predicting the 1-, 3-, and 5-year survival in the validation cohort were 0.635, 0.612, and 0.643, respectively (Fig. 4G). Compared with other prognostic models published in recent years, the PDIS exhibited higher AUCs and C-index in predicting BC OS, indicative of its superior predictive performance (Fig. 4H–M). Furthermore, univariate and multivariate Cox analyses of clinical factors, PIK3CA mutation status, and risk scores showed that, except for the PDIS, other factors could not independently predict the prognosis of BC patients (Fig. 4N, O). In view of the significant difference in immune activity between HR + and HR- BC, we analyzed the differences in immune activity between samples with different ER and PR status in the high- and low-risk groups defined by PDIS (Table S4–5). We found that there was no significant difference between ER+ and ER−, PR+ and PR− BC samples in the high- and the low-risk group, indicating that PDIS had similar characteristics in tumors with different HR states (Fig. S3).
Prognostic value of PIK3CA mutations
We explored survival differences based on PIK3CA mutations in the whole BC, high-, and low-risk groups to test whether PIK3CA mutations could predict PDIS-based risk subgroup prognosis. KM survival analysis indicated that neither PIK3CA mutation status (Fig. S4A–C), nor mutation regions (Fig. S4D–F) could accurately stratify BC patients, which also confirmed the greater prognostic value of the PDIS compared to PIK3CA mutation alone.
Ability of the nomogram to quantitatively predict prognosis
Common prognostic predictors (pathological stage, ER, PR, HER2) and PDIS-based risk scores were combined to construct a nomogram to quantitatively predict the OS of BC patients (Fig. 5A). The calibration curve indicated that nomogram-predicted survival was approximately consistent with the actual observed probability, suggesting that the nomogram could accurately predict the 1-, 3-, and 5-year OS (Fig. 5B). Moreover, the predictive ability of the nomogram was higher than that of the PDIS and other predictors based on the AUCs and DCA curves (Fig. 5C–F). In summary, the nomogram can predict BC patient prognosis with greater accuracy.
Establishment and verification of predictive nomogram. A The total points for each item on the nomogram predicted the survival probability at 3- and 5-year. B Calibration curves to assess the consistency between actual survival probability and 1-, 3-, 5-year OS predicted by nomogram. ROC curves of nomogram, PDIS and other prognostic factors for predicting C 1-year, D 3-year, and E 5-year OS. F The clinical efficacy of the nomogram, PDIS, and other prognostic factors in predicting 5-year OS was assessed by DCA
Biological function analysis
We screened 816 DEGs between PDIS-based risk subgroups, of which 671 were upregulated in the low-risk group, and 145 were upregulated in the high-risk group (Fig. S5). Thereafter, GO and KEGG analysis were employed to explore the biological functions of DEGs. GO analysis revealed that DEGs were significantly enriched in immune-related biological processes, such as immune response-activated cell surface receptor signaling pathway and humoral immune response, among others (Fig. 6A). Similarly, KEGG analysis indicated that DEGs were markedly enriched in a series of signaling pathways (such as cytokine-cytokine receptor interaction, chemokine signaling pathway) that regulate immune, inflammatory, and proliferative processes (Fig. 6B). Subsequently, GSEA was used to determine functional differences between high- and low-risk groups. We found that patients in the high-risk group were enriched in the autoimmunity-related systemic lupus erythematosus pathway, while pathways enriched in the low-risk group were mostly related to chemokines and cytokine-mediated pathways (Fig. 6C, D). Moreover, the two risk groups were significantly associated with the regulation of cell proliferation and differentiation pathway, which is expected to play a role in the development of BC (Fig. 6E, F).
Mutational landscape of high- and low-risk groups
The waterfall plots of the top 10 mutated genes in the two risk groups indicated that PIK3CA had the highest mutation frequency in both risk groups, with missense mutations being the most common (Fig. 7A, B). PIK3CA, CDH1, KMT2C, and HMCN1 were more frequently mutated in the low-risk group than in the high-risk group, while the opposite was true for TP53, TTN, GATA3, MUC16, and MAP3K1. The mean TMB value in the high-risk group was significantly higher than that in the low-risk group (Fig. 7C). Taking the median TMB as the cut-off (0.87 mut/MB), KM analysis revealed no significant difference in OS between patients with a high and low TMB (P = 0.679).
Differences in mutation profiles and immune infiltration between low- and high-risk groups. Frequency and type of mutations in the top 10 genes of A low- and B high-risk groups. C Difference in TMB levels between different risk groups. D Infiltration heatmap of 22 immune cell subtypes and 7 immune-related pathways in low- and high-risk groups. E Relationship between risk scores and stromal scores, immune scores, and estimated scores. Immune cell types with more infiltration fraction in F low- and G high-risk groups. H The expression levels of nine common immune checkpoint genes in different risk groups. Heatmap of the correlation between PDIS-based risk scores and I immune-activating genes, J immunosuppressive genes, K chemokines, L chemokine receptors and MHC genes (*P < 0.05, **P < 0.01, and ***P < 0.001)
Immune infiltration profiles of PDIS-based subgroups
To investigate whether the PDIS reflects the TIME, we analyzed immune cell infiltration and related pathways between high- and low-risk groups. As per the generated heatmap, patients of the low-risk group exhibited greater immune infiltration than those from the high-risk group (Fig. 7D). ESTIMATE analysis also showed that the stromal score, immune score, and ESTIMATE score in the low-risk group were higher than those in the high-risk group (Fig. 7E). Moreover, low-risk tumors harbored greater numbers of naive B cells, activated NK cells, CD8 T cells, follicular helper T cells, and regulatory T cells (Fig. 7F), while M0 and M2 macrophages, neutrophils, as well as resting NK cells were more abundant in the high-risk group (Fig. 7G). These results confirmed that patients in the low-risk group exhibited greater immune cell infiltration. Furthermore, we analyzed the expression of immune checkpoint genes in the two risk subgroups. The expression of PD-L1, PD-1, CTLA4, CD28, CD27, LAG3, IDO1, BTLA, and TNFRSF9 was significantly higher in the low-risk group than in the high-risk group, suggesting that low-risk patients were more likely to benefit from ICIs (Fig. 7H). Notably, most immune-activating genes, immunosuppressive genes, chemokines, and chemokine receptors were negatively correlated with risk scores, while all MHC genes were positively correlated with the risk scores, providing a direction for future studies into the potential mechanisms underlying the observed differences in immune cell infiltration between the high- and low-risk groups (Fig. 7I–M). In addition, PDIS stratification was applied to patients such as TNBC who regularly use ICIs, and different immune activities were observed, low-risk TNBC groups harbored greater numbers of CD8 T cells, regulatory T cells, and M1 macrophages, while M2 macrophages were more abundant in the high-risk TNBC groups (Fig. S6A). The expression of CD27, BTLA and LAG3 in low-risk TNBC patients was significantly higher than that in high-risk group, suggesting that TNBC patients in low-risk group were more likely to benefit from ICI treatment (Fig. S6B).
Drug sensitivity analysis for immunotherapy, chemotherapy, and targeted therapy
The response of patients to ICIs in the TCGA-BRCA cohort was predicted by calculating TIDE scores (Table S6). TIDE scores were higher in the high-risk group (Fig. 8A). Based on TIDE scores, patients were divided into responsive and non-responsive groups, and we found that scores in the high-risk group indicated significantly lower efficacy (Fig. 8B, C). This implied that low-risk patients were more likely to benefit from ICI therapy, while the high-risk patients may exhibit greater tumor immune escape and immunotherapy resistance.
Drug sensitivity analysis of immunotherapy, chemotherapy, and targeted therapies. A TIDE scores for different risk groups. B Correlation between risk scores and response to immunotherapy. C Proportion of non-responder and responder cases to immunotherapy in low- and high-risk groups. The estimated IC50 of some common chemotherapeutic and targeted drugs, such as D cisplatin, E paclitaxel, F doxorubicin, G vinorelbine, H 5-fluorouracil, I etoposide, J cabotinib, K lapatinib (*P < 0.05, **P < 0.01, and ***P < 0.001)
According to IC50 levels, risk groups showed differential drug sensitivity to some common chemotherapeutic drugs (cisplatin, paclitaxel, doxorubicin, vinorelbine, 5-fluorouracil, etoposide) and targeted agents (cabotinib, lapatinib). Compared with the high-risk group, patients in the low-risk group were more sensitive to the above-listed chemotherapeutics, suggesting that they may respond better to chemotherapy (Fig. 8D–I). Among targeted agents, low-risk patients were more sensitive to cabotinib (Fig. 8J), while lapatinib was more suitable for high-risk patients (Fig. 8K).
Discussion
BC is a heterogeneous disease. The molecular subtyping of immunohistochemical markers (ER, PR, HER2, Ki67) has greatly improved prognosis prediction and treatment decision-making (Yeo and Guan 2017). With the development of high-throughput sequencing technology, molecular subtype-based models, such as Oncotype DX, MammaPrint, RecurIndex, Endopredict, and PAM50, have been developed and applied in clinical trials as well as in clinical practice for BC diagnosis, individualized treatment, and survival prediction (Barzaman et al. 2020; Nicolini et al. 2018; Sun et al. 2021). However, these widely used models often neglect the effect of genetic differences on TIME heterogeneity. TIME refers to all immune components within the tumor microenvironment (TME), which plays an established central role in cancer development and progression, having predictive value for BC prognosis and immunotherapy response (Baxevanis et al. 2021; Byrne and Savas 2020; Xu et al. 2021). Tumor genetic heterogeneity, including single-nucleotide variants, short indels, and copy number variants, is involved in establishing TIME heterogeneity (Jia et al. 2022). As the most frequently mutated gene in BC, PIK3CA has gradually become the focus of targeted therapy, but the signature of PIK3CA-driven immune activity driven has not yet been studied.
In the present study, we observed that PIK3CA was most frequently mutated gene in the TCGA-BRCA cohort, with 34.49% samples harboring PIK3CA mutations. H1047R, H1047L, E542K, and E545K accounted for 61.95% of the identified mutations, among which H1047R was the most common, which was basically consistent with the conclusions of other studies (A et al. 2021; Chang et al. 2021; Martínez-Sáez et al. 2020). The clinical relevance of PIK3CA mutations has been extensively studied and is thought to be associated with favorable clinicopathological factors, such as smaller tumor size, HR positivity, and lower grade (Loi, et al. 2013; Sabine et al. 2014; Zardavas et al. 2018). However, no significant association between PIK3CA mutations and clinicopathological features was observed in the TCGA cohort. The characteristics of PIK3CA mutations were validated in clinical samples from a local hospital. The mutation frequency of PIK3CA was 40.83%, with H1047R once again accounting for the highest proportion of mutations. Interestingly, although there remained no link between PIK3CA mutation status and clinicopathological features, we found significant differences in ER, PR status, and molecular subtypes between PIK3CA mutation exon 9 and exon 20, which may provide a basis for individualized endocrine and targeted therapy for patients with different subtypes.
Many previous studies have reported that PIK3CA mutation status may be related to the prognosis of BC patients, but its predictive significance has remained controversial (Baselga et al. 2017; Di Leo et al. 2018; Loi, et al. 2013; Mosele, et al. 2020). Hence, we identified DEGs between PIK3CAMUT and PIK3CAWILD tumors. Two immune subtypes (Immune-H and Immune-L) with differential immune infiltration were distinguished via consensus clustering. The key modules and genes were screened via WGCNA analysis and overlapped with PIK3CA-mutated DEGs to yield PDIGs. After univariate Cox regression, LASSO, and multivariate Cox regression analyses, 16 PDIGs closely related to prognosis were selected to establish PDIS. Validation in the GEO database and comparison with the ROC and C-index of other immune (Chen et al. 2021b; Peng et al. 2021; Zhang et al. 2021), ferroptosis (Liu et al. 2021), and glycolysis-related (Zhang et al. 2020) prognostic models published in recent years demonstrated the prognostic value of the PDIS. Analysis of various prognosis-associated factors revealed that neither PIK3CA mutation status, nor factors commonly employed in clinical practice could independently predict BC prognosis. Meanwhile, the PDIS was an independent predictor of prognosis. Taken together with previous studies, the impact of PIK3CA mutation status or regions on BC prognosis remains largely uncertain to date. Similarly to our work, a number of studies have reported no meaningful association between the two (Loibl et al. 2014; Papaxoinis et al. 2015; Sabine, et al. 2014; Zardavas, et al. 2018). For example, a large research pooling 19 early-stage BC studies revealed that although PIK3CA mutations were associated with better OS in a univariate analysis, this relationship was no longer significant after correction via multivariate analysis, and prognosis was not significantly different between patients carrying mutations in the helical versus kinase domain (Zardavas, et al. 2018). Paradoxically, PIK3CA mutations were reported to predict both a favorable or unfavorable prognosis, varying based on molecular subtype and tumor stage (Deng et al. 2015; Elfgen et al. 2019; Liu et al. 2014; Mosele, et al. 2020; Pang et al. 2014). Still, certain studies suggest biological differences between tumors harboring mutations in the helical or kinase domain, which may be associated with different degrees of invasiveness, yet the specific prognostic value remains undetermined (Barbareschi et al. 2007; Lai et al. 2008; Lerma et al. 2008; Mangone et al. 2012; Pang et al. 2009). In summary, compared with PIK3CA mutation status alone, PDIS exhibits a higher predictive ability for BC patient survival. Moreover, by combining PDIS with ER, PR, HER2 status, and stage, the nomogram constructed can predict the survival probability of patients more accurately and quantitatively than PDIS alone.
Functional analysis revealed that DEGs in the high- and low-risk groups were associated with immune-related biological processes and pathways, as well as that there were differences in the functional pathways involved in the two risk groups that may be associated with BC development. Therefore, we analyzed the immunological profiles of PDIS-based risk groups. The low-risk group possessed a high proportion of infiltrating immunostimulatory cells, such as activated NK cells, CD8 T cells, and follicular helper T cells, which play a role in anti-tumor immunity. Among them, CD8 T cells and follicular helper T cells are important TILs, whose abundant infiltration is widely recognized as a marker of favorable prognosis in BC (Salemme et al. 2021). Conversely, regulatory T cells exhibited relatively abundant infiltration in the low-risk group, exerting an immunosuppressive effect, which has been previously reported as associated with poor prognosis (Martinez et al. 2019; Shash et al. 2021). Immune cells, represented by M2 macrophages, exhibited more abundant infiltration in the high-risk group. Some data suggest that M2 contributes to poor prognosis by stimulating angiogenesis and inflammation, enhancing tumor growth, invasion, and metastasis, whilst promoting immunosuppression (Choi et al. 2018; Mehta et al. 2021). Taken together, it is reasonable to speculate that low-risk patients exhibit a stronger anti-tumor immune response and may respond better to immunotherapy.
ICIs, which block immune checkpoint-mediated suppression to trigger robust anti-tumor immunity, have emerged as one of the most effective forms of immunotherapy, with proven benefit in a variety of cancers (Bagchi et al. 2021; Himmel et al. 2020). However, the low immunogenicity of BC often compromises ICI efficacy, highlighting the need for identifying patients expected to benefit. Herein, we observed a significant increase in the expression of nine common immune checkpoint factors in low-risk patients, including PD-L1, PD-1, and CTLA4, suggesting that this subgroup might benefit from ICIs. It is worth noting that it is recommended to combine PDIS with other indicators commonly used in ICI sensitivity evaluation during BC treatment (including ER, BR, HER2 receptor status, pathological stage, etc.) to play a more valuable role. For example, ICIs are usually used in the treatment of TNBC patients, but the therapeutic effect is limited. TNBC patients stratified by PDIS showed different immune activity. Among the low-risk TNBC population, they may be more likely to benefit from the treatment of ICIs, adding new evidence to the patient’s treatment plan and increasing the patient’s confidence. TMB is considered as a valuable biomarker for predicting ICI response, yet its predictive value varies significantly across cancers (Chan et al. 2019). Our results showed that TMB was generally low in BC, with only 1.8% (17/ 922) exhibiting a TMB greater than 10 mut/Mb in the high-risk group. Further, there was no significant difference in OS between H-TMB and L-TMB, which is consistent with previous findings in BC (McGrail et al. 2021; Wang et al. 2020). H-TMB is associated with a favorable ICI response and prognosis, presumably because mutations give rise to neoantigens that are immunogenic and thus are more likely to trigger T cell responses (Zheng 2022). A threshold of 13 mut/Mb is commonly used in H-TMB cancers, such as melanoma and non-small cell lung cancer. However, as an immunologically “cold” tumor with a low TMB, BC is extremely sensitive to the selection based on a TMB cutoff. In our study, using the median TMB (0.87 mut/Mb) was thus not appropriate. Large prospective studies are needed to determine appropriate cutoff that will allow the effective use of TMB in predicting the response to ICIs and prognosis in BC patients. The value of our PDIS for predicting response immunotherapy and chemotherapy was further explored. The TIDE was reported as an accurate predictor of the response to anti-PD-1 or -CTLA4 when compared to other indicators (e.g., PD-L1 levels and TMB) (Jiang, et al. 2018). We found that TIDE scores were higher in the high-risk group and were associated with poorer ICIs efficacy. The sensitivity of different risk groups to some common chemotherapeutics and targeted therapy drugs varied. Therefore, we have reason to believe that despite the shortcomings of PDIS, it is undeniable that it provides a new potential perspective for the treatment of BC. PDIS is not only an effective biomarker for predicting immunotherapy response, but may also help guide devising chemotherapy regimens in BC patients.
Conclusions
In the present study, we described PIK3CA mutation characteristics based on TCGA data and cases from a local hospital. The PDIS allowed accurate risk stratification of BC patients with good prognostic predictive power and was validated in the GEO database. The nomogram constructed by combining PDIS and other prognostic factors showed higher clinical efficacy. We also found that the PDIS could assess immune infiltration and help select patients that may benefit from ICIs. Differences in response to immune and chemotherapeutic agents in PDIS-based risk groups suggested that PDIS will be a promising tool to guide clinical treatment decisions and enable individualized therapy.
Availability of data and materials
The data generated and analysed during the current study are available in the TCGA (https://portal.gdc.cancer.gov/) and GEO databases (https://www.ncbi.nlm.nih.gov/geo/).
Abbreviations
- TIME:
-
Tumor immune microenvironment
- BC:
-
Breast cancer
- TCGA:
-
The Cancer Genome Atlas
- PDIG:
-
PIK3CA mutation-driven immune gene
- PDIS:
-
PIKC3CA mutation-driven immune signature
- TMB:
-
Tumor mutation burden
- H-TMB:
-
High tumor mutation burden
- L-TMB:
-
Low tumor mutation burden
- GEO:
-
Gene Expression Omnibus
- PI3K:
-
Phosphatidylinositol 3-kinase
- HR:
-
Hormone receptor
- ICI:
-
Immune checkpoint inhibitor
- CTLA4:
-
Cytotoxic T lymphocyte-associated protein 4
- PD-1:
-
Programmed cell death-1
- PD-L1:
-
Programmed cell death ligand-1
- TIL:
-
Tumor-infiltrating lymphocyte
- DEG:
-
Differentially expressed gene
- GSEA:
-
Gene set enrichment analysis
- ssGSEA:
-
Single-sample gene set enrichment analysis
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
References
A, M.P., K, A.G., M, M.T., Κ. Ibragimova, M., A, V.V., Y, G.Z., D, V.P., S, A.K., and N, V.L. (2021) Investigation of somatic PIK3CA gene mutations in breast cancer patients. J Buon 26(3):747–752
Adams S, Gray RJ, Demaria S, Goldstein L, Perez EA, Shulman LN, Martino S, Wang M, Jones VE, Saphner TJ, Wolff AC, Wood WC, Davidson NE, Sledge GW, Sparano JA, Badve SS (2014) Prognostic value of tumor-infiltrating lymphocytes in triple-negative breast cancers from two phase III randomized adjuvant breast cancer trials: ECOG 2197 and ECOG 1199. J Clin Oncol 32(27):2959–2966. https://doi.org/10.1200/jco.2013.55.0491
Adams S, Loi S, Toppmeyer D, Cescon DW, De Laurentiis M, Nanda R, Winer EP, Mukai H, Tamura K, Armstrong A, Liu MC, Iwata H, Ryvo L, Wimberger P, Rugo HS, Tan AR, Jia L, Ding Y, Karantza V, Schmid P (2019) Pembrolizumab monotherapy for previously untreated, PD-L1-positive, metastatic triple-negative breast cancer: cohort B of the phase II KEYNOTE-086 study. Ann Oncol 30(3):405–411. https://doi.org/10.1093/annonc/mdy518
Bagbudar S, Karanlik H, Cabioglu N, Bayram A, Ibis K, Aydin E, Yavuz E, Onder S (2022) Prognostic implications of immune infiltrates in the breast cancer microenvironment: the role of expressions of CTLA-4, PD-1, and LAG-3. Appl Immunohistochem Mol Morphol 30(2):99–107. https://doi.org/10.1097/pai.0000000000000978
Bagchi S, Yuan R, Engleman EG (2021) Immune checkpoint inhibitors for the treatment of cancer: clinical impact and mechanisms of response and resistance. Annu Rev Pathol 16:223–249. https://doi.org/10.1146/annurev-pathol-042020-042741
Barbareschi M, Buttitta F, Felicioni L, Cotrupi S, Barassi F, Del Grammastro M, Ferro A, Dalla Palma P, Galligioni E, Marchetti A (2007) Different prognostic roles of mutations in the helical and kinase domains of the PIK3CA gene in breast carcinomas. Clin Cancer Res 13(20):6064–6069. https://doi.org/10.1158/1078-0432.ccr-07-0266
Barzaman K, Karami J, Zarei Z, Hosseinzadeh A, Kazemi MH, Moradi-Kalbolandi S, Safari E, Farahmand L (2020) Breast cancer: biology, biomarkers, and treatments. Int Immunopharmacol 84:106535. https://doi.org/10.1016/j.intimp.2020.106535
Baselga J, Im SA, Iwata H, Cortés J, De Laurentiis M, Jiang Z, Arteaga CL, Jonat W, Clemons M, Ito Y, Awada A, Chia S, Jagiełło-Gruszfeld A, Pistilli B, Tseng LM, Hurvitz S, Masuda N, Takahashi M, Vuylsteke P, Hachemi S, Dharan B, Di Tomaso E, Urban P, Massacesi C, Campone M (2017) Buparlisib plus fulvestrant versus placebo plus fulvestrant in postmenopausal, hormone receptor-positive, HER2-negative, advanced breast cancer (BELLE-2): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol 18(7):904–916. https://doi.org/10.1016/s1470-2045(17)30376-5
Baxevanis CN, Fortis SP, Perez SA (2021) The balance between breast cancer and the immune system: challenges for prognosis and clinical benefit from immunotherapies. Semin Cancer Biol 72:76–89. https://doi.org/10.1016/j.semcancer.2019.12.018
Byrne A, Savas P (2020) Tissue-resident memory T cells in breast cancer control and immunotherapy responses. Nat Rev Clin Oncol 17(6):341–348. https://doi.org/10.1038/s41571-020-0333-y
Chan TA, Yarchoan M, Jaffee E, Swanton C, Quezada SA, Stenzinger A, Peters S (2019) Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann Oncol 30(1):44–56. https://doi.org/10.1093/annonc/mdy495
Chang DY, Ma WL, Lu YS (2021) Role of alpelisib in the treatment of PIK3CA-mutated breast cancer: patient selection and clinical perspectives. Ther Clin Risk Manag 17:193–207. https://doi.org/10.2147/tcrm.s251668
Chen K, Pan G, Cheng G, Zhang F, Xu Y, Huang Z, Fan Y (2021a) Immune microenvironment features and efficacy of PD-1/PD-L1 blockade in non-small cell lung cancer patients with EGFR or HER2 exon 20 insertions. Thorac Cancer 12(2):218–226. https://doi.org/10.1111/1759-7714.13748
Chen L, Dong Y, Pan Y, Zhang Y, Liu P, Wang J, Chen C, Lu J, Yu Y, Deng R (2021b) Identification and development of an independent immune-related genes prognostic model for breast cancer. BMC Cancer 21(1):329. https://doi.org/10.1186/s12885-021-08041-x
Choi J, Gyamfi J, Jang H, Koo JS (2018) The role of tumor-associated macrophage in breast cancer biology. Histol Histopathol 33(2):133–145. https://doi.org/10.14670/hh-11-916
Collins NB, Al Abosy R, Miller BC, Bi K, Zhao Q, Quigley M, Ishizuka JJ, Yates KB, Pope HW, Manguso RT, Shrestha Y, Wadsworth M, Hughes T, Shalek AK, Boehm JS, Hahn WC, Doench JG, Haining WN (2022) PI3K activation allows immune evasion by promoting an inhibitory myeloid tumor microenvironment. J Immunother Cancer. https://doi.org/10.1136/jitc-2021-003402
De Mattos-Arruda L (2020) PIK3CA mutation inhibition in hormone receptor-positive breast cancer: time has come. ESMO Open. https://doi.org/10.1136/esmoopen-2020-000890
Deng L, Chen J, Zhong XR, Luo T, Wang YP, Huang HF, Yin LJ, Qiu Y, Bu H, Lv Q, Zheng H (2015) Correlation between activation of PI3K/AKT/mTOR pathway and prognosis of breast cancer in Chinese women. PLoS ONE 10(3):e0120511. https://doi.org/10.1371/journal.pone.0120511
Denkert C, von Minckwitz G, Darb-Esfahani S, Lederer B, Heppner BI, Weber KE, Budczies J, Huober J, Klauschen F, Furlanetto J, Schmitt WD, Blohmer JU, Karn T, Pfitzner BM, Kümmel S, Engels K, Schneeweiss A, Hartmann A, Noske A, Fasching PA, Jackisch C, van Mackelenbergh M, Sinn P, Schem C, Hanusch C, Untch M, Loibl S (2018) Tumour-infiltrating lymphocytes and prognosis in different subtypes of breast cancer: a pooled analysis of 3771 patients treated with neoadjuvant therapy. Lancet Oncol 19(1):40–50. https://doi.org/10.1016/s1470-2045(17)30904-x
Di Leo A, Johnston S, Lee KS, Ciruelos E, Lønning PE, Janni W, O’Regan R, Mouret-Reynier MA, Kalev D, Egle D, Csőszi T, Bordonaro R, Decker T, Tjan-Heijnen VCG, Blau S, Schirone A, Weber D, El-Hashimy M, Dharan B, Sellami D, Bachelot T (2018) Buparlisib plus fulvestrant in postmenopausal women with hormone-receptor-positive, HER2-negative, advanced breast cancer progressing on or after mTOR inhibition (BELLE-3): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol 19(1):87–100. https://doi.org/10.1016/s1470-2045(17)30688-5
Dirican E, Akkiprik M, Özer A (2016) Mutation distributions and clinical correlations of PIK3CA gene mutations in breast cancer. Tumour Biol 37(6):7033–7045. https://doi.org/10.1007/s13277-016-4924-2
Elfgen C, Reeve K, Moskovszky L, Güth U, Bjelic-Radisic V, Fleisch M, Tausch C, Varga Z (2019) Prognostic impact of PIK3CA protein expression in triple negative breast cancer and its subtypes. J Cancer Res Clin Oncol 145(8):2051–2059. https://doi.org/10.1007/s00432-019-02968-2
Gaynor N, Crown J, Collins DM (2022) Immune checkpoint inhibitors: key trials and an emerging role in breast cancer. Semin Cancer Biol 79:44–57. https://doi.org/10.1016/j.semcancer.2020.06.016
Hänzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform 14:7. https://doi.org/10.1186/1471-2105-14-7
Herberts C, Murtha AJ, Fu S, Wang G, Schönlau E, Xue H, Lin D, Gleave A, Yip S, Angeles A, Hotte S, Tran B, North S, Taavitsainen S, Beja K, Vandekerkhove G, Ritch E, Warner E, Saad F, Iqbal N, Nykter M, Gleave ME, Wang Y, Annala M, Chi KN, Wyatt AW (2020) Activating AKT1 and PIK3CA mutations in metastatic castration-resistant prostate cancer. Eur Urol 78(6):834–844. https://doi.org/10.1016/j.eururo.2020.04.058
Himmel ME, Saibil SD, Saltman AP (2020) Immune checkpoint inhibitors in cancer immunotherapy. CMAJ 192(24):E651. https://doi.org/10.1503/cmaj.191231
Hu H, Zhu J, Zhong Y, Geng R, Ji Y, Guan Q, Hong C, Wei Y, Min N, Qi A, Zhang Y, Li X (2021) PIK3CA mutation confers resistance to chemotherapy in triple-negative breast cancer by inhibiting apoptosis and activating the PI3K/AKT/mTOR signaling pathway. Ann Transl Med 9(5):410. https://doi.org/10.21037/atm-21-698
Jia Q, Wang A, Yuan Y, Zhu B, Long H (2022) Heterogeneity of the tumor immune microenvironment and its clinical relevance. Exp Hematol Oncol 11(1):24. https://doi.org/10.1186/s40164-022-00277-y
Jiang P, Gu S, Pan D, Fu J, Sahu A, Hu X, Li Z, Traugh N, Bu X, Li B, Liu J, Freeman GJ, Brown MA, Wucherpfennig KW, Liu XS (2018) Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med 24(10):1550–1558. https://doi.org/10.1038/s41591-018-0136-1
Jin N, Keam B, Cho J, Lee MJ, Kim HR, Torosyan H, Jura N, Ng PK, Mills GB, Li H, Zeng Y, Barbash Z, Tarcic G, Kang H, Bauman JE, Kim MO, VanLandingham NK, Swaney DL, Krogan NJ, Johnson DE, Grandis JR (2021) Therapeutic implications of activating noncanonical PIK3CA mutations in head and neck squamous cell carcinoma. J Clin Invest. https://doi.org/10.1172/jci150335
Kalinsky K, Jacks LM, Heguy A, Patil S, Drobnjak M, Bhanot UK, Hedvat CV, Traina TA, Solit D, Gerald W, Moynahan ME (2009) PIK3CA mutation associates with improved outcome in breast cancer. Clin Cancer Res 15(16):5049–5059. https://doi.org/10.1158/1078-0432.ccr-09-0632
Lai YL, Mau BL, Cheng WH, Chen HM, Chiu HH, Tzen CY (2008) PIK3CA exon 20 mutation is independently associated with a poor prognosis in breast cancer patients. Ann Surg Oncol 15(4):1064–1069. https://doi.org/10.1245/s10434-007-9751-7
Langfelder P, Horvath S (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinform 9:559. https://doi.org/10.1186/1471-2105-9-559
Lerma E, Catasus L, Gallardo A, Peiro G, Alonso C, Aranda I, Barnadas A, Prat J (2008) Exon 20 PIK3CA mutations decreases survival in aggressive (HER-2 positive) breast carcinomas. Virchows Arch 453(2):133–139. https://doi.org/10.1007/s00428-008-0643-4
Liu YR, Jiang YZ, Zuo WJ, Yu KD, Shao ZM (2014) PIK3CA mutations define favorable prognostic biomarkers in operable breast cancer: a systematic review and meta-analysis. Onco Targets Ther 7:543–552. https://doi.org/10.2147/ott.s60115
Liu Q, Ma JY, Wu G (2021) Identification and validation of a ferroptosis-related gene signature predictive of prognosis in breast cancer. Aging (albany NY) 13(17):21385–21399. https://doi.org/10.18632/aging.203472
Liu Z, Liu L, Weng S, Guo C, Dang Q, Xu H, Wang L, Lu T, Zhang Y, Sun Z, Han X (2022) Machine learning-based integration develops an immune-derived lncRNA signature for improving outcomes in colorectal cancer. Nat Commun 13(1):816. https://doi.org/10.1038/s41467-022-28421-6
Loi S, Michiels S, Lambrechts D, Fumagalli D, Claes B, Kellokumpu-Lehtinen PL, Bono P, Kataja V, Piccart MJ, Joensuu H, Sotiriou C (2013) Somatic mutation profiling and associations with prognosis and trastuzumab benefit in early breast cancer. J Natl Cancer Inst 105(13):960–967. https://doi.org/10.1093/jnci/djt121
Loibl S, von Minckwitz G, Schneeweiss A, Paepke S, Lehmann A, Rezai M, Zahm DM, Sinn P, Khandan F, Eidtmann H, Dohnal K, Heinrichs C, Huober J, Pfitzner B, Fasching PA, Andre F, Lindner JL, Sotiriou C, Dykgers A, Guo S, Gade S, Nekljudova V, Loi S, Untch M, Denkert C (2014) PIK3CA mutations are associated with lower rates of pathologic complete response to anti-human epidermal growth factor receptor 2 (her2) therapy in primary HER2-overexpressing breast cancer. J Clin Oncol 32(29):3212–3220. https://doi.org/10.1200/jco.2014.55.7876
Lv M, Mao Y, Song Y, Wang Y, Liu X, Wang X, Nie G, Wang H (2020) Clinical features and survival of single hormone receptor-positive breast cancer: a population-based study of 531,605 patients. Clin Breast Cancer 20(5):e589–e599. https://doi.org/10.1016/j.clbc.2020.04.010
Mangone FR, Bobrovnitchaia IG, Salaorni S, Manuli E, Nagai MA (2012) PIK3CA exon 20 mutations are associated with poor prognosis in breast cancer patients. Clinics (sao Paulo) 67(11):1285–1290. https://doi.org/10.6061/clinics/2012(11)11
Martinez LM, Robila V, Clark NM, Du W, Idowu MO, Rutkowski MR, Bos PD (2019) Regulatory T cells control the switch from in situ to invasive breast cancer. Front Immunol 10:1942. https://doi.org/10.3389/fimmu.2019.01942
Martínez-Sáez O, Chic N, Pascual T, Adamo B, Vidal M, González-Farré B, Sanfeliu E, Schettini F, Conte B, Brasó-Maristany F, Rodríguez A, Martínez D, Galván P, Rodríguez AB, Martinez A, Muñoz M, Prat A (2020) Frequency and spectrum of PIK3CA somatic mutations in breast cancer. Breast Cancer Res 22(1):45. https://doi.org/10.1186/s13058-020-01284-9
McGrail DJ, Pilié PG, Rashid NU, Voorwerk L, Slagter M, Kok M, Jonasch E, Khasraw M, Heimberger AB, Lim B, Ueno NT, Litton JK, Ferrarotto R, Chang JT, Moulder SL, Lin SY (2021) High tumor mutation burden fails to predict immune checkpoint blockade response across all cancer types. Ann Oncol 32(5):661–672. https://doi.org/10.1016/j.annonc.2021.02.006
Mehta AK, Kadel S, Townsend MG, Oliwa M, Guerriero JL (2021) Macrophage biology and mechanisms of immune suppression in breast cancer. Front Immunol 12:643771. https://doi.org/10.3389/fimmu.2021.643771
Miller TJ, Anyaegbu CC, Lee-Pullen TF, Spalding LJ, Platell CF, McCoy MJ (2021) PD-L1+ dendritic cells in the tumor microenvironment correlate with good prognosis and CD8+ T cell infiltration in colon cancer. Cancer Sci 112(3):1173–1183. https://doi.org/10.1111/cas.14781
Mosele F, Stefanovska B, Lusque A, Tran Dien A, Garberis I, Droin N, Le Tourneau C, Sablin MP, Lacroix L, Enrico D, Miran I, Jovelet C, Bièche I, Soria JC, Bertucci F, Bonnefoi H, Campone M, Dalenc F, Bachelot T, Jacquet A, Jimenez M, André F (2020) Outcome and molecular landscape of patients with PIK3CA-mutated metastatic breast cancer. Ann Oncol 31(3):377–386. https://doi.org/10.1016/j.annonc.2019.11.006
Nicolini A, Ferrari P, Duffy MJ (2018) Prognostic and predictive biomarkers in breast cancer: past, present and future. Semin Cancer Biol 52(Pt 1):56–73. https://doi.org/10.1016/j.semcancer.2017.08.010
Oliver AJ, Davey AS, Keam SP, Mardiana S, Chan JD, von Scheidt B, Beavis PA, House IG, Van Audernaerde JR, Darcy PK, Kershaw MH, Slaney CY (2019) Tissue-specific tumor microenvironments influence responses to immunotherapies. Clin Transl Immunol 8(11):e1094. https://doi.org/10.1002/cti2.1094
Pang H, Flinn R, Patsialou A, Wyckoff J, Roussos ET, Wu H, Pozzuto M, Goswami S, Condeelis JS, Bresnick AR, Segall JE, Backer JM (2009) Differential enhancement of breast cancer cell motility and metastasis by helical and kinase domain mutations of class IA phosphoinositide 3-kinase. Cancer Res 69(23):8868–8876. https://doi.org/10.1158/0008-5472.can-09-1968
Pang B, Cheng S, Sun SP, An C, Liu ZY, Feng X, Liu GJ (2014) Prognostic role of PIK3CA mutations and their association with hormone receptor expression in breast cancer: a meta-analysis. Sci Rep 4:6255. https://doi.org/10.1038/srep06255
Pang J, Pan H, Yang C, Meng P, Xie W, Li J, Li Y, Xiao SY (2021) Prognostic value of immune-related multi-IncRNA signatures associated with tumor microenvironment in esophageal cancer. Front Genet 12:722601. https://doi.org/10.3389/fgene.2021.722601
Papaxoinis G, Kotoula V, Alexopoulou Z, Kalogeras KT, Zagouri F, Timotheadou E, Gogas H, Pentheroudakis G, Christodoulou C, Koutras A, Bafaloukos D, Aravantinos G, Papakostas P, Charalambous E, Papadopoulou K, Varthalitis I, Efstratiou I, Zaramboukas T, Patsea H, Scopa CD, Skondra M, Kosmidis P, Pectasides D, Fountzilas G (2015) Significance of PIK3CA mutations in patients with early breast cancer treated with adjuvant chemotherapy: a Hellenic Cooperative Oncology Group (HeCOG) study. PLoS ONE 10(10):e0140293. https://doi.org/10.1371/journal.pone.0140293
Peng Y, Yu H, Jin Y, Qu F, Ren H, Tang Z, Zhang Y, Qu C, Zong B, Liu S (2021) Construction and validation of an immune infiltration-related gene signature for the prediction of prognosis and therapeutic response in breast cancer. Front Immunol 12:666137. https://doi.org/10.3389/fimmu.2021.666137
Ren X, Ma L, Wang N, Zhou R, Wu J, Xie X, Zhang H, Liu D, Ma X, Dang C, Kang H, Zhou Z (2021) Antioxidant gene signature impacts the immune infiltration and predicts the prognosis of kidney renal clear cell carcinoma. Front Genet 12:721252. https://doi.org/10.3389/fgene.2021.721252
Ren X, Cui H, Wu J (2022) Identification of a combined apoptosis and hypoxia gene signature for predicting prognosis and immune infiltration in breast cancer. Cancer Med. https://doi.org/10.1002/cam4.4755
Rupp T, Genest L, Babin D, Legrand C, Hunault M, Froget G, Castagné V (2022) Anti-CTLA-4 and anti-PD-1 immunotherapies repress tumor progression in preclinical breast and colon model with independent regulatory T cells response. Transl Oncol 20:101405. https://doi.org/10.1016/j.tranon.2022.101405
Sabine VS, Crozier C, Brookes CL, Drake C, Piper T, van de Velde CJ, Hasenburg A, Kieback DG, Markopoulos C, Dirix L, Seynaeve C, Rea DW, Bartlett JM (2014) Mutational analysis of PI3K/AKT signaling pathway in tamoxifen exemestane adjuvant multinational pathology study. J Clin Oncol 32(27):2951–2958. https://doi.org/10.1200/jco.2013.53.8272
Salemme V, Centonze G, Cavallo F, Defilippi P, Conti L (2021) The crosstalk between tumor cells and the immune microenvironment in breast cancer: implications for immunotherapy. Front Oncol 11:610303. https://doi.org/10.3389/fonc.2021.610303
Samuels Y, Diaz LA Jr, Schmidt-Kittler O, Cummins JM, Delong L, Cheong I, Rago C, Huso DL, Lengauer C, Kinzler KW, Vogelstein B, Velculescu VE (2005) Mutant PIK3CA promotes cell growth and invasion of human cancer cells. Cancer Cell 7(6):561–573. https://doi.org/10.1016/j.ccr.2005.05.014
Schmid P, Cruz C, Braiteh FS, Eder JP, Tolaney S, Kuter I, Nanda R, Chung C, Cassier P, Delord J-P, Gordon M, Li Y, Liu B, O’Hear C, Fasso M, Molinero L, Emens LA (2017) Abstract 2986: Atezolizumab in metastatic TNBC (mTNBC): long-term clinical outcomes and biomarker analyses. Cancer Res 77(13 Supplement):2986–2986. https://doi.org/10.1158/1538-7445.am2017-2986
Seiler M, Huang CC, Szalma S, Bhanot G (2010) ConsensusCluster: a software tool for unsupervised cluster discovery in numerical data. OMICS 14(1):109–113. https://doi.org/10.1089/omi.2009.0083
Shao W, Yang Z, Fu Y, Zheng L, Liu F, Chai L, Jia J (2021) The pyroptosis-related signature predicts prognosis and indicates immune microenvironment infiltration in gastric cancer. Front Cell Dev Biol 9:676485. https://doi.org/10.3389/fcell.2021.676485
Shash LS, Kamal KAM, Abd Raboh NM (2021) Exploring the role CD163-labeled TAMs and FOXP3-labeled Tregs in different types of breast cancer: reflections and putative benefits. Indian J Pathol Microbiol 64(1):28–37. https://doi.org/10.4103/ijpm.ijpm_210_20
Siegel RL, Miller KD (2022) Cancer statistics 2022. CA Cancer J Clin 72(1):7–33. https://doi.org/10.3322/caac.21708
Sun L, Wu A, Bean GR, Hagemann IS, Lin CY (2021) Molecular testing in breast cancer: current status and future directions. J Mol Diagn 23(11):1422–1432. https://doi.org/10.1016/j.jmoldx.2021.07.026
Sung H, Ferlay J, Siegel RL (2021) Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3):209–249. https://doi.org/10.3322/caac.21660
Taylor NA, Vick SC, Iglesia MD, Brickey WJ, Midkiff BR, McKinnon KP, Reisdorf S, Anders CK, Carey LA, Parker JS, Perou CM, Vincent BG, Serody JS (2017) Treg depletion potentiates checkpoint inhibition in claudin-low breast cancer. J Clin Invest 127(9):3472–3483. https://doi.org/10.1172/jci90499
Tian Z, He W, Tang J, Liao X, Yang Q, Wu Y (2020) Identification of important modules and biomarkers in breast cancer based on WGCNA. OncoTargets Ther 13:6805–6817. https://doi.org/10.2147/ott.s258439
Ugai T, Zhao M, Shimizu T, Akimoto N, Shi S, Takashima Y, Zhong R, Lau MC, Haruki K, Arima K, Fujiyoshi K, Langworthy B, Masugi Y, da Silva A, Nosho K, Baba Y, Song M, Chan AT, Wang M, Meyerhardt JA, Giannakis M, Väyrynen JP (2021) Association of PIK3CA mutation and PTEN loss with expression of CD274 (PD-L1) in colorectal carcinoma. Oncoimmunology 10(1):1956173. https://doi.org/10.1080/2162402x.2021.1956173
Wan Q, Liu C, Liu C, Liu W, Wang X, Wang Z (2020) Discovery and validation of a metastasis-related prognostic and diagnostic biomarker for melanoma based on single cell and gene expression datasets. Front Oncol 10:585980. https://doi.org/10.3389/fonc.2020.585980
Wang J, Zhang X, Li J, Ma X, Feng F, Liu L, Wu J, Sun C (2020) ADRB1 was identified as a potential biomarker for breast cancer by the co-analysis of tumor mutational burden and immune infiltration. Aging (albany NY) 13(1):351–363. https://doi.org/10.18632/aging.104204
Xu Q, Chen S, Hu Y, Huang W (2021) Landscape of immune microenvironment under immune cell infiltration pattern in breast cancer. Front Immunol 12:711433. https://doi.org/10.3389/fimmu.2021.711433
Yeo SK, Guan JL (2017) Breast cancer: multiple subtypes within a tumor? Trends Cancer 3(11):753–760. https://doi.org/10.1016/j.trecan.2017.09.001
Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW, Levine DA, Carter SL, Getz G, Stemke-Hale K, Mills GB, Verhaak RG (2013) Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun 4:2612. https://doi.org/10.1038/ncomms3612
Yu G, Wang LG, Han Y, He QY (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16(5):284–287. https://doi.org/10.1089/omi.2011.0118
Zardavas D, Phillips WA, Loi S (2014) PIK3CA mutations in breast cancer: reconciling findings from preclinical and clinical data. Breast Cancer Res 16(1):201. https://doi.org/10.1186/bcr3605
Zardavas D, Te Marvelde L, Milne RL, Fumagalli D, Fountzilas G, Kotoula V, Razis E, Papaxoinis G, Joensuu H, Moynahan ME, Hennessy BT, Bieche I, Saal LH, Stal O, Iacopetta B, Jensen JD, O’Toole S, Lopez-Knowles E, Barbaraeschi M, Noguchi S, Azim HA Jr, Lerma E, Bachelot T, Wang Q, Perez-Tenorio G, Can de Velde CJH, Rea DW, Sabine V, Bartlett JMS, Sotiriou C, Michiels S, Loi S (2018) Tumor PIK3CA genotype and prognosis in early-stage breast cancer: a pooled analysis of individual patient data. J Clin Oncol 36(10):981–990. https://doi.org/10.1200/jco.2017.74.8301
Zhang B, Horvath S (2005) A general framework for weighted gene co-expression network analysis. Stat Appl Genet Mol Biol. https://doi.org/10.2202/1544-6115.1128
Zhang D, Zheng Y, Yang S, Li Y, Wang M, Yao J, Deng Y, Li N, Wei B, Wu Y, Zhu Y, Li H, Dai Z (2020) Identification of a novel glycolysis-related gene signature for predicting breast cancer survival. Front Oncol 10:596087. https://doi.org/10.3389/fonc.2020.596087
Zhang Y, Di X, Chen G, Liu J, Zhang B, Feng L, Cheng S, Wang Y (2021) An immune-related signature that to improve prognosis prediction of breast cancer. Am J Cancer Res 11(4):1267–1285
Zheng M (2022) Tumor mutation burden for predicting immune checkpoint blockade response: the more, the better. J Immunother Cancer. https://doi.org/10.1136/jitc-2021-003087
Zhu Y, Zhu X, Tang C, Guan X, Zhang W (2021) Progress and challenges of immunotherapy in triple-negative breast cancer. Biochim Biophys Acta Rev Cancer 1876(2):188593. https://doi.org/10.1016/j.bbcan.2021.188593
Acknowledgements
We would like to thank TCGA and GEO databases for their support for the original data, as well as Editage (www.editage.cn) for its language embellishment.
Funding
This study was supported by the Key Research and Development Plan of Shaanxi Provincial Department of Science and Technology (No. 2022SF-001) and the International Science and Technology Cooperation Program Project of Shaanxi Province, China (2022KW-01).
Author information
Authors and Affiliations
Contributions
HK and XM put forward the central idea and guided the research design. XR, HC and LD retrieved and analyzed original data. LC, DL, WY and XZ interpreted the results. XR wrote the first draft of the paper and then HK and XM modified it. All authors discussed the results and revised the manuscript. All authors reviewed and approved the submission of final manuscripts.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests in this section.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ren, X., Cui, H., Dai, L. et al. PIK3CA mutation-driven immune signature as a prognostic marker for evaluating the tumor immune microenvironment and therapeutic response in breast cancer. J Cancer Res Clin Oncol 150, 119 (2024). https://doi.org/10.1007/s00432-024-05626-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00432-024-05626-4