Abstract
Idiopathic pulmonary fibrosis (IPF) is a chronic, progressive form of pulmonary fibrosis of unknown etiology. Despite ongoing research, there is currently no cure for this disease. Recent studies have highlighted the significance of competitive endogenous RNA (ceRNA) regulatory networks in IPF development. Therefore, this study investigated the ceRNA network associated with IPF pathogenesis. We obtained gene expression datasets (GSE32538, GSE32537, GSE47460, and GSE24206) from the Gene Expression Omnibus (GEO) database and analyzed them using bioinformatics tools to identify differentially expressed messenger RNAs (DEmRNAs), microRNAs (DEmiRNAs), and long non-coding RNAs (DElncRNA). For DEmRNAs, we conducted an enrichment analysis, constructed protein–protein interaction networks, and identified hub genes. Additionally, we predicted the target genes of differentially expressed mRNAs and their interacting long non-coding RNAs using various databases. Subsequently, we screened RNA molecules with ceRNA regulatory relations in the lncACTdb database based on the screening results. Furthermore, we performed disease and functional enrichment analyses and pathway prediction for miRNAs in the ceRNA network. We also validated the expression levels of candidate DEmRNAs through quantitative real-time reverse transcriptase polymerase chain reaction and analyzed the correlation between the expression of these candidate DEmRNAs and the percent predicted pre-bronchodilator forced vital capacity [%predicted FVC (pre-bd)]. We found that three ceRNA regulatory axes, specifically KCNQ1OT1/XIST/NEAT1-miR-20a-5p-ITGB8, XIST-miR-146b-5p/miR-31-5p- MMP16, and NEAT1-miR-31-5p-MMP16, have the potential to significantly affect IPF progression. Further examination of the underlying regulatory mechanisms within this network enhances our understanding of IPF pathogenesis and may aid in the identification of diagnostic biomarkers and therapeutic targets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Idiopathic pulmonary fibrosis (IPF) is a diffuse lung disease characterized by progressive lung injury and fibrosis, significantly impairing lung structure and function (King et al. 2011). The exact cause of IPF remains unknown, and its prognosis is generally poor, with a median survival rate of only 2–3 years (Kreuter et al. 2017). The diagnosis of IPF relies on imaging techniques and histopathology, specifically identifying usual interstitial pneumonia manifestations (American Thoracic Society 2000). Currently, the available therapeutic options for IPF are limited, and nintedanib and pirfenidone are the only approved treatments. However, these drugs can only delay disease progression and provide limited lung function improvement without curing IPF (Dempsey et al. 2021). Challenges in diagnosing and treating IPF stem from our incomplete understanding of the disease and the absence of precise biomarkers and therapeutic targets. Therefore, there is an urgent need to develop novel diagnostic and therapeutic approaches. Recent advances in high-throughput sequencing technology have shed light on the significant role of non-coding RNAs (ncRNAs) in the pathogenesis of IPF, offering new insights into the disease mechanisms and potential targets for therapeutic intervention.
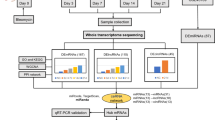
ncRNAs such as microRNAs (miRNAs) and long non-coding RNAs (lncRNAs) are transcribed from the human genome but do not produce proteins (Kaikkonen et al. 2011). However, lncRNAs have been identified as significant disease regulators, affecting the development and treatment of various diseases by controlling gene expression and cellular processes (Morris and Mattick 2014). Specifically, lncRNAs regulate gene expression at the transcriptional and translational levels through different mechanisms, whereas miRNAs primarily regulate the expression of protein-coding genes by inhibiting messenger RNAs (mRNAs) (Bartel 2018). A growing body of research has shown that abnormally expressed ncRNAs play crucial roles in the pathogenesis of IPF (Hadjicharalambous and Lindsay 2020; Zhang et al. 2021a). For instance, miRNAs, such as miR-21 and miR-33, are abnormally expressed in the lung tissues of patients with IPF and influence disease progression by regulating fibroblast activation and inflammation (Ahangari et al. 2023; Liu et al. 2010). Another study has confirmed that lncRNA MALAT1 was highly expressed in patients with IPF and affected the development of IPF by regulating gene expression, inflammation, and other pathways (Lai et al. 2022). Additionally, studies have identified complex regulatory interactions among ncRNA molecules. lncRNAs can interact with miRNAs as competing endogenous RNAs (ceRNAs) or function as reservoirs, thereby affecting gene expression (Zhang et al. 2021c). The ceRNA mechanism is particularly important in the pathogenesis of IPF, in which RNA molecules, including mRNAs, miRNAs, and lncRNAs, compete for the same miRNA-binding sites, thereby influencing their expression levels (Song et al. 2014; Hadjicharalambous et al. 2018). Experiments have demonstrated that certain lncRNAs can act as ceRNAs, sequester miRNAs, and affect the expression of target genes, ultimately leading to IPF. Song et al. showed that lncRNA H19 binds to miR-196a through a ceRNA mechanism and increases the expression of COL1A1, resulting in pulmonary fibrosis (Lu et al. 2018). However, the specific regulatory mechanisms of lncRNAs, miRNAs, and mRNAs in the development of IPF through the ceRNA mechanism are still unclear. Further studies are needed to uncover how the molecules in these networks affect cellular functions and signaling pathways. In addition, current studies on IPF often rely on cell lines or animal models that may not fully replicate the complex pathology of IPF in humans. Therefore, additional human lung tissue specimens must be collected to develop more accurate models. Furthermore, most studies lack clinical value assessment and validation of abnormally expressed ncRNAs, which may overlook important insights into the pathogenesis of IPF and limit the development of more effective therapeutic approaches. Hence, in this study, we used high-throughput sequencing technology and bioinformatic analysis to identify abnormally expressed lncRNAs, miRNAs, mRNAs, and their interacting ceRNA networks in IPF. Disease, functional, and pathway enrichment analyses were conducted to validate the gene expression of ncRNA molecules in the networks. We analyzed the correlation between these networks and human clinical features to identify ceRNA networks with significant regulatory roles in IPF. In conclusion, we examined three ceRNA regulatory axes. The workflow employed in our study is depicted in Fig. 1. This study establishes a solid theoretical foundation for future functional biology experiments. Additionally, the identification of aberrant ncRNA molecules and their interaction networks in IPF provides valuable insights into disease pathogenesis and facilitates the development of novel diagnostic and therapeutic strategies.
Materials and Methods
Data Sources
We searched the Gene Expression Omnibus (GEO) genomics data repository (http://www.ncbi.nlm.nih.gov/geo) to identify gene expression datasets relevant to IPF. The search was performed using the keywords “idiopathic pulmonary fibrosis,” “Homo sapiens,” and “Expression profiling by array” or “Non-coding RNA profiling by array.” Following a systematic review, two mRNA expression datasets (GSE32537 and GSE47460) were selected and downloaded. GSE32537 and GSE47460 were based on the GPL6244 and GPL6480 datasets, respectively. Additionally, one lncRNA expression dataset (GSE24206) was downloaded based on GPL570. One miRNA expression dataset (GSE32538) was selected and downloaded based on GPL8786. The specific dataset information is presented in Table 1.
Differentially Expressed Genes (DEG) Identification
The GEO2R tool, available at http://www.ncbi.nlm.nih.gov/geo/geo2r, was used to identify DEGs between patients with IPF and healthy control tissues. |log fold change|> 1 and adjusted P-value < 0.05 were considered statistically significant.
Enrichment analysis of the Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) for DEGs
The KEGG database is a valuable resource for elucidating the overarching characteristics and effects of biological systems (Kanehisa et al. 2016). GO is a prominent bioinformatics tool that facilitates gene annotation based on its functional roles in biological processes (BP), molecular functions (MF), and cellular components (CC) (Gene Ontology Consortium 2015). To identify the distinctive attributes of DEGs, the Metascape platform was employed (Zhou et al. 2019), with specific screening criteria of a minimum overlap equal to 3 and minimum richness equal to 1.5. Statistical significance was set at 0.01.
Construction of Protein–Protein Interaction (PPI) network and screening of hub genes
The filtered DEGs were submitted to the STRING database (http://www.bork.embl-heidelberg.de/STRING/) for further analyses (von Mering et al. 2003). PPI pairs with a combined score greater than 0.4 were extracted. The degrees of all nodes were calculated using the cytoHubba plugin in Cytoscape version 3.7.2 (http://www.cytoscape.org/). In this experiment, the genes with the 10 highest degree values were identified as hub genes.
Establishment of the ceRNA Network
The construction of this network involved the following three steps. First, an mRNA-miRNA network was constructed by identifying DEmiRNAs from the GSE32538 dataset and predicting their downstream target genes using the miRTarBase database (Huang et al. 2020). The predicted genes were then compared with DEmRNAs from the GSE32537 and GSE47460 datasets to identify candidate target genes. A miRNA-mRNA regulatory network was established by considering the regulatory relationships between miRNAs and mRNAs. Second, a miRNA-lncRNA network was constructed using the Starbase (Li et al. 2014) and Lncbase databases (Karagkouni et al. 2020) to predict lncRNAs that may interact with the DEmiRNAs. These predicted lncRNAs were then compared with DElncRNAs from the GSE24206 dataset to obtain a list of candidate target lncRNAs through intersection analysis. Finally, candidate miRNA-mRNA and miRNA-lncRNA networks were screened for ceRNA networks using the lncACTdb database (Wang et al. 2015, 2019, 2022). The intersection of DEmRNAs, mRNAs, and predicted lncRNAs was integrated and processed using the Cytoscape software.
Animal Model
Male C57BL/6J mice were procured from SiPeiFu Biotechnology (http://www.spfbiotech.com). Following a one-week acclimatization period, 12 mice were randomly assigned to two groups using a random number table, with six mice in each group. The groups were designated as control [phosphate-buffered saline (PBS)] and bleomycin (BLM) groups. On the day of modeling, all mice were intraperitoneally injected with pentobarbital sodium (50mg/kg) to induce anesthesia. Upon achieving anesthesia, mice in the BLM group were administered an intratracheal injection of 1.5 U/kg of BLM dissolved in 50 μL of PBS, while the PBS group received an equivalent volume of PBS solution. After 28 days, humane euthanasia was performed on all mice by intraperitoneal injection of pentobarbital sodium (150mg/kg), and lung tissues were collected for further experimentation. The study was conducted in accordance with the Declaration of Helsinki and approved by the Animal Experimental Inspection Form of Guizhou Medical University (number:12304822).
Cell Culture
The human type II alveolar epithelial (A549) and mouse embryonic fibroblast (3T3) cell lines were obtained from Procell (China) and cultured according to the manufacturer’s instructions. A549 cells were cultured in Ham’s F-12 K medium (Gibco, Thermo Fisher Scientific, USA), whereas 3T3 cells were cultured in Dulbecco’s Modified Eagle’s medium (Gibco, Thermo Fisher Scientific, USA). Both media were supplemented with 10% fetal bovine serum (Gibco, Thermo Fisher Scientific, USA) and 1% penicillin–streptomycin. The cells were maintained at 37°C and 5% CO2. The transforming growth factor beta 1 (TGF-β1) treatment group was exposed to TGF-β1 (10 ng/mL; MedChemExpress, USA) to induce lung fibroblast activation, while the control group received an equivalent volume of solvent. After 72 h of culture, the cells were harvested.
Quantitative Real-Time Reverse Transcriptase Polymerase Chain Reaction (qRT-PCR)
Total RNA was extracted from 3T3 cells, A549 cells, and mouse lung tissue using the TRIzol reagent (Takara, Japan) according to the manufacturer’s instructions. Subsequently, complementary DNA was synthesized using a Primer Script RT kit (Takara, Japan). Real-time PCR was performed using the Quant Studio 1 real-time PCR system (Thermo Fisher Scientific, USA) and TB Green Premix Ex TaqTM kit (Takara, Japan). Glyceraldehyde-3-phosphate dehydrogenase was used as a stable internal control for gene expression analysis in lung tissues during pulmonary fibrosis. Primer sequences used in the experiments are listed in Tables 2 and 3. Primer sequences for the mRNAs were obtained from Sangon Biotech (Shanghai, China).
Statistical Analysis
The 2−ΔΔCt method was employed to determine the relative expression levels of the relevant genes. The obtained data are presented as mean ± standard error of the mean and analyzed using GraphPad PRISM 9.5.0 software (USA). Differences between the two groups were determined using a t-test. p < 0.05 was regarded as statistically significant. Correlation among the candidate DEmRNAs was evaluated using Pearson's correlation analysis.
Results
Identification of DEmRNAs
After conducting a reanalysis of the two gene expression profiles, it was determined that GSE32537 exhibited 313 upregulated and 118 downregulated genes with significant differences in their expression. Volcano plots, uniform manifold approximations and projections (UMAPs), and data boxplots were used to visually represent these findings (Fig. 2a–c). Similarly, GSE47460 displayed 826 upregulated and 544 downregulated genes (Fig. 2d–f). Subsequently, bioinformatic tools were used to identify common DEmRNAs between the GSE32537 and GSE47460 datasets. The analysis revealed 176 upregulated and 60 downregulated genes, as shown in Fig. 4 (a, b, c). The demographic characteristics of the GSE47460 dataset are presented in Table 4.
Identification of DEmiRNAs
Four genes were identified as being upregulated, whereas 42 miRNAs were found to be downregulated in the GSE32538 dataset. Volcano plots, UMAPs, and data boxplots were employed (Fig. 3a–c).
Identification of DElncRNAs
We identified 157 upregulated and 79 downregulated lncRNAs in the GSE24206 dataset. To visually represent our findings, we utilized Volcano plots, UMAPs, and data boxplots, as shown in Fig. 3d–f.
Identification of Hub Genes and Construction of PPI Network of DEmRNAs
The MCODE plugin for Cytoscape was used to identify densely connected regions within the PPI network. The resulting networks are shown in Fig. 4d. The top 10 genes that exhibited the highest number of interactions, referred to as hub genes, were identified from these networks. These hub genes included DNAH9, DNAI1, TTC25, ARMC4, CCDC39, RSPH4A, DRC1, RSPH1, LRRC6, and DNAH5, as shown in Fig. 4e.
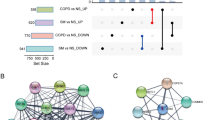
Venn diagrams of the DEmRNAs, a cross areas indicate the upregulated DEmRNAs, b cross areas indicate the downregulated DEmRNAs, and c cross areas indicate the common DEmRNAs, Enrichment analysis, PPI networks, and hub genes about DEmRNAS. d Construction of PPI network of DEmRNAs. e Identified hub genes. (DEmRNA differentially expressed mRNAs, PPI protein–protein interaction, KEGG Kyoto Encyclopedia of Genes and Genomes)
Construction of ceRNA Network
In this study, DEmiRNAs were used to screen miRNA-targeted mRNAs in the miRTarBase database. The identified miRNA-targeted mRNAs were compared with DEmRNAs previously identified in the GSE32537 and GSE47460 datasets. This comparison led to the identification of 14 candidate mRNAs (Fig. 5a). GO and KEGG pathway enrichment analyses indicated that the candidate mRNAs were significantly associated with several BP, such as extracellular structure organization and cell–matrix adhesion. Enrichment of MFs was observed in various binding activities, such as binding with cell adhesion molecules. Furthermore, the analysis revealed enriched CC in the collagen-containing extracellular matrix, endoplasmic reticulum lumen, blood microparticles, platelet alpha granules, and platelet alpha granule lumens (Fig. 5b). Additionally, KEGG pathway analysis demonstrated significant enrichment of DEmRNAs in four pathways: platelet activation, Wnt signaling, cell adhesion molecules, and complement and coagulation cascades (Fig. 5c). Furthermore, the interaction between DEmiRNAs and lncRNAs was predicted using StarBase and lncBase v3.0 databases. The results of this prediction were then cross-referenced with the DElncRNAs to obtain a list of candidate lncRNAs (Fig. 5d, e). Candidate miRNA-mRNA and miRNA-lncRNA networks were screened for ceRNA networks using the lncACTdb database. The DEmiRNAs, mRNAs, and predicted lncRNAs involved in the ceRNA network are presented in Table 5. For instance, lncRNAs such as KCNQ1OT1, XIST, and NEAT1 are involved in different ceRNA regulatory axes, and the same mRNA is regulated by multiple lncRNAs, including ITGB8, MMP16, and COL3A1. Specific ncRNAs were associated with the following ceRNA regulatory axes: KCNQ1OT1-miR-130a-3p/miR-130b-3p-PDGFRA, KCNQ1OT1-miR-15b-5p-PEBP4/TRIM29, KCNQ-1OT1/XIST/NEAT1/GABPB1-AS1-miR-20a-5p-ITGB8, XIST-miR-146b-5p/miR-31-5p-MMP16, NEAT1-miR-205-5p-ACSL1, THUMPD3-AS1/TUG1-miR-29a-3p-COL3A1, TRG-AS1-let-7b-59-COL3A1, and NEAT1/XIST-miR-31-5p-MMP16/SPRY4.To visually represent the results, a Cytoscape network and Sankey plots were constructed (Fig. 6a-b).
miRNA-mRNA network. a Cross areas indicate the candidate DEmRNAs in the miRNA-mRNA network. b and c Bubble plot of BP, CC, MF and KEGG pathway analysis for candidate DEmRNAs in the miRNA-mRNA network. d Cross areas indicate the candidate DELncRNAs in the miRNA-lncRNA network. e The Volcano plots of the candidate DElncRNAs. (miRNA microRNA, mRNA messenger, BP biological processes, CC cellular components, MF molecular functions, KEGG Kyoto Encyclopedia of Genes and Genomes, lncRNA long non-coding RNA, DELncRNA differentially expressed lncRNA)
Enrichment Analysis of Candidate DEmiRNAs
The results of GO and KEGG pathway enrichment analyses demonstrated a significant association between the candidate DEmiRNAs involved in the ceRNA network and 15 diseases, including breast neoplasms, acute myeloid leukemia, heart failure, and lung carcinoma (Fig. 7a). Additionally, the candidate DEmiRNAs were significantly enriched in 10 MF, including inflammation and apoptosis (Fig. 7b). KEGG pathway analysis further revealed that the candidate DEmiRNAs were primarily enriched in the focal adhesion, Hippo signaling, and FoxO signaling pathways (Fig. 7c).
Correlation Analysis Between the Expression of Candidate DEmRNAs and %Predicted FVC (pre-bd)
To evaluate the clinical significance of the DEmRNAs, we analyzed to determine whether there was a relation between the expression levels of these DEmRNAs and the %predicted FVC (pre-bd). This analysis used the clinical information available in the GSE47460 database and the gene expression data of candidate DEmRNAs. The findings revealed that the expression of CLDN1, COL3A1, MMP16, SFRP2, SPRY4, and ITGB8 correlated with the %predicted FVC (pre-bd). Specifically, SPRY4 expression was positively correlated with %predicted FVC (pre-bd), whereas the remaining genes were negatively correlated with %predicted FVC (pre-bd). No significant correlation was observed between the expression of ACSL1, FGG, PDGFRA, and TRIM29 and %predicted FVC (pre-bd) (Fig. 8).
Validation of Candidate DEmRNAs Using qRT-PCR
In this study, we used qRT-PCR to evaluate the expression of candidate DEmRNAs in 3T3 and A549 cells following activation induced by TGF-β1. Our findings revealed that the expression levels of ACSL1, FGG, SPRY4, and PEBP were significantly decreased in the TGF-β1 group compared with the control group. Conversely, the expression levels of CLDN1, ITGB8, PDGFRA, MMP16, TRIM29, SFRP2, and COL3A1 were significantly increased (Fig. 9a-b). Similar trends were observed in the BLM-treated mice (Fig. 9c).
Expression levels of candidate DEmRNAs were assessed using qRT-PCR. (a and b): In 3T3 and A549 cells, the expression of ACSL1, FGG, SPRY4, and PEBP was downregulated in the TGF-β1 group compared with the Control group, while the expression of CLDN1, ITGB8, PDGFRA, MMP16, TRIM29, SFRP2, and COL3A1 was upregulated. (c): In C57BL mice, the expression of ACSL1, FGG, SPRY4, and PEBP was downregulated in the BLM group compared with the PBS group, while the expression of CLDN1, ITGB8, PDGFRA, MMP16, TRIM29, SFRP2, and COL3A1 was upregulated. (**P < 0.01, *** P < 0.001, **** P < 0.001). (qRT-PCR quantitative real-time reverse transcriptase polymerase chain reaction, PBS phosphate-buffered saline, BLM bleomycin)
Discussion
IPF is a highly lethal and rapidly progressive disease with increasing incidence worldwide. Its occurrence is now comparable to that of cancers, such as gastric, liver, testicular, and cervical cancers (Hutchinson et al. 2015). Early diagnosis and treatment options for IPF are limited, and there is currently no known cure. Therefore, it is crucial to identify specific diagnostic markers and therapeutic targets in patients with IPF. In recent years, there has been a growing recognition of the role of ncRNAs in the development of various diseases, including IPF. It has been discovered that lncRNAs can act as ceRNAs in the pathogenesis of IPF. The ceRNA mechanism establishes a connection between mRNAs and ncRNAs that encode proteins, thereby influencing the occurrence and progression of IPF. However, the role of lncRNA-mediated ceRNA networks in the pathogenesis of IPF has not been well studied, and the detailed mechanism requires further exploration. Therefore, to elucidate the potential pathogenesis of IPF, we constructed a ceRNA regulatory network based on four microarray datasets.
This study identified seven candidate DElncRNAs. Among them, TRG-AS1, THUMPD3-AS1, and GABPB1-AS1 have not been studied in fibrotic diseases but have been documented in other conditions such as lung cancer, gastric cancer, and osteosarcoma. These lncRNAs have been shown to regulate cell proliferation, invasion, epithelial-mesenchymal transition, and inflammatory response (Chen et al. 2022; Zhang et al. 2021b, 2023). Additionally, KCNQ1OT1, XIST, NEAT1, and TUG1 were found to be associated with multiple ceRNA regulatory networks, potentially exerting crucial roles in the pathophysiology of idiopathic pulmonary fibrosis (IPF) by affecting numerous target genes and participating in different biological processes or signaling pathways. While there are reports of these DElncRNAs in fibrotic diseases, for instance, Yang et al. demonstrated that KCNQ1OT1 regulates lung fibrosis by modulating Rtn3 expression in a lipopolysaccharide-induced acute lung injury mouse model, the specific pathways and cellular functions involved in this process require further investigation (Yang et al. 2022).Similarly, Wang et al. reported that XIST influences lung fibrosis development by modulating β-catenin expression through miR-139(Wang et al. 2017b). However, validation of these gene regulatory molecules in IPF patient lung tissues and their correlation with clinical features remain to be explored comprehensively. Regarding NEAT1, Zhang et al. reported that NEAT1 positively regulates NFATc3 expression by directly targeting miR-29a (Zhang et al. 2022), but this finding was only validated in cellular models and requires in vivo confirmation. These findings underscore the potential indispensable role of these DElncRNAs in the overall regulatory network of IPF. Given the limited research on related regulatory networks, exploring the regulatory mechanisms of these three candidate DElncRNAs is particularly important. Screening of ceRNA networks can provide clues to their regulatory networks.
For the candidate DEmRNA molecules in the ceRNA network, we validated their expressions and analyzed their correlation with the predicted FVC (pre-bd) % to determine whether these candidate DEmRNAs have clinical significance. The %predicted FVC (pre-bd) is a crucial indicator for assessing lung function and disease progression in IPF patients. Our results indicate differential expression of these DEmRNA molecules, among which the expressions of CLDN1, COL3A1, MMP16, SFRP2, SPRY4, and ITGB8 correlate with % predicted FVC (pre-bd). Interestingly, we found that MMP16 and ITGB8 are also involved in multiple ceRNA regulatory networks. Does this imply that these two mRNAs may be key factors in IPF regulation? ITGB8, a transmembrane heterodimer, serves as the primary receptor for extracellular matrix (ECM) adhesion and recognition (Takada et al. 2007). Integrin exhibits sustained activation in abnormally activated lung fibroblasts, leading to fibrosis progression. Downregulation of ITGB8 expression significantly alleviates fibrosis in the liver, lungs, and kidneys (Bagnato et al. 2018; McCarty 2020). For instance, Minagawa et al. confirmed that inhibiting TGF-β activation using avβ8 antibodies effectively prevents fibroblast activation in the airways of patients with chronic obstructive pulmonary disease (Minagawa et al. 2014). MMP16 belongs to the calcium-zinc-dependent metalloproteinase family, responsible for ECM maintenance. It plays a crucial role in various pathological processes such as atherosclerosis, tumor invasion, and fibrosis (Pittayapruek et al. 2016). Studies have shown higher expression of MMP16 in allergic asthma mice compared to normal mice, and regulating MMP16 expression can reduce fibrosis (Zhong et al. 2022). IPF is characterized by excessive extracellular matrix deposition, highlighting the significance of in-depth research on ITGB8 and MMP16 in ECM regulation. However, current research on this topic is limited, and the specific regulatory pathways and involved biological processes remain unknown. With the construction of ceRNA networks, new clues can be provided for their upstream regulatory relationships, and further investigation into regulatory mechanisms can offer new insights for targeted IPF therapy.
Based on the aforementioned data, it can be inferred that the ceRNA network containing lncRNA KCNQ1OT1, XIST, and NEAT1, as well as mRNA ITGB and MMP16, may play a crucial regulatory role in the development of IPF. These networks include hsa-miR-146b-5p, hsa-miR-20a-5p, and hsa-miR-31-5p. Hsa-miR-20a-5p is a member of the has-mir-17 cluster. Studies have indicated a reduction in the expression of miR-20a in IPF patients, while the miR-17 ~ 92 cluster is associated with lung fibrosis development (Dakhlallah et al. 2013). However, the specific regulatory mechanisms remain unclear. Another miRNA, hsa-miR-146b-5p, is downregulated in lung tissue samples from IPF patients. Subsequent studies by Shuai et al. found that miR-146b-5p can inhibit lung fibrosis through the Notch1/PDGFRβ/ROCK1 pathway (Mullenbrock et al. 2018). However, the reliability and reproducibility of this mechanism are compromised due to imperfect experimental design. Although hsa-miR-31-5p has not been specifically studied in the context of lung fibrosis, it is highly expressed in hypertrophic scar fibroblasts. Knocking out miR-31-5p significantly inhibits fibroblast proliferation under hypoxic conditions, promotes cell invasion, and suppresses the expression of collagen I, collagen III, and fibronectin (Wang et al. 2017a). Therefore, investigating the role of these miRNAs in fibrotic diseases is worthwhile.
However, this study has several limitations. First, it relies solely on data from the GEO database. Although four different IPF microarray datasets were integrated, the sample size remained limited, which may have affected the statistical significance and generalizability of the findings. Second, there was a lack of validation of the expression levels of the identified candidate DEmiRNAs and DElncRNAs and a lack of validation of the expression levels of candidate DEmRNAs in human lung tissues. Additionally, the assessment of the clinical value of the candidate DEmRNAs was based solely on correlation analysis without any validation. Overall, this study is still in the exploratory stage, and the underlying regulatory mechanisms of the ceRNA network have not been fully elucidated through the observed level changes and bioinformatics analyses. Further functional biology experiments with larger sample sizes are required to validate these mechanisms.
Conclusion
This study has constructed three ceRNA regulatory networks (KCNQ1OT1/XIST/NEAT1-miR-20a-5p-ITGB8, XIST-miR-146b-5p/miR-31-5p-MMP16, and NEAT1-miR-31-5p-MMP16). These findings contribute to a deeper understanding of the pathogenesis of IPF and provide clues for identifying potential therapeutic targets and further research directions.
Data Availability
The row data included in this study are available in GEO [https://www.ncbi.nlm.nih.gov/geo/, (accessed on 25 May 2022)].
References
Ahangari F, Price NL, Malik S et al (2023) microRNA-33 deficiency in macrophages enhances autophagy, improves mitochondrial homeostasis, and protects against lung fibrosis. JCI Insight. https://doi.org/10.1172/jci.insight.158100
Anonymous. American Thoracic Society (2000) Idiopathic pulmonary fibrosis: diagnosis and treatment. International Consensus Statement. American Thoracic Society (ATS), and the European Respiratory Society (ERS). Am J Respir Crit Care Med. 161(2 Pt 1):646–664. https://doi.org/10.1164/ajrccm.161.2.ats3-00.
Bagnato GL, Irrera N, Pizzino G et al (2018) Dual Αvβ3 and Αvβ5 blockade attenuates fibrotic and vascular alterations in a murine model of systemic sclerosis. Clin Sci (Lond) 132(2):231–242. https://doi.org/10.1042/CS20171426
Bartel DP (2018) Metazoan microRNAs. Cell 173(1):20–51. https://doi.org/10.1016/j.cell.2018.03.006
Chen J, Bian M, Pan L et al (2022) GABPB1-AS1 promotes the development of osteosarcoma by targeting SP1 and activating the Wnt/β-catenin pathway. J Oncol 2022:8468896. https://doi.org/10.1155/2022/8468896
Dakhlallah D, Batte K, Wang Y et al (2013) Epigenetic regulation of miR-17~92 contributes to the pathogenesis of pulmonary fibrosis. Am J Respir Crit Care Med 187(4):397–405. https://doi.org/10.1164/rccm.201205-0888OC
Dempsey TM, Payne S, Sangaralingham L, Yao X, Shah ND, Limper AH (2021) Adoption of the antifibrotic medications pirfenidone and nintedanib for patients with idiopathic pulmonary fibrosis. Ann Am Thorac Soc 18(7):1121–1128. https://doi.org/10.1513/AnnalsATS.202007-901OC
Gene Ontology Consortium (2015) Gene Ontology Consortium: going forward. Nucleic Acids Res 43(Database issue):D1049–D1056. https://doi.org/10.1093/nar/gku1179.
Hadjicharalambous MR, Lindsay MA (2020) Idiopathic pulmonary fibrosis: pathogenesis and the emerging role of long non-coding RNAs. Int J Mol Sci 21(2):524. https://doi.org/10.3390/ijms21020524
Hadjicharalambous MR, Roux BT, Feghali-Bostwick CA et al (2018) Long non-coding RNAs are central regulators of the IL-1β-induced inflammatory response in normal and idiopathic pulmonary lung fibroblasts. Front Immunol 9:2906. https://doi.org/10.3389/fimmu.2018.02906
Huang HY, Lin YC et al (2020) miRTarBase 2020: updates to the experimentally validated microRNA-target interaction database. Nucleic Acids Res 48(D1):D148–D154. https://doi.org/10.1093/nar/gkz896
Hutchinson J, Fogarty A, Hubbard R et al (2015) Global incidence and mortality of idiopathic pulmonary fibrosis: a systematic review. Eur Respir J 46(3):795–806. https://doi.org/10.1183/09031936.00185114
Kaikkonen MU, Lam MTY, Glass CK (2011) Non-coding RNAs as regulators of gene expression and epigenetics. Cardiovasc Res 90(3):430–440. https://doi.org/10.1093/cvr/cvr097
Karagkouni D, Paraskevopoulou MD et al (2020) DIANA-LncBase v3: indexing experimentally supported miRNA targets on non-coding transcripts. Nucleic Acids Res 48(D1):D101–D110. https://doi.org/10.1093/nar/gkz1036
Kanehisa M, Sato Y, Kawashima M et al (2016) KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res 44(D1):D457-462. https://doi.org/10.1093/nar/gkv1070
King TE, Pardo A, Selman M (2011) Idiopathic pulmonary fibrosis. Lancet (London, England) 378(9807):1949–1961. https://doi.org/10.1016/S0140-6736(11)60052-4
Kreuter M, Swigris J, Pittrow D et al (2017) Health related quality of life in patients with idiopathic pulmonary fibrosis in clinical practice: insights-IPF registry. Respir Res 18(1):139. https://doi.org/10.1186/s12931-017-0621-y
Lai X, Zhong J, Zhang A et al (2022) Focus on long non-coding RNA MALAT1: insights into acute and chronic lung diseases. Front Genet 13:1003964. https://doi.org/10.3389/fgene.2022.1003964
Li J-H, Liu S, Zhou H et al (2014) starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-seq data. Nucleic Acids Res 42(Database issue):92–97. https://doi.org/10.1093/nar/gkt1248
Liu G, Friggeri A, Yang Y et al (2010) miR-21 mediates fibrogenic activation of pulmonary fibroblasts and lung fibrosis. J Exp Med 207(8):1589–1597. https://doi.org/10.1084/jem.20100035
Lu Q, Guo Z, Xie W et al (2018) The lncRNA H19 mediates pulmonary fibrosis by regulating the miR-196a/COL1A1 axis. Inflammation 41(3):896–903. https://doi.org/10.1007/s10753-018-0744-4
McCarty JH (2020) Αvβ8 integrin adhesion and signaling pathways in development, physiology and disease. J Cell Sci 133(12):9434. https://doi.org/10.1242/jcs.239434
von Mering C, Huynen M, Jaeggi D et al (2003) STRING: a database of predicted functional associations between proteins. Nucleic Acids Res 31(1):258–261
Minagawa S, Lou J, Seed RI et al (2014) Selective targeting of TGF-β activation to treat fibroinflammatory airway disease. Sci Transl Med 6(241):241–279. https://doi.org/10.1126/scitranslmed.3008074
Morris KV, Mattick JS (2014) The RISE OF REGULATORY RNA. Nat Rev Genet 15(6):423–437. https://doi.org/10.1038/nrg3722
Mullenbrock S, Liu F, Szak S et al (2018) Systems analysis of transcriptomic and proteomic profiles identifies novel regulation of fibrotic programs by miRNAs in pulmonary fibrosis fibroblasts. Genes. https://doi.org/10.3390/genes9120588
Pittayapruek P, Meephansan J, Prapapan O et al (2016) Role of matrix metalloproteinases in photoaging and photocarcinogenesis. Int J Mol Sci 17(6):868. https://doi.org/10.3390/ijms17060868
Song X, Cao G, Jing L et al (2014) Analysing the relationship between lncRNA and protein-coding gene and the role of lncRNA as ceRNA in pulmonary fibrosis. J Cell Mol Med 18(6):991–1003. https://doi.org/10.1111/jcmm.12243
Takada Y, Ye X, Simon S (2007) The Integrins. Genome Biol 8(5):215. https://doi.org/10.1186/gb-2007-8-5-215
Wang P, Li X et al (2019) LncACTdb 20: an updated database of experimentally supported ceRNA interactions curated from low- and high-throughput experiments. Nucleic Acids Res 47(D1):D121–D127. https://doi.org/10.1093/nar/gky1144
Wang P, Guo Q et al (2022) LncACTdb 3.0: an updated database of experimentally supported ceRNA interactions and personalized networks contributing to precision medicine. Nucleic Acids Res. 50(D1):D183–D189. https://doi.org/10.1093/nar/gkab1092
Wang P, Ning S, Zhang Y et al (2015) Identification of lncRNA-associated competing triplets reveals global patterns and prognostic markers for cancer. Nucleic Acids Res 43(7):3478–3489. https://doi.org/10.1093/nar/gkv233
Wang X, Zhang Y, Jiang BH et al (2017a) Study on the role of Hsa-miR-31-5p in hypertrophic scar formation and the mechanism. Exp Cell Res 361(2):201–209. https://doi.org/10.1016/j.yexcr.2017.09.009
Wang Y, Liang Y, Luo J et al (2017b) XIST/miR-139 axis regulates bleomycin (BLM)-induced extracellular matrix (ECM) and Pulmonary Fibrosis through β-Catenin. Oncotarget 8(39):65359–65369. https://doi.org/10.18632/oncotarget.18310
Yang S, Liu F, Wang D (2022) Long noncoding RNA Kcnq1ot1 prompts lipopolysaccharide-induced acute lung injury by microRNA-7a-5p/Rtn3 axis. Eur J Med Res 27(1):46. https://doi.org/10.1186/s40001-022-00653-8
Zhang H, Song M, Guo J et al (2021a) The function of non-coding RNAs in idiopathic pulmonary fibrosis. Open Med (Warsaw, Poland) 16(1):481–490. https://doi.org/10.1515/med-2021-0231
Zhang M, Zhu W, Haeryfar M et al (2021b) Long non-coding RNA TRG-AS1 promoted proliferation and invasion of lung cancer cells through the miR-224-5p/SMAD4 axis. Onco Targets Ther 14:4415–4426. https://doi.org/10.2147/OTT.S297336
Zhang S, Chen H, Yue D et al (2021c) Long non-coding RNAs: promising new targets in pulmonary fibrosis. J Gene Med 23(3):e3318. https://doi.org/10.1002/jgm.3318
Zhang X, Duan X-J, Li L-R et al (2022) lncRNA NEAT1 promotes hypoxia-induced inflammation and fibrosis of alveolar epithelial cells via targeting miR-29a/NFATc3 Axis. Kaohsiung J Med Sci 38(8):739–748. https://doi.org/10.1002/kjm2.12535
Zhang Z, Li Y, Fan L et al (2023) LncRNA THUMPD3-AS1 promotes invasion and EMT in gastric cancer by regulating the miR-1297/BCAT1 pathway. iScience 26(9):7673. https://doi.org/10.1016/j.isci.2023.107673
Zhong J, Liu M, Chen S et al (2022) Study of the regulatory mechanism of miR-26a-5p in allergic asthma. Cells 12(1):38. https://doi.org/10.3390/cells12010038
Zhou Y, Zhou B, Pache L et al (2019) Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 10:1523. https://doi.org/10.1038/s41467-019-09234-6
Acknowledgements
We would like to express their gratitude to all the participants of the study. We acknowledge Science and Technology Department of Guizhou Province for financial support, and all the databases we used in the paper for providing the platform and contributors for uploading their meaningful data sets.
Funding
This study was supported by the Science and Technology Program of Guizhou Province (ZK [2021] General 347).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation by YC, XJ and WL; data collection were performed by MZ and XW; HZ prepared Figs. 1 and 3; WY prepared Figs. 4 and 5; MZ prepared Figs. 6; CF prepared Figs. 7, 8, and 9; XW prepared Tables 1 and 2, and MZ Tables 3 and 4. The first draft of the manuscript was written by MZ and XW, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethical Approval
The study was conducted in accordance with the Declaration of Helsinki and approved by the Guizhou Medical University Ethical Review Committee.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, M., Wu, X., Zhu, H. et al. Construction and Bioinformatics Analysis of ceRNA Regulatory Networks in Idiopathic Pulmonary Fibrosis. Biochem Genet (2024). https://doi.org/10.1007/s10528-024-10853-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10528-024-10853-y