Abstract
Classic and contemporary trophic ecology-based studies have shown that most non-native freshwater fish species (NNS) that integrate into novel environments have the potential to influence the recipient ecosystems’ structure and function. However, the interspecific trophic interactions amongst co-occurring NNS within invaded systems remain poorly studied. Here, we used carbon (δ13C) and nitrogen (δ15N) stable isotope analyses to examine general fish trophic diversity patterns (native and non-native fishes) and to explore trophic niche patterns amongst co-occurring NNS within a flow-modified river system, the Great Fish River (South Africa). The system was characterised by isotopic variation, which revealed spatial differences in trophic complexity from uninvaded headwater tributaries to invaded mainstem and downstream sections. Two of the invaded sections, the upper mainstem of the Great Fish River (UGFR) and the Koonap River, had low isotopic overlaps between NNS and the native fish assemblages. Furthermore, co-occurring NNS in these two invaded sections had variable isotopic niche sizes and low interspecific isotopic niche overlaps, suggesting the potential for trophic differentiation. By comparison, there was evidence of high resource use patterns among NNS within the lower mainstem section of the Great Fish River (LGFR), which likely reflected trophic plasticity. Overall, results of this study provided evidence of both trophic niche differentiation (UGFR and Koonap River) and niche overlap (LGFR) as probable mechanisms of co-occurrences of the non-native fishes within different invaded sections of the Great Fish River system, and underscores the difficulties associated with predicting their trophic impacts.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Biological invasions and habitat degradation have been shown to potentially threaten community structure and function of freshwater ecosystems (Collen et al. 2014; Comte et al. 2016). Several empirical studies have provided insights on invasion impacts in a wide range of freshwater ecosystems, including those invaded by keystone non-native species (Ligtvoet et al. 1991; Johnson et al. 2006; Ficetola et al. 2007; De Vanna et al. 2011), characterised by high local endemism and low invasion resistance (Tedesco et al. 2012; Collen et al. 2014; Weyl et al. 2014; Jordaan et al. 2020) and subject to high habitat-related anthropogenic modifications (Laurenson and Hocutt 1986; Ruhi et al. 2016; Liu et al. 2017). Whilst many of these studies have focused on impacts of one or few non-native species, there is increasing attention on the occurrence of multiple non-native invaders (e.g. Jackson 2015; Pyšek et al. 2020; Guareschi et al. 2021) whose net impacts are poorly known. Understanding how multiple non-native species integrate and impact the invaded freshwater environments is, thus, crucial for managing species invasions in these ecosystems (Comte et al. 2016). One way to understand such impacts is by exploring trophic and food web patterns of the invaded ecosystems (Copp et al. 2017).
From a trophic ecology perspective, non-native species have been shown to disrupt community structure and function either directly through predation and competition (e.g. Kadye and Booth 2012; Bašić et al. 2019; Rogosch and Olden 2020; Murphy et al. 2021) or indirectly by negatively influencing ecological aspects such as habitat associations and the behaviours of native taxa (e.g. Vander Zanden and Rasmussen 1999; Kadye et al. 2020), which may negatively influence aspects of native biota’s life-history attributes such as growth and reproduction (e.g. Britton et al. 2010a; de Araújo et al. 2022). Both classic and contemporary research on the influence of non-native taxa on the trophic structure and function of recipient ecosystems has been largely instrumental in unravelling invasion impacts. Examples of this research include studies on the ecosystem-wide food web dynamics following the localised extirpations of native fauna by non-native piscivores in lentic habitats (Witte et al. 1992; Vander Zanden et al. 2003; Downing et al. 2012), the competitive interactions between native and non-native species (Byres 2002; Martin et al. 2010; Britton et al. 2018), interspecific and synergistic interactions among non-native species within invaded habitats (Britton et al. 2010b; Jackson et al. 2012), and the trophic cascade-induced dynamics, such as changes in primary production and algal biomass due to heavy predation on grazing invertebrates by non-native salmonids and high nutrient cycling by invasive crustaceans in lotic habitats (Crowl et al. 1992; Flecker and Townsend 1994; Nyström et al. 2001; Herbst et al. 2009).
Recently, there have been increasing concerns on the co-occurrence of multiple non-native species and their potential impacts on recipient freshwater ecosystems’ trophic dynamics (Johnson et al. 2009; Jackson and Britton 2013; Sagouis et al. 2015; Liu et al. 2018). Specifically, there is concern on the interactions among non-native invaders, which could potentially result in either complementary, additive and facilitative effects on the structure and function of food webs in invaded systems (Jackson et al. 2014; Jackson 2015). This is because multiple invasive species occurring in sympatry can have broader ecological impacts due to their likelihood of occupying different trophic levels within the food webs of the invaded communities (Johnson et al. 2009; Jackson and Britton 2013). Similarly, non-native species with high diet plasticity and/or broad trophic niches, such as opportunistic omnivorous and generalist predators, could also prey on native organisms across multiple trophic levels resulting in unpredictable impacts within the invaded communities (Jackson and Britton 2013; Liu et al. 2018). Other studies have shown potential synergistic impacts among invaders. Examples include the high predation and subsequent biomass reduction of predatory invertebrates by non-native bluegill Lepomis macrochirus, facilitating the establishment of non-native bullfrog Lithobates catesbeiana due to enhanced survival of their tadpoles (Adams et al. 2003), and the predation by non-native crayfish Orconectes rusticus on native snails (Physa and Lymnaea spp.), which potentially facilitated the establishment of non-native snail Bellamya chinensis due to reduced competitive interactions between the native and non-native snails (Johnson et al. 2009). Despite the emerging evidence of the impacts of multiple invasions (Jackson and Britton 2013; Sagouis et al. 2015), the broad nature of interspecific trophic interactions amongst co-occurring non-native species outside their native ranges is uncertain as these interactions are likely to be context-dependent (Jackson 2015). For example, there have been reports of different interspecific interactions, ranging from weak amensal and commensal interactions (Griffen et al. 2008; Johnson et al. 2009) to strong interactions, such as overt predation and competition impacts (Liu et al. 2018) and non-native species facilitating each other’s establishment (Hohenadler et al. 2018). Therefore, understanding these complex interactions could assist to mitigate the impacts of co-occurring multiple non-native species on native communities (Liu et al. 2018; Balzani et al. 2020).
Stable isotopes have become an important tool to examine community trophic or food web structures (Layman et al. 2007, 2012), and in assessing the ecological impacts of anthropogenic environmental changes (Cucherousset et al. 2012a; Alp and Cucherousset 2022). Specifically, because they provide temporally-integrated assessment of consumer diet (Hershey et al. 2017), stable isotope analyses (SIA) of carbon (δ13C) and nitrogen (δ15N) can be used to assess the energy sources and trophic positions of organisms, respectively (Post 2002; Anderson and Cabana 2007; Vander Zanden et al. 2016). Thus, they can be used to trace energy flow from basal sources (primary producers and detrital sources) through primary to secondary and tertiary consumers (Hershey et al. 2017; Layman et al. 2012). Consequently, stable isotope analyses have been extensively used to evaluate biological invasions-related trophic ecology of aquatic organisms in freshwater ecosystems (Vander Zanden and Fetzer 2007; Jackson et al. 2020), including the assessment of trophic interactions between co-occurring native and non-native fishes (Pennock et al. 2021; Top-Karakuş et al. 2021) and broad trophic impacts of these introductions on recipient communities (Vander Zanden and Rasmussen 1999; Kadye and Booth 2012; Cucherousset et al. 2012a; Fink and Harrod 2013; Jackson and Britton 2013; Bašić et al. 2019). In addition to providing robust estimates of trophic interrelationships, applications of SIA have recently been extended to provide quantitative assessments of stable isotope-based trophic and functional diversity indices, which facilitate comparisons of food webs among different ecosystems (Layman et al. 2007; Cucherousset and Villéger 2015; Rigolet et al. 2015; De Cáceres et al. 2019). Besides the general applications of these diversity indices to assess the community structure and functioning of freshwater ecosystems (Comte et al. 2016; Frossard et al. 2020), they have also been used to evaluate the role of anthropogenic disturbances, including the impacts of biological invasions within these ecosystems (Cucherousset and Villéger 2015; Jackson et al. 2020).
This study focused on the Great Fish River system in South Africa. In addition to hosting multiple non-native fishes, this river system is characterised by modified mainstem habitats due to an inter-basin water transfer scheme (IBWT) that connects the Orange River (the donor system) with the Great Fish River (Laurenson and Hocutt 1986; Kadye and Booth 2013; Mpopetsi and Kadye 2023). The presence of a flow-modified mainstem section, along with tributaries with near-natural flow regimes, have created spatially variable habitats that likely have implications on the structural and functional interrelationships of fish taxa within the Great Fish River system. Despite the spatial variability in the structural nature of these habitats, non-native fishes have established within both the flow-modified mainstem and the unmodified sections of the major tributaries (Kadye and Booth 2013; Sifundza et al. 2021). This raises the need to understand the ecological dynamics associated with these invasions in the different habitats of this river system. Whilst previous research on species-environment relationships revealed that the non-native species assemblage was characterised by taxa with high environmental tolerance, an indication of propensity towards niche opportunism (Kadye and Booth 2020), there is little information regarding the trophic niche patterns of these species. To fill this gap knowledge, this study employed SIA-based approaches as proxy for trophic ecology. Therefore, the primary objectives of this study were to (1) describe the isotopic patterns of the Great Fish River system food web across its broad spatial scale, (2) elucidate the isotopic diversity patterns of non-native and native fishes within the invaded sections and (3) to evaluate the isotopic niche patterns and probable interspecific interactions of non-native fishes to determine whether trophic niche patterns were the likely drivers of non-native fishes’ proliferation within this river system. Due to the likely preponderance of niche opportunism shown by species-environment relationships (e.g. Kadye and Booth 2020), it was, firstly, hypothesised that non-native fishes would exhibit consistent isotopic diversity patterns that would reflect broad resource use patterns compared to native fishes. Secondly, because of the high likelihood of exploiting broad range of trophic resources, as it has been demonstrated elsewhere (e.g. De Santis et al. 2022), it was hypothesised that the non-native fish species would exhibit high interspecific trophic niche overlap that would likely reflect trophic niche generalisation.
Materials and methods
Data collection and sample preparation
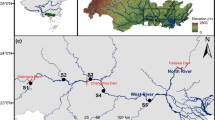
Sampling was done to collect carbon (δ13C) and nitrogen (δ15N) stable isotope data in the Great Fish River mainstem, and its major two tributaries, the Koonap and Kat rivers (Fig. 1a). In total, 67 sites were sampled, with each site being sampled once. In the Great Fish River, sampling was conducted during summer from October 2009 to April 2010 in the flow-altered mainstem, and October 2014–March 2015 in its headwater section. In the Koonap and Kat rivers, sampling was done in October 2018–March 2019. Samples for stable isotope analysis were collected from fishes (Table 1) and all potential prey that included macroinvertebrates and basal sources (Supplementary material 1). Several sampling methods, including electrofishing, fyke netting, gill netting, hand netting and seine netting were used to capture fish from both the tributaries and the mainstem sections of the Great Fish River. For electrofishing, a 12 V battery-powered SAMUS backpack electrofisher in combination with a stop net (4 mm mesh net) were used to sample shallow rocky habitats in the headwaters where neither seine nor fyke nets were efficient. Stunned fish were collected with a hand net. A stop net, secured to the streambed, was used downstream to block electrocuted fish that were missed by hand netting. To standardise the sampling effort, a multiple (three) pass electrofishing was conducted for a maximum of 15 min per site. Marginal and shallow habitats (< 1 m), in headwaters and tributaries, were sampled using an 8 m long seine net with 4 mm mesh size. Two to four seine hauls (average of three) were performed at each site where seine netting was conducted. The seine net was deployed at a distance (the distance varied with river channel size) parallel to the shore and was then rapidly pulled onto the shore. Mainstem sites were sampled using experimental gill nets, fyke nets (single and double ended) and the seine net. Experimental gill nets were 30 m long with three 10 m panels of mesh sizes of 50, 75 and 100 mm. Fyke nets had an 8 m guiding net and a first ring diameter of 55 cm and a 10 mm mesh size. Experimental gill- and fyke-nets were deployed overnight, from late afternoon/evening to morning, with an average soak time of 14 h. Captured fish were identified to species following Skelton (2001) and other regional literature in line with the recent taxonomic changes in some taxa (e.g., Kambikambi et al. 2021). A sample of a maximum of 15 individuals (across species) per site was euthanised by a lethal dose of clove oil after which a small piece of the dorsal muscle tissue was taken using a clean scalpel blade for stable isotope analysis, and the samples were transferred to Eppendorf tubes for storage. For Sandelia bainsii, which is endangered, a small tissue was taken at the tip of the anal fin using a clean pair of scissors.
a The Great Fish River system and its major tributaries; b study system was delineated into nine sections (A–I) based on the presence or absence of non-native species. Red ellipses indicate the nine sections; A The Great Fish River headwaters; B the upper Great Fish River (UGFR); C The lower Great Fish River (LGFR), D The Koonap River headwater tributaries, E the Koonap River upper mainstem, F The lower Koonap River, G, H the Kat River headwater tributaries and I the Kat River upper mainstem section
Macroinvertebrates were collected from instream substratum and submerged vegetation. In flowing habitats, stream substrates were disturbed by kick sampling, two minutes per sample, and the dislodged animals were collected using a 250 µm hand-held scoop net positioned downstream of flow. The net was then progressively moved upstream to catch dislodged macroinvertebrate samples. In vegetated and slow-flowing habitats, macroinvertebrates were captured by sweeping the scoop net for 2 min. Captured macroinvertebrate samples were then rinsed with clean water and then transferred into a collection tray for sorting. Coarse particulate matter (CPOM), which included C3 and C4 plants and other large organic matter debris, was either handpicked or dislodging from the substratum and filtered through a hand-held net. Fine particulate organic matter (FPOM) was collected by filtering 25 L of stream water through a 100 µm net. Epilithic algae was scrapped from coarse substratum (boulders and bedrock) with a scalpel blade and rinsed with distilled water. Free-floating filamentous algae, macrophytes and other organic matter were collected by hand, rinsed and transferred into collection bottles. For basal and macroinvertebrate, three samplings were performed for each taxon at each site. The three samples were then combined to form one sample for that site, and this was then used for stable isotope analysis. All samples for stable isotope analysis were kept on ice in the field and transported to the laboratory, Rhodes University, Makhanda, South Africa, for further processing. For macroinvertebrates, a reference sample (of every taxon collected at each site) was also collected and stored in 70% ethanol, and this was later used in conjunction with frozen samples to confirm the collected samples’ identification in the laboratory.
At the laboratory, sample of macroinvertebrates were thawed, sorted and identified to either family, genus or species level using a dissecting microscope (Labomed CZM4, Labomed, USA) and regional identification guides (Day et al. 2001a, b; Day and De Moor 2002a, b; Day et al. 2002; Gerber and Gerber 2002). All samples were oven-dried at 60 ℃ for 48–72 h after which they were ground into a fine homogenous powder, using a mortar and pestle. The sample material was weighed (1 ± 0.05 mg for animal tissue and 3 ± 0.5 mg for plant tissue) and packed into 8 × 5 mm tin capsules. Stable isotope analysis was done at the Rhodes University and the University of Pretoria, South Africa. At Rhodes University, stable isotope analysis of carbon and nitrogen was performed using a Europa Scientific INTEGRA isotope ratio mass spectrometer at the IsoEnvironmental Lab, Makhanda. At the University of Pretoria, the isotopic analysis was done on a Flash SEA 1112 series coupled to a Delta V Plus stable light isotope ratio mass spectrometer via ConFlo IV system (ThermoFischer, Bremen, Germany). Stable isotope ratios are reported in the δ notation in per mil units (‰) using the following formula:
where X is 13C or 15N and R is the corresponding ration 13C/12C or 15N/14N. The Rstandard values are based on Vienna PeeDee Belemnite (VPDB) for δ13C and atmospheric N2 for δ15N. Analytical precision was evaluated using the following calibrated laboratory standards: Merck Gel (δ13C = – 20.3 ± 0.1‰, δ15N = + 7.9 ± 0.1‰), DL-Valine (δ13C = – 10.6 ± 0.1‰, δ15N = + 6.2 ± 0.1‰), and Casein (δ13C = – 27.0 ± 0.1‰, δ15N = + 5.9 ± 0.1‰). Laboratory standards were calibrated using the following reference material: IAEA-CH-3 (cellulose), IAEA-CH-6 (sucrose), IAEA-CH-7 (polyethylene 241 foil), IAEA N-1 & IAEA N-2 (ammonium sulphate) and IAEA NO-3 (potassium nitrate). Sample precision based on standard deviation of repeated measurements of laboratory standards was 0.1‰ for both δ13C and δ15N. To address potential δ13C lipid-content bias, fish tissue stable isotope values were mathematically normalised based on C/N ratio following Post et al. (2007).
Data analysis
To describe the food web patterns, the study system was firstly delineated into different sections based on the presence or absence of non-native fish species, and with each section being made up of multiple sites (Fig. 1b, Table 1). Thus, the river system was delineated into the following nine sections. (A) The Great Fish River headwaters, which comprised an uninvaded section where one native species occurred. (B) The upper Great Fish River (UGFR), which comprised an invaded upper section of the mainstem. This section had two native and four non-native fishes. (C) The lower Great Fish River (LGFR) that constituted the invaded lower reaches of the mainstem was characterised by both primary freshwater and estuarine-dependent fishes. This section was invaded by six non-native fishes and had 14 native species. (D) The Koonap River headwater tributaries and (E) the Koonap River upper mainstem that were both uninvaded and had one native species. (F) The lower Koonap River where two native fishes and three non-native fishes occurred. Only the uninvaded sections of the Kat River were sampled. This was because the invaded portion of the Kat River comprised sections that were either dominated by silviculture (due to citrus plantations) or heavily polluted by sewage, specifically the section downstream of the Fort Beaufort town. Therefore, the sampled sections of the Kat River included (G) the headwater tributaries where one native species occurred, (H) the headwater tributary where two native fishes occurred, and (I) the upper mainstem section where three native fishes occurred. (Fig. 1b, Table 1).
To elucidate the trophic diversity patterns of non-native and native fish assemblages within the invaded sections, isotopic diversity indices were computed following Cucherousset and Villéger (2015) and Rigolet et al. (2015). Because the sampled isotope data varied both spatially and temporally, the food webs were likely influenced by differences in the stable isotope values for basal sources (e.g., Cucherousset and Villéger 2015). To circumvent this, prior to the computing of diversity indices, stable isotope data were first standardised to z-scores to normalise the effect of baselines following Fry and Davis (2015), and the resultant values were then scaled. The scaling procedure does not affect the distribution of the δ13C and δ15N values, but instead transforms the δ13C and δ15N multidimensional δ-space so that each isotopic axis is scaled to the same range of 0–1 (Cucherousset and Villéger 2015). Thus, the isotopic diversity indices were computed using standardised isotopic data pooled from the three invaded sections, the upper (UGFR) and lower (LGFR) Great Fish River mainstem sections, and the lower Koonap River. During the computation of diversity metrics, stable isotope values were weighed by species relative abundance based on the appropriate catch per unit effort (CPUE) for each section. The isotopic diversity indices were computed as;
-
1.
Isotopic richness (IRic), which provides a quantitative estimate of the isotopic space for the whole community or assemblage. Low IRic values (close to 0) may indicate low functional diversity, possibly due to species loss, limited use of resources and low buffering from disturbances, whereas high IRic values (close to 1) may indicate high use of the trophic space.
-
2.
Isotopic divergence (IDiv), which describes how the isotopic space is occupied based on the degree of isotopic spacing by community members. Low IDiv values (close to 0) likely reflects a community characterised by trophic generalists, whereas high IDiv values (close to 1) likely indicates a high degree of trophic specialisation for the community.
-
3.
Isotopic dispersion (IDis), which reflects the extent to which community members differ in their stable isotope values. Low IDis values (close to 0) indicates that community members have similar stable isotope values, whereas high IDis values (close to 1) reflect dissimilar stable isotope values.
-
4.
Isotopic uniqueness (IUni), which indicate the degree of isotopic dissimilarity among community members. Low IUni values (close to 0) indicate that most of the weight belongs to isotopically similar organisms, whereas high IUni values (close to 1) indicate that most of the organisms are isolated in the stable isotope space.
In addition, isotope overlap between the non-native and native fish assemblages was evaluated based on isotopic nestedness (INes), which provides a quantitative estimate of the degree to which one group or community is a subset of the other (Cucherousset and Villéger 2015). Low INes value indicates no isotopic overlap, whereas high nestedness, which is consistent with high isotopic overlap, reflects that one group is a subset of the other.
Lastly, to evaluate the trophic niche patterns and interspecific interactions of non-native fishes, a Bayesian isotope niche analysis was conducted following Swanson et al. (2015). This was based on a two-pronged approach. Firstly, the isotope niche size for each non-native species was computed based on the joint probability distributions of scaled (0–1) δ13C and δ15N values, resulting in probability distributions (α = 95%) that reflect the most plausible isotopic trophic niche sizes from a multi-dimensional isotopic-space. This was expressed as: \(P(Y \in N_{R} )\), where \(Y\) is the δ13C and δ15N data matrix for a particular species and \(N_{R}\) is the probable isotopic niche region for a particular species. Secondly, interspecific interactions between two species were determined based on the extent of isotopic niche overlap as: \(O_{B}^{A} = P(Y_{A} \in N_{R} (B))\), where \(O_{B}^{A}\) is the probability of species \(A\) overlapping onto the isotopic niche of species \(B\) (Swanson et al. 2015). The isotope niche overlaps were based on Bayesian posterior distributions (posterior means and 95% credible intervals) and were estimated based on Monte Carlo simulations with 1000 iterations. All statistical analyses were performed using R software program version 4.2.2 (R Development Core Team 2022). Isotopic diversity indices were computed using published R code provided by Cucherousset and Villéger (2015). Isotopic niche patterns were computed using the R package nicheROVER (Lysy et al. 2021).
Results
Isotopic variation
The Great Fish River system exhibited spatial variation in its isotopic values from the headwaters and tributaries to the lower section, which appeared consistent with food web complexity (Table 2). Specifically, the Great Fish River mainstem was mostly distinguished by an upstream to downstream differences in the ranges of δ15N, which were small in the headwater section (δ15N = + 6.9 to + 12.2‰), intermediate in the UGFR (δ15N = + 4.8 to + 14.9 ‰) and large in the LGFR (δ15N = + 0.8 to + 18.4‰). Similarly, the Koonap River’s isotopic values were distinguished by δ15N ranges, which were smaller in the tributaries (δ15N = + 1.6 to + 11.2‰) and the upper section (δ15N = – 0.4 to + 12.0‰) than the lower section (δ15N = + 1.8 to + 15.1‰). In comparison, the Kat River isotopic values were mostly distinguished by a large breadth in δ13C values together with upstream to downstream differences in δ15N ranges. Specifically, there was a wide breadth in δ13C for macroinvertebrates in the headwater tributaries, whereas the macroinvertebrates in the mainstem section had both wide δ13C breadth and high δ15N range.
Within the invaded UGFR section, non-native fishes were distinguished by intermediate δ15N values compared to the two native fishes, L. umbratus that had the lowest (δ15N = + 11.6‰) and A. mossambica that had the highest (δ15N = + 14.9‰) values (Table 1). Furthermore, non-native fishes had a wider breadth in δ13C values (– 26.7 to – 22.6‰) compared to those in either the LGFR (– 26.9 to – 23.7 ‰) or the lower Koonap River (– 26.1 to – 24.7‰). By comparison, within the invaded LGFR, the non-native fishes generally had lower δ15N values than most native fishes. In contrast, within the invaded lower Koonap, all non-native fishes were distinguished by higher δ15N values than native fishes (Table 1).
Isotopic diversity patterns of non-native and native fishes
In the UGFR, the non-native and native fish assemblages differed in their isotopic diversity patterns, which also showed no overlap (INes = 0) between the two groups (Fig. 2a). Specifically, non-native fishes were mostly distinguished by high isotopic divergence (IDiv = 0.8) and isotopic dispersion (IDis = 0.7) compared to native species (IDiv = 0.4; IDis = 0.5) (Table 3). This showed that the non-native fishes were characterised by dissimilar isotopic values, and likely comprised species that were characterised by trophic specialisation. By comparison, native fishes were mostly distinguished by high isotopic uniqueness (IUni = 0.7), indicating that these species had different stable isotope values, reflecting different resource utilisation patterns, which was likely consistent with trophic differentiation. In the LGFR, non-native and native fishes were characterised by high isotopic nestedness (INes = 0.9), which depicted high isotopic diversity overlap (Fig. 2b). The non-native fish assemblage’s isotopic diversity, which appeared to be a subset of the native fish assemblage’s isotopic diversity, was distinguished by low isotopic richness (IRic = 0.0) compared to native species (IRic = 0.3). However, non-native species had higher values for other isotopic diversity metrics (i.e., IDiv, IDis and IUni) compared to native species, likely indicating broader resource utilisation pattern among the non-native species. In the Koonap River, although the non-native and native fish assemblages exhibited low isotopic diversity overlap (Fig. 2c), most of their isotopic diversity metrics were comparable (Table 3). Specifically, the two groups were characterised by high isotopic divergence (IDiv = 0.8) and isotopic dispersion (IDis > 0.7). This suggested that although these two groups had species that exhibited different resource patterns, as shown by lack of isotopic diversity overlap, trophic resource utilisation was likely driven by trophic specialisation in both groups.
Isotopic diversity overlap metrics between native (blue) and non-native (red) fish species of the three communities within the Great Fish River system; the upper Great Fish River (UGFR) mainstem (a), the lower Great Fish River (LGFR) mainstem (b) and the Koonap River (c) Isotopic overlap metric, isotopic nestedness (INes), computed from scaled δ13C and δ15N values, is reflected on top of each plot
Isotopic niche patterns and interspecific interactions of non-native fishes
In the UGFR, non-native fishes had variable isotopic niche sizes that ranged from the largest for L. aeneus (scaled niche size (NS) = 0.33 ± 0.06‰2), followed by C. gariepinus (NS = 0.15 ± 0.06‰2) to small for L. capensis (NS = 0.03 ± 0.01‰2) and C. carpio (NS = 0.02 ± 0.00‰2) (Fig. 3a). In general, most pairwise comparisons revealed low isotopic niche overlaps (mean overlap < 20%) (Fig. 4), which showed the likelihood of trophic niche differentiation among these non-native species. Nevertheless, exceptions were shown by the high probabilities of isotopic niche overlaps for C. gariepinus onto the isotopic niche of L. aeneus (mean overlap = 53.24%, 95% CI = 16.0–90.0%) and L. capensis onto the isotopic niche of L. aeneus (mean overlap = 80.23%, 95% CI 32.0–100%). In the LGFR, three non-native species, C. carpio, C. gariepinus and L. aeneus had relatively large niche sizes (NS range = 0.33 ± 0.06–0.48 ± 0.18‰2) compared to those of T. sparrmanii, G. affinis and L. capensis, which had smaller isotopic niche sizes (NS range = 0.05 ± 0.04 – 0.18 ± 0.08‰2) (Fig. 3b). Pairwise comparisons revealed moderate to high probabilities of isotopic niche overlaps (mean overlap > 50%) for 10 of the 30 non-native fishes’ pairings (Fig. 5). In addition, C. gariepinus and L. aeneus had high probabilities of isotopic niche overlaps (mean overlap > 60%) with most non-native species, indicating that these two species were likely dietary generalists in this section. Similarly, G. affinis exhibited high probability of overlap onto the isotopic niche of C. carpio (mean overlap = 97.84%, 95% CI 84–100%), C. gariepinus (mean overlap = 84.73%, 95% CI 62–98%) and L. aeneus (mean overlap = 76.99%, 95% CI 49–96%). In comparison, most species had low probabilities of isotopic niche overlap (mean overlap < 20%) with both G. affinis and L. capensis (Fig. 5). Within the Koonap River, all three non-native species had small isotopic niche sizes (NS < 0.1‰2), with T. sparrmanii having the largest (NS = 0.08 ± 0.02‰2) of the three (Fig. 3c). Most pairwise comparisons revealed low to moderate isotopic niche overlaps (mean overlap = 34.35–37.91%), indicating propensity for niche differentiation amongst species (Fig. 6). These included the overlaps of C. carpio onto the isotopic niche of T. sparrmanii (mean overlap = 34.35%, 95% CI 1–93%) and C. gariepinus onto the isotopic niche of C. carpio (mean overlap = 37.91%, 95% CI 1–97%). Nevertheless, there was a moderately high probability of isotopic niche overlap by C. gariepinus onto the isotopic niche of T. sparrmanii (mean overlap = 58.73%, 95% CI 2–100%).
Isotopic niche sizes (in ‰2) for non-native fishes within the three invaded sections of the Great Fish River system, the upper (a) and the lower (b) Great Fish River mainstem sections and the Koonap River (c). Niche sizes were computed based on the joint probability distributions of δ13C and δ15N values, scaled to range between 0 and 1. Species are colour coded consistently across the three sections and are abbreviated as; Ccar, Cyprinus carpio, Cgar, Clarias gariepinus, Gaff, Gambusia affinis, Laen, Labeobarbus aeneus, Lcap, Labeo capensis; Tspa, Tilapia sparrmanii
Mean niche overlap probabilities (%) of the non-native species within the upper Great Fish River (UGFR). The overlap metric is directional, such that it represents the probability that an individual from a species (row) will overlap onto the isotope niche of the other species (column). The niche size (niche region) was defined as the 95% credibility intervals of isotopic space. Species are abbreviated as Ccar, Cyprinus carpio; Cgar, Clarias gariepinus; Laen, Labeobarbus aeneus; Lcap, Labeo capensis
Mean niche overlap probabilities (%) of the non-native species within the lower Great Fish River (LGFR). The overlap metric is directional, such that it represents the probability that an individual from a species (row) will overlap onto the isotope niche of the other species (column). The niche size (niche region) was defined as the 95% credibility intervals of isotopic space. Species are abbreviated as Ccar, Cyprinus carpio; Cgar, Clarias gariepinus; Gaff, Gambusia affinis; Laen, Labeobarbus aeneus; Lcap, Labeo capensis; Tspa, Tilapia sparrmanii
Mean niche overlap probabilities (%) of the non-native species within the Koonap River. The overlap metric is directional, such that it represents the probability that an individual from a species (row) will overlap onto the isotope niche of the other species (column). The niche size (niche region) was defined as the 95% credibility intervals of isotopic space. Species are colour coded consistently across the three sections and are abbreviated as Ccar, Cyprinus carpio; Cgar, Clarias gariepinus; Tspa, Tilapia sparrmanii
Discussion
The Great Fish River system’s isotopic values in both its mainstem and the tributaries were characterised by longitudinal differences in food web structure, which were generally depicted by high δ15N ranges in downstream sections compared to upstream and headwater sections. These patterns appeared to be congruent with the longitudinal shifts in food web properties, which are generally linked to aspects such as the structural changes in river size and geomorphometry, differences in productivity, and increases in habitat complexity and species richness along the river continuum (e.g. Vannote et al. 1980; Angermeier and Schlosser 1989; Romanuk et al. 2006; Sánchez-Hernández 2023). Similar to other studies elsewhere, which have shown spatial differences in food web characteristics in anthropogenically-modified river systems, including altered flow that influences energy transfer dynamics (e.g. Cross et al. 2013; Brauns et al. 2019) and in invaded systems where non-native fishes influence both trophic chain length and breadth (e.g. Pennock et al. 2021), this study revealed different invasion patterns from a food web perspective. Specifically, relative to native fishes, the non-native species exhibited isotopic variations that ranged from intermediate δ15N values and wide δ13C breadth in the UGFR, low δ15N values and narrow δ13C breadth in the LGFR, and high δ15N values but narrow δ13C breadth in the Koonap River. Although the non-native fish assemblages exhibited variable isotopic diversity patterns that were broadly defined by low overlaps with native species in the UGFR and Koonap River and high isotopic diversity overlap in the LGFR, these non-native fishes were mostly characterised by comparable isotopic diversity metrics across the different sections. Thus, these broad patterns appeared to be consistent with our first hypothesis whereby we postulated consistent patterns that would reflect broad trophic resources utilisation patterns for the non-native fishes. On the other hand, the non-native fishes were characterised by low interspecific isotopic niche overlaps within the invaded UGFR and Koonap River. This posited the likelihood of isotopic niche differentiation among these non-native fishes, which appeared to be inconsistent with our second hypothesis. Nevertheless, in the LGFR, the non-native fishes were mostly characterised by high isotopic niche overlaps, a pattern that appeared to support the second hypothesis. These broad patterns suggest that both the trophic diversity and trophic niche patterns of non-native fishes were likely influenced by the nature of the invaded section, the intraspecific resource use patterns of the non-native species and their probable interactions with the native fish assemblage within the respective invaded sections. The lack of consistent and predictable trophic diversity and niche patterns for non-native fishes across the invaded sections in this study posits the likelihood of trophic plasticity, which is a common phenomenon related to environmental stochasticity, variation in prey availability and interspecific interactions coupled by niche opportunities that has been observed in other studies (Córdova-Tapia et al. 2015; Pennock et al. 2021). Trophic plasticity has been postulated to facilitate establishment of non-native species in freshwater ecosystems (e.g. Pettitt-Wade et al. 2015; Rolla et al. 2020) and to confer a strategy to minimise and/or avoid potential trophic competition (Jackson et al. 2012; Pelage et al. 2022).
In the UGFR and the Koonap River, the low isotopic diversity overlaps (low isotopic nestedness) between non-native and native species and the low isotopic niche overlaps amongst the non-native species, suggest that trophic niche differentiation was a probable mechanism facilitating the proliferation of non-native species in these habitats. Specifically, the results of this study suggested the possibility of trophic niche specialisation due to high isotopic divergence and diversion for the non-native species, and the high isotopic uniqueness for the native species. Furthermore, despite large isotopic niche breadths, most non-native species generally showed low trophic niche overlaps, except for trophic generalists, such as C. carpio and C. gariepinus whose isotopic niche overlapped onto those of other non-native species. This appeared to be consistent with empirical studies elsewhere, which have also shown low isotopic overlaps to reflect trophic niche differentiation among non-native and native species (e.g. Zambrano et al. 2010; Zengeya et al. 2011; Córdova-Tapia et al. 2015; Tarkan et al. 2018; Top-Karakuş et al. 2021). The creation of permanent habitats in the UGFR, as a result of the IBWT, may have facilitated resources by potentially increasing biomass of primary production and that of lower trophic secondary production, such as benthic invertebrates, which in turn could have influenced trophic resource patterns of fishes in this section. For example, O’Keeffe and De Moor (1988) noted that following the creation of permanent flow in the Great Fish River, the system was characterised by a substantial shift towards the dominance of hydropsychid, chironomid and simuliid invertebrate species. These resources likely sustain the proliferation of rheophilic taxa of non-native species such as the benthivorous L. capensis and the generalist invertivorous L. aeneus probably through reduced interspecific interactions with C. carpio, and the generalist piscivore C. gariepinus. Alternatively, it is likely that the low native species richness of the UGFR and Koonap River may have presented vacant niches that were exploited by the different non-native fishes with minimum competitive interactions. This is because species depauperate environments are more likely to be susceptible to invasions due to high availability of vacant niches (Shea and Chesson 2002; Hierro et al. 2005; Leuven et al. 2009; Jeschke 2014).
While trophic niche differentiation is generally considered a common mechanism for coexistence among fishes (Mason et al. 2008; Pilger et al. 2010; Jackson and Britton 2013) and appeared to be most probable in the UGFR and Koonap River, this mechanism appeared less plausible in the LGFR. This is because the non-native fish assemblage’s isotopic diversity largely overlapped with, and appeared to be a subset of, that of the native fish assemblage. Furthermore, these non-native fishes exhibited high isotopic niche overlaps. In contrast to native fishes, the relatively low non-native species’ isotopic diversity within the LGFR suggested less trophic complexity within this species group. While this contradicts other studies that have found non-native species to generally have high isotopic diversity (e.g. Sagouis et al. 2015; De Santis et al. 2022), findings of this study likely reflect the incipient nature of the trophic habits of the predominant non-native species, which mostly comprised generalist piscivores, such as C. gariepinus and species that likely occupied lower trophic levels, including benthivores and invertivores, such as C. carpio, L. aeneus and T. sparrmanii. On the other hand, the generally high isotopic diversity for the native fish assemblage may reflect high trophic complexity that characterises most downstream sections of large rivers, which is generally hypothesised to be maintained by weak consumer-resource interactions (Levin 2000; Kokkoris et al. 2002; Bellmore et al. 2015).
The occurrence of multiple non-native species with relatively low isotopic diversity but high isotopic niche overlap in the LGFR may suggests weak trophic interactions among these species within this invaded section. The establishment of multiple non-native species in this section, however, raises concern on their potential impact. This is because non-native species such as C. gariepinus, C. carpio and L. aeneus, which were found to be characterised by large isotope niche sizes, were likely to exert negative influences on the trophic dynamics of this system due to their foraging habits and possible facilitative interactions. For example, C. carpio, a global invader, is known to influence nutrient availability and to increase water turbidity (Zambrano et al. 2006; Parkos et al. 2011), which may confer an advantage to C. gariepinus, a generalist predator that does not require high visibility for foraging. Thus, there exists a concern about the potential synergistic impacts of these two species on aspects such as facilitated direct predation by C. gariepinus on small- to medium-sized native species, such as the round herring Gilchristella aestuaria, oval moony Monodactylus falciformis and Pseudomyxus capensis. Furthermore, there is potential indirect impacts through interspecific interference and probable competitive interaction for benthic resources by non-native species such as C. carpio and L. capensis, and the invertivorous L. aeneus on species such as the native populations of L. umbratus that feeds on soft sediments and detritus, P. macrolepis that feeds on diatoms grubbed out of bottom sand and the G. callidus that feeds on bottom-living insects and small invertebrates.
General studies on trophic impacts by non-native fishes in lotic habitats have shown that they (non-native fishes) can influence food webs by either increasing the ecosystem’s trophic ranges, such as when there is invasion by non-native top predators (Post and Takimoto 2007; Cucherousset et al. 2012b; Walsworth et al. 2013), expanding the food web sizes, such as when there is invasion by taxa that are located on the edges of the food webs (Sagouis et al. 2015) or shrinking of the food webs and/or compression of the isotopic niche (Vander Zanden and Rasmussen 1999; Walsworth et al. 2013), such as when there is invasion by generalist predators and/or when there is elimination of certain taxa. In this study, some of these aspects appeared to have been reflected as well. For example, the UGFR and the Koonap River appeared to typify food web expansion due to the addition of non-native species, with high δ15N values, on the top or edges of the food webs. Specifically, in the UGFR, the food web appeared to show a wider breadth in carbon sources for the fishes, which was typified by non-native species such as C. carpio with high δ13C value and L. capensis with low δ13C value. In comparison, the Koonap River appeared to reflect an increase in its food web structure, as all the non-native fishes in this section (C. carpio, C. gariepinus and T. sparmanii) had higher δ15N than native fishes.
In conclusion, the results of the present study suggested that isotopic diversity patterns likely reflected trophic niche differentiation among native and non-native fishes. In addition, isotopic niche patterns suggested that trophic niche differentiation was a probable mechanism associated with co-occurrence of different non-native fishes in some invaded sections. Nevertheless, these patterns appeared to vary across the different invaded sections. Specifically, the low isotopic diversity overlaps between native and non-native fishes and low isotopic niche overlap among most non-native fishes in the upper mainstem (UGFR) and invaded tributary (Koonap River) sections posits the likelihood of exploitation of vacant trophic niches by the invaders. In comparison, weak trophic interactions among non-native species were likely responsible for their co-occurrence within the LGFR. The results of this study add to the body of knowledge that seeks to understand how non-native species integrate into disturbed ecosystems and the mechanisms facilitating these integrations.
Availability of data and materials
Data analysed during the current study are available from the corresponding author upon reasonable request.
References
Adams MJ, Pearl CA, Bury RB (2003) Indirect facilitation of an anuran invasion by non-native fishes. Ecol Lett 6:343–351. https://doi.org/10.1046/j.1461-0248.2003.00435.x
Alp M, Cucherousset J (2022) Food webs speak of human impact: using stable isotope-based tools to measure ecological consequences of environmental change. Food Webs 30:e00218. https://doi.org/10.1016/j.fooweb.2021.e00218
Anderson C, Cabana G (2007) Estimating the trophic position of aquatic consumers in river food webs using stable nitrogen isotopes. J N Am Benthol Soc 26:273–285. https://doi.org/10.1899/0887-3593(2007)26[273:ETTPOA]2.0.CO;2
Angermeier PL, Schlosser IJ (1989) Species-area relationships for stream fishes. Ecology 70:1450–1514. https://doi.org/10.2307/1938204
Balzani P, Gozlan RE, Haubrock PJ (2020) Overlapping niches between two co-occurring invasive fish: the topmouth gudgeon Pseudorasbora parva and the common bleak Alburnus alburnus. J Fish Biol 97:1385–1392. https://doi.org/10.1111/jfb.14499
Bašić T, Copp GH, Edmonds-Brown VR, Keskin E, Davison P, Britton R (2019) Trophic consequences of an invasive, small-bodied non-native fish, sunbleak Leucaspius delineatus, for native pond fishes. Biol Invasions 21:261–275. https://doi.org/10.1007/s10530-018-1824-y
Bellmore JR, Baxter CV, Connolly PJ (2015) Spatial complexity reduces interaction strengths in the meta-food web of a river floodplain mosaic. Ecology 96:274–283. https://doi.org/10.1890/14-0733.1
Brauns M, Brabender M, Gehre M, Rincke K, Wietere M (2019) Organic matter resources fuelling food webs in a human-modified lowland river: importance of habitat and season. Hydrobiologia 841:121–131. https://doi.org/10.1007/s10750-019-04011-4
Britton JR, Davies GD, Harrod C (2010a) Trophic interactions and consequent impacts of the invasive fish Pseudorasbora parva in a native aquatic food web: a field investigation in the UK. Biol Invasions 12:1533–1542. https://doi.org/10.1007/s10530-009-9566-5
Britton JR, Harper DM, Oyugi DO, Grey J (2010b) The introduced Micropterus salmoides in an Equatorial Lake: A paradoxical loser in an invasion meltdown scenario? Biol Invasions 12:3439–3448. https://doi.org/10.1007/s10530-010-9742-7
Britton JR, Ruiz-Navarro A, Verreycken H, Amat-Trigo F (2018) Trophic consequences of introduced species: comparative impacts of increased interspecific versus intraspecific competitive interactions. Funct Ecol 32:486–495. https://doi.org/10.1111/1365-2435.12978
Byers JE (2002) Impact of non-indigenous species on natives enhanced by anthropogenic alteration of selection regimes. Oikos 97:449–458. https://doi.org/10.1034/j.1600-0706.2002.970316.x
Collen B, Whitton F, Dyer EE, Baillie JEM, Cumberlidge N, Darwall WRT, Pollock C, Richman NI, Soulsby A-M, Böhm M (2014) Global patterns of freshwater species diversity, threat and endemism. Glob Ecol Biogeogr 23:40–51. https://doi.org/10.1111/geb.12096
Comte L, Cucherousset J, Boulêtreau S, Olden JD (2016) Resource partitioning and functional diversity of worldwide freshwater fish communities. Ecosphere 7:e01356. https://doi.org/10.1002/ecs2.1356
Copp GH, Britton JR, Edmonds-Brown GZ, VR, Pegg J, Vilizzi L, Davison PI, (2017) Trophic consequences of non-native pumpkinseed Lepomis gibbosus for native pond fishes. Biol Invasions 19:25–41. https://doi.org/10.1007/s10530-016-1261-8
Córdova-Tapia F, Contreras M, Zambrano L (2015) Trophic niche overlap between native and non-native fishes. Hydrobiologia 746:291–301. https://doi.org/10.1007/s10750-014-1944-z
Cross WF, Baxter CV, Rosi-Marshall EJ, Hall RO, Kennedy TA, Donner KC, Kelly HAW, Seegert SEZ, Behn KE, Yard MD (2013) Food-web dynamics in a large river discontinuum. Ecol Monogr 83:311–337. https://doi.org/10.1890/12-1727.1
Crowl TA, Townsend CR, McIntosh AR (1992) The impact of introduced brown and rainbow trout on native fish: the case of Australasia. Rev Fish Biol Fish 2:217–241. https://doi.org/10.1007/BF00045038
Cucherousset J, Villéger S (2015) Quantifying the multiple facets of isotopic diversity: new metrics for stable isotope ecology. Ecol Ind 56:152–160. https://doi.org/10.1016/j.ecolind.2015.03.032
Cucherousset J, Blanchet S, Olden SD (2012a) Non-native species promote trophic dispersion of food webs. Front Ecol Environ 10:406–408. https://doi.org/10.1890/12.WB.018
Cucherousset J, Bouletreau S, Martino A, Roussel JM, Santoul F (2012) Using stable isotope analyses to determine the ecological effects of non-native fishes. Fish Manag Ecol 19:111–119. https://doi.org/10.1111/j.1365-2400.2011.00824.x
Day JA, de Moor IJ (eds) (2002a) Guides to the freshwater invertebrates of southern Africa, Vol 5: Non-arthropods. Water Research Commission report no. TT 167/02, Pretoria, South Africa. https://www.wrc.org.za/wp-content/uploads/mdocs/TT-167-02.pdf
Day JA, de Moor IJ (eds) (2002b) Guides to the freshwater invertebrates of southern Africa, Vol 6:Arachnida & mollusca (Araneae, Water Mites & Mollusca). Water Research Commission report no. TT 182/02, Pretoria, South Africa. https://www.wrc.org.za/wp-content/uploads/mdocs/TT-182-02.pdf
Day JA, de Moor IJ, Steward BA, Louw AE (eds) (2001a) Guides to the Freshwater Invertebrates of Southern Africa, Vol 3: Crustacea II. Water Research Commission report no. TT 148/01, Pretoria, South Africa. https://www.wrc.org.za/wp-content/uploads/mdocs/TT-148-01.pdf
Day JA, Stewart BA, de Moor IJ, Louw AE (eds) (2001b) Guides to the Freshwater Invertebrates of Southern Africa. Vol 4: Crustacea III. Water Research Commission report no. TT 141/01, Pretoria, South Africa. https://www.wrc.org.za/wp-content/uploads/mdocs/TT-141-01.pdf
Day JA, Harrison AD, de Moor IJ (eds) (2002) Guides to the freshwater invertebrates of southern Africa: Vol 9: Diptera. Water Research Commission report no. TT 201/02, Pretoria, South Africa. https://www.wrc.org.za/wp-content/uploads/mdocs/TT-201-02.pdf
de Araújo TP, Brighenti LS, Dolabela BM, Ribeiro SP, dos Santos HB, Thomé RG (2022) Can the introduction of non-native fish induce variation in life-history traits of a native species in a neotropical lake? Mar Freshw Res 73:651–661. https://doi.org/10.1071/MF21138
De Cáceres M, Coll L, Legendre P, Allen RB, Wiser SK, Fortin M, Condit R, Hubbell S (2019) Trajectory analysis in community ecology. Ecol Monogr 89:e01350. https://doi.org/10.1002/ecm.1350
De Santis V, Cicala D, Baneschi I, Boschi C, Brignone S, Iaia M, Zaupa S, Volta P (2022) Non-native fish assemblages display potential competitive advantages in two protected small and shallow lakes of northern Italy. Glob Ecol Evol 35:e02082. https://doi.org/10.1016/j.gecco.2022.e02082
De Vanna KM, Bodamer BL, Wellington CG, Hammer E, Mayer CM, Bossenbroek JM (2011) An alternative hypothesis to invasional meltdown in the Laurentian Great Lakes region: general facilitation by Dreissena. J Great Lakes Res 37:632–641. https://doi.org/10.1016/j.jglr.2011.07.005
Downing AS, Van Nes EH, Janse JH, Witte F, Cornelissen IJ, Scheffer M, Mooij WM (2012) Collapse and reorganization of a food web of Mwanza Gulf, Lake Victoria. Ecol Appl 22:229–239. https://doi.org/10.1890/11-0941.1
Ficetola GF, Coïc C, Detaint MB, M, Lorvelec O, Miaud C, (2007) Pattern of distribution of the American bullfrog Rana catesbeiana in Europe. Biol Invasions 9:767–772. https://doi.org/10.1007/s10530-006-9080-y
Flecker AS, Townsend CR (1994) Community - wide consequences of trout introduction in New Zealand streams. Ecol Appl 4:798–807. https://doi.org/10.2307/1942009
Fink P, Harrod C (2013) Carbon and nitrogen stable isotopes reveal the use of pelagic resources by the invasive Ponto-Caspian mysid Limnomysis benedeni. Isot Environ Health Stud 49:312–317. https://doi.org/10.1080/10256016.2013.808197
Fry B, Davis J (2015) Rescaling stable isotope data for standardized evaluations of food webs and species niches. Mar Ecol Prog Ser 528:7–17. https://doi.org/10.3354/meps11293
Frossard V, Vagnon C, Rossignon C, Caudron A (2020) Variability of isotopic partitioning between sympatric brown trout (Salmo trutta) and European grayling (Thymallus thymallus) in the Loue River (France). Ecol Freshw Fish 30:285–295. https://doi.org/10.1111/eff.12583
Gerber A, Gabriel MJM (2002) Aquatic invertebrates of South African rivers: field guide. Institute for water quality studies, Department of water affairs and forestry, Pretoria. https://www.dws.gov.za/iwqs/biomon/aquabugsa/Aquatic_Invertebrates_of_South_African_Rivers_Field_Guide_en.pdf
Griffen BD, Guy T, Buck JC (2008) Inhibition between invasives: a newly introduced predator moderates the impacts of a previously established invasive predator. J Anim Ecol 77:32–40. https://doi.org/10.1111/j.1365-2656.2007.01304.x
Guareschi S, Laini A, England J, Barrett J, Wood PJ (2021) Multiple co-occurrent alien invaders constrain aquatic biodiversity in rivers. Ecol Appl 31:e02385. https://doi.org/10.1002/eap.2385
Herbst DB, Silldorff EL, Cooper SD (2009) The influence of introduced trout on the benthic communities of paired headwater streams in the Sierra Nevada of California. Freshw Biol 54:1324–1342. https://doi.org/10.1111/j.1365-2427.2009.02187.x
Hershey AE, Northington RM, Finlay JC, Peterson BJ (2017) Chapter 23—Stable isotopes in stream food webs. In: Hauer FR, Lamberti GA (eds) Methods in stream ecology, 3rd edn. Academic Press, Amsterdam, pp 3–20. https://doi.org/10.1016/B978-0-12-813047-6.00001-2
Hierro JL, Maron JL, Callaway RM (2005) A biogeographical approach to plant invasions: the importance of studying exotics in their introduced and native range. J Ecol 93:5–15. https://doi.org/10.1111/j.1365-2745.2004.00953.x
Hohenadler MMA, Honka KI, Emde S, Klimpel S, Sures B (2018) First evidence for a possible invasional meltdown among invasive fish parasites. Sci Rep 8:15085. https://doi.org/10.1038/s41598-018-33445-4
Jackson MC (2015) Interactions among multiple invasive animals. Ecology 96:2035–2041. https://doi.org/10.1890/15-0171.1
Jackson MC, Donohue I, Jackson AL, Britton JR, Harper DM, Grey J (2012) Population-level metrics of trophic structure based on stable isotopes and their application to invasion ecology. PLoS ONE 7:e31757. https://doi.org/10.1371/journal.pone.0031757
Jackson MC, Britton JR (2013) Variation in the trophic overlap of invasive Pseudorasbora parva and sympatric cyprinid fishes. Ecol Freshw Fish 22:654–657. https://doi.org/10.1111/eff.12063
Jackson MC, Jones T, Milligan M, Sheath D, Taylor J, Ellis A, England J, Grey J (2014) Niche differentiation among invasive crayfish and their impacts on ecosystem structure and functioning. Freshw Biol 59:1123–1135. https://doi.org/10.1111/fwb.12333
Jackson MC, Fourie HE, Dalu T, Woodford DJ, Wasserman RJ, Zengeya TA, Ellender BR, Kimberg PK, Jordaan MS, Chimimba CT, Weyl OLF (2020) Food web properties vary with climate and land use in South African streams. Funct Ecol 34:1653–1665. https://doi.org/10.1111/1365-2435.13601
Jeschke JM (2014) General hypotheses in invasion ecology. Divers Distrib 20:1229–1234. https://doi.org/10.1111/ddi.12258
Johnson LE, Bossenbroek JM, Kraft CE (2006) Patterns and pathways in the post-establishment spread of non-indigenous aquatic species: the slowing invasion of North American inland lakes by the zebra mussel. Biol Invasions 8:475–489. https://doi.org/10.1007/s10530-005-6412-2
Johnson PTJ, Olden JDT, Solomon CT, Vander Zanden MJ (2009) Interactions among invaders: community and ecosystem effects of multiple invasive species in an experimental aquatic system. Oecologia 159:161–170. https://doi.org/10.1007/s00442-008-1176-x
Jordaan MS, Chakona A, van der Colff D (2020) Protected areas and endemic freshwater fishes of the cape fold ecoregion: Missing the boat for fish conservation? Front Environ Sci 8:502042. https://doi.org/10.3389/fenvs.2020.502042
Kadye WT, Booth AJ (2013) An invader within an altered landscape: one catfish, two rivers and an inter-basin water transfer scheme. River Res Appl 29:1131–1146. https://doi.org/10.1002/rra.2599
Kadye WT, Booth AJ (2012) Integrating stomach content and stable isotope analyses to elucidate the feeding habits of non-native sharptooth catfish Clarias gariepinus. Biol Invasions 14:779–795. https://doi.org/10.1007/s10530-011-0116-6
Kadye WT, Leigh S, Booth AJ (2020) Predator naïve minnows respond to their conspecific alarm substance but not the odour from a non-native predator. Afr J Ecol 58:757–765. https://doi.org/10.1111/aje.12768
Kambikambi M, Kadye W, Chakona A (2021) Allopatric differentiation in the Enteromius ano-plus complex in South Africa, with the revalidation of E. cernuus and E. oraniensis, and description of a new species, E. mandelai (Teleostei: Cyprinidae). J Fish Biol 99:931–954. https://doi.org/10.1111/jfb.14780
Kokkoris GD, Jansen VAA, Loreau M, Troumbis AY (2002) Variability in interaction strength and implications for biodiversity. J Anim Ecol 71:362–371. https://doi.org/10.1046/j.1365-2656.2002.00604.x
Laurenson LB, Hocutt CH (1986) Colonisation theory and invasive biota: the Great Fish River, a case history. Environ Monit Assess 6:71–90. https://doi.org/10.1007/BF00394289
Layman CA, Arrington DA, Montana CG, Post DM (2007) Can stable isotope ratios provide for community wide measures of trophic structure? Ecology 88:42–48. https://doi.org/10.1890/0012-9658(2007)88[42:CSIRPF]2.0.CO;2
Layman CA, Araujo MS, Boucek R, Hammerschlag-Peyer CM, Harrison E, Jud ZR, Matich P, Rosenblatt AE, Vaudo JJ, Yeager LA, Post DM, Bearhop S (2012) Applying stable isotopes to examine food-web structure: an overview of analytical tools. Biol Rev 87:545–562. https://doi.org/10.1111/j.1469-185X.2011.00208.x
Leuven RSEW, van der Velde G, Baijens I, Snijders J, van der Zwart C, Lenders HJR, Bij de Vaate A (2009) The river Rhine: a global highway for dispersal of aquatic invasive species. Biol Invasions 11:1989–2008. https://doi.org/10.1007/s10530-009-9491-7
Levin SA (2000) Multiple scales and the maintenance of biodiversity. Ecosystems 3:498–506. https://doi.org/10.1007/s100210000044
Ligtvoet W, Witte F, Goldschmidt T, Van Oijen M, Wanink JH, Goudswaard PC (1991) Species extinction and concomitant ecological changes in Lake Victoria. Neth J Zool 42:214–232. https://doi.org/10.1163/156854291X00298
Liu C, Comte L, Olden JD (2017) Heads you win, tails you lose: life-history traits predict invasion and extinction risk of the world’s freshwater fishes. Aquat Conserv Mar Freshwat Ecosyst 27:773–779. https://doi.org/10.1002/aqc.2740
Liu X, Wang S, Ke Z, Cheng C, Wang Y, Zhang F, Xu F, Li X, Gao X, Jin C, Zhu W, Yan S, Li Y (2018) More invaders do not result in heavier impacts: the effects of non-native bullfrogs on native anurans are mitigated by high densities of non-native crayfish. J Anim Ecol 87:850–862. https://doi.org/10.1111/1365-2656.12793
Lysy M, Stasko AD, Swanson HK (2021) Niche region and niche overlap metrics for multidimensional ecological niches (Version 1.1.0). Available online https://cran.r-project.org/web/packages/nicheROVER/nicheROVER.pdf
Martin CW, Valentine MM, Valentine JF (2010) Competitive interactions between invasive Nile Tilapia and native fish: the potential for altered trophic exchange and modification of food webs. PLoS ONE 5:e14395. https://doi.org/10.1371/journal.pone.0014395
Mason NWH, Irz P, Lanoiselée C, Mouillot D, Arguillier C (2008) Evidence that niche specialization explains species-energy relationships in lake fish communities. J Anim Ecol 77:285–296. https://doi.org/10.1111/j.1365-2656.2007.01350.x
Mpopetsi PP, Kadye WT (2023) Colonisation theory and invasive biota: the Great Fish River case history, 35 years later. Afr J Aquat Sci 48:84–96. https://doi.org/10.2989/16085914.2022.2142508
Murphy CA, Romer JD, Stertz K, Arismendi I, Emig R, Monzyk F, Johnson SL (2021) Damming salmon fry: evidence for predation by non-native warmwater fishes in reservoirs. Ecosphere 12:1–17. https://doi.org/10.1002/ecs2.3757
Nyström P, Svensson O, Lardner B, Brönmark C, Granéli W (2001) The influence of multiple introduced predators on a littoral pond community. Ecology 82:1023–1039. https://doi.org/10.2307/2679900
O’Keeffe JH, De Moor FC (1988) Changes in the physico-chemistry and benthic invertebrates of the Great Fish River, South Africa, following an interbasin transfer of water. Regul Rivers: Res Manag 2:39–55. https://doi.org/10.1002/rrr.3450020105
Parkos JJ, Santucci VJ, Wahl DH (2011) Effects of adult common carp (Cyprinus carpio) on multiple trophic levels in shallow mescosms. Can J Fish Aquat Sci 60:182–192. https://doi.org/10.1139/f03-011
Pelage L, Lucena-Frédou F, Eduardo LN, Le Loc’h F, Bertrand A, Lira AS, Frédou T (2022) Competing with each other: fish isotopic niche in two resource availability contexts. Front Mar Sci 9:975091. https://doi.org/10.3389/fmars.2022.975091
Pennock CA, Ahrens ZT, McKinstry MC, Budy P, Gido KB (2021) Trophic niches of native and nonnative fishes along a river-reservoir continuum. Sci Rep 11:12140. https://doi.org/10.1038/s41598-021-91730-1
Pettitt-Wade H, Wellband KW, Heath DD, Fisk AT (2015) Niche plasticity in invasive fishes in the Great Lakes. Biol Invasions 17:2565–2580. https://doi.org/10.1007/s10530-015-0894-3
Pilger TJ, Gido KB, Propst DL (2010) Diet and trophic niche overlap of native and nonnative fishes in the Gila River, USA: implications for native fish conservation. Ecol Freshw Fish 19:300–321. https://doi.org/10.1111/j.1600-0633.2010.00415.x
Post DM (2002) Using stable isotopes to estimate trophic position: models, methods, and assumptions. Ecology 83:703–718. https://doi.org/10.1890/0012-9658(2002)083[0703:USITET]2.0.CO;2
Post DM, Takimoto G (2007) Proximate structural mechanisms for variation in food-chain length. Oikos 116:775–782. https://doi.org/10.1111/j.0030-1299.2007.15552.x
Post DM, Layman CA, Arrington DA, Takimoto G, Quattrochi J, Montana CG (2007) Getting to the fat of the matter: models, methods and assumptions for dealing with lipids in stable isotope analyses. Oecologia 152:179–189. https://doi.org/10.1007/s00442-006-0630-x
Pyšek P, Hulme PE, Simberloff D, Bacher S, Blackburn TM, Carlton JT et al (2020) Scientists’ warning on invasive alien species. Biol Rev 95:1511–1534. https://doi.org/10.1111/brv.12627
R Development Core Team (2022) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. www.R-project.org
Rigolet C, Thiébaut É, Brind’Amour A, Dubois SF (2015) Investigating isotopic functional indices to reveal changes in the structure and functioning of benthic communities. Funct Ecol 29:1350–1360. https://doi.org/10.1111/1365-2435.12444
Rogosch JS, Olden JD (2020) Invaders induce coordinated isotopic niche shifts in native fish species. Can J Fish Aquat Sci 77:1348–1358. https://doi.org/10.1139/cjfas-2019-0346
Rolla M, Consuegra S, Garcia de Leaniz C (2020) Trophic plasticity of the highly invasive Topmouth Gudgeon (Pseudorasbora parva) inferred from stable isotope analysis. Front Ecol Evol 8:212. https://doi.org/10.3389/fevo.2020.00212
Romanuk TN, Jackson LJ, Post JR, McCauley E, Martinez ND (2006) The structure of food webs along river networks. Ecography 29:3–10. https://doi.org/10.1111/j.2005.0906-7590.04181.x
Ruhi A, Olden JD, Sabo JL (2016) Declining streamflow induces collapse and replacement of native fish in the American Southwest. Front Ecol Environ 14:465–472. https://doi.org/10.1002/fee.1424
Sagouis A, Cucherousset J, Villéger S, Santoul F, Boulêtreau S (2015) Non-native species modify the isotopic structure of freshwater fish communities across the globe. Ecography 38:979–985. https://doi.org/10.1111/ecog.01348
Sánchez-Hernández J (2023) Fresh perspectives on the river continuum concept require trophic ecology approaches focussed on food web structure and energy mobilisation routes. J Anim Ecol 92:957–964. https://doi.org/10.1111/1365-2656.13928
Shea K, Chesson P (2002) Community ecology theory as a framework for biological invasions. Trends Ecol Evol 17:170–176. https://doi.org/10.1016/S0169-5347(02)02495-3
Sifundza DS, Chakona A, Kadye WT (2021) Distribution patterns and habitat associations of Sandelia bainsii (Teleostei: Anabantidae), a highly threatened narrow-range endemic freshwater fish. J Fish Biol 98:292–303. https://doi.org/10.1111/jfb.14580
Skelton PH (2001) A complete guide to the freshwater fishes of southern Africa, 2nd edn. Struik, Cape Town
Swanson HK, Lysy M, Power M, Stasko AD, Johnson JD, Reist JD (2015) A new probabilistic method for quantifying n-dimensional ecological niches and niche overlap. Ecology 96:318–324. https://doi.org/10.1890/14-0235.1
Tarkan AS, Karakuş U, Tepeköy EG, Top N, Ozdilek SY, Partal N, Britton JR (2018) Trophic interactions of two Ponto-Caspian gobies in the Turkish part of their native range. Turkish J Fish Aquati Sci 18:1279–1286. https://doi.org/10.4194/1303-2712-v18_11_04
Tedesco PA, Leprieur F, Hugueny B, Brosse S, Dürr HH, Beauchard O, Busson F, Oberdorff T (2012) Patterns and processes of global riverine fish endemism. Glob Ecol Biogeogr 21:977–987. https://doi.org/10.1111/j.1466-8238.2011.00749.x
Top-Karakuş N, Karakuş U, Tarkan AS (2021) Niche segregation of a newly introduced invasive and co-occurring native fish species in a productive Shallow Lake (Manyas, NW Anatolia). J Vertebr Biol 70(21043):1–10
Vander Zanden MJ, Rasmussen JB (1999) Primary consumer δ15N and δ13C and the trophic position of aquatic consumers. Ecology 80:1395–1404. https://doi.org/10.1890/0012-9658(1999)080[1395:PCCANA]2.0.CO;2
Vander Zanden MJ, Chandra S, Allen B, Reuter JE (2003) Goldman CR (2003) historical food web structure and restoration of native aquatic communities in the Lake Tahoe (California–Nevada) basin. Ecosystems 6:274–288. https://doi.org/10.1007/s10021-002-0204-7
Vander Zanden M, Fetzer WW (2007) Global patterns of aquatic food chain length. Oikos 116:1378–1388. https://doi.org/10.1111/j.0030-1299.2007.16036.x
Vander Zanden HB, Soto DX, Bowen GJ, Hobson KA (2016) Expanding the isotopic toolbox: applications of hydrogen and oxygen stable isotope ratios to food web studies. Front Ecol Evol 4:1–19. https://doi.org/10.3389/fevo.2016.00020
Vannote RL, Minshall GW, Cummins KW, Sedell JR, Cushing CE (1980) The river continuum concept. Can J Fish Aquat Sci 37:30–137
Walsworth TE, Budy P, Thiede GP (2013) Longer food chains and crowded niche space: effects of multiple invaders on desert stream food web structure. Ecol Freshw Fish 22:439–452. https://doi.org/10.1111/eff.12038
Weyl OLF, Finlayson B, Impson ND, Woodford DJ, Steinkjer J (2014) Threatened endemic fishes in South Africa’s Cape Floristic Region: a new beginning for the Rondegat River. Fisheries 39:270–279. https://doi.org/10.1080/03632415.2014.914924
Witte F, Goldschmidt T, Wanink J, van Oijen M, Goudswaard K, Witte-Maas E, Bouton N (1992) The destruction of an endemic species flock: quantitative data on the decline of the haplochromine cichlids of Lake Victoria. Environ Biol Fishes 34:1–28. https://doi.org/10.1007/BF00004782
Zambrano L, Martínez-Meyer E, Menezes N, Peterson T (2006) Invasive potential of common carp (Cyprinus carpio) and Nile tilapia (Oreochromis niloticus) in American freshwater systems. Can J Fish Aquat Sci 63:1903–1910. https://doi.org/10.1139/f06-088
Zambrano L, Valiente E, Vander Zanden MJ (2010) Food web overlap among native axolotl (Ambystoma mexicanum) and two exotic fishes: carp (Cyprinus carpio) and tilapia (Oreochromis niloticus) in Xochimilco, Mexico City. Biol Invasions 12:3061–3069. https://doi.org/10.1007/s10530-010-9697-8
Zengeya TA, Booth AJ, Bastos ADS, Chimimba CT (2011) Trophic interrelationships between the exotic Nile tilapia, Oreochromis niloticus and indigenous tilapiine cichlids in a sub-tropical African river system (Limpopo River, South Africa). Environ Biol Fishes 92:479–489. https://doi.org/10.1007/s10641-011-9865-4
Acknowledgements
This study was funded by the Rhodes University Sandisa Imbewu Grant and the National Research Foundation (NRF) through the Department of Science and Technology-National Research Foundation (DST-NRF) Innovation Doctoral Scholarship (SFH170705249077) and Research Development Grant (CSRP190416431023). The authors acknowledge that opinions, findings and conclusions or recommendations expressed in this publication generated by the NRF supported research are that of the authors and that the NRF accepts no liability whatsoever in this regard. Martinus Scheepers, Manda Kambikambi, Nonjabulo Matomela and Edith Phala are thanked for their assistance with field surveys.
Funding
Open access funding provided by Rhodes University. This study was funded by the Rhodes University Sandisa Imbewu Grant and the National Research Foundation (NRF) through Department of Science and Technology-National Research Foundation Innovation Doctoral Scholarship (SFH170705249077) and Research Development Grant (CSRP190416431023).
Author information
Authors and Affiliations
Contributions
PPM and WTK were involved in conceptualisation and designing of the study, data acquisition and analysis, as well as the final approval of the final version to be submitted for review. PPM interpreted data and led the writing of the manuscript. WTK critically evaluated manuscript for important intellectual content.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose. The authors have no conflict of interest to declare pertaining to the contents of this article.
Ethical approval
We also confirm that this study was conducted within the norms and regulations of animal welfare as required by the laws of the country. Ethical approval was granted by Rhodes University Animal Research Ethics Committee (Approval number: 2020–2822-4866). Sampling permission was granted by Eastern Cape’s Department of Economic Development, Environmental Affairs and Tourism (Permit number: CRO 130/19CR).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mpopetsi, P.P., Kadye, W.T. Isotopic diversity and niche patterns reveal contrasting resource use among co-occurring non-native fishes within a flow-altered African river system. Biol Invasions 26, 2095–2117 (2024). https://doi.org/10.1007/s10530-024-03297-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-024-03297-3