Abstract
Agronomical traits of crop plants exhibit quantitative variation that is controlled by multiple genes and is dependent on environmental conditions. The main objective of this study was to decipher the genotype-by-environment interaction (GEI) for six yield-related traits of 25 winter oilseed rape (WOSR) genotypes using the additive main effects and multiplicative interaction (AMMI) model. The genotypes chosen included canola cultivars, our newly developed WOSR breeding lines, yellow-seeded, semi-resynthesized and mutant genotypes, together with ogu-INRA F1 hybrids and their parental lines. These were tested in field trials at two locations over three growing seasons. Field experiments were conducted in a randomized block design with four replicates. We recorded the beginning of flowering, seed yield (SY) and SY components, the number of siliques per plant, the length of siliques, the number of seeds per silique, and the weight of 1000 seeds. The average SY in six environments varied from 16.55 to 41.64 dt·ha−1. The AMMI analysis showed significant effects of both G and E, as well as GEI, for the above traits. In this study, we observed that the climate condition, especially precipitation in addition to the soil type were the most influential factors on the SY and SY-trait value. Seed yield was positively correlated with: the number of siliques per plant, the length of siliques, the number of seeds per silique and the weight of 1000 seeds. We also found that our new ogu-INRA F1 hybrids, as well as cultivars Monolit, Mendel, Starter and Sherlock, showed stability for the analyzed traits.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oilseed rape (Brassica napus L. var. oleifera ssp. napus AACC), also known as rapeseed or canola, is an important oil crop worldwide (Hasan et al. 2006; UFOP 2019). Canola seed oil is considered a major source of edible oil as well as an optimal raw material for green fuel production. Rapeseed oil is also used in the pharmaceutical and chemical industries (Friedt and Snowdon 2009). In 2022, annual global canola grain production reached 87 million tons; 19.41 million tons of this was in the European Union (EU27), with 4.51, 4.29 and 3.4 million tons from France, Germany and Poland, respectively. The average yield for winter oilseed rape (WOSR) in Poland ranged from 24.9 dt ha−1 in 2018 to 25.9 dt ha−1 in 2019 and to 32.3 dt ha−1 in 2022 (FAOSTAT 2024).
To feed the growing human population, further increase in canola grain production is needed. Development of WOSR breeding lines is an important goal of rapeseed breeding programs worldwide exhibiting both good oil quality, high agronomic value with satisfactory resistance to biotic and abiotic stresses (Würschum et al. 2012; Chen et al. 2014; Werner et al. 2018). Genetic information on the gene expression of quantitative traits in the early stage of the plant breeding process can help breeders to create an effective strategy for the development of new high yielding cultivars of canola. Many traits of agronomic importance are controlled by multiple gene of small effect, while their phenotypic expression is strongly affected by no-genetic factors as well as genotype by environment interactions (Piepho 1994; Nowosad et al. 2016; Werner et al. 2018; Bocianowski et al. 2019a). Therefore, increasing seed yield (SY) and SY components are the key goal of many studies and canola breeding programs (Würschum et al. 2012; Chen et al. 2014; Werner et al. 2018).
SY and yield-associated traits in crops are difficult to investigate because they are easily affected by changing environmental conditions. The genetic basis of these complex traits has been explored in many experiments on a range of crop plants, including rice (Rahimsoroush et al. 2021), barley (Wang et al. 2019), maize (Pan et al. 2017) and oilseed rape (Chen et al. 2014; Yang et al. 2017). It has been reported that most agronomical traits exhibit quantitative variation, which is controlled by multiple genes and is environmentally dependent. Moreover, Quarrie et al. (2006) pointed out that crop yield reflects the interaction of the environment with all growth and development processes that occur throughout the life cycle. One of the most important processes involved in crop acclimatization, adaptation and productivity is flowering time (FT). This developmental trait is determined by multiple genes that cause variability in the performance of genotypes and diverse responses to the cultivation environment, where important factors include cold exposure, season, day length, temperature, rainfall, and also biotic and abiotic stresses (Lou et al. 2007; Iniguez-Luy and Federico 2011; Fletcher et al. 2015; Raman et al. 2019). Long et al. (2007) demonstrated significant effects of genotype (G), environment (E) and GE interaction (GEI) on the FT of two segregating F2 populations comprising 202 and 404 lines of oilseed rape. Schiessl et al. (2015) showed that day-length and climatic conditions had an impact on the FT of 158 inbred lines of European winter-type B. napus. Moreover, high oil content in different genotypes was correlated with early flowering and a shorter flowering period.
Leon and Becker (1995) pointed out that every developmental process is, to a greater or lesser extent, under genetic control, and also dependent on environmental factors. SY is a typical quantitative trait in this respect, as it is influenced by many genetic factors as well as by interaction with the environment; consequently, particular genotypes can react differently to changing environmental conditions during plant development (Riaz et al. 2001; Zhang et al. 2013). Yan (2001) reported that SY of a certain genotype in a specific environment is influenced by G main effects, E main effects, and GEI. The authors demonstrated that, in field trials carried out under different agroecological conditions, 80% of yield variation was usually attributable to E, while G and GEI each caused 10% of the variation. Similarly to SY, the expression of SY-related traits, such as the number of branches, the number of siliques per plant, and the number of seeds per silique resulted from the G × year (environment) interactions (Chen et al. 2014; Bocianowski et al. 2017; Łopatyńska et al. 2021). Moreover, changes in climate may significantly influence crop growth and yield, and an understanding of the effects of environmental factors on crop development and growth can improve the selection of specific strains for cultivation in the target regions (Marjanović-Jeromela et al. 2011; Zhang et al. 2013; Bocianowski and Liersch 2021). In particular, comprehensive analyses of SY and yield-associated traits, in addition to their interaction with environmental factors, are crucial for rapeseed yield improvement (Shi et al. 2009; Iniguez-Lay and Federico 2011; Zhang et al. 2013). In addition, Chen et al. (2021) investigated transient daily heat stress during the early reproductive phase of canola under controlled environment. The authors showed that heat stress treatments during the week before first open flower on the main stem or during the first week or second week following first open flower greatly reduced pod number and seed yield. They also defined the conditions for controlled environment screening for heat tolerance in B. napus.
In many crops species the multi-environmental analysis [genotype main effect (G), environment (E) and genotype by environment interaction (GE)] of different breeding materials is widely used for characterization of genotypes from an agronomic point of view. Many researchers tried to define the genotypes suitable for various environments with high adaptability and good stability. For this reason, plant breeders often conduct factorial field experiments to study the influence of GEI on crop yield and yield components. This influence may be assessed by combination analysis of variance (ANOVA), for the G and E main effects, and principal component analysis (PCA), including multiplicative parameters, for GEI effects. The AMMI biplot graphic simultaneously displays both main and interaction effects for genotypes and environments, and enables a single analysis of GEI. Therefore, the AMMI model is also known as ‘interaction PCA’ (1990). This model has been used to analyse GEI in field trials of many crops, such as maize (Branković-Radojčić et al. 2018), wheat (Kumar et al. 2018; Singh et al. 2019), soybean (Sneller and Dombek 1995), ethiopian mustard (Tadesse et al. 2018), mustard and canola (Gunasekera et al. 2006; Alizadeh et al. 2020; Zulfqar et al. 2021; Qasemi et al. 2022; Arinaitwe et al. 2023; Okla et al. 2023; Shojaei et al. 2023; Oghan et al. 2024).
The use of the AMMI model is a tool for identifying the genotypes best suited to specific soil and climatic conditions. Measures of stability allow to distinguish stable and specific genotypes. One of the best measures of genotype stability is the AMMI stability value coefficient based on the first two interacting principal components (Bocianowski et al. 2019b, 2021; Bocianowski and Prażak 2022).
The main objective of this study was to use the AMMI model to estimate GEI for 25 WOSR genotypes in six environments for the following SY-related traits: beginning of flowering (BF), SY and SY components i.e. the number of siliques per plant (NSP), the length of siliques (LS), the number of seeds per silique (NSS), and the weight of 1000 seeds (WTS).
Materials and methods
Plant material and field experiment
A collection of 25 genotypes of WOSR (Brassica napus L.) was used in this study: seven Polish and European cultivars (Monolit, Brendy, Starter, Mendel, Polka, Sherlock, Zornyj), three CMS ogu-INRA hybrids (F1_952, F1_S2, F1_239), six parental forms of the ogu-INRA system, i.e. three CMS lines (CMS_64, CMS_313, CMS_1612) and three DH restorer lines (Rfo_37, Rfo_38, Rfo_39), DH breeding line (DH D × C), double low quality breeding lines (HO_TP, HO_TP_00, 00_SS), a yellow-seeded line (Z 114), a high-oleic mutant line (HO_SS), and a mutant recombinant line with changed unsaturated fatty acid composition in seed oil (i.e., high oleic and low linolenic) (HOLL_SS), as well as a semi-resynthetic (semi-RS) WOSR lines (R43, S1) (Table 1). Hybrid seeds resulting from crosses between the male-sterile CMS lines and the restorer lines has been produced in isolation tents under field conditions. Field trials were performed in four replicates of a randomized block design at the Experimental Station of Plant Breeding Company Strzelce Ltd. in Borowo (E1, E2 and E3, respectively) and Plant Breeding Company Smolice Ltd. in Łagiewniki (E4, E5 and E6, respectively), Poland, during the crop seasons 2014/2015, 2015/2016 and 2016/2017. Genotypes were grown in a four-row plots of 10 m2 (Borowo) and 9.6 m2 (Łagiewniki) with a 0.30 row distance and a sowing density of 70 seeds m−2. Agricultural practice were optimal for local agroecological conditions in both of the investigated locations. Field experiments were conducted on podzolic soil of quality class IIIa (E1, E2) and IVa (E3) in Borowo and on typical brown soil of quality class IIIa in Łagiewniki (E4–E6). The more details of the plant materials and the field trials were described by Liersch et al. (2020). The monthly temperatures and precipitations in each growing season, from the end of August (sowing) until the end of July of the next year (harvest), are presented in Fig. 1.
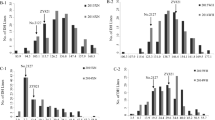
Weather conditions during growing seasons in six environments (E1–E6); dark grey bars, mean monthly precipitation in growing season [mm]; grey bars many-years monthly precipitation [mm]; black solid and grey dashed lines, mean monthly temperatures in growing season and mean many-years monthly temperatures [°C], respectively
Quantitative traits
Experimental data were collected on both morphological and seed features. The beginning of flowering time (days) was defined as the number of the days from the beginning of the year to the beginning of flowering (i.e. when 50% of plants have their first flowers). The number of siliques per plant was the number of well-filled, normally developed siliques from five, randomly selected plants from the middle of each plot. At the maturity stage, siliques from the primary branches of five representative plants from the center of each plot were collected and then dried. Silique length (mm) and the number of seeds per silique were determined on twenty-five well-developed siliques. Seeds were harvested at the seed maturity stage using a plot harvester. Seed yields per plot, which was either 10.0 m2 (Borowo) or 9.6 m2 (Łagiewniki) in area, were measured and converted to dt∙ha−1 for all statistical analyses. The weight of a 1000 seeds (g) was calculated from the average of three measurements from the mixed seeds of each plot.
Statistical analysis
Data were analyzed using the additive main effects and multiplicative interaction model Gauch and Zobel (1990) for each trait, independently. The AMMI model first fits additive effects for the main effects of genotype (G) and environment (E) followed by multiplicative effects for GEI by PCA. The results of AMMI analysis are presented in biplot graphs. The AMMI model (Nowosad et al. 2016) is given by:
where yge is the trait mean of genotype g in environment e, μ is the grand mean, αg are the genotypic mean deviations, βe are the environment mean deviations, N is the number of PCA axis retained in the adjusted model, λn is the eigenvalue of the PCA axis n, γgn is the genotype score for PCA axis n, δen is the score eigenvectors for PCA axis n, Qge is the residuals, including AMMI noise and pooled experimental error. The AMMI stability value (ASV) was used to compare the stability of genotype as described by Purchase et al. (2000):
where SSIPCA1 is the sum of squares for IPCA1, SSIPCA2 is the sum of squares for IPCA2, and the IPCA1 and IPCA2 scores are the genotypic scores in the AMMI model. A lower ASV score indicates a more stable genotype over several environments (Nowosad et al. 2017).
The genotype selection index (GSI) was calculated for each genotype. This incorporates both mean of trait and ASV index in a single criterion (GSIi) as described by Farshadfar and Sutka (2003):
where RMi is the rank of trait mean (from maximal to minimal for SY the NSP, LS, NSS and WTS, and from minimal to maximal for BF) for the i-th genotype, and RAi is the rank of the ASV for the i-th genotype. Finally, the total genotype selection index (TGSI) was calculated for each genotype as the sum of GSIs for all six traits in the study. Correlation analysis between observed traits was performed on genotypic averages. All analyses were conducted using the GenStat v. 18 statistical software package.
Results
We analyzed 25 OSR genotypes including open-pollinated and hybrid cultivars, breeding lines of canola type, Ogura CMS and Rfo canola breeding lines and lines with different C18 mono- and polyunsaturated fatty acid content in seed oil and lines characterized by different seed colour. Mean values of all traits are shown in Table 2.
The three sources of variation (G, E and GEI) were highly significant for all six observed traits of WOSR (Table 2).
Beginning of flowering (BF)
The impact of the environment on BF was relatively low, but significant, representing 10.80% of the total variation. The effects of G explained 67.16% of the variation, while the effects of GEI explained 19.50% of the variation (Table 3). Values for the three principal components were also found to be highly significant and jointly accounted for 92.59% of overall variation in BF. IPCA 1 accounted for 46.75% of the variation caused by the interaction, while IPCA 2 and IPCA 3 accounted for 33.66% and 12.18% of the variation, respectively. Across the 25 genotypes in the six environments, BF varied from 109.5 days for Starter (G03) and Rfo_39 (G09) in E2 (Borowo 2016) to 130.8 days for HO_SS (G13) in E6 (Łagiewniki 2017), with the average BF of 118.5 days (Table 4). Across all environments, the cultivar Starter (G03) was the earliest to flower (the average of 112 days), while the latest was the genotype HO_SS (G13) (the average of 126 days). In each environment, the average BF across all genotypes varied from 117 in E5 (Łagiewniki 2016) to 121.1 days in E3 (Borowo 2017) (Table 4). The stability of the genotypes in the six environments with respect to BF was visualized as a biplot (Fig. 2). The WOSR genotypes interacted differently with respect to the climate conditions in each environment. The genotypes HOLL_SS (G15), S43 (G20) and cultivar Zornyj (G23) interacted positively with environment E5 (Łagiewniki 2016), but negatively with environments E1 (Borowo 2015) and E2 (Borowo 2016) (Fig. 2). The cultivars Starter (G03), Mendel (G04) and Rfo_39 (G09) interacted positively with E4 (Łagiewniki 2015) and E6 (Łagiewniki 2017), but negatively with E2 (Borowo 2016). The analysis showed that some genotypes have a high adaptation capacity; however, most of them have specific adaptability. AMMI stability values (ASVs) revealed variations in BF stability among the genotypes (Table 4). The Z_114 (G05), Brendy (G02) and F1_239 (G25) genotypes, which showed an ASV of 0.198, 0.393 and 0.397, respectively, were found to be the most stable, while HO_SS (G13) and cultivar Zornyj (G23) with an ASV of 3.358 and 2.311, respectively, were the least stable (Table 4). Genotype G05, with a low average BF (115.7 days), had the best ASV (0.198) and genotype selection index (GSI) (6), while the HO_SS (G13) breeding line was characterized by the worst GSI (50).
Biplot for genotype by environment interaction of beginning of flowering in winter oilseed rape genotypes in six environments, showing the effects of primary and secondary components (IPCA 1 and IPCA 2, respectively). E1—Borowo 2015, E2—Borowo 2016, E3—Borowo 2017, E4—Łagiewniki 2015, E5—Łagiewniki 2016, E6—Łagiewniki 2017
Seed yield (SY)
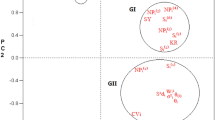
ANOVA showed that the sum of squares for the environment main effects represented 42.15% of the SY variation. The differences between the genotypes explained 36.00% of the SY variation, while the effects of GEI explained 11.52% (Table 3). The values of the three principal components were also found to be highly significant and jointly accounted for 81.61% of the total variation. IPCA 1 accounted for 41.29% of the variation caused by the interaction, while IPCA 2 and IPCA 3 accounted for 22.48 and 17.84%, respectively. SY of the 25 genotypes across the six environments varied from 5.5 dt ha−1 for HO_SS (G13) in Borowo 2017 (E3) to 61.33 dt ha−1 for Starter (G03) in Łagiewniki 2015 (E4), with an average of 29.95 dt ha−1 (Table 5). The cultivar Starter (G03) had the highest average SY (41.08 dt ha−1), while the genotype HO_SS (G13) had the lowest (10.5 dt ha−1). The average SY in each environment varied from 16.55 dt ha−1 in E2 to 41.64 dt ha−1 in E4. The genotypes HO_SS (G13), HOLL_SS (G15), HO_TP (G17), as well as cultivar Zornyj (G23), interacted positively with environment E2 (Borowo 2016), but negatively with E1 (Borowo 2015) and E5 (Łagiewniki 2016) (Fig. 3). Cultivar Mendel (G04) and the S1 breeding line (G21) interacted positively with E4 (Łagiewniki 2015), but negatively with E3 (Borowo 2017) and E6 (Łagiewniki 2017). Genotypes DH_D × C (G05) and CMS_64 (G10) interacted positively with E5 (Łagiewniki 2016). The HO-type cultivar Polka (G16) and the CMS_313 line (G11) returned ASVs of 0.351 and 0.447, respectively, and were thus relatively stable, while genotypes HO_SS (G13) and HOLL_SS (G15), with ASVs of 5.817 and 4.223, respectively, were the least stable (Table 5). Cultivar Brendy (G02) and genotype CMS_313 (G11), with average SYs of, respectively, 36.6 and 33.12 dt ha−1, and ASVs of, respectively, 0.866 and 0.447, were characterized by the best GSI value of 11, compared to the HO_SS (G13) line, which had the worst GSI value (50).
Biplot for genotype by environment interaction of seed yield in winter oilseed rape genotypes in six environments, showing the effects of primary and secondary components (IPCA 1 and IPCA 2, respectively). E1—Borowo 2015, E2—Borowo 2016, E3—Borowo 2017, E4—Łagiewniki 2015, E5—Łagiewniki 2016, E6—Łagiewniki 2017
Number of siliques per plant (NSP)
The three sources of variation (G, E, GEI) were found to be highly significant with respect to NSP. The impact of the environment on this trait was low, albeit significant, representing only 6.26% of the variation. The differences between genotypes explained 57.25% of the variation in NSP, while the effects of GEI explained 24.94% (Table 3). The values of the three principal components (PCs) were also found to be highly significant and jointly accounted for 87.92% of overall variation in NSP. IPCA 1 accounted for 51.51% of the variation caused by the interaction, while IPCA 2 and IPCA 3 accounted for 23.86 and 12.55% of the variation, respectively. Among the tested genotypes, NSP varied across the six environments from 138.4 for HO_SS (G13) in E3 (Borowo 2017) to 392 for F1_239 (G25) in E3 (Borowo 2017), with an average of 263.17 (Table 6). The hybrid F1_S2 (G24) had the highest average NSP (335.2), while genotype HO_SS (G13) had the lowest NSP (166.1). The average NSP in each environment varied from 246.3 in E6 to 283.4 in E1. The cultivar Monolit (G01), as well as the HOLL_SS (G15), S1 (G21) and F1_239 (G25) breeding lines interacted positively with environments E1 (Borowo 2015) and E3 (Borowo 2017), but negatively with E4 (Łagiewniki 2015) and E6 (Łagiewniki 2017) (Fig. 4). The Z114 (G06), CMS_1612 (G12) and HO_SS (G13) breeding lines and Zornyj (G23) interacted positively with E5 (Łagiewniki 2016), but negatively with E2 (Borowo 2016). The F1_952 hybrid (G19) and Rfo_38 (G08) returned an ASV of 0.892 and 1.034, respectively, showing them to be the most stable, while the S43 (G20) and F1_239 hybrid (G25) genotypes, with an ASV of 18.883 and 12.562, respectively, were the least stable (Table 6). The Rfo_38 (G08) breeding line with average NSP equal to 291.2 and an ASV of 1.034 had the best GSI (10), while genotype HO_SS (G13) was characterized by the worst GSI value (45).
Biplot for genotype by environment interaction of the number of siliques per plant in winter oilseed rape genotypes in six environments, showing the effects of primary and secondary components (IPCA 1 and IPCA 2, respectively). E1—Borowo 2015, E2—Borowo 2016, E3—Borowo 2017, E4—Łagiewniki 2015, E5—Łagiewniki 2016, E6—Łagiewniki 2017
Length of siliques (LS)
The three sources of variation were highly significant with respect to LS. However, genotype was the most important factor. The environment main effect represented only 3.99% of the total variation. The differences between genotypes explained 57.04% of the total variation in LS, while the effects of GEI represented 28.44% of the variation (Table 3). The values of the first three PCs were also found to be highly significant and jointly accounted for 79.44% of the overall variation in LS. IPCA 1 explained 31.77% of the variation caused by the interaction, while IPCA 2 and IPCA 3 accounted for 28.29 and 19.37%, respectively. Across the 25 genotypes in the six environments, LS varied from 41.05 mm for the S43 strain (G20) in E1 (Borowo 2015) to 74.5 mm for Zornyj (G23) in E6 (Łagiewniki 2017), with an average of 61.59 mm (Table 7). Cultivar Zornyj (G23) had the highest average LS (69.29 mm), while the S43 (G20) genotype had the lowest (50.79 mm). The average LS in each environment varied from 60.02 mm for Łagiewniki 2015 (E4) to 63.72 mm for Łagiewniki 2016 (E5). The stability of the genotypes in the six environments with respect to LS was visualized as a biplot (Fig. 5). Cultivars Monolit (G01), Polka (G16) and S1 (G21) interacted positively with environment E3 (Borowo 2017), but negatively with E6 (Łagiewniki 2017) (Fig. 5). Cultivar Brendy (G02), together with HO_SS (G13) and HO_TP_00 (G18), interacted positively with environments E1 (Borowo 2015) and E4 (Łagiewniki 2015), but negatively with environments E2 (Borowo 2016) and E5 (Łagiewniki 2016). The Rfo_39 (G09) and Rfo_37 (G07) breeding lines, with an ASV of 0.263 and 0.388, respectively, were identified as the most stable, while the S43 breeding line (G20) and the F1_952 hybrid (G19), with an ASV index of 2.942 and 2.286, respectively, were the least stable (Table 7). The cultivar Sherlock (G22), with a high average LS (66.89 cm) and an ASV index of 0.417, had the best GSI of 7 (Table 7), while the S43 breeding line (G20) had the worst GSI (50).
Biplot for genotype by environment interaction of length of silique in winter oilseed rape genotypes in six environments, showing the effects of primary and secondary components (IPCA 1 and IPCA 2, respectively). E1—Borowo 2015, E2—Borowo 2016, E3—Borowo 2017, E4—Łagiewniki 2015, E5—Łagiewniki 2016, E6—Łagiewniki 2017
Number of seeds per silique (NSS)
The three sources of variation were found to be highly significant with respect to NSS. In ANOVA, the sum of squares for the environment main effect represented 21.86% of the total variation in NSS, while the differences between the genotypes explained 38.15% of the total variation. At the same time, the effects of GEI explained 30.83% of the variation in NSS (Table 3). The values of the three principal components were also found to be highly significant and jointly accounted for 78.94% of the total variation in NSS. IPCA 1 accounted for 38.15% of the variation caused by the interaction, while IPCA 2 and IPCA 3 represented 25.03 and 15.75%, respectively. Of the 25 genotypes across the six environments, NSS varied from 10.1 for the Rfo_39 breeding line (G09) in E2 (Borowo 2016) to 26.02 for the Rfo_39 breeding line (G09) in E4 (Łagiewniki 2015), with an average of 18.24 (Table 8). The cultivar Sherlock (G22) had the highest average NSS (23.19), while the genotype HOLL_SS (G15) had the lowest (13.61). The average NSS in each environment varied from 15.88 in E2 to 21.24 in E4. The stability of the genotypes with respect to NSS was visualized as a biplot (Fig. 6). The genotypes Rfo_37 (G07), Rfo_39 (G09) and HO_SS (G13), as well as S1 (G21), interacted positively with environments E1 (Borowo 2015) and E4 (Łagiewniki 2015), but negatively with E2 (Borowo 2016), E3 (Borowo 2017) and E6 (Łagiewniki 2017) (Fig. 6). Cultivar Monolit (G01), as well as F1 hybrids G19 and G25, interacted positively with E2 (Borowo 2016), but negatively with E5 (Łagiewniki 2016). Cultivars Zornyj (G23) and Starter (G03), as well as the HO_TP (G17) breeding line, with ASVs of 0.075, 0.276 and 0.232, respectively, were found to be the most stable, while the genotypes CMS_1612 (G12) and Rfo_37 (G07) revealed the lowest stability, of 2.365 and 2.348, respectively (Table 8). Cultivar Mendel (G04) and the HO_TP (G17) line revealed a high average NSS (21.47 and 19.8, respectively) and high stability (0.822 and 0.232, respectively). They were also characterized by the best GSI values (8). The Rfo_37 (G07) line had the worst GSI (45).
Biplot for genotype by environment interaction of the number of seeds per silique in winter oilseed rape genotypes in six environments, showing the effects of primary and secondary components (IPCA 1 and IPCA 2, respectively). E1—Borowo 2015, E2—Borowo 2016, E3—Borowo 2017, E4—Łagiewniki 2015, E5—Łagiewniki 2016, E6—Łagiewniki 2017
Weight of 1000 seeds (WTS)
ANOVA showed that the sum of squares for genotype main effects represented 48.26% of the total variation in WTS indicating that the trait depended mostly on G. The differences between the environments explained 27.19% of the total variation in WTS, while the effects of GEI explained 16.42% (Table 3). The values of the three PCs were also found to be highly significant and jointly accounted for 85.91% of the overall effect on variation in WTS. IPCA 1 accounted for 45.57% of the variation caused by the interaction, while IPCA 2 and IPCA 3 represented 24.11% and 16.23%, respectively (Table 3, Fig. 7). Of the 25 genotypes across the six environments, WTS varied from 3.33 g for S43 (G20) in E1 (Borowo 2015) to 6.357 g for the DH_D × C breeding line (G05) in E4 (Łagiewniki 2015), with an average of 5.005 g (Table 9). The F1 hybrid 239 (G25) had the highest average WTS (5.748 g), while the HO_SS breeding line (G13) had the lowest (4.048 g). The average WTS value in each environment varied from 4.63 g in E2 (Borowo 2016) to 5.344 g in E4 (Łagiewniki 2015). The stability of the genotypes in the six environments with respect to WTS was visualized as a biplot (Fig. 7). The breeding lines Rfo_38 (G08) and HOLL_SS (G15) interacted positively with E2 and E5, but negatively with E4 (Fig. 7). The genotypes G06, G07, G20 and G21 interacted positively with E3 (Borowo 2017), but negatively with E1 (Borowo 2015). The F1 hybrids 239 (G25) and 952 (G19), with an ASV of 0.021 and 0.092, respectively, were found to be the most stable, while DH_D × C (G05) and S43 (G20), with an ASV of 1.341 and 1.112, respectively were the least stable (Table 9). A F1 hybrid 239 (G25) with the highest average WTS (5.748 g) and an ASV of 0.021 also had the best GSI (2), whereas the S43 (G20) line had the worst GSI value (48).
Biplot for genotype by environment interaction of weight of 1000 seeds in winter oilseed rape genotypes in six environments, showing the effects of primary and secondary components (IPCA 1 and IPCA 2, respectively). E1—Borowo 2015, E2—Borowo 2016, E3—Borowo 2017, E4—Łagiewniki 2015, E5—Łagiewniki 2016, E6—Łagiewniki 2017
Total genotype selection index (TGSI)
Jointly for the six analyzed traits, the best total genotype selection index value (96.5) was returned for the F1 hybrid 239 (G25) while the HO_SS (G13) line had the worst (260) (Table 2). Therefore, the G25 genotype can be recommended for inclusion in further breeding programs due to its stability and good average values of SY-related traits.
Correlation analysis
Seed yield was positively correlated with: the number of siliques per plant (0.52), the length of siliques (0.55), the number of seeds per silique (0.72) and the weight of 1000 seeds (0.57); while negatively correlated with beginning of flowering (−0.77). In addition, beginning of flowering was negatively correlated with: the number of siliques per plant (−0.43), the number of seeds per silique (−0.48) and the weight of 1000 seeds (−0.53). The length of siliques was positively correlated with the number of seeds per silique (0.69) and the weight of 1000 seeds (0.59) (Fig. 8).
Discussion
The yield of major crop plants is a consequence of plant development, which depends on an appropriate response to environmental and agrotechnical conditions, including adaptation of the plant reproductive system to the prevailing climatic conditions (i.e., vernalization, water supply, day length and temperature), the response to biotic and abiotic stresses, such as storms, in addition to timely and properly performed agrotechnical treatments. In our study we showed a significant influence of G, E and GEI on all rapeseed traits investigated: the beginning of flowering time, seed yield and seed -yield components, i.e., the number of siliques per plant, the length of siliques, the number of seeds per silique and the weight of 1000 seeds.
At both localities, Borowo and Łagiewniki, there was variability in soil type and water supply across the site, i.e., the field experiments were conducted on different types of soil. At Borowo, the various soil types were classified as light, loamy sand and as “medium” or “heavy soil”, while at Łagiewniki the soil was sandy clay loam. The topsoil was slightly acidic, and the pH values ranged from 6.0 (E1, E2, E3) to 6.2 (E4 and E6). Agricultural practices were optimal for the agroecological conditions, and the timing and rate of fertilization were the same in each environment. At the same time, the field trial results demonstrated the impact of weather, especially of temperature and rainfall, on plant development, including germination and wintering (not shown), flowering time, SY and SY-related traits.
Crop-plant species grown under different agroecological conditions reveal phenotypical differences that arise in response to particular environmental parameters, such as soil type, agrotechnical treatments, in addition to weather conditions, such as cold exposure, day length and rainfall. The beginning of flowering time and length of flowering time are determined by multiple genes and quantitative trait loci localized in both the A and C genomes of B. napus, thus leading to a diverse response to the cultivation environment among the various rapeseed genotypes (Long et al. 2007; Iniguez-Luz and Federico 2011; Raman et al. 2011; Fletcher et al. 2015; Schiessl et al. 2015; Ghanbari et al. 2020). Osborn et al. (1997) reported the flowering time as a trait with a high degree of heritability but showing a strong interaction with environmental conditions. In populations of 391 elite OSR and 156 WOSR genotypes, heritability (h2) was 0.91 and 0.84, respectively (Würschum et al. 2012), and ranged from 77 to 90% in other populations (Long et al. 2007). However, the results showed both G and G × E variance in these populations. Similar results were reported by Bocianowski et al. (2017; 2018) for different WOSR cultivars and breeding materials. In our experiment, BF varied considerably with genotype, year, and location, and depended on the temperature in April: BF ranged from 117 days (Łagiewniki 2016) to 121.1 days (Borowo 2017), and flowering began earlier when the average temperature in April was higher (Borowo 2015, 2016 and Łagiewniki 2016).
In this study, the G and GEI variance was significant for SY and yield-related traits of the 25 WOSR genotypes. In both locations, genotype had a significant impact on SY, NSP, LS, NSS and WTS. At the same time, we found a statistically significant influence of E and GEI on the traits analyzed. Studies of G, E and GEI for traits related to plant development and SY in OSR are frequent (Seyis et al. 2006; Spasibionek et al. 2020; Bocianowski and Liersch 2021). Additionally, many reports have pointed out that traits of agronomic importance are controlled by multiple genes, whose expression is the result of the interaction between genotype and environmental factors (Zhao et al. 2005; Long et al. 2007; Shi et al. 2009; Fletcher et al. 2015). Szała et al. (2019) observed a G and E main effect as well as GEI in field trials including fifteen F1 Ogura hybrids and their parental lines in three locations. Both Bocianowski et al. (2017) and Spasibionek et al. (2020) studied SY of different genotypes of WOSR and determined GEI as well the G effect based on the results of field experiments. Nowosad et al. (2016) showed that SY of 25 WOSR genotypes significantly depended on both genetic and environmental factors, where climate conditions were the most influential factors in their experiments in the western part of West Poland. In our study, we observed strong environmental factors, especially precipitation and soil water availability, which depended on the type of soil. A higher SY was observed in Łagiewniki (E4–E5)—respectively 41.64, 30.75 and 28.84 dt ha−1—due to its likely more fertile soil than found in Borowo (E1–E3), where SY was lower, respectively 36.13, 16.55 and 25.84 dt ha−1. Additionally, during the seed maturation period just before harvest, heavy rainfall in July negatively influenced SY in Borowo in 2016 (E2).
NSP and the number of plants per unit area are generally considered to be the most important traits affecting SY in B. napus, followed by NSS and WTS (Wójtowicz 2013). ANOVA indicated that the main effects of genotype and environment, as well as GE interaction, were significant for all yield component traits. Significant differences between the environments (two years) and agronomic traits of WOSR, such as WTS, NSP and SPS, were noted by Chen et al. (2014) and Wolko et al. (2019). ANOVA performed by Shi et al. (2011) revealed that G, E and GEI have significant effects on the performance of 15 yield-correlated traits in rapeseed. Also, Bocianowski et al. (2017), in their field trials of seven different types of WOSR cultivar, observed a significant effect of cultivar × location interaction for LS and BF, and significant effects of cultivar × year × location for the beginning, end and length of flowering, as well as for WTS and oil content. In our research, yield-correlated traits were primarily defined by genotype, and, to a lesser extent, by E and GEI. NSP was higher in Borowo (283.4, 273.2 and 271.6 for E1, E2 and E3, respectively) than in Łagiewniki (255.3, 249.2 and 246.3 for E4, E5 and E6, respectively). The values of other yield components were higher in Łagiewniki. The average LS in Borowo (E1-E3) was respectively 61.02, 60.31 and 62.76 mm, while in Łagiewniki (E4-E6) it was 60.02, 63.72 and 61.68 mm, respectively. The average NSS varied across the genotypes from 18.88 (E2) to 18.54 (E1) in Borowo, and from 17.48 (E6) to 21.24 (E4). At the same time, the mean value of WTS was higher in Łagiewniki (4.969–5.344 g) than in Borowo (4.63–5.141 g). Wójtowicz (2013) concluded in their study that environment has the strongest influence on NSP and the number of plants per unit area, rather than NSS and WTS. At the same time, the author pointed out that water shortage during the flowering period and during the formation of siliques was crucial in limiting the plants’ ability to produce siliques. We noted a similar relationship in our study. The number of generative organs per plant, the number and length of siliques, SY and seed-yield components depend strongly on the humidity conditions and availability of water during the growing season.
In agronomic research, use of the correct statistical model is crucial for revealing how the quantitative traits of the various genotypes studied are affected by environmental factors and agrotechnics (Schmidt et al. 2001; Ma et al. 2004). Environmental conditions may have a different effect on phenotypic traits depending on the particular genotype, thus leading to different GEI. The stability or instability of a genotype can be determined by a series of multi-environment field trials. When phenotypic traits, such as yield and its components, vary under different environmental conditions, G is assessed as unstable. From the breeders’ and farmers’ point of view, G or G with little E influence is the most desirable. In our study, we tested several canola type WOSR accessions with different genetic backgrounds: cultivars of different origin, F1 Ogura hybrids, CMS ogura and restorer lines, semi-resynthesized lines, and yellow-seeded lines, recombinant breeding lines, and lines with different fatty acid composition or breeding lines typically used for canola seed oil production. In plants, all life processes depend on temperature and availability of water, which is used for nutrient uptake and transport, transpiration, and all biochemical and physiological processes. The weather conditions in Poland varied over the three growing seasons in our study (Fig. 1, Fig. S1). In 2014/2015, there were strong abiotic stresses caused by drought, from December 2014 to the end of June 2015. The sum of rainfall in winter and spring 2014/2015 was 191.4 mm in Borowo and 195.8 mm in Łagiewniki, being markedly lower than the average for the period 1957–2016 (318.0 mm in Borowo; 326.0 mm in Łagiewniki, respectively). Also, in 2015/2016 the weather conditions were unfavorable; the lack of snow cover and extremely low temperatures at the beginning of January 2016 caused severe damage to the plants. Moreover, heavy rainfall in July 2016, just before harvest, significantly reduced the SY in Borowo (Fig. 1, Fig. S1). In this study we observed that the weather in addition to the soil type significantly influenced plant development which was reflected in the SY and SY-trait values. At the same time, many authors have shown that quantitative traits in WOSR are affected by the different responses of particular genotypes to varying environmental conditions during development (Shi et al. 2009; Fletcher et al. 2015; Thian et al. 2017; Spasibionek et al. 2020).
In this study, we applied the AMMI model for statistical analyses. This is a two-factor model describing the sum of the main additive effects for G (e.g., varieties, lines, soil cultivation methods), E (year, location or a combination of year and location), and the GEI in a multiplicative form. AMMI models for interpreting GEI have been used in several quality breeding programs and also in plant breeding multi-environmental trials (Vargas et al. 1999; Farshadfar and Sutka 2003; Nowosad et al. 2016). Using this approach, we observed significant GEI for BF and SY-related traits. The TGSI index values allowed us to assess the adaptability and stability of genotypes and to discriminate among the genotypes for particular selection and breeding strategies.
Analysis of genotype stability is a very important part of research and breeding work. Especially when there are large soil and climate variations. A large emphasis on stability studies in research was conducted by: Verma and Singh (2020), Jędzura et al. (2023), Oroian et al., (2023), Filio et al., (2023), Hossain et al. (2023), Kumar et al. (2023), Eradasappa et al. (2024), van der Merwe et al. (2024). In the research presented here, we distinguished genotypes stable through all the environments considered.
Conclusions
This study illustrates how multivariate statistical analysis can describe the interaction effect of a genotype with a particular environment for SY and other quantitative traits in WOSR. AMMI analysis of field-trial data for a collection of 25 WOSR genotypes demonstrated a significant impact of E on SY. We conclude that all traits, with the exception of SY, were influenced mostly by G and to a lesser, but statistically significant, extent by E as well as by GEI.
The F1_239 hybrid (G25) showed significant adaptability and stability with respect to BF and WTS, while hybrid F1_952 (G19) showed adaptability and stability for NSP and WTS. The WOSR cultivars Monolit (G01), Starter (G03), Mendel (G04) and Sherlock (G22), as well as the breeding line HO_TP (G17) and the Ogura hybrid F1_239 (G25), had the best TGSI. Two breeding lines, HO_SS and S43, were the least stable, having a low TGSI.
Data availability
Plant Breeding and Acclimatization Institute-National Research Institute in Radzików (PBAI-NRI), Department of Oilseed Crops, Strzeszyńska 36, 60-479 Poznań, Poland.
Change history
07 August 2024
A Correction to this paper has been published: https://doi.org/10.1007/s10681-024-03396-1
References
Alizadeh B, Pasban Eslam B, Rezaizad A, Yazdandoost Hamadani M, Mostafavirad M (2020) Yield stability assessment of winter oilseed rape lines in cold regions of Iran Using AMMI model. J Plant Prod Res 27(3):85–96. https://doi.org/10.22069/jopp.2020.16557.2509
Arinaitwe U, Clay SA, Nleya T (2023) Growth, yield, and yield stability of canola in the Northern Great Plains of the United States. Agron J 115:744–758. https://doi.org/10.1002/agj2.21269
Bocianowski J, Liersch A (2021) Multi-environmental evaluation of winter oilseed rape genotypic performance using mixed models. Euphytica 217:80. https://doi.org/10.1007/s10681-020-022760-1
Bocianowski J, Prażak R (2022) Genotype by year interaction for selected quantitative traits in hybrid lines of Triticum aestivum L. with Aegilops kotschyi Boiss. and Ae. variabilis Eig. using the additive main effects and multiplicative interaction model. Euphytica 218(2):11
Bocianowski J, Liersch A, Nowosad K, Bartkowiak-Broda I (2017) Variability of the agronomic characters in different types of cultivars of winter oilseed rape (Brassica napus L.). Agronomy 29:3–11
Bocianowski J, Nowosad K, Liersch A, Popławska W, Łącka A (2018) Genotype by environment interaction for lenght of flowering time in winter oilseed rape (Brassica napus L.) using additive main effects and multiplicative interaction model. Colloquium Biometricum 48:27–38
Bocianowski J, Księżak J, Nowosad K (2019a) Genotype by environment interaction for seeds yield in pea (Pisum sativum L.) using additive main effects and multiplicative interaction model. Euphytica 215:191
Bocianowski J, Warzecha T, Nowosad K, Bathelt R (2019b) Genotype by environment interaction using AMMI model and estimation of additive and epistasis gene effects for 1000-kernel weight in spring barley (Hordeum vulgare L.). J Appl Genet 60(2):127–135
Bocianowski J, Radkowski A, Nowosad K, Radkowska I, Zieliński A (2021) The impact of genotype-by-environment interaction on the dry matter yield and chemical composition in timothy (Phleum pratense L.) examined by using the additive main effects and multiplicative interaction model. Grass Forage Sci 76(4):463–484
Branković-Radojčić D, Babić V, Girek Z, Živanović T, Radojčić A, Filipović M, Srdić J (2018) Evaluation of maize grain yield stability by AMMI analysis. Genetika 50(3):1067–1080. https://doi.org/10.2298/GENSR1803067B
Chen B, Xu K, Li J, Li F, Qiao J, Li H, Gao G, Yan G, Wu X (2014) Evaluation of yield and agronomic traits and their genetic variation in 488 global collections of Brassica napus L. Genet Resour Crop Eval 61:979–999. https://doi.org/10.1007/s10722-014-0091-8
Chen S, Stefanova K, Siddique KHM, Cowling WA (2021) Transient daily heat stress during the early reproductive phase disrupts pod and seed development in Brassica napus L. Food Energy Secur 10:e262. https://doi.org/10.1002/fes3.262
Eradasappa E, Mohana GS, Poduval M, Sethi K, Aneesa Rani MS, Lourdusamy IK, Velmurugan S, Manjusha M, Raviprasad TN, Anilkumar C (2024) Analysis of stability for nut yield and ancillary traits in cashew (Anacardium occidentale L.). Sci Rep 14:2127. https://doi.org/10.1038/s41598-024-52030-6
FAOSTAT (2024) Food and Agriculture Organization of the United Nations. Available online: http://www.fao.org/faostat/en/#home
Farshadfar E, Sutka J (2003) Locating QTLs controlling adaptation in wheat using AMMI model. Cereal Res Commun 31:249–256. https://doi.org/10.1007/BF03543351
Filio YL, Maulana H, Aulia R, Suganda T, Ulimaz TA, Aziza V, Concibido V, Karuniawan A (2023) Evaluation of Indonesian Butterfly Pea (Clitoria ternatea L.) using stability analysis and sustainability index. Sustainability 15(3):2459. https://doi.org/10.3390/su15032459
Fletcher RS, Mullen JL, Heiliger A, McKay JK (2015) QTL analysis of root morphology, flowering time, and yield reveals trade-offs in response to drought in Brassica napus. J Exp Bot 66(1):245–256. https://doi.org/10.1093/jxb/eru423
Friedt W, Snowdon RJ (2009) Oilseed Rape. In: Vollmann J, Rajcan I (eds) Handbook of Plant Breeding. Oil Crops, vol 4. Springer, New York, pp 91–120
Gauch HG, Zobel RW (1990) Imputing missing yield trial data. Theor Appl Genet 79:753–761. https://doi.org/10.1007/BF00224240
Ghanbari M, Madhuri P, Möllers C (2020) QTL analysis of shoot elongation before winter in relations to vernalization requirenement in the doubled haploid population L16 × Express617 (Brassica napus L.). Euphytica 216:73. https://doi.org/10.1007/s10681-020-02604-y
Gunasekera CP, Martin LD, Siddique KHM, Walton GH (2006) Genotype by environment interaction of Indian mustard (Brassica juncea L.) and canola (B. napus L.) in Mediterranean-type environments. I. Crop growth and seed yield. Eur J Agron 25:1–12. https://doi.org/10.1016/j.eja.2005.08.002
Hasan M, Seyis F, Badani AG, Pons-Kühnemann J, Friedt W, Lühs W, Snowdon RJ (2006) Analysis of genetic diversity in the Brassica napus L. gene pool using SSR markers. Genet Resour Crop Evol 53:793–802. https://doi.org/10.1007/s10722-004-5541-2
Hossain MA, Sarker U, Azam MG, Kobir MS, Roychowdhury R, Ercisli S, Ali D, Oba S, Golokhvast KS (2023) Integrating BLUP, AMMI, and GGE models to explore GE interactions for adaptability and stability of winter lentils (Lens culinaris Medik.). Plants 12(11):2079. https://doi.org/10.3390/plants12112079
Iniguez-Luy FL, Federico ML (2011) The Genetics of Brassica napus. In: Schmidt R, Bancroft I (eds) Genetics and genomics of the Brassicaceae, vol 9. Springer, New York, pp 291–322. https://doi.org/10.1007/978-1-4419-7118-0-10
Jędzura S, Bocianowski J, Matysik P (2023) The AMMI model application to analyze the genotype–environmental interaction of spring wheat grain yield for the breeding program purposes. Cereal Res Commun 51:197–205. https://doi.org/10.1007/s42976-022-00296-9
Kumar S, Kumari J, Bansai R, Kuri BR, Upadhyay D, Srivastava A, Rana B, Yadav MK, Sengar RS, Singh AK, Singh R (2018) Multi-environmental evaluation of wheat genotypes for drought tolerance. Indian J Genet 78(1):26–35. https://doi.org/10.5958/0975-6906.2018.00004.4
Kumar R, Dhansu P, Kulshreshtha N, Meena MR, Kumaraswamy MH, Appunu C, Chhabra ML, Pandey SK (2023) Identification of salinity tolerant stable sugarcane cultivars using AMMI, GGE and some other stability parameters under multi environments of salinity stress. Sustainability 15(2):1119. https://doi.org/10.3390/su15021119
Leon J, Becker HC (1995) Rapeseed genetics. In: Diepenbrock W, Becker HC (eds) Physiological potentials for yield improvement of annual and protein crops. Advances in Plant Breeding 17. Blackwell Wissenschaftsverlag, Berlin, Germany, pp 53–90
Liersch A, Bocianowski J, Nowosad K, Mikołajczyk K, Spasibionek S, Wielebski F, Matuszczak M, Szała L, Cegielska-Taras T, Sosnowska K, Bartkowiak-Broda I (2020) Effect of genotype x environment interaction for seed traits in winter oilseed rape (Brassica napus L). Agriculture 10:607. https://doi.org/10.3390/agriculture10120607
Long Y, Shi J, Qiu D, Li R, Zhang C, Wang J, Hou J, Zhao J, Shi L, Beom-Seok P, Choi SR, Lim YP, Meng J (2007) Flowering time quantitative trait loci analysis of oilseed Brassica in multiple environments and genom wide alignement with Arabidopsis. Genetics 177:2433–2444. https://doi.org/10.1534/genetics.107.080705
Lou P, Zhao J, Kim JS, Shen S, Del Carpio DP, Song X, Jin M, Vreugdenhil D, Wang X, Koornneef M, Bonnema G (2007) Quantitative trait loci for flowering time and morphological traits in multiple populations of Brassica rapa. J Exp Bot 58(14):4005–4016. https://doi.org/10.1093/jxb/erm255
Łopatyńska A, Bocianowski J, Cyplik A, Wolko J (2021) Multidimensional analysis of diversity in DH lines and hybrid of winter oilseed rape (Brassica napus L.). Agronomy 11:645. https://doi.org/10.3390/agronomy11040645
Ma BL, Yan W, Dwyer LM, Fregeu-Reid J, Voldeng HD, Dion Y, Nass H (2004) Graphic analysis of genotype, environment, nitrogen fertilizater, and their interaction on spring wheat yield. Agron J 96:169–180. https://doi.org/10.2134/agronj2004.1690
Marjanović-Jeromela A, Nagl N, Gvozdanović-Varga J, Hristov N, Kondić-Špika A, Vasić M, Marinković R (2011) Genotype by environment interaction for seed yield per plant in rapeseed using AMMI model. Pesq Agropec Bras 46(2):174–181. https://doi.org/10.1590/S0100-204X2011000200009
Nowosad K, Liersch A, Popławska W, Bocianowski J (2016) Genotype by environment interaction for seed yield in rapeseed (Brassica napus L.) using additive main effects and multiplicative interaction model. Euphytica 208:187–194. https://doi.org/10.1007/s10681-015-1620-z
Nowosad K, Liersch A, Poplawska W, Bocianowski J (2017) Genotype by environment interaction for oil content in winter oilseed rape (Brassica napus L.) using additive main effects and multiplicative interaction model. Indian J Genet Pl Br 77:293–297. https://doi.org/10.5958/0975-6906.2017.00039.6
Oghan HA, Bakhshi B, Rameeh V, Tabrizi HZ, Faraji A, Ghodrati G, Fanaei HR, Askari A, Kiani D, Payghamzadeh K, Sadeghi H, Danaei AK, Kazerani NK, Afrouzi MAAN, Dalili A (2024) Comparative study of univariate and multivariate selection strategies based on an integrated approach applied to oilseed rape breeding. Crop Sci 64:55–73. https://doi.org/10.1002/csc2.21104
Okla MK, Saleem MH, Saleh IA, Zomot N, Perveen S, Parveen A, Abasi F, Ali H, Ali B, Alwasel YA, Abdel-Maksoud MA, Oral MA, Javed S, Ercisli S, Sarfraz MH, Hamed MH (2023) Foliar application of iron-lysine to boost growth attributes, photosynthetic pigments and biochemical defense system in canola (Brassica napus L.) under cadmium stress. BMC Plant Biol 23:648. https://doi.org/10.1186/s12870-023-04672-3
Oroian C, Ugruțan F, Mureșan IC, Oroian I, Odagiu A, Petrescu-Mag IV, Burduhos P (2023) AMMI analysis of Genotype × Environment interaction on sugar beet (Beta vulgaris L.) yield, sugar content and production in Romania. Agronomy 13(10):2549. https://doi.org/10.3390/agronomy13102549
Osborn TC, Kole C, Parkin IAP, Sharpe AG, Kuiper M, Lydiate DJ, Trick M (1997) Comparison of flowering time genes in Brassica rapa, B. Napus and Arabidopsis Thaliana. Genetics 146:1123–1129. https://doi.org/10.1093/genetics/146.3.1123
Pan Q, Xu Y, Li K, Peng Y, Zhan W, Li W, Li L, Yan J (2017) The genetic basis of plant architecture in 10 Maize recombinant inbred line populations. Plant Physiol 175(2):858–873. https://doi.org/10.1104/pp.17.00709
Piepho HP (1994) Best linear unbiased prediction (BLUP) for regional yield trials: a comparison to additive main effects and multiplicative interaction (AMMI) analysis. Theor Appl Genet 89:647–654
Purchase JL, Hatting H, van Deventer CS (2000) Genotype × environment interaction of winter wheat (Triticum aestivum L.) in South Africa: II. Stability analysis of yield performance. S Afr J Plant Soil 17:101–107. https://doi.org/10.1080/02571862.2000.10634878
Qasemi SH, Mostafavi K, Khosroshahli M, Bihamta MR, Ramshini H (2022) Genotype and environment interaction and stability of grain yield and oil content of rapeseed cultivars. Food Sci Nutr 10:4308–4318. https://doi.org/10.1002/fsn3.3023
Quarrie S, Pekic QS, Radosevic R, Kaminska R, Barnes JD, Leverington M, Ceoloni C, Dodig D (2006) Dissecting a wheat QTL for yield present in a range of environments: from QTL to candidate gene. J Exp Bot 57:2627–2637. https://doi.org/10.1093/jxb/erl026
Rahimsoroush H, Nazarian-Forouzabadi F, Chaloshtari MH, Ismaili A, Ebadi AA (2021) Identification of main and epistatis QTLs and QTL through environment interactions for eating and cooking quality in Iranian rice. Euphytica 217:25. https://doi.org/10.1007/s10681-020-02759-8
Raman H, Pragnell R, Eckermann P, Edwards D, Batley J, Coombers N, Taylor B, Wratten N, Luckett D, Dennis L (2011) Genetic dissection of natural variation for flowering time in rapeseed. In Proceedings of the 13th International Rapeseed Congress, Prague, Czech Republic, 05-09.06.2011, CD-ROM, 67–70 Available online: www.irc2011.org
Raman H, Raman R, Qiu Y, Yadav AS, Sureshkumar BL, Rohan M, Wheeler D, Owen O, Menz I, Surehkumar B (2019) GWAS hints at pleiotropic roles for FLOWERING LOCUS T in flowering time and yield-related traits in canola. BMC Genom 20:636. https://doi.org/10.1186/s12864-019-5964-y
Riaz A, Li G, Queresh Z, Swati MS, Quiros CF (2001) Genetic diversity of oilseed Brassica napus inbred lines based on sequence-related amplified polymorphism and its relations to hybrid performance. Plant Breed 120:411–415. https://doi.org/10.1046/j.1439-0523.2001.00636.x
Schiessl S, Iniguez-Luy F, Qian W, Snowdon RJ (2015) Diverse regulatory factors associate with flowering time and yield responses in winter-type Brassica napus. BMC Genom 16:737. https://doi.org/10.1186/s12864-015-1950-1
Schmidt JP, Lamb JA, Schmitt MA, Randall GW, Orf JH, Gollany HT (2001) Soybean varietal response to liquid swine manure application. Agron J 93:358–363. https://doi.org/10.2134/agronj2001.932358x
Seyis F, Friedt W, Lüth W (2006) Yield of Brasscia napus L. hybrids developed using resynthesized rapeseed material sown in different locations. Field Crops Res 96:176–180. https://doi.org/10.1016/j.fcr.2005.06.005
Shi J, Li R, Qiu D, Jiang C, Long Y, Morgan C, Bancroft I, Zhao J, Meng J (2009) Unraveling the complex trait of crop yield with quantitative trait loci mapping in Brassica napus. Genetics 182:581–861. https://doi.org/10.1534/genetics.109.101642
Shi J, Li R, Zou J, Meng J (2011) A dynamic and complex network regulates the heterosis of yield-correlated traits in rapeseed (Brassica napus L.). PLoS ONE 6(7):e21645. https://doi.org/10.1371/journal.pone.0021645
Shojaei SH, Mostafavi K, Ghasemi SH, Bihamta MR, Illés Á, Bojtor C, Nagy J, Harsányi E, Vad A, Széles A, Mousavi SMN (2023) Sustainability on different Canola (Brassica napus L.) cultivars by GGE biplot graphical technique in multi-environment. Sustainability 15(11):8945. https://doi.org/10.3390/su15118945
Singh C, Gupta A, Gupta V, Kumar P, Sendhil R, Tyagi BS, Singh G, Chatrath R, Singh GP (2019) Genotype × environment interaction analysis of multi-environment wheat trials in India using AMMI and GGE biplot models. Crop Breed Appl Biotechnol 19(3):309–318. https://doi.org/10.1590/1984-70332019v19n3a43
Sneller CH, Dombek D (1995) Comparing soybean cultivar ranking and selection for yield with AMMI and full-data performance estimates. Crop Sci 35(6):1536–1541. https://doi.org/10.2135/cropsci1995.0011183X003500060003x
Spasibionek S, Mikołajczyk K, Ćwiek-Kupczyńska H, Piętka T, Krótka K, Matuszczak M, Nowakowska J, Michalski K, Bartkowiak-Broda I (2020) Marker assisted selection of new high oleic and low linolenic winter oilseed rape (Brassica napus L.) inbred lines revealing good agricultural value. PLoS ONE 15(6):e0233959. https://doi.org/10.1371/journal.pone.0233959
Szała L, Kaczmarek Z, Popławska W, Liersch A, Wójtowicz M, Matuszczak M, Biliński ZR, Sosnowska K, Stefanowicz M, Cegielska-Taras T (2019) Estimation of seed yield in oilseed rape to identify the potential of semi-resynthesized parents for the development of new hybrid cultivars. PLoS ONE 14(4):e0215661. https://doi.org/10.1371/journal.pone.0215661
Tadesse T, Sefera G, Tekalign A (2018) Genotype × Environment interaction analysis for Ethiopian mustard (Brassica carinata L.) genotypes using AMMI model. J Plant Breed Crop Sci 10(4):86–92. https://doi.org/10.5897/JPBCS2017.0701
Thian HY, Channa SA, Hu SW (2017) Relationships between genetic distance, combining ability and heterosis in rapeseed (Brassica napus L.). Euphytica 213:1. https://doi.org/10.1007/s10681-016-1788-x
UFOP (2019) Union for the Promotion of Oil and Protein Plants (UFOP). UFOP Report on Global Market Supply 2018/2019. Available online: https://www.ufop.de/
van der Merwe R, Labuschagne MY, Smit A (2024) Cultivar variability and stability of vegetable-type soybean for seed yield and pod shattering. S Afr J Bot 166:106–115. https://doi.org/10.1016/j.sajb.2024.01.034
Vargas W, Crossa J, van Eeuwijk FA, Ramirez E, Sayre K (1999) Using partial least squares regression, factorial regression and AMMI models for interpreting genotype-by-environment interaction. Crop Sci 93:955–967. https://doi.org/10.2135/cropsci1999.0011183X003900040002x
Verma A, Singh GP (2020) Combining AMMI and mean yield of wheat genotypes evaluated under rainfed conditions of Northern Hills zone for stability analysis. Int J Bio-Resour Stress Manag 11(6):590–600. https://doi.org/10.23910/1.2020.2162b
Wang O, Sun G, Ren X, Du B, Cheng Y, Wang Y, Li C, Sun D (2019) Dissecting the genetic basis of grain size and weight in Barley (Hordeum vulgare L.) by QTL and comparative genetic analyses. Front Plant Sci 10:469. https://doi.org/10.3389/fpls.2019.00469
Werner CR, Voss-Fels KP, Miller CN, Qian W, Hua W, Guan CY, Snowdon RJ, Qian L (2018) Effective genomic selection in a narrow-gene pool crop with low-density markers: Asian rapeseed as an example. Plant Genome 11(2):170084. https://doi.org/10.3835/plantgenome2017.09.0084
Wolko J, Dobrzycka A, Bocianowski J, Bartkowiak-Broda I (2019) Estimation of heterosis for yield-related traits for single cross and three-way cross hybrids of oilseed rape (Brassica napus L.). Euphytica 215:156. https://doi.org/10.1007/s10681-019-2482-6
Wójtowicz M (2013) Effect of environmental and agronomical factors on quantity and quality of yield of winter oilseed rape (Brassica napus L.). Dissertation PBAI-NRI, Radzików 45:7–111. (in Polish)
Würschum T, Liu W, Mauer HP, Abel S, Reif JC (2012) Dissecting the genetic architecture of agronomic traits in multiple segregating populations in rapeseed (Brassica napus L.). Theor Appl Genet 124:153–161. https://doi.org/10.1007/s00122-011-1694-5
Yan W (2001) GGE biplot – a windows application for graphical analysis of multienvironment trail data and order types of two-way data. Agron J 93:1111–1118. https://doi.org/10.2134/agronj2001.9351111x
Yang Y, Zhan J, Shi J, Wang X, Liu G, Wang H (2017) Genetic and cytological analyses of the natural variation of seed number per pod in rapeseed (Brassica napus L.). Front Plant Sci 8:1890. https://doi.org/10.3389/fpls.2017.01890
Zhao J, Becker HC, Zhang D, Zhang Y, Ecke W (2005) Oil content in an European × Chinese rapeseed population: QTL with additive and epistatic effects and their genotype-environment interactions. Crop Sci 45:51–59. https://doi.org/10.2135/cropsci2005.0051a
Zhang HP, Berger JD, Milroy P (2013) Genotype × environment interaction studies highlight the role of phenology in specific adaptation of canola (Brassica napus) to contrasting Mediterranean climates. Field Crop Res 144:77–88. https://doi.org/10.1016/j.fcr.2013.01.006
Zulfqar M, Mustafa HSB, Ejaz-Ul-Hasan SS, Qamar R, Gill AN, Mahmood T, Ud D, Ahsan M, Kalyar MTA, Ali S, Hameed A, Salim J, Wakeel A (2021) Quantitative evaluation of commercial canola cultivars through G × E analysis under different agro-climatic conditions. Plant Cell Biotechnol Mol Biol 22(71–72):469–480
Acknowledgements
The authors would like to thank the colleagues at PBAI-NRI: Wiesława Popławska for F1 hybrids and their parental genotypes development and breeding, as well as Katarzyna Śliwińska, Sławomir Hoffa and Jacek Kwiatek for their excellent field work.
Funding
The field trials were funded by the Polish Ministry of Agriculture and Rural Development (https://www.gov.pl/web/rolnictwo); program titled “Biological Progress in Plant Production, 2014–2020”, task no. 48.
Author information
Authors and Affiliations
Contributions
Material preparations were performed by SS, JN, MM, LS, TC-T and KS. Data collection and analysis were performed by AL, JB, KM, SS and FW. JB performed statistical analyses. The first draft of the manuscript was written by AL, JB and IB-B. KM commented on previous versions of the manuscript and all these authors edited and revised the manuscript. B-B, KM and AL supervised the experiment. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
Authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animal performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original version of this article was revised: the Department name in the first affiliation was placed at the incorrect position.
Supplementary Information
Below is the link to the electronic supplementary material.
Fig. S1
Weather conditions during stem elongation, flowering, development of siliques and seed maturing in six environments (E1-E6); dark grey bars, precipitation [mm]; black solid and dashed lines, minimum and maximum temperatures [°C], respectively.
Supplementary file1 (PNG 482 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liersch, A., Bocianowski, J., Spasibionek, S. et al. Evaluation of the stability of quantitative traits of winter oilseed rape (Brassica napus L.) by AMMI analysis. Euphytica 220, 130 (2024). https://doi.org/10.1007/s10681-024-03375-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-024-03375-6