Abstract
Background
The potential effects of insulin therapy on osteoarthritis (OA) risk are poorly understood. This study aimed to explore the causal relationship between insulin therapy and OA.
Methods
Mendelian randomization (MR) analysis was performed to examine the association between genetically proxied inhibition of insulin targets and the risk of overall, hip (HOA) and knee OA (KOA). We then performed univariable MR using summary statistics regarding insulin target genes derived from the DrugBank database. Data related to blood glucose reduction levels were used as a proxy for insulin levels. Two phenotypes, type 2 diabetes, and glycosylated hemoglobin levels, were selected as positive controls to confirm the direction and validity of the proxies. The OA datasets were derived from the UK Biobank cohort. Multivariable MR was adjusted for body mass index, sedentary behavior, cigarette smoking, frequency of alcohol intake, age, and genetic sex.
Results
Genetically proxied insulin therapy was associated with an increased risk of overall OA [odds ratio (OR):1.2595; 95% confidence interval (CI):1.0810–1.4675] and HOA (OR:1.4218; 95%CI:1.1240–1.7985), which remained consistent across multiple MR methods. After adjusting for confounders, we found evidence supporting a significant causal link with a higher risk of overall OA and HOA. A further two-step MR analysis revealed no significant mediation effects from the six mediators in the associations.
Conclusion
There was a causal association between genetically proxied insulin therapy and a higher risk of OA, especially HOA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Osteoarthritis (OA) is a chronic inflammatory joint disease that affects over 300 million individuals worldwide, representing a significant public health burden (GBD Diseases and Injuries Collaborators 2018). Mainstream therapy primarily consists of pain management and, in severe cases, joint replacement (Chen et al. 2012). First-line therapeutic drugs include nonsteroidal anti-inflammatory drugs (NSAIDs) and acetaminophen. However, OA often leads to chronic disability due to pain and impaired joint function, and many patients receiving drug therapy fail to experience significant improvements in physical function and quality of life (Bennell et al. 2012; Towheed et al. 2006).
The pathogenesis of OA is multifactorial, involving mechanical, genetic, and metabolic factors. Among these, the relationships between obesity-related insulin resistance, diabetes mellitus, and OA are critical areas of current research. Endogenous insulin secretion responds to metabolic signals that prompt pancreatic β cells to secrete the precise amount of insulin needed to maintain euglycemia. Insulin therapy for diabetes aims to replicate this complex physiological process (Kramer et al. 2021). Current pharmacotherapies, such as NSAIDs and antioxidants, have shown effectiveness in mitigating the inflammatory effects induced by insulin resistance. These drugs work by inhibiting the activity of Tumor Necrosis Factor (TNF)-α and Interleukin (IL)-6, which are known to play critical roles in the pathogenesis of osteoarthritis, thereby reducing inflammation and preventing further cartilage degradation. For instance, NSAIDs are widely used to manage pain and inflammation in OA by blocking the cyclooxygenase (COX) enzymes, which play a role in the synthesis of pro-inflammatory prostaglandins (Hotamisligil 2006). Moreover, antioxidants such as curcumin have also been shown to possess anti-inflammatory properties that can be beneficial in OA management.
However, insulin therapy has been identified to be associated with potential adverse effects, including articular cartilage degeneration in 82% of OA patients with elevated insulin levels (Ribeiro et al. 2016; Veronese et al. 2019). Insulin has been shown to increase the production of pro-inflammatory cytokines such as TNF-α and IL-6. The chronic low-grade inflammation associated with insulin resistance exacerbates the degradation of joint cartilage (Boden 2011; Kapoor et al. 2011; Scheller et al. 2014). Insulin resistance is linked to increased oxidative stress and the activation of matrix metalloproteinases (MMPs), which contribute to the breakdown of cartilage in osteoarthritis. This process highlights the intersection between metabolic dysfunction and joint degeneration (Ribeiro et al. 2016). Despite this, clinical and in vitro findings on the relationship between insulin and OA risk are inconsistent, with some studies reporting null or protective associations (Cai et al. 2002; Courties and Sellam 2016). Interpreting epidemiological data is challenging due to several factors. Pharmacoepidemiological studies are often affected by residual confounders caused by unmeasured or imprecisely measured variables, including those related to indication (McMahon 2003). Additionally, prior medication use can influence study covariates, resulting in bias due to “survivors” in the early period of pharmacotherapy (Ray 2003). Detection bias, caused by stricter monitoring and clinical tests for patients in the therapeutic group, can also produce upward bias in this group.
Further research leveraging Mendelian randomization (MR) can enhance our understanding of how genetic variants influence the effectiveness of pharmacotherapies. Personalized treatment strategies can be developed by identifying individuals who are genetically predisposed to higher inflammatory responses due to insulin resistance. Therefore, this MR analysis utilizes natural variations in genes encoding drug targets to serve as proxies for targets of insulin therapy, thereby examining the impact of therapy manipulation on disease outcomes (Smith and Ebrahim 2003). These proxies are less likely to be affected by confounders and reverse causation. MR analysis allows for the examination of long-term modulation of drug targets on OA risk, including overall OA, knee OA (KOA), and hip OA (HOA). Drug-target MR mimics the pharmacological modulation of a drug target in clinical trials, making it a useful tool for predicting the potential benefits and adverse effects of clinical therapy interventions (Ference et al. 2015; Plenge et al. 2013; Swerdlow et al. 2015). In this study, we applied an MR approach to investigate the effect of long-term inhibition of insulin therapy drug targets on the risk of overall OA, KOA, and HOA.
Methods
Study design
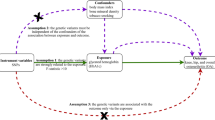
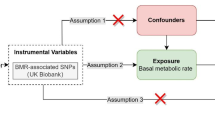
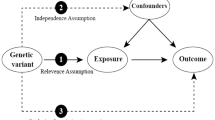
This research was conducted in accordance with the Strengthening the Reporting of Observational Studies in Epidemiology-Mendelian Randomization (STROBE-MR) guidelines. The causal relationships between genetic proxies for insulin targets and OA subtypes were assessed using a drug-target MR approach. Three fundamental assumptions were followed: (1) Relevance: The instrumental variables (IVs) were strongly correlated with exposure; (2) Exclusion Restriction: The relationship between the IVs and the outcome is exclusively mediated by exposure; (3) Independence: IVs are independent of unobserved confounding factors. Genetic variants of target genes linked to blood glucose (BG) reduction were identified and used as proxies for drug-target effects. Two phenotypes, type 2 diabetes and glycosylated hemoglobin levels, were selected as positive controls to confirm the direction and validity of the proxies. The causal impact of genetic proxies on OA and its subtypes was evaluated using various MR techniques. All data utilized in this study were publicly accessible, summarized, and validated by the IEU openGWAS (https://gwas.mrcieu.ac.uk/) and GWAS catalog databases (https://www.ebi.ac.uk/) and did not require additional ethical approval. The detailed research flowchart is presented in Fig. 1.
Analysis Process and Verification of Key MR Assumptions Flowchart. We utilize different colors to represent the relationship between the analysis methods and the three core assumptions of MR. Yellow indicates the relevance assumption and the methods used to verify it; green represents the independence assumption and the methods used to verify it; blue denotes the exclusion restriction assumption and the methods used for its verification. UVMR univariable Mendelian randomization, MVMR multi-variable Mendelian randomization, BG blood glucose levels, T2DM type 2 diabetes, HbA1C glycated hemoglobin levels, OA osteoarthritis, KOA knee osteoarthritis, HOA hip osteoarthritis, ACL frequency of alcohol intake, AGE age at recruitment, BMI body mass index, SBs sedentary behaviors, SEX genetic sex, SMO cigarette smoking
Data sources and instrumental variable selection
The largest datasets of OA and its subtypes from the openGWAS database, derived from the UK Biobank cohort (Tachmazidou et al. 2019; Yengo et al. 2018), were used as outcomes to enhance statistical power. OA diagnosis was based on clinical evidence of disease requiring joint replacement or radiographic evidence of the disease (Kellgren-Lawrence grade ≥ 2). A total of 417,596 participants, comprising 39,427 patients and 378,169 controls, were included in the OA analysis. The KOA cohort included 403,124 participants, with 24,955 cases and 378,169 controls, and the HOA cohort included 393,873 participants, with 15,704 cases and 378,169 controls, respectively.
T2DM data included 212,351 participants, 29,166 cases, and 183,185 controls from the FinnGen Biobank Analysis Consortium database (https://finngen.gitbook.io/documentation/). Diagnosis was based on the International Classification of Diseases (ICD-10) criteria. The HbA1c dataset was obtained from a European cohort in a transancestral GWAS study (J. Chen et al. 2021), comprising 146,806 participants.
Considering the complex clinical and genetic background and pathogenesis of OA, we further used multivariable MR (MVMR) and two-step mediation MR analyses to validate the Independence Assumption. Since we used BG as a proxy to obtain IVs, six potential confounders known to be associated with both OA and blood glucose levels (Cao et al. 2023; Leyland et al. 2021; Szilagyi et al. 2022) were incorporated into a MVMR and mediation MR model to adjust for mediation effects. These included body mass index (BMI; sample of 681,275 participants) (Yengo et al. 2018), sedentary behaviors (SBs, 437,887 participants) (van de Vegte et al. 2020), cigarette smoking (SMO, 249,752 participants) (Liu et al. 2019), frequency of alcohol intake (ACL, 462,346 participants), age at recruitment (AGE, 361,194 participants), and genetic sex (SEX, 361,194 participants). ACL, AGE, and SEX data were sourced from the UK Biobank. Data for all mediators were collated from different databases with those used for outcome to avoid sample overlap impacts.
Information regarding target genes with defined pharmacological actions was acquired from the DrugBank database (Wishart et al. 2018). Data related to BG levels were used as proxies for insulin effects, with a sample size of 200,622. SNPs located in or near (± 200 kb) the corresponding target genes, which were associated with BG at a significance level of p < 1 × 10−4, were selected as proxies for insulin as described previously (Huang et al. 2021). F-statistics were calculated for each IVs and only those with F > 10 were included to minimize weak instrumental bias (Shim et al. 2015). SNPs were clumped according to a liberal linkage disequilibrium (LD) threshold of r2 < 0.1, using the European reference panel from the 1000 Genomes Project. MR Steiger filtering was used to exclude any IVs accounting for lower variance in the outcome than in the exposure and to determine a causal direction for a given SNP between exposure and outcome by comparing the proportion of variance explained in each (Hemani et al. 2017). Data were cross-matched with the phenome-wide association studies database (pheWAS) using a threshold of p < 5 × 10^-6 to mitigate potential links between SNPs and confounding factors. All estimates are presented per one standard deviation (SD) unit increase, and effect sizes are presented as odds ratios (OR) with a 95% confidence interval (CI). All study participants were of European ancestry to control for potential mixed population effects.
Statistical approach
R software (version 4.1.3) with R packages, Two-Sample MR (version 0.5.6)(Hemani et al. 2018), MRPRESSO (version 1.0)(Verbanck et al. 2018), MVMR (version 0.3.0)(Sanderson et al. 2021), and MendelianRandomization (version 0.5.1)(Sanderson et al. 2021; Yavorska and Burgess 2017) were used. When an outcome association SNP was missing in the harmonization process of the IVs, a proxy with r2 ≥ 0.8 was used, and SNPs without proxies were removed. Six statistical methods were employed to assess the causal effects of insulin and OA in univariable MR (UVMR). The primary method was inverse-variance weighted (IVW)(Slob and Burgess 2020), which combines Wald estimates of causality for each IV to provide a relatively accurate assessment with a precise confidence interval, even under the assumption of invalid genetic instruments (Burgess et al. 2013; Staley and Burgess 2017). The MR-Egger method quantifies pleiotropy across IVs using the slope and intercept of the MR-Egger regression and offers an adjusted, robust estimate independent of IV validity (Bowden et al. 2015; Burgess and Thompson 2017). The MR-PRESSO method identifies and adjusts for distorted outliers that contribute to significant pleiotropy and heterogeneity, providing a corrected causal-effect estimate (Verbanck et al. 2018). The weighted-median method yields consistent, valid inferences when over 50% of the instrumental variables are valid (Bowden et al. 2016). Bayesian Weighted Mendelian Randomization (BWMR) corrects for pleiotropy violations and polygenic weak effect uncertainties within a Bayesian weighting framework to give reliable causal inferences (Zhao et al. 2020). The MR-Robust Adjusted Profile Score (MRAPS) method increases the statistical power to provide robust estimates under conditions of significant weak instrumental bias and horizontal pleiotropy (Zhao et al. 2018). Bonferroni correction was applied with a corrected p < 0.0083 considered significant in UVMR to prevent multiple comparison errors. Four methods, IVW, MR-Egger, MR-PRESSO, and MRAPS, were used to analyze the positive control group, and an independent Bonferroni corrected significance threshold of p < 0.0125 was adopted.
Multivariable random-effects IVW and MR-Egger methods were employed within the MVMR model, and all six mediators were concurrently adjusted to derive a consolidated, corrected evaluation. Two-step mediation MR analysis, based on different methods, quantifies the mediation effects and mechanisms for each mediator (Sanderson 2021). The total effect (beta0) denotes the direct causal impact of exposure on the outcome and is derived using the IVW method employed in UVMR. The step1 effect (beta1) indicates the causal effect value of the exposure on the mediator, and the step2 effect (beta2) represents the causal effect value of the mediator on the outcome. The association between exposure and outcome was deemed to be fully mediated by a given mediator if beta0 was non-significant, whereas both beta1 and beta2 were significant. Conversely, significant values for beta0, beta1, and beta2 indicate that the exposure-to-outcome association was partially mediated by a factor. Mediators were deemed not to influence the exposure-to-outcome association if beta0 was significant, whereas either beta1 or beta2 were non-significant. Substantial mediation effects were analyzed to identify the direct or indirect nature of the effect and calculate the proportion mediated (Burgess et al. 2017). A p-value less than 0.05 was considered statistically significant in the MVMR and mediation MR analysis.
Sensitivity analysis
Conditional F-statistics were utilized to evaluate instrument strength in the MVMR analysis, testing the robustness of each SNP's prediction of exposure conditional on other exposures. A conditional F-value of less than 10 signifies a substantial risk of weak instrument bias.
Heterogeneity arising from the invalidity of the IVs was measured using Cochran’s Q-statistic. A p-value of less than 0.05 was deemed indicative of significant heterogeneity, and a random-effect IVW model was used if heterogeneity could not be rectified using the MR-PRESSO method (Greco et al. 2015). The MR-Egger and MR-PRESSO methods were used to test the violation of the second IV assumption, prompted by direction pleiotropy, and Cochran's Q-statistics to assess heterogeneity. Statistical power was evaluated using the binary-outcome model from the mRnd tools (https://shiny.cnsgenomics.com/mRnd/). A power below 80% indicated insufficient statistical power, and MRAPS results were prioritized. Leave-one-out analysis was conducted to identify unstable SNPs with an individual disproportionate influence on the results under the Bonferroni-corrected threshold. Such SNPs were omitted, and the results were reassessed accordingly (Burgess and Thompson 2017). Colocalization analysis using the 'coloc' R package was performed to evaluate potential violations of the exclusion restriction assumption. The likelihood of traits sharing causal variants was assessed to infer potential causality between traits that did not account for horizontal pleiotropy or the direction of the association. Thus, the probability of shared causal variants across SNP-drug target analyses was evaluated. The 'coloc' package utilizes an approximation of Bayes factor computations to produce posterior probabilities for different configurations of trait associations within a specified region. Colocalization analysis was performed within a ± 300 kb window of the top instrumental SNP for each identified drug target. Because the number of proxy SNPs was limited, reducing the statistical power, a more lenient threshold of a posterior probability greater than 0.6 was used to increase sensitivity and indicate substantial support for a specific configuration.
Results
Genetic instruments and validation
Six pharmacological target genes—LRP2, IGFBP7, CPE, NOV, IGF1R, and INSR—were identified in the DrugBank database. Proxy IVs were found for three loci: 134 for LRP2, 1 for IGFBP7, and 66 for IGF1R. However, no suitable SNPs within or in the vicinity of CPE, NOV, or INSR were found (S.Table 1). The positive control group was used as the reference set to align and screen proxies. Sixteen IVs with F-statistics greater than 10 were selected, suggesting a minimal risk of weak instrument bias. MR Steiger filtering indicated that all IVs maintained the correct causal direction from exposure to outcome (S.Table 2–3).
Only LRP2 and IGF1R had sufficient proxy SNPs to facilitate additional colocalization analyses. The probability of shared IVs between the exposure and outcome was low (PP.H4: 0.098–0.258). Conversely, IVs were likely exclusively linked to exposure (PP.H1: 0.738–0.799). The integrity of the exclusion restriction assumption was maintained, verifying its non-violation. Details of the sensitivity analysis are presented in Table 1. No significant heterogeneity or pleiotropy was found between the insulin genetic proxies and positive controls or OA. All outcomes showed strong statistical power, ranging from 98 to 100%. Only one IV, rs508506, had a disproportionate influence over the results in the leave-one-out test of insulin and overall OA (S.Fig. 1). Scatter plots of UVMR analysis of the effects of insulin and OA are shown in S.Fig. 2.
The SNPs used for validation analyses of the positive controls are listed in S.Table 4. Positive control analyses showed that genetic proxies of insulin had a pronounced protective effect on both T2DM (ORIVW: 0.6911; 95% CI: 0.5499, 0.8687; p = 1.53E−03) and HbA1c (ORIVW: 0.6531; 95% CI: 0.6318, 0.6751; p = 6.32E−140). These outcomes are consistent with findings from clinical trials, validating the instrumental SNPs, as depicted in Fig. 2 and S. Table 5.
Drug-target UVMR analysis
The accuracy of the MR-Egger method is compromised by the restricted number of SNPs, resulting in broader confidence intervals. Hence, IVW results were primarily used for UVMR, with reference to the MR-Egger findings only when considerable heterogeneity was detected. The genetic proxies for insulin showed a significant causal association with overall OA risk (ORIVW: 1.2595; 95% CI: 1.0810–1.4675; p = 3.08E−03), consistent with estimates from the weighted median (OR: 1.3576; 95% CI: 1.1014–1.6735; p = 4.17E−03), BWMR (OR: 1.2626; 95% CI: 1.0818–1.4735; p = 3.09E−03), MR-PRESSO (OR: 1.2595; 95% CI: 1.0961–1.4473; p = 5.33E−03), and MR.RAPS (OR: 1.2647; 95% CI: 1.0793–1.4818; p = 3.68E−03). A causal link was also detected between the genetic proxies of insulin and HOA (ORIVW: 1.4218; 95% CI: 1.1240–1.7985; p = 3.34E−03), which remained consistent across many MR methods. Estimates were 1.5214 (95% CI: 1.1137, 2.0785; p = 8.28E−03) by weighted median, 1.4256 (95% CI: 1.1245, 1.8072; p = 3.39E−03) by BWMR, 1.4218 (95% CI:1.2060- 1.6761; p = 7.88E−04) by MR-PRESSO, and 1.4238 (95% CI: 1.1156, 1.8172; p = 4.52E−03) by MR.RAPS. However, no significant association was found between genetic proxies for insulin and KOA (ORIVW: 1.1234; 95% CI: 0.9317, 1.3547; p = 2.29E−01). A forest plot of the UVMR is presented in Fig. 3.
Drug-target MVMR and Mediation MR analysis
The six potential confounders were adjusted in the multivariable IVW and MR-Egger models. Weak instrumental strength was reported for the genetic proxies for insulin, with conditional F-statistics = 2.7430, likely due to the disproportionate number of IVs relative to other mediators.
Genetic proxies for insulin maintained a significant causal link with a higher risk of overall OA (ORMVMR-IVW: 1.5872; 95% CI: 1.1600, 2.1719; p = 0.004) and HOA (ORMVMR-IVW: 1.6438; 95% CI: 1.0169, 2.6570; p = 0.043) but not with KOA (ORMVMR-IVW: 1.4579; 95% CI: 0.9948, 2.1366; p = 0.0540), aligning with the UVMR findings (Fig. 4 and S.Table 6). No significant mediation effects from the six mediators of the associations between genetic proxies for insulin and overall OA or HOA were detected during two-step MR analyses (Table 2), implying that the causal relationship was predominantly due to direct effects.
Forest plot of the multivariable Mendelian randomization analyses exploring genetically determined association of the effect of insulin with osteoarthiritis and its subtypes adjusted for confounding traits (alcohol intake, cigarettes consumption, age at recruitment, genetic sex, sedentary behaviors, and body mass index). OR odds ratio, CIs confidence intervals, OA osteoarthritis, KOA knee osteoarthritis, HOA hip osteoarthritis
Discussion
In our MR analysis of up to 39,427 OA cases and 378,169 controls, genetically proxied insulin therapy was associated with an increased risk of overall OA and HOA. This association remained significant even after adjusting for age, sex, BMI, alcohol consumption, and smoking status.
Previous studies have not directly observed this association between insulin therapy and OA risk. For example, Konstari et al. (Konstari et al. 2021) did not find that elevated plasma fasting glucose levels or diagnosed diabetes without insulin therapy predicted a higher incidence of knee OA; instead, elevated fasting plasma glucose levels were related to a reduced risk of incident knee OA. Additionally, males with diabetes, especially those requiring insulin, reported more severe joint pain and higher analgesic use compared to males without diabetes or those using other diabetes medications (Szilagyi et al. 2022). No previous study has reported an association between insulin use and HOA.
The mechanism underlying the association between genetically proxied insulin and OA risk remains unclear, and the role of insulin in OA is controversial. High insulin levels are associated with insulin resistance and T2DM (Xu et al. 2023). Distinguishing between the effects of insulin per se and insulin resistance is challenging. Excess insulin in T2DM patients may damage cartilage, and various complications are thought to result from high insulin levels in obesity and metabolic syndrome. Insulin is known to prolong chondrocyte survival, stimulate chondrocyte proliferation, and suppress differentiation, which could impair cartilage formation by inhibiting chondrocyte maturation (Alarid et al. 1992; Torres et al. 2003). Moreover, hyperinsulinemia reduces circulating T4 and its conversion to T3, mimicking hypothyroidism (Crunkhorn and Patti 2008; Farasat et al. 2012; Ortiz-Caro et al. 1984; Roos et al. 2007), and reduces the signal from thyroid hormones necessary for chondrocyte maturation, resulting in a predisposition to OA (Williams 2013).
Our study found that genetic predisposition to insulin therapy increases OA risk independently of age, sex, BMI, alcohol consumption, and smoking status. This suggests that insulin may have a direct genetic association with OA, rather than a mediated link through factors such as obesity, which predisposes individuals to OA. The inflammatory mechanisms potentially mediating this relationship may involve the upregulation of pro-inflammatory cytokines and the alteration of cartilage metabolism under chronic hyperinsulinemic conditions. Chronic hyperinsulinemia can exacerbate inflammation by promoting the production of pro-inflammatory cytokines such as IL-6 and TNF-α, which have been shown to contribute to cartilage degradation and increased synovial inflammation, important components in the pathophysiology of OA (Goldring and Otero 2011). Furthermore, insulin's impact on cartilage cells could involve adverse effects on chondrocyte proliferation and extracellular matrix production, leading to compromised joint integrity (Y. Chen et al. 2018). These insights support a more nuanced understanding of insulin's role in OA, suggesting a potential causal link between insulin therapy and increased OA risk, mediated through both metabolic and inflammatory pathways. This underscores the importance of considering the inflammatory side effects of insulin therapy in OA management and suggests avenues for therapeutic intervention that may involve modulating insulin levels or action.
There are several strengths to our MR study. First, the inclusion of participants from various European ancestries minimizes potential bias from population stratification. Additionally, various sensitivity analyses were employed to identify violations of MR assumptions. The use of summarized-level MR methods allowed us to leverage large-scale genetic information from the GWAS dataset, improving the power of the test and the reliability of causal inference. No heterogeneity was detected within the IVs in the MR analysis, indicating an absence of bias due to pleiotropy. Moreover, germline IVs were used as proxies for insulin therapy targets, facilitating the evaluation of long-term inhibition, which is a more realistic analogy of the typically decades-long use of insulin therapy compared to the short-term effects assessed in conventional observational studies and randomized trials.
Our study also has several limitations. First, only the on-target effects of insulin therapy were predicted owing to the inclusion of well-documented protein targets. Off-target drug effects not exerted by these proteins were not captured by the MR models. Second, the genetic prediction of drug effects might be different from those found in clinical therapy. An exposure instrumented by genetic variants was considered to be present from birth and last throughout the lifetime. Thus, the analyses only assessed the modulating effects of drug target proteins over the long term. Moreover, considering the lifelong genetic effects, the effects of exposure to insulin during a specific lifetime period cannot be accommodated. Third, no key gene (h4 > 0.60 was found in the colocalization analysis, suggesting that insulin-induced high OA risk was not caused by a single gene, but by the combined effects of multiple genes. The limited number of proxy SNPs meant that only LRP2 and IGF1R were included in further colocalized analysis, and CPE, NOV, INSR, and IGFBP7 with ≤ 1 SNPs were excluded. This observation may also contribute to the failure to identify key genes. The size of the BG dataset that matched the proxy SNPs was a limiting factor, and further verification would be enabled by the availability of larger and higher-quality BG datasets. Finally, the genetic data were restricted to those of European ancestry and may not be generalizable to those of different heritages.
Conclusion
Our MR analyses suggest that genetically proxied long-term insulin therapy is associated with an increased risk of OA, particularly HOA. Evaluating insulin use in randomized controlled trials with long-term follow-up data is essential to determine the long-term safety of insulin therapy. Future studies should aim to clarify the mechanistic pathways underlying the association between insulin therapy and OA. This study also exemplifies the use of MR design to provide insights into the indications or potential contraindications for approved drugs.
Availability of data and materials
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Alarid ET, Schlechter NL, Russell SM, Nicoll CS (1992) Evidence suggesting that insulin-like growth factor-I is necessary for the trophic effects of insulin on cartilage growth in vivo. Endocrinology 130(4):2305–2309. https://doi.org/10.1210/endo.130.4.1547741
Bennell KL, Hunter DJ, Hinman RS (2012) Management of osteoarthritis of the knee. BMJ 345:e4934. https://doi.org/10.1136/bmj.e4934
Boden G (2011) Obesity, insulin resistance and free fatty acids. Curr Opin Endocrinol Diabetes Obes 18(2):139–143. https://doi.org/10.1097/MED.0b013e3283444b09
Bowden J, Davey Smith G, Burgess S (2015) Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol 44(2):512–525. https://doi.org/10.1093/ije/dyv080
Bowden J, Davey Smith G, Haycock PC, Burgess S (2016) Consistent estimation in mendelian randomization with some invalid instruments using a weighted median estimator. Genet Epidemiol 40(4):304–314. https://doi.org/10.1002/gepi.21965
Burgess S, Thompson SG (2017) Interpreting findings from Mendelian randomization using the MR-Egger method. Eur J Epidemiol 32(5):377–389. https://doi.org/10.1007/s10654-017-0255-x
Burgess S, Butterworth A, Thompson SG (2013) Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol 37(7):658–665. https://doi.org/10.1002/gepi.21758
Burgess S, Thompson DJ, Rees JMB, Day FR, Perry JR, Ong KK (2017) Dissecting causal pathways using Mendelian randomization with summarized genetic data: application to age at menarche and risk of breast cancer. Genetics 207(2):481–487. https://doi.org/10.1534/genetics.117.300191
Cai L, Okumu FW, Cleland JL, Beresini M, Hogue D, Lin Z, Filvaroff EH (2002) A slow release formulation of insulin as a treatment for osteoarthritis. Osteoarthritis Cartilage 10(9):692–706. https://doi.org/10.1053/joca.2002.0813
Cao Z, Li Q, Li Y, Wu J (2023) Causal association of leisure sedentary behavior with arthritis: a Mendelian randomization analysis. Semin Arthritis Rheum 59:152171. https://doi.org/10.1016/j.semarthrit.2023.152171
Chen A, Gupte C, Akhtar K, Smith P, Cobb J (2012) The global economic cost of osteoarthritis: how the UK compares. Arthritis 2012:698709. https://doi.org/10.1155/2012/698709
Chen Y, Hu Y, Yu YE, Zhang X, Watts T, Zhou B et al (2018) Subchondral trabecular rod loss and plate thickening in the development of osteoarthritis. J Bone Miner Res 33(2):316–327. https://doi.org/10.1002/jbmr.3313
Chen J, Spracklen CN, Marenne G, Varshney A, Corbin LJ, Luan J et al (2021) The trans-ancestral genomic architecture of glycemic traits. Nat Genet 53(6):840–860. https://doi.org/10.1038/s41588-021-00852-9
Courties A, Sellam J (2016) Osteoarthritis and type 2 diabetes mellitus: what are the links? Diabetes Res Clin Pract 122:198–206. https://doi.org/10.1016/j.diabres.2016.10.021
Crunkhorn S, Patti ME (2008) Links between thyroid hormone action, oxidative metabolism, and diabetes risk? Thyroid 18(2):227–237. https://doi.org/10.1089/thy.2007.0249
Farasat T, Cheema AM, Khan MN (2012) Hyperinsulinemia and insulin resistance is associated with low T3/T4 ratio in pre diabetic euthyroid Pakistani subjects. J Diabetes Complications 26(6):522–525. https://doi.org/10.1016/j.jdiacomp.2012.05.017
Ference BA, Majeed F, Penumetcha R, Flack JM, Brook RD (2015) Effect of naturally random allocation to lower low-density lipoprotein cholesterol on the risk of coronary heart disease mediated by polymorphisms in NPC1L1, HMGCR, or both: a 2 × 2 factorial Mendelian randomization study. J Am Coll Cardiol 65(15):1552–1561. https://doi.org/10.1016/j.jacc.2015.02.020
GBD Diseases and Injuries Collaborators (2018) Global, regional, and national incidence, prevalence, and years lived with disability for 354 diseases and injuries for 195 countries and territories, 1990–2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet 392(10159):1789–1858. https://doi.org/10.1016/s0140-6736(18)32279-7
Goldring MB, Otero M (2011) Inflammation in osteoarthritis. Curr Opin Rheumatol 23(5):471–478. https://doi.org/10.1097/BOR.0b013e328349c2b1
Greco MF, Minelli C, Sheehan NA, Thompson JR (2015) Detecting pleiotropy in Mendelian randomisation studies with summary data and a continuous outcome. Stat Med 34(21):2926–2940. https://doi.org/10.1002/sim.6522
Hemani G, Tilling K, Davey Smith G (2017) Orienting the causal relationship between imprecisely measured traits using GWAS summary data. PLoS Genet 13(11):e1007081. https://doi.org/10.1371/journal.pgen.1007081
Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D et al (2018) The MR-Base platform supports systematic causal inference across the human phenome. Elife. https://doi.org/10.7554/eLife.34408
Hotamisligil GS (2006) Inflammation and metabolic disorders. Nature 444(7121):860–867. https://doi.org/10.1038/nature05485
Huang W, Xiao J, Ji J, Chen L (2021) Association of lipid-lowering drugs with COVID-19 outcomes from a Mendelian randomization study. Elife. https://doi.org/10.7554/eLife.73873
Kapoor M, Martel-Pelletier J, Lajeunesse D, Pelletier JP, Fahmi H (2011) Role of proinflammatory cytokines in the pathophysiology of osteoarthritis. Nat Rev Rheumatol 7(1):33–42. https://doi.org/10.1038/nrrheum.2010.196
Konstari S, Sääksjärvi K, Heliövaara M, Rissanen H, Knekt P, Arokoski JPA, Karppinen J (2021) Associations of metabolic syndrome and its components with the risk of incident knee osteoarthritis leading to hospitalization: A 32-year follow-up study. Cartilage 13(1_suppl):1445–1456. https://doi.org/10.1177/1947603519894731
Kramer CK, Retnakaran R, Zinman B (2021) Insulin and insulin analogs as antidiabetic therapy: a perspective from clinical trials. Cell Metab 33(4):740–747. https://doi.org/10.1016/j.cmet.2021.03.014
Leyland KM, Gates LS, Sanchez-Santos MT, Nevitt MC, Felson D, Jones G et al (2021) Knee osteoarthritis and time-to all-cause mortality in six community-based cohorts: an international meta-analysis of individual participant-level data. Aging Clin Exp Res 33(3):529–545. https://doi.org/10.1007/s40520-020-01762-2
Liu M, Jiang Y, Wedow R, Li Y, Brazel DM, Chen F et al (2019) Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat Genet 51(2):237–244. https://doi.org/10.1038/s41588-018-0307-5
McMahon AD (2003) Approaches to combat with confounding by indication in observational studies of intended drug effects. Pharmacoepidemiol Drug Saf 12(7):551–558. https://doi.org/10.1002/pds.883
Ortiz-Caro J, Obregón MJ, Pascual A, Jolin T (1984) Decreased T4 to T3 conversion in tissues of streptozotocin-diabetic rats. Acta Endocrinol (copenh) 106(1):86–91. https://doi.org/10.1530/acta.0.1060086
Plenge RM, Scolnick EM, Altshuler D (2013) Validating therapeutic targets through human genetics. Nat Rev Drug Discov 12(8):581–594. https://doi.org/10.1038/nrd4051
Ray WA (2003) Evaluating medication effects outside of clinical trials: new-user designs. Am J Epidemiol 158(9):915–920. https://doi.org/10.1093/aje/kwg231
Ribeiro M, López de Figueroa P, Blanco FJ, Mendes AF, Caramés B (2016) Insulin decreases autophagy and leads to cartilage degradation. Osteoarthritis Cartilage 24(4):731–739. https://doi.org/10.1016/j.joca.2015.10.017
Roos A, Bakker SJ, Links TP, Gans RO, Wolffenbuttel BH (2007) Thyroid function is associated with components of the metabolic syndrome in euthyroid subjects. J Clin Endocrinol Metab 92(2):491–496. https://doi.org/10.1210/jc.2006-1718
Sanderson E (2021) Multivariable Mendelian randomization and mediation. Cold Spring Harb Perspect Med 11:2. https://doi.org/10.1101/cshperspect.a038984
Sanderson E, Spiller W, Bowden J (2021) Testing and correcting for weak and pleiotropic instruments in two-sample multivariable Mendelian randomization. Stat Med 40(25):5434–5452. https://doi.org/10.1002/sim.9133
Scheller J, Garbers C, Rose-John S (2014) Interleukin-6: from basic biology to selective blockade of pro-inflammatory activities. Semin Immunol 26(1):2–12. https://doi.org/10.1016/j.smim.2013.11.002
Shim H, Chasman DI, Smith JD, Mora S, Ridker PM, Nickerson DA et al (2015) A multivariate genome-wide association analysis of 10 LDL subfractions, and their response to statin treatment, in 1868 Caucasians. PLoS One 10(4):e0120758. https://doi.org/10.1371/journal.pone.0120758
Slob EAW, Burgess S (2020) A comparison of robust Mendelian randomization methods using summary data. Genet Epidemiol 44(4):313–329. https://doi.org/10.1002/gepi.22295
Smith GD, Ebrahim S (2003) “Mendelian randomization”: can genetic epidemiology contribute to understanding environmental determinants of disease? Int J Epidemiol 32(1):1–22. https://doi.org/10.1093/ije/dyg070
Staley JR, Burgess S (2017) Semiparametric methods for estimation of a nonlinear exposure-outcome relationship using instrumental variables with application to Mendelian randomization. Genet Epidemiol 41(4):341–352. https://doi.org/10.1002/gepi.22041
Swerdlow DI, Preiss D, Kuchenbaecker KB, Holmes MV, Engmann JE, Shah T et al (2015) HMG-coenzyme A reductase inhibition, type 2 diabetes, and bodyweight: evidence from genetic analysis and randomised trials. Lancet 385(9965):351–361. https://doi.org/10.1016/s0140-6736(14)61183-1
Szilagyi IA, Waarsing JH, Schiphof D, van Meurs JBJ, Bierma-Zeinstra SMA (2022) Towards sex-specific osteoarthritis risk models: evaluation of risk factors for knee osteoarthritis in males and females. Rheumatology (oxford) 61(2):648–657. https://doi.org/10.1093/rheumatology/keab378
Tachmazidou I, Hatzikotoulas K, Southam L, Esparza-Gordillo J, Haberland V, Zheng J et al (2019) Identification of new therapeutic targets for osteoarthritis through genome-wide analyses of UK Biobank data. Nat Genet 51(2):230–236. https://doi.org/10.1038/s41588-018-0327-1
Torres ES, Andrade CV, Fonseca EC, Mello MA, Duarte ME (2003) Insulin impairs the maturation of chondrocytes in vitro. Braz J Med Biol Res 36(9):1185–1192. https://doi.org/10.1590/s0100-879x2003000900007
Towheed TE, Maxwell L, Judd MG, Catton M, Hochberg MC, Wells G (2006) Acetaminophen for osteoarthritis. Cochrane Database Syst Rev 2006(1):CD004257. https://doi.org/10.1002/14651858.CD004257.pub2
van de Vegte YJ, Said MA, Rienstra M, van der Harst P, Verweij N (2020) Genome-wide association studies and Mendelian randomization analyses for leisure sedentary behaviours. Nat Commun 11(1):1770. https://doi.org/10.1038/s41467-020-15553-w
Verbanck M, Chen CY, Neale B, Do R (2018) Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet 50(5):693–698. https://doi.org/10.1038/s41588-018-0099-7
Veronese N, Cooper C, Reginster JY, Hochberg M, Branco J, Bruyère O et al (2019) Type 2 diabetes mellitus and osteoarthritis. Semin Arthritis Rheum 49(1):9–19. https://doi.org/10.1016/j.semarthrit.2019.01.005
Williams GR (2013) Thyroid hormone actions in cartilage and bone. Eur Thyroid J 2(1):3–13. https://doi.org/10.1159/000345548
Wishart DS, Feunang YD, Guo AC, Lo EJ, Marcu A, Grant JR et al (2018) DrugBank 5.0: a major update to the DrugBank database for 2018. Nucleic Acids Res 46(D1):D1074-d1082. https://doi.org/10.1093/nar/gkx1037
Xu R, Wang K, Yao Z, Zhang Y, Jin L, Pang J et al (2023) BRSK2 in pancreatic β cells promotes hyperinsulinemia-coupled insulin resistance and its genetic variants are associated with human type 2 diabetes. J Mol Cell Biol. https://doi.org/10.1093/jmcb/mjad033
Yavorska OO, Burgess S (2017) MendelianRandomization: an R package for performing Mendelian randomization analyses using summarized data. Int J Epidemiol 46(6):1734–1739. https://doi.org/10.1093/ije/dyx034
Yengo L, Sidorenko J, Kemper KE, Zheng Z, Wood AR, Weedon MN et al (2018) Meta-analysis of genome-wide association studies for height and body mass index in ∼700000 individuals of European ancestry. Hum Mol Genet 27(20):3641–3649. https://doi.org/10.1093/hmg/ddy271
Zhao J, Ming J, Hu X, Chen G, Liu J, Yang C (2020) Bayesian weighted Mendelian randomization for causal inference based on summary statistics. Bioinformatics 36(5):1501–1508. https://doi.org/10.1093/bioinformatics/btz749
Zhao Q, Chen Y, Wang J, Small DS (2018) Powerful genome-wide design and robust statistical inference in two-sample summary-data Mendelian randomization
Acknowledgements
The authors express their gratitude to EditSprings (https://www.editsprings.com/) for providing expert linguistic services. This manuscript is not under review by any other journal and has not been previously published. Ethical approval: Ethics approval was not required, as this study collected primary data from human participants or animals.
Funding
This study was supported by the National Key R&D Program of China (No. 2019YFA0112100). The study funders/sponsors had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the manuscript; or decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
Conceptualization, Yajia Li and Jianhuang Wu; methodology, Ziqin Cao and Yajia Li; software, Qiangxiang Li, Ziqin Cao, and Yajia Li; formal analysis, Ziqin Cao and Yajia Li; data curation, Qiangxiang Li and Jianhuang Wu; investigation, Ziqin Cao and Yajia Li; writing—original draft preparation, Ziqin Cao and Yajia Li; writing—review and editing, all authors. All the authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they do not have any competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cao, Z., Li, Q., Wu, J. et al. Genetic proxies for therapy of insulin drug targets and risk of osteoarthritis: a drug-target Mendelian randomization analysis. Inflammopharmacol (2024). https://doi.org/10.1007/s10787-024-01542-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10787-024-01542-8