Abstract
Aromatic side chains are attractive probes of protein dynamics on the millisecond time scale, because they are often key residues in enzyme active sites and protein binding sites. Further they allow to study specific processes, like histidine tautomerization and ring flips. Till now such processes have been studied by aromatic 13C CPMG relaxation dispersion experiments. Here we investigate the possibility of aromatic 1H CPMG relaxation dispersion experiments as a complementary method. Artifact-free dispersions are possible on uniformly 1H and 13C labeled samples for histidine δ2 and ε1, as well as for tryptophan δ1. The method has been validated by measuring fast folding–unfolding kinetics of the small protein CspB under native conditions. The determined rate constants and populations agree well with previous results from 13C CPMG relaxation dispersion experiments. The CPMG-derived chemical shift differences between the folded and unfolded states are in good agreement with those obtained directly from the spectra. In contrast, the 1H relaxation dispersion profiles in phenylalanine, tyrosine and the six-ring moiety of tryptophan, display anomalous behavior caused by 3J 1H–1H couplings and, if present, strong 13C–13C couplings. Therefore they require site-selective 1H/2H and, in case of strong couplings, 13C/12C labeling. In summary, aromatic 1H CPMG relaxation dispersion experiments work on certain positions (His δ2, His ε1 and Trp δ1) in uniformly labeled samples, while other positions require site-selective isotope labeling.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Proteins are dynamic entities that continuously undergo dynamic processes on various time scales. Especially conformational transitions on the millisecond time scale are often linked to biological function (Mittermaier and Kay 2009) and transiently populated high-energy states play important roles in enzyme catalysis (Boehr et al. 2006; Cole and Loria 2002; Eisenmesser et al. 2002) or ligand binding (Demers and Mittermaier 2009; Malmendal et al. 1999). Such transitions between different conformations generally lead to a modulation of NMR parameters as the chemical shift (Gutowsky and Saika 1953) or residual dipolar couplings (Igumenova et al. 2007; Vallurupalli et al. 2007), resulting in exchange contributions to transverse relaxation rate constants. This can be probed by NMR relaxation dispersion methods from which one can gain unique information on the structures, thermodynamics and dynamics of the underlying processes (Palmer 2004; Palmer et al. 2001). To date, Carr–Purcell–Meiboom–Gill (CPMG) experiments (Carr and Purcell 1954; Meiboom and Gill 1958), that cover the millisecond time scale, have been designed for amide 15N (Loria et al. 1999a, b) and 1H (Ishima and Torchia 2003), backbone CO (Lundström et al. 2008) and Cα (Lundström et al. 2009a), aliphatic side chain Cβ (Lundström et al. 2009b) and 1H (Hansen et al. 2012), side chain CO (Hansen and Kay 2011), methyl groups (Baldwin et al. 2010; Mulder et al. 2002; Otten et al. 2010; Weininger et al. 2012b) and aromatic side chain 13C (Weininger et al. 2012c).
Aromatic side chains are bulky and responsible for a significant proportion of the protein hydrophobic core. They typically form pairs or clusters where they make specific aromatic–aromatic interactions (Burley and Petsko 1985, 1989). They are prevalent in protein binding interfaces (Trp is four times enriched in binding sites, relative to their natural occurrence, Tyr > 2 times, the only other amino acid > 2 is Arg), where they contribute significantly to the binding free energy (Birtalan et al. 2010; Bogan and Thorn 1998; Lo Conte et al. 1999). His and Tyr (18% and 6% of all catalytic residues) also play prominent roles in enzyme catalysis (Bartlett et al. 2002). All this makes aromatic side chains interesting and useful probes for studying protein dynamics. Even more, Phe and Tyr undergo frequent 180° rotations of the χ2 angle (‘ring flips’) and thereby provide unique information of transient ‘breathing’ processes of proteins (Li et al. 1999; Wagner 1980; Wagner et al. 1976). His can exist in three different states, one protonated and two different neutral tautomeric forms. Transient changes between these states affect hydrogen bonding patterns around the histidine. Thus, it is of great interest to monitor the dynamics of aromatic residues. With easy and robust labeling protocols to achieve site-selective 13C labeling (Lundström et al. 2007; Teilum et al. 2006; Weininger 2017) studies of dynamics on aromatic side chains have come into focus. Improved methods of obtaining relaxation rate constants have been developed (Weininger et al. 2012a) and the first studies of order parameters have been reported (Boyer and Lee 2008; Kasinath et al. 2013, 2015). 13C relaxation dispersion experiments for the study of dynamics on the ms (Weininger et al. 2012c) and µs (Weininger et al. 2014a) time scale have been developed and applied on the characterization of ring flips (Weininger et al. 2013, 2014b) and transient histidine tautomerization (Weininger et al. 2017). Furthermore, residual dipolar couplings have been obtained (Sathyamoorthy et al. 2013).
So far, all this new studies of aromatic side chain dynamics are based on 13C. Here we investigate the possibility of 1H CPMG relaxation dispersion experiments in aromatic side chains, as a complementary method. Some processes will more reflect in 1H others in 13C CPMG relaxation dispersion profiles. 1H CPMG relaxation dispersion experiments can be affected by 3J 1H–1H couplings and 13C–13C strong couplings. Artifact-free relaxation dispersion profiles can be obtained in uniformly 1H and 13C labeled samples in case of His δ2, His ε1 and Trp δ1, since 3J 1H–1H couplings are sufficiently small and 13C–13C strong couplings do not occur in these positions. Other positions require site-selective 13C labeling in case of 13C–13C strong couplings and site-selective deuteration in all cases. 1H CPMG relaxation dispersion experiments were applied and verified on unfolding of CspB (Weininger et al. 2012c; Zeeb and Balbach 2005) for His δ2, His ε1 and Trp δ1, therefore providing a complementary approach to 13C CPMG relaxation dispersion experiments.
Materials and methods
Protein samples
Uniformly 15N13C labeled ubiquitin at pH 6.5 was purchased from ASLA. Uniformly 15N13C labeled and site-selective 13C labeled, using 1-13C and 2-13C glucose (Lundström et al. 2007; Teilum et al. 2006), GB1 (QDD variant) at pH 7.0 in water was expressed and purified as described in Lindman et al. (2006). Site-selective 13C labeled, using 1-13C glucose, SlyD at pH 7.4 in 20 mM HEPES was purified and described as in Löw et al. (2010). Uniformly 13C labeled CspB at pH 7.0 in 10 mM HEPES was expressed and purified as described in Schindler et al. (1995). Uniformly 13C labeled human carbonic anhydrase II, at pH 7.7 in 50 mM TRIS/HCl was expressed and purified as described in Michalczyk et al. (2015).
NMR spectroscopy
All experiments were acquired on a Bruker Avance III spectrometer at a static magnetic field of 14.1 T and 298 K. Additionally, relaxation dispersion experiments on CspB were recorded on Bruker Avance III spectrometer at a static magnetic field of 18.8 T. Samples contained 1–3 mM protein (except carbonic anhydrase, which was below 100 µM) and 10% (v/v) D2O. 1H CPMG relaxation dispersion experiments were performed with refocusing frequencies between 100 and 1000 Hz (2000 Hz in case of CspB) and B1 field strengths for the CPMG pulses between 13 and 15 kHz. Spectra were processed with NMRPipe (Delaglio et al. 1995) and analyzed with PINT (Ahlner et al. 2013).
Data analysis
CPMG relaxation dispersion experiments were fitted globally to the Carver–Richards equation (Carver and Richards 1972; Davis et al. 1994). Data modeling utilized the Levenberg–Marquardt (Press et al. 2002) nonlinear least-squares optimization algorithm implemented in MATLAB. Derived Δδ values were compared with 1H shift differences between native and extrapolated unfolded signals from a NMR urea titration (Weininger et al. 2012c).
Density matrix calculations
The evolution of the spin density operator in Liouville space during the 1H CPMG experiment was simulated by way of the homogeneous master equation (Jeener 1982; Levitt and Dibari 1992) as implemented in Qsim (Helgstrand and Allard 2004). Cross relaxation effects were included. The simulations covered the full CPMG relaxation period. Simulations were performed for 3- or 4-spin systems for the directly coupled 1H-13C spin pair of interest to one 1H, two 1H, or one 1H bound to 13C, for 3J 1H–1H coupling constants of 2 Hz or 8 Hz. The 1H carrier was set 1000 Hz away from the proton of interest. The loss of magnetization was used to calculate an artificial Rex contribution.
Results and discussion
Couplings in aromatic side chains
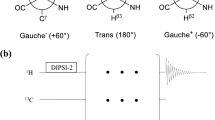
Acquiring artifact-free 1H relaxation dispersion profiles in aromatic side chains (Fig. 1) can potentially be prevented by different J-couplings within the aromatic ring. First and foremost by 3J 1H–1H couplings of a certain size. 3J 1H–1H couplings in Phe, Tyr and the six-ring moiety of Trp are around 8 Hz as evaluated from 1H spectra of the free amino acids. In contrast, no J-couplings between carbon bound protons and nitrogen bound protons in His (δ2 and ε1) and the five-ring moiety of Trp (δ1) could be detected. It was therefore concluded that these couplings are 2 Hz or smaller. Additionally, 1H–1H or 13C–13C strong couplings (chemical shift difference of the coupling nuclei is not sufficiently larger than the coupling constant) could affect the relaxation dispersion profiles. Strong couplings can lead to severe additional losses of magnetization, at high or low CPMG refocusing frequencies (Weininger et al. 2013). 1H–1H strong coupling can be expected in Phe, Tyr and the six-ring moiety of Trp, but not in His (δ2 and ε1) and the five-ring moiety of Trp (δ1), because the chemical shift of the nitrogen bound proton is too different (Ulrich et al. 2008). Additionally, the underlying 3J coupling constant of 2 Hz or below is too small to matter (Weininger et al. 2013). 13C–13C strong coupling can be expected in Phe, Tyr (Cγ–Cδ) and the six-ring moiety of Trp, but again not in His (δ2 and ε1) and the five-ring moiety of Trp (δ1), as evaluated from possible chemical shifts (Ulrich et al. 2008). In Trp, possible strong couplings can directly be evaluated from the spectra. In Phe and Tyr, were signals are most often averaged because of fast ring flips (Wagner et al. 1976), this is often not possible.
Acquiring 1H CPMG relaxation dispersion profiles
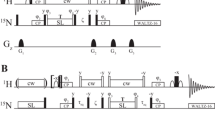
The pulse sequence for measurement of 1H CPMG relaxation dispersions in aromatic side chains is shown in Fig. 2. It uses the relaxation compensation approach between proton inphase and proton–carbon antiphase magnetization (Loria et al. 1999a). To account for different 1H–13C J-couplings in aromatic side chain purging 90° pulses are applied on both sides of the block converting inphase to antiphase magnetization (Vallurupalli et al. 2007; Weininger et al. 2012c). 2D detection can be achieved in a constant-time or non constant-time approach, while water is suppressed by selection gradients (GC and GH). Since the TROSY effect for aromatic protons is neglectable, no TROSY selection of the relaxation rate constant was applied (Loria et al. 1999b; Weininger et al. 2012c). Also shaped CPMG pulses on proton are not practical, because of their long duration, which would allow only low refocusing frequencies. In summary 1H CPMG relaxation dispersions can be recorded in a simple, robust and straightforward way.
Pulse sequence of the 1H CPMG relaxation dispersion experiment for measuring conformational exchange of aromatic side chains. All pulses are applied along the x-axis unless otherwise indicated. Narrow (wide) solid bars indicate rectangular high-power 90° (180°) pulses. Wide grey bars indicate 180° pulses in the CPMG elements, which have attenuated power. The wide semi-ellipse on 13C represents a REBURP (Geen and Freeman 1991) pulse with a bandwidth of 40 ppm. All proton pulses after (a) are applied on resonant on the aromatics, after (b) on-resonant on water. The delay τ can be set to 1.6 ms (Phe and Tyr), 1.35 ms (all aromatics) or 1.25 ms (His). Ta and Tb are 1.1 ms and 1.1 ms + t1/2 for non constant-time detection and 4.464 ms − t1/4 and 4.464 ms + t1/4 for constant-time detection, respectively. The pulses flanking the CPMG blocks purge non-refocused magnetization remaining as a consequence of the variation among aromatic sites in the 1JHC coupling constant (Vallurupalli et al. 2007; Weininger et al. 2012c). The phase cycle is: ϕ1 = (x, − x), ϕ2 = (x, x, − x, − x), ϕrec = (x, − x, − x, x). Pulsed field gradients G1–3 are employed to suppress unwanted coherences and artifacts, while GC and GH are encoding and decoding gradients, respectively, for echo/anti-echo coherence selection, obtained by inverting the signs of GH (Kay et al. 1992). For every second t1 increment ϕ1 and the receiver were incremented. Gradient durations (in ms) and relative power levels (in %) are set to (duration, power level) G1 = (1.0, 13), G2 = (0.5, 10), G3 = (1.0, 90), GC = (1.0, 80), GH = (1.0, − 20.1)
Simulation of 3J 1H–1H couplings
In order to quantify the disturbance of aromatic 1H CPMG relaxation dispersion profiles caused by 3J 1H–1H couplings, density operator simulations of the CPMG relaxation period were performed, including cross relaxation effects. For His δ2, His ε1 and Trp δ1 simulations were performed assuming a maximal J-coupling of 2 Hz. The effect of this J-coupling on the relaxation dispersion is minimal (Fig. 3a). Relaxation rate constants are not affected at all for most refocusing frequencies, resulting in a rate constant of zero. Only for refocusing frequencies of 400 Hz and 500 Hz a small increase of 0.5 Hz is observed, which should be well within the experimental noise. Also the simulation is already the worst case scenario, assuming 2 Hz coupling. In Phe, Tyr and the six-ring moiety of Trp, the J-coupling is 8 Hz. Here a clear artificial contribution to the relaxation dispersion profile is observed (Fig. 3b). In case of one coupling proton the artificial dispersion step is 2 s−1, for two coupling protons it is 4 s−1. In case the coupling proton is connected to 13C, the effect is slightly more severe, than for just one coupling proton. In summary the simulations show that 3J 1H–1H couplings should not be a problem for His δ2, His ε1 and Trp δ1, but lead to certain artifacts in other positions.
Simulation of additional R2 contributions in aromatic 1H CPMG relaxation dispersion profiles caused by 3J 1H–1H couplings. Simulations were performed with Qsim (Helgstrand and Allard 2004). a Assuming a J-coupling of 2 Hz, which is the maximal J-coupling in case of His δ2, His ε1 and Trp δ1 (RMSD 0.12 s−1). b Assuming a J-coupling of 8 Hz, which is the coupling in Phe, Tyr and the six-ring moiety of Trp. Here, simulations were performed for one coupling 1H (black circles), two coupling 1H (black squares) or one coupling 1H which itself is directly coupling to 13C (red circles)
Off-resonance effects
In order for the CPMG block to work as intended, it is often crucial that the pulses are applied close to on-resonant. This was investigated on His 68 of ubiquitin. One CPMG relaxation dispersion experiment was performed with the 1H carrier during the CPMG block at 7.5 ppm, a second with the carrier at 7.0 ppm. The first experiment is within 150 Hz on-resonant on H68 ε1, the second on H68 δ2. Accordingly the other experiments are performed within 400–500 Hz off-resonant to the respective His position. The on-resonant cases (Fig. 4a, b, black symbols) result in flat dispersion profiles, as one can expect for positions with low 3J 1H–1H couplings in a system with known absence of ms exchange. In contrast already for somewhat off-resonant cases (Fig. 4a, b, red symbols) the relaxation dispersion profile is affected. The severe influence of the 1H carrier during the CPMG block is unexpected and not in agreement with the simulation. Although this effect is not understood, artifact-free relaxation dispersion profiles can be acquired if CPMG pulses are applied close to on-resonant, in our experience within 300 Hz to the signals of interest.
Aromatic 1H CPMG relaxation dispersion profiles of ubiquitin H68 δ2 (a), H68 ε1 (b) and GB1 W43 δ1 (c). Black symbols represent a close to on-resonant carrier during the CPMG block, red symbols an off-resonant carrier. RMSD values for on-resonant experiments (black symbols) are 0.18 s−1, 0.12 s−1, 0.26 s−1
Artifact-free 1H CPMG dispersion profiles of His δ2, His ε1 and Trp δ1
Density operator simulations, as well as considerations of strong couplings, lead to the conclusion, that artifact-free aromatic 1H CPMG relaxation dispersion profiles can be obtained for His δ2, His ε1 and Trp δ1. This was validated experimentally by studying two model proteins with known absence of ms dynamics at 25 °C, ubiquitin for His δ2 and ε1 (H68), and GB1 for Trp δ1 (W43). They all show artifact-free flat dispersion profiles, as one should expect for the absence of ms dynamics (Fig. 4, black symbols). Hence, these positions are suitable for aromatic 1H CPMG relaxation dispersion experiments in uniformly or site-selectively 13C labeled proteins.
Effect of couplings and strong couplings in aromatic six-ring moieties
Density operator simulations of aromatic six-ring moieties (3J 1H–1H of 7–8 Hz) show an artificial increase of R2 in the dispersion profiles with increasing refocusing frequencies caused by the 3J 1H–1H coupling (Fig. 3b). Experimental relaxation dispersion profiles of uniformly 13C labeled ubiquitin also show this behavior for refocusing frequencies of 300–1000 Hz, but also display additional very high R2 values for refocusing frequencies of 100 and 200 Hz (Fig. 5a–c). In contrast, relaxation dispersion profiles of site-selectively 13C labeled tSlyD only showed the profile expected from the simulation (Fig. 5d–f). The low frequency artifact in ubiquitin can be addressed to 13C–13C strong couplings. Strong couplings of the nuclei directly coupled to the nuclei investigated by CPMG relaxation experiments can have dramatic effects on the dispersion profiles (Weininger et al. 2013). Phe (Fig. 5a, b) are likely to display 13C–13C strong couplings, while Tyr δ (Fig. 5c) can display 13C–13C strong couplings to Tyr γ (Ulrich et al. 2008). In order to elaborate this further, relaxation dispersion profiles were acquired on uniformly and site-selectively 13C labeled GB1 (Fig. 5g–l). Trp ε3 and ζ3 (Fig. 5g, h) are strongly 13C–13C coupled without a doubt, as directly seen from the spectra. They show extremely high R2 values at low refocusing frequencies in case of uniformly, but not for site-selectively 13C labeling. Trp η2 (Fig. 5i) is not strongly 13C–13C coupled, as seen from the spectra, and shows the expected profile even with uniform 13C labeling. Additionally, for Phe δ (Fig. 5j) the low frequency artifact can be observed for uniform, but not for site-selective 13C labeling and Tyr does also show the expected profiles independent of the labeling. In summary, we experimentally found profiles similar to the simulated ones, that are caused by a sizable 3J 1H–1H coupling. R2 values increase with higher refocusing frequencies. Additionally, 13C–13C strong couplings cause very high R2 values at low refocusing frequencies. 1H–1H strong couplings in tSlyD F79 (Weininger et al. 2013; Fig. 5d) only cause a constant shift of R2 values independent of the refocusing frequency.
Aromatic 1H CPMG relaxation dispersion profiles of ubiquitin (a–c) F4 δ* (a), F4 ε* (b), Y59 δ* (c), tSlyD (d–f) F79 δ* (d), F91 δ* (e), F117 δ* (f), GB1 (g–l) W43 ε3 (g), W43 ζ3 (h), W43 η2 (i), F52 δ* (j), Y33 δ* (k) and Y33 ε* (l). Red symbols represent measurements on uniformly 13C labeled samples, black symbols measurements on site-selective 13C labeled samples
Unfolding of CspB by 1H CPMG relaxation dispersion
Aromatic 1H CPMG relaxation dispersion experiments were applied on CspB from Bacillus subtilis (Fig. 6). CspB is folding/unfolding on the ms time scale and has been used as a model system for CPMG relaxation dispersion experiments (Weininger et al. 2012c; Zeeb and Balbach 2005). It contains one Trp and one His, that allow artifact-free 1H CPMG relaxation dispersion experiments on H29δ2, H29ε1 and W8δ1. The corresponding dispersion were fitted globally to a two-state exchange model with kex = 690 ± 65 s−1 and pu = 1.8 ± 0.1% (Fig. 7). The resulting rate constant of unfolding, which is more or less independent from buffer conditions, is ku = 12 ± 1 s−1. This is in excellent agreement with kex = 528 ± 52 s−1, pu = 2.3 ± 0.2% and ku = 12 ± 3 s−1 derived from aromatic 13C CPMG relaxation dispersion experiments under the same conditions (Weininger et al. 2012c). kex and pu are somewhat dependent on the stabilizing effect of salt, resulting in lower pu and higher kex (because of an increased kf). Chemical shift differences derived from the relaxation dispersion experiments match chemical shift differences derived from a urea transition, back extrapolated to 0 M urea (Fig. 8). Taken together, the unfolding of CspB can be described accurately by aromatic 1H CPMG relaxation dispersion experiments on Hδ2, Hε1 and Wδ1.
Ribbon representation of CspB using 1csp.pdb. The ribbon is colored in blue, side chains of W8 and are shown as sticks, labeled and colored in red. The figure was generated using PyMOL (Schrodinger 2010)
Aromatic 1H CPMG relaxation dispersion profiles acquired on a 1.8 mM sample of CspB in 10 mM HEPES pH 7.0 at 25 °C and static magnetic field strengths of 14.1 T (blue) and 18.8 T (red). Data are shown for H29 δ2 (a), H29 ε1 (b) and W8 δ1 (c). Solid lines represent the global fit to the experimental data
Correlation of 1H chemical shift differences between the folded and unfolded states of CspB derived from CPMG relaxation dispersion experiments under native conditions and measured directly from 1H–13C HSQC spectra in an urea titration experiment. The solid black line represents the ideal correlation
Outlook: larger systems
We determined 1H R2 values at high CPMG refocusing frequencies of various Hε1 on human carbonic anhydrase II, a 29 kDa enzyme. These rate constants vary between 10 and 50 s−1, which are workable values from a background relaxation perspective. Thus such systems can be investigated by aromatic 1H CPMG relaxation dispersion experiments, if enough sensitivity is provided.
Conclusions
We have investigated experimentally and theoretically the possibility of acquiring 1H CPMG relaxation dispersion experiments in aromatic side chains complementary to 13C CPMG relaxation dispersion experiments.. Positions with high (7–8 Hz) 3J 1H–1H couplings (Fδ, Fε, Fζ, Yδ, Yε, Wε3, Wζ3, Wη2 and Wζ2) do not allow to acquire artifact-free relaxation dispersion profiles in uniformly protonated samples. They can be impacted further by 13C–13C strong couplings in uniformly 13C labeled samples. In contrast, Hδ2, Hε1 and Wδ1 only have small (< 2 Hz) 3J 1H–1H couplings and no 13C–13C strong couplings, thus, providing artifact-free relaxation dispersion profiles. Therefore, correct parameters of CspB unfolding could be derived by 1H CPMG relaxation dispersion experiments on a uniformly 1H and 13C labeled sample.
References
Ahlner A, Carlsson M, Jonsson BH, Lundström P (2013) PINT: a software for integration of peak volumes and extraction of relaxation rates. J Biomol NMR 56:191–202. https://doi.org/10.1007/s10858-013-9737-7
Baldwin AJ, Religa TL, Hansen DF, Bouvignies G, Kay LE (2010) (CHD2)-C-13 methyl group probes of millisecond time scale exchange in proteins by H-1 relaxation dispersion: an application to proteasome gating residue dynamics. J Am Chem Soc 132:10992–10995
Bartlett GJ, Porter CT, Borkakoti N, Thornton JM (2002) Analysis of catalytic residues in enzyme active sites. J Mol Biol 324:105–121
Birtalan S, Fisher RD, Sidhu SS (2010) The functional capacity of the natural amino acids for molecular recognition. Mol Biosyst 6:1186–1194. https://doi.org/10.1039/b927393j
Boehr DD, McElheny D, Dyson HJ, Wright PE (2006) The dynamic energy landscape of dihydrofolate reductase catalysis. Science 313:1638–1642
Bogan AA, Thorn KS (1998) Anatomy of hot spots in protein interfaces. J Mol Biol 280:1–9
Boyer JA, Lee AL (2008) Monitoring aromatic picosecond to nanosecond dynamics in proteins via C-13 relaxation: expanding perturbation mapping of the rigidifying core mutation, V54A, in eglin C. Biochemistry 47:4876–4886
Burley SK, Petsko GA (1985) Aromatic-aromatic interaction: a mechanism of protein structure stabilization. Science 229:23–28
Burley SK, Petsko GA (1989) Electrostatic interactions in aromatic oligopeptides contribute to protein stability. Trends Biotechnol 7:354–359
Carr HY, Purcell EM (1954) Effects of diffusion on free precession in nuclear magnetic resonance. Exp Phys Rev 94:630–638
Carver JP, Richards RE (1972) General 2-site solution for chemical exchange produced dependence of T2 upon Carr-purcell pulse separation. J Magn Reson 6:89. https://doi.org/10.1016/0022-2364(72)90090-X
Cole R, Loria JP (2002) Evidence for flexibility in the function of ribonuclease A. Biochemistry 41:6072–6081
Davis DG, Perlman ME, London RE (1994) Direct measurements of the dissociation-rate constant for inhibitor-enzyme complexes via the T-1-Rho and T-2 (Cpmg) methods. J Magn Reson B 104:266–275. https://doi.org/10.1006/jmrb.1994.1084
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) Nmrpipe—a multidimensional spectral processing system based on unix pipes. J Biomol NMR 6:277–293
Demers JP, Mittermaier A (2009) Binding mechanism of an SH3 domain studied by NMR and ITC. J Am Chem Soc 131:4355–4367. https://doi.org/10.1021/ja808255d
Eisenmesser EZ, Bosco DA, Akke M, Kern D (2002) Enzyme dynamics during catalysis. Science 295:1520–1523
Geen H, Freeman R (1991) Band-selective radiofrequency pulses. J Magn Reson 93:93–141. https://doi.org/10.1016/0022-2364(91)90034-Q
Gutowsky HS, Saika A (1953) Dissociation, chemical exchange, and the proton magnetic resonance in some aqueous electrolytes. J Chem Phys 21:1688–1694
Hansen AL, Kay LE (2011) Quantifying millisecond time-scale exchange in proteins by CPMG relaxation dispersion NMR spectroscopy of side-chain carbonyl groups. J Biomol NMR 50:347–355
Hansen AL, Lundstrom P, Velyvis A, Kay LE (2012) Quantifying millisecond exchange dynamics in proteins by CPMG relaxation dispersion NMR using side-chain H-1 probes. J Am Chem Soc 134:3178–3189
Helgstrand M, Allard P (2004) QSim, a program for NMR simulations. J Biomol NMR 30:71–80. https://doi.org/10.1023/B:Jnmr.0000042962.00088.F4
Igumenova TI, Brath U, Akke M, Palmer AG (2007) Characterization of chemical exchange using residual dipolar coupling. J Am Chem Soc 129:13396–13396+
Ishima R, Torchia DA (2003) Extending the range of amide proton relaxation dispersion experiments in proteins using a constant-time relaxation-compensated CPMG approach. J Biomol NMR 25:243–248
Jeener J (1982) Superoperators in magnetic-resonance. Adv Magn Reson 10:1–51
Kasinath V, Valentine KG, Wand AJ (2013) A C-13 labeling strategy reveals a range of aromatic side chain motion in calmodulin. J Am Chem Soc 135:9560–9563
Kasinath V, Fu YN, Sharp KA, Wand AJ (2015) A sharp thermal transition of fast aromatic-ring dynamics in ubiquitin. Angew Chem Int Ed 54:102–107
Kay LE, Keifer P, Saarinen T (1992) Pure absorption gradient enhanced heteronuclear single quantum correlation spectroscopy with improved sensitivity. J Am Chem Soc 114:10663–10665. https://doi.org/10.1021/Ja00052a088
Levitt MH, Dibari L (1992) Steady-state in magnetic-resonance pulse experiments. Phys Rev Lett 69:3124–3127. https://doi.org/10.1103/PhysRevLett.69.3124
Li H, Yamada H, Akasaka K (1999) Effect of pressure on the tertiary structure and dynamics of folded basic pancreatic trypsin inhibitor. Biophys J 77:2801–2812
Lindman S, Xue WF, Szczepankiewicz O, Bauer MC, Nilsson H, Linse S (2006) Salting the charged surface: pH and salt dependence of protein G B1 stability. Biophys J 90:2911–2921. https://doi.org/10.1529/biophysj.105.071050.
Lo Conte L, Chothia C, Janin J (1999) The atomic structure of protein-protein recognition sites. J Mol Biol 285:2177–2198
Loria JP, Rance M, Palmer AG (1999a) A relaxation-compensated Carr-Purcell-Meiboom-Gill sequence for characterizing chemical exchange by NMR spectroscopy. J Am Chem Soc 121:2331–2332
Loria JP, Rance M, Palmer AG (1999b) A TROSY CPMG sequence for characterizing chemical exchange in large proteins. J Biomol NMR 15:151–155
Löw C et al (2010) Crystal structure determination and functional characterization of the metallochaperone SlyD from Thermus thermophilus. J Mol Biol 398:375–390
Lundström P et al (2007) Fractional C-13 enrichment of isolated carbons using [1-C-13]- or [2-C-13]-glucose facilitates the accurate measurement of dynamics at backbone C-alpha and side-chain methyl positions in proteins. J Biomol NMR 38:199–212
Lundström P, Hansen DF, Kay LE (2008) Measurement of carbonyl chemical shifts of excited protein states by relaxation dispersion NMR spectroscopy: comparison between uniformly and selectively (13)C labeled samples. J Biomol NMR 42:35–47
Lundström P, Hansen DF, Vallurupalli P, Kay LE (2009a) Accurate measurement of alpha proton chemical shifts of excited protein states by relaxation dispersion NMR spectroscopy. J Am Chem Soc 131:1915–1926
Lundström P, Lin H, Kay LE (2009b) Measuring (13)C(beta) chemical shifts of invisible excited states in proteins by relaxation dispersion NMR spectroscopy. J Biomol NMR 44:139–155
Malmendal A, Evenas J, Forsen S, Akke M (1999) Structural dynamics in the C-terminal domain of calmodulin at low calcium levels. J Mol Biol 293:883–899
Meiboom S, Gill D (1958) Modified spin-echo method for measuring nuclear relaxation times. Rev Sci Instrum 29:688–691
Michalczyk R et al (2015) Joint neutron crystallographic and NMR solution studies of Tyr residue ionization and hydrogen bonding: implications for enzyme-mediated proton transfer. Proc Natl Acad Sci USA 112:5673–5678
Mittermaier AK, Kay LE (2009) Observing biological dynamics at atomic resolution using NMR trends. Biochem Sci 34:601–611
Mulder FAA, Hon B, Mittermaier A, Dahlquist FW, Kay LE (2002) Slow internal dynamics in proteins: application of NMR relaxation dispersion spectroscopy to methyl groups in a cavity mutant of T4 lysozyme. J Am Chem Soc 124:1443–1451
Otten R, Villali J, Kern D, Mulder FAA (2010) Probing microsecond time scale dynamics in proteins by methyl H-1 Carr-Purcell-Meiboom-Gill relaxation dispersion nmr measurements. Application to activation of the signaling protein NtrC(r). J Am Chem Soc 132:17004–17014
Palmer AG (2004) NMR characterization of the dynamics of biomacromolecules. Chem Rev 104:3623–3640
Palmer AG, Kroenke CD, Loria JP (2001) Nuclear magnetic resonance methods for quantifying microsecond-to-millisecond motions in biological macromolecules. Methods Enzymol 339:204–238
Press WH, Teukolsky SA, Vetterling WT, Flannery BP (2002) Numerical recipes in C++: the art of scientific computing, 2nd edn. Cambridge University Press, Cambridge
Sathyamoorthy B, Singarapu KK, Garcia AE, Szyperski T (2013) Protein conformational space populated in solution probed with aromatic residual dipolar C-13-H-1 couplings. ChemBioChem 14:684–688
Schindler T, Herrler M, Marahiel MA, Schmid FX (1995) Extremely rapid protein-folding in the absence of intermediates. Nat Struct Biol 2:663–673
Schrodinger LLC (2010) The PyMOL Molecular Graphics System, Version 1.3r1
Teilum K, Brath U, Lundstrom P, Akke M (2006) Biosynthetic C-13 labeling of aromatic side chains in proteins for NMR relaxation measurements. J Am Chem Soc 128:2506–2507
Ulrich EL et al (2008) BioMagResBank. Nucleic Acids Res 36:D402–D408
Vallurupalli P, Hansen DF, Stollar E, Meirovitch E, Kay LE (2007) Measurement of bond vector orientations in invisible excited states of proteins. Proc Natl Acad Sci USA 104:18473–18477
Wagner G (1980) Activation volumes for the rotational motion of interior aromatic rings in globular-proteins determined by high-resolution H-1-Nmr at variable pressure. FEBS Lett 112:280–284
Wagner G, Demarco A, Wuthrich K (1976) Dynamics of aromatic amino-acid residues in globular conformation of basic pancreatic trypsin-inhibitor (Bpti).1. H-1 Nmr-studies. Biophys Struct Mech 2:139–158
Weininger U (2017) Site-selective 13C labeling of proteins using erythrose. J Biomol NMR 67:191–200. https://doi.org/10.1007/s10858-017-0096-7
Weininger U, Diehl C, Akke M (2012a) C-13 relaxation experiments for aromatic side chains employing longitudinal- and transverse-relaxation optimized NMR spectroscopy. J Biomol NMR 53:181–190
Weininger U, Liu Z, McIntyre DD, Vogel HJ, Akke M (2012b) Specific 12CbD212CgD2S13CeHD2 isotopomer labeling of methionine to characterize protein dynamics by 1H and 13C NMR relaxation dispersion. J Am Chem Soc 134:18562–18565. https://doi.org/10.1021/ja309294u
Weininger U, Respondek M, Akke M (2012c) Conformational exchange of aromatic side chains characterized by L-optimized TROSY-selected C-13 CPMG relaxation dispersion. J Biomol NMR 54:9–14
Weininger U, Respondek M, Low C, Akke M (2013) Slow aromatic ring flips detected despite near-degenerate NMR frequencies of the exchanging nuclei. J Phys Chem B 117:9241–9247
Weininger U, Brath U, Modig K, Teilum K, Akke M (2014a) Off-resonance rotating-frame relaxation dispersion experiment for C-13 in aromatic side chains using L-optimized TROSY-selection. J Biomol NMR 59:23–29
Weininger U, Modig K, Akke M (2014b) Ring flips revisited: C-13 relaxation dispersion measurements of aromatic side chain dynamics and activation barriers in basic pancreatic trypsin inhibitor. Biochemistry 53:4519–4525
Weininger U, Modig K, Geitner AJ, Schmidpeter PAM, Koch JR, Akke M (2017) Dynamics of aromatic side chains in the active site of FKBP. Biochemistry 56(12):334–343. https://doi.org/10.1021/acs.biochem.6b01157
Zeeb M, Balbach J (2005) NMR spectroscopic characterization of millisecond protein folding by transverse relaxation dispersion measurements. J Am Chem Soc 127:13207–13212
Acknowledgements
This research was supported by the Deutsche Forschungsgemeinschaft (WE 5587/1-1).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Raum, H.N., Dreydoppel, M. & Weininger, U. Conformational exchange of aromatic side chains by 1H CPMG relaxation dispersion. J Biomol NMR 72, 105–114 (2018). https://doi.org/10.1007/s10858-018-0210-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-018-0210-5