Abstract
These cross-sectional studies reported the occurrence, genetic characteristics, and factors associated with the distribution of Listeria species on cattle farms and beef abattoirs in Gauteng Province, South Africa. A total of 328 samples (faeces, feeds, silage, and drinking water) were collected from 23 cattle farms (communal, cow-calf, and feedlot), and 262 samples (faeces, carcass swabs, and effluents) from 8 beef abattoirs (low throughput and high throughput) were processed using standard bacteriological and molecular methods to detect Listeria species. The factors associated with the prevalence of Listeria species were investigated, and multiplex polymerase chain reaction (mPCR) was used to determine Listeria species, the pathogenic serogroups, and the carriage of eight virulence-associated genes by Listeria monocytogenes. The overall prevalence of Listeria species in cattle farms was 14.6%, comprising Listeria innocua (11.3%), Listeria monocytogenes (3.4%), Listeria welshimeri (0.0%) compared with 11.1%, comprising Listeria innocua (5.7%), Listeria monocytogenes (4.6%), Listeria welshimeri (0.8%) for beef abattoirs. Of the three variables (area, type of farm/abattoir, and sample type) investigated, only the sample types at abattoirs had a significant (P < 0.001) effect on the prevalence of L. innocua and L. welshimeri. The frequency of distribution of the serogroups based on 11 L. monocytogenes isolated from farms was 72.7% and 27.3% for the serogroup 1/2a-3a and 4b-4d-4e, respectively, while for the 12 L. monocytogenes isolates recovered from abattoirs, it was 25%, 8.3%, 50% and 16.7% for the serogroup 1/2a-3a, 1/2b-3b, 1/2c-3c, and 4b-4d-4e respectively (P < 0.05). All (100%) isolates of L. monocytogenes from the farms and abattoirs were positive for seven virulence genes (hlyA, inlB, plcA, iap, inlA, inlC, and inlJ). The clinical and food safety significance of the findings cannot be ignored.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Listeria monocytogenes is a major cause of ruminant listeriosis, even though it can infect various animal species. Listeriosis in ruminants can also be caused by L. ivanovii, which is non-pathogenic for other animal species and humans (Kaur and Balgir 2021). Cattle and other livestock are exposed to these pathogens through improperly produced silage, feeds, and contaminated water (Rodriguez et al. 2021). Some clinical manifestations of listeriosis in ruminant animals include encephalitis. Listeriosis in cattle has been reported in several countries, such as Latvia (Terentjeva et al. 2021), the USA (Nightingale et al. 2004), Ireland (Hilliard et al. 2018), Jordan (Obaidat et al. 2020), Spain (Hurtado et al. 2017), England (McLauchlin et al. 2020) and Nigeria (Chuku et al. 2019). The Listeria species have been isolated from slaughterhouses/abattoirs in China (Zhao et al. 2021), Japan (Takahashi et al. 2007), Turkey (Al et al. 2022), Belgium (Demaître et al. 2020) and Nigeria (Aiyedun et al. 2020). Regarding the beef chain and the epidemiology of L. monocytogenes, the organism has been reported to be present on cattle farms, including feedlots and cow-calf operations (Mohammed et al. 2010). Slaughter cattle at abattoirs have been shown to shed L. monocytogenes in their faeces, making this an important source of infection in the meat value chain. The sources of L. monocytogenes on cattle farms have been reported to be feeds, including spoilt silages, faeces, and farm environments (Palacios-Gorba et al. 2021). At the abattoirs, L. monocytogenes has been isolated pre-slaughter from the faeces, peri-anal areas, and skins and carcasses of cattle (Foerster et al. 2012; Zhao et al. 2021).
Listeriosis is an important emerging foodborne disease, causing life-threatening infections in humans, including abortion and stillbirth in pregnant women, septicemia, encephalitis, meningitis, gastroenteritis, and perinatal infections (Dhama et al. 2015). The foodborne bacteria, Listeria. monocytogenes and other Listeria species are highly adaptable pathogens that can persist in various environmental and food-related chains (NicAogáin and O’Byrne, 2016). It can acclimatize and live in extensive stressed conditions such as low water activity, temperature, and pH, making it problematic for producers who depend on these stresses for conservation (Chapin et al. 2014).
Some characteristics of L. monocytogenes, including their serogroups and their carriage of virulence genes, have been associated with the ability of the microorganism to cause listeriosis. The serogroups of L. monocytogenes commonly detected on cattle farms, beef abattoirs, or slaughterhouses include 4b-4d-4e and 1/2b-3b (Castro et al. 2018), and 1/2a-3a, 1/2b-3b, 1/2c-3c and 4b-4d-4e (Demaître et al. 2021). These serogroups are of public health importance and are commonly associated with animal and human listeriosis cases. The predominant serogroups detected in listeriosis were 1/2a-3a, 1/2b-3b, 1/2c-3c, and 4b-4d-4e (Oh et al. 2018; Obaidat et al. 2020). Virulence genes, hly, sigB, plcA, inlB, inlC actA, inlA, inlB, plcB, hlyA, and inlJ have been documented in L. monocytogenes strains isolated from cattle farms, beef abattoirs and in human cases of listeriosis (Al et al. 2022; Oh et al. 2016; Pournajaf et al. 2016). However, the predominant virulence genes reported are plcA, prfA, hlyA, inlB, inlA, inlC, inlJ, actA, and iap (Kayode and Okoh 2022). Some of the modes of action of the prevalent virulence genes are the predominant activation of hlyA and plcA within the phagosomal compartment and actA and inlC in the host cell cytosol (Bubert et al. 1999), while prfA is activated by transcription of the listeriolysin gene (Chakraborty et al. 1992). Some of the virulence genes are involved in adherence to and internalization by the host cell (inlA, inlB, and inlJ), escape from the vacuoles (hly, plcA, and plcB), intracellular replication (htp), and cellular movement (actA) (Kastbjerg et al. 2010).
In South Africa, there is a dearth of information on epidemiological data on the samples assessed for contamination by L. monocytogenes and Listeria species, the risk posed to cattle, beef, and beef products, especially since ‘Polony’, a meat product, was responsible for the large listeriosis outbreak in the country; and the species of Listeria, other than L. monocytogenes, in beef and beef products (Allam et al. 2018). Recently, Matle et al. (2019) conducted a study on raw intact meat, ready-to-eat (RTE) meat products, and raw processed meat in the country’s nine provinces and reported a prevalence of 14.7% for L. monocytogenes. Therefore, the current study was conducted to determine the occurrence, genetic characteristics (pathogenic serogroups and virulence profiles), and the independent factors (area, type of cattle farms/abattoirs, and sample types) associated with the distribution of Listeria species on cattle farms and beef abattoirs in Gauteng Province, South Africa.
Material and methods
Study design and sample size determination
The cross-sectional study was conducted on 23 cattle farms and eight abattoirs in Gauteng province, South Africa, to determine the occurrence and characteristics of Listeria species in cattle farms and abattoirs. Gauteng province is one of the nine provinces and the smallest in South Africa, with approximately 15.81 million people.
A sample size of 328 and 262 for cattle farms and beef abattoirs, respectively, was determined using a formula by Thrusfield (2007); n = [1.962 Pexp (1 − Pexp)]/d2, where n = required minimum sample size, Pexp = estimated prevalence of listeriosis and d = desired absolute precision.
Selection of cattle farms and beef abattoirs, sources, and types of samples, and transportation to the laboratory for processing
The study was designed to randomly select cattle farms and abattoirs from the list made available by the Gauteng Department of Agriculture and Rural Development and Environment (GDARDE). Once the owners and managers of cattle farms and beef abattoirs were unwilling to participate in the study because of the COVID-19 pandemic, the next farm on the list was selected by systematic random sampling.
For the cattle farms consisting of feedlots (n = 3), cow-calf operations (n = 10), and communal farms (n = 10) in Gauteng, South Africa, samples were aseptically collected as described by Onyeka et al. (2021). Faecal (rectal faecal grab or freshly voided faeces) of individual cattle, and environmental samples inclusive of pooled faeces from areas where cattle congregated, drinking water in troughs, feeds (grains and grass), and silage in feeding troughs.
At the beef abattoirs consisting of high throughput, HT (n = 6), and low throughput, LT (n = 2), the following samples were collected using the procedure described by Zweifel et al. (2005). Carcass sampling was done according to the European Union Decision 2001/471/EC. Swab samples were obtained from vertical and horizontal streaks by applying gentle pressure using a swab rinse kit (SRK) (Copan Diagnostics, Inc., UK). The sources and types of samples were as follows: Pre-slaughter faeces in the lairage, carcass swabs [pre-evisceration, post-evisceration, 24 h post-chilled carcasses (-4 °C—20 °C)], abattoir effluents (environmental).
All the cattle farms and beef abattoir samples were collected between 2020 and 2021.
The collected samples from the cattle farms and beef abattoirs were transported to the ARC-Onderstepoort Veterinary Institute’s Feed and Food Laboratory, ice-cooled within 12 h of collection, and processed within 48 h.
Enrichment of samples collected from cattle farms and beef abattoirs and PCR detection of Listeria species
For faecal and swab samples, sterile spoons were used to scoop faecal samples (farm samples) from the cups to sterile Petri dishes to weigh 10 g of the faecal samples, which were transferred aseptically into stomacher bags that contained 90 ml ONE Broth-Listeria (Thermo Fisher, South Africa). The samples were homogenized (Stomacher Laboratory Blender 400, Seward Ltd., West Sussex, UK) at normal speed for 2 min, followed by 48 h aerobic incubation at 35˚ C. For the swab samples from abattoirs, we used a swab rinse kit (SRK) (Copan Diagnostics, Inc., UK), and one millilitre (1 ml) of sample in the SRK was removed into 90 ml tubes of ONE Broth-Listeria (Thermo Fisher, South Africa) for 48 h aerobic incubation at 35˚C. Feed samples were aseptically withdrawn from the cup using forceps, and 10 g of the feed samples (grass and grain) was weighed using a weighing balance and transferred into a stomacher bag which contained 90 ml of ONE Broth-Listeria (Thermo Fisher, South Africa), which was followed by homogenization and aerobic incubation at 35 ˚C for 48 h. The water centrifugation method was used to isolate Listeria species from water and effluent samples for drinking and effluent samples. For each sample, 100 ml was aliquoted into four 25 ml amounts in centrifuge bottles and then spun down at 15,493 × g for five minutes. The pellets were pooled from the four bottles and inoculated into 9 ml of ONE Broth-Listeria (Thermo Fisher, South Africa) for enrichment, followed by aerobic incubation at 35 ˚C for 48 h. The enriched broth was used to inoculate Brilliance-Listeria agar (BLA) (Thermo Fisher Scientific, South Africa) plates to isolate Listeria species. A loopful of enriched broth culture growth in ONE Broth-Listeria (Th Thermo Fisher, South Africa) was inoculated onto Brilliance Listeria Agar (BLA) plates and streaked for isolation. The inoculated plates were incubated aerobically at 35˚C for 48 h. Single colonies of suspected Listeria species (colonies that appeared blue without a halo) and L. monocytogenes (blue colonies with a white/cream halo) were phenotypically identified, as described by Jamali et al. (2013). PCR was used to confirm the isolates of Listeria species. DNA was extracted from enriched broth cultures and isolates by the boiling-centrifugation method, as described by Soumet et al. (1994). The DNA extracts used as templates in the mPCR assays were prepared as described by Soumet et al. (1994). All enriched broth samples were screened by multiplex PCR for Listeria species (Listeria genus), using the prs gene as a target marker (Supplementary Table S1). Screening by PCR was performed utilizing an mPCR assay that targets the genes listed in Supplementary Table S1, as described by Doumith et al. (2004). To detect the different species of Listeria, the DNA extracts used as templates in the mPCR assays were prepared as described by Soumet et al. (1994). The primers utilized in the current study are shown in Supplementary Table S2. The multiplex PCR mix was prepared as recommended (Doumith et al. 2004). PCR amplicons were electrophoresed on a 3.0% agarose gel using 1 × Tris–acetate-EDTA (TAE) buffer and stained with ethidium bromide (Ryu et al. 2013).
Determination of the serogroups of L. monocytogenes isolates
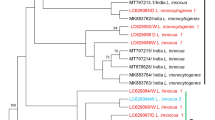
The same mPCR assay method was used to detect the Listeria genus and to characterize L. monocytogenes regarding their serogroups. The mPCR targeting five gene fragments of L. monocytogenes, namely, imo1118, imo0737, orf2110, orf2819, and prs (specific for Listeria genus) was used to determine the serogroups of L. monocytogenes as previously described by Doumith et al. (2004). The five primers used to classify the strains into serogroups are shown in supplementary data, Table S1.
Detection of virulence genes in L. monocytogenes isolates
The presence of selected virulence genes in the isolates of L. monocytogenes was determined as earlier described (Lomonaco et al. 2012; Pournajaf et al. 2016). Multiplex PCR was used to detect eight virulence-associated genes of L. monocytogenes: plcA, hlyA, actA, inIB, lap, inlA, inlC, and inlJ in two multiplexes. Multiplex 1 (mPCR 1) contained 5 primer sets (plcA, hlyA, actA, inIB, and iap). In comparison, Multiplex 2 (mPCR 2) consisted of three primer sets (inlA, inlC, and inlJ) (Supplementary Table S3).
Data analysis
Laboratory data generated for the occurrence of the six species of Listeria (L. monocytogenes, L. innocua, L. welshineri, L. grayi, L. ivanovii, and L. seeligeri), serogroups, and virulence-associated genes from beef and beef products collected at cattle farms and beef abattoirs in the current study were entered into Microsoft Excel 2016. The data were analyzed using Epi Info software (Version 7.0), and the association of variables (independent factors, e.g., area, type of farms and abattoirs, and types of samples) with the detection of Listeria or selected characteristics (dependent factors) was determined using Fishers Exact and Chi-square. The significant difference was evaluated using (P-value < 0.05), and percentages were calculated at a 95% confidence interval. Epi Info was also employed to generate percentages for categorical data on the prevalence of the six species of Listeria by the geographical location of farms and abattoirs, type of farms and abattoirs, and sample types.
Results
Occurrence of Listeria species on cattle farms and abattoirs
Overall, 14.6% (48/328) of the samples collected from the cattle farms were positive for the genus Listeria, while for the beef abattoirs, it was 11.1% (29/262). The difference was not statistically significant (P = 0.201). Of a total of 590 samples processed from the cattle farms and beef abattoirs, 77 (13.1%), 23 (3.9%), 52 (8.8%), and 2 (0.3%) were positive for Listeria species (Listeria isolates that could not be identified to species level), L. monocytogenes, L. innocua, and L. welshimeri, respectively (P < 0.001). Listeria ivanovii, L. grayi, and L. seeligeri were not detected in any of the samples from this study.
Prevalence of L. monocytogenes on cattle farms and beef abattoirs and associated factors
The prevalence of L. monocytogenes on cattle farms and the univariate analysis of associated factors are shown in Table 1. The prevalence of L. monocytogenes was 3.4% (11/328). The three variables assessed (area, type of farm, and type of samples) had no statistically significant (P > 0.05) on the prevalence of L. monocytogenes. Table 2 shows the prevalence of L. monocytogenes in abattoirs and the univariate analysis of associated factors. The prevalence of L. monocytogenes on beef abattoirs was 4.6% (12/262). The location of the abattoirs, type of abattoir, and type of samples tested did not significantly (P > 0.05) affect the prevalence of L. monocytogenes. In comparison, although the prevalence of L. monocytogenes in samples collected from cattle farms, 3.4%, was lower than found in beef abattoirs, 4.6%, the difference was not statistically significant (P = 0.444).
Prevalence of L. innocua on cattle farms and beef abattoirs and associated factors
The prevalence of L. innocua in cattle farms and the univariate analysis of associated factors are shown in Table 1. The overall prevalence of L. innocua on cattle farms was 11.3% (37/328). The three variables (area, type of abattoir, and type of samples) did not have a statistically significant (P > 0.05) effect on the prevalence of L. innocua. At the abattoirs, L. innocua was detected in 5.7% (15/262) of the samples processed (Table 2). Statistically significant (P < 0.001) differences were detected only among the type of samples tested, with a range from 0.0% (chilled carcass swabs) to 31.8% (environmental samples). Comparatively, the prevalence of L. innocua in the samples collected from cattle farms (11.5%) was statistically significantly higher (P = 0.018) than that detected in abattoir samples (5.7%).
Prevalence of L. welshimeri in cattle farms and beef abattoirs
All the 328 samples collected from cattle farms were negative for L. welshimeri, with a prevalence of 0.0%. The prevalence of L. welshimeri in abattoirs and the univariate analysis of associated factors are shown in Table 2. The overall prevalence of the organism was 0.76% (2/262). The prevalence of L. welshimeri varied significantly (P < 0.001) across sample types only (environmental samples, 9.1% versus other types of samples, 0.0%). The difference between the prevalence of L. welshimeri on cattle farms (0%) and beef abattoirs (0.76%) was not statistically significant (P = 0.197).
Frequency of the serogroups of L. monocytogenes isolated from cattle farms and beef abattoirs
The frequency distribution of the serogroups of L. monocytogenes isolated from farms was 72.7% (8/11) and 27.3% (3/11) for 1/2a-3a and 4b-4d-4e, respectively. The difference was statistically significant (P = 0.033). The distribution of the serogroups among isolates of L. monocytogenes recovered from cattle farms by area, type of farms, and type of samples is shown in Table 3. Statistically significant differences were detected in the frequency of L. monocytogenes serogroup 1/2a-3a according to the variables assessed as follows: area (P = 0.001), type of farm (P = 0.030) and type of sample (P = 0.030).
For beef abattoir samples, the frequency of distribution of the serogroups among the L. monocytogenes isolates was 25% (3/12), 8.3% (1/12), 50% (6/12), and 16.7% (2/12) for the serogroup 1/2a-3a, 1/2b-3b, 1/2c-3c, and 4b-4d-4e respectively (Table 4). The area, type of farms, and type of samples did not significantly affect the serogroups of L. monocytogenes (P > 0.05).
Among the serogroups of L. monocytogenes, statistically significant differences were detected in their frequencies between farm and abattoir isolates only for 1/2a-1/3a (P = 0.022) (72.7% versus 25%) and 1/2c-3c (P = 0.014) (0% versus 50%).
Frequency of detection of virulence genes in L. monocytogenes isolates
For the 11 L. monocytogenes isolates from cattle farms and for the eight virulence genes assayed, seven genes (hlyA, inlB, plcA, iap, inlA, nlC, and inlJ), were detected in each of the isolates, 100% (11/11), while actA was detected in 9 of 110 (81.8%) isolates. Furthermore, of the total 8 farm isolates of L. monocytogenes that belonged to serogroup 1/2a-3a, only 6 (75%) were positive for virulence gene actA compared with the three isolates that belonged to serogroup 4b-4d-4e, which were all (100%) for the 8 virulence genes assayed, as shown in Table 5.
Similarly, among the 12 isolates of L. monocytogenes recovered from abattoirs and for the eight virulence genes assayed, the frequency of seven genes (hlyA, inlB, plcA, iap, inlA, nlC, and inlJ) in each isolate was 100% (12/12), but for actA, the frequency was 83.3% (10/12).
Overall, the differences in the frequencies of virulence genes among isolates of L. monocytogenes recovered from cattle farms and beef abattoirs were not statistically significant (P > 0.05).
Frequency of detection of virulence genes in L. monocytogenes isolates according to demography and serogroups
All the isolates of L. monocytogenes from the three types of farms (communal, cow-calf, and feedlot) were carriers of the seven virulence genes, but one isolate each from communal farms and cow-calf operation was negative for gene actA. A total of seven virulence genes were detected in the two serogroups (1/2a-3a and 4b-4d-4e), but the actA gene was detected in 75% (6/8) of the isolates in serogroup 1/2a-3a as shown Table 5.
For the L. monocytogenes isolates from beef abattoirs, all (100%) the isolates of L. monocytogenes were positive for seven genes (hlyA, inlB, plcA, iap, inlA, nlC, and inlJ) except for virulence gene, actA, detected in 10 (83.3%). The area, type of abattoir, and type of samples did not significantly (P > 0.05) affect the detection frequency of virulence-associated genes (Table 6).
For the total of six isolates of L. monocytogenes recovered from the abattoirs that belonged to three serogroups (1/2a-3a, 3 isolates; 1/2b-3b, 1 isolate, and 4b-4d-4e, 2 isolates), all (100%; 6/6) were positive for seven virulence genes ((hlyA, inlB, plcA, iap, inlA, nlC, and inlJ). However, for the six isolates that belonged to serogroup 1/2c-3c, only 4 (66.7%) were positive for virulence gene actA (Table 6).
Overall, there were no statistically significant differences (P > 0.05) in the frequencies of detection of virulence-associated genes in farm and abattoir isolates of L. monocytogenes.
Discussion
For the first time, our study documented the prevalence and characteristics of Listeria species on cattle farms and beef abattoirs in South Africa. This is relevant and significant because the cattle farms (production sector) and beef abattoirs (processing sector) constitute parts of the beef production chain in the country, and ‘polony’, a ready-to-eat beef product, was implicated in the world’s largest known outbreak of human listeriosis, experienced by the country (Allam et al. 2018). Our report of Listeria prevalence and characteristics from the production and processing sectors adds to the prevalence data of L. monocytogenes in beef and beef products (retailing sector), which was earlier reported by Matle et al. (2019).
Although L. monocytogenes is the main species of Listeria responsible for livestock and human listeriosis (Hilliard et al. 2018; Koopmans et al. 2023), L. ivanovii has been associated primarily with livestock listeriosis (Arslan and Baytur 2019; Chand and Sadana 1999), and L. innocua, generally considered a non-pathogen, has been implicated in listeriosis in immunocompromised humans (Moura et al. 2019; Perrin et al. 2003). Hence, in the current study, these three Listeria species were investigated in addition to L. seeligeri and L. grayi. Of the five species of Listeria (L. monocytogenes, L. innocua, L. seeligeri, L. ivanovii, L. grayi, and L. welshimeri) investigated, only two (L. monocytogenes and L. innocua) were detected on the cattle farms. It is important that we detected L. monocytogenes in 3.4% of the samples collected from the cattle farms. This prevalence is higher than the 0.5% found on cattle farms in China but considerably lower than the range of 19% to 42.3% reported in other countries ( Nightingale et al. 2004; Obaidat and Stringer 2019; Hurtado et al. 2017).
In our study, the prevalence of L. monocytogenes according to the types of farms was low. It did not vary significantly across farms (communal farm: 3.4%, cow-calf: 3.4%, feedlot: 3.1%), which were slightly different from the findings of 3.1% and 0.3% reported in cow-calf and feedlots, respectively on cattle farms in central and southern California, USA (Mohammed et al. 2010); and was considerably lower than the farm prevalence of 11% reported in Latvia by Terentjeva et al. (2021). The low farm prevalence of L. monocytogenes found in our study is an indication that cattle farms in Gauteng province may not be important sources of L. monocytogenes to cause cattle listeriosis or to the contamination of slaughterhouses or abattoirs when the cattle are slaughtered.
Interestingly, all the silage samples processed in the current study were negative for L. monocytogenes, which may have contributed to the relatively low prevalence of the pathogen detected on the farms. It has been documented that silage, mainly if poorly fermented or of poor quality, harbour Listeria spp. (Mohammed et al. 2010; Rodriguez et al. 2021), and its consumption has been associated with livestock listeriosis, thus posing a threat to public health (Driehuis et al. 2018; Peng et al. 2022; Queiroz et al. 2018). The prevalences of L. monocytogenes in the other sample types reflect the carriage of the pathogen (pooled faeces: 2.7%) and the cattle’s risk of exposure to the pathogen through communal consumption of (feed, 11.1%) and water (3.3%) in troughs. Compared to our study, where the carriage and faecal shedding of L. monocytogenes was 2.7%, others have documented higher frequencies of 7.1% in Jordan (Obaidat et al. 2020), 18.2% in Slovenia (Bandelj et al. 2018), 28% in Latvia (Terentjeva et al. 2021). Similarly, there was a considerably higher prevalence of L. monocytogenes in mixed feed from the feeding troughs and hay (29%) and in drinking water troughs (28%) on cattle farms in Latvia by Terentjeva et al. (2021). Variability in the farm prevalence of L. monocytogenes may be partly explained by differences in the faecal shedding of the pathogen, the contamination of the feeds, drinking water, farm environments, and management practices (Ferreira et al. 2014; Hurtado et al. 2017; Stipetic et al. 2016; Terentjeva et al. 2021).
In our study, L. innocua was detected with a higher farm prevalence than L. monocytogenes (11.3% versus 3.4%) and from each sample type (faeces, feeds, silage, and drinking water) collected from cattle farms in our study. These findings agree with published reports that L. innocua has a broader distribution on cattle farms elsewhere (Gradovska et al. 2023; Gana et al. 2023). However, considering the organism is viewed as a non-pathogen, the risk of causing livestock and human listeriosis is minimal.
Unlike the farm samples, three Listeria species (L. monocytogenes, L. innocua, and L. welshimeri) were recovered from the abattoirs in our study at a prevalence of 4.6% (L. monocytogenes), 5.7% (L. innocua), 0.8% (L. welshineri). Compared with a similar study conducted in abattoirs in Jos, Nigeria, Dunka et al. (2021) reported a prevalence of 2.5%, 33.6%, 4.4%, and 1.7% for L. monocytogenes, L. ivanovii, L. grayi, and L. seeligeri, respectively. The prevalence of L. monocytogenes in the abattoir samples in the current study is considerably lower than reported for abattoirs in other countries (Al et al. 2022; Demaître et al. 2021).
The differences in the types and frequencies of Listeria species detected in abattoirs across countries may be partly due to the prevalence of L. monocytogenes in slaughtered cattle and sanitary practices during slaughter, which affect cross-contamination of carcasses by L. monocytogenes and other pathogens (Demaître et al. 2020; 2021; Mpundu et al. 2022a; Onyeka et al. 2021). Contrary to the lack of any significant effect of the three variables on the prevalence of L. monocytogenes in our study, others have documented the impact of the regional location of abattoirs (Demaître et al. 2021), type of abattoirs, HT versus LT (Onyeka et al. 2021), and types of samples (Dunka et al. 2021; Matle et al. 2019) on the prevalence of L. monocytogenes.
The comparatively slightly higher prevalence of L. monocytogenes detected at abattoirs (4.6%) than in cattle farms (3.4%) may be explained, in part, by the types of samples processed, which are exposed to a variable degree of cross-contamination. This is because, on cattle farms, the samples collected (silage, feces, feeds, water, and effluents) experience limited cross-contamination compared with abattoir samples (pre-slaughter faecal samples, pre- and post-evisceration swab samples, and chilled carcass swabs) which are subjected to cross-contamination. Cross-contamination of carcasses by L. monocytogenes, Salmonella spp., Shiga-toxin Escherichia coli (STEC), and other pathogens have been reported in the abattoir settings in South Africa and elsewhere (Manqele 2018; Onyeka et al. 2020; Rhoades et al. 2009).
Listeria welshimeri was isolated at a frequency of only 0.8% from beef abattoir samples in our study, a frequency considerably lower than reported by others, ranging from 3.8% to the 22% recorded in abattoirs in other countries (Al et al. 2022; Mpundu et al. 2022b; Mpundu et al. 2022b). Although L. welshimeri is being documented in abattoir samples for the first time in the country, it is generally considered non-pathogenic (Korsak and Szuplewska 2016).
It is of zoonotic significance that our study detected two pathogenic serogroups of L. monocytogenes, 1/2a-3a and 4b-4d-4e, in cattle farm isolates. Serogroup 1/2a-3a was predominantly detected (72.7%), which agrees with a study conducted in the USA, where 1/2a was the predominant serotype recovered from cattle farms (Borucki et al. 2005). The pathogenicity of serotype 1/2a has been attributed to its ability to form biofilms (Huang et al. 2018) and its high resistance to sanitizer and bacteriocins (Orsi et al. 2011). This is indicative that, although the overall prevalence of L. monocytogenes is low (3.4%), cattle exposed to the pathogenic serogroups of L. monocytogenes may be at risk of listeriosis. On the contrary, four pathogenic serogroups, 1/2a-3a, 1/2b-3b, 1/2c-3c, and 4b-4d-4e, were found in our abattoir isolates. These findings agree with the three serogroups (1/2a-3a, 1/2b-3b, and 4b-4d-4e) isolated from Belgian cattle slaughterhouses by Demaître et al. (2021) as these serogroups are commonly isolated from ruminants and widespread in the environment. Furthermore, Wieczorek et al. (2012) indicated that L. monocytogenes contaminated beef carcasses during the slaughter process predominantly harboured serotypes 1/2a, 1/2c, 4b, 1/2b, which also agree with the serogroups detected in the current study. It is also pertinent to mention that these serotypes are the dominant serotypes for food strains, accounting for human and ruminant listeriosis (Wang et al. 2015). Even though we did not classify the isolates into serotypes but, serogroups containing some of the known pathogenic serotypes, the majority harbored genes responsible for virulence in L. monocytogenes, highlighting the pathogenic potential of these isolates. Therefore, our findings of pathogenic serogroups on carcasses in the abattoirs sampled provide helpful information about the dominant serogroups of L. monocytogenes in beef abattoirs and their potential food safety implication in Gauteng province, South Africa, for policymaking, surveillance, and biosecurity. The detection of different serogroups and frequencies in the isolates of L. monocytogenes recovered from cattle farms and beef abattoirs in Gauteng province may be attributed to factors such as the fact that the study design is cross-sectional and not longitudinal, the samples collected and assessed at the farm level originated from only 23 farms in the country, which did not represent the origins of the cattle slaughtered at the eight abattoirs included in the current study.
It is of food safety and public health significance that all the 23 isolates of L. monocytogenes recovered from cattle farms and abattoirs in the current study were carriers of seven virulence genes (hlyA, inlB, plcA, iap, inlA, inlC, and inlJ)) while 19 (82.6%) were positive for the actA gene. This is because the pathogenicity of L. monocytogenes has been associated with the possession of virulence genes, especially those present in the Listeria Pathogenicity Islands (LIPIs) (Lopez-Valladares et al. 2018; Wiktorczyk-Kapischke et al. 2023). Three (plcA, hlyA, and actA) of the virulence genes detected in our study belong to the LIPI-1 cluster genes known to be involved in the infectious life cycle and survival in the food processing environment (Koopmans et al. 2023; Lopez-Valladares et al. 2018). Of relevance is the documentation that virulence genes, including those detected in our study, perform different roles and functions in the pathogenesis of L. monocytogenes and have been implicated in human listeriosis (Koopmans et al. 2023). The high frequency of the eight virulence genes assayed, and the detection of pathogenic serogroups in our L. monocytogenes may increase the pathogenicity of the L. monocytogenes we detected on cattle farms and abattoirs in our study. In agreement with our findings, Matle et al. (2019) detected the same eight virulence at similar frequencies from L. monocytogenes isolates from meat and meat products in South Africa. Varying types and frequencies of virulence genes have been reported for farm and abattoir isolates of L. monocytogenes in other countries (Ayaz et al. 2018; Obaidat et al. 2020; Wieczorek et al. 2012). Most virulence genes detected in our abattoir isolates of L. monocytogenes have been associated with human listeriosis (Arslan and Baytur 2019; Koopmans et al. 2023; Soni et al. 2015). The detected high frequency (83.3%-100.0%) of the eight virulence genes in pathogenic serogroups of the L. monocytogenes isolates recovered in the current study could pose a health risk to humans if contaminated beef and beef products from these sources are consumed.
Conclusions and recommendations
For the first time, the current study demonstrated the presence and distribution of L. monocytogenes, L. innoua, and L. welshimeri in various sample types collected from cattle farms and beef abattoirs in South Africa. The prevalence of L. monocytogenes in samples collected at both the farm (production industry) and abattoir (processing industry) of the beef chain has food safety and public health significance because they belong to pathogenic serogroups and carry virulence genes associated with ruminant and human listeriosis. The detection of L. innocua from cattle farms and abattoirs indicates contamination. It can potentially cause listeriosis in immunocompromised humans should the strains enter the food chain in the farm-abattoir sector of the human food chain.
It is recommended that the contamination of feed and water at the farm level and carcasses of slaughtered animals by L. monocytogenes be reduced through good sanitary practices at both levels to prevent the entry of the pathogen into the human food chain. It is also recommended that a comprehensive risk assessment of the three variables (area, type/size of farm/abattoir, and sample types) investigated in the current study and other variables be conducted using a higher number of farms and abattoirs across the country’s eight provinces, to determine their importance as risk factors for the occurrents of L. monocytogenes and other Listeria species on cattle farms and abattoirs, in the country.
Data availability
All the data are contained within the article.
References
Aiyedun, J.O., Olatoye, O.I., Oludairo, O.O., Adesope, A.O., Ogundijo, O., 2020. Occurrence, Antimicrobial Susceptibility and Biofilm Production in Listeria monocytogenes Isolated from Pork and other Meat Processing Items at Oko- Oba Abattoir, Lagos State, Nigeria. Sahel Journal of Veterinary Sciences, 17(4), pp.24-30.
Al, S., Disli, H.B., Hizlisoy, H., Ertas Onmaz, N., Yildirim, Y., Gonulalan, Z. 2022. Prevalence and molecular characterization of Listeria monocytogenes isolated from wastewater of cattle slaughterhouses in Turkey. Journal of Applied Microbiology, 132(2), 1518-1525
Allam, M., Tau, N., Smouse, S. L., Mtshali, P. S., Mnyameni, F., Khumalo, Z. T., Smith, A. M. 2018. Whole-genome sequences of Listeria monocytogenes sequence type 6 isolates associated with a large foodborne outbreak in South Africa, 2017 to 2018. Genome announcements, 6(25), e00538-18. https://doi.org/10.1128/genomeA.00538-18
Arslan, S. and Baytur, S., 2019. Prevalence and antimicrobial resistance of Listeria species and subtyping and virulence factors of Listeria monocytogenes from retail meat. Journal of Food Safety, 39(1), e12578. https://doi.org/10.1111/jfs.12578
Ayaz, N.D., Onaran, B., Cufaoglu, G., Goncuoglu, M., Ormanci, F.S., Erol, I., 2018. Prevalence and characterization of Listeria monocytogenes isolated from beef and sheep carcasses in Turkey with the characterization of locally isolated listeriophages as a control measure. Journal of Food Protection, 81(12), 2045-2053
Bandelj, P., Jamnikar‐Ciglenecki, U., Ocepek, M., Blagus, R., Vengust, M., 2018. Risk factors associated with fecal shedding of Listeria monocytogenes by dairy cows and calves. Journal of Veterinary Internal Medicine, 32(5), 1773-1779.
Bubert, A., Sokolovic, Z., Chun, S.K., Papatheodorou, L., Simm, A., Goebel, W., 1999. Differential expression of Listeria monocytogenes virulence genes in mammalian host cells. Molecular and General Genetics MGG, 261(2), 323-336.
Borucki, M.K., Gay, C.C., Reynolds, J., McElwain, K.L., Kim, S.H., Call, D.R., Knowles, D.P., 2005. Genetic diversity of Listeria monocytogenes strains from a high-prevalence dairy farm. Applied and Environmental Microbiology, 71(10), 5893-5899.
Castro, H., Jaakkonen, A., Hakkinen, M., Korkeala, H., Lindström, M., 2018. Occurrence, persistence, and contamination routes of Listeria monocytogenes genotypes on three Finnish dairy cattle farms: a longitudinal study. Applied and Environmental Microbiology, 84(4), e02000-17.
Chakraborty, T., Leimeister-Wächter, M., Domann, E., Hartl, M., Goebel, W., Nichterlein, T., Notermans, S., 1992. Coordinate regulation of virulence genes in Listeria monocytogenes requires the product of the prfA gene. Journal of Bacteriology, 174(2), 568-574.
Chapin, T.K., Nightingale, K.K., Worobo, R.W., Wiedmann, M., Strawn, L.K., 2014. Geographical and meteorological factors associated with the isolation of Listeria species in New York State produce production and natural environments. Journal of Food Protection, 77(11), 1919-1928.
Chand, P., Sadana, J. R., 1999. Outbreak of Listeria ivanovii abortion in sheep in India. The Veterinary Record, 145(3), 83-84.
Chuku, A., Obande, G.A., Eya, S.B., 2019. Listerial contamination of raw beef and chevon in north-central Nigeria. IMC Journal of Medical Science, 13(2), pp.1-8.
Demaître, N., Van Damme, I., De Zutter, L., Geeraerd, A.H., Rasschaert, G., De Reu, K., 2020. Occurrence, distribution, and diversity of Listeria monocytogenes contamination on beef and pig carcasses after slaughter. Meat Science, 169, 108177
Demaître, N., De Reu, K., Haegeman, A., Schaumont, D., De Zutter, L., Geeraerd, A., Rasschaert, G., 2021. Study of the transfer of Listeria monocytogenes during the slaughter of cattle using molecular typing. Meat Science, 175, 108450.
Dhama, K., Karthik, K., Tiwari, R., Shabbir, M.Z., Barbuddhe, S., Malik, S.V.S., Singh, R.K., 2015. Listeriosis in animals, its public health significance (food-borne zoonosis) and advances in diagnosis and control: a comprehensive review. Veterinary Quarterly, 35(4), 211-235.
Doumith, M., Buchrieser, C., Glaser, P., Jacquet, C., Martin, P., 2004. Differentiation of the major Listeria monocytogenes serovars by multiplex PCR. Journal of Clinical Microbiology 42, 3819-3822.
Driehuis, F., Wilkinson, J.M., Jiang, Y., Ogunade, I., Adesogan, A.T., 2018. Silage review: animal and human health risks from silage. Journal of Dairy Science, 101(5), 4093-4110.
Dunka, H. I., Bello, M., Lawan, M. K., 2021. Prevalence and antibiogram of Listeria monocytogenes contamination of liver, spleen, ruminal content and effluent in Jos, Nigeria. Journal of Veterinary Medicine and Animal Sciences, 4(1), 1072.
Ferreira, V., Wiedmann, M., Teixeira, P., Stasiewicz, M. J., 2014. Listeria monocytogenes persistence in food-associated environments: epidemiology, strain characteristics, and implications for public health. Journal of Food Protection, 77(1), 150-170.
Foerster, C., Vidal, L., Troncoso, M., Figueroa, G., 2012. Characterization of Listeria monocytogenes isolates from cattle and ground beef by pulsed-field gel electrophoresis. Revista Argentina de Microbiología, 44(3), 195-200.
Gana, J., Gcebe, N., Pierneef, R.E., Chen, Y., Moerane, R., Adesiyun, A.A., 2023. Genomic Characterization of Listeria innocua Isolates Recovered from Cattle Farms, Beef Abattoirs, and Retail Outlets in Gauteng Province, South Africa. Pathogens, 12(8), p.1062.
Gradovska, S., Šteingolde, Ž., Ķibilds, J., Meistere, I., Avsejenko, J., Streikiša, M., Alksne, L., Terentjeva, M., Bērziņš, A., 2023. Genetic diversity and known virulence genes in Listeria innocua strains isolated from cattle abortions and farm environment. Veterinary and Animal Science, 19, p.100276.
Huang, Y., Morvay, A. A., Shi, X., Suo, Y., Shi, C., Knøchel, S., 2018. Comparison of oxidative stress response and biofilm formation of Listeria monocytogenes serotypes 4b and 1/2a. Food Control, 85, 416-422
Hurtado, A., Ocejo, M., Oporto, B., 2017. Salmonella spp. and Listeria monocytogenes shedding in domestic ruminants and characterization of potentially pathogenic strains. Veterinary Microbiology, 210, 71-76
Hilliard, A., Leong, D., O’Callaghan, A., Culligan, E.P., Morgan, C.A., DeLappe, N., Hill, C., Jordan, K., Cormican, M., Gahan, C.G., 2018. Genomic characterization of Listeria monocytogenes isolates associated with clinical listeriosis and the food production environment in Ireland. Genes, 9(3), p.171
Jamali, H., Chai, L.C., Thong, K.L., 2013. Detection and isolation of Listeria spp. and Listeria monocytogenes in ready-to-eat foods with various selective culture media. Food Control, 32(1), 19–24.
Kaur, T., Balgir, P.P., 2021. Listeriosis in Animals: Prevalence and Detection. Advances in Animal Disease Diagnosis, pp.147–167.
Kastbjerg, V.G., Larsen, M.H., Gram, L., Ingmer, H., 2010. Influence of sublethal concentrations of common disinfectants on expression of virulence genes in Listeria monocytogenes. Applied and Environmental Microbiology, 76(1), 303-309.
Kayode, A. J., Okoh, A. I., 2022. Assessment of the molecular epidemiology and geneticmultiplicity of Listeria monocytogenes recovered from ready-to-eat foods following the South African listeriosis outbreak. Scientific Reports, 12(1), 20129.
Koopmans, M. M., Brouwer, M. C., Vázquez-Boland, J. A., van de Beek, D., 2023. Human listeriosis. Clinical Microbiology Reviews, 36(1), e00060-19.
Korsak, D., and Szuplewska, M., 2016. Characterization of nonpathogenic Listeria species isolated from food and food processing environment. International Journal of Food Microbiology, 238, 274-280.
Lopez-Valladares, G., Danielsson-Tham, M. L., Tham, W., 2018. Implicated food products for listeriosis and changes in serovars of Listeria monocytogenes affecting humans in recent decades. Foodborne pathogens and disease, 15(7), 387-397.
Lomonaco, S., Patti, R., Knabel, S.J., Civera, T., 2012. Detection of virulence-associated genes and epidemic clone markers in Listeria monocytogenes isolates from PDO Gorgonzola cheese. International Journal of Food Microbiology, 160(1), 76-79.
Manqele, A., 2018. Prevalence and characterization of Salmonella spp. in slaughter cattle, abattoir environment, and meat sold at retail outlets in Gauteng, South Africa. M.Sc. Thesis, University of Pretoria,
Matle, I., Mbatha, K. R., Lentsoane, O., Magwedere, K., Morey, L., Madoroba, E. 2019. Occurrence, serotypes, and characteristics of Listeria monocytogenes in meat and meat products in South Africa between 2014 and 2016. Journal of Food Safety, 39(4), e12629.
Mohammed, H.O., Atwill, E., Dunbar, L., Ward, T., McDonough, P., Gonzalez, R., Stipetic, K., 2010. The risk of Listeria monocytogenes infection in beef cattle operations. Journal of Applied Microbiology, 108(1), 349-356
Moura, A., Disson, O., Lavina, M., Thouvenot, P., Huang, L., Leclercq, A., Lecuit, M. 2019. Atypical hemolytic Listeria innocua isolates are virulent, albeit less than Listeria monocytogenes: Infection and Immunity, 87(4), e00758–18.
Mpundu, P., Muma, J. B., Mukumbuta, N., Mukubesa, A. N., Muleya, W., Kapila, P., Munyeme, M., 2022a. Isolation, discrimination, and molecular detection of Listeria species from slaughtered cattle in Namwala District, Zambia. BMC Microbiology, 22(1), 1-12.
Mpundu, P., Muma, J. B., Mukubesa, A. N., Kainga, H., Mudenda, S., Bumbangi, F. N., Munyeme, M., 2022b. Antibiotic Resistance Patterns of Listeria Species Isolated from Broiler Abattoirs in Lusaka, Zambia. Antibiotics, 11(5), 591.
McLauchlin, J., Grant, K.A., Amar, C.F.L., 2020. Human foodborne listeriosis in England and Wales, 1981 to 2015. Epidemiology & Infection, 148.
NicAogáin, K., O’Byrne, C.P., 2016. The role of stress and stress adaptations in determining the fate of the bacterial pathogen Listeria monocytogenes in the food chain. Frontiers in microbiology, 7, p.1865.
Nightingale, K.K., Schukken, Y.H., Nightingale, C.R., Fortes, E.D., Ho, A.J., Her, Z., Grohn, Y.T., McDonough, P.L., Wiedmann, M., 2004. Ecology and transmission of Listeria monocytogenes infecting ruminants and in the farm environment. Applied and Environmental Microbiology, 70(8), 4458-4467.
Obaidat, M.M. and Stringer, A.P., 2019. Prevalence, molecular characterization, and antimicrobial resistance profiles of Listeria monocytogenes, Salmonella enterica, and Escherichia coli O157: H7 on dairy cattle farms in Jordan. Journal of Dairy Science, 102(10), 8710-8720.
Obaidat, M.M., Kiryluk, H., Rivera, A., Stringer, A.P., 2020. Molecular serogrouping and virulence of Listeria monocytogenes from local dairy cattle farms and imported beef in Jordan. LWT, 127, p.109419
Oh, H., Kim, S., Lee, S., Lee, H., Ha, J., Lee, J., Coi, Y., Choi, K.H., Yoon, Y., 2016. Prevalence and genetic characteristics of meatborne Listeria monocytogenes isolates from livestock farms in Korea. Korean Journal for Food Science of Animal Resources, 36(6), 779-786.
Oh, H., Kim, S., Lee, S., Lee, H., Ha, J., Lee, J., Yoon, Y., 2018. Prevalence, serotype diversity,genotype and antibiotic resistance of Listeria monocytogenes isolated from carcasses and human in Korea. Korean Journal for Food Science of Animal Resources, 38(5), 851.
Onyeka, L.O., Adesiyun, A.A., Keddy, K.H., Madoroba, E., Manqele, A., Thompson, P.N., 2020. Shiga toxin–producing Escherichia coli contamination of raw beef and beef-based ready-to-eat products at retail outlets in Pretoria, South Africa. Journal of Food Protection, 83(3), 476-484.
Onyeka, L. O., Adesiyun, A. A., Keddy, K. H., Manqele, A., Madoroba, E., Thompson, P. N., 2021. Prevalence, risk factors and molecular characteristics of Shiga toxin-producing Escherichia coli in beef abattoirs in Gauteng, South Africa. Food Control, 123, 107746
Orsi, R.H., den Bakker, H.C., Wiedmann, M., 2011. Listeria monocytogenes lineages: genomics, evolution, ecology, and phenotypic characteristics. International Journal of Medical Microbiology, 301(2), 79-96
Palacios‐Gorba, C., Moura, A., Gomis, J., Leclercq, A., Gómez‐Martín, Á., Bracq‐Dieye, H., Mocé, M.L., Tessaud‐Rita, N., Jiménez‐Trigos, E., Vales, G., García‐Muñoz, Á., 2021. Ruminant‐associated Listeria monocytogenes isolates belong preferentially to dairy‐associated hypervirulent clones: A longitudinal study in 19 farms. Environmental Microbiology, 23(12), pp.7617-7631.
Perrin, M., Bemer, M., Delamare, C., 2003. Fatal case of Listeria innocua bacteremia. Journal of Clinical Microbiology, 41(11), 5308-5309. https://doi.org/10.1128/JCM.41.11.5308-5309.2003
Peng, X., Ed-Dra, A., Yue, M. 2022. Whole genome sequencing for the risk assessment of probiotic lactic acid bacteria. Critical Reviews in Food Science and Nutrition, 1–19. https://doi.org/10.1080/10408398.2022.2087174.
Pournajaf, A., Rajabnia, R., Sedighi, M., Kassani, A., Moqarabzadeh, V., Lotfollahi, L., Ardebilli, A., Emadi, B., Irajian, G., 2016. Prevalence, and virulence determination of Listeria monocytogenes strains isolated from clinical and non-clinical samples by multiplex polymerase chain reaction. Revista da Sociedade Brasileira de Medicina Tropical, 49, 624-627.
Queiroz, O.C.M., Ogunade, I.M., Weinberg, Z., Adesogan, A.T., 2018. Silage review: Foodborne pathogens in silage and their mitigation by silage additives. Journal of Dairy Science, 101(5), 4132-4142.
Rhoades, J. R., Duffy, G., Koutsoumanis, K. 2009. Prevalence and concentration of verocytotoxigenic Escherichia coli, Salmonella enterica and Listeria monocytogenes in the beef production chain: a review. Food Microbiology, 26(4), 357-376.
Rodriguez, C., Taminiau, B., García-Fuentes, E., Daube, G., Korsak, N., 2021. Listeria monocytogenes dissemination in farming and primary production: Sources, shedding and control measures. Food Control, 120, p.107540.
Ryu, J., Park, S.H., Yeom, Y.S., Shrivastav, A., Lee, S.H., Kim, Y.R., Kim, H.Y., 2013. Simultaneous detection of Listeria species isolated from meat processed foods using multiplex PCR. Food Control 32, 659-664
Soni, D. K., Singh, D. V., Dubey, S. K., 2015. Pregnancy-associated human listeriosis: Virulence and genotypic analysis of Listeria monocytogenes from clinical samples. Journal of Microbiology, 53(9), 653-660.
Soumet, C., Ermel, G., Fach, P., Colin, P., 1994. Evaluation of different DNA extraction procedures for the detection of Salmonella from chicken products by polymerase chain reaction. Letters in Applied Microbiology, 19(5), 294-298.
Stipetic, K., Chang, Y. C., Peters, K., Salem, A., Doiphode, S. H., McDonough, P. L., Mohammed, H. O. 2016. The risk of carriage of Salmonella spp. and Listeria monocytogenes in food animals in dynamic populations. Veterinary Medicine and Science, 2(4), 246–254.
Takahashi, T., Ochiai, Y., Matsudate, H., Hasegawa, K., Segawa, T., Fukuda, M., Hondo, R., Ueda, F., 2007. Isolation of Listeria monocytogenes from the skin of slaughtered beef cattle. Journal of Veterinary Medical Science, 69(10), 1077-1079.
Terentjeva, M., Šteingolde, Ž., Meistere, I., Elferts, D., Avsejenko, J., Streikiša, M., Gradovska, S., Alksne, L., Ķibilds, J. and Bērziņš, A. 2021. Prevalence, genetic diversity and factors associated with distribution of Listeria monocytogenes and other Listeria spp. in cattle farms in Latvia. Pathogens, 10(7), p.851.
Thrusfield, M. 2007. Sample size determination. Veterinary epidemiology, 3, 185-189.
Wang, Y., Jiao, Y., Lan, R., Xu, X., Liu, G., Wang, X., Zhang, L., Pang, H., Jin, D., Dai, H., Yuan, X., 2015. Characterization of Listeria monocytogenes isolated from human listeriosis cases in China. Emerging Microbes and Infections, 4(1), 1-3.
Wieczorek, K., Dmowska, K., Osek, J., 2012. Characterization and antimicrobial resistance of Listeria monocytogenes isolated from retail beef meat in Poland. Foodborne Pathogens and Disease, 9, 681-685.
Wiktorczyk-Kapischke, N., Skowron, K., Wałecka-Zacharska, E., 2023. Genomic and pathogenicity islands of Listeria monocytogenes—overview of selected aspects. Frontiers in Molecular Biosciences, 10, 1161486.
Zhao, Q., Hu, P., Li, Q., Zhang, S., Li, H., Chang, J., Jiang, Q., Zheng, Y., Li, Y., Liu, Z., Ren, H., 2021. Prevalence and transmission characteristics of Listeria species from ruminants in farm and slaughtering environments in China. Emerging Microbes and Infections, 10(1), 356-364.
Zweifel, C., Baltzer, D., Stephan, R. 2005. Microbiological contamination of cattle and pig carcasses at five abattoirs determined by swab sampling in accordance with EU Decision 2001/471/EC. Meat Science, 69(3), 559-566.
Acknowledgements
We are grateful to the Agricultural Research Council-Onderstepoort Veterinary Institute (ARC-OVI) technical staff (Kuda Jwamba, Makhado Lavhesani, Carol Matau, and Lesego Mashiane) for their assistance and the laboratory supervision provided by Dr. I. Matle during the study. The cooperation and help of fellow graduate students (Nduduzo Mtshali and Khomotso C. Moabelo) are appreciated. We are sincerely grateful to the owners and managers of the cattle farms and beef abattoirs, who, despite being in the middle of the global COVID-19 outbreak, facilitated our access to their facilities to collect samples for the study. The project would not have been possible without their cooperation and assistance.
Funding
Open access funding provided by University of Pretoria. This study was funded by the Red Meat Research and Development, South Africa (RMRD-SA) grant # 2018–12-20, December 2018, which enabled us to conduct the study.
Author information
Authors and Affiliations
Contributions
AAA and NG conceived and designed research. JG, TT, and KM collected samples, JG and NG conducted laboratory experiments, and YBN, JG, and AAA analyzed data. JG, AAA, and NG wrote the first draft of the manuscript. All authors (AAA, NG, JG, RM, YBN, TT, and KM) read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Ethical approval
Before the commencement of the study, approvals were obtained from the following bodies and committees: Research Ethics Committee (REC) of the Faculty of Veterinary Science, University of Pretoria, South Africa (REC 138–19), Animal Ethics Committee (AEC) of the Faculty of Veterinary Science, University of Pretoria, South Africa (REC 138–19), and Sect. 20 according to Act 35 of 1984 by the Director of Animal Health at the Department of Agriculture, Forestry and Fisheries (DAFF), [Number: 12/11/1/1/8(1131)] South Africa.
Informed consent
Managers or owners of the cattle farms and beef abattoirs from where cattle and beef carcasses and environmental samples were collected for the study consented.
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gana, J., Gcebe, N., Moerane, R. et al. A comparative study on the occurrence, genetic characteristics, and factors associated with the distribution of Listeria species on cattle farms and beef abattoirs in Gauteng Province, South Africa. Trop Anim Health Prod 56, 88 (2024). https://doi.org/10.1007/s11250-024-03934-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11250-024-03934-y