Abstract
The curation and preservation of Dutch apple germplasm depends on reliable accession level information. However, many accessions of Dutch heirloom apple cultivars maintained publicly by the “Centre for Genetic Resources, the Netherlands” (CGN) and privately by Dutch pomological societies lack information regarding true-to-typeness and pedigree ancestry. The aim of this study was to address this knowledge gap by genotyping 652 apple accessions maintained in the CGN collection and Dutch private collections, compare their genotypic information to each other and to a large database of apple cultivars from around the world to identify genotypic duplicates and pedigree relationships for the Dutch apple cultivars. Towards this aim, accessions were genotyped on the 20 K Illumina Infinium(R) apple SNP array and with 15 SSR markers. Each accession was assigned to a genotypic profile code (MUNQ codes, as used in previous studies) facilitating communication regarding genotypic duplicates. There were 211 (51.1%) genotypic profiles in the Dutch germplasm which were not identified in other germplasm collections. Private collections maintained many of these unique accessions, including important pedigree ancestors. The study identified a number of common pedigree ancestors of Dutch cultivars, such as ‘Herfst Bloem Soete’, ‘Huismanszoet’ (2), and ‘Reinette Rouge Étoilée’. The duplicate and pedigree reconstruction results and relevant literature descriptions were used to pomologically verify the identity of relevant accessions. The results of this study resolved identity disputes, helped to decide which accessions should be retained or included in the CGN collection, and benefited ongoing pomological studies in ancestry and provenance of traditional Dutch cultivars.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Apples (Malus domestica Borkh.) have been an important and celebrated part of the culture in areas where they have been grown since times immemorial. In the past, each apple growing region had traditional cultivars that were suited to the local growing conditions and for specific culinary end uses. However, in recent times, production and consumption worldwide have largely relied on the production of a limited number of closely interrelated cultivars (Bannier 2011; Migicovsky et al. 2021), used primarily for fresh eating. This change in production and consumption habits has resulted in a reduction in the growing of traditional cultivars, which in turn poses risks to the loss of both cultural traditions and adapted germplasm (Fowler and Mooney 1990; Goland and Bauer 2004).
To combat these risks, private and public organizations have made efforts to conserve traditional cultivars in germplasm collections and to make them available for use in breeding, for genetic studies, and for aiding in the preservation and passing on of cultural traditions (Volk and Bramel 2021). However, information about the identities, origins, and relationships of accessions held in these collections is often either unknown or has not been validated, resulting in a lack of information about the interrelations between accessions and which are duplicates. This lack of accession-level information hampers effective and efficient germplasm curation, consequently hindering effective curation and limiting the end use potential of available germplasm.
In the Netherlands, historical and regionally important cultivars are maintained in collections publicly at the Wageningen University and Research “Centre for Genetic Resources, the Netherlands” (CGN) and privately by Dutch pomological societies such as the Pomologische Vereniging Noord-Holland (POMVNH) and the Noordelijke Pomologische Vereniging (NPV). The goal of the CGN collection is to maintain and make publicly available the most important cultivars with recorded origin or historical significance in the Netherlands (e.g. Knoop 1758; Noort 1830; Berghuis and van Hall 1868). To meet this goal, the CGN has partnered with the Dutch pomological society and the Nederlands Fruit Netwerk for pomological verification of accessions in the CGN collection, determining true-to-typeness, and to select cultivars suitable for future conservation in the CGN collection.
To assist curation, conservation, and pomological efforts, accessions in the CGN collection and accessions in the Pomologische Vereniging Noord-Holland, the Noordelijke Pomologische Vereniging, and several private Dutch collection orchards were previously genotyped with a set of 16 single sequence repeat (SSR) markers (van Treuren et al. 2010). That study identified many duplicates and helped resolve some identity disputes but had two major limitations. First, the SSRs chosen for use did not enable sufficient comparison to collections held in other countries. A large-scale cross-collection comparison enabled by an SSR genotyping project initially described by (Urrestarazu et al. 2016). Those efforts subsequently resulted in the ongoing Malus UNiQue genotype (MUNQ) coding system (Denancé et al. 2020; Muranty et al. 2020), enabling an improved curation of the collections involved. However the CGN collection has so far not been connected to this project. Second, the use of SSRs in genotypic audits of apple collections have more recently been superseded by the use of single nucleotide polymorphism (SNP) marker array technology. SNP arrays offer a more standardized set of markers for conducting duplicate analyses and the resulting genotypic data is more readily integratable across studies. Additionally, SNP array data can be used for more accurate and detailed pedigree reconstruction (e.g., as in Muranty et al. 2020; Skytte af Sätra et al. 2020; Luby et al. 2022), which is presently being compiled in a large collaborative study (Howard et al. 2018). Pedigree reconstruction results can be used to resolve cultivar identity disputes and uncover previously unknown histories of cultivars. In the context of the goals of the CGN, such pedigree information could be used to validate or identify which cultivars held in private collections are particularly worthy of inclusion in the CGN collection to ensure preservation of Dutch apple heritage.
The aim of the current study was to genotype apple accessions in the CGN collection and various Dutch pomological group collections and compare them to each other and to a large database of cultivars. This was done to facilitate the ongoing efforts of the CGN and Dutch pomological groups to curate their collections effectively and to build a centralised collection of traditional Dutch cultivars to be preserved at the CGN. Specifically, accessions were assigned MUNQ codes to facilitate communication regarding genotypic duplicates, pedigree reconstruction was performed on each unique genotypic profile, relevant literature descriptions were used to pomologically verify the identity of accessions, and the resulting data was used to help characterize accessions and resolve identity disputes, which in turn was used to decide which accessions should be included in the CGN collection.
Materials and methods
Plant material
A set of 652 apple accessions was genotyped in this study, which included 196 accessions from the CGN, and 456 accessions selected by the Dutch pomological societies based on their accession names in various collections maintained by pomological societies and private persons in the Netherlands (Table S1). The samples were harvested as young leaves, which were immediately placed on silica gel and thereafter freeze dried.
Genotyping
DNA extraction was performed directly in 96-well plates in which the dry leaf samples were initially grinded on a ball mill before manually performing the extraction using the Qiagen DNeasy Plant Mini Kit (Qiagen®, Hilden, Germany) following the manufacturer’s protocol. The 96-well plates with DNA aliquots were used for both SSR and SNP genotyping. A set of 16 SSR markers (Urrestarazu et al. 2016) were used for SSR genotyping. The use of this set allowed for a comparison of all genotypic profiles in the Dutch dataset to genotypic profiles in the SSR genotyping project (Denancé et al. 2020; Muranty et al. 2020) and ascribe each accession to a MUNQ code.
SNP genotyping was performed using the 20 K Infinium® apple SNP array (Bianco et al. 2014) according to the standard Illumina protocol (Chagné et al. 2012). The iScan data was analysed in GenomeStudio (GS) v2.0 (Illumina Inc.) as described by Vanderzande et al. (2019), with the modifications and cluster definitions identified by Howard et al. (2021c). The genetic positions for the “robust” SNPs identified by Howard et al. (2021c) were used and included the set of 10,321 SNPs used by Volk et al. (2022). All genotypic profiles were compared to those in the collaborative large-scale apple pedigree reconstruction project (Howard et al. 2018). Accessions from that project that were relevant to this study were recorded (Table S2).
SNP data processing and pedigree reconstruction
The intensity data was evaluated in GS utilizing the B-allele frequency to ensure that all samples were meeting the quality criteria defined by Vanderzande et al. (2019). Genotypic duplicates were considered true for accessions with more than 99.5% identical calls for the highly curated SNP calls obtained though the pedigree reconstruction project. This threshold was deemed appropriate given the high repeatability previously reported for this array and set of SNPs (Howard et al. 2021c). The ploidy level of each accession was determined accessing the B-allele frequency in GS as described by Chagné et al. (2015).
Pedigree reconstruction for diploid accessions was conducted as described in Vanderzande et al. (2019); Howard et al. (2017b, 2021b), whereas pedigree reconstruction involving triploid accessions was conducted as described by Howard et al. (2023). Parent-offspring duo relationships were ordered utilizing the parent-offspring order resolution (POR) tests, POR-1 and POR-2 as previously described (Howard et al. 2022). In the large-scale, ongoing pedigree reconstruction project being conducted concurrent with this study, dummy parents were recorded and in some cases imputed following the methods of Howard et al. (2017a, 2021b, 2023). These dummy parents were given names, reflecting the nature of their relationships. Dummy parents that were identified as being the offspring of two previously existing genotypic profiles and being the parent of less than three individuals were not imputed and given the prefix “UP_”, standing for “Unknown Parent”, followed by the name of their identified offspring. Dummy parents that were deduced unreduced gamete donating parents of triploids (Howard et al. 2022), and the parent of less than three diploids (or the reduced gamete donating parent of triploids) were partially imputed via the homozygous calls of their triploid offspring. These were given the name “UGDP_”, standing for “Unreduced Gamete Donating Parent”, followed by the name of their triploid offspring. Dummy parents that had at least five offspring or that were the unreduced gamete donating parent of at least two triploids and that also had diploid offspring were partially to fully imputed. These were given the name “Unknown_Founder_X”, where “X” was a sequential number, following the first “Unknown Founder” genotypically described in Howard et al. (2021b).
Following pedigree reconstruction, a pedigree network was generated using the ggraph (Pedersen 2024) package in R (R core team 2024).
Evaluating true-to-typeness
The true-to-typeness of each accession was evaluated by the Dutch pomological societies utilizing duplicate and pedigree reconstruction results in combination with passport data, pedigree records, and pomological cultivar descriptions in the literature (e.g. Knoop 1758; Noort 1830). In addition, morphological traits in the field collections were, as far as was possible, compared with pomological cultivar descriptions for evaluating the true-to-typeness. Based on these evaluations, each genotypic profile was given a “preferred name” that best suited the accession/cultivar’s provenance (see Table S3). In case a Dutch accession name did not match any cultivar description in the literature, the identity of a genotype was in some cases considered resolved if the same genotype was found under another name in another collection that had been pomologically verified.
Results
Genotypic duplicates
The 652 genotyped accessions represented 413 unique genotypic profiles (i.e., 413 distinct MUNQ codes; Table S3). The genotypic profiles were identified and distinguished equally efficiently with the 16 SSRs and with the 10 K SNPs. There were 283 genotypes that were only represented by one accession in the Dutch dataset. The duplicate groups represented 130 genotypes including 11 groups that were represented by five or more accessions. There were 211 unique genotypic profiles in this study that were not previously represented in the MUNQ database or the apple pedigree reconstruction project.
Of the 413 unique genotypic profiles from this study, 236 (57.4%) were not present in the CGN germplasm collection. The 196 accessions from the CGN collection represented 177 genotypic profiles, of which 88 were not present in other Dutch collections. There were 16 duplicate groups in the CGN collection which were each represented by at least two accessions. There were few examples in which identical genotypic profiles with distinct accession names represented selected colour sports, such as ‘Gronsvelder Klumpke’, which is red sport of ‘Rheinischer Krummstiel’.
True-to-typeness analysis
There were 107 accessions that were confidently deemed not true to type, of which 20 accessions were obtained from the CGN collection (Table S1). In some cases, the identity was resolved using information about genotypic duplicates (MUNQs) in other collections, two of these being the rootstock cultivars M.26 and MM.111. In these two cases, the original scions probably had died but the rootstocks lived and grew leaves, which were sampled. There were six accessions with different names that each matched the genotypic profile of ‘Dubbele Zoete Aagt’ (MUNQ 293), which similarly may have been due to the death of scions, as ‘Dubbele Zoete Aagt’ has sometimes been used as an interstock. In other cases, such as for accession Ros04-109, the accession’s identity was previously unknown and was assigned to a genotypic profile group temporarily named ‘Hollandse Bellefleur’, though the correct name and provenance is currently under investigation. Pedigree reconstruction results were useful for resolving a number of identity disputes or for classifying accessions as not being true to type, for example for the accessions BG8Ap015, NPV-K14, and NPV-T39 where the SNP verified pedigrees were inconsistent with records or due to inconsistencies in the seniority between parent and offspring.
Pedigree reconstruction
Both extant parents were identified for 148 (35.8%) and one parent was identified for 106 (25.7%) of the 413 unique genotypic profiles, excluding dummy parents (Table S3). One genotypic profile had two unknown, partially imputed genotypic profiles as parents and 38 genotypic profiles had one unknown, partially imputed genotypic profile as one of their parents. None of the parents were identified for 159 (38.5%) of the genotypic profiles.
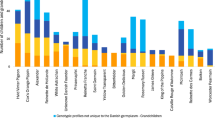
Numerous common pedigree ancestors of the individuals were identified (Table 1; Fig. 1). Some of the most common pedigree ancestors are commonly found in the pedigrees of modern commercial cultivars, such as ‘Cox’s Orange Pippin’, ‘Jonathan’, and ‘Golden Delicious’, while other common pedigree ancestors are older, regionally important cultivars that are not in the pedigrees of modern commercial cultivars. Some of these had been genotyped in previous studies, such as ‘Brabantse Bellefleur’, ‘Reinette Rouge Étoilée’, ‘Groninger Kroon’, and ‘Keswick Codlin’.
Relationship network diagram depicting parent-offspring relationships among genotypic profiles of Dutch accessions evaluated. Arrows depict the direction of the relationship. Squares represent genotypic profiles present among the Dutch accessions sampled. Triangles represent genotypic profiles that were not present among the Dutch accessions sampled. Circles represent imputed and ungenotyped (dummy) profiles. Lighter red points with numbers represent genotypic profiles with at least six offspring and the numbers correspond to those noted in Table 1. Offspring of Dutch genotypic profiles that were not present in Dutch collections and genotypic profiles lacking identified pedigree relationships were not included in this figure
Some important pedigree ancestors of Dutch germplasm, such as ‘Herfst Bloem Soete’ and ‘Huismanszoet’ (2) were SNP and SSR genotyped for the first time in this study. ‘Herfst Bloem Soete’ was identified as the parent of nine genotypic profiles and a grandparent of 20 genotypic profiles in Dutch germplasm (Fig. 2; Table S3; Table S4). It was also the parent and a parent of ‘Broholm Rosenæble’ from The Pometum (accession DNK0017, MUNQ 5954), ‘Krügers Dickstiel’ from the Ökowerk collection in Germany (accession Oek_330-1, MUNQ 1009), and ‘Reinette Rouge Étoilée’, which itself had 12 direct offspring in the Dutch germplasm.
Discussion
The results of this study were successfully used to improve curation of the CGN collection by removing genotypic duplicates, updating the identities of accessions deemed not true to type, and identifying culturally or genotypically significant Dutch cultivars in the CGN collection. The assignment of MUNQ codes to each accession will be useful in facilitating communication regarding genotypic duplicates within and between collections. Pedigree reconstruction results were used to characterize accessions, verify true-to-typeness, and help identify several previously unknown pedigree ancestors of Dutch heritage cultivars.
Curation of apple collections
The results of the genotypic duplicate information from this study are useful in aiding current ongoing curation efforts in the CGN collection and private collections. The CGN collection had a relatively low number of genotypic duplicates (9.7%) compared to other North European collections of historical apple cultivars, such as The Pometum collection in Denmark (23%) (Larsen et al. 2017) or the Swedish Central Collection (14%) (Skytte af Sätra et al. 2020). The relatively low number of genotypic duplicates in the CGN collection was likely due to the fact that several duplicates were previously identified and removed from the CGN collection by utilizing genotypic information obtained through SSR genotyping the GGN accessions previously performed (van Treuren et al. 2010). Most of the current duplicates were from newer accessions in the CGN collection that had not been genetically evaluated since that earlier study. These newer results thus will further help remove unnecessary duplication. The efforts of van Treuren et al. (2010) were useful for identifying genotypic duplicates in the CGN collection but did not allow to compare CGN accessions with accessions from other collection sites. Similar efforts performing SSR or SNP based fingerprinting or pedigree reconstruction based on accessions from a single genebank collections been done in various other North-European apple genebank collections (e.g. Larsen et al. (2017); Skytte af Sätra et al. (2020); Gilpin et al. (2023).
The current study allowed for the first time genotypic comparison of CGN accessions with accessions from other germplasm collections in The Netherlands as well as other parts of the world. Similar efforts utilizing genotypic information on apple germplasm across collection sites including SNP-based pedigree reconstruction was also performed by Muranty et al. (2020) and Luby et al. (2022). However, this is the first study ascribing MUNQ codes to the complete set of accessions maintained in a genebank collection and performing pedigree reconstruction utilizing all genotypic profiles represented in the genebank and in the pedigree reconstruction project. The MUNQ and pedigree reconstruction results were in several cases useful to verify cultivar identities, to solve accession identity discrepancies in the CGN collection, reveal previously unknown pedigree relations, or to highlight important Dutch germplasm from other collection sites which were not present in the CGN collection.
Duplicates between the CGN and the private Dutch collections helped resolve many identity disputes in cases where cultivar descriptions in the literature were either non-existing or too poor to be used for identity verification. Examples of these identity dispute resolutions include the CGN accessions ‘Zure Renette’ (GEN061), which matched the genotypic profile of ‘Brabantse Bellefleur’, ‘Pomme d’Orange’ (GEN277) matching the genotypic profile of ‘Zigeunerin’, and ‘Spekappel’ (GEN197) matching the genotypic profile of ‘Notarisappel’. More than half (57.4%) of the genotypes identified in this study were not present in the CGN collection, including the historically important Dutch cultivar ‘Schone van Boskoop’ and the three important Dutch pedigree ancestor cultivars, ‘Herfst Bloem Soete’, ‘Huismanszoet’ (2), and ‘Reinette de Hollande’. These three pedigree ancestors were only identified in private Dutch collections and are now being propagated to be preserved in the CGN collection. Some CGN accessions were labelled with a Dutch accession name but were genotypically identical to accessions in other collections and were not considered to be Dutch traditional cultivars. For example, the accession ‘Zoete Crombach b1’ (GEN047) which was pomologically verified as ‘Pomme d’Or’ (MUNQ 618) in collaboration with the Frensh pomologist Henri Fourey from the Croqueurs de Pommes.
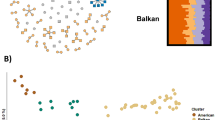
Integrating the genotypic profiles of Dutch germplasm into the databases the MUNQ project and the pedigree reconstruction project were also very useful for resolving identities or identifying accessions that were not true to type. For example, the CGN accession GEN083, named ‘Zaailing de Jongh’, was identical to ‘Geflammter Kardinal’ (MUNQ 770), a German cultivar that was sampled for the project described in Howard et al. (2018) from the private orchard of noted German pomologist Hans Joachim Bannier, who had pomologically verified the accession. In another example, three genotypically identical accessions labelled ‘Stichtsche Bellefleur’ (AK502), ‘Zoete Rode Ossekop’ (FF53), and ‘Jan Steen’ (Ros14-022) (Table S1) were found to be identical to the American cultivar Spartan (MUNQ 309), of which accessions exist from the Julius Kühn Institute (accession DEU_JKI_MD_0105) and the USDA collection in the USA (PI_588871). Another example is the CGN accession, ‘Zure van Driebergen’ (GEN076). This accession name does not refer to any literature description, but the identity was resolved through the identification of a genotypically identical accession held in the Danish germplasm collection “The Pometum” named ‘Hvid Vinter Pigeon’ (MUNQ 1509) pomologically described by Bredsted (1893). ‘Hvid Vinter Pigeon’ was found to be a pedigree ancestor of two genotypic profiles in the Dutch dataset.
True-to-typeness validation via pedigree reconstruction
The pedigree reconstruction results were useful in aiding ongoing germplasm collection curation efforts. Some accessions were deemed not true to type due to their pedigrees not matching historical records. For example, accession NPV-K18, recorded as the name ‘Zoete Holaart’, was determined not to be true to type because its identified parent, ‘Lunterse Pippeling’, is younger than the pedigree records, whereas the identical accessions AK506 and GEN267 were considered to truly represent ‘Zoete Holaart’ because their pedigrees matched pomological records. Another example involves the true identity of ‘Du Halder’. Two accessions named ‘Du Halder’, Ros11-044 (MUNQ 9674) and Ros20-008 (MUNQ 9683), did not match pedigree records whereas a third accession (Gen212, MUNQ 5473, also named ‘Du Halder’) was consistent with pedigree and pomological records (Berghuis and van Hall 1868).
The SNP verified pedigrees were also used to pinpoint other incorrect accession names where the SNP validated pedigrees did not match pedigree records. The genotypic profile MUNQ 1894 existed under the two accession names ‘Rubens 40/37’ at the CGN and ‘Prins Bernhard’ at the NPV. MUNQ 1894 was also identified as ‘Rubens’ in the National Fruit Collection (NFC) in the UK and has the SNP validated pedigree ‘Cox’s Orange Pippin’ × ‘Reinette Rouge Étoilée’. This pedigree is matching the recorded pedigree of ‘Rubens’, a cultivar released from the Institute for Horticultural Plant Breeding in Wageningen, The Netherlands. The correct identity of MUNQ 1894 was therefore concluded to be ‘Rubens’.
The pedigree reconstruction results enabled true-to-typeness verification for five cultivars developed by the Dutch Laboratory for Horticultural Cultivation, which are held in the CGN. The SNP verified pedigrees were consistent with the pedigree records for ‘Koningin Juliana’ (Reinette Rouge Étoilée’ × ‘Cox’s Orange Pippin’), for ‘Prinses Beatrix’ and ‘Prinses Margriet’ (‘Cox’s Orange Pippin’ × ‘Jonathan’) and ‘Prinses Irene’ and ‘Prinses Marijke’ (‘Jonathan’ × ‘Cox’s Orange Pippin’). An accession name inconsistency was found for the two CGN accessions ‘Prins Bernhard’ (GEN143) and ‘Lucullus’ (GEN284), which were identical and assigned to MUNQ 335. The recorded pedigrees for both accessions were ‘Jonathan’ × ‘Cox’s Orange Pippin’, which matched the pedigree reconstruction results for MUNQ 335. Thus, it was not possible to determine whether ‘Prins Bernhard’ or ‘Lucullus’ was the correct accession name for MUNQ 335.
Highlighting accessions with identity disputes
The results of this study also highlighted many identity disputes and uncovered previously unknown identity inconsistencies. For instance, there are two genotypic profiles that are currently named ‘Hollandsche Bellefleur’ (two accessions) and ‘Hollandse Bellefleur’ (five accessions). These are alternate spellings of what were thought to be the same cultivar but were found to have different genotypic profiles. ‘Hollandsche Bellefleur’ is a SNP verified diploid and ‘Hollandse Bellefleur’ is a SNP verified triploid. Additional pomological work would be needed to resolve this dispute.
There was also identify uncertainty uncovered for 10 pairs of accessions that had the same accession names but different genotypic profiles: ‘Huismanszoet’ (1) and ‘Huismanszoet’ (2), ‘Peterselieappel’ (1) and ‘Peterselieappel’ (2), ‘Rode Kroonsappel’ (1) and ‘Rode Kroonsappel’ (2), ‘Roem van Dekker’ (1) and ‘Roem van Dekker’ (2), ‘Roosappel’ (1) and ‘Roosappel’ (2), ‘Witte Wijnappel’ (1) and ‘Witte Wijnappel’ (2), ‘Zoete Kroon’ (1) and ‘Zoete Kroon’ (2), ‘Zoete Pippeling’ (1) and ‘Zoete Pippeling’ (2), ‘Zoete Princesse Noble’ (1) and ‘Zoete Princesse Noble’ (2), and ‘Zoete van de Bent’ (1) and ‘Zoete van de Bent’ (2). In each of these cases, the true identity may never be determined, owing to vague historical pomological descriptions and inconsistent naming structures used by pomologists and fruit growers.
Pedigree ancestors of Dutch apple cultivars
The study allowed for the identification of some previously unknown common pedigree ancestors of Dutch heritage cultivars. The traditional Dutch cultivar, ‘Herfst Bloem Soete’, which was included in the Dutch pomology by Knoop (1758), was identified for the first time as a major ancestor of Dutch cultivars. This cultivar was parent of the historically important ‘Reinette Rouge Étoilée’, which had 19 direct or multigeneration descendants in the Dutch germplasm. Some of these descendants originated from breeding efforts performed after the 1930s at the Dutch Laboratory for Horticultural Cultivation, which used ‘Reinette Rouge Étoilée’ as a parent (Vermeulen 1962). Another newly identified common pedigree ancestor was ‘Huismanszoet’ (2), which was a parent of seven cultivars, including heirloom cultivars such as ‘Anjelierappel’ and ‘Jasappel’. According to literature descriptions this could be the correct ‘Huismanszoet’, even though the available cultivar descriptions are relatively vague. ‘Reinette de Hollande’ had 27 first- or second-generation descendants in the Dutch germplasm. The cultivar was identified as an important parent to a large assemble of European germplasm by Muranty et al. (2020), who previously identified six of the nine direct pedigree relations found in this study. The name ‘Reinette de Hollande’ suggests a Dutch origin, though the cultivar is rarely grown in The Netherlands and its origin is unknown.
The two most common parents in Dutch germplasm, ‘Cox’s Orange Pippin’ and ‘Jonathan’ have been highly popular cultivars for more than a century in north-western and central Europe where they have been extensively included in breeding programmes (Smith et al. 1971; Bannier 2011). ‘Cox’s Orange Pippin’ was also found to be the most common founder in The Pometum collection in Denmark (Larsen et al. 2017) and in the Ökowerk collection in Northern Germany (Howard et al. 2021a). Other common parents of Dutch germplasm such as ‘Brabantse Bellefleur’, ‘Groninger Kroon’, and ‘Reinette Rouge Étoilée’ were more specific ancestors of the Dutch heritage material and had little or no importance as parents in The Pometum and Ökowerk collections. ‘Kasseler Renette’ and ‘Keswick Codlin’ which were identified as pedigree ancestors by Muranty et al. (2020) were also common parents of Dutch germplasm.
These pedigree reconstruction results provide new information about pedigree ancestors and provenance of Dutch apple germplasm. The results will benefit the preservation of Dutch heritage cultivars and allow for the improvement of the curation of the CGN collection.
References
Bannier H-J (2011) Moderne apfelzüchtung: Genetische verarmung und tendenzen zur inzucht. Erwerbs-Obstbau 52:85–110
Berghuis S, van Hall HC (1868) De Nederlandsche boomgaard. J.B. Wolters
Bianco L, Cestaro A, Sargent DJ, Banchi E, Derdak S, Di Guardo M, Salvi S, Jansen J, Viola R, Gut I, Laurens F, Chagné D, Velasco R, van de Weg E, Troggio M (2014) Development and validation of a 20K single nucleotide polymorphism (SNP) whole genome genotyping array for Apple (Malus × Domestica Borkh). PLoS ONE 9:e110377. https://doi.org/10.1371/journal.pone.0110377
Bredsted HC (1893) Haandbog i dansk Pomologi, 2. æbler. Hempelske Bog- og Papirhandels Forlag, Odense
Chagné D, Crowhurst RN, Troggio M, Davey MW, Gilmore B, Lawley C, Vanderzande S, Hellens RP, Kumar S, Cestaro A, Velasco R, Main D, Rees JD, Iezzoni A, Mockler T, Wilhelm L, van de Weg E, Gardiner SE, Bassil N, Peace C (2012) Genome-wide SNP detection, validation, and development of an 8K SNP array for Apple. PLoS ONE 7. https://doi.org/10.1371/journal.pone.0031745
Chagné D, Kirk C, Whitworth C, Erasmuson S, Bicknell R, Sargent DJ, Kumar S, Troggio M (2015) Polyploid and aneuploid detection in apple using a single nucleotide polymorphism array. Tree Genet Genomes 11:1–6
Denancé C, Muranty H, Durel C-E (2020) MUNQ - Malus UNiQue genotype code for grouping apple accessions corresponding to a unique genotypic profile. V1 edn. Portail Data INRAE. https://doi.org/10.15454/HKGMAS
Fowler C, Mooney PR (1990) Shattering: food, politics, and the loss of genetic diversity. University of Arizona
Gilpin L, Røen D, Schubert M, Davik J, Rumpunen K, Gardli KA, Hjeltnes SH, Alsheikh M (2023) Genetic characterization of the Norwegian Apple Collection. Horticulturae 9:575
Goland C, Bauer S (2004) When the apple falls close to the tree: local food systems and the preservation of diversity. Renewable Agric Food Syst 19:228–236
Howard NP, van de Weg E, Bedford DS, Peace CP, Vanderzande S, Clark MD, Teh SL, Cai L, Luby JJ (2017a) Elucidation of the ‘Honeycrisp’ pedigree through haplotype analysis with a multi-family integrated SNP linkage map and a large apple (Malusxdomestica) pedigree-connected SNP data set. Hortic Res 4:17003. https://doi.org/10.1038/hortres.2017.3
Howard NP, van de Weg E, Tillman J, Tong CBS, Silverstein KAT, Luby JJ (2017b) Two QTL characterized for soft scald and soggy breakdown in apple (Malus × Domestica) through pedigree-based analysis of a large population of interconnected families. Tree Genet Genomes 14:2. https://doi.org/10.1007/s11295-017-1216-y
Howard NP, Albach DC, Luby JJ (2018) The identification of apple pedigree information on a large diverse set of apple germplasm and its application in apple breeding using new genetic tools. Foerdergemeinschaft Oekologischer Obstbau e. V. (FOEKO), Weinsberg, pp 88–91
Howard NP, Luby JJ, van de Weg E, Durel CE, Denancé C, Muranty H, Larsen B, Troggio M, Ristel M, Albach DC (2021a) Applications of SNP-based apple pedigree identification to regionally specific germplasm collections and breeding programs, 1307 edn. International Society for Horticultural Science (ISHS), Leuven, Belgium, pp 231–238
Howard NP, Peace C, Silverstein KAT, Poets A, Luby JJ, Vanderzande S, Durel C-E, Muranty H, Denancé C, van de Weg E (2021b) The use of shared haplotype length information for pedigree reconstruction in asexually propagated outbreeding crops, demonstrated for apple and sweet cherry. Hortic Res 8:202. https://doi.org/10.1038/s41438-021-00637-5
Howard NP, Troggio M, Durel C-E, Muranty H, Denancé C, Bianco L, Tillman J, van de Weg E (2021c) Integration of Infinium and Axiom SNP array data in the outcrossing species Malus × Domestica and causes for seemingly incompatible calls. BMC Genomics 22:246. https://doi.org/10.1186/s12864-021-07565-7
Howard NP, van de Weg E, Luby JJ (2022) A new method to reconstruct the direction of parent-offspring duo relationships using SNP array data and its demonstration on ancient and modern cultivars in the outcrossing species malus × Domestica. Hortic Res 9. https://doi.org/10.1093/hr/uhab069
Howard NP, Micheletti D, Luby JJ, Durel CE, Denancé C, Muranty H, Ordidge M, Albach DC (2023) Pedigree reconstruction for triploid apple cultivars using single nucleotide polymorphism array data. Plants, People, Planet 5:98–111
Knoop JH (1758) Pomologia: dat is, Beschryvingen en afbeeldingen van de beste soorten van appels en peeren, welke in Nederen Hoog-Duitsland, Frankryk, Engelland en elders geagt zyn, en tot dien einde gecultiveert worden. Ferwerda
Larsen B, Toldam-Andersen TB, Pedersen C, Ørgaard M (2017) Unravelling genetic diversity and cultivar parentage in the Danish apple gene bank collection. Tree Genet Genomes 13:14. https://doi.org/10.1007/s11295-016-1087-7
Luby JJ, Howard NP, Tillman JR, Bedford DS (2022) Extended pedigrees of apple cultivars from the University of Minnesota breeding program elucidated using SNP array markers. HortScience 57:472–477
Migicovsky Z, Gardner KM, Richards C, Thomas Chao C, Schwaninger HR, Fazio G, Zhong G-Y, Myles S (2021) Genomic consequences of apple improvement. Hortic Res 8
Muranty H, Denancé C, Feugey L, Crépin J-L, Barbier Y, Tartarini S, Ordidge M, Troggio M, Lateur M, Nybom H, Paprstein F, Laurens F, Durel C-E (2020) Using whole-genome SNP data to reconstruct a large multi-generation pedigree in apple germplasm. BMC Plant Biol 20:2. https://doi.org/10.1186/s12870-019-2171-6
Noort Mv (1830) Pomologia Batava van Noort
Pedersen T (2024) ggraph: An Implementation of Grammar of Graphics for Graphs and Networks. R package version 2.2.1.9000. https://github.com/thomasp85/ggraph
R Core Team (2024) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Skytte af Sätra J, Troggio M, Odilbekov F, Sehic J, Mattisson H, Hjalmarsson I, Ingvarsson P, Gustavsson L (2020) Genetic status of the Swedish Central collection of heirloom apple cultivars. Sci Hort 272:109599. https://doi.org/10.1016/j.scienta.2020.109599
Smith MWG, Great Britain. Ministry of, Agriculture F (1971) Food National Apple Register of the United Kingdom. Ministry of Agriculture, Fisheries and Food
Urrestarazu J, Denance C, Ravon E, Guyader A, Guisnel R, Feugey L, Poncet C, Lateur M, Houben P, Ordidge M, Fernandez-Fernandez F, Evans KM, Paprstein F, Sedlak J, Nybom H, Garkava-Gustavsson L, Miranda C, Gassmann J, Kellerhals M, Suprun I, Pikunova AV, Krasova NG, Torutaeva E, Dondini L, Tartarini S, Laurens F, Durel CE (2016) Analysis of the genetic diversity and structure across a wide range of germplasm reveals prominent gene flow in apple at the European level. BMC Plant Biol 16:130. https://doi.org/10.1186/s12870-016-0818-0
van Treuren R, Kemp H, Ernsting G, Jongejans B, Houtman H, Visser L (2010) Microsatellite genotyping of apple (Malus domestica Borkh.) Genetic resources in the Netherlands: application in collection management and variety identification. Genet Resour Crop Evol 57:853–865
Vanderzande S, Howard NP, Cai L, Da Silva Linge C, Antanaviciute L, Bink MCAM, Kruisselbrink JW, Bassil N, Gasic K, Iezzoni A, Van de Weg E, Peace C (2019) High-quality, genome-wide SNP genotypic data for pedigreed germplasm of the diploid outbreeding species apple, peach, and sweet cherry through a common workflow. PLoS ONE 14:e0210928. https://doi.org/10.1371/journal.pone.0210928
Vermeulen AJ (1962) De Sprenger-appelrassen. Laboratorium Voor Tuinbouwplantenteelt. Landbouwhogeschool Wageningen Report of Proef 148
Volk GM, Bramel P (2021) Apple Genetic resources: Diversity and Conservation. Apple Genome :33–45
Volk GM, Peace CP, Henk AD, Howard NP (2022) DNA profiling with the 20K apple SNP array reveals Malus domestica hybridization and admixture in M. Sieversii, M. Orientalis, and M. Sylvestris genebank accessions. Front Plant Sci 13
Acknowledgements
Several accessions used for the genotyping were generously provided by M. van Lienden, E. Puijk, J. Zandbergen, G. van Santvoort, H. Rossel, A. R. Kleefstra, and M. Jostmeijer, who also provided valuable help resolving the identities of specific accessions for which we are deeply thankful. The staff at the GENTYANE genotyping platform (doi:https://doi.org/10.15454/1.5572409592543596E12) (especially Dr. Charles Poncet, Carole Confolent and Lydia Jaffrelo, INRAE, Clermont-Ferrand, France) are warmly acknowledged for producing SSR genotyping data. SNP data that was used for comparison of the Dutch germplasm SNP dataset was obtained through the pedigree reconstruction project. Part of this 20 K SNP data came from the FruitBreedomics project no 265582: “Integrated approach for increasing breeding efficiency in fruit tree crops” DOI: https://doi.org/10.1038/s41438-018-0016-3, which was co-funded by the EU seventh Framework Programme. Other 20 K SNP data that was used for comparison was provided by the Fondazione Edmund Mach. SNP data obtained through genotyping leaf samples from the Seed Savers Exchange, USA were also included.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding authors states that there is no conflict of interest.
Data Archiving Statement
The data used in this study are deposited in the Genome Database for Rosaceae (rosaceae.org) under the accession number tfGDR1079.
Additional information
Communicated by Communicated by M. Troggio.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Larsen, B., van Dooijeweert, W., Durel, CE. et al. SNP genotyping Dutch heritage apple cultivars allows for germplasm characterization, curation, and pedigree reconstruction using genotypic data from multiple collection sites across the world. Tree Genetics & Genomes 20, 21 (2024). https://doi.org/10.1007/s11295-024-01655-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-024-01655-9