Abstract
Periodically, the European Space Agency (ESA) updates scientific roadmaps in consultation with the scientific community. The ESA SciSpacE Science Community White Paper (SSCWP) 9, “Biology in Space and Analogue Environments”, focusses in 5 main topic areas, aiming to address key community-identified knowledge gaps in Space Biology. Here we present one of the identified topic areas, which is also an unanswered question of life science research in Space: “How to Obtain an Integrated Picture of the Molecular Networks Involved in Adaptation to Microgravity in Different Biological Systems?” The manuscript reports the main gaps of knowledge which have been identified by the community in the above topic area as well as the approach the community indicates to address the gaps not yet bridged. Moreover, the relevance that these research activities might have for the space exploration programs and also for application in industrial and technological fields on Earth is briefly discussed.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
Humanity has been living and working in space for more than 60 years. Accordingly, we now know that spaceflight induces a variety of biological changes. These have recently been characterized as key biological features of spaceflight which include mitochondrial dysregulation, epigenetic changes, DNA damage, altered telomeres, microbiome shifts, and oxidative stress1. It has been suggested that matching these features to the known hazards of spaceflight (which include altered radiation, altered gravity, being in a hostile and closed environment, being confined, and being at a distance from Earth) is the main area where translational space biology should focus1.
Much as others have recently consolidated the key biological features of spaceflight, the European Space Agency (ESA) periodically consolidates ESA scientific community views on strategic priorities for ESA Space Biology. Here we present the ESA SciSpacE Science Community White Paper views on the topic of “How to Obtain an Integrated Picture of the Molecular Networks Involved in Adaptation to Microgravity in Different Biological Systems”. This topic emerged as a new topic for ESA as the result of the United States National Aeronautics and Space Administration (NASA) formation of GeneLab and the ESA formation of Topical Teams focused on Space Omics and Personalized Medicine.
Historically, space biology experiments have been divided between sub-disciplines such that it is not always clear if results from one species or cell type relate or translate to another species or cell type. For example, some microbes grow faster in space while others grow slower2,3 or some physiological systems show large alterations in space whereas others do not1,4,5. Despite these obvious differences, accumulated data points to the fact that altered metabolism appears to largely be a conserved across species response to spaceflight6,7,8,9,10,11 and that specific signals such as reactive oxygen species and peptide signals also appear to be altered across species (including animals and plants)12,13,14. Thus, while continued reductionist, mechanistically hypothesis-driven research remains critical for advancing our knowledge of space biology, it is also imperative that we use large data analysis methods to re-explore already accumulated data and to accumulate a more accurate picture of common versus distinct cellular and biological system responses to spaceflight15,16. For example, the ESA Space Omics Topical Team has recently reviewed ESA Space Omics research17 and has used this analysis to make recommendations for supporting future Space Omics work18. Similarly, the ESA Space Omics Topical Team has reviewed all space omics data for C. elegans and used this data19, in conjunction with other cross species data analysis by NASA GeneLab, to propose and fly a new C. elegans spaceflight experiment to test predicted causes of reproducible gene expression changes (the UK Space Agency’s recent Molecular Muscle Experiment 2).
Key knowledge gaps
Despite the growing interest of the space science community depicting the mechanisms relying on human susceptibility to the space environment, there are still significant information gaps which need to be investigated deeper. As a result, in order to enable extended human missions in space, particularly for operations beyond low Earth orbit (LEO), the next scientific questions should be aimed at the prediction and modelling of long-term effects of spaceflight. Thus, it will be essential to obtain an integrated picture of the molecular and cellular networks (e.g. intercellular communication, extracellular vesicles, etc.) involved in adaptation to altered gravity within the organism and farther compare different biological systems. This increased knowledge is also key to implement strategies that effectively prevent and counteract short- and long-term effects of adaptation to spaceflight by identifying novel diagnostic markers and therapeutic targets that could be also useful to support medical research on Earth. A key issue for ESA is that NASA and the Japanese Aerospace Exploration Agency (JAXA) are already moving forward with these knowledge gaps and ESA risks falling behind if they do not add this area of research to their portfolio.
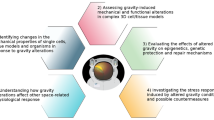
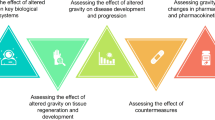
Three key issues have been identified (Fig. 1):
-
1.
Since altered gravity conditions affect many levels of cellular response, thus affecting both human physiology and pathophysiology20, it will be pivotal to compare the effects of the space environment in different types of cells in the same organism and/or a single cell type in different organisms. To provide a wider picture of space related effects on cells and tissues, it will be necessary to resort to the application of -omics approaches.
-
2.
To handle big data coming from -omics analysis, it will be required to use artificial intelligence and/or machine learning modelling for developing hypotheses concerning genes and pathways involved (common to the entire organism or specific to an individual system).
-
3.
To develop bioinformatics tools to allow real-time analysis and comparison within genomic and pathway databases.
The SSCWP made recommendations for work/goals in these three areas. In the short term, network biology approaches should be applied to compare changes within and across species. In the near term, AI/ML approaches should be incorporated into network biology approaches. In the longer term, network biology approaches should be employed in real time in flight. Additional details and references are provided in the text.
Proposed research activities to fulfill open scientific question
In the effort to fill the above gaps of knowledge, thus contributing to answer the question “How to obtain an integrated picture of the molecular networks involved in adaptation to microgravity in different biological systems?” three main classes of research activities have been identified:
-
1.
To investigate cell and tissue response to both simulated and real space conditions, proposed research activities should be aimed at a thorough analysis of biological samples which will allow to connect biology, physiology and evolutionary anthropology. A more comprehensive approach, human-scaled experiments, as well as basic science research investigations are required in this context to help unveil the origins of (mal)adaptations to space in the context of human evolution.
The International Standards for Space Omics Processing group has recently reviewed the pros and cons of various omic model systems for spaceflight21, and we encourage use of these systems. For example, omics have recently been employed with physiologic measures to suggest mechanisms underlying strength decline in flight for C. elegans22. Importantly, as ESA does not have on-orbit rodent research facilites, we encourage ESA community researchers to use rodent data collected by NASA and JAXA. For example, using such data, a recent ESA Space Omics Topical Team study has confirmed cross-tissue coordination underlying omic changes in mice in flight23. In terms of human subjects, the ESA Space Omics Topical Team has recently suggested that this should be a priority19,24, a view we support. Further, cross-species analysis should be encouraged in order to facilitate translational relevance. For example, a forthcoming analysis showing C. elegans, rodents, humans all display alterations in insulin controlled genes in response to spaceflight25.
-
2.
To implement the use of artificial intelligence (AI) and/or machine learning (ML) tools for establishing hypotheses about the biological response to the space environment, the main goal of proposed research activities will be aimed at: (i) developing bioinformatics models to highlight both differential gene expression and pathway activation in altered gravity conditions (for ML-centric examples see26,27); introducing predictive modelling as a tool for actively supporting ground and space crews in tackling health status and therapeutic treatment anomalies; (ii) promoting new capabilities for space omics research in Europe; (iii) collaborating more intensively with international ISS partners, NASA GeneLab (for example the AI/ML Analysis Working Group), International Standards for Space Omics Processing or similar space omics international consortiums.
-
3.
To enable real-time analysis and comparison of genomic and pathway databases generated by the integration of biological tests and artificial intelligence, proposed research ideas should be oriented towards: the development of in space -omics data processing coupled with bioinformatics analysis; ranking of cross-correlated biomarkers relative to intertangled morbidities and co-morbidities.
The timeline needed to obtain outcomes from research activities belonging to class 1 and class 2 is expected to range from short to medium, while the implementation of the activities belonging to class 3 requires a timeline ranging from medium to long. For all three classes of research activities, the suitable testbed environment to develop the related research programs and experiments spans from simulation of altered gravity conditions to be performed on ground to in vitro and in vivo tests to be performed both in low Earth orbit (LEO) and beyond LEO, as well as human-scoped tests to be handled in LEO and beyond LEO. Table 1 summarises open scientific questions and identified research activities in the context of the above-mentioned action points (including testbed environment and space relevance) across short-, middle-, and long-term timeframes.
Priorities for the Space Programme (microgravity and/or exploration relevance)
From a microgravity perspective, comparing the -omics response of different cell types, will enable the creation of the space equivalent of a cell atlas much as for human disease and physiology on Earth21,28. This will enable more accurate work on modelling where existing data from similar endeavours on Earth can be used to inform hypotheses concerning genes and pathways involved in the response to spaceflight. This could be achieved via partnership with the European Molecular Biology Laboratory (EMBL) or various precision medicine groups across Europe. From an exploration perspective, identifying pathways involved in (mal)adaptation to spaceflight will enable a database of -omics changes that are and are not related to spaceflight maladaptation. This knowledge base forms a foundation of future personalised medicine initiatives and real time analysis of -omics changes in space. This could be achieved not only using ESA astronaut data but also via partnership with NASA, JAXA, and commercial spaceflight providers. The Space Omics and Medical Atalas (SOMA)29 provides an example of such a partnership opportunity.
Benefit for Earth and industrial relevance
Improved understanding of the molecular basis of (mal)adaption to spaceflight should translate into better understanding of the molecular basis of many chronic diseases commonly observed on Earth30,31. The possibility of identifying changes that are common to single cell types or physiological systems in space adds an important datapoint to terrestrial -omics studies that aim to elucidate the molecular basis of diseases or health conditions via the use of large, unique, multi-data sets. Importantly, achieving the objective to perform real-time analysis, by online measuring of identified pathways, on, in, or beyond low Earth orbit (LEO) missions poses a technical challenge that is similar to that of remote bedside monitoring32. Thus, much as on-site gene sequencing enables research and diagnostics both on the International Space Station33 and on ground (currently in the context of infectious disease management)34, future technologies developed for use in space should have important spin out uses on Earth.
Conclusions
While different cell types and different species show differences in their sensitivity to microgravity, recent studies demonstrated that there are common features in microgravity-induced alterations observed across different cell types and different species. While the molecular mechanisms underlying these features can be studied through specific experiments using suitable in vitro, ex vivo, in vivo models, and also focused clinical trials, the application of large data analysis methods to process data sets from new and previous studies can allow us to obtain a more holistic view of the molecular networks involved in adaptation to microgravity in different biological systems and a clearer picture of common versus distinct responses to spaceflight at cellular and systemic level. By folding these community-identified knowledge gaps into its own reseach portfolio, ESA can ensure that it joins NASA and JAXA at the forefront of space biolgy research as the field continues its rapid shift in paradigm towards a ‘big data’ era.
Data availability
No datasets were generated or analysed during the current study.
References
Afshinnekoo, E. et al. Fundamental biological features of spaceflight: advancing the field to enable deep-space exploration. Cell 183, 1162–1184 (2020).
Benoit, M. R. & Klaus, D. M. Microgravity, bacteria, and the influence of motility. Adv. Space Res. 39, 1225–1232 (2007).
Huang, B., Li, D.-G., Huang, Y. & Liu, C.-T. Effects of spaceflight and simulated microgravity on microbial growth and secondary metabolism. Mil. Med Res. 5, 18 (2018).
Weronika, E. & Łukasz, K. Tardigrades in Space Research - Past and Future. Orig. Life Evol. Biosph. 47, 545–553 (2017).
Aubert, A. E. et al. Towards human exploration of space: the THESEUS review series on cardiovascular, respiratory, and renal research priorities. NPJ Microgravity 2, 16031 (2016).
Ma, L., Ma, J. & Xu, K. Effect of spaceflight on the circadian rhythm, lifespan and gene expression of Drosophila melanogaster. PLoS One 10, e0121600 (2015).
Sharma, G. & Curtis, P. D. The impacts of microgravity on bacterial metabolism. Life 12, 774 (2022).
Higashibata, A. et al. Microgravity elicits reproducible alterations in cytoskeletal and metabolic gene and protein expression in space-flown Caenorhabditis elegans. NPJ Microgravity 2, 15022 (2016).
Stein, T. P. et al. Energy expenditure and balance during spaceflight on the space shuttle. Am. J. Physiol. 276, R1739–R1748 (1999).
Altman, P. L. & Talbot, J. M. Nutrition and metabolism in spaceflight. J. Nutr. 117, 421–427 (1987).
da Silveira, W. A. et al. Comprehensive multi-omics analysis reveals mitochondrial stress as a central biological hub for spaceflight impact. Cell 183, 1185–1201.e20 (2020).
Yang, J., Zhang, G., Dong, D. & Shang, P. Effects of Iron Overload and Oxidative Damage on the Musculoskeletal System in the Space Environment: Data from Spaceflights and Ground-Based Simulation Models. Int. J. Mol. Sci. 19, (2018).
Goodwin, T. J. & Christofidou-Solomidou, M. Oxidative Stress and Space Biology: An Organ-Based Approach. Int J Mol Sci 19, (2018).
Sugimoto, M. et al. Genome-wide expression analysis of reactive oxygen species gene network in Mizuna plants grown in long-term spaceflight. BMC Plant Biol. 14, 4 (2014).
Ray, S. et al. GeneLab: Omics database for spaceflight experiments. Bioinformatics 35, 1753–1759 (2019).
Scott, R. T. et al. Advancing the Integration of Biosciences Data Sharing to Further Enable Space Exploration. Cell Rep. 33, 108441 (2020).
Deane, C. S. et al. Space omics research in Europe: Contributions, geographical distribution and ESA member states funding schemes. iScience 25, 103920 (2022).
Manzano, A. et al. Enhancing European capabilities for application of multi-omics studies in biology and medicine space research. iScience 25, 107289 (2022).
Scott, A. et al. C. elegans in microgravity: An omics perspective. iScience 25, 107180 (2022).
Bradbury, P. et al. Modeling the impact of microgravity at the cellular level: implications for human disease. Front Cell Dev. Biol. 8, 96 (2020).
Rutter, L. et al. A new era for space life science: international standards for space omics processing. Patterns 1, 100148 (2020).
Soni, P. et al. Spaceflight Induces Strength Decline in Caenorhabditis elegans. Cells 12, 2470 (2023).
Vitry, G. et al. Muscle atrophy phenotype gene expression during spaceflight is linked to a metabolic crosstalk in both the liver and the muscle in mice. iScience 25, 105213 (2022).
Cope, H. et al. Routine omics collection is a golden opportunity for European human research in space and analogue enviornments. iScience 25, 100550 (2022).
Mathyk, B. A. et al. Spaceflight alters insulin and estrogen signaling pathways. Research square https://doi.org/10.21203/rs.3.rs-2362750/v1 (2023).
Li, K. et al. Explainable machine learning identifies multi-omics signatures of muscle response to spaceflight in mice. npj Microgravity 9, 90 (2023).
Cope, H. et al. More than a Feeling: Dermatological Changes Impacted by Spaceflight. Research square https://doi.org/10.21203/rs.3.rs-2367727/v1 (2023).
Regev, A. et al. The Human Cell Atlas. Elife 6, (2017).
Overbey, E. G. Collection of Biospecimens from the Inspiration4 Mission Establishes the Standards fro the Space Omics and Medical Atlas (SOMA). bioRxiv https://doi.org/10.1101/2023.05.02.539108 (2023).
Strollo, F. et al. Recent Progress in Space Physiology and Aging. Front Physiol. 9, 1551 (2018).
Vernikos, J. & Schneider, V. S. Space, gravity and the physiology of aging: parallel or convergent disciplines? A mini-review. Gerontology 56, 157–166 (2010).
Nangalia, V., Prytherch, D. R. & Smith, G. B. Health technology assessment review: remote monitoring of vital signs–current status and future challenges. Crit. Care 14, 233 (2010).
Castro-Wallace, S. L. et al. Nanopore DNA sequencing and genome assembly on the international space station. Sci. Rep. 7, 18022 (2017).
Mongan, A. E., Tuda, J. S. B. & Runtuwene, L. R. Portable sequencer in the fight against infectious disease. J. Hum. Genet 65, 35–40 (2020).
Acknowledgements
This manuscript has been written in the frame of the activities for the preparation and dissemination of the ESA-ROADMAPS FOR FUTURE RESEARCH which define strategic goals for future space research on the ISS and supporting research platforms. Support was granted to JIB and AC by the Federal Ministry of Economics and Technology/Climate Action [BMWi/K, 50WB1931 and 50WB2222].
Author information
Authors and Affiliations
Contributions
Conceptualization, Monica Monici (M.M.) and Nathaniel J. Szewczyk (N.J.S.); writing, Craig R. G. Willis (C.R.G.W.), Marco Calvaruso (M.C.), N.J.S. and M.M., these authors contributed equally to the preparation of the manuscript; writing review and editing, C.R.G.W., M.C., M.M. and N.J.S.; supervision, M.M. and N.J.S. The other authors contributed to write the SSCWP.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Willis, C.R.G., Calvaruso, M., Angeloni, D. et al. How to obtain an integrated picture of the molecular networks involved in adaptation to microgravity in different biological systems?. npj Microgravity 10, 50 (2024). https://doi.org/10.1038/s41526-024-00395-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41526-024-00395-3
- Springer Nature Limited