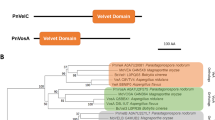
Abstract
The fungus Parastagonospora nodorum infects wheat through the use of necrotrophic effector (NE) proteins that cause host-specific tissue necrosis. The Zn2Cys6 transcription factor PnPf2 positively regulates NE gene expression and is required for virulence on wheat. Little is known about other downstream targets of PnPf2. We compared the transcriptomes of the P. nodorum wildtype and a strain deleted in PnPf2 (pf2-69) during in vitro growth and host infection to further elucidate targets of PnPf2 signalling. Gene ontology enrichment analysis of the differentially expressed (DE) genes revealed that genes associated with plant cell wall degradation and proteolysis were enriched in down-regulated DE gene sets in pf2-69 compared to SN15. In contrast, genes associated with redox control, nutrient and ion transport were up-regulated in the mutant. Further analysis of the DE gene set revealed that PnPf2 positively regulates twelve genes that encode effector-like proteins. Two of these genes encode proteins with homology to previously characterised effectors in other fungal phytopathogens. In addition to modulating effector gene expression, PnPf2 may play a broader role in the establishment of a necrotrophic lifestyle by orchestrating the expression of genes associated with plant cell wall degradation and nutrient assimilation.
Similar content being viewed by others
Introduction
The fungus Parastagonospora nodorum causes septoria nodorum blotch (SNB) of wheat. P. nodorum uses necrotrophic effectors (NEs) to cause tissue necrosis and facilitate infection of hosts possessing dominant susceptibility genes. The genes encoding three of these NEs are known: SnToxA, SnTox1, and SnTox3. SnToxA encodes a 13.2 kDa mature protein that causes necrosis on wheat cultivars that possess the dominant susceptibility gene Tsn11,2. Near-identical copies of ToxA have been found in two other wheat fungal pathogens, Pyrenophora tritici-repentis (Ptr)3 and Bipolaris sorokiniana4. These may have been horizontally acquired, presumably from P. nodorum1. SnTox1 encodes a 10.3 kDa cysteine-rich mature protein that causes necrosis and confers virulence on wheat cultivars possessing Snn15. SnTox3 is also a cysteine-rich NE. Sensitivity to the effector is conferred by either Snn3-B1 or Snn3-D1 located on wheat chromosomes 5BS and 5DS, respectively6,7. Genetic studies and protein purification assays indicate that P. nodorum possesses many more unidentified effectors associated with SNB8.
SnToxA, SnTox1 and SnTox3 are highly expressed during early infection but their expression is greatly decreases during saprophytic growth on the necrotised host tissue9. However, else was known about factors affecting their regulation until recently. Studies of TFs in P. nodorum have also provided some insights into effector gene regulation. Deletion of the APSES-class TF gene SnStuA in P. nodorum resulted in mutants with abnormal vegetative growth, loss of sporulation and a complete loss of virulence on wheat10. The expression of SnTox3 was significantly down-regulated in the mutant, though the loss in virulence is likely attributable to pleotropic effects incurred by the mutation. A C2H2 zinc finger TF PnCon7 that binds to the promoter region of SnTox3 was identified using a combination of yeast-1-hybrid (Y1H) and DNase footprinting, suggesting that PnCon7 may directly regulate SnTox3 expression11. Silencing of PnCon7 drastically reduced SnTox3 expression, suggesting that PnCon7 may be a direct regulator11.
Cho et al.12 identified and characterised a Pleosporales-specific zinc-finger TF gene Abpf2 from Alternaria brassicicola using gene knockout methods. Mutants lacking AbPf2 were non-pathogenic on various brassica hosts. Gene expression analysis using RNAseq identified eight putative candidate effector genes that were positively regulated by AbPf2. A BLAST search of AbPf2 against the P. nodorum predicted protein set identified a conserved homolog, PnPf29. Functional analysis revealed that PnPf2 is a positive regulator of SnToxA and SnTox3 expression and mutants lacking PnPf2 were only infective on Snn1 wheat lines9. Based on all evidence observed, we hypothesise that PnPf2 regulates the expression of novel effectors in P. nodorum. Firstly, P. nodorum SN15 carrying SnToxA, SnTox1 and SnTox3 deletions (toxa13) retained the ability to produce culture filtrate that cause host-specific chlorosis13 and remained highly pathogenic on many modern bread wheat lines14. Secondly, genetic analysis revealed new quantitative trait loci for SNB were detected on wheat mapping populations14,15. It is possible that these QTL may be associated with novel dominant susceptibility genes8. Lastly, SN15 carrying deletions in both PnPf2 and SnTox1 lost the ability to infect all wheat lines tested including those that demonstrated susceptibility to P. nodorum toxa139. This strongly suggests that PnPf2 positively regulates the expression of novel effector genes. To investigate this hypothesis and dissect other biochemical aspects of PnPf2 regulation, we used RNAseq to compare the gene expression profiles of a P. nodorum pnpf2 mutant with the wildtype strain under conditions that are conducive for effector gene expression.
Results
PnPf2 is required for full hyphal proliferation during host infection
The transcriptome of the P. nodorum reference wildtype strain SN15 was compared to the PnPf2-deleted strain pf2-69 grown under two conditions. Firstly, we sampled RNA during early infection at three days in planta (ip) where PnPf2, SnToxA, SnTox1 and SnTox3 are maximally expressed. Wheat cv. Halberd (Tsn1, Snn1, Snn3) was used as a host as it is susceptible to SN15 and pf2-699. Secondly, SN15 and pf2-69 were grown for three days in vitro (iv) in Fries 3 broth which is conducive for SnTox1 and SnTox3 production9. Vegetative growth of pf2-69 was comparable to SN159. Paired-end Illumina HiSeq technology was used as an RNAseq sequencing platform. The latest SN15 genome revision produced 13,563 predicted genes16. Deep sequencing produced more than 90% fungal transcripts that aligned to predicted genes for all samples (Supplementary Data S1 and Table 1). In vitro and ip samples returned an average of 24 million and 290 million read pair fragments (including plant reads), respectively. Between 18 and 22 million read pairs, representing an average of 6.94% of the total reads, aligned to the SN15 genome for the SN15 ip treatment (Supplementary Table S1). Between 3.4 and 5.9 million reads (average 1.57% of total) from pf2-69 ip growth aligned to the SN15 genome. The low proportion of fungal reads from pf2-69 suggests reduced biomass during infection. Quantitative PCR of genomic DNA extracted from three days post infected wheat cv. Halberd confirmed that the biomass of pf2-69 was significantly lower than strains carrying a functional copy of PnPf2 (Fig. 1a,b).
Infection, biomass and RNAseq analysis. (a) The onset of chlorotic symptoms was observed for SN15, pf2-69, Ect and pf2::PnPf2-infected wheat at three dpi. (b) Q-PCR quantification of biomass via fungal gDNA. Average biomass levels determined from PCR amplification of actin and α-tubulin not connected by the same letter are significantly different (P < 0.05) based on ANOVA (n = 3). (c) Comparing transcriptomes of SN15 and pf2-69 sampled in vitro and in planta using PCA from DESeq. 2 normalised reads (Supplementary Data S2). PnPf2 plays a bigger regulatory role during infection compared to in vitro growth. PC1 and PC2 explains 71% and 12% of the total variance, respectively. (d) Comparative RNAseq expression profiling of SnToxA, SnTox1 and SnTox3 in SN15 and pf2-69 under in vitro (iv) and in planta (ip) conditions. Bars show mean FPKM estimated by Cufflinks for each sample (n = 4), and error bars indicate standard deviation. Expression of all three effectors was reduced in the in planta pf2-69 samples compared with SN15.
Analysis of differentially expressed (DE) genes
Genes were considered DE in a contrast of isolate or treatment if tests of absolute log2 fold change >1 were consistently significant (Padj < 0.05) for three test methods described below (Supplementary Data S3). Because pf2-69 ip samples had considerably fewer reads than other samples, an additional filter requiring pf2-69 ip samples to have > = 10 counts per million (CPM) for a gene to be called down-regulated was used for high-confidence DE prediction sets. For SN15 ip and iv treatments, 1,889 genes were up-regulated and 1393 were down-regulated ip (Supplementary Table S2). A total of 1,736 genes were up-regulated and 706 genes were down-regulated between the pf2-69 ip and iv treatments. For ip comparisons, 303 genes were significantly reduced whereas 449 were up-regulated in pf2-69 over SN15. Additional DE genes were observed using relaxed criteria, allowing genes where any of the three tests are significant (<3 tests) (Supplementary Table S2). The main difference between the three tests results were in how they handle contrasts involving samples with few or no reads aligned to the gene. Additional genes involving pf2-69 ip samples with <10 CPM were identified using the same relaxed criteria. In total, 269 gene were down-regulated in pf2-69 ip compared to iv growth and had fewer than 10 CPM in pf2-69 during ip growth. Similarly, 163 genes were down-regulated in pf2-69 during infection compared to SN15 and had fewer than 10 CPM in pf2-69 during ip growth (Supplementary Table S2).
A principal component analysis (PCA) plot for PC1 and PC2 was constructed based on normalised fragment counts per gene to describe the variation between and within each treatment (Fig. 1c). The biological replicates tightly clustered together, with each treatment strongly differentiated from the others. This indicates that sample treatment and sequencing did not contribute to systematic biases that could not be removed by normalisation. PC1 captured 71% of the total variance and discriminated iv from ip samples. PC2 captured 12% of the variance and discriminated SN15 from pf2-69.
We then examined SnToxA, SnTox1 and SnTox3 expression profiles (Fig. 1d). As expected, the expression of SnToxA and SnTox3 was almost abolished in pf2-69 ip. SnTox3 expression was also highly reduced in pf2-69 iv. SnToxA is poorly expressed in SN15 and pf2-69 during iv growth. SnTox1 expression was significantly higher in SN15 compared to pf2-69. SnTox1 is still strongly expressed during ip growth and had the lowest fold change difference between SN15 and pf2-69 ip compared to SnToxA and SnTox3.
PnPf2 regulates genes that encode effector-like proteins
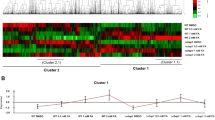
To identify candidate effector genes positively regulated by PnPf2, we analysed genes that were down-regulated in pf2-69 that possessed a secretory signal peptide (but no transmembrane domains outside of the signal peptide) and were predicted to be effector-like by EffectorP17. Twelve genes that showed a similar expression profile to SnToxA (ie. down-regulated in pf2-69 ip compared to SN15 ip and up-regulated ip in both strains) were identified (Fig. 2). In contrast, SnTox1 and SnTox3 were the only effector genes categorised in their respective expression profile categories (Fig. 2). The expression profiles of these candidate effector genes in SN15 and pf2-69 three days post-infection were validated using qRT-PCR (Supplementary Fig. S1). Apart from SNOG_10736, SNOG_13939 and SNOG_02980, the qRT-PCR-based expression profile of all other candidate effector genes between SN15 and pf2-69 was consistent with findings from the RNAseq data. The expression profiles of the 12 candidate effector genes in SN15 were examined between three and 10 days post-infection using available microarray gene expression data18 and qRT-PCR analyses performed in this study (Supplementary Fig. S2). SNOG_08150, SNOG_13939, SNOG_30077, SNOG_30352 and SNOG_30359 demonstrated similar expression profiles to SnToxA, SnTox1 and SnTox3 where gene expression peaked at three dpi and decreased to almost non-detectable levels at seven and 10 dpi, coinciding with host tissue necrosis.
Identification of SN15 candidate effector genes positively regulated by PnPf2. (a) An UpSet plot demonstrating the number of candidate effector genes that displayed similar expression profiles. Rows in the matrix represents sets of differentially expressed effector candidates in a contrast, with the solid triangles indicating the direction of expression change. Lines connecting rows of the matrix indicate an intersection between the sets, and the vertical bar chart indicates the number of genes that are common to those sets. Set intersections containing known effectors are indicated with colour. (b) A heatmap showing mean FPKM (n = 4) profiles of the candidate effector genes that share a common expression profile with SnToxA.
Four of the 12 candidate effectors possess Pfam domains (Supplementary Data S4 and Table 1). SNOG_01146 and SNOG_15270 possess a glycosyl hydrolase family domain. SNOG_02980 and SNOG_14243 both possess a hydrolase-type esterase family domain. A BLAST search of PHIbase19 indicated that SNOG_01146 displays significant amino acid sequence similarities to MoCDIP4 (Magnaporthe oryzae cell death–inducing protein P4) of the rice blast fungus M. oryzae20 whereas SNOG_15270 is similar to the Botrytis cinerea partial virulence determinant gene Xyn11A which encodes a xylanase21. Pfam domains were not observed for the other six candidates (Supplementary Data S4 and Table 1) but SNOG_08150, 12350, 30352, 30359 and 30077 encode small cysteine-rich (<20 kDa) proteins and BlastP analyses of SNOG_02755. 08150, 10736, 12350 and 13939 revealed significant hits to other fungal hypothetical proteins, whereas SNOG_30352, 30359 and 30077 appear to be unique to P. nodorum based on tBlastN searches.
P. nodorum SN79-1087 is non-pathogenic on wheat and lacks SnToxA, 1 and 35. We decided to investigate if these 12 candidate effectors are present or altered in SN79-108722. BlastP and tBlastN analysis revealed five genes were identical between SN15 and SN79-1087. SNOG_02755 and 10736 are also present in SN79-1087, but both have in-frame deletions in low-complexity amino acid repeat regions. Changes in amino acid sequence were observed for seven gene homologs in SN79-1087 (Table 1). Frame shifts or premature stop codons were not observed for these genes.
PnPf2 regulates depolymerase and nutrient assimilation gene expression in planta
To investigate changes in overall biochemical processes between SN15 and pf2-69 during iv and ip growth, we assessed DE genes for enrichment of GO terms23 (Fig. 3). GO terms were assigned to all genes where possible using InterProScan24 and dbCAN25.
An illustrated summary of gene ontology (GO) term enrichment analysis between pf2-69 and SN15 in vitro (a,b) and in planta (c and d). Bubble plots illustrate GO terms that were over-represented in differentially expressed (DE) genes as the sum of all log2 fold changes of pf2-69 relative to SN15 and the statistical significance of enrichment tests for GO terms (a,c). Bubble size indicates the number of genes annotated with that GO term. The relationships between significantly over-represented GO terms were highlighted using network analysis (b,d). Nodes represent a single GO term and are connected if a DE gene is annotated with both terms, with the shade of the edges indicating the proportion of genes with both GO terms that are differentially expressed. Node sizes indicate the statistical significance of GO term enrichment tests. Detailed GO term analysis is deposited as Supplementary Data S5. Interactive GO enrichment and network plots are deposited as Supplementary Data S6.
During iv growth, genes categorised under oxidoreductase activities, flavin adenine dinucleotide binding and catalytic activity were significantly up-regulated in pf2-69 (Fig. 3a). The majority of these genes encode cytochrome P450s, FAD binding proteins and oxidases (Supplementary Data S3 and S5). GO network analysis revealed that differentially expressed genes associated with oxidoreductase activities are central to biological processes related to respiratory electron transport chain, steroid metabolism, redox, carbohydrate metabolism and generation of precursor metabolites and energy (Fig. 3b).
During ip growth, molecular functions (MFs) associated with hydrolase, glucosidase and peptidase activities tended to be down-regulated in pf2-69 (Fig. 3c). The MF hydrolase activity associated with hydrolysing N-glycosyl compounds consisted of 155 genes. Of these, the expression of 39 genes were significantly lower in pf2-69. Similarly, the MF associated with another hydrolase activity associated with hydrolysing N-glycosyl compounds consisted of 32 genes, of which eight genes were significantly down-regulated in pf2-69 compared to SN15. The MF hydrolase activity consisted of 1,168 genes. Of these, 64 were expressed at lower levels in pf2-69. The majority of genes annotated encode plant cell wall degrading enzymes (CWDEs) and other carbohydrate depolymerases such as β-xylosidases, acetyl xylan esterases, glucanases and glucosidases (Supplementary Data S3 & S5). Arabinose is a major constituent of the plant cell wall. GO enrichment indicates that PnPf2 regulates arabinose metabolism in P. nodorum. Of the six genes associated with α-L-arabinofuranosidase activity, five were expressed at lower levels in pf2-69 (Fig. 3c,d).
For protein degradation, 240 genes encode proteins with predicted peptidase activity were differentially expressed (GO:0008233) (Fig. 3c). Of these, 29 were down-regulated in pf2-69 compared to SN15. Additionally, 63 genes encoding proteins with putative metallopeptidase activity were identified from the genome. Of these, 14 were down-regulated in pf2-69 ip compared to SN15. The MF associated with metallocarboxypeptidase activity (GO:0004181) consisted of nine genes where the expression of seven was reduced in pf2-69. For the MF associated with serine-type peptidase activity, 22 of 131 genes were expressed at lower levels in pf2-69. CAZyme and Interpro analyses of genes classified under GO:0008233, 0004181, 0008237 and 0008236 indicate that most encode peptidases and esterases (Supplementary Data S3 and S5).
GO analysis revealed that cellular redox potential in pf2-69 was perturbed during ip growth in addition to a similar defect observed during iv growth. A MF associated with oxidoreductase (GO:0016491) activity was enriched in up-regulated genes in pf2-69 ip compared to SN15 (Fig. 3c). The majority of DE genes encoding oxidases, cytochrome P450s, reductases and dehydrogenases (Supplementary Data S3 and S5) are associated with a biological role in carbohydrate metabolism (Fig. 3d). In addition, MFs linked to transport activities were enriched with genes that were similarly up-regulated in pf2-69 ip compared to SN15 (Fig. 3c). Genes associated with the transport function encode sugar and amino acid transporters (Fig. 3d; Supplementary Data S3 and S5).
Identification of DNA motifs enriched in the promoters of PnPf2-regulated genes
We hypothesised that a shared pf2-69 DE patterns implied a common transcriptional regulator. Therefore, promoters of these gene sets may harbor over-represented motif(s) functioning as potential PnPf2 transcription factor binding site(s) (TFBS). Analysis of the promoters from the respective pf2-69 DE gene groupings revealed three such motifs (Fig. 4). The motif WMGGVCCGAA, enriched in pf2-69 iv and ip down-regulated gene promoters, is similar to an enriched motif associated with AbPf2 down-regulated genes in A. brassicicola12 and is characteristic of a Zn2Cys6 TFBS26,27. A second motif resembling a C2H2 TFBS (RTSYGGGGWA) was significantly enriched in pf2-69 ip down-regulated gene promoters. The third motif (CTGYGCCGCA) also resembled a C2H2 TFBS and was enriched in pf2-69 iv up-regulated gene promoters. The identification of unique enriched motifs in the separate datasets suggests that PnPf2 may act as an indirect regulator or its binding site specificity can be influenced by other regulators of target genes.
Identification of motifs displaying enrichment in promoters of DE genes. Promoter sets used to model the motifs are listed in the second column. The third column contains motif matches to known fungal TFBSs in the JASPAR 2018 non-redundant database and their associated TF family. The fourth column lists the treatment groups displaying enrichment of the respective motif in the promoter set. Motif frequency and statistical analysis are described in Supplementary Data S7.
Absence of interaction between PnPf2 and the putative consensus motif on SnToxA and SnTox3 promoters
Inspection of the SnToxA and SnTox3 promoter region revealed at least one occurrence of the WMGGVCCGAA motif consensus sequences that was absent from SnTox1. For SnToxA, the consensus sequence was identified at 218, 364 and 416 bp upstream of the transcriptional start site. The consensus sequence was also observed at two sites in the PtrToxA promoter of Ptr. For SnTox3, the consensus sequence was identified at 679 bp upstream of the transcriptional start site. This consensus sequence was not observed in the promoter region of SnTox1. Therefore, it was hypothesised that WMGGVCCGAA functions as a PnPf2 binding site (Pf2BS). A yeast 1-hybrid (Y1H) assay was performed in order to determine whether PnPf2 can directly interact with the putative binding site represented in the SnToxA promoter. No significant interaction was observed between PnPf2 and four tandem repeats of the Pf2BS (Fig. 5a). Western blot analysis confirmed the presence of the PnPf2 protein indicating that the absence of Y1H interaction was not the result of the lack of protein (Fig. 5b).
Y1H analysis of PnPf2 and putative promoter motif interaction. (a) yPf2BS expressing PnPf2 grew on the -Leu medium; however, was not able to grow on the -Leu -His medium. 50 mM 3-AT was added to the -Leu -His plate for preventing possible histidine leakage. Mutated PnPf2 binding site (pf2bs) and empty vector were manipulated as negative controls. p53 interaction was used as positive control. Dilutions of yeast cells are indicated. (b) Western blots using HA antibody and coomassie blue (CB) staining of yeast cell extracts. 1: yp53BS x p53. 2: yPf2BS x empty vector. 3: yPf2BS x PnPf2. 4: ypf2bs x PnPf2. The PnPf2-GAL4AD-HA tag protein band is indicated (*). These are cropped images from different gel and blot photos. Original photos are supplied as Supplementary Fig. S3 where possible.
Identification of DE TF genes
We then screened for putative TF genes that were DE between SN15 and pf2-69 from the high confidence DE gene set to explore the possibility that PnPf2 operates indirectly. We limited our search to genes that encode proteins with TF domains found in fungi28. A total of 20 DE putative TFs were identified covering both iv and ip treatments. Based on distinct InterPro classifications24, this set consisted of five basic leucine zippers, one zinc knuckle, one myc-type, one CCHC-type, one p53-like, one C2H2, one homeodomain-like, six fungal specific Zn2Cys6 and three unspecified fungal TFs (Table 2). A BLAST search of PHIbase19 revealed that seven of these DE TF genes have strong matches to other fungal TFs associated with virulence (Table 2). Three of these belong to the fungal-specific Zn2Cys6 class (SNOG_03490, 07307 and 08440), one homeodomain-like (SNOG_08237) and three basic-leucine zippers (SNOG_04486, 13689 and 16487).
Discussion
Regulation of downstream target genes including those that encode effector-like proteins by members of the Pf2 Zn2Cys6 family was first reported in Cho et al.12 in A. brassisicola. The comparative RNAseq approach employed in that study derived from A. brassicicola-infected A. thaliana tissue which yielded a total of 8.5 to 9.3 × 105 reads from the WT and abpf2 mutant sample (approximately 0.5% of total reads) that mapped to the A. brassicola genome, respectively. Much higher fungal read counts were obtained in this study through the use of deep sequencing across four biological replicates resulting in more read information to exhaustively identify DE genes between SN15 and pf2-69 during ip growth (Supplementary Table S1).
RNAseq confirmed SnToxA and SnTox3 down-regulation in pf2-69 but the expression of SnTox1 was significantly higher in SN15 than pf2-69 than our previous observation9. The possibility that PnPf2 plays a minor regulatory role in SnTox1 regulation requires further investigation. Culture filtrates derived from pnpf2 mutants caused chlorosis on Snn1 wheat lines although the symptom was slightly weaker than with SN159. Nevertheless, SnTox1 is still strongly expressed in pf2-69 during infection and is sufficient to produce detectable SnTox1 activity in the culture filtrate and confer virulence on Snn1 wheat lines9.
It is not known if A. brassicicola uses effectors to modulate host infection. However, Cho et al.12 identified eight genes that encode small-secreted proteins with effector-like hallmarks positively regulated by AbPf2. Candidate effector genes were identified in this study that showed the same differential expression patterns as SnToxA and have effector-like properties. One of the effector candidates displayed significant sequence similarities to a known effector and pathogenicity factor. SNOG_01146 possesses a glycosyl hydrolase 61 domain and showed amino acid similarity to MoCDIP4. MoCDIP4 was identified as an apoplastic effector secreted by M. oryzae that causes cell death in rice20. Moreover, MoCDIP4 also induces cell death in non-host eudicots. In addition, these effectors are small, cysteine rich and expressed highly during early infection. SNOG_15270 is an homolog of Xyn11A which encodes an endo-ß-1,4-xylanase in B. cinerea. Deletion of Xyn11A in B. cinerea caused a significant reduction in virulence and growth on xylan21. All 12 candidate effector genes are also present in SN79-1087. Seven of these candidate proteins encode altered protein sequences in SN79-1087, which may explain some difference in pathogenicity. Five proteins possess changes in amino acid residues. It was previously observed that ToxA isoforms differ greatly in necrosis-inducing activities on Tsn1 wheats and affect the speed of asexual sporulation29. It is interesting to note that SNOG_02755 and 10736 polypeptides contain short amino acid sequence repeats that are partially deleted in SN79-1087. Several well-studied fungal and oomycete effectors contain repeats that possess functional roles in cellular localisation, host recognition and plant cell wall binding30. Additionally, recent studies have indicated that differential expression of effector genes between P. nodorum isolates affect their contributions to SNB of wheat15,31,32. The expression of these candidate genes in SN79-1087 requires further study.
GO enrichment revealed that PnPf2 functions as a positive regulator of a large subset of plant CWDEs and proteases during infection. Additionally, the removal of PnPf2 resulted in a general up-regulation in expression of nutrient transporter genes during infection. It is still not known whether this change is caused directly by the absence of PnPf2, or indirectly via another mechanism regulated by PnPf2. Comparative transcriptomic analysis of A. brassicicola identified only 13 genes that encode hydrolytic enzymes including two pectate lyases, were regulated by AbPf212. Deep sequencing used in this study provided a higher resolution insight into CAZyme regulation exerted by the Pf2 Zn2Cys6 class. Quantifying the contributions of plant CWDEs to phytopathogenicity is difficult because many fungal phytopathogens possess expanded gene families that result in functional redundancies33. For example, early studies on the causal agent of northern leaf spot of maize Cochliobolus carbonum (eg.34,35,36,37) did not find a clear role for CWDEs in fungal virulence. This is not to imply that CWDEs are dispensable for fungal virulence. It was reported that feruloyl esterases from Valsa mali38, a AbPf2-regulated pectate lyase from A. brassicicola39 and an endo-β-1,4-xylanase from B. cinerea21 function as virulence factors. Since plant CWDEs deconstruct the plant cell wall and liberate simple carbohydrates for assimilation and growth, it remains to be determined if SN15 can outcompete pf2-69 during co-infection on Snn1 wheats as the former can express a much larger repertoire of extracellular hydrolytic enzymes. RNAseq read counts suggested that pf2-69 accumulated much less biomass than SN15 at three dpi. This is surprising as pf2-69 retained the ability to causes lesions on Snn1 wheat lines comparable to SN15 as previously observed9. It is probable that SnTox1 secreted by pf2-69 during infection is the main cause of necrosis rather than the accumulation of fungal biomass at the lesion.
Analysis of the pf2-69 DE gene sets identified three distinct over-represented motifs (Fig. 4). The most notable of these is the WMGGVCCGAA motif associated with genes under PnPf2 positive regulation, as this motif was observed at multiple sites along the Sn/PtrToxA and SnTox3 promoters and also enriched in AbPf2-regulated gene promoters12. We hypothesised that it functions as a PnPf2 binding site as it resembles a Zn2Cys6 TFBS26,27. However, Y1H assay indicated that PnPf2 did not bind to the motif. This suggests either PnPf2 does not function as a direct regulator of SnToxA, SnTox3 and DE genes through interaction with the WMGGVCCGAA motif, or that necessary PnPf2 post-translational modifications/interactions are not compatible with the Y1H system. It was noted however that six other Zn2Cys6-type TF genes were differentially expressed between pf2-69 and SN15 (Table 2). Of these, only two were down-regulated but may serve as alternate candidates for direct regulation targeting the WMGGVCCGAA motif. A BLAST search of these against PHIbase revealed pathogenicity-associated functions in fungal homologues. SNOG_03490 is 52% identical to GzZC232 of Fusarium graminearum, the causal agent of fusarium head blight of wheat and is required for full virulence40. SNOG_08440 is homologous to a Zn2Cys6-type TF gene AtrR of Aspergillus fumigatus, an opportunistic fungal pathogen of mammals41. AtrR is a regulator of ergosterol biosynthesis pathway genes most notably Cyp51, a target for fungicide control. Deletion of AtrR resulted in impaired fungal growth and attenuated virulence on mice41. The other enriched motifs were characteristic of C2H2 binding sites27 however, only one DE TF of this class was identified - SNOG_15627 (Table 2). SNOG_15627 expression was up-regulated in pf2-69 under iv condition but remained unchanged during ip growth. SNOG_15627 demonstrated weak similarity to a characterised TF in F. graminearum shown to be dispensable for pathogenicity on wheat40. As the CTGYGCCGCA motif was enriched in the pf2-69 iv up-regulated gene promoters, it is possible that SNOG_15627 functions as a direct regulator. PnCon7 is the only characterised C2H2 TF in P. nodorum involved in SnTox3-mediated disease and direct regulation11. However, the cis-regulatory element of PnCon7 differs to both predicted C2H2 binding sites observed in this study.
We propose a model to explain the role of PnPf2 during early host infection based on evidence observed in this study (Fig. 6). The removal of PnPf2 drastically diminishes effector expression and so restricts the number of hosts on which P. nodorum is virulent9 (Fig. 6a). Both mutant and wild type strains are able to infect but the reduced ability to produce effectors and cell wall degrading enzymes means that pf2-69 is delayed in accessing bulk nutrients that come from the early stages of cell necrosis (Fig. 6b). The mutant has reduced access to nutrients stored as complex carbohydrates or compartmentalised in plant cells leading to a reduction in growth during host infection. Increased expression of transporter proteins may be an attempt to scavenge freely available nutrients possibly from the apoplastic space42 (Fig. 6c). In addition, we have identified candidate effector genes that are homologous to virulence factors and effectors in other phytopathogens. It is evident that PnPf2 functions to coordinate the expression of a subset of DE genes identified in this study through other TFs. Studies are currently under way to functionally characterise effector candidates and DE TF genes for their role in effector regulation and pathogenicity on wheat.
The proposed model for the role of PnPf2 during infection. (a) The deletion of PnPf2 diminishes effector expression and effector-triggered susceptibility in wheat carrying Tsn1 and Snn3. (b) In addition, PnPf2 functions as a positive regulator of CWDE expression in planta. (c) Without the full complement of CWDEs being produced, pf2-69 has a reduced ability to breakdown plant polymers and complex carbohydrates for assimilation needed during in planta growth.
Methods
Infection assays
Whole plant infection assay on two week-old wheat seedlings was performed as previously described43. Disease severity was visually determined and scored. A score of zero indicates no disease symptoms. A score of nine indicates a fully necrotised plant. Detached leaf infection assays on two-week old wheat cv. Halberd leaves was performed as previously described43,44.
Biomass analysis using quantitative (Q)-PCR
Q-PCR was to determine fungal biomass from infected wheat. Wheat cv. Halberd was infected with P. nodorum pycnidiospores as described above. Infection was allowed to develop for three days prior to sampling. The inoculated leaf section was excised and collected. Following this, gDNA was extracted using a Biosprint genomic DNA extraction kit (Qiagen, Venlo, Netherlands). Q-PCR was essentially carried out as described in Brouwer et al.45 using the primer pair alTubulinqPCRf/r and ActinqPCRhp2F/R (Supplementary Table S3).
RNA extraction and handling
RNA isolation and in planta gene expression analyses were performed as described in Rybak et al.9 using three day post-infected lesions excised from detached wheat. Library construction and sequencing was performed by the Ramaciotti Centre for Genomics (The University of NSW, Australia). Briefly, the TruSeq Stranded mRNA-seq method was used to prepare all libraries. Following this, sequencing was performed on an Illumina HiSeq. 2500 platform (San Diego, CA, USA) to generate 125 bp paired-end reads. Deep sequencing of all in planta samples were carried on individual lanes in the flowcell to ensure maximum sequence data was obtained from low fungal biomass. Samples derived from in vitro growth conditions were multiplexed into a single lane. The experiment was performed with four biological replicates.
RNAseq QC and read trimming
The quality of reads in the FastQ files were assessed using FastQC v0.11.5 (http://www.bioinformatics.babraham.ac.uk/projects/fastqc) before trimming adapter sequences using cutadapt v1.1246. Adapter trimmed reads were then filtered into sets belonging to SN15 and wheat using BBSplit v36.67 (https://sourceforge.net/projects/bbmap) using the P. nodorum genome16. Fungal reads were aligned to the SN15 genome using STAR v2.5.0a47. Novel splice sites were identified in a first pass alignment of the adapter-trimmed reads of all samples combined. Sample reads were then aligned individually using the novel splice sites identified in the first pass.
Determining differential gene expression in RNAseq
Fragments overlapping annotated features in the genome were counted using the SubRead featureCount v1.5.1 program using the union mode48. Differentially expressed (DE) genes were determined using the R packages EdgeR v3.16.449, DESeq. 2 v1.14.150 and Limma v3.30.651. DE genes were determined from tests of log2 fold changes (LFC) against the null hypothesis \(-1\le LFC\le 1\) (i.e. \({H}_{a}=|LFC| > 1\)) using a BH-adjusted P-value significance threshold of 0.05. Tests were also performed against the null hypothesis \(LFC\ne 0\), to be used where greater sensitivity (but lower confidence) was required. Unless otherwise specified, all results refer to thresholded tests (\(|LFC| > 1)\). Genes that were determined to be DE from tests by all three programs were taken as high-confidence DE sets. For contrasts involving samples with fewer than 10 million fragments (pf2-69 ip), genes from these samples were required to have a minimum of 10 CPM to be considered as differentially expressed in the high confidence sets. Normalised FPKM statistics were obtained using Cufflinks v2.2.152. Genes with mean FPKM > 100 were considered to be highly expressed.
Functional annotation
Functional annotations for existing genes were determined using InterProScan v5.19-58.024. Additionally, carbohydrate active enzymes were predicted using HMMER v3.1b2 (hmmer.org) and dbCAN v525. GO terms23 for each gene were found from combined dbCAN and InterProScan results, including matches from: Pfam53, TIGRFAM54, SMART55, PIRSF56, PANTHER57, HAMAP58, Prosite59, ProDom60, PRINTS61, and CATH-Gene3D62. Likely protein locations were determined using SignalP v4.163, TargetP v1.164, and TMHMM v2.0c65. Predicted proteins with a signal peptide and no transmembrane domains outside of the first 27 amino acids were considered to be secreted. Proteins with effector-like properties were determined using EffectorP v1.066 and were considered to be effector-like if they were also predicted to be secreted using the criteria above. Candidate genes were searched for in SN79-1087 (NCBI, GCA_002267025.1) using Spaln v2.3.367. Overlapping SN79-1087 genes were extracted and protein sequences were aligned using the needle command using EMBOSS68.
Functional enrichment of differentially expressed genes
Over-representation of GO terms in high-confidence differentially expressed gene sets were performed using the R package Goseq v1.26.069. Due to differences in the ability of DESeq. 2, EdgeR, and Limma to handle features with few aligned fragments; enrichment of effector-like or secreted transcripts were determined using the union of differentially expressed genes from all three prediction packages.
QRT-PCR determination of gene expression
Total RNA extraction from infected wheat cv. Halberd and P. nodorum mycelia from in vitro growth was extracted as described earlier. QRT-PCR was performed using a Quantitect SYBR Green RT-PCR kit (Qiagen, Valencia, CA, USA) and a Bio-Rad (Hercules, CA, USA) CFX96 system. P. nodorum SN15 gDNA was used as a quantitative standard. The expression value of each gene was normalised against the housekeeping gene actin (Act1) using the primer pair ActinqPCRf and ActinqPCRr70.
Analysis of promoters for enriched motifs
Common DNA motifs were discovered from the promoter regions 1.5 kbp upstream (or to the next annotated gene) of predicted transcription start sites of DE genes. Weeder 2.071 was used to search for enriched motifs in these promoters. A full set of SN15 predicted gene promoters was used for background frequencies with the redundancy filter set at 0.5. Utilising the consensus option in MEME v5.0.172, position weight matrices (PWMs) for top non-redundant motifs from each subset were derived for downstream analysis with MEMEsuite tools73. Each PWM motif was assessed for overrepresentation in pf2-69 DE subsets similar to Cho et al.12. Motif occurrences were first counted using FIMO74 and promoters with at least one occurrence were regarded as positive. Significance of over-representation in DE gene promoter sets was determined using Fisher’s exact test with Bonferroni corrected P-values (Padj < 0.05)75 as compared with the full promoter set of SN15. TOMTOM76 was used to search the JASPAR NR 2018 databases for matches (E < 1) to published fungal TFBSs in order to characterise the over-represented motifs.
Y1H assay
The construction of yeast reporter strain and Y1H screening was carried out based on the method of Ouwerkerk and Meijer77 with modifications. Y1H bait constructs were prepared by cloning three repeats of the p53 binding site (p53BS) (5′-AGACATGCCT-3′) using the primer pair p53BS-F1/R178, four repeats of the putative SnToxA PnPf2 binding site (Pf2BS) (5′-AAGGACCGA-3′) using the primer pair Pf2BS-F1/R1 and four repeats of pf2bs (5′-AAGGAAATA-3′) using the primer pair pf2bs-F1/R1 into pINT1-HIS3NB (provided by Dr. P.B.F Ouwerkerk, Leiden University) (Supplementary Table S3). Repeats of binding sites were cloned into pINT1-HIS3NB. Each construct was linearised, transformed into the yeast strain Y187 (Clontech, CA, USA) and selected on YPAD supplemented with G418. Bait strains were grown on selective media (-His) containing 3-amino-1,2,4-triazole (Sigma-Aldrich, MO, USA). Mating of the yeast bait strains with the prey strains was conducted by mixing the two strains together and grown on YPAD medium. Confirmation of the specific interaction between the bait sequence and the target protein was performed by reintroduce the prey construct into the bait strain. The prey construct pGADT7-p53 was built by cloning partial p53 from pGBKT7-53 (Clontech, CA, USA) into pGADT7. Similarly, PnPf2 was amplified from cDNA using the primer pair Pf2-F2/R3 and ligated into pGADT7.
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files)
References
Friesen, T. L. et al. Emergence of a new disease as a result of interspecific virulence gene transfer. Nature Genetics 38, 953–956 (2006).
Faris, J. D. et al. A unique wheat disease resistance-like gene governs effector-triggered susceptibility to necrotrophic pathogens. Proceedings of the National Academy of Sciences USA 107, 13544–13549 (2010).
Ciuffetti, L. M., Tuori, R. P. & Gaventa, J. M. A single gene encodes a selective toxin causal to the development of tan spot of wheat. Plant Cell 9, 135–144 (1997).
McDonald, M. C., Ahren, D., Simpfendorfer, S., Milgate, A. & Solomon, P. S. The discovery of the virulence gene ToxA in the wheat and barley pathogen Bipolaris sorokiniana. Mol. Plant Pathol. 19, 432–439 (2018).
Liu, Z. et al. The cysteine rich necrotrophic effector SnTox1 produced by Stagonospora nodorum triggers susceptibility of wheat lines harboring Snn1. PLoS Pathogens 8, e1002467 (2012).
Liu, Z. et al. SnTox3 acts in effector triggered susceptibility to induce disease on wheat carrying the Snn3 gene. PLoS Pathogens 5, e1000581 (2009).
Zhang, Z. et al. Two putatively homoeologous wheat genes mediate recognition of SnTox3 to confer effector-triggered susceptibility to Stagonospora nodorum. The Plant Journal 65, 27–38 (2011).
Friesen, T. L., Faris, J. D., Solomon, P. S. & Oliver, R. P. Host-specific toxins: effectors of necrotrophic pathogenicity. Cellular Microbiology 10, 1421–1428 (2008).
Rybak, K. et al. A functionally conserved Zn2 Cys6 binuclear cluster transcription factor class regulates necrotrophic effector gene expression and host-specific virulence of two major Pleosporales fungal pathogens of wheat. Mol. Plant Pathol. 18, 420–434 (2017).
IpCho, S. V. et al. The transcription factor StuA regulates central carbon metabolism, mycotoxin production, and effector gene expression in the wheat pathogen Stagonospora nodorum. Eukaryot. Cell 9, 1100–1108 (2010).
Lin, S. Y., Chooi, Y. H. & Solomon, P. S. The global regulator of pathogenesis PnCon7 positively regulates Tox3 effector gene expression through direct interaction in the wheat pathogen Parastagonospora nodorum. Mol. Microbiol (2018).
Cho, Y., Ohm, R. A., Grigoriev, I. V. & Srivastava, A. Fungal-specific transcription factor AbPf2 activates pathogenicity in Alternaria brassicicola. The Plant Journal 75, 498–514 (2013).
Tan, K. C. et al. Functional redundancy of necrotrophic effectors - consequences for exploitation for breeding. Frontiers in Plant Science 6, 501 (2015).
Phan, H. T. T. et al. Novel sources of resistance to Septoria nodorum blotch in the Vavilov wheat collection identified by genome-wide association studies. Theor Appl Genet (2018).
Phan, H. T. et al. Differential effector gene expression underpins epistasis in a plant fungal disease. The Plant Journal 87, 343–354 (2016).
Syme, R. A. et al. Comprehensive annotation of the Parastagonospora nodorum reference genome using next-generation genomics, transcriptomics and proteogenomics. PLoS One 11, e0147221 (2016).
Sperschneider, J. et al. EffectorP: predicting fungal effector proteins from secretomes using machine learning. New Phytologist 210, 743–761 (2016).
Ipcho, S. V. et al. Transcriptome analysis of Stagonospora nodorum: gene models, effectors, metabolism and pantothenate dispensability. Mol. Plant Pathol. 13, 531–545 (2012).
Urban, M. et al. PHI-base: a new interface and further additions for the multi-species pathogen-host interactions database. Nucleic Acids Res. 45, D604–D610 (2017).
Chen, S. et al. Identification and characterization of in planta-expressed secreted effector proteins from Magnaporthe oryzae that induce cell death in rice. Mol Plant Microbe Interact 26, 191–202 (2013).
Brito, N., Espino, J. J. & Gonzalez, C. The endo-beta-1,4-xylanase Xyn11A is required for virulence in Botrytis cinerea. Mol. Plant-Microbe Interact. 19, 25–32 (2006).
Richards, J. K., Wyatt, N. A., Liu, Z., Faris, J. D. & Friesen, T. L. Reference Quality Genome Assemblies of Three Parastagonospora nodorum Isolates Differing in Virulence on Wheat. G3 (Bethesda) 8, 393–399 (2018).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25, 25–29 (2000).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240 (2014).
Yin, Y. et al. dbCAN: a web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 40, W445–451 (2012).
MacPherson, S., Larochelle, M. & Turcotte, B. A fungal family of transcriptional regulators: the zinc cluster proteins. Microbiology and molecular biology reviews: MMBR 70, 583–604 (2006).
Khan, A. et al. JASPAR 2018: update of the open-access database of transcription factor binding profiles and its web framework. Nucleic Acids Res. 46, D1284 (2018).
Shelest, E. Transcription Factors in Fungi: TFome Dynamics, Three Major Families, and Dual-Specificity TFs. Front Genet 8, 53 (2017).
Tan, K. C. et al. Quantitative variation in effector activity of ToxA isoforms from Stagonospora nodorum and Pyrenophora tritici-repentis. Mol. Plant-Microbe Interact. 25, 515–522 (2012).
Mesarich, C. H., Bowen, J. K., Hamiaux, C. & Templeton, M. D. Repeat-containing protein effectors of plant-associated organisms. Front Plant Sci 6, 872 (2015).
Faris, J. D., Zhang, Z., Rasmussen, J. B. & Friesen, T. L. Variable expression of the Stagonospora nodorum effector SnToxA among isolates is correlated with levels of disease in wheat. Molecular plant-microbe interactions: MPMI 24, 1419–1426 (2011).
Peters, A. R., Zhang, Z., Richards, J. K., Friesen, T. L. & Faris, J. D. Genetics of variable disease expression conferred by inverse gene-for-gene interactions in the wheat-Parastagonospora nodorum pathosystem. Plant Physiol (2019).
Kubicek, C. P., Starr, T. L. & Glass, N. L. Plant Cell Wall-Degrading Enzymes and Their Secretion in Plant-Pathogenic Fungi. Annual Review of Phytopathology, Vol 52 52, 427–451 (2014).
Apel-Birkhold, P. C. & Walton, J. D. Cloning, disruption, and expression of two endo-beta 1, 4-xylanase genes, XYL2 and XYL3, from Cochliobolus carbonum. Appl. Environ. Microbiol. 62, 4129–4135 (1996).
Kim, H. et al. Mutational analysis of beta-glucanase genes from the plant-pathogenic fungus Cochliobolus carbonum. Mol Plant Microbe Interact 14, 1436–1443 (2001).
Ahn, J. H., Sposato, P., Kim, S. I. & Walton, J. D. Molecular cloning and characterization of cel2 from the fungus Cochlibolus carbonum. Biosci. Biotech. Bioch. 65, 1406–1411 (2001).
Gorlach, J. M., Van Der Knaap, E. & Walton, J. D. Cloning and targeted disruption of MLG1, a gene encoding two of three extracellular mixed-linked glucanases of Cochliobolus carbonum. Appl. Environ. Microbiol. 64, 385–391 (1998).
Xu, M. et al. The feruloyl esterase genes are required for full pathogenicity of the apple tree canker pathogen Valsa mali. Mol. Plant Pathol. 19, 1353–1363 (2018).
Cho, Y. et al. A Pectate Lyase-Coding Gene Abundantly Expressed during Early Stages of Infection Is Required for Full Virulence in Alternaria brassicicola. PLoS One 10, e0127140 (2015).
Son, H. et al. A phenome-based functional analysis of transcription factors in the cereal head blight fungus, Fusarium graminearum. PLoS Pathogens 7, e1002310 (2011).
Hagiwara, D. et al. A Novel Zn2-Cys6 Transcription Factor AtrR Plays a Key Role in an Azole Resistance Mechanism of Aspergillus fumigatus by Co-regulating cyp51A and cdr1B Expressions. PLoS Pathog 13, e1006096 (2017).
Solomon, P. S. & Oliver, R. P. The nitrogen content of the tomato leaf apoplast increases during infection by Cladosporium fulvum. Planta 213, 241–249 (2001).
Solomon, P. S., Tan, K. C. & Oliver, R. P. Mannitol 1-phosphate metabolism is required for sporulation in planta of the wheat pathogen Stagonospora nodorum. Mol. Plant-Microbe Interact. 18, 110–115 (2005).
Benedikz, P. W., Mappledoram, C. J. & Scott, P. R. A laboratory technique for screening cereals for resistance to Septoria nodorum using detached seedling leaves. Trans. Br. Mycol. Soc. 77, 667–668 (1981).
Brouwer, M. et al. Quantification of disease progression of several microbial pathogens on Arabidopsis thaliana using real-time fluorescence PCR. FEMS Microbiol. Lett. 228, 241–248 (2003).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal, http://journal.embnet.org/index.php/embnetjournal/article/view/200/458 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq. 2. Genome Biol 15, 550 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7, 562–578 (2012).
Punta, M. et al. The Pfam protein families database. Nucleic Acids Res. 40, D290–301 (2012).
Haft, D. H. et al. TIGRFAMs: a protein family resource for the functional identification of proteins. Nucleic Acids Research 29, 41–43 (2001).
Letunic, I., Doerks, T. & Bork, P. SMART: recent updates, new developments and status in 2015. Nucleic Acids Res. 43, D257–260 (2015).
Wu, C. H. et al. PIRSF: family classification system at the Protein Information Resource. Nucleic Acids Res. 32, D112–114 (2004).
Mi, H., Poudel, S., Muruganujan, A., Casagrande, J. T. & Thomas, P. D. PANTHER version 10: expanded protein families and functions, and analysis tools. Nucleic Acids Res. 44, D336–342 (2016).
Pedruzzi, I. et al. HAMAP in 2015: updates to the protein family classification and annotation system. Nucleic Acids Res. 43, D1064–1070 (2015).
Sigrist, C. J. A. et al. New and continuing developments at PROSITE. Nucleic Acids Research 41, E344–E347 (2013).
Bru, C. et al. The ProDom database of protein domain families: more emphasis on 3D. Nucleic Acids Res. 33, D212–215 (2005).
Attwood, T. K. et al. The PRINTS database: a fine-grained protein sequence annotation and analysis resource–its status in 2012. Database (Oxford) 2012, bas019 (2012).
Dawson, N. L. et al. CATH: an expanded resource to predict protein function through structure and sequence. Nucleic Acids Research 45, D289–D295 (2017).
Petersen, T. N., Brunak, S., von Heijne, G. & Nielsen, H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8, 785–786 (2011).
Emanuelsson, O., Nielsen, H., Brunak, S. & von Heijne, G. Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. Journal of Molecular Biology 300, 1005–1016 (2000).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. L. Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes. Journal of Molecular Biology 305, 567–580 (2001).
Sperschneider, J. et al. EffectorP: predicting fungal effector proteins from secretomes using machine learning. The New phytologist (2015).
Gotoh, O. A space-efficient and accurate method for mapping and aligning cDNA sequences onto genomic sequence. Nucleic Acids Res. 36, 2630–2638 (2008).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet 16, 276–277 (2000).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11, R14 (2010).
Tan, K. C. et al. A signaling-regulated, short-chain dehydrogenase of Stagonospora nodorum regulates asexual development. Eukaryot. Cell 7, 1916–1929 (2008).
Pavesi, G., Mereghetti, P., Mauri, G. & Pesole, G. Weeder Web: discovery of transcription factor binding sites in a set of sequences from co-regulated genes. Nucleic Acids Res. 32, W199–203 (2004).
Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–208 (2009).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Armstrong, R. A. When to use the Bonferroni correction. Ophthal Physl Opt 34, 502–508 (2014).
Gupta, S., Stamatoyannopoulos, J. A., Bailey, T. L. & Noble, W. S. Quantifying similarity between motifs. Genome Biology 8 (2007).
Ouwerkerk, P. B. & Meijer, A. H. Yeast one-hybrid screens for detection of transcription factor DNA interactions. Methods Mol Biol 678, 211–227 (2011).
Wang, Y., Schwedes, J. F., Parks, D., Mann, K. & Tegtmeyer, P. Interaction of p53 with its consensus DNA-binding site. Mol. Cell. Biol. 15, 2157–2165 (1995).
Lu, J., Cao, H., Zhang, L., Huang, P. & Lin, F. Systematic analysis of Zn2Cys6 transcription factors required for development and pathogenicity by high-throughput gene knockout in the rice blast fungus. PLoS Pathogens 10, e1004432 (2014).
Tanaka, A. et al. ProA, a transcriptional regulator of fungal fruiting body development, regulates leaf hyphal network development in the Epichloe festucae-Lolium perenne symbiosis. Mol. Microbiol. 90, 551–568 (2013).
Kim, S. et al. Homeobox transcription factors are required for conidiation and appressorium development in the rice blast fungus Magnaporthe oryzae. PLoS Genet 5, e1000757 (2009).
Pereira Silva, L. et al. Genome-wide transcriptome analysis of Aspergillus fumigatus exposed to osmotic stress reveals regulators of osmotic and cell wall stresses that are SakA(HOG1) and MpkC dependent. Cell Microbiol 19 (2017).
Li, X., Wu, Y., Liu, Z. & Zhang, C. The function and transcriptome analysis of a bZIP transcription factor CgAP1 in Colletotrichum gloeosporioides. Microbiol Res 197, 39–48 (2017).
Acknowledgements
This study was supported by the Centre for Crop and Disease Management, a joint initiative of Curtin University and the Grains Research and Development Corporation [research grant CUR00023 (Programme 3)]. EJ was supported by the Australian Government Research Training Program Scholarship. We thank Dr. James Hane for bioinformatic discussions.
Author information
Authors and Affiliations
Contributions
K.C.T. conceived the experiment. D.A.B.K., E.J., K.R., H.T.T.P. and S.Y.L. performed the experiment. K.B.S., P.S.S. and R.P.O. provided additional experimental design ideas and critical feedbacks. K.C.T., D.A.B.J. and E.J. wrote the paper. D.A.B.J., E.J., K.B.S., P.S.S., R.P.O. and K.C.T. edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jones, D.A.B., John, E., Rybak, K. et al. A specific fungal transcription factor controls effector gene expression and orchestrates the establishment of the necrotrophic pathogen lifestyle on wheat. Sci Rep 9, 15884 (2019). https://doi.org/10.1038/s41598-019-52444-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-52444-7
- Springer Nature Limited
This article is cited by
-
The Velvet transcription factor PnVeA regulates necrotrophic effectors and secondary metabolism in the wheat pathogen Parastagonospora nodorum
BMC Microbiology (2024)
-
MoSET1-dependent transcription factors regulate different stages of infection-related morphogenesis in Pyricularia oryzae
Journal of General Plant Pathology (2023)
-
Chromosome-level genome assembly and manually-curated proteome of model necrotroph Parastagonospora nodorum Sn15 reveals a genome-wide trove of candidate effector homologs, and redundancy of virulence-related functions within an accessory chromosome
BMC Genomics (2021)
-
GWAS analysis reveals distinct pathogenicity profiles of Australian Parastagonospora nodorum isolates and identification of marker-trait-associations to septoria nodorum blotch
Scientific Reports (2021)
-
Genetic response to nitrogen starvation in the aggressive Eucalyptus foliar pathogen Teratosphaeria destructans
Current Genetics (2021)