Abstract
Ovarian cancer remains a dismal disease with diagnosing in the late, metastatic stages, therefore, there is a growing realization of the critical need to develop effective biomarkers for understanding underlying mechanisms. Although existing evidences demonstrate the important role of the single genetic abnormality in pathogenesis, the perturbations of interactors in the complex network are often ignored. Moreover, ovarian cancer diagnosis and treatment still exist a large gap that need to be bridged. In this work, we adopted a network-based survival-associated approach to capture a 12-gene network module based on differential co-expression PPI network in the advanced-stage, high-grade ovarian serous cystadenocarcinoma. Then, regulatory genes (protein-coding genes and non-coding genes) direct interacting with the module were found to be significantly overlapped with cell death genes. More importantly, these overlapping genes tightly clustered together pointing to the module, deciphering the crosstalk between network-based survival-associated module and cell death in ovarian cancer.
Similar content being viewed by others
Introduction
Ovarian cancer, the most lethal gynecological malignancy, is the fifth most common causes of cancer death in women1. Generally, ovarian cancer patients have a poor prognosis with 5-year survival rate of 43.7% and even 63% who are diagnosed with metastasis disease only have the 5-year survival rate of 26.9%2 attributed to late detection and chemo-resistance. Therefore, it is of critical importance to identify clinical biomarkers responsible for monitoring ovarian cancer treatment, which may lead to development of novel therapeutic targets and eventually decrease the risk of death in ovarian cancer patients.
In recent decades, unprecedented multi-level omics data provide a convenient way to identify biomarkers for ovarian cancer with rapid development of high-throughput technologies, however, the findings in these studies still lack some successful applications for ovarian cancers so far3,4. Mostly, only rare genes without functional relationships are selected and treated as a potential biomarker of ovarian cancer3,5,6. Furthermore, owing to the highly clinically and genetically heterogeneous nature of cancer, several irrelevant abnormalities that are either low frequency of occurrences in all patients or rarely sufficient to cause cancer3,5,6,7 are probably not the ideal candidate considering the high reproducibility and sensitivity of biomarker. These difficulties will lead to a fatal problem influencing their application. However, considering the complex nature of the interaction between genes, single genetic abnormality can spread along the links of the complex intracellular network to alter a series of common gene products’ activities in either a direct or an indirect manner8,9,10. Just as Taylor IW et al. emphasized that changes in the biochemical wiring of oncogenic cells drives phenotypic transformations that can directly affect disease outcome11. Hence, it has recent emerged several network-based studies utilizing interaction information between genes in ovarian cancer12,13. However, these studies primarily fall into the category of static network analysis ignoring dynamics of network on both temporal and spatial specificity. Therefore, network-based dynamic modularity analysis for survival-associated biomarker discovery in longer-versus shorter-survival patients will provide more robust insights into preclinical therapeutic modality development on ovarian cancer.
More importantly, to a better understanding of outcome and optimal treatment of ovarian cancer, researchers are not fully satisfied with the challenges of obtaining biomarkers, thus, substantial interests have arisen to decipher their biological function. Currently, it gradually became clear that biomarker genes are mainly involved in these processes of immune, inflammatory, cell cycle and cell death14,15,16. Specifically for cell death, as a fundamental biological process, it plays an important role during the development, maintenance of tissue homeostasis and elimination of damaged cells17,18. Growing evidences have shown that excessive or defective cell death contributes to a broad spectrum of human diseases, including ovarian cancer19,20,21. Insights into the molecular mechanisms involved in cell death will likely have important implications and offer the opportunity to target this process for therapeutic purpose22,23,24,25. However, the rational treatment design and selection are often precluded due to the lack of the elaborate wiring diagram of biomarkers and cell death. Therefore, it is necessary to dissect and decipher the crosstalk between biomarkers and cell death. Shaping this roadmap will definitely provide more benefit for a more accurate outcome prediction and personalized management of ovarian cancer.
In this paper, based on constructed weighted survival and differential co-expression network between longer- versus shorter- survival patients using survival information, protein-protein interactions (PPI) in STRING26 and gene expression data from TCGA ovarian cancer, we adopted a network-based approach to capture a 12-gene network module. Survival analysis showed that this module was significantly related to overall survival of patients in ovarian cancer, whose prognostic ability was further confirmed in internal and external independent datasets. To elucidate the underlying mechanisms of this module in ovarian cancer, we further explored the genes significantly regulating the 12-gene module. Functional annotation of these genes showed a close correlation with cell death. Specifically, these significantly regulatory genes direct interacting with the module were significantly overlapped with cell death genes from our miRDeathDB27,28 and HADB29 and DeathBase30. More importantly, these overlapping genes were found to be tightly clustered together pointing to the 12-gene module. These findings highlighted that deciphering the crosstalk between network-based survival-associated module and cell death in ovarian cancer not only sheds light on its mechanism of action, but may also contribute to biomarker-guided preclinical and clinical therapeutic modality development.
Results
Identification of network-based survival-associated module in ovarian cancer
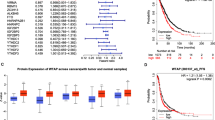
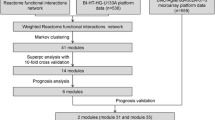
To identify module biomarkers of ovarian cancer, we first adopted a network-based simulated annealing approach to search putative modules by integrating survival information, PPI network and gene expression. Under the criteria with module score ranked in the top 1% (module score > 6.15) and p < 0.01, a total of 71 modules were identified in the constructed survival and differential co-expression PPI network between longer- versus shorter- survival patients. Then, for each module, we evaluated its predictive ability for survival of ovarian cancer patients, as described in the Materials and Methods. Notably, 27 of 71 modules were found to be significantly associated with overall survival of ovarian cancer patients in the training dataset (p < 0.1). Among all the survival-associated modules, the predictive ability of only a 12-gene module (Fig. 1, Table 1 and Supplementary Table S1), was further confirmed in independent internal dataset (In training dataset, log-rank p = 2.09E-3; In test dataset, log-rank p = 0.014). Gene Ontology functional annotation on the 12-gene module was presented in Supplementary Table S2. The distribution of the module genes’ risk scores and heatmap of the module genes’ expression profiles were shown in Fig. 2.
The 12-gene module biomarker and Kaplan-Meier estimates of overall survival in ovarian cancer patients according to this module biomarker.
(A) The interaction wiring of the 12-gene module. The nodes in red or blue indicated whether the genes have been verified as being related to ovarian cancer or not, respectively. The node sizes indicated the significance of association between the genes with cancer survival. The width of edges indicated the extent of differential co-expression of two genes. (B) Kaplan-Meier estimates of overall survival in the training and internal test datasets.
12-gene module risk score analysis of ovarian cancer.
(A) The distribution of the 12-gene module risk score. Patients were divided into a high-risk group (Red) or a low-risk group (Blue) using the median risk score as the cutoff point. (B) Heatmap of the module genes’ expression profiles. Rows and columns represented genes and patients, respectively.
By using Cytoscape31, interaction wiring of the module genes was visualized in Fig. 1A. And it can be clearly seen that the module genes exhibited a context-specific PPI pattern, reflecting the dynamic feature of module facing to different malignant extent of ovarian cancer patients. Following this, the clinical relevance of the module genes was further examined. As shown in Table 1, except for CD8B, TRAT1 and SYK, all the other module genes were found to be significantly associated with survival of ovarian cancer patients (p < 0.05). More importantly, the CD8B, TRAT1 and SYK genes were here included in this module because of owning the larger differential co-expression with their interaction neighbors, the survival-associated genes between longer- and shorter- survival patients (Supplementary Table S1). For instance, CD8B through interacting with the survival-associated CD3G, CD3E, CD3D and ZAP70 genes was recruited in this module. Although no significant survival association was observed in these three genes, their potential biological or clinical relevance were confirmed by the evidences excavated from related researches. Especially, cancer genes, TRAT1 and SYK, have been validated as being significantly associated with lethal ovarian cancer, driving malignant transformation of ovarian cancer32,33,34. Taken together, the network-based module analysis has the ability to provide a deeper understanding of the characteristics of biomarker beyond biomarker discovery.
Independent external validation of the 12-gene module biomarker in ovarian cancer
To further assess the predictive ability of the 12-gene module biomarker, we here used an independent external data for ovarian cancer35, in which only those patients with advanced-stage, high-grade ovarian serous cancer were analyzed. After dividing the patients into two subgroups according to the median value of EM scores in training dataset, we found that the 12-gene module was significantly related to overall survival of ovarian cancer patients using the log-rank test (log-rank p = 5.9E-3; Fig. 3). The hazard ratio of high-risk versus low-risk groups was 0.49 (95% CI: 0.29–0.82, p = 7.12E-3). Specifically, the patients with low EM scores resided in the high-risk group with a shorter survival. The median survival time for low-risk group was 105 months, whereas the median survival time for high-risk group was only 46 months. To further verify the correlation between the 12-gene module and survival, the predictive ability of the module in the independent external data was further confirmed and presented in Supplementary Table S3.
Prognostic value of the 12-gene module for assessing clinical outcome of ovarian cancer
After further adjusting for age, grade, stage and residual tumor size, as shown in Table 2, the univariate and multivariate analysis indicated that the 12-gene module biomarker, as an independent risk factor, was significantly associated with overall survival of ovarian cancer patients in the training (HR = 0.47, 95% CI: 0.38–0.73, p = 1.58E-4) and internal test (HR = 0.63, 95% CI: 0.41-0.97, p = 0.038) data. In addition, multivariate analysis also demonstrated that the designation of high- and low-risk groups remained statistically significant in the independent external data (HR = 0.51, 95% CI: 0.30-0.88, p = 0.014). Taken together, these analyses demonstrated the capacity of the 12-gene module biomarker to add value in a prognostic setting.
Crosstalk between the 12-gene module and cell death
Considering that the 12-gene module biomarker only provided a starting point for improving the decision making process, further research will be necessary to elucidate the underlying mechanisms of this module in ovarian cancer. Here, we examined the network wiring around the module genes and extracted their significant interacting neighbors. In total, 151 significantly regulating genes including miRNA genes were identified (p < 0.1, hypergeometric test; Supplementary Table S4) as described in Materials and Methods. Then, annotation enrichment analysis using DAVID36,37 showed that 146 of 151 genes were annotated to 51 KEGG pathways (all p < 0.1) and detailed description of these pathways was presented in Supplementary Table S5. Specifically, the above enrichment analysis demonstrated a close correlation between the regulating genes and cell death for apoptosis pathway (p = 4.2E-6).
Notably, cell death, an established cancer hallmark, might serve as a promising candidate in prevention and treatment of ovarian cancer. We further explored the relationships between the 12-gene module biomarker and cell death genes from our miRDeathDB27,28 and HADB29 and DeathBase30. A nonrandom amount of overlap was observed between the 151 significantly regulating genes and 727 cell death genes (p = 1.12E-5, hypergeometric test; Fig. 4A), suggesting potential clinical benefit for tumor suppression via regulating cell death. For example, STAT3, contributing to oncogenesis by inhibition of apoptosis, interacts with LCK leading to T-cell transformation by Herpesvirus saimiri (HVS)38. Specifically, 18 of 21 overlapping genes were found to be tightly clustered together pointing to the module biomarker (Fig. 4B). Moreover, majority of the overlapping genes were known to be cancer genes, whose close association with ovarian cancer have been confirmed as presented in detail in Table 3. For example, BCL2L1, as a key protein in regulating programmed cell death or apoptosis, was found to be dysregulated in ovarian cancer cell lines and specimens that promoted cancer progression39. The hsa-miR-335-5p was regarded as an invasion suppressor, whose dysregulation drove cancer transformation by targeting Bcl-w24,40.
Overlap and wiring diagram between the regulatory genes and cell death genes.
(A) Venn plot, showing a nonrandom amount of overlap between the 151 significantly regulatory genes and 727 cell death genes. (B) The interaction wiring of 21 cell death genes and their wiring connections on the 12-gene module. The nodes in red or blue indicated whether the genes have been verified as being related to ovarian cancer or not, respectively. The node sizes indicated the significance of the genes in regulating the 12-gene module.
Taken together, these results provided the additional evidence to support our findings from the aspect of biological importance, demonstrating that based on integrating survival information and differential co-expression between longer- and shorter- survival patients, the network-based survival-associated module biomarker has guidance for the treatment of ovarian cancer, excepting for the diagnosis of ovarian cancer.
Discussion
Ovarian cancer, as a complex disease, is characterized by dysregulation of multiple cellular functions that interact in a complex network environment10. Furthermore, gene intersections and their dynamic wiring, as essential components of network, underlie the orchestration of biological processes10, hence, it is reasonable to perform network-based dynamic modularity analysis for biomarker discovery. Different from the traditional network-based analyses that usually ignore the patients’ survival hazards or the correlations existing between gene expressions41,42, we here facilitated the dynamic responded-intersections based on survival and differential co-expression PPI network between longer- and shorter-survival patients and identified a 12-gene module biomarker for ovarian cancer and further confirmed its predictive ability in internal and external independent datasets. Despite the 12-gene module biomarker was shown to be an independent risk factor for ovarian cancer from age, grade, stage and residual tumor size, the influence of drug on the survival of ovarian cancer patients should be further excluded. Nevertheless, this analysis is consistent with a clinical viewpoint that the rationale behind the biomarker discovery is to find robust and effective biomarker, given that modules play a central role in maintaining network stability.
When the transition from normal state into disease state, network will subject to many forms of disruption and network modules as response element of the disease, which can confer cellular functions. Thus, the functional analysis of module biomarker will enhance our understanding of the underlying mechanisms of the disease. As for the identified 12-gene module biomarker, we further deciphered the underlying biological mechanisms of the biomarker and dissected the wiring diagram between the biomarker and cell death and found the module genes having close interaction with cell death.
Notably, the wiring analysis of module biomarker demonstrated that its 151 regulating interacting neighbors were significantly overlapped with cell death genes, of which 21 overlapping cell death genes interacted closely with the 12-gene module. More specifically, 4 of 21 overlapped cell death genes were miRNAs, which was overwhelming (4/5 = 80%) in the significant regulatory miRNAs, implying that cell death related non-coding RNAs might play an important role in regulating the 12-gene module. And 18 overlapped cell death genes clustered together pointing to the 12-gene module, implying the cooperative behavior for the benefit to increase evades and prevents cell death. These analyses begin to bridge the gap between cancer diagnosis and treatment and pave a clear path from cancer diagnosis to treatment.
In conclusion, our analyses demonstrated the effectiveness and robustness of network-based module analysis for biomarker discovery by integrating survival information and differential co-expression between longer- and shorter- survival patients, highlighting the importance of functional analysis in understanding biomarker prediction and monitoring treatment. Especially, the wiring diagram discovery between biomarker and cell death has made an important step towards transforming from preclinical to clinical assessments.
Methods
Gene expression and clinical data
Gene expression data (Level 3) generated by Agilent platform and clinical data of 419 patients with advanced-stage (stages III and IV), high-grade (grades 3 and 4) ovarian serous cystadenocarcinoma were downloaded from TCGA repository (http://cancergenome.nih.gov/). Notably, this dataset was divided into two subsets: one training set consisting of Batches 9, 11–15 and 17 and one test set consisting of Batches 18–19, 21–22, 24, 27 and 40, as described in detail in Table 4. Separately for each set, the patients were further stratified into the longer- and shorter- survival groups according to the criteria that a 37-month median survival time in ovarian cancer identified by Macmillan Cancer Support (http://www.macmillan.org.uk/) derived from the researches of the Cancer Research UK Cancer Survival Group at the London School of Hygiene and Tropical Medicine. Additionally, a cohort of 129 ovarian cancer patients from GEO database (Accession No. GSE32062)35 was used as an independent external test set. Microarray data was median-normalized and replicate genes were combined by averaging their expression values43,44.
Protein-protein interactions (PPI), RNA-protein interactions and cell death genes
Protein-protein interactions were retrieved from STRING v9.126. To minimize the impact of network size, only high-confidence interactions with String-score ≥0.90 were extracted to construct network, involving 76, 709 interactions12. RNA-protein interactions were integrated from our RAID45 and miRTarBase46 databases.
Cell death genes were manually reviewed from literatures and collected from our miRDeathDB27,28 and HADB29 and DeathBase30 databases. After removing the redundant and unrecognized genes, a total of 727 cell death genes were used for subsequent analysis.
Construction of weighted PPI network
A weighted PPI network was constructed, in which each node (gene  ) was assigned a weight,
) was assigned a weight,  , on the basis of its association with survival of patients using Cox proportional hazards regression model, as follows:
, on the basis of its association with survival of patients using Cox proportional hazards regression model, as follows:

where  represented the significance of association between each gene expression and survival of patients, calculated from univariable Cox proportional hazards regression model.
represented the significance of association between each gene expression and survival of patients, calculated from univariable Cox proportional hazards regression model.  represented the inverse standard normal cumulative distribution function (CDF)41,47. Thus,
represented the inverse standard normal cumulative distribution function (CDF)41,47. Thus,  followed a standard normal distribution, with a smaller p-value corresponding to a larger z-score value.
followed a standard normal distribution, with a smaller p-value corresponding to a larger z-score value.
And each edge (interaction  ) was assigned a weight,
) was assigned a weight,  , on the basis of its degree of differential co-expression of a pair genes (genes
, on the basis of its degree of differential co-expression of a pair genes (genes  and
and  ) between longer- and shorter- survival samples, as follows:
) between longer- and shorter- survival samples, as follows:
First, Pearson correlation coefficient  (or
(or  ) of genes
) of genes  and
and  between patients in longer-survival group (or shorter-survival group) in the training dataset was calculated as
between patients in longer-survival group (or shorter-survival group) in the training dataset was calculated as

where  and
and  represented the expression levels of gene
represented the expression levels of gene  and gene
and gene  in patient
in patient  of longer-survival group (or shorter-survival group);
of longer-survival group (or shorter-survival group);  and
and represented the average expression levels of gene
represented the average expression levels of gene  and gene
and gene  in longer-survival group (or shorter-survival group);
in longer-survival group (or shorter-survival group);  represented the number of patients in longer-survival group (or shorter-survival group).
represented the number of patients in longer-survival group (or shorter-survival group).
Second, the Pearson correlation coefficient ( ) was further transformed into z-score value by using Fisher’s Z transformation
) was further transformed into z-score value by using Fisher’s Z transformation

Then, the degree of differential co-expression of this pair genes (genes  and
and  ) between longer- and shorter-survival groups,
) between longer- and shorter-survival groups,  , was calculated as
, was calculated as

where  and
and  represented the transformed Pearson correlation coefficients in longer- and shorter-survival groups, respectively;
represented the transformed Pearson correlation coefficients in longer- and shorter-survival groups, respectively;  and
and  represented the numbers of patients in longer- and shorter-survival groups, respectively.
represented the numbers of patients in longer- and shorter-survival groups, respectively.
Identification of network-based modules
As we known, the problem of finding the maximal-scoring connected module was NP-hard. To solve this problem and obtain the globally optimal solution, a simulated annealing algorithm47,48 was here introduced to search candidate modules in the weighted network. For each iteration  , the highest-scoring module, denoted as
, the highest-scoring module, denoted as  , was scored by the following formula49:
, was scored by the following formula49:

where  represented the number of genes (
represented the number of genes ( ) and
) and  represented the number of interactions (
represented the number of interactions ( ) in module
) in module  .
.
Those modules overlapping to a very high extent, more than 80%, in comparison to their sizes were further merged concurrently rather than sequentially in order to avoid recalculating the overlap of modules50,51.
To exclude the effect of module sizes on their scores, we randomly sampled gene sets of size  , a permutation experiment using random resampling of 10,000 times was performed to estimate the score mean
, a permutation experiment using random resampling of 10,000 times was performed to estimate the score mean  and standard deviation
and standard deviation  and then
and then  was further adjusted as follows:
was further adjusted as follows:

Survival analysis and module biomarker selection
Survival curves were estimated by the Kaplan-Meier method and compared with log-rank test. Univariate and multivariate survival analyses were performed using the Cox proportional hazard model.
For each candidate module, we calculated an eigengene of the module (EM) value for each sample as the weighted average of gene expression levels of their first principal components52 and then assessed its predictive ability as a predictor of survival after dividing the patients into two subgroups based on the median value of EM values.
Regulating genes identification
According to PPI or RNA-protein interactions, genes direct interacting with module genes were examined by using hypergeometric test and those genes with p < 0.1 were defined as significant regulating genes.
Additional Information
How to cite this article: Jin, N. et al. Network-based survival-associated module biomarker and its crosstalk with cell death genes in ovarian cancer. Sci. Rep. 5, 11566; doi: 10.1038/srep11566 (2015).
References
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2015. CA. Cancer J. Clin. 65, 5–29; 10.3322/caac.21254 (2015).
Slomski, A. Screening women for ovarian cancer still does more harm than good. JAMA 307, 2474–2475; 10.1001/jama.2012.5646 (2012).
Integrated genomic analyses of ovarian carcinoma. Nature 474, 609–615; 10.1038/nature10166 (2011).
Bast, R. C., Jr., Hennessy, B. & Mills, G. B. The biology of ovarian cancer: new opportunities for translation. Nat. Rev. Cancer 9, 415–428; 10.1038/nrc2644 (2009).
Risch, H. A. et al. Population BRCA1 and BRCA2 mutation frequencies and cancer penetrances: a kin-cohort study in Ontario, Canada. J. Natl. Cancer Inst. 98, 1694–1706; 10.1093/jnci/djj465 (2006).
Pal, T. et al. BRCA1 and BRCA2 mutations account for a large proportion of ovarian carcinoma cases. Cancer 104, 2807–2816; 10.1002/cncr.21536 (2005).
Ahmed, A. A. et al. Driver mutations in TP53 are ubiquitous in high grade serous carcinoma of the ovary. J. Pathol. 221, 49–56; 10.1002/path.2696 (2010).
Cantor, J. R. & Sabatini, D. M. Cancer cell metabolism: one hallmark, many faces. Cancer discov. 2, 881–898; 10.1158/2159-8290.CD-12-0345 (2012).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674; 10.1016/j.cell.2011.02.013 (2011).
Barabasi, A. L., Gulbahce, N. & Loscalzo, J. Network medicine: a network-based approach to human disease. Nat. Rev. Genet. 12, 56–68; 10.1038/nrg2918 (2011).
Taylor, I. W. et al. Dynamic modularity in protein interaction networks predicts breast cancer outcome. Nat. Biotechnol. 27, 199–204; 10.1038/nbt.1522 (2009).
Hofree, M., Shen, J. P., Carter, H., Gross, A. & Ideker, T. Network-based stratification of tumor mutations. Nat. Methods 10, 1108–1115; 10.1038/nmeth.2651 (2013).
Ideker, T. & Krogan, N. J. Differential network biology. Mol. Syst. Biol. 8, 565; 10.1038/msb.2011.99 (2012).
Bonnefond, M. L. et al. Calcium signals inhibition sensitizes ovarian carcinoma cells to anti-Bcl-xL strategies through Mcl-1 down-regulation. Apoptosis 20, 535–550; 10.1007/s10495-015-1095-3 (2015).
Chandler, R. L. et al. Coexistent ARID1A-PIK3CA mutations promote ovarian clear-cell tumorigenesis through pro-tumorigenic inflammatory cytokine signalling. Nat. Commun. 6, 6118; 10.1038/ncomms7118 (2015).
Li, L. et al. MiR-17-5p Up-Regulates YES1 to Modulate the Cell Cycle Progression and Apoptosis in Ovarian Cancer Cell Lines. J. Cell. Biochem. 116, 1050–1059; 10.1002/jcb.25060 (2015).
Knutson, K. L., Karyampudi, L., Lamichhane, P. & Preston, C. Targeted immune therapy of ovarian cancer. Cancer Metastasis Rev. 34, 53–74; 10.1007/s10555-014-9540-2 (2015).
Jacobson, M. D., Weil, M. & Raff, M. C. Programmed cell death in animal development. Cell 88, 347–354 (1997).
Denoyelle, C. et al. miR-491-5p-induced apoptosis in ovarian carcinoma depends on the direct inhibition of both BCL-XL and EGFR leading to BIM activation. Cell death dis. 5, e1445; 10.1038/cddis.2014.389 (2014).
Simonin, K. et al. Platinum compounds sensitize ovarian carcinoma cells to ABT-737 by modulation of the Mcl-1/Noxa axis. Apoptosis 18, 492–508; 10.1007/s10495-012-0799-x (2013).
Mathew, R. et al. Autophagy suppresses tumorigenesis through elimination of p62. Cell 137, 1062–1075; 10.1016/j.cell.2009.03.048 (2009).
Oliveira-Ferrer, L. et al. c-FOS suppresses ovarian cancer progression by changing adhesion. Br. J. Cancer 110, 753–763; 10.1038/bjc.2013.774 (2014).
Wang, H. et al. Effect of miR-335 upregulation on the apoptosis and invasion of lung cancer cell A549 and H1299. Tumour Biol. 34, 3101–3109; 10.1007/s13277-013-0878-9 (2013).
Cao, J. et al. miR-335 represents an invasion suppressor gene in ovarian cancer by targeting Bcl-w. Oncol. Rep. 30, 701–706; 10.3892/or.2013.2482 (2013).
Hafezi, F. et al. The absence of c-fos prevents light-induced apoptotic cell death of photoreceptors in retinal degeneration in vivo. Nat. Med. 3, 346–349 (1997).
Franceschini, A. et al. STRING v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 41, D808–D815; 10.1093/nar/gks1094 (2013).
Li, Y. et al. Connect the dots: a systems level approach for analyzing the miRNA-mediated cell death network. Autophagy 9, 436–439; 10.4161/auto.23096 (2013).
Xu, J. & Li, Y. H. miRDeathDB: a database bridging microRNAs and the programmed cell death. Cell Death Differ. 19, 1571; 10.1038/cdd.2012.87 (2012).
Moussay, E. et al. The acquisition of resistance to TNFalpha in breast cancer cells is associated with constitutive activation of autophagy as revealed by a transcriptome analysis using a custom microarray. Autophagy 7, 760–770; 10.4161/auto.7.7.15454 (2011).
Diez, J., Walter, D., Munoz-Pinedo, C. & Gabaldon, T. DeathBase: a database on structure, evolution and function of proteins involved in apoptosis and other forms of cell death. Cell Death Differ. 17, 735–736; 10.1038/cdd.2009.215 (2010).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504; 10.1101/gr.1239303 (2003).
Jiang, L. et al. Overexpression of proto-oncogene FBI-1 activates membrane type 1-matrix metalloproteinase in association with adverse outcome in ovarian cancers. Mol. cancer 9, 318; 10.1186/1476-4598-9-318 (2010).
Woenckhaus, J. et al. Prognostic value of PIK3CA and phosphorylated AKT expression in ovarian cancer. Virchows Arch. 450, 387–395; 10.1007/s00428-006-0358-3 (2007).
Aya Sultan, C. Y. W., George E., Duran, E., Brian Francisco, Jonathan S., Berek & Branimir, I. Sikic. Abstract 1401: SYK promotes tumor progression in ovarian cancer cell lines. Cancer Res. 71, 1401; 10.1158/1538-7445. (2011).
Yoshihara, K. et al. High-risk ovarian cancer based on 126-gene expression signature is uniquely characterized by downregulation of antigen presentation pathway. Clin. Cancer Res. 18, 1374–1385; 10.1158/1078-0432.CCR-11-2725 (2012).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57; 10.1038/nprot.2008.211 (2009).
Dennis, G., Jr. et al. DAVID: Database for Annotation, Visualization and Integrated Discovery. Genome Biol. 4, P3; 10.1186/gb-2003-4-5-p3 (2003).
Heck, E. et al. T-cell growth transformation by herpesvirus saimiri is independent of STAT3 activation. J. Virol. 79, 5713–5720; 10.1128/JVI.79.9.5713-5720.2005 (2005).
Kurita, T. et al. Mitochondrial transcription factor A regulates BCL2L1 gene expression and is a prognostic factor in serous ovarian cancer. Cancer Sci. 103, 239–444; 10.1111/j.1349-7006.2011.02156.x (2012).
Sorrentino, A. et al. Role of microRNAs in drug-resistant ovarian cancer cells. Gynecol. Oncol. 111, 478–486; 10.1016/j.ygyno.2008.08.017 (2008).
Li, J., Roebuck, P., Grunewald, S. & Liang, H. SurvNet: a web server for identifying network-based biomarkers that most correlate with patient survival data. Nucleic Acids Res. 40, W123–W126; 10.1093/nar/gks386 (2012).
Segura, M. F. et al. Melanoma MicroRNA signature predicts post-recurrence survival. Clin. Cancer Res. 16, 1577–1586; 10.1158/1078-0432.CCR-09-2721 (2010).
Wang, D. et al. Evaluation of cDNA microarray data by multiple clones mapping to the same transcript. OMICS 13, 493–499; 10.1089/omi.2009.0077 (2009).
Bonome, T. et al. A gene signature predicting for survival in suboptimally debulked patients with ovarian cancer. Cancer Res. 68, 5478–5486; 10.1158/0008-5472.CAN-07-6595 (2008).
Zhang, X. et al. RAID: a comprehensive resource for human RNA-associated (RNA-RNA/RNA-protein) interaction. RNA 20, 989–993; 10.1261/rna.044776.114 (2014).
Hsu, S. D. et al. miRTarBase update 2014: an information resource for experimentally validated miRNA-target interactions. Nucleic Acids Res. 42, D78–D85; 10.1093/nar/gkt1266gkt1266 (2014).
Ideker, T., Ozier, O., Schwikowski, B. & Siegel, A. F. Discovering regulatory and signalling circuits in molecular interaction networks. Bioinformatics 18 Suppl 1, S233–S240 (2002).
Kirkpatrick, S., Gelatt, C. D., Jr. & Vecchi, M. P. Optimization by simulated annealing. Science 220, 671–680; 10.1126/science.220.4598.671 (1983).
Kim, Y. et al. Principal network analysis: identification of subnetworks representing major dynamics using gene expression data. Bioinformatics 27, 391–398; 10.1093/bioinformatics/btq670 (2011).
Nepusz, T., Yu, H. & Paccanaro, A. Detecting overlapping protein complexes in protein-protein interaction networks. Nat. Methods 9, 471–472; 10.1038/nmeth.1938 (2012).
Bader, G. D. & Hogue, C. W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinformatics 4, 2; 10.1186/1471-2105-4-2 (2003).
Liu, W., Li, L. & Li, W. Gene co-expression analysis identifies common modules related to prognosis and drug resistance in cancer cell lines. Int. J. Cancer 135, 2795–2803; 10.1002/ijc.28935 (2014).
Das, H. et al. Quantitation of Fas and Fas ligand gene expression in human ovarian, cervical and endometrial carcinomas using real-time quantitative RT-PCR. Br. J. Cancer 82, 1682–1688; 10.1054/bjoc.2000.1118 (2000).
Hein, S. et al. Expression of Jun and Fos proteins in ovarian tumors of different malignant potential and in ovarian cancer cell lines. Oncol. Rep. 22, 177–183; 10.3892/or_00000422 (2009).
Ravikumar, S., Perez-Liz, G., Del Vale, L., Soprano, D. R. & Soprano, K. J. Insulin receptor substrate-1 is an important mediator of ovarian cancer cell growth suppression by all-trans retinoic acid. Cancer Res. 67, 9266–9275; 10.1158/0008-5472.CAN-07-2088 (2007).
Fan, Y. et al. NFKB1 insertion/deletion promoter polymorphism increases the risk of advanced ovarian cancer in a Chinese population. DNA Cell Biol. 30, 241–245; 10.1089/dna.2010.1107 (2011).
Campbell, I. G. et al. Mutation of the PIK3CA gene in ovarian and breast cancer. Cancer Res. 64, 7678–7681; 10.1158/0008-5472.CAN-04-2933 (2004).
Zhang, L. et al. Integrative genomic analysis of phosphatidylinositol 3’-kinase family identifies PIK3R3 as a potential therapeutic target in epithelial ovarian cancer. Clin. Cancer Res. 13, 5314–5321; 10.1158/1078-0432.CCR-06-2660 (2007).
Niesporek, S. et al. [NF-kappaB subunit p65/RelA expression in ovarian carcinoma: prognostic impact and link to COX-2 overexpression]. Verh. Dtsch. Ges. Pathol. 91, 243–249 (2007).
Wiener, J. R. et al. Decreased Src tyrosine kinase activity inhibits malignant human ovarian cancer tumor growth in a nude mouse model. Clin. Cancer Res. 5, 2164–2170 (1999).
Gest, C. et al. Ovarian cancer: Stat3, RhoA and IGF-IR as therapeutic targets. Cancer Lett. 317, 207–217; 10.1016/j.canlet.2011.11.026 (2012).
McCann, G. A. et al. Targeting constitutively-activated STAT3 in hypoxic ovarian cancer, using a novel STAT3 inhibitor. Oncoscience 1, 216–228 (2014).
Miles, G. D., Seiler, M., Rodriguez, L., Rajagopal, G. & Bhanot, G. Identifying microRNA/mRNA dysregulations in ovarian cancer. BMC Res. Notes 5, 164; 10.1186/1756-0500-5-164 (2012).
Acknowledgements
This work was supported by the National High Technology Research and Development Program of China (2014AA021102), the Major State Basic Research Development Program of China (2014CB910504), the National Natural Science Foundation of China (31100901), the China Postdoctoral Science Foundation funded project (2013M531064, 2014T70363), the Heilongjiang Postdoctoral Foundation (LBH-Z12171) and the Scientific Research Fund of Heilongjiang Provincial Education Department (12541426).
Author information
Authors and Affiliations
Contributions
D. Wang, X. Li and K. Li conceived and designed the experiments. N. Jin, H. Wu, Z. Miao, Y. Huang, Y. Hu, X. Bi, D. Wu, K. Qian, L. Wang and C. Wang performed the experiments. D. Wang, H. Wang and N. Jin wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Jin, N., Wu, H., Miao, Z. et al. Network-based survival-associated module biomarker and its crosstalk with cell death genes in ovarian cancer. Sci Rep 5, 11566 (2015). https://doi.org/10.1038/srep11566
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep11566
- Springer Nature Limited
This article is cited by
-
Analyzing the regulation of miRNAs on protein-protein interaction network in Hodgkin lymphoma
BMC Bioinformatics (2019)
-
A computational framework for complex disease stratification from multiple large-scale datasets
BMC Systems Biology (2018)
-
Identification of oral cancer related candidate genes by integrating protein-protein interactions, gene ontology, pathway analysis and immunohistochemistry
Scientific Reports (2017)
-
A VEGF-dependent gene signature enriched in mesenchymal ovarian cancer predicts patient prognosis
Scientific Reports (2016)
-
A powerful weighted statistic for detecting group differences of directed biological networks
Scientific Reports (2016)