Abstract
The progression of cancers and neurodegenerative disorders is largely defined by a set of molecular determinants that are either complementarily deregulated, or share remarkably overlapping functional pathways. A large number of such molecules have been demonstrated to be involved in the progression of both diseases. In this review, we particularly discuss our current knowledge on p53, cyclin D, cyclin E, cyclin F, Pin1 and protein phosphatase 2A, and their implications in the shared or distinct pathways that lead to cancers or neurodegenerative diseases. In addition, we focus on the inter-dependent regulation of brain cancers and neurodegeneration, mediated by intercellular communication between tumor and neuronal cells in the brain through the extracellular microenvironment. Finally, we shed light on the therapeutic perspectives for the treatment of both cancer and neurodegenerative disorders.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
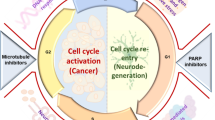
Cancer and neurodegenerative diseases represent one of the most chronic physiological ailments. Aging, characterized by the deterioration of physiological functions necessary for survival and fertility, is considered as a major risk factor for the two disorders [1,2,3]. Cancer has been associated with generalized hallmarks such as sustenance of proliferative signaling, evasion of growth suppressors, resistance to cell death, acquisition of replicative immortality, induction of angiogenesis, and activation of invasion and metastasis. Interestingly, current research has indicated parameters such as deregulated cellular energetics and avoidance of immune destruction, as pertinent hallmarks. These features are effectuated by genome instability, mutations and/ or tumor-promoting inflammation [4,5,6] (Fig. 1). Neurodegeneration is characterized by dysfunction and loss of neurons [7], impairment of synaptic plasticity, proteinopathies, which include misfolded amyloid-β (Aβ) and tau in Alzheimer’s disease (AD), α-synuclein in Parkinson’s disease (PD), and their aggregates [8,9,10,11], as well as progressive muscle atrophy or muscle wasting, which causes memory deficits, cognitive failures, and movement disorders [7] (Fig. 1).
Featured hallmarks of cancer and neurodegenerative diseases. Pathophysiological features of two representative age-related diseases, cancer and neurodegenerative diseases are shown. Mechanisms are inversely undergone in these two diseases to lead to cell survival and cell death in cancer and neurodegenerative diseases, respectively. DNA damage, cell cycle aberrations, redox imbalance, inflammation, and immunity are closely associated as emerging shared characteristics between cancer and neurodegenerative diseases
Inverse comorbidities between cancer and neurodegenerative diseases have been reported by many clinical and epidemiological studies [12,13,14,15,16,17,18,19,20,21,22,23,24]. In this light, molecular mechanisms that operate inversely in these two disorders (one leading to enhanced resistance to cell death and the other to a higher risk of cell death) would form effective means of diagnosis and prognosis at the physiological level (Fig. 1). A recent study, which performed transcriptomic meta-analyses of three neurodegenerative diseases (AD, PD, and schizophrenia) and three kinds of cancers (lung, prostate, and colorectal cancer) reported a significant overlap between the genes upregulated in the neurodegenerative diseases and downregulated in cancer, and between the genes downregulated in the neurodegenerative diseases and upregulated in cancer [16].
Multiple signaling pathways that regulate cell death and survival have been well investigated in tumorigenesis, including DNA damage, cell cycle aberrations, inflammation, immunity, and oxidative stress; these pathways have now been shown to be associated with neurodegenerative diseases as well [18, 25,26,27,28,29,30,31] (Fig. 1). In addition, aberrant expression or mutations in genes such as α-synuclein, phosphatase and tensin homolog (PTEN), PTEN induced kinase 1 (PINK1; parkinsonism associated deglycase 6, PARK6), DJ-1 (PARK7), leucine rich repeat kinase 2 (LRRK2; PARK8), microtubule-associated protein tau (MAPT), amyloid precursor protein (APP), presenilin 1/2 (PSEN1/2), and cyclin-dependent kinase 5 (CDK5), which play essential roles in neurodegeneration, are also observed in cancer [32].
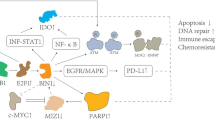
Over the past decade, accumulating evidences have demonstrated an intriguing relationship between cancer and neurodegenerative diseases. Better understanding of the relationship between the two will provide novel avenues for the study of these age-related diseases. In this review, we will discuss the current knowledge on the shared or distinct roles of overlapping molecules that have been significantly correlated with the pathophysiology of both cancer and neurodegenerative diseases, such as p53, cyclin D, cyclin E, cyclin F, peptidyl-prolyl cis–trans isomerase (PPIase) NIMA (Never in Mitosis A)-interacting 1 (Pin1), and protein phosphatase 2A (PP2A) (Fig. 2). In addition, we describe the inter-dependent regulation of brain cancers and neurodegeneration through intercellular communications between tumor and neuronal cells in the brain. Furthermore, this review provides some perspectives into the application towards pharmacological therapeutics for both cancer and neurodegenerative diseases.
Changes in overlapping molecules in cancer and neurodegenerative diseases. Cyclin D and cyclin E are upregulated whereas protein phosphatase 2A (PP2A) is downregulated in both diseases. p53 is downregulated in cancer but inversely upregulated in neurodegenerative diseases. Peptidyl-prolyl cis–trans isomerase NIMA-interacting 1 (Pin1) is mainly upregulated in cancer and Parkinson’s disease (PD) but downregulated in Alzheimer’s disease (AD). Cyclin F is downregulated in cancer, while its mutant form is found in amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD). NIMA never in mitosis A
Overlapping molecules between cancer and neurodegenerative diseases
p53
The transcription factor p53 is an extensively studied tumor suppressor [33,34,35], and is known to be associated with around 50% of all human malignancies. In most of these cases, p53 gene has been reported to contain missense mutations [36,37,38,39]. The mutant p53 proteins no longer have tumor suppressor activity, and obtain several gain-of-functions such as invasion [40,41,42,43,44,45,46,47,48], enhanced migration [42, 49,50,51,52], anchorage-independent growth [53,54,55,56,57,58], propagation of cell cycle [59,60,61,62,63,64,65], cell survival and avoidance of cell death [66,67,68,69,70,71,72,73,74,75,76], genomic instability [77,78,79,80,81,82], and angiogenesis [83,84,85]. A commonly occurring mutant form of p53, p53-R273H, contributes to the impaired detoxification of reactive oxygen species (ROS) by decreasing the nuclear factor erythroid 2 (NF-E2)-related factor 2 (NRF2)-mediated expression of phase 2 ROS-detoxifying enzymes, quinone oxidoreductase 1 (NQO1) and heme oxygenase-1 (HO-1), which resulted in a reduced antioxidant response and imbalance of redox homeostasis in lung or colon cancer cells [70, 86, 87] (Fig. 3). Overexpression of another mutant, p53-G245D, upregulated a transcription factor called forkhead box protein M1 (FOXM1), which exerted oncogenic properties [88]. However, another study showed that the enhanced level of FOXM1 downregulates ROS levels by increasing antioxidant enzymes like superoxide dismutase (SOD) and thioredoxin-dependent peroxide reductase, peroxiredoxin 3 (PRDX3) [89]. These complicated results of mutant p53 on redox homeostasis could warrant more careful considerations when targeting dual factors p53 and redox regulation for the treatment of cancers. Furthermore, mutant p53 proteins are rather reluctant to degradation compared to wild-type p53 proteins, and thus the accumulated mutant p53 proteins are often a major therapeutic target for cancer treatment [36, 37, 85, 90,91,92].
Mechanisms of overlapping molecules in cancer and neurodegenerative diseases. (Down) Mutant p53 inhibits the nuclear factor erythroid 2 (NF-E2)-related factor 2 (NRF2)-mediated antioxidant enzymes, and thus induces reactive oxygen species (ROS) production. Low-molecular-weight (LMW) cyclin E cannot translocate into the nucleus, and forms LMW cyclin E cyclin-dependent kinase 2 (CDK2) complex in the cytoplasm, thereby activating oncogenic functions, such as tumor cell invasion and metastasis. (Top left) Amyloid-β (Aβ), Aβ fibrils and plaques are formed from amyloid precursor protein (APP) via amyloidogenic processing. Cyclin D1 promotes tau phosphorylation and induces apoptosis through a caspase 3-mediated pathway. PP2A dephosphorylates phosphorylated tau, and glycogen synthase kinase 3 beta (GSK3β) phosphorylates tau. Reduced activity of superoxide dismutase (SOD) and glutathione reductase (GR) induces the increase of ROS production, which leads to the conformational change of p53, and this unfolded p53 is also observed in AD. (Top right) Mutant parkin and glutamate excitotoxicity increase cyclin E accumulation, which induces apoptosis. In addition, p53 also induces the upregulation of apoptotic proteins, such as Bcl-2 associated X (Bax) and caspases 3, which is observed in the PD brain. The interaction between Pin1 and synphilin-1 (an α-synuclein-binding protein) enhances the formation of α-synuclein inclusions, and this α-synuclein inclusion formation can be inhibited by PP2A. ER endoplasmic reticulum, HO-1 heme oxygenase-1, NQO1 quinone oxidoreductase 1, PM plasma membrane. Molecule name marked in purple indicates studies that involve exogenous proteins
Unlike the role of p53 in cancer, the level and activity of p53 in neurodegenerative diseases have been shown to be substantially increased [93,94,95]. Brains of AD patients and model mice showed increased levels of p53 [96,97,98,99] and apoptotic neuronal cell death [100,101,102]. In double transgenic AD mice that express the mutants of amyloid precursor protein/presenilin (APP/PS) and accumulate Aβ [103], cerebral gray matter displayed a positive correlation between the p53 level and accumulated Aβ level [104]. In addition, the triggering receptor expressed on myeloid cells 2 (TREM2), an AD-associated molecule, was reported to be upregulated positively in a p53-dependent manner in vitro [105]. Similar to AD, p53 level and activity were also increased in the brains of PD patients and model mice [106]. Specifically, the caudate nucleus, but not the substantia nigra, putamen, and cerebral cortex, of the PD patient brains showed a significantly enhanced p53 protein level [106]. The p53-dependent apoptosis-related proteins, such as Bcl-2 associated X (Bax) and caspase-3, were increased in the PD brains [107, 108].
Genetic mutations in p53 have not been reported for neurodegenerative diseases. However, functionally compromised variants of p53 such as those with an altered tertiary structure (called unfolded p53 or conformational mutant p53) have been distinctly observed in AD patients [109, 110] and in older APPswe/PS1-A246E AD transgenic mice [111], but not in non-AD individuals, including PD patients [109]. In human neuroblastoma cell line SH-SY5Y overexpressing APP, an increased level of unfolded p53 was observed. This was shown to be associated with a lack of p53 pro-apoptotic activity (Fig. 3) and an impairment in neuronal responses against acute cytotoxic injury [112].
Oxidative imbalance has also been demonstrated as a distinctive feature in neurodegenerative diseases [113,114,115,116,117,118,119]. This imbalance has been observed to be mediated by reduced activities of SOD and glutathione reductase during AD [113]. Importantly, it was noted that the reduced enzyme activity of SOD corresponded with increased levels of unfolded p53, suggesting that ROS possibly contributes to p53 conformational change in AD [113] (Fig. 3). In a PD model induced by the treatment of 1-methyl-4-phenylpyridinium (MPP +), the expression of sestrin-2, an antioxidant protein, was increased by MPP + -induced p53 activation, and such enhanced expression of sestrin-2 protects cells against ROS, suggesting a novel role of p53 in PD [119]. Therefore, counteracting oxidative stress or improving cellular antioxidative properties would provide effective therapeutic alternatives for neurodegenerative disorders.
Cyclins
Many studies have demonstrated that dysregulated destruction of cell cycle regulators, many of which play a role in either tumor suppression or tumorigenesis, is tightly linked to cancer initiation and progression. Cyclins are known to regulate cell cycle by modulating the activity of cyclin-dependent kinases (CDKs). Deregulation of the cell cycle through changes in the activity of cell cycle CDKs or their regulators form essential determinants of human cancers [120,121,122,123,124,125,126,127].
Beyond their role in cell cycle regulation, cyclins contribute immensely to the cellular aspects of the terminally differentiated neurons [128]. Cell cycle-independent roles of cyclins, including cyclin E and cyclin Y, have been reported in postmitotic neuronal physiology [129,130,131]. Cyclin E deficiency has been shown to reduce the number of synapses and spine volume, and also to impair long-term potentiation and memory [131]. Knockdown of cyclin Y in hippocampal neurons has been reported to enhance activity-dependent synaptic delivery of α-amino-3-hydroxy-5-methyl-4-isoxazolepropionate (AMPA) receptors and long-term potentiation [129]. Interestingly, such aberrations in the cell cycle components and the resulting neuronal cell death have also been found in neurodegeneration and neurodegenerative diseases [132,133,134,135].
Cyclin D
Cyclin D, which controls the entry from the quiescence (G0) to G1 phase of the cell cycle [136], and its associated CDKs, CDK4 and CDK6, are overexpressed and hyperactive, respectively, in most tumors [124, 137] (Fig. 4). Mutant cyclin D1 knockin mice, in which CDK4 and CDK6 are not activated, were resistant to breast tumors initiated by the activated erbB-2 oncogene [122, 127]. Mice lacking cyclin D1 were also resistant to breast cancers induced by the erbB-2 and ras oncogenes, but were sensitive to breast cancers induced by other oncogenes like c-myc or Wnt-1. Furthermore, knockdown of cyclin D1 induced oxidative imbalance by elevating intracellular ROS levels in cancer cells, which promoted the senescence of cancer cells through a retinoblastoma-independent pro-senescence pathway [138]. These investigations suggest that anti-breast cancer therapy targeting cyclin D1 could be very specific to breast cancers depending on the activated pathways [127].
Cell cycle and cyclins, such as cyclin D, cyclin E, and cyclin F in cancer and neurodegenerative diseases. Dysregulated cell cycle is tightly related with cancer initiation as well as neurodegeneration. Cyclin D1, associated with CDK4 and CDK6, modulates the entry from quiescence (G0) to the G1 phase of the cell cycle, and is also linked to the progression of neurodegeneration. Cyclin E plays a role in the initiation of DNA replication at the G1/S checkpoint, and also causes neurodegeneration in the postmitotic neurons. Ubiquitin-specific peptidase 27 (USP27), a cyclin E interactor, increases the stability of cyclin E by inhibiting the ubiquitination and subsequent degradation of cyclin E. SCF-cyclin F and APC/C-Cdh1 are controlled by a negative reciprocal feedback circuit, which controls S phase entry. In addition, cyclin F interacts with stem-loop binding protein (SLBP) and promotes SLBP degradation during G2. Blockade of the interaction between cyclin F and SLBP increases apoptosis upon genotoxic stress in G2 phase. Pin1 is a positive regulator of cyclin D1, and both Pin1 and cyclin D1 are upregulated in many cancers. Pin1 expression is negatively regulated by small non-coding microRNAs (miRNAs), including miR-200b, miR-296-5p, miR-874-3p, miR-140-5p, and miR-370. SCF Skp1–Cul1–F-box, APC/C anaphase promoting complex/cyclosome, Chd1 Cdc20 homologue 1. Molecule names marked in purple indicate studies that involve exogenous proteins
CDK4, a cyclin D-associated CDK, was observed to be increased in the brains of AD patients [139]. In addition, the upregulation of cyclin D was reported to be linked to increases of phosphorylated tau and caspase 3, which led to apoptosis in cultured hippocampal neurons, suggesting that cyclin D increment could be a crucial factor in the progression of neurodegenerative pathology [140] (Fig. 3). Conversely, in cyclin D1-deficient mutant mice, phenotypically characterized by small eyes with thin retinas, a reduced proliferation of retina cells and an increased photoreceptor cell death were observed [141]. In addition, stimuli inducing cortical neuronal degeneration reduced the protein level of cyclin D1 [142]. Recently, neuronal gain- or loss-of-function of cyclin D/CDK4 in Drosophila caused neurodegeneration [143]. In addition, cyclin D/CDK4-mediated neurodegeneration was shown to be mediated by altered mitochondrial function and an accompanying increase in ROS [143].
Cyclin E
Cyclin E plays a role in the initiation of DNA replication at the G1/S checkpoint [144] and is a regulatory subunit of CDK2 [145]. Cyclin E was reported to be overexpressed in many types of cancers, such as breast cancer [146], non-small-cell lung cancer [147], colorectal carcinomas [148], lymphomas [149], acute myelogenous leukemia [150], gastric carcinomas [151], and osteosarcoma [152]. Over 10% of female transgenic mice overexpressing cyclin E developed mammary carcinoma at around 8–13 months of age [153]. Tumorigenesis mediated by cyclin E overexpression likely involves genomic instability [154,155,156,157] as cyclin D1 overexpression also induces genomic instability [158]. Ubiquitin-specific peptidase 27 (USP27) was recently identified as a cyclin E interactor, and was reported to increase cyclin E stability through the negative regulation of cyclin E ubiquitination [159] (Fig. 4). Depletion of USP27 inhibited the migration and metastasis of hepatocellular carcinoma and the tumor growth, suggesting that USP27 is a novel therapeutic target for cancers involving cyclin E [159].
Besides the 50 kDa full-length cyclin E, tumor-specific low-molecular-weight (LMW) cyclin E isoforms, ranging from 33 to 45 kDa, have been found to be accumulated in cancer cells [146, 160,161,162,163,164,165,166]. LMW cyclin E has mostly lost its N-terminal nuclear localization signal [167], which results in the cytoplasmic accumulation of cyclin E [168] (Fig. 3). It exerts oncogenic functions through properties such as hyperactivation of CDK2 due to more stable LMW cyclin E/CDK2 complex formation [169], the resistance of LMW cyclin E/CDK2 complex to inhibitors p21 and p27 [170], altered substrate interactions [171], and cytoplasmic novel interactions that differ from the full-length cyclin E [172]. Like cytoplasmic LMW cyclin E, excessive cytoplasmic cyclin D1 also exerts oncogenic functions by promoting tumor cell invasion and metastasis [173,174,175] through cytoplasmic interactions [175, 176].
Cyclin E is a substrate of the parkin E3 ubiquitin ligase. Mutations in the parkin gene are the most common cause of PD, and upregulates cyclin E and CDK2 [177,178,179]. Glutamate excitotoxicity has also been implicated in PD [180, 181]. Like parkin mutations, the glutamatergic excitotoxin kainate treatment increased cyclin E accumulation in cultured neurons, and this increase was further enhanced in parkin knockdown neurons, which resulted in the promotion of apoptosis (Fig. 3). Conversely, parkin overexpression retarded the cyclin E accumulation in cultured neurons treated with excitotoxin, and protected the neurons from the kainate-induced excitotoxicity [182]. Cyclin E expression is also related to AD [183,184,185], for example, expression of cyclin E in the brain induced cell cycle activation and led to neurodegeneration of postmitotic neurons in a Drosophila tauopathy AD model [184] (Fig. 4). In addition, Aβ treatment in cultured neurons increased ROS production and activated the mammalian target of rapamycin (mTOR) complex 1, thereby leading to the expression of cell cycle regulatory proteins such as cyclin D1/CDK4 and cyclin E/CDK2 [185].
Cyclin F
Cyclin F, also known as F-box only 1 (FBXO1), was first reported as the F-box family of proteins, which contain an F-box motif [186]. F-box proteins are the substrate-recognition subunits of Skp1–Cul1–F-box (SCF) E3 ubiquitin ligase complexes, and thus in the SCF-cyclin F complex, cyclin F recognizes target proteins and mediates the ubiquitination of target proteins for proteolysis [187, 188]. Cyclin F is also a member of the cyclin family; however, unlike other cyclins, which regulate the cell cycle in concert with the activity of their associated CDKs, cyclin F does not require CDK activity to regulate cell cycle-associated functions. The SCF-cyclin F complex controls the cell cycle through a tightly regulated ubiquitin-mediated proteolysis of centrosomal protein of 110 kDa (CP110), which prevents centrosomal duplications [189]. In addition, cyclin F in S phase regulates the ubiquitination and subsequent degradation of Cdc20 homologue 1 (Chd1), a substrate adaptor protein of the anaphase promoting complex/cyclosome (APC/C), while cyclin F in G1 phase is ubiquitinated and subsequently degraded by the APC/C-Chd1 complex [190] (Fig. 4). In addition, cyclin F interacts with stem-loop binding protein (SLBP) through Arg97 (R97) and Leu99 (L99) in SLBP, and regulates SLBP degradation during G2 [191]. Disruption of the interaction between SLBP and cyclin F by expressing SLBP (RL97/99AA) in G2 led to increased apoptosis upon genotoxic stress [191]. The crucial roles of F-box proteins, including cyclin F, in tumorigenesis are gradually becoming acknowledged owing to their pivotal roles in such cases of cell cycle regulation and genome stability [189, 192]. Recently, it was discovered that cyclin F is upregulated under metabolic stress conditions and inhibits tumorigenesis mediated by an oncogenic mutant form of isocitrate dehydrogenase 1 (IDH1), IDH1-R132H, in glioma [193]. Indeed, it was also reported that cyclin F is downregulated in hepatocellular carcinomas, and low cyclin F expression is correlated with poor survival and recurrence-free survival of hepatocellular carcinoma patients [194], supporting that, unlike other cyclins, cyclin F acts as a tumor suppressor and could be further investigated as a promising prognostic marker for hepatocellular carcinoma.
A recent study using whole-exome sequencing identified mutations in the cyclin F gene, ccnf, in the relatives of patients with amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD) [195], which have a common pathological feature of aberrant accumulation of ubiquitinated transactive response (TAR) DNA-binding protein 43 (TDP-43) [196,197,198]. These findings drive the need to investigate whether TDP-43 is a substrate of the SCF-cyclin F E3 ubiquitin ligase complex. An ALS/FTD-causing pathogenic mutation in cyclin F at amino acid position 621 from serine to glycine (Cyclin F-S621G) was shown to increase the specific ubiquitination at lysine-48 of proteins, which led to the accumulation of lysine-48-ubiquitinated proteins and the impairment of autophagic degradation [199], indicating autophagy to be a degradative mechanism underlying the pathogenesis of ALS/FTD. The roles of cyclin F, which acts as a cyclin as well as an F-box protein, have not been explored in this context, and thus can be further investigated to understand their relevance in mediating neurodegenerative diseases.
Pin1
Pin1 is a regulatory protein of cyclin D1, which is a major regulator of G1 checkpoint progression. Like cyclin D1, Pin1 has been reported to be overexpressed in various cancers, including breast, colon, liver, and lung cancers. Further, cyclin D1 and Pin1 expression levels have been shown to correlate positively in such cancers [200, 201], thereby indicating Pin1 as a potential tumor-promoting factor. Accordingly, Pin1 expression and tumor progression have also been positively correlated in brain, breast, cervical, colon, liver, and prostate cancers [201,202,203,204].
Pin1 is a transcriptional target of the E2 factor (E2F). The E2F promotes Pin1 expression by binding to the E2F-binding sites of the Pin1 gene promoter [205]. In addition, several studies have demonstrated that small non-coding microRNAs (miRNAs), including miR-200b in breast tumor [206], miR-296-5p in prostate cancer [207], miR-874-3p [208] and miR-140-5p [209] in hepatocellular carcinoma, and miR-370 in esophageal squamous cell carcinoma [210], negatively regulate Pin1 expression (Fig. 4). In other words, suppression of such Pin1-targeting miRNAs leads to Pin1 overexpression in various cancers.
Alongside the tumor-promoting function, Pin1 has also been suggested to bear conditional tumor suppressor activity. Pin1 binds to and negatively regulates the protein expression levels of cyclin E [211, 212], whose overexpression mediates tumorigenesis and involves genomic instability [154,155,156,157] (Fig. 4). Many therapeutic studies for treating cancers have developed Pin1 inhibitors based on the fact that Pin1 is a generally recognized tumor-promoting factor [213,214,215,216,217,218,219,220,221,222,223,224,225,226]. Considering that Pin1 has also been reported to have a tumor suppressing function, Pin1-directed inhibitors must be carefully implicated in cancers.
In another paradigm, Pin1 has been known to bind to phosphorylated tau in normal and AD brain extracts, and soluble Pin1 protein has been found to be negligible in AD brains [227]. Pin1 facilitates the dephosphorylation of tau by PP2A [228]. Accordingly, Pin1 expression is inversely correlated with neurofibrillary hyperphosphorylated tau aggregates in AD [229]. Furthermore, Pin1 knockout mice caused tau hyperphosphorylation and tau filament formation [229], and also enhanced amyloidogenic APP processing and selectively increased insoluble Aβ in brains [230]. Similarly in the synaptopathy aspect, Pin1 proteins are decreased in the synapses of AD patients and AD mice brains, and blocking Pin1 activity causes the degradation of a major postsynaptic density organizer, Shank3, resulting in the disruption of synapse structure and thus plasticity. These data indicate that loss of Pin1 activity could lead to deficits in synapse function and plasticity during AD development [231]. Taken together, Pin1 plays a pivotal role in protecting neurodegeneration, and thus could be used as a promising therapeutic target for AD.
Unlike in AD, Pin1 is upregulated in a 1-methyl-4-phenyl-1,2,3,6-tetrahydropyridine (MPTP)-induced PD mouse model and human PD brains [232]. Pin1 was reported to be involved in Lewy body formations in PD [233]; it locates to 50–60% of the Lewy bodies, which are cytosolic inclusions containing α-synuclein aggregates, in PD patient brains [233]. In addition, Pin1 interacts with an α-synuclein-binding protein synphilin-1 [234], which resultantly enhances the interaction between α-synuclein and synphilin-1 and thus the formation of α-synuclein inclusions [233] (Fig. 3). Given that Pin1 is downregulated and upregulated in AD and PD, respectively, precise modulations of Pin1 levels depending on the biological contexts might be one of the crucial factors to be considered in differential therapeutic strategies for treating AD and PD.
PP2A
PP2A, a member of serine/threonine protein phosphatase, is a tumor suppressor [235,236,237,238] and a master regulator of the cell cycle known to dephosphorylate over 300 substrates related to the cell cycle [239]. Partial reduction of PP2A-Aα subunit expression to ~ 50% of normal levels induced anchorage-independent growth and tumorigenicity, whereas over 63% reduction of PP2A-Aα expression resulted in apoptosis [236]. In addition, a tenfold reduction of PP2A-Aα expression level was observed in almost half of the glioma samples studied [240]. A cancer-associated mutation in the PP2A-Aα subunit, PP2A Aα-E64D, increased the incidence of lung cancer by 50–60% in mice [241], further supportive of PP2A as a tumor suppressor (Fig. 2). Another mutation of PP2A-Aα, PP2A-Aα-W257G, was shown to promote cancer cell migration [242]. Apart from the PP2A-Aα subunit, mutations in the PP2A-B55α regulatory subunit have been identified in prostate cancer [243].
The phosphorylation level of proteins maintained by the activity of kinases and phosphatases is an important factor for regulating brain function, and PP2A is the most important phosphatase in the brain [244, 245]. One of the main hallmarks of AD is tau hyperphosphorylation [244, 246], which has 3–4-fold higher levels of tau phosphorylation compared to control brains [247]. Consistent with this notion, the reduced activity and expression of ABαC subunit of PP2A, the major tau phosphatase [248,249,250], which consists of a scaffolding A subunit, a regulatory B subunit, and a catalytic C subunit [251], were observed in AD brains, but not in non-AD dementias [252, 253]. In contrast, enhanced activation of glycogen synthase kinase 3 beta (GSK3β), a major tau kinase [248, 250, 254], was found to increase tau phosphorylation [250] (Fig. 3). Decreased activity and expression of PP2A-C subunit in AD were not only reported to be involved in tau hyperphosphorylation, but also suggested to be responsible for the activation of c-jun N-terminal kinase (JNK), which could lead to Aβ overproduction [255, 256]. Accordingly, two endogenous inhibitors of PP2A, I1(PP2A) and I2(PP2A), were upregulated in the neocortex by in situ hybridization in AD brains, and were suggested to be involved in the hyperphosphorylation of tau in AD [257].
The PP2A-B55α regulatory subunit serves as a major phosphatase for α-synuclein and prevents its accumulation; thereby restricting the key element of PD pathology [258, 259]. In addition, it has been demonstrated that α-synuclein regulates PP2A activity [260,261,262,263], and low activity of PP2A was reported in PD [259, 264] (Fig. 3). Taken together, balanced phosphorylation and dephosphorylation of proteins are critical for physiology, and in particular, reverting PP2A activity ultimately to dephosphorylate tau or α-synuclein could be a promising therapeutic strategy for AD or PD treatment [265, 266].
The shared mechanisms in both cancer and neurodegeneration involve activating kinases and inactivating protein phosphatases. For instance, in brain tumors, PP2A-Aα subunit levels have been found to be reduced in 8 out of 23 glioblastomas, 10 out of 19 oligodendrogliomas, and 7 out of 16 anaplastic oligodendrogliomas [240]. Further, PP2A-Aα subunit mutations were found to contribute to cancer development and tumorigenicity [236]. The most frequent PP2A-Aα mutation, R183W, has been shown to lack the ability to suppress tumor growth, and lead to decreased sensitivity of tumors towards MEK inhibitors [267]. Alongside, PPP2R2C, which encodes a gamma isoform of the subunit B55 subfamily, was also reported to be downregulated in various glioma cell lines and glioma patients [268]. Overexpression of PPP2R2C suppressed cancer cell proliferation by inhibiting the activity of S6K in the mTOR pathway, and further promoting the binding of PP2A-C with S6K, indicating PP2A as a potent tumor suppressor in human brain cancer [268]. Further investigation on the detailed mechanisms underlying PP2A downregulation in gliomas, which leads to neurodegeneration would provide better information for PP2A-based drug development for cancer and neurodegenerative diseases.
Brain tumors and neurodegeneration: Intercellular communications between cancer and neuronal cells
As discussed in the previous section, molecules such as p53, cyclins, Pin1 and PP2A play important roles in the deregulation of homeostatic pathways in respective cell types, leading to cancers and neurodegeneration. Besides the intracellular actions of such molecules, recent studies have shed light on another viewpoint that cancer and neurodegeneration can affect each other by the communication between tumor cells and neuronal cells in the brain.
Several studies have demonstrated that malignant primary brain tumor glioma cells secrete excessive glutamate via the cystine/glutamate antiporter xCT (SLC7A11) [269], which generates a toxic microenvironment for the neurons lying in the vicinity of the glioma, thereby inducing excitotoxicity, neuronal cell death, and neurodegeneration [270,271,272,273] (Fig. 5). In addition, gliomas implanted into the striata of adult rats have shown high glutamate release, rapid growth of the glioma, and neuronal degeneration in the vicinity of the tumor [272]. This effect was found to be reduced by blocking the glutamate receptor, N-methyl-d-aspartate (NMDA) receptor, with its antagonist MK801 or memantine [272], indicating that glutamate-releasing glioma cells mediate neurodegeneration by generating excessive glutamate excitotoxicity in the vicinity of the glioma. With this knowledge, potential therapeutic effects of antagonizing tumor-secreted glutamate or its receptors can be considered. These findings are further supportive of a positive correlation between brain tumors and AD [274, 275].
Reciprocal regulation between brain cancer and neuronal cells in the brain. Excessive glutamate (Glu) secreted from glioma cells leads to neurodegeneration as well as glioma progression. NMDA receptors (NMDARs) on the metastasized breast cancer cells receive glutamate, and promote breast-to-brain cancer metastasis. Neuroligin-3 (NLGN-3) is cleaved by ADAM10 and released from a postsynaptic neuron. This secreted soluble neuroligin-3 (sNLGN-3) acts as a mitogen for glioma cells, thereby fostering glioma progression through a FAK and PI3K–mTOR signaling pathway. Transforming growth factor β (TGF-β)1-induced anti-apoptotic factor (TIAF1) aggregates, which can form a peritumor capsule, cause neurotoxicity while suppressing tumor progression through the interaction with Smad4, WW domain-containing oxidoreductase (WWOX) and p53. Neurotoxicity can also be caused by the formation of TIAF1-Aβ complex. ADAM10 a disintegrin and metalloproteinase 10, AMPAR AMPA receptor, APP amyloid precursor protein, Cys cystine, FAK focal adhesion kinase, mTOR mammalian target of rapamycin, NRXN neurexin, Smad4 mothers against decapentaplegic homolog 4
Besides the cancer cell-driven regulation of neurodegeneration, studies suggesting the neuronal regulation of brain cancer have also been demonstrated [276,277,278,279]. A recent study reported that presynapse-releasing glutamate is implicated in invasive tumor growth in breast cancer metastasized to the brain [279]. Breast cancer metastasized to the brain express NMDA receptors that may be activated by glutamate released from the presynapse, and aid in promoting the breast-to-brain cancer metastasis [279]. The paracrine action of glutamate is achieved by forming pseudo-tripartite synapses, composed of a cancer cell, a presynapse, and a postsynapse [279] (Fig. 5). In a patient-derived pediatric glioblastoma xenograft model, optogenetically induced neuronal activity promoted the proliferation and growth of the glioma in vivo [276, 278]. This glioma growth was found to be mediated by activity-dependent cleavage and secretion of the synaptic adhesion molecule neuroligin-3 from a postsynaptic neuron or oligodendrocyte precursor cell. This cleavage is carried out by a disintegrin and metalloproteinase, ADAM10 [278]. The secreted soluble neuroligin-3 can act as a mitogen for the glioma, inducing focal adhesion kinase (FAK), phosphoinositide 3-kinase (PI3K)-mTOR pathway, expression of neuroligin-3 and other synapse genes, leading to the proliferation of the glioma cells [276,277,278] (Fig. 5). Interestingly, patient-derived glioma xenografted into the CA1 region of hippocampal circuit was found to exhibit AMPA receptor-mediated excitatory postsynaptic currents (EPSCs) on the glioma, and form structural synapses with neurons [277]. The glioma progression was promoted through the integration of electrical and synaptic features of the glioma into neural circuits in the brain [277].
The microenvironment of brain cancer is also governed by transforming growth factor β (TGF-β)1-induced anti-apoptotic factor (TIAF1), found to be aggregated at the interface between metastatic cancer cells, such as metastatic small-cell lung cancer cells and metastatic lung adenocarcinoma, forming a protective peritumor capsule, that can be toxic to neurons [280] (Fig. 5). TIAF1 aggregates have been found in the hippocampi of both non-demented humans and AD patients, along with Aβ and tumor suppressors, such as Smad4 and WW domain-containing oxidoreductases (WWOX or WOX1) [280, 281]. TIAF1 aggregation suppresses anchorage-independent growth, metastasis, and tumor progression, while inducing apoptosis and cell death, which may lead to neurodegeneration [280, 281]. Consistently, a TIAF1/WWOX/p53 triad was found to suppress cancer progression [280, 282], but caused brain protein aggregation in the brain due to functional antagonism of p53 to WWOX-mediated cancer suppression, which lead to neurodegeneration [282]. Unlike TIAF aggregates, zinc finger-like protein that regulates apoptosis (Zfra) and bind tau and Aβ in the AD hippocampus, was reported to suppress melanoma-mediated neurodegeneration in the hippocampus and cortex [283]. More detailed investigation on the intracellular, extracellular or intercellular mechanisms of where and how TIAF1 and Zfra exert their actions in establishing communication between brain cancer cells and neuronal cells would be interesting.
Discussion and perspectives
Many epidemiological studies have demonstrated an inverse correlation between the two age-related diseases, cancer and neurodegenerative diseases [22,23,24], and this intriguing correlation was restricted to certain types of cancers and neurodegenerative diseases. Indeed, in the case of schizophrenia, varying degrees of risk for different types of cancers have been reported [284, 285]. For instance, patients with schizophrenia have shown an increased, marginal, and decreased risk in colon, breast, and respiratory cancer, respectively [285]. Many studies, as described in this review, have revealed the shared roles of overlapping molecules involved in both cancers and neurodegenerative diseases. However, the underlying mechanisms for the two are very distinct, wherein cancers escape cell death while neurodegeneration occurs towards cell death (Fig. 1). Therefore, it would be conceivable that individuals afflicted with a neurodegenerative disease may have a reduced chance of developing certain types of cancers and vice versa. Given the molecular overlap of both diseases, studies in the fields of cancer and neurodegeneration would provide mutual benefits for each other. Because both diseases are closely associated with genetic mutations, it would be valuable to investigate the correlations of the genetic mutations which are found in one disease and also affect the other disease. For this, large amounts of intensive epidemiological studies investigating the incidence of one disease in the population that is affected by the other disease would need to be performed.
Besides clinical and epidemiological studies, which indicate an inverse association between cancer and neurodegenerative diseases [14, 18, 24, 31, 286, 287], studies on shared molecular mechanisms between cancer and neurodegenerative diseases are increasing [16, 288]. Further in-depth investigations into the cellular and molecular mechanisms related to distinct or shared features targeting the molecular crosstalk between cancer and neurodegeneration will assist in the development of additional biomarkers and new therapeutics. Because of the inverse associations between the two diseases with shared molecules in their pathological processes, the therapeutic development in cancer research may lead to the identification of prognostic markers even for both cancer and neurodegeneration, which could potentially result in improved treatments for both disorders. Indeed, over the last decade, drug repositioning from anticancer agents to medicine for neurodegenerative diseases or in the opposite direction has been applied to develop novel therapeutics to overcome these two aging-related diseases with success or failure [289, 290].
Cyclin D and cyclin E are upregulated in both cancer and neurodegenerative diseases, while PP2A is downregulated in both diseases. In addition, cyclin F is downregulated in cancer, and functionally mutated cyclin F is found in neurodegenerative diseases. p53 is downregulated in cancer but upregulated in neurodegenerative diseases, while Pin1 is upregulated in cancer and PD but downregulated in AD (Fig. 2). Overall, it seems that such overlapping molecules between cancer and neurodegenerative diseases may play important roles in pathophysiology and physiological functions differentially in various contexts, depending on the stage or severity of disease and molecular characteristics.
Several studies have demonstrated that inhibition of Pin1 effectively suppresses the growth of various cancer cells [291, 292], and is considered as a promising target for cancer treatment. Indeed, many small molecule inhibitors targeting Pin1 have been developed [215, 293, 294], that exhibit anticancer activities [225, 295]. All-trans retinoic acid (ATRA), a target drug used for acute promyelocytic leukemia (APL), binds to the substrate binding site of Pin1 and thus inhibits Pin1 activity in breast cancer [225]. Juglone, a compound produced by walnut trees, covalently modifies the catalytic core of Pin1 [215, 293], and inhibits multiple cancer cells [291, 296]. API-1, a small molecule targeting the PPIase domain of Pin1, suppresses the proliferation and migration of hepatocellular carcinoma cells [292]. KPT-6566 covalently binds to the PPIase catalytic core of Pin1, and selectively inhibits and degrades Pin1 [213]. Such Pin1 inhibitors that reduce Pin activity would not be directly applicable to treat AD because Pin1 deficiency contributes to AD, and Pin1 expression is inversely correlated with tauopathy and AD [230]. However, a compensatory activation or upregulation of Pin1 has been found in mild cognitive impairment, critically indicating that Pin1-based therapeutics needs to be considered depending on the course of AD [297].
In addition, intracellular organelles that regulate the balance between cell survival and death, are also governed by Pin1 in cancer and apoptotic neurons [298,299,300]. Activated p53, under genotoxic stress, regulates apoptosis-related Bax and Puma expression [301]. Pin1 binds to the activated p53 in the cytoplasm, which promotes the translocation of Pin1 to the mitochondrial membrane, where Pin1 binds to the Bcl-2 homology 3 (BH3)-only protein, Bcl-2-interacting mediator of cell death (Bim)-extralong (BimEL), and mediates neural-specific mitochondrial pro-apoptotic activity [299, 300]. As discussed, Pin1 can be either pro- or anti-apoptotic depending on the cellular context, and therefore, the role of Pin1 in mitochondria-driven apoptosis could provide a direct mechanical link between cancer and neurodegeneration. Therefore, future research in the field should prioritize the investigation of the sophisticated cellular and molecular mechanistic details between cancer and neurodegenerative diseases. Such work will provide a detailed checklist for the development and repositioning of therapeutics, and by unraveling the inverse association between cancer and neurodegenerative diseases, ultimately contribute to personalized medicine and treatment.
Abbreviations
- Aβ:
-
Amyloid-β
- AD:
-
Alzheimer’s disease
- ALS:
-
Amyotrophic lateral sclerosis
- AMPA:
-
α-Amino-3-hydroxy-5-methyl-4-isoxazolepropionate
- APC/C:
-
Anaphase promoting complex/cyclosome
- APP:
-
Amyloid precursor protein
- ATRA:
-
All-trans retinoic acid
- APL:
-
Acute promyelocytic leukemia
- Bax:
-
Bcl-2 associated X
- BH3:
-
Bcl-2 homology 3
- Bim:
-
Bcl-2-interacting mediator of cell death
- BimEL:
-
Bim-extra long
- CDK:
-
Cyclin-dependent kinase
- Chd1:
-
Cdc20 homologue 1
- CP110:
-
Centrosomal protein of 110 kDa
- E2F:
-
E2 factor
- EPSCs:
-
Excitatory postsynaptic currents
- ER:
-
Endoplasmic reticulum
- ERK:
-
Extracellular signal-regulated kinase
- FAK:
-
Focal adhesion kinase
- FBXO1:
-
F-box only 1
- FOXM1:
-
Forkhead box protein M1
- FTD:
-
Frontotemporal dementia
- GR:
-
Glutathione reductase
- GSK3β:
-
Glycogen synthase kinase 3 beta
- HD:
-
Huntington’s disease
- HO-1:
-
Heme oxygenase-1
- IDH1:
-
Isocitrate dehydrogenase 1
- JNK:
-
C-jun N-terminal kinase
- LMW:
-
Low-molecular-weight
- LRRK2:
-
Leucine rich repeat kinase 2
- MAPT:
-
Microtubule-associated protein tau
- miRNA:
-
MicroRNA
- MPP + :
-
1-Methyl-4-phenylpyridinium
- MPTP:
-
1-Methyl-4-phenyl-1,2,3,6-tetrahydropyridine
- mTOR:
-
Mammalian target of rapamycin
- NRF2:
-
Nuclear factor erythroid 2 (NF-E2)-related factor 2
- NIMA:
-
Never in mitosis A
- NMDA:
-
N-methyl-d-aspartate
- NQO1:
-
Quinone oxidoreductase 1
- PARK6:
-
Parkinsonism associated deglycase 6
- PD:
-
Parkinson’s disease
- Pin1:
-
Peptidyl-prolyl cis–trans isomerase NIMA-interacting 1
- PI3K:
-
Phosphoinositide 3-kinase
- PINK1:
-
PTEN induced kinase 1
- PM:
-
Plasma membrane
- PPIase:
-
Peptidyl-prolyl cis–trans isomerase
- PP2A:
-
Protein phosphatase 2A
- PRDX3:
-
Peroxiredoxin 3
- PSEN1/2:
-
Presenilin 1/2
- PTEN:
-
Phosphatase and tensin homolog
- ROS:
-
Reactive oxygen species
- SCF:
-
Skp1–Cul1–F-box
- SLBP:
-
Stem-loop binding protein
- Smad4:
-
Mothers against decapentaplegic homolog 4
- SOD:
-
Superoxide dismutase
- TAR:
-
Transactive response
- TDP-43:
-
TAR DNA-binding protein 43
- TGF-β:
-
Transforming growth factor β
- TIAF1:
-
TGF-β1-induced anti-apoptotic factor
- TREM2:
-
Triggering receptor expressed on myeloid cells 2
- USP27:
-
Ubiquitin-specific peptidase 27
- WWOX or WOX1:
-
WW domain-containing oxidoreductase
- Zfra:
-
Zinc finger-like protein that regulates apoptosis
References
Ambrose M, Gatti RA (2013) Pathogenesis of ataxia-telangiectasia: the next generation of ATM functions. Blood 121(20):4036–4045. https://doi.org/10.1182/blood-2012-09-456897
Hekmatimoghaddam S, Zare-Khormizi MR, Pourrajab F (2017) Underlying mechanisms and chemical/biochemical therapeutic approaches to ameliorate protein misfolding neurodegenerative diseases. BioFactors 43(6):737–759. https://doi.org/10.1002/biof.1264
Lenart P, Novak J, Bienertova-Vasku J (2018) PIWI-piRNA pathway: setting the pace of aging by reducing DNA damage. Mech Ageing Dev 173:29–38. https://doi.org/10.1016/j.mad.2018.03.009
Crusz SM, Balkwill FR (2015) Inflammation and cancer: advances and new agents. Nat Rev Clin Oncol 12(10):584–596. https://doi.org/10.1038/nrclinonc.2015.105
Fisher JC (1958) Multiple-mutation theory of carcinogenesis. Nature 181(4609):651–652. https://doi.org/10.1038/181651b0
Hanahan D, Weinberg RA (2011) Hallmarks of cancer: the next generation. Cell 144(5):646–674. https://doi.org/10.1016/j.cell.2011.02.013
Gao HM, Hong JS (2008) Why neurodegenerative diseases are progressive: uncontrolled inflammation drives disease progression. Trends Immunol 29(8):357–365. https://doi.org/10.1016/j.it.2008.05.002
Avila J (2010) Common mechanisms in neurodegeneration. Nat Med 16(12):1372. https://doi.org/10.1038/nm1210-1372a
Ganguly G, Chakrabarti S, Chatterjee U, Saso L (2017) Proteinopathy, oxidative stress and mitochondrial dysfunction: cross talk in Alzheimer's disease and Parkinson's disease. Drug Des Devel Ther 11:797–810. https://doi.org/10.2147/DDDT.S130514
Pilato F, Profice P, Ranieri F, Capone F, Di Iorio R, Florio L, Di Lazzaro V (2012) Synaptic plasticity in neurodegenerative diseases evaluated and modulated by in vivo neurophysiological techniques. Mol Neurobiol 46(3):563–571. https://doi.org/10.1007/s12035-012-8302-9
Walker LC, LeVine H 3rd (2012) Corruption and spread of pathogenic proteins in neurodegenerative diseases. J Biol Chem 287(40):33109–33115. https://doi.org/10.1074/jbc.R112.399378
Bajaj A, Driver JA, Schernhammer ES (2010) Parkinson's disease and cancer risk: a systematic review and meta-analysis. Cancer Causes Control 21(5):697–707. https://doi.org/10.1007/s10552-009-9497-6
Driver JA (2014) Inverse association between cancer and neurodegenerative disease: review of the epidemiologic and biological evidence. Biogerontology 15(6):547–557. https://doi.org/10.1007/s10522-014-9523-2
Driver JA, Beiser A, Au R, Kreger BE, Splansky GL, Kurth T, Kiel DP, Lu KP, Seshadri S, Wolf PA (2012) Inverse association between cancer and Alzheimer's disease: results from the Framingham Heart Study. BMJ 344:e1442. https://doi.org/10.1136/bmj.e1442
Ferreira JJ, Neutel D, Mestre T, Coelho M, Rosa MM, Rascol O, Sampaio C (2010) Skin cancer and Parkinson's disease. Mov Disord 25(2):139–148. https://doi.org/10.1002/mds.22855
Ibanez K, Boullosa C, Tabares-Seisdedos R, Baudot A, Valencia A (2014) Molecular evidence for the inverse comorbidity between central nervous system disorders and cancers detected by transcriptomic meta-analyses. PLoS Genet 10(2):e1004173. https://doi.org/10.1371/journal.pgen.1004173
Jespersen CG, Norgaard M, Borre M (2016) Parkinson's disease and risk of prostate cancer: a Danish population-based case-control study, 1995–2010. Cancer Epidemiol 45:157–161. https://doi.org/10.1016/j.canep.2016.11.002
Klus P, Cirillo D, Botta Orfila T, Gaetano Tartaglia G (2015) Neurodegeneration and cancer: where the disorder prevails. Sci Rep 5:15390. https://doi.org/10.1038/srep15390
Lin CY, Lane HY, Chen TT, Wu YH, Wu CY, Wu VY (2013) Inverse association between cancer risks and age in schizophrenic patients: a 12-year nationwide cohort study. Cancer Sci 104(3):383–390. https://doi.org/10.1111/cas.12094
Musicco M, Adorni F, Di Santo S, Prinelli F, Pettenati C, Caltagirone C, Palmer K, Russo A (2013) Inverse occurrence of cancer and Alzheimer disease: a population-based incidence study. Neurology 81(4):322–328. https://doi.org/10.1212/WNL.0b013e31829c5ec1
Roe CM, Behrens MI, Xiong C, Miller JP, Morris JC (2005) Alzheimer disease and cancer. Neurology 64(5):895–898. https://doi.org/10.1212/01.WNL.0000152889.94785.51
Tacik P, Curry S, Fujioka S, Strongosky A, Uitti RJ, van Gerpen JA, Diehl NN, Heckman MG, Wszolek ZK (2016) Cancer in Parkinson's disease. Parkinsonism Relat Disord 31:28–33. https://doi.org/10.1016/j.parkreldis.2016.06.014
Vanacore N, Spila-Alegiani S, Raschetti R, Meco G (1999) Mortality cancer risk in parkinsonian patients: a population-based study. Neurology 52(2):395–398. https://doi.org/10.1212/wnl.52.2.395
White RS, Lipton RB, Hall CB, Steinerman JR (2013) Nonmelanoma skin cancer is associated with reduced Alzheimer disease risk. Neurology 80(21):1966–1972. https://doi.org/10.1212/WNL.0b013e3182941990
Acharya A, Das I, Chandhok D, Saha T (2010) Redox regulation in cancer: a double-edged sword with therapeutic potential. Oxid Med Cell Longev 3(1):23–34. https://doi.org/10.4161/oxim.3.1.10095
Aramillo Irizar P, Schauble S, Esser D, Groth M, Frahm C, Priebe S, Baumgart M, Hartmann N, Marthandan S, Menzel U, Muller J, Schmidt S, Ast V, Caliebe A, Konig R, Krawczak M, Ristow M, Schuster S, Cellerino A, Diekmann S, Englert C, Hemmerich P, Suhnel J, Guthke R, Witte OW, Platzer M, Ruppin E, Kaleta C (2018) Transcriptomic alterations during ageing reflect the shift from cancer to degenerative diseases in the elderly. Nat Commun 9(1):327. https://doi.org/10.1038/s41467-017-02395-2
Li JM, Liu C, Hu X, Cai Y, Ma C, Luo XG, Yan XX (2014) Inverse correlation between Alzheimer's disease and cancer: implication for a strong impact of regenerative propensity on neurodegeneration? BMC Neurol 14:211. https://doi.org/10.1186/s12883-014-0211-2
Liu Z, Zhou T, Ziegler AC, Dimitrion P, Zuo L (2017) Oxidative stress in neurodegenerative diseases: from molecular mechanisms to clinical applications. Oxid Med Cell Longev 2017:2525967. https://doi.org/10.1155/2017/2525967
Madabhushi R, Pan L, Tsai LH (2014) DNA damage and its links to neurodegeneration. Neuron 83(2):266–282. https://doi.org/10.1016/j.neuron.2014.06.034
Sankowski R, Mader S, Valdes-Ferrer SI (2015) Systemic inflammation and the brain: novel roles of genetic, molecular, and environmental cues as drivers of neurodegeneration. Front Cell Neurosci 9:28. https://doi.org/10.3389/fncel.2015.00028
Shafi O (2016) Inverse relationship between Alzheimer's disease and cancer, and other factors contributing to Alzheimer's disease: a systematic review. BMC Neurol 16(1):236. https://doi.org/10.1186/s12883-016-0765-2
Plun-Favreau H, Lewis PA, Hardy J, Martins LM, Wood NW (2010) Cancer and neurodegeneration: between the devil and the deep blue sea. PLoS Genet 6(12):e1001257. https://doi.org/10.1371/journal.pgen.1001257
Beckerman R, Prives C (2010) Transcriptional regulation by p53. Cold Spring Harb Perspect Biol 2(8):a000935. https://doi.org/10.1101/cshperspect.a000935
Funk WD, Pak DT, Karas RH, Wright WE, Shay JW (1992) A transcriptionally active DNA-binding site for human p53 protein complexes. Mol Cell Biol 12(6):2866–2871. https://doi.org/10.1128/mcb.12.6.2866
Lane DP (1992) Cancer. p53, guardian of the genome. Nature 358(6381):15–16. https://doi.org/10.1038/358015a0
Bieging KT, Mello SS, Attardi LD (2014) Unravelling mechanisms of p53-mediated tumour suppression. Nat Rev Cancer 14(5):359–370. https://doi.org/10.1038/nrc3711
Duffy MJ, Synnott NC, Crown J (2017) Mutant p53 as a target for cancer treatment. Eur J Cancer 83:258–265. https://doi.org/10.1016/j.ejca.2017.06.023
Kandoth C, McLellan MD, Vandin F, Ye K, Niu B, Lu C, Xie M, Zhang Q, McMichael JF, Wyczalkowski MA, Leiserson MDM, Miller CA, Welch JS, Walter MJ, Wendl MC, Ley TJ, Wilson RK, Raphael BJ, Ding L (2013) Mutational landscape and significance across 12 major cancer types. Nature 502(7471):333–339. https://doi.org/10.1038/nature12634
Leroy B, Fournier JL, Ishioka C, Monti P, Inga A, Fronza G, Soussi T (2013) The TP53 website: an integrative resource centre for the TP53 mutation database and TP53 mutant analysis. Nucleic Acids Res 41(Database issue):D962–969. https://doi.org/10.1093/nar/gks1033
Ahn JH, Kim TJ, Lee JH, Choi JH (2017) Mutant p53 stimulates cell invasion through an interaction with Rad21 in human ovarian cancer cells. Sci Rep 7(1):9076. https://doi.org/10.1038/s41598-017-08880-4
Dong P, Xu Z, Jia N, Li D, Feng Y (2009) Elevated expression of p53 gain-of-function mutation R175H in endometrial cancer cells can increase the invasive phenotypes by activation of the EGFR/PI3K/AKT pathway. Mol Cancer 8:103. https://doi.org/10.1186/1476-4598-8-103
Kang N, Wang Y, Guo S, Ou Y, Wang G, Chen J, Li D, Zhan Q (2018) Mutant TP53 G245C and R273H promote cellular malignancy in esophageal squamous cell carcinoma. BMC Cell Biol 19(1):16. https://doi.org/10.1186/s12860-018-0167-y
Muller PA, Caswell PT, Doyle B, Iwanicki MP, Tan EH, Karim S, Lukashchuk N, Gillespie DA, Ludwig RL, Gosselin P, Cromer A, Brugge JS, Sansom OJ, Norman JC, Vousden KH (2009) Mutant p53 drives invasion by promoting integrin recycling. Cell 139(7):1327–1341. https://doi.org/10.1016/j.cell.2009.11.026
Muller PA, Trinidad AG, Timpson P, Morton JP, Zanivan S, van den Berghe PV, Nixon C, Karim SA, Caswell PT, Noll JE, Coffill CR, Lane DP, Sansom OJ, Neilsen PM, Norman JC, Vousden KH (2013) Mutant p53 enhances MET trafficking and signalling to drive cell scattering and invasion. Oncogene 32(10):1252–1265. https://doi.org/10.1038/onc.2012.148
Noll JE, Jeffery J, Al-Ejeh F, Kumar R, Khanna KK, Callen DF, Neilsen PM (2012) Mutant p53 drives multinucleation and invasion through a process that is suppressed by ANKRD11. Oncogene 31(23):2836–2848. https://doi.org/10.1038/onc.2011.456
Schofield HK, Zeller J, Espinoza C, Halbrook CJ, Del Vecchio A, Magnuson B, Fabo T, Daylan AEC, Kovalenko I, Lee HJ, Yan W, Feng Y, Karim SA, Kremer DM, Kumar-Sinha C, Lyssiotis CA, Ljungman M, Morton JP, Galban S, Fearon ER, Pasca di Magliano M (2018) Mutant p53R270H drives altered metabolism and increased invasion in pancreatic ductal adenocarcinoma. JCI Insight. https://doi.org/10.1172/jci.insight.97422
Shakya R, Tarulli GA, Sheng L, Lokman NA, Ricciardelli C, Pishas KI, Selinger CI, Kohonen-Corish MRJ, Cooper WA, Turner AG, Neilsen PM, Callen DF (2017) Mutant p53 upregulates alpha-1 antitrypsin expression and promotes invasion in lung cancer. Oncogene 36(31):4469–4480. https://doi.org/10.1038/onc.2017.66
Yoshikawa K, Hamada J, Tada M, Kameyama T, Nakagawa K, Suzuki Y, Ikawa M, Hassan NM, Kitagawa Y, Moriuchi T (2010) Mutant p53 R248Q but not R248W enhances in vitro invasiveness of human lung cancer NCI-H1299 cells. Biomed Res 31(6):401–411
Basu S, Gnanapradeepan K, Barnoud T, Kung CP, Tavecchio M, Scott J, Watters A, Chen Q, Kossenkov AV, Murphy ME (2018) Mutant p53 controls tumor metabolism and metastasis by regulating PGC-1alpha. Genes Dev 32(3–4):230–243. https://doi.org/10.1101/gad.309062.117
Dong P, Karaayvaz M, Jia N, Kaneuchi M, Hamada J, Watari H, Sudo S, Ju J, Sakuragi N (2013) Mutant p53 gain-of-function induces epithelial-mesenchymal transition through modulation of the miR-130b-ZEB1 axis. Oncogene 32(27):3286–3295. https://doi.org/10.1038/onc.2012.334
Wang W, Xiong Y, Ding X, Wang L, Zhao Y, Fei Y, Zhu Y, Shen X, Tan C, Liang Z (2019) Cathepsin L activated by mutant p53 and Egr-1 promotes ionizing radiation-induced EMT in human NSCLC. J Exp Clin Cancer Res 38(1):61. https://doi.org/10.1186/s13046-019-1054-x
Yeudall WA, Vaughan CA, Miyazaki H, Ramamoorthy M, Choi MY, Chapman CG, Wang H, Black E, Bulysheva AA, Deb SP, Windle B, Deb S (2012) Gain-of-function mutant p53 upregulates CXC chemokines and enhances cell migration. Carcinogenesis 33(2):442–451. https://doi.org/10.1093/carcin/bgr270
Dittmer D, Pati S, Zambetti G, Chu S, Teresky AK, Moore M, Finlay C, Levine AJ (1993) Gain of function mutations in p53. Nat Genet 4(1):42–46. https://doi.org/10.1038/ng0593-42
Iwanicki MP, Chen HY, Iavarone C, Zervantonakis IK, Muranen T, Novak M, Ince TA, Drapkin R, Brugge JS (2016) Mutant p53 regulates ovarian cancer transformed phenotypes through autocrine matrix deposition. JCI Insight. https://doi.org/10.1172/jci.insight.86829
Shi XB, Nesslinger NJ, Deitch AD, Gumerlock PH, deVere White RW (2002) Complex functions of mutant p53 alleles from human prostate cancer. Prostate 51(1):59–72
Sun Y, Nakamura K, Wendel E, Colburn N (1993) Progression toward tumor cell phenotype is enhanced by overexpression of a mutant p53 tumor-suppressor gene isolated from nasopharyngeal carcinoma. Proc Natl Acad Sci USA 90(7):2827–2831. https://doi.org/10.1073/pnas.90.7.2827
Xie TX, Zhou G, Zhao M, Sano D, Jasser SA, Brennan RG, Myers JN (2013) Serine substitution of proline at codon 151 of TP53 confers gain of function activity leading to anoikis resistance and tumor progression of head and neck cancer cells. Laryngoscope 123(6):1416–1423. https://doi.org/10.1002/lary.23846
Yeudall WA, Wrighton KH, Deb S (2013) Mutant p53 in cell adhesion and motility. Methods Mol Biol 962:135–146. https://doi.org/10.1007/978-1-62703-236-0_11
Bossi G, Lapi E, Strano S, Rinaldo C, Blandino G, Sacchi A (2006) Mutant p53 gain of function: reduction of tumor malignancy of human cancer cell lines through abrogation of mutant p53 expression. Oncogene 25(2):304–309. https://doi.org/10.1038/sj.onc.1209026
Datta A, Ghatak D, Das S, Banerjee T, Paul A, Butti R, Gorain M, Ghuwalewala S, Roychowdhury A, Alam SK, Das P, Chatterjee R, Dasgupta M, Panda CK, Kundu GC, Roychoudhury S (2017) p53 gain-of-function mutations increase Cdc7-dependent replication initiation. EMBO Rep 18(11):2030–2050. https://doi.org/10.15252/embr.201643347
Gurtner A, Starace G, Norelli G, Piaggio G, Sacchi A, Bossi G (2010) Mutant p53-induced up-regulation of mitogen-activated protein kinase kinase 3 contributes to gain of function. J Biol Chem 285(19):14160–14169. https://doi.org/10.1074/jbc.M109.094813
Scian MJ, Stagliano KE, Deb D, Ellis MA, Carchman EH, Das A, Valerie K, Deb SP, Deb S (2004) Tumor-derived p53 mutants induce oncogenesis by transactivating growth-promoting genes. Oncogene 23(25):4430–4443. https://doi.org/10.1038/sj.onc.1207553
Scian MJ, Stagliano KE, Ellis MA, Hassan S, Bowman M, Miles MF, Deb SP, Deb S (2004) Modulation of gene expression by tumor-derived p53 mutants. Cancer Res 64(20):7447–7454. https://doi.org/10.1158/0008-5472.CAN-04-1568
Singh S, Vaughan CA, Frum RA, Grossman SR, Deb S, Palit Deb S (2017) Mutant p53 establishes targetable tumor dependency by promoting unscheduled replication. J Clin Invest 127(5):1839–1855. https://doi.org/10.1172/JCI87724
Vaughan CA, Singh S, Windle B, Sankala HM, Graves PR, Andrew Yeudall W, Deb SP, Deb S (2012) p53 mutants induce transcription of NF-kappaB2 in H1299 cells through CBP and STAT binding on the NF-kappaB2 promoter and gain of function activity. Arch Biochem Biophys 518(1):79–88. https://doi.org/10.1016/j.abb.2011.12.006
Alexandrova EM, Xu S, Moll UM (2017) Ganetespib synergizes with cyclophosphamide to improve survival of mice with autochthonous tumors in a mutant p53-dependent manner. Cell Death Dis 8(3):e2683. https://doi.org/10.1038/cddis.2017.108
Edwards ZC, Trotter EW, Torres-Ayuso P, Chapman P, Wood HM, Nyswaner K, Brognard J (2017) Survival of head and neck cancer cells relies upon LZK kinase-mediated stabilization of mutant p53. Cancer Res 77(18):4961–4972. https://doi.org/10.1158/0008-5472.CAN-17-0267
Foggetti G, Ottaggio L, Russo D, Mazzitelli C, Monti P, Degan P, Miele M, Fronza G, Menichini P (2019) Autophagy induced by SAHA affects mutant P53 degradation and cancer cell survival. Biosci Rep. https://doi.org/10.1042/BSR20181345
Hui L, Zheng Y, Yan Y, Bargonetti J, Foster DA (2006) Mutant p53 in MDA-MB-231 breast cancer cells is stabilized by elevated phospholipase D activity and contributes to survival signals generated by phospholipase D. Oncogene 25(55):7305–7310. https://doi.org/10.1038/sj.onc.1209735
Kalo E, Kogan-Sakin I, Solomon H, Bar-Nathan E, Shay M, Shetzer Y, Dekel E, Goldfinger N, Buganim Y, Stambolsky P, Goldstein I, Madar S, Rotter V (2012) Mutant p53R273H attenuates the expression of phase 2 detoxifying enzymes and promotes the survival of cells with high levels of reactive oxygen species. J Cell Sci 125(Pt 22):5578–5586. https://doi.org/10.1242/jcs.106815
Kawamata H, Omotehara F, Nakashiro K, Uchida D, Shinagawa Y, Tachibana M, Imai Y, Fujimori T (2007) Oncogenic mutation of the p53 gene derived from head and neck cancer prevents cells from undergoing apoptosis after DNA damage. Int J Oncol 30(5):1089–1097
Li R, Sutphin PD, Schwartz D, Matas D, Almog N, Wolkowicz R, Goldfinger N, Pei H, Prokocimer M, Rotter V (1998) Mutant p53 protein expression interferes with p53-independent apoptotic pathways. Oncogene 16(25):3269–3277. https://doi.org/10.1038/sj.onc.1201867
Lim LY, Vidnovic N, Ellisen LW, Leong CO (2009) Mutant p53 mediates survival of breast cancer cells. Br J Cancer 101(9):1606–1612. https://doi.org/10.1038/sj.bjc.6605335
Zalcenstein A, Weisz L, Stambolsky P, Bar J, Rotter V, Oren M (2006) Repression of the MSP/MST-1 gene contributes to the antiapoptotic gain of function of mutant p53. Oncogene 25(3):359–369. https://doi.org/10.1038/sj.onc.1209061
Zhu H, Mao Q, Lin Y, Yang K, Xie L (2011) RNA interference targeting mutant p53 inhibits growth and induces apoptosis in DU145 human prostate cancer cells. Med Oncol 28(Suppl 1):S381–387. https://doi.org/10.1007/s12032-010-9679-9
Zhu HB, Yang K, Xie YQ, Lin YW, Mao QQ, Xie LP (2013) Silencing of mutant p53 by siRNA induces cell cycle arrest and apoptosis in human bladder cancer cells. World J Surg Oncol 11:22. https://doi.org/10.1186/1477-7819-11-22
Hanel W, Moll UM (2012) Links between mutant p53 and genomic instability. J Cell Biochem 113(2):433–439. https://doi.org/10.1002/jcb.23400
Hermsen R, Toonen P, Kuijk E, Youssef SA, Kuiper R, van Heesch S, de Bruin A, Cuppen E, Simonis M (2015) Lack of major genome instability in tumors of p53 null rats. PLoS ONE 10(3):e0122066. https://doi.org/10.1371/journal.pone.0122066
Mackay HL, Moore D, Hall C, Birkbak NJ, Jamal-Hanjani M, Karim SA, Phatak VM, Pinon L, Morton JP, Swanton C, Le Quesne J, Muller PAJ (2018) Genomic instability in mutant p53 cancer cells upon entotic engulfment. Nat Commun 9(1):3070. https://doi.org/10.1038/s41467-018-05368-1
Murphy KL, Dennis AP, Rosen JM (2000) A gain of function p53 mutant promotes both genomic instability and cell survival in a novel p53-null mammary epithelial cell model. FASEB J 14(14):2291–2302. https://doi.org/10.1096/fj.00-0128com
Samassekou O, Bastien N, Yan J, Mai S, Drouin R (2018) Study of telomere dysfunction in TP53 mutant LoVo cell lines as a model for genomic instability. Methods Mol Biol 1769:209–230. https://doi.org/10.1007/978-1-4939-7780-2_14
Song H, Hollstein M, Xu Y (2007) p53 gain-of-function cancer mutants induce genetic instability by inactivating ATM. Nat Cell Biol 9(5):573–580. https://doi.org/10.1038/ncb1571
Capponcelli S, Pedrini E, Cerone MA, Corti V, Fontanesi S, Alessio M, Bachi A, Soddu S, Ribatti D, Picci P, Helman LJ, Cantelli-Forti G, Sangiorgi L (2005) Evaluation of the molecular mechanisms involved in the gain of function of a Li-Fraumeni TP53 mutation. Hum Mutat 26(2):94–103. https://doi.org/10.1002/humu.20192
Fontemaggi G, Dell'Orso S, Trisciuoglio D, Shay T, Melucci E, Fazi F, Terrenato I, Mottolese M, Muti P, Domany E, Del Bufalo D, Strano S, Blandino G (2009) The execution of the transcriptional axis mutant p53, E2F1 and ID4 promotes tumor neo-angiogenesis. Nat Struct Mol Biol 16(10):1086–1093. https://doi.org/10.1038/nsmb.1669
Fu S, Hou MM, Naing A, Janku F, Hess K, Zinner R, Subbiah V, Hong D, Wheler J, Piha-Paul S, Tsimberidou A, Karp D, Araujo D, Kee B, Hwu P, Wolff R, Kurzrock R, Meric-Bernstam F (2015) Phase I study of pazopanib and vorinostat: a therapeutic approach for inhibiting mutant p53-mediated angiogenesis and facilitating mutant p53 degradation. Ann Oncol 26(5):1012–1018. https://doi.org/10.1093/annonc/mdv066
Eriksson SE, Ceder S, Bykov VJN, Wiman KG (2019) p53 as a hub in cellular redox regulation and therapeutic target in cancer. J Mol Cell Biol 11(4):330–341. https://doi.org/10.1093/jmcb/mjz005
Lisek K, Campaner E, Ciani Y, Walerych D, Del Sal G (2018) Mutant p53 tunes the NRF2-dependent antioxidant response to support survival of cancer cells. Oncotarget 9(29):20508–20523. https://doi.org/10.18632/oncotarget.24974
Tanaka N, Zhao M, Tang L, Patel AA, Xi Q, Van HT, Takahashi H, Osman AA, Zhang J, Wang J, Myers JN, Zhou G (2018) Gain-of-function mutant p53 promotes the oncogenic potential of head and neck squamous cell carcinoma cells by targeting the transcription factors FOXO3a and FOXM1. Oncogene 37(10):1279–1292. https://doi.org/10.1038/s41388-017-0032-z
Park HJ, Carr JR, Wang Z, Nogueira V, Hay N, Tyner AL, Lau LF, Costa RH, Raychaudhuri P (2009) FoxM1, a critical regulator of oxidative stress during oncogenesis. EMBO J 28(19):2908–2918. https://doi.org/10.1038/emboj.2009.239
Blandino G, Di Agostino S (2018) New therapeutic strategies to treat human cancers expressing mutant p53 proteins. J Exp Clin Cancer Res 37(1):30. https://doi.org/10.1186/s13046-018-0705-7
Mantovani F, Collavin L, Del Sal G (2019) Mutant p53 as a guardian of the cancer cell. Cell Death Differ 26(2):199–212. https://doi.org/10.1038/s41418-018-0246-9
Schulz-Heddergott R, Moll UM (2018) Gain-of-function (GOF) mutant p53 as actionable therapeutic target. Cancers (Basel). https://doi.org/10.3390/cancers10060188
de la Monte SM, Sohn YK, Ganju N, Wands JR (1998) P53- and CD95-associated apoptosis in neurodegenerative diseases. Lab Invest 78(4):401–411
Jazvinscak Jembrek M, Slade N, Hof PR, Simic G (2018) The interactions of p53 with tau and Ass as potential therapeutic targets for Alzheimer's disease. Prog Neurobiol 168:104–127. https://doi.org/10.1016/j.pneurobio.2018.05.001
Nakanishi A, Minami A, Kitagishi Y, Ogura Y, Matsuda S (2015) BRCA1 and p53 tumor suppressor molecules in Alzheimer's disease. Int J Mol Sci 16(2):2879–2892. https://doi.org/10.3390/ijms16022879
Cenini G, Sultana R, Memo M, Butterfield DA (2008) Elevated levels of pro-apoptotic p53 and its oxidative modification by the lipid peroxidation product, HNE, in brain from subjects with amnestic mild cognitive impairment and Alzheimer's disease. J Cell Mol Med 12(3):987–994. https://doi.org/10.1111/j.1582-4934.2008.00163.x
Kitamura Y, Shimohama S, Kamoshima W, Matsuoka Y, Nomura Y, Taniguchi T (1997) Changes of p53 in the brains of patients with Alzheimer's disease. Biochem Biophys Res Commun 232(2):418–421. https://doi.org/10.1006/bbrc.1997.6301
Ohyagi Y, Asahara H, Chui DH, Tsuruta Y, Sakae N, Miyoshi K, Yamada T, Kikuchi H, Taniwaki T, Murai H, Ikezoe K, Furuya H, Kawarabayashi T, Shoji M, Checler F, Iwaki T, Makifuchi T, Takeda K, Kira J, Tabira T (2005) Intracellular Abeta42 activates p53 promoter: a pathway to neurodegeneration in Alzheimer's disease. FASEB J 19(2):255–257. https://doi.org/10.1096/fj.04-2637fje
Turnquist C, Horikawa I, Foran E, Major EO, Vojtesek B, Lane DP, Lu X, Harris BT, Harris CC (2016) p53 isoforms regulate astrocyte-mediated neuroprotection and neurodegeneration. Cell Death Differ 23(9):1515–1528. https://doi.org/10.1038/cdd.2016.37
Anderson AJ, Stoltzner S, Lai F, Su J, Nixon RA (2000) Morphological and biochemical assessment of DNA damage and apoptosis in Down syndrome and Alzheimer disease, and effect of postmortem tissue archival on TUNEL. Neurobiol Aging 21(4):511–524
Sajan FD, Martiniuk F, Marcus DL, Frey WH 2nd, Hite R, Bordayo EZ, Freedman ML (2007) Apoptotic gene expression in Alzheimer's disease hippocampal tissue. Am J Alzheimers Dis Other Demen 22(4):319–328. https://doi.org/10.1177/1533317507302447
Su JH, Satou T, Anderson AJ, Cotman CW (1996) Up-regulation of Bcl-2 is associated with neuronal DNA damage in Alzheimer's disease. NeuroReport 7(2):437–440
Holcomb L, Gordon MN, McGowan E, Yu X, Benkovic S, Jantzen P, Wright K, Saad I, Mueller R, Morgan D, Sanders S, Zehr C, O'Campo K, Hardy J, Prada CM, Eckman C, Younkin S, Hsiao K, Duff K (1998) Accelerated Alzheimer-type phenotype in transgenic mice carrying both mutant amyloid precursor protein and presenilin 1 transgenes. Nat Med 4(1):97–100
Dorszewska J, Oczkowska A, Suwalska M, Rozycka A, Florczak-Wyspianska J, Dezor M, Lianeri M, Jagodzinski PP, Kowalczyk MJ, Prendecki M, Kozubski W (2014) Mutations in the exon 7 of Trp53 gene and the level of p53 protein in double transgenic mouse model of Alzheimer's disease. Folia Neuropathol 52(1):30–40
Zajkowicz A, Gdowicz-Klosok A, Krzesniak M, Janus P, Lasut B, Rusin M (2018) The Alzheimer's disease-associated TREM2 gene is regulated by p53 tumor suppressor protein. Neurosci Lett 681:62–67. https://doi.org/10.1016/j.neulet.2018.05.037
Mogi M, Kondo T, Mizuno Y, Nagatsu T (2007) p53 protein, interferon-gamma, and NF-kappaB levels are elevated in the parkinsonian brain. Neurosci Lett 414(1):94–97. https://doi.org/10.1016/j.neulet.2006.12.003
Burguillos MA, Deierborg T, Kavanagh E, Persson A, Hajji N, Garcia-Quintanilla A, Cano J, Brundin P, Englund E, Venero JL, Joseph B (2011) Caspase signalling controls microglia activation and neurotoxicity. Nature 472(7343):319–324. https://doi.org/10.1038/nature09788
Tatton NA (2000) Increased caspase 3 and Bax immunoreactivity accompany nuclear GAPDH translocation and neuronal apoptosis in Parkinson's disease. Exp Neurol 166(1):29–43. https://doi.org/10.1006/exnr.2000.7489
Lanni C, Racchi M, Mazzini G, Ranzenigo A, Polotti R, Sinforiani E, Olivari L, Barcikowska M, Styczynska M, Kuznicki J, Szybinska A, Govoni S, Memo M, Uberti D (2008) Conformationally altered p53: a novel Alzheimer's disease marker? Mol Psychiatry 13(6):641–647. https://doi.org/10.1038/sj.mp.4002060
Lanni C, Uberti D, Racchi M, Govoni S, Memo M (2007) Unfolded p53: a potential biomarker for Alzheimer's disease. J Alzheimers Dis 12(1):93–99
Serrano J, Fernandez AP, Martinez-Murillo R, Martinez A (2010) High sensitivity to carcinogens in the brain of a mouse model of Alzheimer's disease. Oncogene 29(15):2165–2171. https://doi.org/10.1038/onc.2009.503
Buizza L, Prandelli C, Bonini SA, Delbarba A, Cenini G, Lanni C, Buoso E, Racchi M, Govoni S, Memo M, Uberti D (2013) Conformational altered p53 affects neuronal function: relevance for the response to toxic insult and growth-associated protein 43 expression. Cell Death Dis 4:e484. https://doi.org/10.1038/cddis.2013.13
Buizza L, Cenini G, Lanni C, Ferrari-Toninelli G, Prandelli C, Govoni S, Buoso E, Racchi M, Barcikowska M, Styczynska M, Szybinska A, Butterfield DA, Memo M, Uberti D (2012) Conformational altered p53 as an early marker of oxidative stress in Alzheimer's disease. PLoS ONE 7(1):e29789. https://doi.org/10.1371/journal.pone.0029789
Cheignon C, Tomas M, Bonnefont-Rousselot D, Faller P, Hureau C, Collin F (2018) Oxidative stress and the amyloid beta peptide in Alzheimer's disease. Redox Biol 14:450–464. https://doi.org/10.1016/j.redox.2017.10.014
Chinta SJ, Andersen JK (2008) Redox imbalance in Parkinson's disease. Biochim Biophys Acta 1780(11):1362–1367. https://doi.org/10.1016/j.bbagen.2008.02.005
Covarrubias-Pinto A, Moll P, Solis-Maldonado M, Acuna AI, Riveros A, Miro MP, Papic E, Beltran FA, Cepeda C, Concha II, Brauchi S, Castro MA (2015) Beyond the redox imbalance: oxidative stress contributes to an impaired GLUT3 modulation in Huntington's disease. Free Radic Biol Med 89:1085–1096. https://doi.org/10.1016/j.freeradbiomed.2015.09.024
Musgrove RE, Helwig M, Bae EJ, Aboutalebi H, Lee SJ, Ulusoy A, Di Monte DA (2019) Oxidative stress in vagal neurons promotes parkinsonian pathology and intercellular alpha-synuclein transfer. J Clin Invest. https://doi.org/10.1172/JCI127330
Xie H, Hou S, Jiang J, Sekutowicz M, Kelly J, Bacskai BJ (2013) Rapid cell death is preceded by amyloid plaque-mediated oxidative stress. Proc Natl Acad Sci USA 110(19):7904–7909. https://doi.org/10.1073/pnas.1217938110
Zhou D, Zhan C, Zhong Q, Li S (2013) Upregulation of sestrin-2 expression via P53 protects against 1-methyl-4-phenylpyridinium (MPP+) neurotoxicity. J Mol Neurosci 51(3):967–975. https://doi.org/10.1007/s12031-013-0081-x
Gao A, Sun T, Ma G, Cao J, Hu Q, Chen L, Wang Y, Wang Q, Sun J, Wu R, Wu Q, Zhou J, Liu L, Hu J, Dong JT, Zhu Z (2018) LEM4 confers tamoxifen resistance to breast cancer cells by activating cyclin D-CDK4/6-Rb and ERalpha pathway. Nat Commun 9(1):4180. https://doi.org/10.1038/s41467-018-06309-8
Gennaro VJ, Stanek TJ, Peck AR, Sun Y, Wang F, Qie S, Knudsen KE, Rui H, Butt T, Diehl JA, McMahon SB (2018) Control of CCND1 ubiquitylation by the catalytic SAGA subunit USP22 is essential for cell cycle progression through G1 in cancer cells. Proc Natl Acad Sci USA 115(40):E9298–E9307. https://doi.org/10.1073/pnas.1807704115
Landis MW, Pawlyk BS, Li T, Sicinski P, Hinds PW (2006) Cyclin D1-dependent kinase activity in murine development and mammary tumorigenesis. Cancer Cell 9(1):13–22. https://doi.org/10.1016/j.ccr.2005.12.019
Malumbres M, Barbacid M (2009) Cell cycle, CDKs and cancer: a changing paradigm. Nat Rev Cancer 9(3):153–166. https://doi.org/10.1038/nrc2602
Musgrove EA, Caldon CE, Barraclough J, Stone A, Sutherland RL (2011) Cyclin D as a therapeutic target in cancer. Nat Rev Cancer 11(8):558–572. https://doi.org/10.1038/nrc3090
Sittithumcharee G, Suppramote O, Vaeteewoottacharn K, Sirisuksakun C, Jamnongsong S, Laphanuwat P, Suntiparpluacha M, Matha A, Chusorn P, Buraphat P, Kakanaporn C, Charngkaew K, Silsirivanit A, Korphaisarn K, Limsrichamrern S, Tripatara P, Pairojkul C, Wongkham S, Sampattavanich S, Okada S, Jirawatnotai S (2019) Dependency of cholangiocarcinoma on cyclin D-dependent kinase activity. Hepatology. https://doi.org/10.1002/hep.30704
VanArsdale T, Boshoff C, Arndt KT, Abraham RT (2015) Molecular pathways: targeting the cyclin D-CDK4/6 axis for cancer treatment. Clin Cancer Res 21(13):2905–2910. https://doi.org/10.1158/1078-0432.CCR-14-0816
Yu Q, Geng Y, Sicinski P (2001) Specific protection against breast cancers by cyclin D1 ablation. Nature 411(6841):1017–1021. https://doi.org/10.1038/35082500
Ghose A, Shashidhara LS (2011) Cyclin beyond the cell cycle: new partners at the synapse. Dev Cell 21(4):601–602. https://doi.org/10.1016/j.devcel.2011.09.015
Cho E, Kim DH, Hur YN, Whitcomb DJ, Regan P, Hong JH, Kim H, Ho Suh Y, Cho K, Park M (2015) Cyclin Y inhibits plasticity-induced AMPA receptor exocytosis and LTP. Sci Rep 5:12624. https://doi.org/10.1038/srep12624
Joe IS, Kim JH, Kim H, Hong JH, Kim M, Park M (2017) Cyclin Y-mediated transcript profiling reveals several important functional pathways regulated by Cyclin Y in hippocampal neurons. PLoS ONE 12(2):e0172547. https://doi.org/10.1371/journal.pone.0172547
Odajima J, Wills ZP, Ndassa YM, Terunuma M, Kretschmannova K, Deeb TZ, Geng Y, Gawrzak S, Quadros IM, Newman J, Das M, Jecrois ME, Yu Q, Li N, Bienvenu F, Moss SJ, Greenberg ME, Marto JA, Sicinski P (2011) Cyclin E constrains Cdk5 activity to regulate synaptic plasticity and memory formation. Dev Cell 21(4):655–668. https://doi.org/10.1016/j.devcel.2011.08.009
Bonda DJ, Bajic VP, Spremo-Potparevic B, Casadesus G, Zhu X, Smith MA, Lee HG (2010) Review: cell cycle aberrations and neurodegeneration. Neuropathol Appl Neurobiol 36(2):157–163. https://doi.org/10.1111/j.1365-2990.2010.01064.x
Hoglinger GU, Breunig JJ, Depboylu C, Rouaux C, Michel PP, Alvarez-Fischer D, Boutillier AL, Degregori J, Oertel WH, Rakic P, Hirsch EC, Hunot S (2007) The pRb/E2F cell-cycle pathway mediates cell death in Parkinson's disease. Proc Natl Acad Sci U S A 104(9):3585–3590. https://doi.org/10.1073/pnas.0611671104
McShea A, Lee HG, Petersen RB, Casadesus G, Vincent I, Linford NJ, Funk JO, Shapiro RA, Smith MA (2007) Neuronal cell cycle re-entry mediates Alzheimer disease-type changes. Biochim Biophys Acta 1772(4):467–472. https://doi.org/10.1016/j.bbadis.2006.09.010
Rao HV, Thirumangalakudi L, Desmond P, Grammas P (2007) Cyclin D1, cdk4, and Bim are involved in thrombin-induced apoptosis in cultured cortical neurons. J Neurochem 101(2):498–505. https://doi.org/10.1111/j.1471-4159.2006.04389.x
Sherr CJ (1994) G1 phase progression: cycling on cue. Cell 79(4):551–555. https://doi.org/10.1016/0092-8674(94)90540-1
Malumbres M, Barbacid M (2001) To cycle or not to cycle: a critical decision in cancer. Nat Rev Cancer 1(3):222–231. https://doi.org/10.1038/35106065
Laphanuwat P, Likasitwatanakul P, Sittithumcharee G, Thaphaengphan A, Chomanee N, Suppramote O, Ketaroonrut N, Charngkaew K, Lam EW, Okada S, Panich U, Sampattavanich S, Jirawatnotai S (2018) Cyclin D1 depletion interferes with oxidative balance and promotes cancer cell senescence. J Cell Sci. https://doi.org/10.1242/jcs.214726
McShea A, Harris PL, Webster KR, Wahl AF, Smith MA (1997) Abnormal expression of the cell cycle regulators P16 and CDK4 in Alzheimer's disease. Am J Pathol 150(6):1933–1939
Atabay KD, Karabay A (2012) Pin1 inhibition activates cyclin D and produces neurodegenerative pathology. J Neurochem 120(3):430–439. https://doi.org/10.1111/j.1471-4159.2011.07259.x
Ma C, Papermaster D, Cepko CL (1998) A unique pattern of photoreceptor degeneration in cyclin D1 mutant mice. Proc Natl Acad Sci USA 95(17):9938–9943. https://doi.org/10.1073/pnas.95.17.9938
Small DL, Monette R, Comas T, Fournier M, Morley P (1999) Loss of cyclin D1 in necrotic and apoptotic models of cortical neuronal degeneration. Brain Res 842(2):376–383. https://doi.org/10.1016/s0006-8993(99)01852-1
Icreverzi A, de la Cruz AF, Walker DW, Edgar BA (2015) Changes in neuronal CycD/Cdk4 activity affect aging, neurodegeneration, and oxidative stress. Aging Cell 14(5):896–906. https://doi.org/10.1111/acel.12376
Caruso JA, Duong MT, Carey JPW, Hunt KK, Keyomarsi K (2018) Low-molecular-weight cyclin E in human cancer: cellular consequences and opportunities for targeted therapies. Cancer Res 78(19):5481–5491. https://doi.org/10.1158/0008-5472.CAN-18-1235
Ekholm SV, Reed SI (2000) Regulation of G(1) cyclin-dependent kinases in the mammalian cell cycle. Curr Opin Cell Biol 12(6):676–684
Keyomarsi K, Pardee AB (1993) Redundant cyclin overexpression and gene amplification in breast cancer cells. Proc Natl Acad Sci USA 90(3):1112–1116. https://doi.org/10.1073/pnas.90.3.1112
Fukuse T, Hirata T, Naiki H, Hitomi S, Wada H (2000) Prognostic significance of cyclin E overexpression in resected non-small cell lung cancer. Cancer Res 60(2):242–244
Leach FS, Elledge SJ, Sherr CJ, Willson JK, Markowitz S, Kinzler KW, Vogelstein B (1993) Amplification of cyclin genes in colorectal carcinomas. Cancer Res 53(9):1986–1989
Erlanson M, Landberg G (2001) Prognostic implications of p27 and cyclin E protein contents in malignant lymphomas. Leuk Lymphoma 40(5–6):461–470. https://doi.org/10.3109/10428190109097645
Iida H, Towatari M, Tanimoto M, Morishita Y, Kodera Y, Saito H (1997) Overexpression of cyclin E in acute myelogenous leukemia. Blood 90(9):3707–3713
Akama Y, Yasui W, Yokozaki H, Kuniyasu H, Kitahara K, Ishikawa T, Tahara E (1995) Frequent amplification of the cyclin E gene in human gastric carcinomas. Jpn J Cancer Res 86(7):617–621. https://doi.org/10.1111/j.1349-7006.1995.tb02442.x
Molendini L, Benassi MS, Magagnoli G, Merli M, Sollazzo MR, Ragazzini P, Gamberi G, Ferrari C, Balladelli A, Bacchini P, Picci P (1998) Prognostic significance of cyclin expression in human osteosarcoma. Int J Oncol 12(5):1007–1011
Bortner DM, Rosenberg MP (1997) Induction of mammary gland hyperplasia and carcinomas in transgenic mice expressing human cyclin E. Mol Cell Biol 17(1):453–459. https://doi.org/10.1128/mcb.17.1.453
Akli S, Zheng PJ, Multani AS, Wingate HF, Pathak S, Zhang N, Tucker SL, Chang S, Keyomarsi K (2004) Tumor-specific low molecular weight forms of cyclin E induce genomic instability and resistance to p21, p27, and antiestrogens in breast cancer. Cancer Res 64(9):3198–3208
Hinchcliffe EH, Li C, Thompson EA, Maller JL, Sluder G (1999) Requirement of Cdk2-cyclin E activity for repeated centrosome reproduction in Xenopus egg extracts. Science 283(5403):851–854. https://doi.org/10.1126/science.283.5403.851
Sutter T, Dansranjavin T, Lubinski J, Debniak T, Giannakudis J, Hoang-Vu C, Dralle H (2002) Overexpression of cyclin E protein is closely related to the mutator phenotype of colorectal carcinoma. Int J Colorect Dis 17(6):374–380. https://doi.org/10.1007/s00384-002-0390-y
Willmarth NE, Albertson DG, Ethier SP (2004) Chromosomal instability and lack of cyclin E regulation in hCdc4 mutant human breast cancer cells. Breast Cancer Res 6(5):R531–539. https://doi.org/10.1186/bcr900
Casimiro MC, Crosariol M, Loro E, Ertel A, Yu Z, Dampier W, Saria EA, Papanikolaou A, Stanek TJ, Li Z, Wang C, Fortina P, Addya S, Tozeren A, Knudsen ES, Arnold A, Pestell RG (2012) ChIP sequencing of cyclin D1 reveals a transcriptional role in chromosomal instability in mice. J Clin Invest 122(3):833–843. https://doi.org/10.1172/JCI60256
Dong L, Yu L, Bai C, Liu L, Long H, Shi L, Lin Z (2018) USP27-mediated Cyclin E stabilization drives cell cycle progression and hepatocellular tumorigenesis. Oncogene 37(20):2702–2713. https://doi.org/10.1038/s41388-018-0137-z
Bales E, Mills L, Milam N, McGahren-Murray M, Bandyopadhyay D, Chen D, Reed JA, Timchenko N, van den Oord JJ, Bar-Eli M, Keyomarsi K, Medrano EE (2005) The low molecular weight cyclin E isoforms augment angiogenesis and metastasis of human melanoma cells in vivo. Cancer Res 65(3):692–697
Corin I, Di Giacomo MC, Lastella P, Bagnulo R, Guanti G, Simone C (2006) Tumor-specific hyperactive low-molecular-weight cyclin E isoforms detection and characterization in non-metastatic colorectal tumors. Cancer Biol Ther 5(2):198–203. https://doi.org/10.4161/cbt.5.2.2356
Davidson B, Skrede M, Silins I, Shih Ie M, Trope CG, Florenes VA (2007) Low-molecular weight forms of cyclin E differentiate ovarian carcinoma from cells of mesothelial origin and are associated with poor survival in ovarian carcinoma. Cancer 110(6):1264–1271. https://doi.org/10.1002/cncr.22918
Harwell RM, Porter DC, Danes C, Keyomarsi K (2000) Processing of cyclin E differs between normal and tumor breast cells. Cancer Res 60(2):481–489
Milne AN, Carvalho R, Jansen M, Kranenbarg EK, van de Velde CJ, Morsink FM, Musler AR, Weterman MA, Offerhaus GJ (2008) Cyclin E low molecular weight isoforms occur commonly in early-onset gastric cancer and independently predict survival. J Clin Pathol 61(3):311–316. https://doi.org/10.1136/jcp.2006.042648
Porter DC, Zhang N, Danes C, McGahren MJ, Harwell RM, Faruki S, Keyomarsi K (2001) Tumor-specific proteolytic processing of cyclin E generates hyperactive lower-molecular-weight forms. Mol Cell Biol 21(18):6254–6269. https://doi.org/10.1128/mcb.21.18.6254-6269.2001
Wang XD, Rosales JL, Magliocco A, Gnanakumar R, Lee KY (2003) Cyclin E in breast tumors is cleaved into its low molecular weight forms by calpain. Oncogene 22(5):769–774. https://doi.org/10.1038/sj.onc.1206166
Moore JD, Kornbluth S, Hunt T (2002) Identification of the nuclear localization signal in Xenopus cyclin E and analysis of its role in replication and mitosis. Mol Biol Cell 13(12):4388–4400. https://doi.org/10.1091/mbc.e02-07-0449
Delk NA, Hunt KK, Keyomarsi K (2009) Altered subcellular localization of tumor-specific cyclin E isoforms affects cyclin-dependent kinase 2 complex formation and proteasomal regulation. Cancer Res 69(7):2817–2825. https://doi.org/10.1158/0008-5472.CAN-08-4182
Wingate H, Puskas A, Duong M, Bui T, Richardson D, Liu Y, Tucker SL, Van Pelt C, Meijer L, Hunt K, Keyomarsi K (2009) Low molecular weight cyclin E is specific in breast cancer and is associated with mechanisms of tumor progression. Cell Cycle 8(7):1062–1068. https://doi.org/10.4161/cc.8.7.8119
Wingate H, Zhang N, McGarhen MJ, Bedrosian I, Harper JW, Keyomarsi K (2005) The tumor-specific hyperactive forms of cyclin E are resistant to inhibition by p21 and p27. J Biol Chem 280(15):15148–15157. https://doi.org/10.1074/jbc.M409789200
Bagheri-Yarmand R, Nanos-Webb A, Biernacka A, Bui T, Keyomarsi K (2010) Cyclin E deregulation impairs mitotic progression through premature activation of Cdc25C. Cancer Res 70(12):5085–5095. https://doi.org/10.1158/0008-5472.CAN-09-4095
Lucenay KS, Doostan I, Karakas C, Bui T, Ding Z, Mills GB, Hunt KK, Keyomarsi K (2016) Cyclin E associates with the lipogenic enzyme ATP-citrate lyase to enable malignant growth of breast cancer cells. Cancer Res 76(8):2406–2418. https://doi.org/10.1158/0008-5472.CAN-15-1646
Body S, Esteve-Arenys A, Miloudi H, Recasens-Zorzo C, Tchakarska G, Moros A, Bustany S, Vidal-Crespo A, Rodriguez V, Lavigne R, Com E, Casanova I, Mangues R, Weigert O, Sanjuan-Pla A, Menendez P, Marcq B, Picquenot JM, Perez-Galan P, Jardin F, Roue G, Sola B (2017) Cytoplasmic cyclin D1 controls the migration and invasiveness of mantle lymphoma cells. Sci Rep 7(1):13946. https://doi.org/10.1038/s41598-017-14222-1
Fuste NP, Fernandez-Hernandez R, Cemeli T, Mirantes C, Pedraza N, Rafel M, Torres-Rosell J, Colomina N, Ferrezuelo F, Dolcet X, Gari E (2016) Cytoplasmic cyclin D1 regulates cell invasion and metastasis through the phosphorylation of paxillin. Nat Commun 7:11581. https://doi.org/10.1038/ncomms11581
Fuste NP, Ferrezuelo F, Gari E (2016) Cyclin D1 promotes tumor cell invasion and metastasis by cytoplasmic mechanisms. Mol Cell Oncol 3(5):e1203471. https://doi.org/10.1080/23723556.2016.1203471
Meng H, Tian L, Zhou J, Li Z, Jiao X, Li WW, Plomann M, Xu Z, Lisanti MP, Wang C, Pestell RG (2011) PACSIN 2 represses cellular migration through direct association with cyclin D1 but not its alternate splice form cyclin D1b. Cell Cycle 10(1):73–81. https://doi.org/10.4161/cc.10.1.14243
Kitada T, Asakawa S, Hattori N, Matsumine H, Yamamura Y, Minoshima S, Yokochi M, Mizuno Y, Shimizu N (1998) Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism. Nature 392(6676):605–608. https://doi.org/10.1038/33416
Morris LG, Veeriah S, Chan TA (2010) Genetic determinants at the interface of cancer and neurodegenerative disease. Oncogene 29(24):3453–3464. https://doi.org/10.1038/onc.2010.127
Verdaguer E, Garcia-Jorda E, Canudas AM, Dominguez E, Jimenez A, Pubill D, Escubedo E, Pallas JC, Camins A (2002) Kainic acid-induced apoptosis in cerebellar granule neurons: an attempt at cell cycle re-entry. NeuroReport 13(4):413–416. https://doi.org/10.1097/00001756-200203250-00010
Olanow CW, Tatton WG (1999) Etiology and pathogenesis of Parkinson's disease. Annu Rev Neurosci 22:123–144. https://doi.org/10.1146/annurev.neuro.22.1.123
Schulz JB, Matthews RT, Klockgether T, Dichgans J, Beal MF (1997) The role of mitochondrial dysfunction and neuronal nitric oxide in animal models of neurodegenerative diseases. Mol Cell Biochem 174(1–2):193–197
Staropoli JF, McDermott C, Martinat C, Schulman B, Demireva E, Abeliovich A (2003) Parkin is a component of an SCF-like ubiquitin ligase complex and protects postmitotic neurons from kainate excitotoxicity. Neuron 37(5):735–749
Copani A, Condorelli F, Caruso A, Vancheri C, Sala A, Giuffrida Stella AM, Canonico PL, Nicoletti F, Sortino MA (1999) Mitotic signaling by beta-amyloid causes neuronal death. FASEB J 13(15):2225–2234
Khurana V, Lu Y, Steinhilb ML, Oldham S, Shulman JM, Feany MB (2006) TOR-mediated cell-cycle activation causes neurodegeneration in a Drosophila tauopathy model. Curr Biol 16(3):230–241. https://doi.org/10.1016/j.cub.2005.12.042
Lee KH, Lee SJ, Lee HJ, Choi GE, Jung YH, Kim DI, Gabr AA, Ryu JM, Han HJ (2017) Amyloid beta1-42 (Abeta1-42) induces the CDK2-mediated phosphorylation of tau through the activation of the mTORC1 signaling pathway while promoting neuronal cell death. Front Mol Neurosci 10:229. https://doi.org/10.3389/fnmol.2017.00229
Bai C, Richman R, Elledge SJ (1994) Human cyclin F. EMBO J 13(24):6087–6098
Bai C, Sen P, Hofmann K, Ma L, Goebl M, Harper JW, Elledge SJ (1996) SKP1 connects cell cycle regulators to the ubiquitin proteolysis machinery through a novel motif, the F-box. Cell 86(2):263–274. https://doi.org/10.1016/s0092-8674(00)80098-7
Galper J, Rayner SL, Hogan AL, Fifita JA, Lee A, Chung RS, Blair IP, Yang S (2017) Cyclin F: a component of an E3 ubiquitin ligase complex with roles in neurodegeneration and cancer. Int J Biochem Cell Biol 89:216–220. https://doi.org/10.1016/j.biocel.2017.06.011
D'Angiolella V, Donato V, Vijayakumar S, Saraf A, Florens L, Washburn MP, Dynlacht B, Pagano M (2010) SCF(Cyclin F) controls centrosome homeostasis and mitotic fidelity through CP110 degradation. Nature 466(7302):138–142. https://doi.org/10.1038/nature09140
Choudhury R, Bonacci T, Arceci A, Lahiri D, Mills CA, Kernan JL, Branigan TB, DeCaprio JA, Burke DJ, Emanuele MJ (2016) APC/C and SCF(cyclin F) constitute a reciprocal feedback circuit controlling S-phase entry. Cell Rep 16(12):3359–3372. https://doi.org/10.1016/j.celrep.2016.08.058
Dankert JF, Rona G, Clijsters L, Geter P, Skaar JR, Bermudez-Hernandez K, Sassani E, Fenyo D, Ueberheide B, Schneider R, Pagano M (2016) Cyclin F-mediated degradation of SLBP limits H2A.X accumulation and apoptosis upon genotoxic stress in G2. Mol Cell 64(3):507–519. https://doi.org/10.1016/j.molcel.2016.09.010
D'Angiolella V, Donato V, Forrester FM, Jeong YT, Pellacani C, Kudo Y, Saraf A, Florens L, Washburn MP, Pagano M (2012) Cyclin F-mediated degradation of ribonucleotide reductase M2 controls genome integrity and DNA repair. Cell 149(5):1023–1034. https://doi.org/10.1016/j.cell.2012.03.043
Deshmukh RS, Sharma S, Das S (2018) Cyclin F-dependent degradation of RBPJ inhibits IDH1(R132H)-mediated tumorigenesis. Cancer Res 78(22):6386–6398. https://doi.org/10.1158/0008-5472.CAN-18-1772
Fu J, Qiu H, Cai M, Pan Y, Cao Y, Liu L, Yun J, Zhang CZ (2013) Low cyclin F expression in hepatocellular carcinoma associates with poor differentiation and unfavorable prognosis. Cancer Sci 104(4):508–515. https://doi.org/10.1111/cas.12100
Williams KL, Topp S, Yang S, Smith B, Fifita JA, Warraich ST, Zhang KY, Farrawell N, Vance C, Hu X, Chesi A, Leblond CS, Lee A, Rayner SL, Sundaramoorthy V, Dobson-Stone C, Molloy MP, van Blitterswijk M, Dickson DW, Petersen RC, Graff-Radford NR, Boeve BF, Murray ME, Pottier C, Don E, Winnick C, McCann EP, Hogan A, Daoud H, Levert A, Dion PA, Mitsui J, Ishiura H, Takahashi Y, Goto J, Kost J, Gellera C, Gkazi AS, Miller J, Stockton J, Brooks WS, Boundy K, Polak M, Munoz-Blanco JL, Esteban-Perez J, Rabano A, Hardiman O, Morrison KE, Ticozzi N, Silani V, de Belleroche J, Glass JD, Kwok JB, Guillemin GJ, Chung RS, Tsuji S, Brown RH Jr, Garcia-Redondo A, Rademakers R, Landers JE, Gitler AD, Rouleau GA, Cole NJ, Yerbury JJ, Atkin JD, Shaw CE, Nicholson GA, Blair IP (2016) CCNF mutations in amyotrophic lateral sclerosis and frontotemporal dementia. Nat Commun 7:11253. https://doi.org/10.1038/ncomms11253
Kabashi E, Valdmanis PN, Dion P, Spiegelman D, McConkey BJ, Vande Velde C, Bouchard JP, Lacomblez L, Pochigaeva K, Salachas F, Pradat PF, Camu W, Meininger V, Dupre N, Rouleau GA (2008) TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat Genet 40(5):572–574. https://doi.org/10.1038/ng.132
Mackenzie IR, Rademakers R (2008) The role of transactive response DNA-binding protein-43 in amyotrophic lateral sclerosis and frontotemporal dementia. Curr Opin Neurol 21(6):693–700. https://doi.org/10.1097/WCO.0b013e3283168d1d
Rutherford NJ, Zhang YJ, Baker M, Gass JM, Finch NA, Xu YF, Stewart H, Kelley BJ, Kuntz K, Crook RJ, Sreedharan J, Vance C, Sorenson E, Lippa C, Bigio EH, Geschwind DH, Knopman DS, Mitsumoto H, Petersen RC, Cashman NR, Hutton M, Shaw CE, Boylan KB, Boeve B, Graff-Radford NR, Wszolek ZK, Caselli RJ, Dickson DW, Mackenzie IR, Petrucelli L, Rademakers R (2008) Novel mutations in TARDBP (TDP-43) in patients with familial amyotrophic lateral sclerosis. PLoS Genet 4(9):e1000193. https://doi.org/10.1371/journal.pgen.1000193
Lee A, Rayner SL, De Luca A, Gwee SSL, Morsch M, Sundaramoorthy V, Shahheydari H, Ragagnin A, Shi B, Yang S, Williams KL, Don EK, Walker AK, Zhang KY, Yerbury JJ, Cole NJ, Atkin JD, Blair IP, Molloy MP, Chung RS (2017) Casein kinase II phosphorylation of cyclin F at serine 621 regulates the Lys48-ubiquitylation E3 ligase activity of the SCF((cyclin F)) complex. Open Biol. https://doi.org/10.1098/rsob.170058
Nakashima M, Meirmanov S, Naruke Y, Kondo H, Saenko V, Rogounovitch T, Shimizu-Yoshida Y, Takamura N, Namba H, Ito M, Abrosimov A, Lushnikov E, Roumiantsev P, Tsyb A, Yamashita S, Sekine I (2004) Cyclin D1 overexpression in thyroid tumours from a radio-contaminated area and its correlation with Pin1 and aberrant beta-catenin expression. J Pathol 202(4):446–455. https://doi.org/10.1002/path.1534
Wulf GM, Ryo A, Wulf GG, Lee SW, Niu T, Petkova V, Lu KP (2001) Pin1 is overexpressed in breast cancer and cooperates with Ras signaling in increasing the transcriptional activity of c-Jun towards cyclin D1. EMBO J 20(13):3459–3472. https://doi.org/10.1093/emboj/20.13.3459
Ayala G, Wang D, Wulf G, Frolov A, Li R, Sowadski J, Wheeler TM, Lu KP, Bao L (2003) The prolyl isomerase Pin1 is a novel prognostic marker in human prostate cancer. Cancer Res 63(19):6244–6251
Bao L, Kimzey A, Sauter G, Sowadski JM, Lu KP, Wang DG (2004) Prevalent overexpression of prolyl isomerase Pin1 in human cancers. Am J Pathol 164(5):1727–1737. https://doi.org/10.1016/S0002-9440(10)63731-5
Pang R, Yuen J, Yuen MF, Lai CL, Lee TK, Man K, Poon RT, Fan ST, Wong CM, Ng IO, Kwong YL, Tse E (2004) PIN1 overexpression and beta-catenin gene mutations are distinct oncogenic events in human hepatocellular carcinoma. Oncogene 23(23):4182–4186. https://doi.org/10.1038/sj.onc.1207493
Ryo A, Liou YC, Wulf G, Nakamura M, Lee SW, Lu KP (2002) PIN1 is an E2F target gene essential for Neu/Ras-induced transformation of mammary epithelial cells. Mol Cell Biol 22(15):5281–5295. https://doi.org/10.1128/mcb.22.15.5281-5295.2002
Zhang X, Zhang B, Gao J, Wang X, Liu Z (2013) Regulation of the microRNA 200b (miRNA-200b) by transcriptional regulators PEA3 and ELK-1 protein affects expression of Pin1 protein to control anoikis. J Biol Chem 288(45):32742–32752. https://doi.org/10.1074/jbc.M113.478016
Lee KH, Lin FC, Hsu TI, Lin JT, Guo JH, Tsai CH, Lee YC, Lee YC, Chen CL, Hsiao M (1843) Lu PJ (2014) MicroRNA-296-5p (miR-296-5p) functions as a tumor suppressor in prostate cancer by directly targeting Pin1. Biochim Biophys Acta 9:2055–2066. https://doi.org/10.1016/j.bbamcr.2014.06.001
Leong KW, Cheng CW, Wong CM, Ng IO, Kwong YL, Tse E (2017) miR-874-3p is down-regulated in hepatocellular carcinoma and negatively regulates PIN1 expression. Oncotarget 8(7):11343–11355. https://doi.org/10.18632/oncotarget.14526
Yan X, Zhu Z, Xu S, Yang LN, Liao XH, Zheng M, Yang D, Wang J, Chen D, Wang L, Liu X, Liu J, Chen RH, Zhou XZ, Lu KP, Liu H (2017) MicroRNA-140-5p inhibits hepatocellular carcinoma by directly targeting the unique isomerase Pin1 to block multiple cancer-driving pathways. Sci Rep 7:45915. https://doi.org/10.1038/srep45915
Chen F, Feng Z, Zhu J, Liu P, Yang C, Huang R, Deng Z (2018) Emerging roles of circRNA_NEK6 targeting miR-370-3p in the proliferation and invasion of thyroid cancer via Wnt signaling pathway. Cancer Biol Ther 19(12):1139–1152. https://doi.org/10.1080/15384047.2018.1480888
Yeh ES, Lew BO, Means AR (2006) The loss of PIN1 deregulates cyclin E and sensitizes mouse embryo fibroblasts to genomic instability. J Biol Chem 281(1):241–251. https://doi.org/10.1074/jbc.M505770200
Yeh ES, Means AR (2007) PIN1, the cell cycle and cancer. Nat Rev Cancer 7(5):381–388. https://doi.org/10.1038/nrc2107
Campaner E, Rustighi A, Zannini A, Cristiani A, Piazza S, Ciani Y, Kalid O, Golan G, Baloglu E, Shacham S, Valsasina B, Cucchi U, Pippione AC, Lolli ML, Giabbai B, Storici P, Carloni P, Rossetti G, Benvenuti F, Bello E, D'Incalci M, Cappuzzello E, Rosato A, Del Sal G (2017) A covalent PIN1 inhibitor selectively targets cancer cells by a dual mechanism of action. Nat Commun 8:15772. https://doi.org/10.1038/ncomms15772
Guo C, Hou X, Dong L, Dagostino E, Greasley S, Ferre R, Marakovits J, Johnson MC, Matthews D, Mroczkowski B, Parge H, Vanarsdale T, Popoff I, Piraino J, Margosiak S, Thomson J, Los G, Murray BW (2009) Structure-based design of novel human Pin1 inhibitors (I). Bioorg Med Chem Lett 19(19):5613–5616. https://doi.org/10.1016/j.bmcl.2009.08.034
Hennig L, Christner C, Kipping M, Schelbert B, Rucknagel KP, Grabley S, Kullertz G, Fischer G (1998) Selective inactivation of parvulin-like peptidyl-prolyl cis/trans isomerases by juglone. Biochemistry 37(17):5953–5960. https://doi.org/10.1021/bi973162p
Kanaoka R, Kushiyama A, Seno Y, Nakatsu Y, Matsunaga Y, Fukushima T, Tsuchiya Y, Sakoda H, Fujishiro M, Yamamotoya T, Kamata H, Matsubara A, Asano T (2015) Pin1 inhibitor juglone exerts anti-oncogenic effects on LNCaP and DU145 cells despite the patterns of gene regulation by Pin1 differing between these cell lines. PLoS ONE 10(6):e0127467. https://doi.org/10.1371/journal.pone.0127467
Kozono S, Lin YM, Seo HS, Pinch B, Lian X, Qiu C, Herbert MK, Chen CH, Tan L, Gao ZJ, Massefski W, Doctor ZM, Jackson BP, Chen Y, Dhe-Paganon S, Lu KP, Zhou XZ (2018) Arsenic targets Pin1 and cooperates with retinoic acid to inhibit cancer-driving pathways and tumor-initiating cells. Nat Commun 9(1):3069. https://doi.org/10.1038/s41467-018-05402-2
Liao XH, Zhang AL, Zheng M, Li MQ, Chen CP, Xu H, Chu QS, Yang D, Lu W, Tsai TF, Liu H, Zhou XZ, Lu KP (2017) Chemical or genetic Pin1 inhibition exerts potent anticancer activity against hepatocellular carcinoma by blocking multiple cancer-driving pathways. Sci Rep 7:43639. https://doi.org/10.1038/srep43639
Mori T, Hidaka M, Lin YC, Yoshizawa I, Okabe T, Egashira S, Kojima H, Nagano T, Koketsu M, Takamiya M, Uchida T (2011) A dual inhibitor against prolyl isomerase Pin1 and cyclophilin discovered by a novel real-time fluorescence detection method. Biochem Biophys Res Commun 406(3):439–443. https://doi.org/10.1016/j.bbrc.2011.02.066
Russo Spena C, De Stefano L, Palazzolo S, Salis B, Granchi C, Minutolo F, Tuccinardi T, Fratamico R, Crotti S, D'Aronco S, Agostini M, Corona G, Caligiuri I, Canzonieri V, Rizzolio F (2018) Liposomal delivery of a Pin1 inhibitor complexed with cyclodextrins as new therapy for high-grade serous ovarian cancer. J Control Rel 281:1–10. https://doi.org/10.1016/j.jconrel.2018.04.055
Subedi A, Shimizu T, Ryo A, Sanada E, Watanabe N, Osada H (2016) Discovery of novel selenium derivatives as Pin1 inhibitors by high-throughput screening. Biochem Biophys Res Commun 474(3):528–533. https://doi.org/10.1016/j.bbrc.2016.04.124
Tatara Y, Lin YC, Bamba Y, Mori T, Uchida T (2009) Dipentamethylene thiuram monosulfide is a novel inhibitor of Pin1. Biochem Biophys Res Commun 384(3):394–398. https://doi.org/10.1016/j.bbrc.2009.04.144
Uchida T, Takamiya M, Takahashi M, Miyashita H, Ikeda H, Terada T, Matsuo Y, Shirouzu M, Yokoyama S, Fujimori F, Hunter T (2003) Pin1 and Par14 peptidyl prolyl isomerase inhibitors block cell proliferation. Chem Biol 10(1):15–24
Wang J, Liu K, Wang XF, Sun DJ (2017) Juglone reduces growth and migration of U251 glioblastoma cells and disrupts angiogenesis. Oncol Rep 38(4):1959–1966. https://doi.org/10.3892/or.2017.5878
Wei S, Kozono S, Kats L, Nechama M, Li W, Guarnerio J, Luo M, You MH, Yao Y, Kondo A, Hu H, Bozkurt G, Moerke NJ, Cao S, Reschke M, Chen CH, Rego EM, Lo-Coco F, Cantley LC, Lee TH, Wu H, Zhang Y, Pandolfi PP, Zhou XZ, Lu KP (2015) Active Pin1 is a key target of all-trans retinoic acid in acute promyelocytic leukemia and breast cancer. Nat Med 21(5):457–466. https://doi.org/10.1038/nm.3839
Zheng M, Xu H, Liao XH, Chen CP, Zhang AL, Lu W, Wang L, Yang D, Wang J, Liu H, Zhou XZ, Lu KP (2017) Inhibition of the prolyl isomerase Pin1 enhances the ability of sorafenib to induce cell death and inhibit tumor growth in hepatocellular carcinoma. Oncotarget 8(18):29771–29784. https://doi.org/10.18632/oncotarget.15967
Lu PJ, Wulf G, Zhou XZ, Davies P, Lu KP (1999) The prolyl isomerase Pin1 restores the function of Alzheimer-associated phosphorylated tau protein. Nature 399(6738):784–788. https://doi.org/10.1038/21650
Zhou XZ, Kops O, Werner A, Lu PJ, Shen M, Stoller G, Kullertz G, Stark M, Fischer G, Lu KP (2000) Pin1-dependent prolyl isomerization regulates dephosphorylation of Cdc25C and tau proteins. Mol Cell 6(4):873–883
Liou YC, Sun A, Ryo A, Zhou XZ, Yu ZX, Huang HK, Uchida T, Bronson R, Bing G, Li X, Hunter T, Lu KP (2003) Role of the prolyl isomerase Pin1 in protecting against age-dependent neurodegeneration. Nature 424(6948):556–561. https://doi.org/10.1038/nature01832
Pastorino L, Sun A, Lu PJ, Zhou XZ, Balastik M, Finn G, Wulf G, Lim J, Li SH, Li X, Xia W, Nicholson LK, Lu KP (2006) The prolyl isomerase Pin1 regulates amyloid precursor protein processing and amyloid-beta production. Nature 440(7083):528–534. https://doi.org/10.1038/nature04543
Xu L, Ren Z, Chow FE, Tsai R, Liu T, Rizzolio F, Boffo S, Xu Y, Huang S, Lippa CF, Gong Y (2017) Pathological role of peptidyl-prolyl isomerase Pin1 in the disruption of synaptic plasticity in Alzheimer's disease. Neural Plast 2017:3270725. https://doi.org/10.1155/2017/3270725
Ghosh A, Saminathan H, Kanthasamy A, Anantharam V, Jin H, Sondarva G, Harischandra DS, Qian Z, Rana A, Kanthasamy AG (2013) The peptidyl-prolyl isomerase Pin1 up-regulation and proapoptotic function in dopaminergic neurons: relevance to the pathogenesis of Parkinson disease. J Biol Chem 288(30):21955–21971. https://doi.org/10.1074/jbc.M112.444224
Ryo A, Togo T, Nakai T, Hirai A, Nishi M, Yamaguchi A, Suzuki K, Hirayasu Y, Kobayashi H, Perrem K, Liou YC, Aoki I (2006) Prolyl-isomerase Pin1 accumulates in lewy bodies of parkinson disease and facilitates formation of alpha-synuclein inclusions. J Biol Chem 281(7):4117–4125. https://doi.org/10.1074/jbc.M507026200
Engelender S, Kaminsky Z, Guo X, Sharp AH, Amaravi RK, Kleiderlein JJ, Margolis RL, Troncoso JC, Lanahan AA, Worley PF, Dawson VL, Dawson TM, Ross CA (1999) Synphilin-1 associates with alpha-synuclein and promotes the formation of cytosolic inclusions. Nat Genet 22(1):110–114. https://doi.org/10.1038/8820
Cayla X, Ballmer-Hofer K, Merlevede W, Goris J (1993) Phosphatase 2A associated with polyomavirus small-T or middle-T antigen is an okadaic acid-sensitive tyrosyl phosphatase. Eur J Biochem 214(1):281–286. https://doi.org/10.1111/j.1432-1033.1993.tb17922.x
Chen W, Arroyo JD, Timmons JC, Possemato R, Hahn WC (2005) Cancer-associated PP2A Aalpha subunits induce functional haploinsufficiency and tumorigenicity. Cancer Res 65(18):8183–8192. https://doi.org/10.1158/0008-5472.CAN-05-1103
Sablina AA, Hector M, Colpaert N, Hahn WC (2010) Identification of PP2A complexes and pathways involved in cell transformation. Cancer Res 70(24):10474–10484. https://doi.org/10.1158/0008-5472.CAN-10-2855
Westermarck J, Hahn WC (2008) Multiple pathways regulated by the tumor suppressor PP2A in transformation. Trends Mol Med 14(4):152–160. https://doi.org/10.1016/j.molmed.2008.02.001
Wlodarchak N, Xing Y (2016) PP2A as a master regulator of the cell cycle. Crit Rev Biochem Mol Biol 51(3):162–184. https://doi.org/10.3109/10409238.2016.1143913
Colella S, Ohgaki H, Ruediger R, Yang F, Nakamura M, Fujisawa H, Kleihues P, Walter G (2001) Reduced expression of the Aalpha subunit of protein phosphatase 2A in human gliomas in the absence of mutations in the Aalpha and Abeta subunit genes. Int J Cancer 93(6):798–804. https://doi.org/10.1002/ijc.1423
Ruediger R, Ruiz J, Walter G (2011) Human cancer-associated mutations in the Aalpha subunit of protein phosphatase 2A increase lung cancer incidence in Aalpha knock-in and knockout mice. Mol Cell Biol 31(18):3832–3844. https://doi.org/10.1128/MCB.05744-11
Jeong AL, Han S, Lee S, Su Park J, Lu Y, Yu S, Li J, Chun KH, Mills GB, Yang Y (2016) Patient derived mutation W257G of PPP2R1A enhances cancer cell migration through SRC-JNK-c-Jun pathway. Sci Rep 6:27391. https://doi.org/10.1038/srep27391
Cheng Y, Liu W, Kim ST, Sun J, Lu L, Sun J, Zheng SL, Isaacs WB, Xu J (2011) Evaluation of PPP2R2A as a prostate cancer susceptibility gene: a comprehensive germline and somatic study. Cancer Genet 204(7):375–381. https://doi.org/10.1016/j.cancergen.2011.05.002
Javadpour P, Dargahi L, Ahmadiani A, Ghasemi R (2019) To be or not to be: PP2A as a dual player in CNS functions, its role in neurodegeneration, and its interaction with brain insulin signaling. Cell Mol Life Sci 76(12):2277–2297. https://doi.org/10.1007/s00018-019-03063-y
Sontag JM, Sontag E (2014) Protein phosphatase 2A dysfunction in Alzheimer's disease. Front Mol Neurosci 7:16. https://doi.org/10.3389/fnmol.2014.00016
Cho E, Park M (2016) Palmitoylation in Alzheimer's disease and other neurodegenerative diseases. Pharmacol Res 111:133–151. https://doi.org/10.1016/j.phrs.2016.06.008
Kopke E, Tung YC, Shaikh S, Alonso AC, Iqbal K, Grundke-Iqbal I (1993) Microtubule-associated protein tau. Abnormal phosphorylation of a non-paired helical filament pool in Alzheimer disease. J Biol Chem 268(32):24374–24384
Hu W, Wu F, Zhang Y, Gong CX, Iqbal K, Liu F (2017) Expression of tau pathology-related proteins in different brain regions: a molecular basis of tau pathogenesis. Front Aging Neurosci 9:311. https://doi.org/10.3389/fnagi.2017.00311
Martin L, Latypova X, Wilson CM, Magnaudeix A, Perrin ML, Terro F (2013) Tau protein phosphatases in Alzheimer's disease: the leading role of PP2A. Ageing Res Rev 12(1):39–49. https://doi.org/10.1016/j.arr.2012.06.008
Qian W, Shi J, Yin X, Iqbal K, Grundke-Iqbal I, Gong CX, Liu F (2010) PP2A regulates tau phosphorylation directly and also indirectly via activating GSK-3beta. J Alzheimers Dis 19(4):1221–1229. https://doi.org/10.3233/JAD-2010-1317
Sangodkar J, Farrington CC, McClinch K, Galsky MD, Kastrinsky DB, Narla G (2016) All roads lead to PP2A: exploiting the therapeutic potential of this phosphatase. FEBS J 283(6):1004–1024. https://doi.org/10.1111/febs.13573
Gong CX, Singh TJ, Grundke-Iqbal I, Iqbal K (1993) Phosphoprotein phosphatase activities in Alzheimer disease brain. J Neurochem 61(3):921–927. https://doi.org/10.1111/j.1471-4159.1993.tb03603.x
Sontag E, Luangpirom A, Hladik C, Mudrak I, Ogris E, Speciale S, White CL 3rd (2004) Altered expression levels of the protein phosphatase 2A ABalphaC enzyme are associated with Alzheimer disease pathology. J Neuropathol Exp Neurol 63(4):287–301. https://doi.org/10.1093/jnen/63.4.287
Martin L, Latypova X, Wilson CM, Magnaudeix A, Perrin ML, Yardin C, Terro F (2013) Tau protein kinases: involvement in Alzheimer's disease. Ageing Res Rev 12(1):289–309. https://doi.org/10.1016/j.arr.2012.06.003
Colombo A, Bastone A, Ploia C, Sclip A, Salmona M, Forloni G, Borsello T (2009) JNK regulates APP cleavage and degradation in a model of Alzheimer's disease. Neurobiol Dis 33(3):518–525. https://doi.org/10.1016/j.nbd.2008.12.014
Zhang JW, Jing LJ, Jian G, Dong GC (2015) Mechanism of protein phosphatase-2Aregulating phosphorylation of amyloid precursor proteosome and Abeta generation. Bratisl Lek Listy 116(3):184–190
Tanimukai H, Grundke-Iqbal I, Iqbal K (2005) Up-regulation of inhibitors of protein phosphatase-2A in Alzheimer's disease. Am J Pathol 166(6):1761–1771. https://doi.org/10.1016/S0002-9440(10)62486-8
Lee KW, Chen W, Junn E, Im JY, Grosso H, Sonsalla PK, Feng X, Ray N, Fernandez JR, Chao Y, Masliah E, Voronkov M, Braithwaite SP, Stock JB, Mouradian MM (2011) Enhanced phosphatase activity attenuates alpha-synucleinopathy in a mouse model. J Neurosci 31(19):6963–6971. https://doi.org/10.1523/JNEUROSCI.6513-10.2011
Park HJ, Lee KW, Park ES, Oh S, Yan R, Zhang J, Beach TG, Adler CH, Voronkov M, Braithwaite SP, Stock JB, Mouradian MM (2016) Dysregulation of protein phosphatase 2A in parkinson disease and dementia with lewy bodies. Ann Clin Transl Neurol 3(10):769–780. https://doi.org/10.1002/acn3.337
Liu G, Chen M, Mi N, Yang W, Li X, Wang P, Yin N, Li Y, Yue F, Chan P, Yu S (2015) Increased oligomerization and phosphorylation of alpha-synuclein are associated with decreased activity of glucocerebrosidase and protein phosphatase 2A in aging monkey brains. Neurobiol Aging 36(9):2649–2659. https://doi.org/10.1016/j.neurobiolaging.2015.06.004
Lou H, Montoya SE, Alerte TN, Wang J, Wu J, Peng X, Hong CS, Friedrich EE, Mader SA, Pedersen CJ, Marcus BS, McCormack AL, Di Monte DA, Daubner SC, Perez RG (2010) Serine 129 phosphorylation reduces the ability of alpha-synuclein to regulate tyrosine hydroxylase and protein phosphatase 2A in vitro and in vivo. J Biol Chem 285(23):17648–17661. https://doi.org/10.1074/jbc.M110.100867
Peng X, Tehranian R, Dietrich P, Stefanis L, Perez RG (2005) Alpha-synuclein activation of protein phosphatase 2A reduces tyrosine hydroxylase phosphorylation in dopaminergic cells. J Cell Sci 118(Pt 15):3523–3530. https://doi.org/10.1242/jcs.02481
Wu J, Lou H, Alerte TN, Stachowski EK, Chen J, Singleton AB, Hamilton RL, Perez RG (2012) Lewy-like aggregation of alpha-synuclein reduces protein phosphatase 2A activity in vitro and in vivo. Neuroscience 207:288–297. https://doi.org/10.1016/j.neuroscience.2012.01.028
Du TT, Chen YC, Lu YQ, Meng FG, Yang H, Zhang JG (2018) Subthalamic nucleus deep brain stimulation protects neurons by activating autophagy via PP2A inactivation in a rat model of Parkinson's disease. Exp Neurol 306:232–242. https://doi.org/10.1016/j.expneurol.2018.05.017
Schweiger S, Matthes F, Posey K, Kickstein E, Weber S, Hettich MM, Pfurtscheller S, Ehninger D, Schneider R, Krauss S (2017) Resveratrol induces dephosphorylation of Tau by interfering with the MID1-PP2A complex. Sci Rep 7(1):13753. https://doi.org/10.1038/s41598-017-12974-4
Yang CC, Kuai XX, Gao WB, Yu JC, Wang Q, Li L, Zhang L (2016) Morroniside-induced PP2A activation antagonizes tau hyperphosphorylation in a cellular model of neurodegeneration. J Alzheimers Dis 51(1):33–44. https://doi.org/10.3233/JAD-150728
O'Connor CM, Leonard D, Wiredja D, Avelar RA, Wang Z, Schlatzer D, Bryson B, Tokala E, Taylor SE, Upadhyay A, Sangodkar J, Gingras AC, Westermarck J, Xu W, DiFeo A, Brautigan DL, Haider S, Jackson M, Narla G (2019) Inactivation of PP2A by a recurrent mutation drives resistance to MEK inhibitors. Oncogene. https://doi.org/10.1038/s41388-019-1012-2
Fan YL, Chen L, Wang J, Yao Q, Wan JQ (2013) Over expression of PPP2R2C inhibits human glioma cells growth through the suppression of mTOR pathway. FEBS Lett 587(24):3892–3897. https://doi.org/10.1016/j.febslet.2013.09.029
Kim JY, Kanai Y, Chairoungdua A, Cha SH, Matsuo H, Kim DK, Inatomi J, Sawa H, Ida Y, Endou H (2001) Human cystine/glutamate transporter: cDNA cloning and upregulation by oxidative stress in glioma cells. Biochim Biophys Acta 1512(2):335–344. https://doi.org/10.1016/s0005-2736(01)00338-8
Behrens PF, Langemann H, Strohschein R, Draeger J, Hennig J (2000) Extracellular glutamate and other metabolites in and around RG2 rat glioma: an intracerebral microdialysis study. J Neurooncol 47(1):11–22. https://doi.org/10.1023/a:1006426917654
Savaskan NE, Fan Z, Broggini T, Buchfelder M, Eyupoglu IY (2015) Neurodegeneration and the brain tumor microenvironment [corrected]. Curr Neuropharmacol 13(2):258–265. https://doi.org/10.2174/1570159x13666150122224158
Takano T, Lin JH, Arcuino G, Gao Q, Yang J, Nedergaard M (2001) Glutamate release promotes growth of malignant gliomas. Nat Med 7(9):1010–1015. https://doi.org/10.1038/nm0901-1010
Ye ZC, Sontheimer H (1999) Glioma cells release excitotoxic concentrations of glutamate. Cancer Res 59(17):4383–4391
Lehrer S (2010) Glioblastoma and dementia may share a common cause. Med Hypoth 75(1):67–68. https://doi.org/10.1016/j.mehy.2010.01.031
Lehrer S (2018) Glioma and Alzheimer's disease. J Alzheimers Dis Rep 2(1):213–218. https://doi.org/10.3233/ADR-180084
Venkatesh HS, Johung TB, Caretti V, Noll A, Tang Y, Nagaraja S, Gibson EM, Mount CW, Polepalli J, Mitra SS, Woo PJ, Malenka RC, Vogel H, Bredel M, Mallick P, Monje M (2015) Neuronal activity promotes glioma growth through neuroligin-3 secretion. Cell 161(4):803–816. https://doi.org/10.1016/j.cell.2015.04.012
Venkatesh HS, Morishita W, Geraghty AC, Silverbush D, Gillespie SM, Arzt M, Tam LT, Espenel C, Ponnuswami A, Ni L, Woo PJ, Taylor KR, Agarwal A, Regev A, Brang D, Vogel H, Hervey-Jumper S, Bergles DE, Suva ML, Malenka RC, Monje M (2019) Electrical and synaptic integration of glioma into neural circuits. Nature 573(7775):539–545. https://doi.org/10.1038/s41586-019-1563-y
Venkatesh HS, Tam LT, Woo PJ, Lennon J, Nagaraja S, Gillespie SM, Ni J, Duveau DY, Morris PJ, Zhao JJ, Thomas CJ, Monje M (2017) Targeting neuronal activity-regulated neuroligin-3 dependency in high-grade glioma. Nature 549(7673):533–537. https://doi.org/10.1038/nature24014
Zeng Q, Michael IP, Zhang P, Saghafinia S, Knott G, Jiao W, McCabe BD, Galvan JA, Robinson HPC, Zlobec I, Ciriello G, Hanahan D (2019) Synaptic proximity enables NMDAR signalling to promote brain metastasis. Nature 573(7775):526–531. https://doi.org/10.1038/s41586-019-1576-6
Chang JY, Chiang MF, Lin SR, Lee MH, He H, Chou PY, Chen SJ, Chen YA, Yang LY, Lai FJ, Hsieh CC, Hsieh TH, Sheu HM, Sze CI, Chang NS (2012) TIAF1 self-aggregation in peritumor capsule formation, spontaneous activation of SMAD-responsive promoter in p53-deficient environment, and cell death. Cell Death Dis 3:e302. https://doi.org/10.1038/cddis.2012.36
Lee MH, Lin SR, Chang JY, Schultz L, Heath J, Hsu LJ, Kuo YM, Hong Q, Chiang MF, Gong CX, Sze CI, Chang NS (2010) TGF-beta induces TIAF1 self-aggregation via type II receptor-independent signaling that leads to generation of amyloid beta plaques in Alzheimer's disease. Cell Death Dis 1:e110. https://doi.org/10.1038/cddis.2010.83
Chou PY, Lin SR, Lee MH, Schultz L, Sze CI, Chang NS (2019) A p53/TIAF1/WWOX triad exerts cancer suppression but may cause brain protein aggregation due to p53/WWOX functional antagonism. Cell Commun Signal 17(1):76. https://doi.org/10.1186/s12964-019-0382-y
Lee MH, Shih YH, Lin SR, Chang JY, Lin YH, Sze CI, Kuo YM, Chang NS (2017) Zfra restores memory deficits in Alzheimer's disease triple-transgenic mice by blocking aggregation of TRAPPC6ADelta, SH3GLB2, tau, and amyloid beta, and inflammatory NF-kappaB activation. Alzheimers Dement (N Y) 3(2):189–204. https://doi.org/10.1016/j.trci.2017.02.001
Chou FH, Tsai KY, Wu HC, Shen SP (2016) Cancer in patients with schizophrenia: what is the next step? Psychiatry Clin Neurosci 70(11):473–488. https://doi.org/10.1111/pcn.12420
Hippisley-Cox J, Vinogradova Y, Coupland C, Parker C (2007) Risk of malignancy in patients with schizophrenia or bipolar disorder: nested case-control study. Arch Gen Psychiatry 64(12):1368–1376. https://doi.org/10.1001/archpsyc.64.12.1368
Ma LL, Yu JT, Wang HF, Meng XF, Tan CC, Wang C, Tan L (2014) Association between cancer and Alzheimer's disease: systematic review and meta-analysis. J Alzheimers Dis 42(2):565–573. https://doi.org/10.3233/JAD-140168
Realmuto S, Cinturino A, Arnao V, Mazzola MA, Cupidi C, Aridon P, Ragonese P, Savettieri G, D'Amelio M (2012) Tumor diagnosis preceding Alzheimer's disease onset: is there a link between cancer and Alzheimer's disease? J Alzheimers Dis 31(1):177–182. https://doi.org/10.3233/JAD-2012-120184
Holohan KN, Lahiri DK, Schneider BP, Foroud T, Saykin AJ (2012) Functional microRNAs in Alzheimer's disease and cancer: differential regulation of common mechanisms and pathways. Front Genet 3:323. https://doi.org/10.3389/fgene.2012.00323
Monacelli F, Cea M, Borghi R, Odetti P, Nencioni A (2017) Do cancer drugs counteract neurodegeneration? repurposing for Alzheimer's disease. J Alzheimers Dis 55(4):1295–1306. https://doi.org/10.3233/JAD-160840
Pushpakom S, Iorio F, Eyers PA, Escott KJ, Hopper S, Wells A, Doig A, Guilliams T, Latimer J, McNamee C, Norris A, Sanseau P, Cavalla D, Pirmohamed M (2019) Drug repurposing: progress, challenges and recommendations. Nat Rev Drug Discov 18(1):41–58. https://doi.org/10.1038/nrd.2018.168
Chen Y, Wu YR, Yang HY, Li XZ, Jie MM, Hu CJ, Wu YY, Yang SM, Yang YB (2018) Prolyl isomerase Pin1: a promoter of cancer and a target for therapy. Cell Death Dis 9(9):883. https://doi.org/10.1038/s41419-018-0844-y
Pu W, Li J, Zheng Y, Shen X, Fan X, Zhou JK, He J, Deng Y, Liu X, Wang C, Yang S, Chen Q, Liu L, Zhang G, Wei YQ, Peng Y (2018) Targeting Pin1 by inhibitor API-1 regulates microRNA biogenesis and suppresses hepatocellular carcinoma development. Hepatology 68(2):547–560. https://doi.org/10.1002/hep.29819
Marsolier J, Perichon M, DeBarry JD, Villoutreix BO, Chluba J, Lopez T, Garrido C, Zhou XZ, Lu KP, Fritsch L, Ait-Si-Ali S, Mhadhbi M, Medjkane S, Weitzman JB (2015) Theileria parasites secrete a prolyl isomerase to maintain host leukocyte transformation. Nature 520(7547):378–382. https://doi.org/10.1038/nature14044
Zhang Y, Fussel S, Reimer U, Schutkowski M, Fischer G (2002) Substrate-based design of reversible Pin1 inhibitors. Biochemistry 41(39):11868–11877. https://doi.org/10.1021/bi0262395
Xu GG, Etzkorn FA (2009) Pin1 as an anticancer drug target. Drug News Perspect 22(7):399–407. https://doi.org/10.1358/dnp.2009.22.7.1381751
Min SH, Zhou XZ, Lu KP (2016) The role of Pin1 in the development and treatment of cancer. Arch Pharm Res 39(12):1609–1620. https://doi.org/10.1007/s12272-016-0821-x
Wang S, Simon BP, Bennett DA, Schneider JA, Malter JS, Wang DS (2007) The significance of Pin1 in the development of Alzheimer's disease. J Alzheimers Dis 11(1):13–23. https://doi.org/10.3233/jad-2007-11105
Becker EB, Bonni A (2006) Pin1 mediates neural-specific activation of the mitochondrial apoptotic machinery. Neuron 49(5):655–662. https://doi.org/10.1016/j.neuron.2006.01.034
Sorrentino G, Comel A, Mantovani F, Del Sal G (2014) Regulation of mitochondrial apoptosis by Pin1 in cancer and neurodegeneration. Mitochondrion 19(Pt As):88–96. https://doi.org/10.1016/j.mito.2014.08.003
Sorrentino G, Mioni M, Giorgi C, Ruggeri N, Pinton P, Moll U, Mantovani F, Del Sal G (2013) The prolyl-isomerase Pin1 activates the mitochondrial death program of p53. Cell Death Differ 20(2):198–208. https://doi.org/10.1038/cdd.2012.112
Youle RJ, Strasser A (2008) The BCL-2 protein family: opposing activities that mediate cell death. Nat Rev Mol Cell Biol 9(1):47–59. https://doi.org/10.1038/nrm2308
Acknowledgements
The work in the Park laboratory was supported by the Original Technology Research Program for Brain Science of the National Research Foundation of Korea (NRF) funded by the Korean government (MSIT; no. 2018M3C7A1021848).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that this article content has no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Seo, J., Park, M. Molecular crosstalk between cancer and neurodegenerative diseases. Cell. Mol. Life Sci. 77, 2659–2680 (2020). https://doi.org/10.1007/s00018-019-03428-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-019-03428-3