Abstract
Here we present the synthesis, pharmacological activity, and molecular docking of novel non-competitive antagonists of GluK2 receptor. The compounds concerned are derivatives of indole and carbazole and are the second reported series of non-competitive antagonists of the GluK2 receptor (the first one was also published by our group). The activity of the indole derivatives is in the micromolar range, as in the case of the first series of non-competitive GluK2 receptor antagonists. We have found that designed carbazole derivatives are devoid of activity. Active indole derivatives interact with the transduction domain of the GluK2 receptor, i.e., the domain which links the transmembrane region of the receptor with the agonist-binding domain. The binding pocket is situated within one receptor subunit.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The glutamatergic system is an attractive molecular target for pharmacological intervention (Kaczor and Matosiuk, 2010). Ligands acting on ionotropic glutamate receptors (iGluRs: NMDA, AMPA, and kainate receptors) or metabotropic glutamate receptors (mGluRs) are potential drug candidates for the treatment of neurodegenerative diseases (Alzheimer’s disease, Parkinson’s disease, Huntington’s disease), epilepsy, as well as schizophrenia, anxiety, and memory disorders (Kew and Kemp, 2005). Although only a few glutamate receptor ligands have turned out to be clinically useful (firstly, because of the crucial role of the glutamatergic system in many physiological processes, and secondly, due to the unfavorable psychotropic side effects, traditionally linked with high-affinity NMDA receptor antagonists), ligands of kainate receptors seem to be especially promising. Kainate receptors are involved in epileptogenesis and inducing synaptic plasticity, mainly via the mossy fiber long-term potentiation mechanism. Thus, antagonists of kainate receptors are potential anti-seizure and neuroprotective agents. Moreover, non-competitive antagonists of AMPA receptors are well tolerated in preclinical and clinical studies (Szénási et al., 2008), thus it may be expected that this will also be the case for such ligands of kainate receptors.
Research on non-competitive antagonists of kainate receptors is hindered by the fact that only three series of such compounds have been obtained up to now (Kaczor et al., 2012; Valgeirsson et al., 2003, 2004). Recently, we have reported 1,2,3,5-tetrasubstituted indole derivatives which are among the most active non-competitive antagonists of the GluK1 receptor and are the first known such ligands of the GluK2 receptor, Fig. 1 (Kaczor et al., 2012). We have also suggested a binding site for them in the receptor transduction domain (Kaczor et al., 2014) which was enabled by the construction of whole receptor models (Kaczor et al., 2008, 2009, 2014). Here we present further modifications, 2–7, of the lead compound E099-25011, (1-ethyl-5-methoxy-2-(4-methoxyphenyl)-3-methylindole), 1. The lead compound was identified by searching the internal databases of compounds at the Elbion Institute, Radebul, Germany. 1 is an analog of Zindoxifene, an anti-estrogen, tumor-inhibiting compound (Schneider et al., 1991). We have previously optimized compound 1 by changing substituents in positions 1, 2, 3, and 5 of the indole system (Fig. 1) (Kaczor et al., 2012, 2014). Compounds 3 and 5–7 were tested for their affinity to the GluK2 receptor, and compounds 3 and 5 were found to be non-competitive antagonists at this receptor. Furthermore, we show how novel non-competitive antagonists 3 and 5 of the GluK2 receptor interact with the transduction domain of the previously constructed homology model of this receptor (Kaczor et al., 2014).
Non-competitive antagonists of GluK1/GluK2 receptors (Kaczor et al., 2012)
Materials and methods
Chemistry
Reactions were routinely monitored by thin-layer chromatography (TLC) in silica gel (60 F254 Merck plates) and the products were visualized with ultraviolet light at 254 nm. All NMR spectra were acquired on a Bruker AVANCE III 600 MHz spectrometer equipped with a BBO Z-gradient probe. Spectra were recorded at 25 °C using chloroform as a solvent with a non-spinning sample in 5 mm NMR-tubes. High resolution mass spectra (HRMS) were recorded on a Bruker microTOF-Q II and processed using Compass Data Analysis software. The elementary analysis was performed using a Perkin-Elmer analyzer. Melting points were determined with Boetius apparatus and are uncorrected.
5-methoxy-3-methyl-2-(2-thienyl)indole (2)
Colorless crystalline needles (EtOH). This compound was prepared from 0.05 mol of 4-methoxyphenylhydrazine hydrochloride, 0.05 mol of 1-(2-thienyl)propan-1-one (2-propionylthiophene), 100 ml of anhydrous ethanol, and 10 ml of ethanol saturated with HCl, which were mildly boiled in a round-bottomed flask with a reflux condenser for 4 h. The reaction mixture was left overnight. The precipitation obtained was filtered and purified by crystallization from ethanol and repeated washing with n-hexane. Because of the tendency of the products to photooxidation, they had to be kept in the dark in a refrigerator. Yield: 69 %, mp 100–102 °C. 1H NMR (600 MHz, CDCl3) δ = 10.82 (s, 1H, NH), 7.47 (dd, J = 1.2, 5.3 Hz, 1H, H-para thienyl), 7.25 (d, J = 8.8, 1H, H-7), 7.19 (dd, J = 3.6, 5.3 Hz, 1H, H-meta thienyl), 7.11 (dd, J = 1.2, 3.6 Hz, 1H, H-ortho thienyl), 7.04 (d, J = 2.4, 1H, H-4), 6.93 (dd, J = 2.4, 8.8 Hz, 1H, H-6), 3.78 (s, 3H, 5-OMe), 2.29 (s, 3H, 3-Me); 13C NMR (125 MHz, CDCl3) δ = 151.93(C-5), 132.85 (Cipso thienyl), 131.04 (C-7a), 127.33 (C-2), 124.76 (C-ortho thienyl), 124.05 (C-meta thienyl), 122.95 (C-para thienyl), 122.41 (C-3a), 113.91 (C-6), 110.74 (C-3), 110.23 (C-7), 100.68 (C-4), 55.95 (C-5-OMe), 9.65 (C-3-Me); HRMS (EI) m/z: 243.3278 C14H13NOS (calcd 243.3282); Anal. Calcd for C14H13NOS: C, 69.10; H, 5.38; N, 5.76; S, 13.18. Found: C, 69.16; H, 5.42; N, 5.74; S, 13.14.
1-(2-thienyl)propan-1-one (2-propionylothiophene)
0.25 mol (32.53 g) of propionic acid anhydride and 0.2 mol (16.83 g) of thiophene were heated to 60 °C in a three-necked flask, equipped with a mechanic mixer, air condenser, and thermometer. Next, while still mixing, 1.10 g of 85 % orthophosphoric (V) acid was slowly added. Heating was continued for 2.5 h at 125 °C (with the mixture getting darker). After cooling, the mixture was washed with 50 ml of water and 100 ml of 10 % solution of sodium carbonate. The organic layer was dried with anhydrous sodium sulfate and subjected to vacuum distillation. The fraction boiling at 99–103/14 mmHg was collected. bp 88 °C/7 mmHg (Harthough and Kosak, 1947). Yield 12.90 g (46 %).
5-methoxy-1,3-dimethyl-2-(2-thienyl)indole (3)
Colorless crystalline needles (EtOH). This compound was prepared from 0.4 g of sodium hydride (50 % oil suspension) and 10 ml of anhydrous DMF, which were placed in a three-necked round-bottomed flask, equipped with a mechanic mixer and a thermometer. The mixture was cooled to 0 °C, and then a solution of 0.001 mol of 5-methoxy-3-methyl-2-(2-thienyl)indole (2) in 10 ml of anhydrous DMF was added dropwise. The mixture was stirred for 45 min, and a solution of 0.001 mol of methyl sulfate in 5 ml of anhydrous DMF was added. After 20 min, the ice bath was removed and the mixing was continued for 1.5 h at room temperature. Then a few milliliters of water were carefully added to decompose the excess of sodium hydride. The reaction mixture was filtered, the filtrate was cooled, and 20 ml of water was added to it. The precipitation obtained was purified by crystallization from ethanol and repeated washing with n-hexane. Yield 41 %, mp 71–73 °C. 1H NMR (600 MHz, CDCl3) δ = 7.43 (dd, J = 1.2, 5.3 Hz, 1H, H-para thienyl), 7.21 (d, J = 8.8 Hz, 1H, H-7), 7.15 (dd, J = 3.6, 5.3 Hz, 1H, H-meta thienyl), 7.08 (dd, J = 1.2, 3.6 Hz, 1H, H-ortho thienyl), 7.01 (d, J = 2.4 Hz, 1H, H-4), 6.89 (dd, J = 2.4; 8.8 Hz, 1H, H-6), 3.74 (s, 3H, 5-OMe), 2.25 (s, 3H, 3-Me), 1.24 (s, 3H, 1-Me); 13C NMR (125 MHz, CDCl3) δ = 152.09 (C-5), 132.83 (Cipso thienyl), 131.36 (C-7a), 128.71 (C-2), 122.53(C-ortho thienyl), 123.12 (C-meta thienyl), 123.08 (C-para thienyl), 121.69 (C-3a), 113.18 (C-6), 110.77 (C-3), 110.25 (C-7), 100.73 (C-4), 56.03 (C-5-OMe), 15.42 (N1-Me), 9.61 (C-3-Me); HRMS (EI): m/z 257.3552 C15H15NOS (calcd 257.3553); Anal. Calcd for C15H15NOS: C, 70.01; H, 5.87; N, 5.44; S, 12.46. Found: C, 69.95; H, 5.92; N, 5.48; S, 12.41.
1-(1H-Indol-3-yl)-3-phenylprop-2-en-1-one (4)
Derivative 4 was obtained by means of Friedel–Crafts acylation according to (Guchhait et al., 2011) in 7.5 % yield as a yellowish white solid; mp 225-230 °C. Spectral data according to (Guchhait et al., 2011).
3-[1-(4-chlorobenzyl)-1H-indol-5-yl]-1-phenylprop-2-en-1-one (5)
Yellowish solid (EtOH). This compound was prepared as follows: 0.01 mol of derivative 4 and 30 ml of anhydrous DMF were mixed in a round-bottomed flask equipped with a thermometer and a dropping funnel. The reaction mixture was cooled to 0 °C and 0.8 g of sodium hydride was added (50 % oil suspension). After 30 min of mixing, a solution of 0.012 mol of 4-chlorobenzyl chloride in 20 ml of anhydrous DMF was added dropwise. The reaction was continued at room temperature for 3 h. The mixture was filtered and 10–15 ml of water was added to the filtrate. The resulting resin-like substance was removed and the next portion of water (25–30 ml) was added until the solution becomes opaque. The mixture was kept in refrigeration for two hours and the precipitation obtained was filtered and purified by crystallization from ethanol and repeated washing with n-hexane. Yield 72.0 %, mp 235–238 °C, 1H NMR (600 MHz, CDCl3) δ = 8.64–8.54 (m, 1H, H-5), 7.92 (s, 1H, H-2), 7.85 (d, J = 15.5 Hz, 1H, H-4), 7.65 (s, 2H, CH), 7.46–7.26 (m, 9H, Ar, phenyl + benzyl), 7.13–7.09 (m, 2H, H-6, H-7), 5.35 (s, 2H, CH2); 13C NMR (151 MHz, CDCl3) δ = 184.47 (CO), 141.40 (C-2) 140.71 (Cipso phenyl), 137.11 (CHCHCO), 135.34 (C-7a), 134.51 (Cipso benzyl), 134.37 (C-para benzyl), 134.17 (C-ortho phenyl), 129.94 (C-para phenyl), 129.29 (C-meta benzyl), 128.88 (C-meta phenyl), 128.21 (CHCHCO), 127.09 (C-ortho benzyl), 123.97 (C-3a), 123.85 (C-6), 123.24 (C-5), 122.98 (C-4), 118.43 (C-3), 110.13 (C-7), 50.20 (CH2). HRMS (EI): m/z 371.8434 C24H18NOCl (calcd 371.8591); Anal. Calcd for C24H18NOCl C, 77.51; H, 4.87; N, 3.77; Cl, 9.53. Found: C, 77.55; H, 4.88; N, 3.73; S, 9.49.
9H-4-oxo-1,2,3,4-tetrahydrocarbazole (6)
A solution of 0.1 mol of phenylhydrazine in 150 ml of water was added dropwise for 1.5 h to a solution of 1,3-cyclohexadione in 100 ml of water. The orange precipitation of 1,3-cyclohexadione monophenylhydrazone obtained was filtered. Yield 99 %, mp 173.5 °C (Hester, 1969). 100 g of polyphosphoric acid (PPA) was heated to 80 °C and then 0.025 mol of monophenylhydrazone of 1,3-cyclohexadione was added. The temperature slowly increased to 110 °C due to an exothermic reaction. The reactants were mixed for 30 min and then the reaction mixture was poured onto ice. The precipitation obtained was filtered and crystallized from methanol. Derivative 6 was obtained in a 61.6 % yield as a colorless solid, mp 234–235 °C. Spectral data as described by (Rodriguez et al., 1989).
9-(4-chlorobenzyl)-4-oxo-1,2,3,4-tetrahydrocarbazole (7)
Colorless solid (EtOH). This compound was prepared as follows: 25 ml of DMF, 0.1 ml of water, and 0.013 mol of potassium hydroxide were mixed for 5 min. 0.01 mol of 6 was added and mixing was continued for 1 h. Then a solution of 0.0015 mol of 4-chlorobenzyl chloride in 10 ml of DMF was added dropwise and the reaction was continued under stirring for 2 h. The reaction mixture was kept in a refrigerator overnight. 5 ml of water was added and the first portion of precipitation was obtained and filtered. The second portion of precipitation was obtained after adding a further 15 ml of water. The combined precipitation was crystallized from ethanol. Yield 87.7 %, mp 171–173 °C. 1H NMR (500 MHz, CDCl3) 7.58 (d, 1H, J = 7.8, H-5), 7.33 (d, 1H, J = 8.0, H-8), 7.22 (dd, 1H, J = 7.2; 8.0, H-7), 7.20 (d, 2H, J = 8.4, H-meta benzyl), 7.13 (dd, 1H, J = 7.2;7.8, H-6), 6.87 (d, 2H, J = 8.4, H-ortho benzyl), 5.17 (s, 2H, CH2), 2.90 (m, 2H, H-1), 2.59 (m, 2H, H-3), 2.25 (m, 2H, H-2), 13C NMR (100 MHz, CDCl3) 163.32 (CO), 158.25 (C-9a), 148.73 (Cipso benzyl), 139.25 (C-8a), 127.81 (C-para benzyl), 123.97 (C-meta benzyl), 123.85 (C-ortho benzyl), 116.08 (C-4b), 115.64 (C-7), 113.97 (C-6), 112.82 (C5), 111.09 (C-4a), 105.46 (C-8), 20.95 (CH2), 13.51 (C-3), 13.05 (C-1), 12.73 (C-2). HRMS (EI) m/z: 309.7822 C19H16NOCl (calcd 309.7890); Anal. Calcd for C19H16NOCl C, 73.66; H, 5.21; N, 4.52; Cl, 11.44. Found: C, 73.65; H, 5.22; N, 4.53; S, 11.41.
Pharmacology
The pharmacological studies were performed as described previously (Kaczor et al., 2012). HEK293 lines expressing GluK2 kainate receptors, together with aequorin, a bioluminescent Ca2+ reporter protein, were used to determine the effect of the compounds being investigated on GluK2 receptor activity. The influx of Ca2+ ions through open kainate receptor ion channels led to oxidation of coelenterazine, the cofactor of aequorin, which eventually resulted in the emission of photons. After incubation of the cells with coelenterazine, the culture medium was replaced with an assay buffer (Ringer buffer + 100 mM CaCl2). In a luminometer (LumiStar, BMG, Germany), 275 μM of glutamate was applied to the cells and the luminescence signals were recorded before, during, and after glutamate application.
Molecular modeling
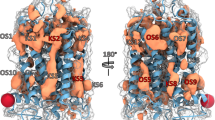
The homology model of the GluK2 receptor was constructed as described previously (Kaczor et al., 2014). The crystal structure of the AMPA GluA2 receptor (PDB ID: 3KG2) (Sobolevsky et al., 2009) was selected as the main template. Additional templates were used for the N-terminal domain (crystal structure of the GluK2/GluK5 NTD tetramer assembly, PDB ID: 3QLV) (Kumar et al., 2011) and the ligand-binding domain (crystal structure of GluK1 ligand-binding domain (S1S2) in complex with an antagonist, PDB ID: 4DLD) (Venskutonytė et al., 2012). Homology modeling was carried out with Modeler v. 9.11 (Eswar et al., 2006). Input conformations of the compounds being investigated were prepared using the LigPrep protocol from the Schrödinger Suite. To sample different protonation states of the ligands in physiological pH, the Epik module was used. The structural and electronic parameters of the ligands were calculated with VegaZZ v.2.4.0.25 (Pedretti et al., 2004), Gausian09 (Frisch et al., 2009), and Discovery Studio 3.1. Molecular docking was performed with Glide from the Schrödinger Suite. Molecular dynamics of ligand-receptor complexes were performed as described previously (Kaczor et al., 2014). Ligand-receptor complexes were inserted into a POPC lipid bilayer and water with a suitable module of Schrödinger suite of programs, and sodium and potassium ions were added to balance the protein charges and then up to a concentration of 0.15 M. The stability of the ligand-receptor complexes was assessed by molecular dynamics simulations with Desmond v. 3.0.3.1 (Bowers et al., 2006) The ligand-receptor complexes in lipid bilayer were minimized and subjected to MD first in the NVT ensemble for 1 ns and then in the NPT ensemble for 20 ns. The following software was also used to visualize the results: Chimera v.1.5.3 (Pettersen et al., 2004), VegaZZ v.2.4.0.25, Yasara Structure v.11.9.18 (Krieger and Vriend, 2002), and PyMol v.0.99 (The PyMOL Molecular Graphics System, Version 0.99, Schrödinger, LLC).
Results and discussion
Chemistry
The synthesis of compounds 2–7 is presented in Fig. 2. Compound 2 was obtained by Fischer indolization reaction. Alkylation of 2 with dimethyl sulfate resulted in compound 3. The lead compound 1 and derivative 2 were previously characterized as anti-estrogens (Masatoshi et al., 1993; von Angerer et al., 1984, 1987, 1990). Compound 3 is a new compound. Compound 4 was obtained in Friedel–Crafts acylation of indole as previously described (Guchhait et al., 2011). Derivative 5 is a new compound and was obtained in alkylation of 4 with 4-chlorobenzyl chloride. Compound 6 was obtained by cyclization of monophenylhydrazone of 1,3-cyclohexadione (obtained from phenylhydrazine and 1,3-cyclohexadione) in PPA and was characterized previously (Rodriguez et al., 1989). Compound 7 is a new compound and was obtained by alkylation of 6 with 4-chlorobenzyl chloride.
Pharmacology
Compounds 3 and 5–7 were tested for their affinity to GluK2 receptors as described previously (Kaczor et al., 2012; 2014). The IC50 values for the compounds being investigated are listed in Table 1. The investigations with the 3H-kainate binding assay showed no inhibition, which makes it possible to conclude that the antagonism for compounds 3 and 5 is of the non-competitive type.
Structural and electronic parameters of novel ligands
In order to address the structure–activity relationship observed, structural and electronic parameters were calculated for compounds 1, 3, 5, 6, and 7. The data are presented in Tables 2 and 3. The data shown in Table 2 show that the lack of activity of compound 6 may be explained by the fact that the molecular volume is too low and the dipole moment too high. The significant difference between the HOMO and LUMO values (Table 3) indicates that the compounds are nucleophilic and may participate as acceptors (through oxygen atoms) in hydrogen bonds with the binding pocket residues; this is in agreement with our earlier studies (Kaczor et al., 2012). Moreover, the novel ligands have more favorable lipophilicity values in comparison to the previous series, with the exception of compound 5 (Kaczor et al., 2012).
Ligand-receptor interactions
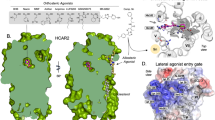
The binding site for non-competitive GluK2 receptor antagonists was identified in the receptor transduction domain, i.e., in the domain which connects the ligand-binding domain and the transmembrane domain (Fig. 3). This assumption was made on the basis of studies by (Balannik et al., 2005) for AMPA receptors as well as on our earlier molecular modeling studies (Kaczor et al., 2009, 2014). The exact binding site was found on the basis of sequence differences between the GluK1 and GluK2 receptors in the transduction domain as reported in our previous studies (Kaczor et al., 2009, 2014). There are no differences in the S1-M1 linker and in the S2-M4 linker. Asp823 and Asn824 in GluK1 correspond to Glu808 and Ser809 in GluK2. The interactions of compounds 3 and 5 with the GluK2 receptor are presented in Fig. 4a, b, c, d, respectively. There are the following residues in the binding pocket: Lys544, Pro545, Asn546, Gly547, Pro667, Asp669, Glu807, Glu808, Lys810, Glu811, and Ala812 which interact with both ligands. Furthermore, in the case of ligand 5, the pocket is extended with the following additional residues: Thr753, Gln754, Ile755, and Gly756. The carbonyl group of ligand 5 forms a hydrogen bond with the side chain of Lys810. The binding pocket is situated within one receptor subunit which is in accordance with our recent studies (Kaczor et al., 2014).
Model of the GluK2 receptor (Kaczor et al., 2014)
Conclusions
In this paper, we have reported the second series of GluK2 receptor non-competitive antagonists. We obtained two indole derivatives with activity in the low micromolar range. Furthermore, we found that the designed carbazole derivatives were not active. The novel non-competitive antagonists interact with the transduction domain of the GluK2 receptor, in the same way as the previously reported series. The binding site is located within one receptor subunit.
References
Balannik V, Menniti FS, Paternain AV, Lerma J, Stern-Bach Y (2005) Molecular mechanism of AMPA receptor noncompetitive antagonism. Neuron 48(2):279–288
Bowers KJ, Chow E, Xu H, Dror RO, Eastwood MP, Gregersen BA, Klepeis JL, Kolossváry I, Moraes MA, Sacerdoti FD, Salmon JK, Shan Y, Shaw DE (2006) Scalable algorithms for molecular dynamics simulations on commodity clusters. Proceedings of the ACM/IEEE conference on supercomputing (SC06), Tampa, Florida, November 11–17
Eswar N, Marti-Renom MA, Webb B, Madhusudhan MS, Eramian D, Shen M, Pieper U, Sali A (2006) Comparative protein structure modeling with MODELLER. Curr Protoc Bioinformatics 15:561–5630
Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery JA Jr, Peralta JE, Ogliaro F, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Staroverov VN, Kobayashi R, Normand J, Raghavachari K, Rendell A, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam NJ, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas Ö, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ (2009) Gaussian 09 Revision D01. Gaussian Inc, Wallingford
Guchhait SK, Kashyap M, Kamble H (2011) ZrCl4-mediated regio- and chemoselective friedel-crafts acylation of indole. J Org Chem 76(11):4753–4758
Harthough HD, Kosak AI (1947) Acylation studies in the thiophene and furan series. IV. Strong inorganic oxyacids as catalysts. J Am Chem Soc 69:3093–3096
Hester JB Jr: (1969) Fr 1 566 173
Kaczor AA, Matosiuk D (2010) Molecular structure of ionotropic glutamate receptors. Curr Med Chem 17(24):2608–2635
Kaczor AA, Kijkowska-Murak UA, Matosiuk D (2008) Theoretical studies on the structure and symmetry of the transmembrane region of glutamatergic GluR5 receptor. J Med Chem 51(13):3765–3776
Kaczor AA, Kijkowska-Murak UA, Kronbach C, Unverferth K, Matosiuk D (2009) Modeling of glutamate GluR6 receptor and its interactions with novel noncompetitive antagonists. J Chem Inf Model 49(4):1094–1104
Kaczor AA, Kronbach C, Unverferth K, Pihlaja K, Wiinämaki K, Sinkkonen J, Kijkowska-Murak U, Wróbel T, Stachal T, Matosiuk D (2012) Novel non-competitive antagonists of kainate gluk1/gluk2 receptors. Lett Drug Design Discov 9:891–898
Kaczor AA, Karczmarzyk Z, Fruziński A, Pihlaja K, Sinkkonen J, Wiinämaki K, Kronbach C, Unverferth K, Poso A, Matosiuk D (2014) Structural studies, homology modeling and molecular docking of novel non-competitive antagonists of GluK1/GluK2 receptors. Bioorg Med Chem 22:787–795
Kew JNC, Kemp JA (2005) Ionotropic and metabotropic glutamate receptor structure and pharmacology. Psychopharmacology 179(1):4–29
Krieger E, Vriend G (2002) Models@Home: distributed computing in bioinformatics using a screensaver based approach. Bioinformatics 18:315–318
Kumar J, Schuck P, Mayer ML (2011) Structure and assembly mechanism for heteromeric kainate receptors. Neuron 71(2):319–331
Masatoshi I, Tadanao S, Jun K, Masako M, Akie T (1993) PCT Int Appl WO 9323374 A1 19931125.
Pedretti A, Villa L, Vistoli G (2004) VEGA - an open platform to develop chemo-bio-informatic applications, using plug-in architecture and script” programming. J Comput Aided Mol Des 18:167–173
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera - a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Rodriguez J-G, Temprano F, Esteban-Calderon C, Martinez-Ripoll M (1989) Synthesis of 4-(N, N- dimethylaminoethyl)-1,2,3,4-tetrahydrocarbazole: molecular structure and reactivity of the 1,2-dihydrocarbazol-4(3H)-one and derivatives. J Chem Soc Perkin Trans 1(11):2117–2122
Schneider MR, Schiller CD, Humm A, von Angerer E (1991) Effect of zindoxifene on experimental prostatic tumours of the rat. J Cancer Res Clin Oncol 117(1):33–36
Sobolevsky AI, Rosconi MP, Gouaux E (2009) X-ray structure, symmetry and mechanism of an AMPA-subtype glutamate receptor. Nature 462(7274):745–756
Szénási G, Vegh M, Szabo G, Kertesz S, Kapus G, Albert M, Greff Z, Ling I, Barkoczy J, Simig G, Spedding M, Harsing LG (2008) 2,3-benzodiazepine-type AMPA receptor antagonists and their neuroprotective effects. Neurochem Int 52:166–183
The PyMOL Molecular Graphics System, Version 0.99, Schrödinger, LLC
Valgeirsson J, Nielsen EØ, Peters D, Varming T, Mathiesen C, Kristensen AS, Madsen U (2003) 2-Arylureidobenzoic acids: selective noncompetitive antagonists for the homomeric kainate receptor subtype GluR5. J Med Chem 46(26):5834–5843
Valgeirsson J, Nielsen EO, Peters D, Mathiesen C, Kristensen AS, Madsen U (2004) Bioisosteric modifications of 2-arylureidobenzoic acids: selective noncompetitive antagonists for the homomeric kainate receptor subtype GluR5. J Med Chem 47(27):6948–6957
Venskutonytė R, Frydenvang K, Valadés EA, Szymańska E, Johansen TN, Kastrup JS, Pickering DS (2012) Structural and pharmacological characterization of phenylalanine-based AMPA receptor antagonists at kainate receptors. ChemMedChem 7(10):1793–1798
Von Angerer E, Strohmeier J (1987) 2-Phenylindoles Effect of N-benzylation on estrogen receptor affinity, estrogenic properties, and mammary tumor inhibiting activity. J Med Chem 30(1):131–136
Von Angerer E, Prekajac J, Strohmeier J (1984) 2-Phenylindoles Relationship between structure, estrogen receptor affinity, and mammary tumor inhibiting activity in the rat. J Med Chem 27(11):1439–1447
Von Angerer E, Knebel N, Kager M, Ganss B (1990) 1-(aminoalkyl)-2-phenylindoles as novel pure estrogen antagonists. J Med Chem 33(9):2635–2640
Acknowledgments
The paper was developed using equipment purchased under the project “The equipment of innovative laboratories doing research on new medicines used in the therapy of civilization and neoplastic diseases” within the Operational Programme Development of Eastern Poland 2007–2013, Priority Axis I Modern Economy, Task I.3 Supporting Innovativeness. The research was partially performed during the postdoctoral fellowship of Agnieszka A. Kaczor at the University of Eastern Finland, Kuopio, Finland as part of a Marie Curie fellowship. The pharmacological investigations presented were funded by European Union EFRE grants and by grants of the Free State of Saxony (Project No. 8093). Computations were performed under a computational grant from the Interdisciplinary Centre for Mathematical and Computational Modelling, Warsaw, Poland, Grant No. G30-18. Calculations with Desmond were carried out using resources of CSC, Finland.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Kaczor, A.A., Wróbel, T., Kronbach, C. et al. Synthesis and molecular docking of novel non-competitive antagonists of GluK2 receptor. Med Chem Res 24, 810–817 (2015). https://doi.org/10.1007/s00044-014-1171-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-014-1171-1