Abstract
Whilst colistin (polymyxin E) represents the last mainstream treatment option for multidrug-resistant Gram-negative pathogens, details of its mechanism of action remain to be fully resolved. In this study, the effects of sub-inhibitory, inhibitory-bactericidal, and supra-bactericidal levels of colistin on the membrane integrity and morphology of Escherichia coli and Pseudomonas aeruginosa were investigated using potassium loss, flow cytometry, and scanning electron microscopy (SEM). Supra-bactericidal colistin concentrations induced just 4–12% intracellular potassium loss from bacteria after 24 h. Flow cytometry data suggested colistin might alter cell arrangement, and SEM confirmed the antibiotic causes bacterial aggregation. Filamentation was not detected in either species at any concentration or time-point up to 24 h. These results argue against the hypotheses that colistin kills bacteria by puncturing the cytoplasmic membrane or disrupting DNA synthesis. The colistin-induced bacterial aggregation detected has implications for the interpretation of MBC, time-kill, and other test results obtained with this antibiotic.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colistin (polymyxin E) is an antibiotic with a spectrum of activity that includes problematic carbapenem-resistant and extensively drug-resistant Gram-negative bacteria such as Pseudomonas aeruginosa and Escherichia coli. Systemic use of colistin largely ceased in the 1970s due to concerns about nephrotoxicity and neurotoxicity, but growing resistance to other antibiotic classes prompted a reassessment of its safety at the turn of the century (Tängdén and Giske 2015; Tran et al. 2016). Under suitable dosage regimens and with careful monitoring, it is now accepted that the risk of colistin-induced kidney or nerve damage can be minimized (Kelesidis and Falagas 2015; Shields et al. 2017), and the antibiotic has been returned to use as a last option or salvage therapy (Poirel et al. 2017). Clinical indications include treatment of ventilator-associated pneumonia and lung infections in cystic fibrosis patients (Gu et al. 2014; Liu et al. 2015) as well as bacteraemia and urinary tract infections (Tängdén and Giske 2015; Bader et al. 2017) caused by extensively drug resistant Gram-negative organisms. Transmissible colistin resistance emerged in 2011 and has spread worldwide, but its prevalence is still quite low (Giske 2015; Liu et al. 2016; Baron et al. 2016).
Because colistin is rapidly bactericidal against Gram-negative but not Gram-positive species, and has a strong binding affinity for lipopolysaccharide (LPS), it is widely accepted that this antibiotic targets the bacterial outer membrane. However, precise details of how colistin exerts its antibacterial effect remain unclear (Tran et al. 2016; Poirel et al. 2017). At the time of its discovery and development in the late 1940s, regulatory and licensing bodies such as the Medicines and Healthcare products Regulatory Agency (MHRA) were not in existence, and the substantial body of microbiological and pharmacological evidence now necessary for licensing an antibiotic was simply not required (Landersdorfer and Nation 2015). Therefore, a relatively limited body of information on the antibacterial effects of colistin is available to inform rational use (Honoré et al. 2014; Theuretzbacher 2014). Elucidation of further details could permit optimization of colistin dosing, prediction of synergistic drug combinations, prediction of drug combinations that reduce the risk of additional resistance emerging, and the development of second-generation polymyxins (Deris et al. 2014; Poirel et al. 2017).
Several recent studies have explored how colistin interacts with the bacterial cell using transmission electron microscopy (TEM), model membranes, and other techniques. Consensus exists that initial electrostatic and hydrophobic interactions occur between colistin molecules, which are positively charged, and the negatively charged LPS layer of the outer membrane. This first step, sometimes referred to as ‘self-promoted uptake’, leads to displacement of the divalent cations (Ca2+ and Mg2+) that normally stabilize the LPS monolayer (Velkov et al. 2013). How colistin then kills bacterial cells is uncertain. One longstanding hypothesis is that colistin molecules disrupt the physical integrity of the cytoplasmic membrane, causing leakage of intracellular contents (Velkov et al. 2010; Poirel et al. 2017). Another hypothesis is that colistin causes the inner layer of the outer membrane and outer layer of the cytoplasmic membrane to come together, resulting in phospholipid exchange, and an osmotic imbalance (Clausell et al. 2003; Velkov et al. 2013). Additional possibilities, not necessarily mutually exclusive, are that colistin inhibits vital respiratory enzymes (type II NADH-quinone oxidoreductases) at the cytoplasmic membrane (Deris et al. 2014), that colistin induces the formation of reactive oxygen species when it crosses the cytoplasmic membrane (Yu et al. 2017) or, in a similar manner to other cationic antimicrobial peptides, that colistin binds to bacterial DNA (Kong et al. 2011) inhibiting replication and transcription.
The purpose of the present study was to gain further insight into the interaction between colistin and bacterial cells including its underlying mechanism of action. Cytoplasmic membrane damage was assessed by measuring potassium loss from colistin-treated cells, as leakage of this intracellular solute is an early sign that membrane integrity has been compromised (Yu et al. 2016; Liang et al. 2016). Colistin-treated bacteria were also examined by flow cytometry and scanning electron microscopy (SEM) as changes in the size and shape of cells can be useful in identifying antibiotic targets too (Peach et al. 2013; Cushnie et al. 2016). To account for the fact that antibiotic-induced morphological changes can vary with antibiotic concentration (Alsteens et al. 2008), inoculum density (Diver and Wise 1986), and test bacterial species (Wojnicz et al. 2007), we examined the effects of sub-inhibitory, inhibitory, and bactericidal concentrations of colistin upon two species, E. coli and P. aeruginosa, using minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) values determined for assay-specific inoculum densities.
Materials and methods
Bacteria
E. coli NCTC 4174, P. aeruginosa NCTC 6750 and Staphylococcus aureus NCTC 6571 were obtained from the National Collection of Type Cultures (Health Protection Agency Culture Collections, UK). These bacteria were stored, sub-cultured, harvested, and washed as described previously (O’Driscoll et al. 2013).
Antibacterial agents, chemical reagents, and growth media
Colistin sulfate, phosphate buffered saline (PBS) tablets, isopropanol (99+%), and sodium phosphate were purchased from Sigma-Aldrich Company (Poole, UK). Potassium dihydrogen orthophosphate SLR and glutaraldehyde were from Fisons Scientific (Loughborough, UK), and sodium carbonate was from BDH (Poole, UK). Sodium chloride (NaCl; general-purpose grade), acetone (technical grade), and formaldehyde were purchased from Fisher Scientific (Loughborough, UK), and nutrient broth and agar were from Oxoid (Basingstoke, UK). The BacLight™ Live/Dead Kit was obtained from Molecular Probes (Invitrogen, Paisley, UK), while flow cytometry Sheath fluid, Flow Check™ fluorospheres and all plastic disposable equipment were from Beckman Coulter (Buckinghamshire, UK).
Determination of MIC and MBC values for colistin
MICs were determined according to the CLSI broth microdilution method (CLSI 2006; Wiegand et al. 2008), but with the following necessary modifications. Rather than using the CLSI-recommended inoculum density of 5 × 105 cfu mL−1 and CLSI-recommended incubation time of 16–20 h, MICs were determined for assay-specific inoculum densities (1 × 106 cfu mL−1 bacteria for the potassium loss and flow cytometry assays, and 1 × 107 cfu mL−1 bacteria for scanning electron microscopy analysis) and assay-specific incubation time of 24 h. Microtitre plates (96-well; Bibby Sterilin, Staffordshire, UK) were sealed with an optically clear, gas-permeable seal (Fisher Scientific) and incubated at 37 °C (Versamax Microplate Reader, Molecular Devices, Berkshire, UK).
MBC values were established by a replica plating procedure, with 1–2 µL from each well transferred to the surface of a 13.5-cm-diameter nutrient agar plate using a 96-pin multi-point replicator (Boekel Scientific, US). The presence or absence of growth was recorded after 24 h aerobic incubation at 37 °C. All assays were conducted in triplicate on six separate occasions.
Quantification of intracellular potassium loss from populations of colistin-treated bacteria
Populations of 1 × 106 cfu mL−1 E. coli and P. aeruginosa were incubated with inhibitory-bactericidal (1 × MIC and 1 × MBC) and supra-bactericidal (10 × MBC) levels of colistin (37 °C; 150 rpm) for various time periods (0, 1, 2, 4, 8 and 24 h), and examined for potassium loss using a flame atomic absorption spectrophotometer (Model AA3110, Perkin Elmer, Beaconsfield, UK) as described previously (O’Driscoll et al. 2013). Untreated cells were used as a negative control. Sub-inhibitory levels of colistin (1/20 × MIC) induced negligible potassium loss during preliminary testing, and were excluded from the study to allow more rapid processing of samples from the other test conditions (i.e. untreated, inhibitory-bactericidal, and supra-bactericidal). Total intracellular potassium content was determined by sonicating each cell sample (Misonix XL 2010 Ultrasonic Liquid Processor; Heat Systems, Farmingdale, US) at 20 kHz for four 30-s pulses, pausing between pulses to place the centrifuge tube on ice. All assays were conducted in triplicate on three separate occasions.
Analysis of colistin-treated bacterial populations by flow cytometry
Populations of 1 × 106 cfu mL−1 E. coli and P. aeruginosa were incubated in nutrient broth containing sub-inhibitory (1/20 × MIC), inhibitory-bactericidal (1 × MIC and 1 × MBC) and supra-bactericidal (10 × MBC) levels of colistin (37 °C for 24 h; 150 rpm) prior to analysis by flow cytometry using BacLight™ Live/Dead Bacterial Viability reagents (Invitrogen). In accordance with the manufacturer’s instructions, bacterial populations incubated in aqueous 0.9% (w/v) NaCl and 70% (v/v) isopropyl alcohol were employed as viable and non-viable controls. All bacterial populations, both control and colistin-treated, were prepared for flow cytometric analysis as follows. Firstly, 10 µL of each bacterial suspension was pipetted into a series of sterile 1.5-mL microcentrifuge tubes containing 987 µL of 0.9% (w/v) NaCl with 1.5 µL of Syto9 and 1.5 µL propidium iodide (PI) fluorescent dye solutions. Tubes were then incubated in the dark for 15 min at room temperature prior to analysis using an Epics®XL MCL flow cytometer (Beckman Coulter). Prior to using the flow cytometer, the discriminator was set at 0.2 µm and the laser alignment was checked and adjusted using Flow Check™ fluorospheres. The test samples were processed once the half-peak co-efficient variation was ≤ 2% on channels FL1, FL2, FL3 and FL4. Channels FL1 and FL3 were set to detect Syto9 and PI, respectively. When being analysed, each sample either had 10,000 events recorded requiring approximately 5 s for controls, or was left for up to 5 min permitting the maximum number of events to be gathered. All assays were conducted in triplicate on three separate occasions.
Examination of the effect of colistin treatment on bacterial cells and populations by scanning electron microscopy
Populations of 1 × 107 cfu mL−1 E. coli and P. aeruginosa were incubated with sub-inhibitory (1/20 × MIC), inhibitory-bactericidal (1 × MIC and 1 × MBC), and supra-bactericidal (10 × MBC) levels of colistin (37 °C; 150 rpm) for various time periods (0, 1, 2, 4, 8, and 24 h), then prepared for and analysed by SEM (LEO S430, Carl Zeiss SMT Ltd., UK) using a protocol described previously (O’Driscoll et al. 2013). A slightly higher bacterial cell density was needed in this assay compared to potassium loss and flow cytometry assays, as cells in untreated control populations of 1 × 106 cfu mL− 1 appear sparse when examined by SEM prior to 24 h incubation (data not shown) and antibiotic-induced morphological changes are known to occur much earlier than 24 h (sometimes within just 1 h of treatment; Cushnie et al. 2016). The size of cells and cell aggregates was determined using LEO S430 software. After extensive examination at a range of magnifications (4000× to 400,000×), micrographs of those fields of view representing typical bacterial characteristics were captured.
Results
Determination of MIC and MBC values for colistin
The MIC and MBC values determined for colistin against the test species of bacteria, at cell densities of 1 × 106 and 1 × 107 cfu mL−1, are shown in Table 1. These results correspond well with previously published MICs and MBCs (Li et al. 2006; Cummins et al. 2009; Alhanout et al. 2010), and confirm the lack of activity of colistin against Gram-positive bacteria.
Quantification of intracellular potassium loss from populations of colistin-treated bacteria
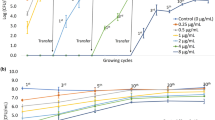
Low level potassium loss was detected in untreated control populations of 1 × 106 cfu mL− 1 E. coli (1.3% at time zero, and 3.6% after 24 h) in this study (Fig. 1a), probably a consequence of shear forces encountered during sample centrifugation (Peterson et al. 2012) and low level osmotic lysis occurring in the hypotonic potassium-free environment (Cushnie et al. 2009). E. coli populations treated with inhibitory-bactericidal (0.28 mg L−1; 1 × MIC and 1 × MBC) and supra-bactericidal (2.8 mg l−1; 10 × MBC) levels of colistin, respectively, lost 5.7 and 5.9% of their total potassium content within 1 h of exposure (Fig. 1a). Potassium loss readings taken at 24 h were not much higher, just 5.9% for 1 × MIC (1 × MBC) treated cells and 11.6% for 10 × MBC treated cells.
Potassium loss from populations of a 1 × 106 cfu mL−1 E. coli and b 1 × 106 cfu mL−1 P. aeruginosa incubated with and without colistin for 24 h (error bars represent standard error of the mean for three assays). The total intracellular potassium content, determined by sonication, was 3.2 mg/L for both E. coli and P. aeruginosa. Triangle: untreated control; unfilled circle: 1 × MIC (equal to 1 × MBC) colistin; unfilled square: 10 × MBC colistin
A similar response was seen with P. aeruginosa (Fig. 1b). Low level potassium loss was detected in untreated control populations of 1 × 106 cfu mL−1 P. aeruginosa (1.4% at time zero, and 3.7% after 24 h). Suspensions of P. aeruginosa treated with inhibitory-bactericidal (1.5 mg L−1; 1 × MIC and 1 × MBC) and supra-bactericidal (15 mg L− 1; 10 × MBC) levels of colistin, respectively, lost 3.8 and 4.0% of their total potassium pool within 1 h, these values remaining unchanged or almost unchanged (3.8 and 4.1%, respectively) when cells were examined at 24 h.
Analysis of colistin-treated bacterial populations by flow cytometry
To further scrutinize the effects of colistin, we sampled and analysed by flow cytometry populations of 1 × 106 cfu mL−1 bacteria exposed to the antibiotic for 24 h. For populations of E. coli treated with sub-inhibitory (1/20 × MIC) levels of colistin (Fig. 2b), there was only a slight difference compared to the untreated control (Fig. 2a). In samples of E. coli treated with inhibitory-bactericidal (1 × MIC and 1 × MBC) and supra-bactericidal (10 × MBC) levels of colistin, by contrast, very few events were detected in the gated population (Fig. 2c, d). The discriminator was set at a threshold of 0.2 µm such that only cells or structures larger than this and within the gate would be detected.
Flow cytometric analysis of populations of 1 × 106 cfu mL− 1 E. coli incubated for 24 h. a Without colistin, b with 1/20 × MIC colistin, c with 1 × MIC (equal to 1 × MBC) colistin, and d with 10 × MBC colistin. FL1 LOG channel measured Syto9 fluorescence and FL3 LOG channel measured propidium iodide (PI) fluorescence
The P. aeruginosa control culture (Fig. 3a) appeared more heterogeneous than the E. coli control culture (Fig. 2a), with two sub-groups apparent within the gated population. These two sub-groups probably reflect diversity of individual bacterial cell size in a normal growing population, and simply indicate that overnight P. aeruginosa cultures contain cells which are in two size groupings. Having established that two groups exist, examination of bacterial populations following treatment with sub-inhibitory (1/20 × MIC) colistin revealed a decrease in number of the smaller-sized cells and a slight increase in number of the larger-sized cells (Fig. 3b). In the P. aeruginosa cultures treated with inhibitory-bactericidal (1 × MIC and 1 × MBC) and supra-bactericidal (10 × MBC) levels of colistin, very few events were detected in the gated population (Fig. 3c, d).
Flow cytometric analysis of populations of 1 × 106 cfu mL−1 P. aeruginosa incubated for 24 h. a Without colistin, b with 1/20 × MIC colistin, c with 1 × MIC (equal to 1 × MBC) colistin, and d with 10 × MBC colistin. FL1 LOG channel measured Syto9 fluorescence and FL3 LOG channel measured propidium iodide (PI) fluorescence
Examination of the effect of colistin treatment on bacterial cells and populations by SEM
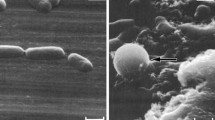
Treatment of populations of 1 × 107 cfu mL−1 E. coli with sub-inhibitory (0.49 mg L−1; 1/20 × MIC), inhibitory-bactericidal (9.8 mg L−1; 1 × MIC and 1 × MBC), and supra-bactericidal (98 mg l−1; 10 × MBC) levels of colistin triggered an immediate response. Within 60 min, profound cell aggregation was observed, these aggregates being spherical in shape and measuring 5–15 µm in diameter depending on the antibiotic concentration (Fig. 4). Aggregates remained visible in the colistin-treated populations at all concentrations [1/20 × MIC, 1 × MIC (1 × MBC), and 20 × MBC] and time-points (1, 2, 4, 8, and 24 h; data shown for 60 minutes only), although in populations treated with sub-inhibitory colistin (1/20 × MIC) the aggregates displayed more wide-ranging sizes (Fig. S1; Supplementary material). No change in cell length was detected at any colistin concentration or any time-point with E. coli. No blebbing of the bacterial cell surface was detected at any concentration or time-point with E. coli either.
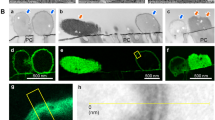
For populations of 1 × 107 cfu mL−1 P. aeruginosa treated with sub-inhibitory (0.25 mg l−1; 1/20 × MIC), inhibitory-bactericidal (4.9 mg L−1; 1 × MIC and 1 × MBC), and supra-bactericidal (49 mg l−1; 10 × MBC) levels of colistin, the morphological changes observed were both concentration- and time-dependent. After 60 min, a decrease in the length of individual cells was detected in populations treated with sub-inhibitory (1/20 × MIC) and inhibitory-bactericidal (1 × MIC and 1 × MBC) levels of colistin (Fig. 5b and c). Compared to control cells which were 2.0–2.5 µm in length, cells treated with sub-inhibitory levels of colistin were just 0.9–1.8 µm in length, and cells treated with inhibitory-bactericidal levels of colistin were only ~ 0.8 µm in length. Aggregation was only detected at the highest colistin concentration (10 × MBC) at 60 min (Fig. 5d), but became apparent for the other two concentrations [1/20 × MIC and 1 × MIC (1 × MBC)] by 2 h (data not shown). This aggregation initially manifested as layers of bacteria interlinked by strands of extracellular material, with spherical clumps not detectable until 8 h and remaining extremely sparse until 24 h (Fig. S2c and S2d; Supplementary material). Blebbing of the bacterial cell surface was detected from 8 h onwards (data not shown).
Discussion
The polymyxin antibiotic colistin has recently been resurrected for the treatment of infections with extensively drug-resistant Gram-negative bacteria. We investigated the impact of a range of colistin concentrations upon E. coli and P. aeruginosa populations to obtain a clearer understanding of how this antibiotic functions. To determine if colistin compromises cytoplasmic membrane integrity as part of its antibacterial mechanism of action, we examined populations of E. coli and P. aeruginosa for loss of the intracellular solute potassium. The data obtained show that, even after 24 h treatment with supra-bactericidal (10 × MBC) levels of colistin, E. coli populations lost less than 12% of their total potassium pool and P. aeruginosa populations lost less than 5%. Genuinely cytoplasmic membrane-active agents such as protegrin and cathelicidin have, by comparison, been shown to induce 90–100% potassium loss in similar studies with E. coli in less than 1 h (Orlov et al. 2002; Bolintineanu et al. 2010). Overall, the fact that colistin-treated populations retain more than 85% of their total potassium pool confirms the cytoplasmic membrane remains intact, if not functional, in most of the treated cells. Results presented here correlate well with two previous studies, the first by Zhang et al. (2000) which showed that colistin exerts a lethal effect on P. aeruginosa without affecting cytoplasmic membrane proton motive force, and the second by Alhanout and colleagues (2010), which showed that 4 h treatment with 4 × MIC colistin led to E. coli and P. aeruginosa populations losing just 4–5% of their total intracellular ATP. Taken together, these findings argue against the longstanding and widely cited hypothesis that colistin kills bacterial cells by puncturing the cytoplasmic membrane.
In the next stage of the study, antibiotic-treated populations of bacteria were examined by flow cytometry, using Syto9 and PI fluorescent dyes to identify intact cells of viable and non-viable bacteria, respectively. For E. coli, treatment with inhibitory-bactericidal (1 × MIC and 1 × MBC) or supra-bactericidal (10 × MBC) levels of colistin resulted in a near-complete loss of signal from the gated population (Fig. 2c, d). In the absence of large-scale potassium loss (Fig. 1a), this result cannot be due to colistin-induced cell lysis. Rather, colistin must be altering the size or arrangement of cells in such a way that they are not being detected. With P. aeruginosa, flow cytometry revealed the presence of two slightly differently sized sub-populations in overnight cultures (Fig. 3a), the number of smaller-sized cells decreasing following treatment with sub-inhibitory (1/20 × MIC) levels of colistin (Fig. 3b). This pattern of results might suggest that a cell cycle event subsequent to septation and cell separation renders bacteria susceptible to colistin-induced cell lysis. Perhaps more likely though, given the accompanying increase in number of larger-sized cells (Fig. 3b), colistin is inhibiting cell separation or causing cell aggregation. As with E. coli, treatment of P. aeruginosa with inhibitory-bactericidal (1 × MIC and 1 × MBC) or supra-bactericidal (10 × MBC) levels of colistin resulted in a near-complete loss of signal (Fig. 3c, d), suggesting colistin modifies cell size or arrangement in such a way that they are no longer detected.
SEM studies were performed next to visualize directly the impact of colistin upon whole populations of E. coli and P. aeruginosa. With E. coli, the observation of colistin-induced aggregation of bacterial cells (Fig. 4 and Fig. S1) may explain the small number of events detected during flow cytometry (Fig. 2c, d) as aggregation would greatly reduce the number of individual cells present. Colistin-induced aggregation of bacteria has not, to our knowledge, been reported previously, but may be more readily detectable by SEM than TEM or atomic force microscopy. Aggregation similar to this has been reported in bacteria treated with other cationic antimicrobial peptides (O’Driscoll et al. 2013), but the reason for this population-level change is not clear. It may be a direct effect of the antibiotic, for example colistin molecules inserting into the outer membrane of cells and promoting intercellular attachment in a manner comparable to that observed with liposomes (Wallace et al. 2012). Alternatively, it may be a coordinated response on the part of the bacterial population to minimize antibiotic exposure. Young suggests the ability of bacteria to form aggregates is an evolutionary response to predation by protozoa (Young 2007) and, as observed with bacterial biofilms (Costerton et al. 1999), aggregation offers protection from antibiotic treatment also. Unlike previous studies with colistin-treated E. coli (Koike et al. 1969), we did not observe any blebbing of the bacterial outer membrane, but this is more readily detectable by TEM than SEM (Cushnie et al. 2016).
With P. aeruginosa, sub-inhibitory (1/20 × MIC) and inhibitory-bactericidal (1 × MIC and 1 × MBC) levels of the antibiotic both caused a decrease in bacterial cell length (Fig. 5 and Fig. S2) within 1 h. These results correlate well with an atomic force microscopy study by Mortensen et al., which found that sub-MBC colistin-treated cells of P. aeruginosa also exhibited reduced cell length (Mortensen et al. 2009). These observations suggest that colistin treatment, either directly or indirectly, leads to inhibition of lateral cell wall formation. Possible targets include the cytoskeletal Mre system (proteins MreB, MreC and MreD) or enzyme PBP2, both of which are essential for lateral peptidoglycan synthesis and elongation of the bacterial cell (Osborn and Rothfield 2007; Varma and Young 2009). Both the Mre system and PBP2 are anchored to the cytoplasmic membrane (Adachi et al. 1987; Osborn and Rothfield 2007), and would be vulnerable to any colistin-induced conformational changes taking place at this locale. Colistin-induced aggregation of P. aeruginosa cells, whilst slightly different in appearance to that visualised with E. coli, was observed (Fig. 5 and Fig. S2) and may again account for the small number of events detected during flow cytometry (Fig. 3c, d). Blebbing of colistin-treated P. aeruginosa was also detected in this study, an observation reported previously by Koike et al. (1969) and Alhanout et al. (2010). This ultrastructural change, because it is induced by multiple membrane-active agents including chlorhexidine and EDTA, is considered an indicator of possible outer membrane disruption (Cushnie et al. 2016).
When the SEM results for E. coli and P. aeruginosa are viewed conjointly, it is clear there are both similarities and differences in how colistin affects these bacteria. Filamentation was conspicuous by its absence in both species. Filamentation is a hallmark of DNA damage, induced via the SOS response by a diverse range of physical and chemical agents including UV, cosmomycin D, metronidazole, mitomycin C, the fluoroquinolones, novobiocin, and zidovudine (Cushnie et al. 2016). The failure of colistin to induce filamentation at any concentration or time-point in either species argues strongly against the hypothesis that colistin kills bacterial cells through DNA damage or disruption of DNA synthesis. Colistin-induced cell aggregation was, conversely, detected in both species. Whilst it is not clear how, or even if, this aggregation relates to colistin mechanism of action, it could have important implications for diagnostic laboratories. Colistin-induced clumping of cells would reduce colony forming unit (cfu) numbers of bacteria in MBC and time-kill assays, resulting in artificially low MBC values for the antibiotic. Colistin-induced cell aggregation could also complicate how the course of infections is monitored because, in patients receiving this antibiotic, false-negative blood and urine cultures would be more likely to occur. Moving on to differences in how colistin affects the bacteria, the most noticeable of these was the reduction in cell size observed with P. aeruginosa but not E. coli. This might be due to slight variations in the cell envelope composition or PBP2/Mre protein conformation of the two species. Ultimately, this and other observed interspecies variations probably reflect indirect or peripheral aspects of colistin’s mechanism of action as both bacterial species are susceptible to the antibiotic.
In conclusion, our study has shown that colistin induces minimal potassium loss and no filamentation in populations of E. coli or P. aeruginosa, findings which indicate the antibiotic does not exert its antibacterial effect by perforating the cytoplasmic membrane or disrupting DNA synthesis. Future studies should therefore focus on exploring the three remaining hypotheses, namely that colistin kills bacterial cells by (1) inducing phospholipid exchange between the outer and cytoplasmic membranes, (2) inhibiting respiratory enzymes, and/or (iii) inducing reactive oxygen species formation. Colistin-induced aggregation of bacterial cells hindered our attempt to study this antibiotic’s effects by flow cytometry, but represents an interesting observation in itself, one with implications for the interpretation of MBC, time-kill, and other microbiology assay results.
References
Adachi H, Ohta T, Matsuzawa H (1987) A water-soluble form of penicillin-binding protein 2 of Escherichia coli constructed by site-directed mutagenesis. FEBS Lett 226:150–154
Alhanout K, Malesinki S, Vidal N, Peyrot V, Rolain J-M, Brunel JM (2010) New insights into the antibacterial mechanism of action of squalamine. J Antimicrob Chemother 65:1688–1693
Alsteens D, Verbelen C, Dague E, Raze D, Baulard AR, Dufrêne YF (2008) Organization of the mycobacterial cell wall: a nanoscale view. Pflugers Arch Eur J Physiol 456:117–125
Bader MS, Loeb M, Brooks AA (2017) An update on the management of urinary tract infections in the era of antimicrobial resistance. Postgrad Med 129:242–258
Baron S, Hadjadj L, Rolain J-M, Olaitan AO (2016) Molecular mechanisms of polymyxin resistance: knowns and unknowns. Int J Antimicrob Agents 48:583–591
Bolintineanu D, Hazrati E, Davis HT, Lehrer RI, Kaznessis YN (2010) Antimicrobial mechanism of pore-forming protegrin peptides: 100 pores to kill E. coli. Peptides 31:1–8
Clausell A, Pujol M, Alsina MA, Cajal Y (2003) Influence of polymyxins on the structural dynamics of Escherichia coli lipid membranes. Talanta 60:225–234
CLSI (2006) Clinical and Laboratory Standards Institute. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically: approved standard—seventh edition. CLSI document M7-A7. Wayne, PA
Costerton JW, Stewart PS, Greenberg EP (1999) Bacterial biofilms: a common cause of persistent infections. Science 284:1318–1322
Cummins J, Reen FJ, Baysse C, Mooij MJ, O’Gara F (2009) Subinhibitory concentrations of the cationic antimicrobial peptide colistin induce the pseudomonas quinolone signal in Pseudomonas aeruginosa. Microbiology 155:2826–2837
Cushnie TPT, Robertson PKJ, Officer S, Pollard PM, McCullagh C, Robertson JMC (2009) Variables to be considered when assessing the photocatalytic destruction of bacterial pathogens. Chemosphere 74:1374–1378
Cushnie TPT, O’Driscoll NH, Lamb AJ (2016) Morphological and ultrastructural changes in bacterial cells as an indicator of antibacterial mechanism of action. Cell Mol Life Sci 73:4471–4492
Deris ZZ, Akter J, Sivanesan S, Roberts KD, Thompson PE, Nation RL, Li J, Velkov T (2014) A secondary mode of action of polymyxins against Gram-negative bacteria involves the inhibition of NADH-quinone oxidoreductase activity. J Antibiot 67:147–151
Diver JM, Wise R (1986) Morphological and biochemical changes in Escherichia coli after exposure to ciprofloxacin. J Antimicrob Chemother 18:31–41
Giske CG (2015) Contemporary resistance trends and mechanisms for the old antibiotics colistin, temocillin, fosfomycin, mecillinam and nitrofurantoin. Clin Microbiol Infect 21:899–905
Gu W-J, Wang F, Tang L, Bakker J, Liu J-C (2014) Colistin for the treatment of ventilator-associated pneumonia caused by multidrug-resistant Gram-negative bacteria: a systematic review and meta-analysis. Int J Antimicrob Agents 44:477–485
Honoré PM, Jacobs R, de Regt J, van Gorp V, de Waele E, Spapen HD (2014) Colistin dosing for treatment of multidrug-resistant Pseudomonas in critically ill patients - please, be adequate! Crit Care 18:412
Kelesidis T, Falagas ME (2015) The safety of polymyxin antibiotics. Expert Opin Drug Saf 14:1687–1701
Koike M, Iida K, Matsuo T (1969) Electron microscopic studies on mode of action of polymyxin. J Bacteriol 97:448–452
Kong L, Liu Z, Hu X, Liu S (2011) Interaction of polymyxin B with ds-DNA, and determination of DNA or polymyxin B via resonance Rayleigh scattering and resonance non-linear scattering spectra. Microchim Acta 173:207–213
Landersdorfer CB, Nation RL (2015) Colistin: how should it be dosed for the critically ill? Semin Respir Crit Care Med 36:126–135
Li J, Nation RL, Turnidge JD, Milne RW, Coulthard K, Rayner CR, Paterson DL (2006) Colistin: the re-emerging antibiotic for multidrug-resistant Gram-negative bacterial infections. Lancet Infect Dis 6:589–601
Liang J, Deng J, Li M, Tong M (2016) Bactericidal activity and mechanism of AgI/AgBr/BiOBr0.75I0.25 under visible light irradiation. Colloids Surf B Biointerfaces 138:102–109
Liu D, Zhang J, Liu H-X, Zhu Y-G, Qu J-M (2015) Intravenous combined with aerosolised polymyxin versus intravenous polymyxin alone in the treatment of pneumonia caused by multidrug-resistant pathogens: a systematic review and meta-analysis. Int J Antimicrob Agents 46:603–609
Liu Y-Y, Wang Y, Walsh TR, Yi L-X, Zhang R, Spencer J, Doi Y, Tian G et al (2016) Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis 16:161–168
Mortensen NP, Fowlkes JD, Sullivan CJ, Allison DP, Larsen NB, Molin S, Doktycz MJ (2009) Effects of colistin on surface ultrastructure and nanomechanics of Pseudomonas aeruginosa cells. Langmuir 25:3728–3733
O’Driscoll NH, Labovitiadi O, Cushnie TPT, Matthews KH, Mercer DK, Lamb AJ (2013) Production and evaluation of an antimicrobial peptide-containing wafer formulation for topical application. Curr Microbiol 66:271–278
Orlov DS, Nguyen T, Lehrer RI (2002) Potassium release, a useful tool for studying antimicrobial peptides. J Microbiol Methods 49:325–328
Osborn MJ, Rothfield L (2007) Cell shape determination in Escherichia coli. Curr Opin Microbiol 10:606–610
Peach KC, Bray WM, Winslow D, Linington PF, Linington RG (2013) Mechanism of action-based classification of antibiotics using high-content bacterial image analysis. Mol BioSyst 9:1837–1848
Peterson BW, Sharma PK, van der Mei HC, Busscher HJ (2012) Bacterial cell surface damage due to centrifugal compaction. Appl Environ Microbiol 78:120–125
Poirel L, Jayol A, Nordmann P (2017) Polymyxins: antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes. Clin Microbiol Rev 30:557–596
Shields RK, Anand R, Clarke LG, Paronish JA, Weirich M, Perone H, Kieserman J, Freedy H et al (2017) Defining the incidence and risk factors of colistin-induced acute kidney injury by KDIGO criteria. PLOS ONE 12:e0173286
Tängdén T, Giske CG (2015) Global dissemination of extensively drug-resistant carbapenemase-producing Enterobacteriaceae: clinical perspectives on detection, treatment and infection control. J Intern Med 277:501–512
Theuretzbacher U (2014) Product information for parenteral colistin varies substantially across Europe. J Antimicrob Chemother 69:1987–1992
Tran TB, Velkov T, Nation RL, Forrest A, Tsuji BT, Bergen PJ, Li J (2016) Pharmacokinetics/pharmacodynamics of colistin and polymyxin B: are we there yet? Int J Antimicrob Agents 48:592–597
Varma A, Young KD (2009) In Escherichia coli, MreB and FtsZ direct the synthesis of lateral cell wall via independent pathways that require PBP2. J Bacteriol 191:3526–3533
Velkov T, Thompson PE, Nation RL, Li J (2010) Structure-activity relationships of polymyxin antibiotics. J Med Chem 53:1898–1916
Velkov T, Roberts KD, Nation RL, Thompson PE, Li J (2013) Pharmacology of polymyxins: new insights into an ‘old’ class of antibiotics. Future Microbiol 8:711–724
Wallace SJ, Li J, Nation RL, Prankerd RJ, Boyd BJ (2012) Interaction of colistin and colistin methanesulfonate with liposomes: colloidal aspects and implications for formulation. J Pharm Sci 101:3347–3359
Wiegand I, Hilpert K, Hancock REW (2008) Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat Protoc 3:163–175
Wojnicz D, Kłak M, Adamski R, Jankowski S (2007) Influence of subinhibitory concentrations of amikacin and ciprofloxacin on morphology and adherence ability of uropathogenic strains. Folia Microbiol 52:429–436
Young KD (2007) Bacterial morphology: why have different shapes? Curr Opin Microbiol 10:596–600
Yu L, Zhu M, Liu E, Yang T, Chen X, Wang X (2016) Studies of the mechanism of an antibacterial peptide (cecropinA-magainin) on methicillin-resistant Staphylococcus aureus membranes. Biotechnol Appl Biochem 63:805–811
Yu Z, Zhu Y, Qin W, Yin J, Qiu J (2017) Oxidative stress induced by polymyxin E is involved in rapid killing of Paenibacillus polymyxa. BioMed Res Int (Article 5437139). https://doi.org/10.1155/2017/5437139
Zhang L, Dhillon P, Yan H, Farmer S, Hancock REW (2000) Interactions of bacterial cationic peptide antibiotics with outer and cytoplasmic membranes of Pseudomonas aeruginosa. Antimicrob Agents Chemother 44:3317–3321
Acknowledgements
This work was supported in part by an award (G08/10) from Tenovus Scotland. The authors would like to thank Dr Derry Mercer for helpful discussions, Mrs Emily Hunter and Mr Iain Tough for assistance with operation of the SEM, Dr Tim King for valuable assistance with interpretation of SEM images, Mr Steve Darwood from Beckman Coulter Ltd. for ongoing help and advice with flow cytometry, and Professor Paul Kong Thoo Lin for constructive comments during drafting of the manuscript. Also we wish to thank our friend and colleague Miss Vivienne Hamilton, who very sadly passed away during this project, for all her technical support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Erko Stackebrandt.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
O’Driscoll, N.H., Cushnie, T.P.T., Matthews, K.H. et al. Colistin causes profound morphological alteration but minimal cytoplasmic membrane perforation in populations of Escherichia coli and Pseudomonas aeruginosa. Arch Microbiol 200, 793–802 (2018). https://doi.org/10.1007/s00203-018-1485-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-018-1485-3