Abstract
Repairing DNA damage is one of the most important functions of the ‘housekeeping’ proteins, as DNA molecules are constantly subject to different kinds of damage. An important mechanism of DNA repair is the mismatch repair system (MMR). In eukaryotes, it is more complex than it is in bacteria or Archaea due to an inflated number of paralogues produced as a result of an extensive process of gene duplication and further specialization upon the evolution of the first eukaryotes, including an important part of the meiotic machinery. Recently, the discovery and sequencing of Asgard Archaea allowed us to revisit the MMR system evolution with the addition of new data from a group that is closely related to the eukaryotic ancestor. This new analysis provided evidence for a complex evolutionary history of eukaryotic MMR: an archaeal origin for the nuclear MMR system in eukaryotes, with subsequent acquisitions of other MMR systems from organelles.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The DNA is relatively unstable and subject to frequent breaks and several kinds of damage. DNA damage may be a result of different physical and chemical agents, including UV radiation, gamma rays, free oxygen radicals, replication errors, among others (Lindahl 1993). The consequences of DNA damage range from point mutations to impairment of DNA function, compromising RNA transcription and protein synthesis, sometimes leading to cellular failure and death (Jackson and Bartek 2009). The evolution of DNA repair machineries allowed for efficient and specific DNA repair according to damage type and rescue of genetic functions. Such machineries became a fundamental part of cellular housekeeping genes. For each kind of damage, there is a specific repair system (Sancar et al. 2004). The well-characterized DNA repair systems so far include homologous recombination (Seitz et al. 1998), photolyases (Essen and Klar 2006), base excision repair (Krokan and Bjørås 2013), nucleotide excision repair (Schärer 2013), non-homologous end joining (Chang et al. 2017), mismatch repair (Li 2008) and polymerases (Wood and Shivji 1997). Most of them are widely distributed in all cellular domains (Bacteria, Archaea and Eukarya) and bear a high level of primary sequence conservation, as in the case of the recombinases (recA superfamily) implied in homologous recombination and mutS–mutL involved in mismatch repair (MMR) system (Lin et al. 2006, 2007).
Eukaryotic recombinases have been proposed to be related to Archaeal recombinases, which underwent ancestral duplication events in order to provide the several eukaryotic paralogues (Lin et al. 2006), one of them a component of the meiosis-specific machinery (Bishop et al. 1992). There is a second kind of recombinase in eukaryotes, this one with a bacterial signature. In this case, their origin can be traced back to both Alpha-proteobacteria and Cyanobacteria. They are a result of endosymbiotic gene transfers from mitochondria and chloroplasts to eukaryotes and because they possess a targeting domain, are thought to be active only inside endosymbiotic organelles (Hofstatter et al. 2016). Similarly, the eukaryotic MMR system proteins MutS and MutL homologues evolved by means of a series of gene duplications yielding several paralogues in the ancestral of eukaryotes (Lin et al. 2007), some of which present meiosis-specific functions as well (Hollingsworth et al. 1995). However, differently from the recombinases, the MutS–MutL system implied in nuclear MMR in eukaryotes has been proposed to be a contribution from the mitochondrial ancestor to early eukaryotes (Lin et al. 2007). Interestingly, bacterial mutS and mutL occur in some Archaeal groups as well, probably as a result of horizontal gene transfers (HGT) from Bacteria to Archaea (Lin et al. 2007). Recent evidence further suggests that two kinds of MMR systems evolved independently in the ancestors of both Bacteria and Archaea after their divergence, the NucS system in Archaea and mutS–mutL system in Bacteria with some level of exchange of these genes later (Castañeda-García et al. 2017; Takemoto et al. 2018). The MutS dimer works together with MutL dimer in order to perform DNA repair. A dimer of MutS proteins identifies and is attracted to a site of mismatch on a DNA molecule and recruits a dimer of MutL, which cuts and removes the mismatch, allowing the action of DNA polymerases and DNA ligases to fill the gap and finish the process (Modrich and Lahue 1996). Some bacterial groups (e.g. Escherichia coli) also present MutH, which bias the nicking and removing the mismatched site towards the unmethylated strand, i.e. the newly synthesized one (Smith and Modrich 1996). Although the system bears a clear archaeal signature, NucS occurs in Actinobacteria, probably due to an early HGT from Archaea to this bacterial group. In addition, both systems may occur in some archaeal groups (Castañeda-García et al. 2017). Even though archaeal in nature, NucS homologues do not occur in eukaryotes and were probably lost in the transition from Archaea to the first eukaryotes.
The bacterial MMR system composed of mutS and mutL was acquired by eukaryotes and ancestrally duplicated several times producing the eukaryotic paralogues that occur in most eukaryotic lineages (Modrich and Lahue 1996; Lin et al. 2007). Six mutS eukaryotic paralogues are widespread in eukaryotes, namely MSH1-6; additionally, there are four mutL paralogues, namely MLH1-4. All of them interact in specific ways with each other. Paralogues MSH4 and MSH5 are meiosis-specific and do not realize MMR anymore, but participate of crossing-over resolution during meiosis (Hollingsworth et al. 1995). The evolution of MMR in eukaryotes is linked to the evolution of meiosis itself. All the eukaryotic mutS homologues are highly conserved and operate MMR not only inside the nucleus, but also inside organelles as well. The activity of mutS homologues inside organelles, the bacterial signature of this DNA repair system, and phylogenetic patterns observed by Lin and colleagues (Lin et al. 2007) led to the assumption that the whole eukaryotic MMR system was acquired by eukaryotes laterally from Alpha-proteobacteria upon the mitochondrial endosymbiotic event. However, the discovery of a new group of Archaea closely related to eukaryotes, the Asgard Archaea, offered the opportunity to revisit this matter (Zaremba-Niedzwiedzka et al. 2017). In this study, we revisit the subject with more comprehensive data and more advanced techniques in order to verify the putative mitochondrial contribution to meiosis in eukaryotes.
Results
Contrary to previous conclusions (Lin et al. 2007), the main group of eukaryotic mutS paralogues active inside the nucleus (paralogues MSH2-6) are not a mitochondrial contribution but are a result of gene duplications of a mutS gene present in the archaeal ancestor. The archaeal ancestor in turn, probably acquired the mutS laterally from a bacterial group very long ago, before the evolution of the first eukaryotes (Fig. 1, Suppl. Tree 1). In Asgard Archaea, mutS and mutL occur in tandem in the same way they occur in Firmicutes. This fact makes Firmicutes a good candidate for a donor of the bacterial MMR to Asgard group. The same reconstructions reveal the mitochondrial origin of eukaryotic MSH1, which has an Alpha-proteobacterial signature. Still in the same reconstructions, there is a third group of eukaryotic mutS with cyanobacterial origin, acquired from the chloroplast ancestor in some photosynthesizing groups.
mutS protein tree. The eukaryotic homologues of bacterial mutS have three different sources: one associated with Asgard Archaea (nuclear mutS homologues MSH2, MSH3, MSH4, MSH5, MSH6, MSH7), one associated with Alpha-proteobacteria (mitochondrial MSH1), and another one associated with Cyanobacteria (chloroplastic MSH1)
The same applies to mutL: mutS and mutL were probably acquired together by the archaeal group that gave origin to eukaryotes and then both underwent gene duplications that are an important step in early eukaryotic evolution. Systematic reconstructions of both mutS and mutL proteins exhibit similar patterns. Patterns observed in the mutL reconstructions are similar to the ones observed in the case of mutS (Fig. 2, Supple. Tree 2). The nuclear paralogues MLH1-4 exhibit the same patterns of the nuclear MHS2-6. Both mutS and mutL must have been acquired together, what are expected as both work together as part of the same complex. The mitochondrial mutL seems to have been lost and probably replaced by some nuclear paralogue. The chloroplastic mutL was seemingly lost in the green algae, but kept by red algae. A concatenation of mutS and mutL provides a similar result regarding the evolution of both proteins when treated alone, but at this time, all Heimdallarchaea are attracted towards the nuclear eukaryotic group (Suppl. Fig. 1, Suppl. Tree 3). This may be interpreted as an increase in the signal of the reconstruction, assuming both genes evolved in concerted evolution.
mutL protein tree. The eukaryotic mutL homologues have two distinguishable sources: the nuclear mutL homologues (MLH1, MLH3, MLH3, MLH4) are associated with Asgard Archaea, and another branch of eukaryotic mutL are associated with Cyanobacteria (chloroplast). No mitochondria-associated MutL proteins could be traced back to Alpha-proteobacteria
Discussion
The importance of the MMR repair for maintenance of the integrity of the DNA and the high level of conservation of its components across all cellular domains have raised questions about the evolution of the whole MMR system, specially in eukaryotes (Lin et al. 2007). However, by revisiting this matter, we have found a very different history concerning mutS and mutL evolution. Back then, a mitochondrial origin for all eukaryotic paralogues was proposed, including both nuclear and mitochondrial forms. The chloroplastic ones could not be distinguished with those data. The introduction of Asgard data was crucial for retelling this history. Asgard is an archaeal group closely related to eukaryotes, or at least, is the closest known archaeal group to eukaryotes that was sampled to this day (Zaremba-Niedzwiedzka et al. 2017).
Eukaryotic mutS homologues were acquired three times independently by eukaryotes: one vertically from archaeal ancestors, then laterally from the mitochondrial ancestor, and again laterally from the chloroplastic ancestor in the photosynthesizing lineages (Fig. 3).
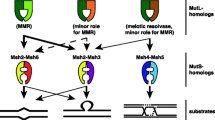
Main events in the evolution of mismatch repair machinery composed by mutS–mutL system: Eukaryotic mutS and mutL were inherited vertically from archaeal ancestors and then acquired laterally twice upon endosymbiotic events. The transition from Archaea to eukaryotes was characterized by large-scale duplication events to produce all the nuclear paralogues; later, an acquisition from the mitochondrial ancestor, which is encoded in the nuclear genome, but is active inside the organelles, and again from the chloroplast in the photosynthesizing eukaryotes, also encoded in the nuclear genome, but active inside de organelles. Heimdallarchaeota is sister group to all eukaryotic nuclear paralogues. Archaeal groups themselves seem to have laterally acquired the mutS–mutL system from some bacterial donor once or more
The discovery that the nuclear MMR duties are carried out in eukaryotes by proteins of archaeal origin is more plausible than a mitochondrial origin for this process because other kinds of DNA repair systems that occur in eukaryotes are also of archaeal origin (Lin et al. 2006; Malik et al. 2007). Not only DNA repair systems, but also eukaryotic DNA replication and transcription machineries are of archaeal origin (Kwapisz et al. 2008; Makarova et al. 2014). Contributions of the archaeal ancestor tend to be ‘informational’, while mitochondrial (and chloroplastic) contributions tend to be ‘operational’ in nature (Rivera et al. 1998). Proteins active inside organelles that are synthesized in the cell cytoplasm and imported by specialized mechanisms into mitochondria and chloroplasts (Wiedemann et al. 2004). Mitochondria and chloroplasts import the products of genes that were endosymbiotically transferred to the nuclear genome by means of a system of transit peptides that precede active sites of the mature active proteins implied in the process (Neupert and Herrmann 2007). Earlier analyses suggest a mitochondrial contribution of Alpha-proteobacteria to eukaryotic DNA repair system and meiosis (Lin et al. 2007); the new results support a very different history, which implies no mitochondrial contribution known so far to eukaryotic nuclear processes. In addition, the importance of the mitochondrial endosymbiont to meiosis was overestimated. As the mitochondrial variable seems to have been isolated here, the understanding of the evolution of meiosis may be developed independently from the mitochondrial symbiosis event. In this scenario, sex itself may be older than the mitochondrion in eukaryotes. The discovery of the Asgard Archaea group played a fundamental role at establishing a new version for the evolutionary history of the MMR in eukaryotes.
Material and Methods
A strategic sampling of mutS and mutL representatives was performed in order to maximize the number of bacterial and archaeal phyla in the sampling, while only a few eukaryotic representatives were selected. The mismatch repair system based on MutS–MutL proteins is almost ubiquitous in Eubacteria, except for Actinobacteria, where it was seemingly replaced by NucS (Castañeda-García et al. 2017). Thus, a handful of representatives of every major bacterial phylum was included. Preference was given to well-characterized species with whole genome sequenced of Proteobateria, Cyanobacteria, Chloroflexi, Chlamydia, Deinococcus, Firmicutes, Thermotogae and Spirochaetes, which account for the majority of extant bacterial lineages and include the most probable ancestors of eukaryotic organelles (mitochondria belong to Alpha-proteobacteria and chloroplasts belong to Cyanobacteria). A similar approach was applied to the case of Archaea with a special attention given to the Asgard Archaea group because of its recently proposed close relationship to eukaryotes (Zaremba-Niedzwiedzka et al. 2017). In the case of eukaryotes, very distantly related eukaryotic representatives have been selected in order to represent a less biased sampling of the group: Octopus (Metazoa), Arabidopsis (green plants), Saccharomyces (Fungi), Dictyostelium (Amoebozoa) and Monocercomonoides (a flagellated protist assumed to be early branching among eukaryotes).
For the construction of HMMER profiles, we employed sequences from mismatch repair obtained and characterized from eukaryotic model organisms, such as Saccharomyces, Arabidopsis or Homo. They were used as a starting point. Different eukaryotic paralogues were aligned with 100 iterations of MAFFT (Katoh and Standley 2013) aligner for precision. The aligned set of proteins was given as input for HMMER tool hmmbuild (Eddy 2011), which was employed to build both MutS and MutL profiles used in the searches. Profiles were employed by HMMER tool hmmsearch (Eddy 2011) to search in the databases (built with strategic sampling) for MutS and MutL. Best hits were extracted from databases with HMMER tool esl-sfetch (Eddy 2011) for further processing. Again, resulting sequences were aligned with 100 iterations of MAFFT (Katoh and Standley 2013); the resulting matrix trimmed with Trimal (Capella-Gutiérrez et al. 2009) where the sites had more than 50% indels/unaligned positions. IQ-Tree (Nguyen et al. 2015) was chosen as a state-of-art algorithm for reconstructions. Heavy mixture models (LG + C60 + F + G) were set for reconstructions for both MutS- and MutL-trimmed matrices. Standard number of bootstrap replicates (1000) was applied. An additional concatenated MutS–MutL matrix was produced for a third tree (in this case, eukaryotic MLH1 and MSH6 were arbitrarily chosen). In addition, Asgard data were manually checked to verify the relative position of mutS and mutL homologues in relation to each other in the available Asgard contigs.
References
Bishop DK, Park D, Xu L, Kleckner N (1992) DMC1: a meiosis-specific yeast homolog of E. coli recA required for recombination, synaptonemal complex formation, and cell cycle progression. Cell 69:439–456
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973. https://doi.org/10.1093/bioinformatics/btp348
Castañeda-García A, Prieto AI, Rodríguez-Beltrán J et al (2017) A non-canonical mismatch repair pathway in prokaryotes. Nat Commun 8:14246. https://doi.org/10.1038/ncomms14246
Chang HHY, Pannunzio NR, Adachi N, Lieber MR (2017) Non-homologous DNA end joining and alternative pathways to double-strand break repair. Nat Rev Mol Cell Biol 18:495–506. https://doi.org/10.1038/nrm.2017.48
Eddy SR (2011) Accelerated Profile HMM Searches. PLOS Comput Biol 7:e1002195. https://doi.org/10.1371/journal.pcbi.1002195
Essen LO, Klar T (2006) Light-driven DNA repair by photolyases. Cell Mol Life Sci CMLS 63:1266–1277. https://doi.org/10.1007/s00018-005-5447-y
Hofstatter PG, Tice AK, Kang S et al (2016) Evolution of bacterial recombinase A (recA) in eukaryotes explained by addition of genomic data of key microbial lineages. Proc Biol Sci 283:20161453. https://doi.org/10.1098/rspb.2016.1453
Hollingsworth NM, Ponte L, Halsey C (1995) MSH5, a novel MutS homolog, facilitates meiotic reciprocal recombination between homologs in Saccharomyces cerevisiae but not mismatch repair. Genes Dev 9:1728–1739. https://doi.org/10.1101/gad.9.14.1728
Jackson SP, Bartek J (2009) The DNA-damage response in human biology and disease. Nature 461:1071–1078. https://doi.org/10.1038/nature08467
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Krokan HE, Bjørås M (2013) Base excision repair. Cold Spring Harb Perspect Biol 5:a012583. https://doi.org/10.1101/cshperspect.a012583
Kwapisz M, Beckouët F, Thuriaux P (2008) Early evolution of eukaryotic DNA-dependent RNA polymerases. Trends Genet 24:211–215. https://doi.org/10.1016/j.tig.2008.02.002
Li G-M (2008) Mechanisms and functions of DNA mismatch repair. Cell Res 18:85–98. https://doi.org/10.1038/cr.2007.115
Lin Z, Kong H, Nei M, Ma H (2006) Origins and evolution of the recA/RAD51 gene family: evidence for ancient gene duplication and endosymbiotic gene transfer. Proc Natl Acad Sci USA 103:10328–10333. https://doi.org/10.1073/pnas.0604232103
Lin Z, Nei M, Ma H (2007) The origins and early evolution of DNA mismatch repair genes–multiple horizontal gene transfers and co-evolution. Nucleic Acids Res 35:7591–7603. https://doi.org/10.1093/nar/gkm921
Lindahl T (1993) Instability and decay of the primary structure of DNA. Nature 362:709–715. https://doi.org/10.1038/362709a0
Makarova KS, Krupovic M, Koonin EV (2014) Evolution of replicative DNA polymerases in archaea and their contributions to the eukaryotic replication machinery. Front Microbiol 5:354. https://doi.org/10.3389/fmicb.2014.00354
Malik S-B, Ramesh MA, Hulstrand AM, Logsdon JM (2007) Protist homologs of the meiotic Spo11 gene and topoisomerase VI reveal an evolutionary history of gene duplication and lineage-specific loss. Mol Biol Evol 24:2827–2841. https://doi.org/10.1093/molbev/msm217
Modrich P, Lahue R (1996) Mismatch repair in replication fidelity, genetic recombination, and cancer biology. Annu Rev Biochem 65:101–133. https://doi.org/10.1146/annurev.bi.65.070196.000533
Neupert W, Herrmann JM (2007) Translocation of proteins into mitochondria. Annu Rev Biochem 76:723–749. https://doi.org/10.1146/annurev.biochem.76.052705.163409
Nguyen L-T, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Rivera MC, Jain R, Moore JE, Lake JA (1998) Genomic evidence for two functionally distinct gene classes. Proc Natl Acad Sci USA 95:6239–6244. https://doi.org/10.1073/pnas.95.11.6239
Sancar A, Lindsey-Boltz LA, Unsal-Kaçmaz K, Linn S (2004) Molecular mechanisms of mammalian DNA repair and the DNA damage checkpoints. Annu Rev Biochem 73:39–85. https://doi.org/10.1146/annurev.biochem.73.011303.073723
Schärer OD (2013) Nucleotide excision repair in eukaryotes. Cold Spring Harb Perspect Biol 5:a012609. https://doi.org/10.1101/cshperspect.a012609
Seitz EM, Brockman JP, Sandler SJ et al (1998) RadA protein is an archaeal RecA protein homolog that catalyzes DNA strand exchange. Genes Dev 12:1248–1253
Smith J, Modrich P (1996) Mutation detection with MutH, MutL, and MutS mismatch repair proteins. Proc Natl Acad Sci USA 93:4374–4379
Takemoto N, Numata I, Su’etsugu M, Miyoshi-Akiyama T (2018) Bacterial EndoMS/NucS acts as a clamp-mediated mismatch endonuclease to prevent asymmetric accumulation of replication errors. Nucleic Acids Res 46:6152–6165. https://doi.org/10.1093/nar/gky481
Wiedemann N, Frazier AE, Pfanner N (2004) The protein import machinery of mitochondria. J Biol Chem 279:14473–14476. https://doi.org/10.1074/jbc.R400003200
Wood RD, Shivji MK (1997) Which DNA polymerases are used for DNA-repair in eukaryotes? Carcinogenesis 18:605–610. https://doi.org/10.1093/carcin/18.4.605
Zaremba-Niedzwiedzka K, Caceres EF, Saw JH et al (2017) Asgard archaea illuminate the origin of eukaryotic cellular complexity. Nature 541:353–358. https://doi.org/10.1038/nature21031
Acknowledgements
This study was partially developed at Dalhousie University, and we are grateful to Dayana E. Salas-Leiva and Andrew J. Roger from CGEB. In addition, we would like to thank Gokilavani Thangavel from MPIPZ for further support.
Funding
Open Access funding enabled and organized by Projekt DEAL.. Open Access funding enabled and organized by Projekt DEAL.. This study was supported by the Fundação de Amparo à Pesquisa do Estado de São Paulo (Fapesp), Project Number: 2017/04391-5.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Arturo Becerra.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hofstatter, P.G., Lahr, D.J.G. Complex Evolution of the Mismatch Repair System in Eukaryotes is Illuminated by Novel Archaeal Genomes. J Mol Evol 89, 12–18 (2021). https://doi.org/10.1007/s00239-020-09979-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-020-09979-5