Abstract
Sphagnum mosses are keystone plant species in the peatland ecosystems that play a crucial role in the formation of peat, which shelters a broad diversity of endophytic bacteria with important ecological functions. In particular, methanotrophic and nitrogen-fixing endophytic bacteria benefit Sphagnum moss hosts by providing both carbon and nitrogen. However, the composition and abundance of endophytic bacteria from different species of Sphagnum moss in peatlands of different nutrient statuses and their drivers remain unclear. This study used 16S rRNA gene amplicon sequencing to examine endophytic bacterial communities in Sphagnum mosses and measured the activity of methanotrophic microbial by the 13C-CH4 oxidation rate. According to the results, the endophytic bacterial community structure varied among Sphagnum moss species and Sphagnum capillifolium had the highest endophytic bacterial alpha diversity. Moreover, chlorophyll, phenol oxidase, carbon contents, and water retention capacity strongly shaped the communities of endophytic bacteria. Finally, Sphagnum palustre in Hani (SP) had a higher methane oxidation rate than S. palustre in Taishanmiao. This result is associated with the higher average relative abundance of Methyloferula an obligate methanotroph in SP. In summary, this work highlights the effects of Sphagnum moss characteristics on the endophytic bacteriome. The endophytic bacteriome is important for Sphagnum moss productivity, as well as for carbon and nitrogen cycles in Sphagnum moss peatlands.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Peatlands is a unique type of wetland ecosystem that occupies approximately 3% of the land area and stores about 600 Pg carbon, identifying it as an important carbon pool [1, 2]. However, global warming and changes of precipitation patterns have substantially undermined peatland stability [3,4,5]. For example, the speed of peat soil decomposition has accelerated [6] and emissions of methane (CH4) and carbon dioxide (CO2) have increased [7]. It is well known Sphagnum mosses are the “engineers” of peatlands; they largely increase long term carbon sequestration and greatly accelerate biogeochemical cycling. As a dominant bryophyte, Sphagnum mosses influences the surrounding environments as well as provides primary productivity to peatland ecosystems [8,9,10,11,12].
Sphagnum mosses are among the oldest nonvascular terrestrial plants and have adapted to highly acidic and nutrient deprived habitats that are often waterlogged and anoxic. By releasing H+ to acidify their surroundings or by generating phenolics and releasing them into the environment, Sphagnum mosses create an unfavorable environment for their competitors [11, 13]. These biochemical compounds can suppress the activity of extracellular enzymes of microorganisms and thus prevent the Sphagnum moss from being degraded [14], resulting in consistent accumulation of Sphagnum moss residues. Moreover, Sphagnum moss can activate aluminum and iron oxides in soils, thus facilitating the accumulation of mineral-associated organic carbon which is more stable in carbon poor environments [15]. Sphagnum mosses not only shape favorable microhabitats for their own growth and to maintain primary productivity [12], but also the Sphagnum-associated microbiome plays a critical role in the acquisition of nutrients and the protection against pathogenic bacteria [16, 17]. Water-filled hyaline cells are a special cell type accounting for ~90% of the volume of Sphagnum moss. Hyaline cells are essential for Sphagnum moss, as they provide the space for interactions between plant and the microbial community [18].
Sphagnum mosses possess a diverse microbiome including endophytic and epiphytic microorganisms [19, 20]. Especially, endophytes inhabiting hyaline cells are crucial to moss development and ecosystem function [21]. For example, Sphagnum-associated methanotrophic bacteria in hyaline cells can oxidize CH4 and provide additional carbon (C) to green cells engaged in photosynthesis [22,23,24]. In addition, N2-fixing prokaryotes fix nitrogen (N) from the atmosphere which is supplied to the plant host to compensate for the nitrogen deprived ecosystem [25,26,27,28]. Therefore, it is vital to better understand the relationship between Sphagnum mosses and microbiomes, especially endophytes that are not only involved in nutrient acquisition, but also those that impact C and N cycling in Sphagnum-dominated peatland ecosystems.
Several studies have shown that ecological factors are the main drivers that control the structure and function of the microbiome, such as the effectiveness of nutrients, pH, and temperature; these factors have a distinct association with the community composition of the microbiome [20, 29,30,31]. Moreover, a recent study reported that Sphagnum moss metabolites are an important predictor of the microbial community [32]. Physiological and biochemical properties of Sphagnum mosses can also shape the microbiome. However, many studies regarded both the endophytes and epiphytic microorganisms of Sphagnum mosses as a whole, and little is known about the specific role of endophytic bacteria. This study focused on the endophytic bacteria of different species of Sphagnum mosses, and clarified the microbiome community composition in detail. A previous study reported that plant species drive microbial communities [33], and the genotype of Sphagnum mosses may result in different structures and functions of endophytic bacteria. Moreover, endophytic bacteria are influenced by soil nutrient loading, temperature, and precipitation [34, 35]. Feather mosses are a dominant order in boreal forests and have an important nitrogen fixation capacity; however, moss-associated bacteria have been shown to vary between two dominant species, shaped by climatic, environmental, and nutritional factors [36]. However, it still remains whether endophytic microbial communities are affected by different types of peatlands, soil nutrients, or moss species, and which factors are crucial. To better understand the key factors affecting endophytic bacterial communities and the mechanism of how the endophytic bacteriome drives carbon cycling, an experiment was carried out involving two different typical peatlands. The dominant endophytic bacterial communities of Sphagnum mosses were analyzed, and the influences of Sphagnum moss physiological and biochemical factors as well as soil factors on these bacterial communities were assessed. Further, metagenomic metabolic functions and the C/N cycling the endophytic bacterial community is involved in were predicted.
A previous study suggested that the CH4 oxidizing ability varied between different species of Sphagnum mosses, although the inter-species influence was found to be small compared to that of habitats [37]. Furthermore, all Sphagnum moss species were shown to have the ability to support methanotrophic bacteria, but the methanotrophic activity was influenced by the water level [38]. However, these studies shared the limitation that sampling areas were located in the same climate and environmental zones. Therefore, one of the species in this study, Sphagnum palustre, was sampled in both ombrotrophic peatlands and poor minerotrophic fens. This experimental design was used to identify whether the same species of Sphagnum moss growing in different types of peatlands possesses the same CH4 oxidation activity, which was determined by activity assay. This approach better unlocks an understanding of how peatland types govern methane-oxidizing bacteria communities and their activity. The objectives of this study were to (1) examine the composition and structure Sphagnum moss endophytic bacteriome among different peatland types and species; (2) assess the influence of plant (Sphagnum moss physiological and biochemical) characteristics on these bacterial communities; (3) examine CH4 oxidation activity of the same species of Sphagnum moss between different peatland types (those sampled from Hani (SP) and those sampled from Taishanmiao (HSP)). In addition, PICRUST and FAPROTAX were used to predict the function of endophytic bacteria based on 16S rRNA genes.
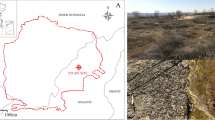
To reach these objectives, five Sphagnum moss species were collected: S. palustre, S. magellanicum, S. fuscum, and S. capillifolium were sampled from the Hani peatland, which is a temperate poor fen in the Changbai mountains, northeast China; S. palustre was sampled from Taishanmiao, which is a subtropical bog in southwest Hubei province, central China (Fig. 1). The endophytic microbial community was analyzed based on 16S rRNA marker genes, utilizing their metabolic and C/N element cycle functions. In addition, the incorporation of 13C-CH4 was measured to assess the activity of methanotrophic bacteria in S. palustre.
Materials and Methods
Study Sites
The Sphagnum moss samples were collected from two different types of peatlands (Fig. 1). Hani peatland is a poor minerotrophic peatland (i.e., a poor fen, where precipitation and underground water are the main sources of nutrients), located in the Changbai Mountains region of northeast China (42°13′05″N, 126°31′05″E, 900 m above sea level). Hani peatland is situated in the continental monsoon climate with a mean annual temperature of 3.9 °C and mean annual precipitation of 780 mm, respectively. The vegetation of Hani peatland includes shrubs, herbs, graminoids, and Sphagnum mosses. The dominant shrub is Betula fruticose Pall. var. ruprechtiana Trautv and herbs include Eriophorum polystachion L., Carex lasiocarpa Ehrh., and Smilacina japonica A. Gray [39]. Taishanmiao is an ombrotrophic peatland (i.e., a bog, where precipitation is the main source of nutrients) located in western Hubei province of central China (30°7′44″N, 109°47′12″E, 1800–1920 m above sea level), located in the subtropical subalpine region with a mean annual temperature of 7.20–8.27 °C, and a mean annual precipitation of 1768 mm. At Taishanmiao, vegetation types are mainly divided into shrubs, herbs, and mosses. Enkianthus serrulatus, Rhododendron auriculatum, Carex taliensis, and Juncus effusus are dominant shrub and herb species [40].
Sample Collection and Processing
Because of the different development periods and environmental conditions, the dominant Sphagnum moss species in the two peatlands differed. S. palustre, S. magellanicum, S. fuscum, and S. capillifolium dominate Hani peatland [41], while Taishanmiao is dominated by S. palustre only. The habitats of these Sphagnum mosses are located in hummocks or midway between hummocks and hollows.
Sphagnum moss samples were collected into sterile zip lock bags and were transported to the laboratory with ice bag. Sphagnum moss samples were collected in triplicate. Upon arrival at laboratory, every sample was divided into three parts. In one part, the green parts were cut into pieces of 3–5 cm, the surface of which was sterilized for DNA extraction and sequencing; the other two parts were used to determine physicochemical indexes and stable isotope labeling activity assays. In addition, each kind of Sphagnum moss was incubated in a culture tray in an artificial climate chamber.
Peat soil samples (0–20 cm) were collected below the surface of growing Sphagnum mosses as the background value in two peatlands. The specific approach was to remove the vegetation cover and take peat cores by drilling; then, 0–20m peat soil cores were sliced and put into sterile bags. Peat soils samples were collected in three replicates in both Hani and Taishanmiao. Each sample was homogenized and stored at 4 °C until determination of physicochemical characteristics.
To extrac the DNA of Sphagnum moss endophytes, the samples were processed according to the following steps: First, the Sphagnum moss sample surface was slowly washed with tab water. Then, samples were placed into a beaker of 75% alcohol for 1 min and rinsed in sterile deionized water 5–6 times. Next, samples were immersed in 1% NaClO for 1 min and rinsed in sterile deionized water 7–8 times. Finally, using sterilized filter paper, the surface moisture of samples was dried. The final sterile water that had been used to rinse the samples was used to inoculate Rudolph culture medium. Sphagnum moss materials were frozen at −80 °C until DNA extraction [42, 43].
Water-Holding Capacity
The water-holding capacity of Sphagnum mosses was characterized by measuring the water absorption until saturation. Three 2 cm × 2 cm quadrats of each species were placed into culture trays, and the green parts of the Sphagnum mosses capitulum were cut (approximately 3 cm). Samples were placed into a 50-ml centrifuge tube with 20 ml deionized water to adequately absorb water. Then, samples were taken out upside down until water dripping ceased, and the sample weight (W1) was recorded. Then, samples were placed in a drying oven at 72 °C and were dried to a constant weight 48 h and record dry weight (W2). Water-holding capacity = (W1 − W2)/W2 × 100%. The method has been described before [44] and adjustments were minor.
Physiological and Biochemical Index
Carbon and Nitrogen Contents
Sphagnum moss samples (green segments of collected Sphagnum mosses) were dried in an oven at 105 °C for 30 min. Then, the temperature was adjusted to 72 °C, drying was continued for 48 h, and dried samples were ground with a ball mill. The dried and ground Sphagnum moss samples were used for subsequent analyses. C and N contents were determined using an elemental analyzer (CNS analyzer, EA 1110 Carlo Erba, Thermo Fisher Scientific, Waltham, MA, USA).
Chlorophyll Content, Total Phenols, Total Carbohydrates, Malondialdehyde, and Proline
Chlorophyll content was determined by the spectrophotometric method [45]. Briefly, fresh green capitulum of Sphagnum mosses (0.1 g) was put into a sterilized centrifuge tube with 80% (v/v) acetone solution (5 ml) to extract chlorophyll. The tube was wrapped with foil to protect it from light until the sample was extracted completely, followed by centrifugation for 10 min at 4 °C and 8000 rpm; then, the supernatant was absorbed. The supernatant (200 µl) was transferred into a 96-well plate to measure the absorbance value at 645 nm and 663 nm with a microplate reader (Thermo Scientific™ Varioskan™ LUX, Thermo Fisher Scientific, Waltham, MA, USA). Chlorophyll content was expressed in milligram of chlorophyll per a fresh weight (mg/g FW).
The total phenol content was determined by the Folin-Ciocalteu method [46]. First, dried Sphagnum moss sample (0.10 g) was extracted by 60% ethanol (2.5 ml) for 30 min and ultrasonicated for 30 min at 60 °C, followed by centrifugation for 10 min at 4 °C and 8000 rpm to obtain the supernatant. Then, Folin-Ciocalteu (250 µl) and 20% Na2CO3 (250 µl) were added to the supernatant (50 µl) of each sample. After incubation for 10 min, the absorbance was determined at 760 nm. Gallic acid was used as standard to calculate the total phenol content.
The total carbohydrates content was determined using the phenol-sulfuric acid method with minor modifications [47]. Fresh frozen Sphagnum moss samples (40 mg) were ground in liquid N; then, 50% methanol solution (4 ml) was added and the mixture was ultrasonicated for 40 min at 20 °C and 8000 rpm to obtain extracts. These extracts were shaken in an orbital shaker for 18 h at room temperature and 150 rpm, after which, the previous ultrasonication extraction step was repeated, followed by centrifugation for 5 min at 4 °C and 8000 rpm and absorption of the supernatant (final extract). Absorbed extracts (20 µl) were put into an Eppendorf tube to which distilled water (200 ml) was added for dilution, then 5% phenol solution (200 µl) and 98% H2SO4 (1 ml) were added. After incubation for 30 min and absorbance was determined at 490 nm. Glucose was prepared to draw the standard curve.
Malondialdehyde (MDA) is the product of lipid peroxidation in plants and the contents of MDA reflects the degree of lipid peroxidation. MDA was determined by the thiobarbituric acid method [48]. Fresh frozen Sphagnum moss samples (0.1 g) were ground in liquid N; then, 10% trichloroacetic acid (5 ml) was added, and the mixture was centrifugated for 10 min at 4 °C and 8000 rpm. Then, the supernatant (2 ml) was added to thiobarbituric acid (2 ml). After 15 min in the water bath, followed by centrifugation for 10 min at 4 °C and 8000 rpm, the absorbance was determined at 600, 532, and 450 nm.
Proline is a kind of osmoregulatory substance and the acid ninhydrin method was used to measure the proline content [49]. Briefly, the proline was extracted from dried Sphagnum moss samples (0.025 g) by 3% sulfosalicylic acid (1 ml), followed by centrifugation for 10 min at 4 °C and 4500 rpm to obtain the extracts. The supernatant (700 µl) was added to 5-ml glass tubes, and then the acid ninhydrin (2 ml) was added into the tube. In a water bath, samples were shaken gently to achieve uniform mixing. Absorbance was determined at 520 nm using l-proline as standard to calculate proline contents.
Enzyme Activity
The oxidative enzyme activities of phenoloxidase (PO O2) (PPO) and peroxidase (PO H2O2) (PER) were quantified [50, 51], using O2 and H2O2 as acceptors, respectively. First, enzymes were extracted: Green parts of fresh Sphagnum moss samples (3 g) were submerged in CaCl2 (50 ml, 0.10 M) with Tween 80 (0.05%) and polyvinylpolypyrrolidone (20 g) and the mixture was shaken for 1 h at room temperature. After centrifugation for 10 min at 4 °C and 10000 rpm, the supernatant was filtrated (1.2 µm, Waterman GF/D filters). Then, the enzyme activity was quantified. For PPO, concentrated extracts (150 µl) with 2,7-diaminofluorene (2 µl) in 96 wells microtiter plate. For PER, in addition to the above, 0.3% H2O2 (10 µl) was added to the rection system. PPO and PER were monitored at 600 nm by a microplate reader (Thermo Scientific™ Varioskan™ LUX, Thermo Fisher Scientific, Waltham, MA, USA). The enzyme activity was expressed as 1 nmol of substrate oxidized per min per mg of dry mass. In addition, another method (guaiacol method) was also used to determine peroxidase, expressed in POD [52]. Fresh sample (0.5 g) was added to 5 ml phosphate butter (pH = 5.6, containing 1% polyvinylpolypyrrolidone and 0.1% mercaptoethanol), followed by grinding in an ice bath. After centrifugation for 10 min at 4 °C and 10000 rpm, enzymes extracted with phosphate butter, 2% H2O2 and 2% guaiacol were determined by a spectrophotometer (FC-1100, Thermo Fisher Scientific, Waltham, MA, USA) at 470 nm. The absorbance was recorded every minute for five times in total. The enzyme activity was obtained by measuring the change in absorbance.
Glutamine synthetase activity was determined by the appropriate enzyme activity detection kit (Qiyi Biological Technology, Shanghai, China).
Soil Physicochemical Analysis
After soil samples were processed including air-drying, grinding, and passing through a 2-mm sieve, total carbon (TC) and total nitrogen (TN) concentrations were determined by elemental analyzer (CNS analyzer, EA 1110 Carlo Erba, Thermo Fisher Scientific, Waltham, MA, USA). Before determining the soil total phosphorus (TP) of samples by elemental analyzer, the nitric acid and perchloric acid digestion method was used first to extract phosphorus from the sample to obtain an extraction solution. The dissolved organic carbon (DOC) concentration was determined by a total organic carbon analyzer (LIOYIL TOCII, Elementar, Germany). The peat soil water content was determined by gravimetric method. The pH value of the peat soil sample was measured by an acidity meter (OxyScan 300, UMS GmbH & Co. KG, Germany). Peat soil physiochemical properties are showed in Table S1.
CH 4 Oxidation Activity Assay
To estimate the CH4 oxidation activity of Sphagnum-associated methane microbes in SP an HSP, the incorporation of the stable isotope (13C-CH4) in Sphagnum moss samples was measured [53]. Briefly, all test samples were in consistent vigorous growth condition (3-cm length of Sphagnum mosses capitulum). Into sterilized glass bottles (100 ml) with airtight plugs, three plantlets (of which the fresh weight was determined) were added and each species had three replicates. To all bottles, 5 ml (0.05%) 13C-CH4 was added to the headspace and controls were incubated without any labelled gassed. All samples were incubated in an artificial climate chamber for 48 h at 24 °C, including light/dark treatment (16 h with light and 8 h without light) and dark treatment (without light for 48 h), respectively. After this incubating step, samples were placed in oven at 72 °C for 48 h until a constant weight was reached (the dry weight) and ground by a ball mill (Retsch: MM 400). Then, the dried mass of each sample was weighted (approximately 4 mg) and put into tin cups. An elemental analyzer (CNS analyzer, Thermo Fisher Scientific) coupled to an isotopic ratio mass spectrometer (Finnigan Delta Plus, Thermo Fisher Scientific (Bremen) GmbH, Germany) was used to determine the fraction of 13C that had been incorporated into each sample. The activity of CH4 oxidation was expressed in nmol CH4 g−1 DW d−1.
DNA Extraction, Amplification, and 16S rRNA Sequencing
Sterilized frozen samples were used for DNA extraction. The total genomic DNA of endophytic bacteria from Sphagnum moss samples was extracted using the OMEGA DNA Kit (M5635-02) (Omega Bio-Tek, Norcross, GA, USA) according to the manufacturer’s instructions. Using a NanoDrop NC200 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA), the quantity and quality of samples were measured and 0.8% agarose gel electrophoresis was used to measure the integrity of the extracted DNA. The V5 and V7 regions of the microbial 16S rRNA gene were amplified using 799F (5′-AACMGGATTAGATACCCKG-3′)/1193R (5′-ACGTCATCCCCACCTTCC-3′) [54, 55]. PCR amplicons were purified with Vazyme Vahtstm DNA Clean Beads and quantified the by the Quant-iT PicoGreen dsDNA Assay Kit (Invitrogen, Carlsbad, CA, USA). Amplicons were pooled in equal amounts and pair-end 2250 bp sequences were used, obtained by Illumina NovaSeq platform with NovaSeq 6000 SP Reagent Kit (500 cycles) at Shanghai Personal Biotechbology Co., Ltd (Shanghai, China).
Bioinformatics Analysis
The raw data were stored in FASTQ format and reads were performed with QIIME2 (2019.4) [56]. Raw paired-end reads primers and chimera were removed by the DADA2 plugin [57]. Quality filtering, denoising, and merging were also performed using DADA2. The sequence was processed by the above steps and dereplication. Using the cluster size module in Vsearch (v2.13.4_linux_x86_64) [58], the sequences with ≥ 97% similarity were clustered. The amplicon sequenced variants (ASVs) were aligned by MAFFT [59] and further used for annotation. Prokaryotic taxonomy was assigned with QIIME (2019.4), utilizing the Greengens database [60] (http://greengenes.secondgenome.com/) and Silva database [61] (http://www.arb-silva.de). ASV matrices were rarefied according to the sample with the least reads and depth was set to 95% of the lowest sequencing. All subsequent analyses were based on these rarefied data.
The Phylogenetic Investigation of Communities by Reconstruction of Unobserved States (PICRUSt2) [62] was used to predict the metabolic functions of marker gene sequence abundance in the samples based on the Kyoto Encyclopedia of Genes and Genomes (KEGG) database [63] (http://www.genome.jp/kegg/pathway.html). The database of Functional Annotation of Prokaryotic Taxa (FAPROTAX) [64] was also used to predict the function of prokaryote in Sphagnum mosses, with a specific focus on the processes of C and N elements cycling the members of the endophytic bacterial community are involved in. All original 16S rRNA sequences data were uploaded to the Sequence Read Archive of the National Center for Biotechnology Information (NCBI) database (accession number: PRJNA1006557).
Statistical Analysis
Differences between Sphagnum moss endophytic microbiome (such as richness, Shannon’s diversity index, and Pileou’s evenness) were calculated in R with the vegan package. Non-metric multidimensional scaling (NMDS) with Bray-Curtis dissimilarity was used to identify the dissimilarities in endophytic bacterial communities. An analysis of similarities (ANOSIM) was conducted to examine the clustering of bacterial communities. Permutational multivariate analysis of variance model (PERMANOVA) was used to examine species-level differentiation in communities of endophytic bacteria. Canonical correspondence analysis (CCA) was performed to study the relationship between the relative abundance of the dominant endophytic bacterial communities as well as soil and plant variables (see Tables 1 and S1). Mantel test was used to examine the relationship between plant factors and the average relative abundance of the top 10 phylum, family, and genus endophytic bacteria communities. LEfSe analysis was used to identify biomarkers of endophytic microbial community in different Sphagnum mosses. The above analysis was carried out in R (4.1.3) using phyloseq, DESeq2, ggcor, geosphere, tidyverse, microbiomeViz, ggtree, phyloseq, and vegan packages, and figures were drawn using the ggplot2 package.
Spearman’s correlation test was calculated by SPSS 18.0. Soil and plant parameters were normality tested before analysis. Differences in physiological characteristics between different species of Sphagnum mosses were evaluated using one-way ANOVA and multiple comparisons in SPSS 18.0. The results of predicting prokaryote C/N element cycling and metabolic pathway function were tested with the Kruskal-Wallis test.
Results
Physiological and Biochemical Characteristics of Sphagnum Mosses
The physiological and biochemical characteristics of Sphagnum mosses showed significant differences among Sphagnum mosses, especially the contents of C, N and total carbohydrate (Table 1). C and N in Sphagnum palustre sampled from Taishanmiao (HSP) were 43.10% and 1.86%, respectively, and were the highest among these samples (P < 0.05, Table 1); while C and N in Sphanum palustre sampled from Hani (SP) were 23% and 30%, respectively (Table 1). The chlorophyll content of Sphagnum fuscum (SF) was 4.23 mg/g, which was significantly higher compared to other species of Sphagnum mosses (P < 0.05, Table 1). Sphagnum capillifolium (SC) had the highest concentration of total carbohydrates (134.80 mg/g) while HSP had the lowest concentration (63.35 mg/g) (Table 1).
Sphagnum species from Hani had similar water-holding capacities except for SF (29.44%) (P < 0.05, Table 1) and SP (58.44%) which had higher capacities than HSP (33.35%). These results showed that the contents of proline and MDA did not vary significantly between species. The activity of POD had clear differences among different moss species, where the highest was SM with 4.11 mmol·min−1·mg−1, followed by SF (3.38 mmol·min−1·mg−1), and SC was the lowest (0.33 mmol·min−1·mg−1) (Table 1).
Endophytic Bacterial Community Structure and Composition in Different Sphagnum Species
In total, 33,481 ASVs were obtained in all samples after quality control and rarefaction. The rarefaction curves across all samples almost reached stable values (Fig. S1), indicating that most of the endophytic bacteria in Sphagnum moss samples were captured.
Based on ASVs, alpha diversity indexes of endophytic bacteria were estimated using Shannon, Chao1 (Fig. 2a, b) and Pielous evenness indexes (Table S2). Alpha diversity was the highest in SC, followed by SF and SP had the lowest alpha diversity (P < 0.05, Fig. 2 and Table S2). Between HSP and SP, Chao1 and Shannon diversity indexes were significantly different (P < 0.05, Fig. 2a, b).
Endophytic bacterial communities in Sphagnum moss. Alpha diversity of 16S rRNA gene in different species Sphagnum mosses including Chao1 index (a) and Shannon index (b). Non metric multidimensional scaling (NMDS) ordinations of prokaryotic ASV compositions among different Sphagnum moss species (c). Different color points represent different species of Sphagnum mosses. Significant differences (p < 0.05) are indicated with lowercase letters. SM, Sphagnum magellanicum; SF, Sphagnum fuscum; SC, Sphagnum capillifolium; HSP, Sphagnum palustre (sampled from Taishanmiao); SP, Sphagnum palustre (sampled from Hani)
NMDS analysis showed the dissimilarities in endophytic bacterial communities based on the Bray-Curtis matrix. Different Sphagnum moss species were distinguished by differences in their prokaryotic ASV composition (Fig. 2c), and communities of the same Sphagnum moss species clustered together. Notably, the endophytic communities of HSP were separated from the other four groups on the first axis, and the endophytic communities of SP were completely separated from the other three species from Hani on the second axis (Fig. 2c). ANOSIM test showed that the endophytic communities differed significantly between Sphagnum moss species (R = 0.9481, P = 0.001). PERMANOVA was further used to examine species-level differentiation in communities of endophytic bacteria (F = 3.832, P < 0.001).
To clarify the composition of the endophytic bacteria community among the sampled Sphagnum mosses, the relative abundances of the top 10 bacterial phyla, families, and genera were analyzed. In total, 36 phyla were identified from all sequences and the most abundant top 10 phyla are depicted in Fig. 3a. The endophytic bacterial community in Sphagnum mosses was mainly composed of Proreobacteria, Actinobacteria, Acidobacteria, Firmictes, and Bacteroidetes; however, the proportions differed (Fig. 3a and Table S3). Proteobacteria was more abundant in SP than in other mosses, accounting for 86.00% of the total relative abundance, while Bacteroidetes and Firmicutes were the lowest among all moss samples (Fig. 3a and Table S3). The average relative abundance of Acidobacteria in HSP was 0.40% lower than in other species, while Chlamydiae were significantly more abundant in HSP compared to other species (P < 0.05, Table S3). At the family level, the average relative abundance of Acetobacteraceae was higher in SP (P < 0.05, Fig. 3b and Table S3). Conversely, Rhizobiaceae were significantly higher in HSP (10.05%) than in other Sphagnum mosses (P < 0.05, Fig. 3b and Table S3). The average relative abundance of Beijerinckiaceae was 5.10% in SP, which is significantly higher compared to HSP (P < 0.05, Fig. 3b and Table S3). Moreover, the endophytic microbiome between SP and HSP had distinct genera (Fig. S2); an example can be found in the average relative abundances of the Acidocella and Bacteroides (P < 0.05, Table S3).
Sphagnum moss endophytic bacterial community composition. Average relative abundance of the top 10 phylum (a) and family (b) in different species Sphagnum mosses. Taxonomic cladogram (c) through linear discriminant analyzes effect size (LEfSe) and biomarker of endophytic bacterial communities in different species Sphagnum mosses with LDA SCORE > 3 (d). Significant discriminant taxa of SP (Sphagnum palustre was sampled from Hani), SM (Sphagnum magellanicum), SF (Sphagnum fuscum), SC (Sphagnum capillifolium), and HSP (Sphagnum palustre was sampled from Taishanmiao) are colored in brown, pink, purple, orange, and blue, separately. Colorless nodes represent taxa that do not significantly discriminate among Sphagnum mosses. The dimension of nodes is positively correlated with the relative abundance of the taxon
To identify biomarkers of the endophytic microbial community in different Sphagnum moss, LEfSe analysis was performed (Fig. 3c, d). According to the results of LDA score (LDA > 3, P < 0.01, Fig. 3d), SF had five discriminative biomarkers from phylum to genus, which were affiliated with phylum Firmicutes, class Bacilli, order Lactobacillales, family Lactobacillaceae, and genus Lactobacillus. In SC samples, two biomarkers were family Beijerinckiaceae and genus Roseiarcus. HSP and SP had different biomarkers; according to the evolutionary clustering analyses diagram, Rhizobiaceae were abundant in the blue parts, representing HSP, while Aquabacterium were most abundant in the brown parts, representing SP (Fig. 3c).
Relationship Between Phytochemical Parameters and Endophytic Bacterial Abundances in Sphagnum Mosses
To better understand the linkages between plant physiological and biochemical characteristics and endophytic bacteria communities, CCA and Mantel test were used to detect important environmental factors that influence the relative abundance of the top 10 phylum, family, and genera of endophytic bacteria. CCA analysis showed that the correlation between chlorophyll contents, water-holding capacity, and PPO activity and endophytic bacteria communities at the phylum level (R2 = 0.7231, Fig. 4a). Similarly, chlorophyll contents and water-holding capacity also shaped the relative abundance of the top 10 family and genera endophytic bacteria in Sphagnum mosses (Fig. S3, Table S5and Table S6). Significant Spearman’s correlations were found between the relative abundance of Proteobacteria and water-holding capacity, while Firmicutes were negatively correlated with water retention capacity (P < 0.05, Table S4). Chlorophyll contents were identified as a crucial factor affecting the endophytic bacterial communities at the family level. The relative abundance of Acetobacteraceae and Muribaculaceae were significantly positively correlated with chlorophyll contents (P < 0.05, Table S5).
Correlation between plant phytochemical parameters and bacterial communities. Canonical correspondence analysis of the relative abundance of the top 10 phyla endophytic bacteria in association Sphagnum moss characteristics (a) and correlation between Sphagnum moss physiological and biochemical characteristics and the relative abundance of the top 10 phyla, families, and genera of endophytic bacteria (b). C, carbon contents; N, nitrogen contents; Chl, chlorophyll contents; Tp, total phenol contents; Tc, total carbohydrates; Pro, proline; MDA, malondialdehyde; Wr, water retention capacity; PPO, phenol oxidase (PO O2); PER, peroxidase (PO H2O2); POD, peroxidase; GS, glutamine synthetase
Moreover, the Mantel test also presented correlations between plant parameters and the microbial community (Fig. 4b). Significant correlation were found between the relative abundance of top the 10 genera endophytic bacteria and C contents (Mantel’s r = 0.303, P = 0.017), and the relationships with chlorophyll content (Mantel’s r = 0.459, P = 0.006); further, the chlorophyll content correlated significantly with the top 10 families of endophytic bacteria communities (Mantel’s r = 0.492, P = 0.003), and correlated significantly with the relative abundance of the top 10 phylum endophytic bacteria and PPO (Mantel’s r = 0.448, P = 0.01).
Endophytic Bacterial Communities Enrich Functional Roles Related to C Cycling
Based on 16S rRNA gene taxonomic analysis, the functions of putative methanotrophic prokaryotic taxa are discussed. Methanotrophic communities at the family or genus level varied between different Sphagnum moss species (Fig. 5a and Table S7). The relative abundance of Beijerinckiaceae family comprised 5.10% of the total endophytic bacteria communities in SP; at the genus level, SP had the highest relative abundance of Methyloferula (family Beijerinckiaceae). The relative abundance of Methyloferula in Sphagnum mosses from Hani was higher than in mosses from Taishanmiao. However, the genus of Methylobacteriu accounted for 1.24% in HSP, which was the highest among all Sphagnum mosses. Moreover, methanotrophic communities showed differences between HSP and SP, as depicted in Fig. 5b.
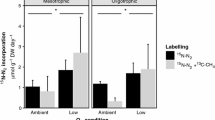
Comparison of the relative abundances of methanotrophic prokaryotic taxa functions. The relative abundances of methanotrophic in five Sphagnum mosses (a), Sphagnum palustre was sampled from Hani (SP) and Taishanmiao (HSP), respectively (b). Methane oxidation activity (represente by 13C-CH4 incorporation rate) of Sphagnum palustre was sampled from Hani (SP) and Taishanmiao (HSP) at light/dark (c) and dark treatments (d), respectively. Light/dark treatment represents samples kept in artificial climate chamber for 16 h with light and 8 h without light at 24 °C. Dark treatment represents samples kept in artificial climate chamber without light for 48 h at 24°C
Because the methanotrophic communities show differences between SP and HSP (see Fig. 5b and Table S9), whether these differences influence the methane oxidation efficiency was examined. An activity experiment was conducted to explore whether there were differences between SP and HSP (Fig. 5c, d). The CH4 oxidation rate of methanotrophic communities in SP (3.0702 ± 0.2554 nmol CH4 g−1 DW d−1) was higher than that of HSP (2.2752 ± 0.8289 nmol CH4 g−1 DW d−1) (Fig. 5c and Table S9). Similarly, SP was also higher than HSP in the dark treatment (Fig. 5d and Table S9). However, no significant difference was found between HSP and SP.
Element Cycling (C and N) and Prediction of Metabolic Pathway Function
Given the key role endophytic microbes play in Sphagnum mosses, combined with functions and activity assays of putative methanotrophic prokaryotic taxa, functional annotation of prokaryotic taxa was performed (FAPROTAX) to analyze the putative function of the Sphagnum microbiome (Fig. 7a). Most of the functional genes showed similar abundances across different Sphagnum species. The average functional abundances of methylotrophy, methanol oxidation, and ureolysis were higher in HSP than in other mosses. The average functional abundance methanotrophy in SP was the highest among all Sphagnum moss samples, followed by SF. Specifically, the number of ASVs of methanotrophy in SP was approximately twice that of HSP (Fig. 6a).
In addition, microbial functions were also involved in N cycling (e.g., nitrogen fixation, nitrification, and nitrate reduction). Nitrogen fixation differed significantly among sampled mosses; the average functional abundance of nitrogen fixation was higher in HSP than in other species (Fig. 7a), and the number of ASVs involved in nitrogen fixation was also higher in HSP (Fig. 6b). Combined with putative diazotrophic communities based on 16S rRNA gene taxonomic analysis (Fig. S4), diazotrophic communities included Magnetospirillaceae, Sphingomonadaceae, Rhizobiaceae, Pseudomonadaceae, and Beijerinckiaceae. The relative abundance of Rhizobiaceae was the highest in HSP among all sampled mosses, and the relative abundance of Sphingomonadaceae within all endophytic bacterium communities was 3.11 ± 0.01% in SP (Table S5). Furthermore, the relative abundance of Beijerinckiaceae (including C and N cycling functional microorganisms) was also the highest in SP. Correlation analysis was conducted between the function predicted by FAPROTAX and the relative abundance of the top 10 family endophytic bacteria (Table S10). These results showed that Rhizobiaceae exhibited a highly positive Spearman’s correlation with the functions of N fixation (P < 0.01), ureolysis (P < 0.01), and methanol oxidation (P < 0.01, Table S10).
Element cycling functional genes and potential metabolic pathways prediction. Heatmap of C and N cycling functions of endophytic bacteria based on FAPROTAX prediction (a). Heatmap of metabolism and genetic information process pathways (KEGG II) of endophytic bacterium between Sphagnum mosses by PICRUSt2 (b). Data were based on number of ASVs of each function in sphagnum moss
PICRUSt2 was used to predict the potential metabolic functions of the endophytic microbial community. The results showed that different species of Sphagnum mosses had similar relative abundances on the KEGG level I pathway (Fig. S5). Metabolism and genetic information process pathways differed significantly between Sphagnum mosses (P < 0.05) (Table S11). Metabolic pathways including 12 pathways in level II, the relative abundance of carbohydrate metabolism, amino acid metabolism, as well as the cofactors and vitamins metabolism were elevated in every species moss (Fig. 7b). Metabolism of terpenoids and polyketides differed significantly between species (P < 0.05, Kruskal-Wallis test, Table S11,) and SC was the highest.
Discussion
Endophytic bacteria have great implications to the growth and development of Sphagnum mosses, and the mosses, in turn, influence the C and N cycles in the peatland ecosystem [11, 65]. This study demonstrated the differences of endophytic bacteria communities in Sphagnum mosses across two sites and various species. These differences have important implications for clarifying the endophytic microbial community structure, thus contributing to a better understanding of the function of endophytic microbes. This knowledge furthers research on how to promote Sphagnum moss growth and its development under climate change, as well as on how to evaluate C and N cycling functions in peatlands. Previous studies reported that plant-associated microbiomes are influenced by both biotic and abiotic factors [25, 66, 67]. Similarly, endophytic bacteria are also influenced by many factors including environmental conditions, plant species or genotypes, and even growth periods [68, 69].
In this study, differences were found among the sampled Sphagnum moss species regarding physiological and biochemical characteristics. S. palustre (HSP and SP) grown in different types of peatlands also showed physiological differences (Table 1). Sphagnum moss species also differed in morphological traits, for example, the shape and size of leaves in the capitate branch, which is a critical water retention characteristic for mosses [18]. Microbiomes are shaped by plant host species, microbial interactions, and environmental factors including latitude, precipitation, and soil nutrients [70,71,72]. These factors also affect the endophytic microbiome to a certain extent. Indeed, the characteristics of the sampled Sphagnum moss species significantly affected the structure of endophytic bacterial communities (Fig. 4). Sphagnum moss gametophytes can acquire specific microbiomes from sporophytes which feature abundant bacterial diversity [19], emphasizing that host specificity and genetic factors are determinants of the moss plant microbiome. The identity of host species influences bacterial gene expression, and it was proposed [73] that feather moss can upregulate certain genes, thus promoting cyanobacterial abundance and growth to a certain degree. Except for species identity, in this study, remarkable correlation between the relative abundance of the top 10 phyla or families of endophytic bacterial communities in Sphagnum moss and water-holding capacity, chlorophyll contents, and the activity of PPO (Fig. 4). These factors vary with Sphagnum moss species, and can further influence endophytic bacterial communities. Chlorophyll, water, and CO2 are indispensable for photosynthesis. On the one hand, abundant bacteria associated with methane oxidation in both the green photosynthetic cells and hyaline cells of Sphagnum mosses provide additional CO2 thus guaranteeing supplementation with sufficient CO2 [24, 74]. On the other hand, sufficient water showed that the Sphagnum moss hyaline cells perform well. Porous hyaline cells not only play a vital function as water storage organs and transport water to adjacent green photosynthetic cell but also provide a habitat for an abundance of microbial communities [18, 19]. Hence, this also reflects the positive correlation between the dominant microbiome at the phylum and family levels with chlorophyll contents. Water-holding capacity was another important factor for shifts in endophytic bacterial communities. The results of Spearman correlation test indicated that there is a positive relationship between water-holding capacity and the family of Burkholderiaceae (Spearman’s ρ = 0.564, P < 0.05). The members of Burkholderiaceae are distributed throughout a variety of habitats including animals, plants, and soil [75], and this community adapted acidic peatlands [76]. Previous studies have shown that in Sphagnum moss, both endophytic and entophytic bacteria, possess a high diversity of the genus of Burkholderia. Burkholderia was dominant in S. magellanicum and S. fuscum [17], and ingredients with antifungal activities were detected in Burkholderia species [77].
Global warming will change the stability and carbon sequestration may suffer from a series of influences. These influences can be partly attributed to the change of the activity of extracellular phenol oxidases which could degrade complex polyphenols and play important roles in the soil carbon cycle in the peatland ecosystem [78]. In this study, the PPO activity did not show differences between the sampled Sphagnum moss species, while the activities of PER between SC and SF were significantly different (Table 1). Previous research examined the activities of PPO and PER in S. fuscum in Sphagnum-dominated peatlands under simulated warming climate; the findings showed that warming treatment also did not alter PPO activity but increased PER activities in the living top segments [79]. Overall, the differences in the phenol oxidases activity between different species of Sphagnum mosses may be related to differences of endophytic bacteria. Activity is also affected by abiotic factors, and phenol oxidase activity was positively related to the moisture content [80].
Among these Sphagnum moss species, HSP and SP belong to the same species but grow in different peatland types and show differences in bacterial composition at the phylum level. For example, the relative abundances of Bacteroidetes (HSP, 16.46%; SP, 0.43%) and Firmicutes (HSP, 9.97%; SP, 0.40%) (Fig. 3a and Supplemental Table S3). The LEfSe results also showed that Rhizobiaceae were abundant in HSP while Aquabacterium were the most abundant in SP (Fig. 3c, d). Prior studies reporting that as long as the same species of moss was sampled in the same ecological amplitude, the moss-associated microbial composition was similar [19, 26]. Moreover, in comparison to other species sampled in Hani (i.e., SC, SF, and SM), SP still showed differences. According to a previous observation, S. fuscum (hummock) and S. angustifolium (broader ecology), which grow in the same bog ecosystem, show a high degree of similarity [17, 20]. These discrepancies can partially be explained as follows: in each of the experiments, the research object differ. More than just endophytes in sampled Sphagnum mosses bacteriome in their study, the diversity of the microbial community on plant surface is generally higher than the diversity of the endophytic bacteria [81].
In this study, active methanotrophic endophytic bacteria in Sphagnum mosses include Methyloferula and Methylobacterium. A previous study identified Methylferula as one of the most active methanotrophs associated with Sphagnum mosses in peatland [82]. The relative abundance of Methyloferula in SP was higher than that of other mosses and the activity assay results further showed that the CH4 oxidation rate was higher in SP than in HSP (Fig. 5c, d). This result suggests that Methyloferula has mainly methanotrophs function in the Sphagnum moss endophytic bacteriome. Beyond Methyloferula, the relative abundance of Beijerinckiaceae was higher in SP than in other species including HSP (Table S7). Beijerinckiaceae comprise obligate methanotrophs, facultative methanotrophs, and facultative methylotrophs. Moreover, in line with previous studies, the higher abundance and diversity diazotrophs could contribute to higher N2-fixation rates, emphasizing that the higher microbiome diversity will promote the microbiome function [36, 83, 84]. At the same time, methanotrophs have been shown to provide C derived from atmospheric CH4 to Sphagnum mosses [38], but the rate of oxidation varied among Sphagnum moss species. However, it has been suggested that CH4-oxidizing bacteria associated with Sphagnum mosses are not influenced by the Sphagnum moss species [85], but rather, are controlled by abiotic factors [38, 86]. In the present study, both HSP and SP were collected from a moderately rich fen in northeast China and a poor nutrient bog located in a subtropical alpine region (Fig. 1), respectively; the results highlight the differences between SP and HSP (Fig. 5c, d). In addition, the rate of CH4 oxidation was higher in the dark than in the light/dark treatment, which differed from previously published results showing that light can stimulate of CH4 oxidation activity [87]. Moreover, physiologic characteristics of Sphagnum moss can influence the number of endophytic CH4-oxidizing bacteria in plant tissue and thus the CH4 oxidation capacity [37]; the reason is that two different species of Sphagnum mosses with physiologic characteristics differences growing at the same site showed different CH4 consumption levels. Microbial communities growing in similar environments were found to have similar functions [88] in Sphagnum mosses; despite, these differences in microbiome, functional redundancy in CH4 oxidation is a widespread mechanism in peatland, the higher rate in SP may be the result of the higher diversity and activity of functional endophytic microbes in cells.
Based on the relative abundances results of nitrogen fixation prokaryotic taxa in the endophytic bacteriome of Sphagnum mosses, it can be inferred that the N fixing efficiency differed among the sampled mosses. It has been shown that the order of Rhizobiales contains nitrogen-fixing or methane-oxidation microbiomes [89, 90] that play a crucial role in the growth and development of Sphagnum mosses, especially in N and C acquisition [22, 24, 65, 91,92,93]. The results of this study show that HSP had the highest average relative abundance of Rhizobiaceae, which may be related to the lack of N in the ombrotrophic peatland ecosystem; therefore, the Sphagnum-associated microorganisms that are linked to N acquisition increase. The genus Methyloferula belongs to Beijerinckiaceae, which also contribute to nitrogen fixation [29, 36, 65, 94]. Follow-up work is required where the 15N-N2 incorporation method is used to verify which bacterial taxa plays a role in N fixation. In addition, cyanobacterial diazotrophs also play certain roles in N fixation in Sphagnum moss [25, 29]; however, in this study, Cyanobacteria were not detected in the endophytic bacteriome of the sampled Sphagnum mosses. Except for microorganisms with C and N cycling function, chemo-organoheterotrophs were also found in Sphagnum mosses. For example, Granulicella and Acidisoma are known to degrade arabinose and polysaccharides, the main components of cell walls [95, 96]. These genera that are associated with Sphagnum mosses and peat soils were isolated from peatland.
Combined with metabolic functional predictions of endophytic bacterial communities, the results showed that both metabolism and genetic information processing pathways were more abundant among the sampled Sphagnum mosses (Fig. S5). The results show that the abundances of terpenoid, amino acid, and carbohydrate metabolism genes in the endophytic bacterial community of Sphagnum mosses were elevated (Fig. 7b). These pathways are associated with plant growth, photosynthesis, and survival [97]. In a recent study, researchers have identified Sphagnum moss metabolites as key factors for microbial structure and characteristics [32]. It remains unclear whether the metabolic pathways with high abundance were related to the “host” release of an array of metabolites to affect the colonization of endophytes or to recruit specific a microbiome [98, 99] and further influence endophytic bacterial metabolic pathways. Therefore, metabolites need to be considered in further experiments to study the relationship with endophytic bacterial in Sphagnum mosses.
Conclusion
In this study, the endophytic microbial community structure and function of different species of Sphagnum mosses were examined in two different types of peatlands in China. Significant differences were found in endophytic bacterial communities among different Sphagnum mosses, including the same species of Sphagnum moss under different types of peatlands. Moreover, methane oxidation rates of methanotrophic populations of S. palustre sampled from Hani were higher than those sampled from Taishanmiao. Correspondingly, the average relative abundance of Methyloferula (an obligate methanotroph) was higher in SP. Moreover, diazotrophic taxa at the ASV level were also analyzed, and the average relative abundance of Rhizobiaceae was the highest in HSP while other diazotrophs were not present among all mosses. In addition, physiological and biochemical characteristics of mosses driving endophytic bacterial communities were also determined. The remarkable correlation between chlorophyll contents, water-holding capacity, and followed by C contents and endophytic bacterial communities at the relative abundance level of the top 10 phylum, family, and genus. These results enhance the current understanding of dominant endophytic bacteria among Sphagnum moss species in Hani and Taishanmiao peatland ecosystems. Many endophytic bacteria in Sphagnum mosses are not well known and their functions remain uncertain; therefore, 15N-N2 fixation experiments are required. Further work should include more species of Sphagnum mosses inhabiting different microhabitats, to elucidate the endophytic microbiome, including fungi and archaea. Such knowledge will improve the available understanding of their ecological roles in peatland ecosystems in the face of global climate change.
Data Availability
All 16S rRNA sequences data have uploaded to the Sequence Read Archive of the National Center for Biotechnology Information (NCBI) database and BioProject accession number PRJNA1006557.
References
Page SE, Rieley JO, Banks CJ (2011) Global and regional importance of the tropical peatland carbon pool. Global Change Biology 17:798–818. https://doi.org/10.1111/j.1365-2486.2010.02279.x
Yu Z, Loisel J, Brosseau DP, Beilman DW, Hunt SJ (2010) Global peatland dynamics since the Last Glacial Maximum. Geophys Res Lett 37. https://doi.org/10.1029/2010gl043584
Hopple AM, Wilson RM, Kolton M, Zalman CA, Chanton JP, Kostka J, Hanson PJ, Keller JK, Bridgham SD (2020) Massive peatland carbon banks vulnerable to rising temperatures. Nat Commun 11. https://doi.org/10.1038/s41467-020-16311-8
Waddington JM, Morris PJ, Kettridge N, Granath G, Thompson DK, Moore PA (2015) Hydrological feedbacks in northern peatlands. Ecohydrology 8. https://doi.org/10.1002/eco.1493
Wilson RM, Hopple AM, Tfaily MM, Sebestyen SD, Schadt CW, Pfeifer-Meister L, Medvedeff C, McFarlane KJ, Kostka JE, Kolton M, Kolka RK, Kluber LA, Keller JK, Guilderson TP, Griffiths NA, Chanton JP, Bridgham SD, Hanson PJ (2016) Stability of peatland carbon to rising temperatures. Nat Commun 7. https://doi.org/10.1038/ncomms13723
Strakova P, Penttila T, Laine J, Laiho R (2012) Disentangling direct and indirect effects of water table drawdown on above- and belowground plant litter decomposition: consequences for accumulation of organic matter in boreal peatlands. Global Change Biol 18:322–335. https://doi.org/10.1111/j.1365-2486.2011.02503.x
Updegraff K, Bridgham SD, Pastor J, Weishampel P, Harth C (2001) Response of CO2 and CH4 emissions from peatlands to warming and water table manipulation. Ecol Appl 11:311–326
Bengtsson F, Rydin H, Baltzer JL, Bragazza L, Bu ZJ, Caporn SJM, Dorrepaal E, Flatberg KI, Galanina O, Gałka M, Ganeva A, Goia I, Goncharova N, Hájek M, Haraguchi A, Harris LI, Humphreys E, Jiroušek M, Kajukało K, Karofeld E, Koronatova NG, Kosykh NP, Laine AM, Lamentowicz M, Lapshina E, Limpens J, Linkosalmi M, Ma JZ, Mauritz M, Mitchell EAD, Munir TM, Natali SM, Natcheva R, Payne RJ, Philippov DA, Rice SK, Robinson S, Robroek BJM, Rochefort L, Singer D, Stenøien HK, Tuittila ES, Vellak K, Waddington JM, Granath G, Aerts R (2020) Environmental drivers of Sphagnum growth in peatlands across the Holarctic region. J Ecol 109:417–431. https://doi.org/10.1111/1365-2745.13499
Shaw AJ, Cox CJ, Boles SB (2003) Global patterns in peatmoss biodiversity. Mole Ecol 12:2553–2570. https://doi.org/10.1046/j.1365-294X.2003.01929.x
Shaw AJ, Devos N, Cox CJ, Boles SB, Shaw B, Buchanan AM, Cave L, Seppelt R (2010) Peatmoss (Sphagnum) diversification associated with Miocene Northern Hemisphere climatic cooling? Mole Phylogen Evol 55:1139–1145. https://doi.org/10.1016/j.ympev.2010.01.020
Turetsky MR (2003) The role of bryophytes in carbon and nitrogen cycling. Bryologist 106:395–409
Turetsky MR, Bond-Lamberty B, Euskirchen E, Talbot J, Frolking S, McGuire AD, Tuittila ES (2012) The resilience and functional role of moss in boreal and arctic ecosystems. New Phytol 196:49–67. https://doi.org/10.1111/j.1469-8137.2012.04254.x
Vanbreemen N (1995) How sphagnum bogs down other plants. Trends Ecol Evol 10:270–275. https://doi.org/10.1016/0169-5347(95)90007-1
Freeman C, Ostle N, Kang H (2001) An enzymic ‘latch’ on a global carbon store - a shortage of oxygen locks up carbon in peatlands by restraining a single enzyme. Nature 409:149–149. https://doi.org/10.1038/35051650
Zhao Y, Liu C, Li X, Ma L, Zhai G, Feng X (2023) Sphagnum increases soil’s sequestration capacity of mineral-associated organic carbon via activating metal oxides. Nat Commun 14:5052–5052. https://doi.org/10.1038/s41467-023-40863-0
Opelt K, Berg G (2004) Diversity and antagonistic potential of bacteria associated with bryophytes from nutrient-poor habitats of the Baltic Sea coast. Appl Environ Microbiol 70:6569–6579. https://doi.org/10.1128/aem.70.11.6569-6579.2004
Opelt K, Chobot V, Hadacek F, Schonmann S, Eberl L, Berg G (2007) Investigations of the structure and function of bacterial communities associated with Sphagnum mosses. Environ Microbiol 9:2795–2809. https://doi.org/10.1111/j.1462-2920.2007.01391.x
Kostka JE, Weston DJ, Glass JB, Lilleskov EA, Shaw AJ, Turetsky MR (2016) The Sphagnum microbiome: new insights from an ancient plant lineage. New Phytol 211:57–64. https://doi.org/10.1111/nph.13993
Bragina A, Berg C, Cardinale M, Shcherbakov A, Chebotar V, Berg G (2012) Sphagnum mosses harbour highly specific bacterial diversity during their whole lifecycle. ISME J 6:802–813. https://doi.org/10.1038/ismej.2011.151
Bragina A, Maier S, Berg C, Mueller H, Chobot V, Hadacek F, Berg G (2012) Similar diversity of Alphaproteobacteria and nitrogenase gene amplicons on two related Sphagnum mosses. Front Microbiol 3. https://doi.org/10.3389/fmicb.2011.00275
Bragina A, Oberauner-Wappis L, Zachow C, Halwachs B, Thallinger GG, Mueller H, Berg G (2014) The Sphagnum microbiome supports bog ecosystem functioning under extreme conditions. Mol Ecol 23:4498–4510. https://doi.org/10.1111/mec.12885
Kip N, van Winden JF, Pan Y, Bodrossy L, Reichart G-J, Smolders AJP, Jetten MSM, Damste JSS, Op den Camp HJM (2010) Global prevalence of methane oxidation by symbiotic bacteria in peat-moss ecosystems. Nat Geosci 3:617–621. https://doi.org/10.1038/ngeo939
Larmola T, Leppanen SM, Tuittila E-S, Aarva M, Merila P, Fritze H, Tiirola M (2014) Methanotrophy induces nitrogen fixation during peatland development. Proc Nat Acad Sci United States Am 111:734–739. https://doi.org/10.1073/pnas.1314284111
Raghoebarsing AA, Smolders AJP, Schmid MC, Rijpstra WIC, Wolters-Arts M, Derksen J, Jetten MSM, Schouten S, Damste JSS, Lamers LPM, Roelofs JGM, den Camp H, Strous M (2005) Methanotrophic symbionts provide carbon for photosynthesis in peat bogs. Nature 436:1153–1156. https://doi.org/10.1038/nature03802
Berg A, Danielsson A, Svensson BH (2013) Transfer of fixed-N from N-2-fixing cyanobacteria associated with the moss Sphagnum riparium results in enhanced growth of the moss. Plant Soil 362:271–278. https://doi.org/10.1007/s11104-012-1278-4
A Bragina C Berg H Mueller D Moser G Berg 2013 Insights into functional bacterial diversity and its effects on Alpine bog ecosystem functioning Sci Rep 3 https://doi.org/10.1038/srep01955
Leppanen SM, Rissanen AJ, Tiirola M (2015) Nitrogen fixation in Sphagnum mosses is affected by moss species and water table level. Plant Soil 389:185–196. https://doi.org/10.1007/s11104-014-2356-6
Lindo Z, Nilsson MC, Gundale MJ (2013) Bryophyte-cyanobacteria associations as regulators of the northern latitude carbon balance in response to global change. Glob Chang Biol 19:2022–2035. https://doi.org/10.1111/gcb.12175
Carrell AA, Kolton M, Glass JB, Pelletier DA, Warren MJ, Kostka JE, Iversen CM, Hanson PJ, Weston DJ (2019) Experimental warming alters the community composition, diversity, and N-2 fixation activity of peat moss (Sphagnum fallax) microbiomes. Global Chang Biol 25:2993–3004. https://doi.org/10.1111/gcb.14715
Carrell AA, Velickovic D, Lawrence TJ, Bowen BP, Louie KB, Carper DL, Chu RK, Mitchell HD, Orr G, Markillie LM, Jawdy SS, Grimwood J, Shaw AJ, Schmutz J, Northen TR, Anderton CR, Pelletier DA, Weston DJ (2022) Novel metabolic interactions and environmental conditions mediate the boreal peatmoss-cyanobacteria mutualism. ISME J 16:1074–1085. https://doi.org/10.1038/s41396-021-01136-0
Jassey VEJ, Lamentowicz L, Robroek BJM, Gazbka M, Rusinska A, Lamentowicz M (2014) Plant functional diversity drives niche-size-structure of dominant microbial consumers along a poor to extremely rich fen gradient. J Ecol 102:1150–1162. https://doi.org/10.1111/1365-2745.12288
Sytiuk A, Cereghino R, Hamard S, Delarue F, Guittet A, Barel JM, Dorrepaal E, Kuttim M, Lamentowicz M, Pourrut B, Robroek BJM, Tuittila E-S, Jassey VEJ (2022) Predicting the structure and functions of peatland microbial communities from Sphagnum phylogeny, anatomical and morphological traits and metabolites. J Ecol 110:80–96. https://doi.org/10.1111/1365-2745.13728
Burns JH, Anacker BL, Strauss SY, Burke DJ (2015) Soil microbial community variation correlates most strongly with plant species identity, followed by soil chemistry, spatial location and plant genus. Aob Plants 7. doi: https://doi.org/10.1093/aobpla/plv030
Latz MAC, Kerrn MH, Sorensen H, Collinge DB, Jensen B, Brown JKM, Madsen AM, Jorgensen HJL (2021) Succession of the fungal endophytic microbiome of wheat is dependent on tissue-specific interactions between host genotype and environment. Sci Total Environ 759. https://doi.org/10.1016/j.scitotenv.2020.143804
Lehtonen P, Helander M, Saikkonen K (2005) Are endophyte-mediated effects on herbivores conditional on soil nutrients? Oecologia 142:38–45. https://doi.org/10.1007/s00442-004-1701-5
Renaudin M, Laforest-Lapointe I, Bellenger J-P (2022) Unraveling global and diazotrophicbacteriomes of boreal forest floor feather mosses and their environmental drivers at the ecosystem and at the plant scale in North America. Sci Total Environ 837. https://doi.org/10.1016/j.scitotenv.2022.155761
Basiliko N, Knowles R, Moore TR (2004) Roles of moss species and habitat in methane consumption potential in a northern peatland. Wetlands 24:178–185. https://doi.org/10.1672/0277-5212(2004)024[0178:Romsah]2.0.Co;2
Larmola T, Tuittila E-S, Tiirola M, Nykanen H, Martikainen PJ, Yrjala K, Tuomivirta T, Fritze H (2010) The role of Sphagnum mosses in the methane cycling of a boreal mire. Ecology 91:2356–2365. https://doi.org/10.1890/09-1343.1
Ge L, Chen C, Li T, Bu Z-J, Wang M (2023) Contrasting effects of nitrogen and phosphorus additions on fine root production and morphological traits of different plant functional types in an ombrotrophic peatland. Plant Soil. https://doi.org/10.1007/s11104-023-06087-3
Wang H, Wu L, Xue D, Liu X, Hong L, Mou L, Li X (2020) Distribution and environmental characteristics of Sphagnum peat bogs in Taishanmiao in Enshi City, Hubei Province. Wetland Sci 18:266–274
Bu Z-J, Sundberg S, Feng L, Li H-K, Zhao H-Y, Li H-C (2017) The Methuselah of plant diaspores: Sphagnum spores can survive in nature for centuries. New Phytol 214:1398–1402. https://doi.org/10.1111/nph.14575
Tian W, Xiang X, Ma L, Evers S, Wang R, Qiu X, Wang H (2019) Rare Species Shift the Structure of Bacterial Communities Across Sphagnum Compartments in a Subalpine Peatland. Front Microbiol 10:3138. https://doi.org/10.3389/fmicb.2019.03138
Bulgari D, Casati P, Brusetti L, Quaglino F, Brasca M, Daffonchio D, Bianco PA (2009) Endophytic bacterial diversity in grapevine (Vitis vinifera L.) leaves described by 16S rRNA gene sequence analysis and length heterogeneity-PCR. J Microbiol 47:393–401. https://doi.org/10.1007/s12275-009-0082-1
He C, Zhang Z, Wang Z, Shi K (2020) Water retention characteristics of Sphagnum and their relationship with soil nutrient content in the peatland of Maruo, Upland Guizhou. Plant Sci J 38:618–626
MacKinney G (1941) Absorption of light by chlorophyll solutions. J Biol Chem 140:315–322
Singleton VL, Orthofer R, Lamuela-Raventos RM (1999) Analysis of total phenols and other oxidation substrates and antioxidants by means of Folin-Ciocalteu reagent. Oxidants and Antioxidants, Pt A 299:152–178. https://doi.org/10.1016/S0076-6879(99)99017-1
Masuko T, Minami A, Iwasaki N, Majima T, Nishimura SI, Lee YC (2005) Carbohydrate analysis by a phenol-sulfuric acid method in microplate format. Anal Biochem 339:69–72. https://doi.org/10.1016/j.ab.2004.12.001
Kuk YI, Shin JS, Burgos NR, Hwang TE, Han O, Cho BH, Jung SY, Guh JO (2003) Antioxidative enzymes offer protection from chilling damage in rice plants. Crop Sci 43:2109–2117. https://doi.org/10.2135/cropsci2003.2109
Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207. https://doi.org/10.1007/bf00018060
Criquet S, Tagger S, Vogt G, Iacazio G, Le Petit J (1999) Laccase activity of forest litter. Soil Biol Biochem 31:1239–1244. https://doi.org/10.1016/s0038-0717(99)00038-3
Jassey VEJ, Chiapusio G, Gilbert D, Toussaint M-L, Binet P (2012) Phenoloxidase and peroxidase activities in Sphagnum-dominated peatland in a warming climate. Soil Biol Biochem 46:49–52. https://doi.org/10.1016/j.soilbio.2011.11.011
Fu JM, Huang BR (2001) Involvement of antioxidants and lipid peroxidation in the adaptation of two cool-season grasses to localized drought stress. Environ Exp Botany 45:105–114. https://doi.org/10.1016/s0098-8472(00)00084-8
Kox MAR, Aalto SL, Penttila T, Ettwig KF, Jetten MSM, van Kessel M (2018) The influence of oxygen and methane on nitrogen fixation in subarctic Sphagnum mosses. AMB Express 8:76. https://doi.org/10.1186/s13568-018-0607-2
Bodenhausen N, Horton MW, Bergelson J (2013) Bacterial communities associated with the leaves and the roots of Arabidopsis thaliana. Plos One 8. https://doi.org/10.1371/journal.pone.0056329
Chelius MK, Triplett EW (2001) The diversity of archaea and bacteria in association with the roots of Zea mays L. Microbial Ecology 41:252–263. https://doi.org/10.1007/s002480000087
Bolyen E, Rideout JR, Dillon MR, Bokulich N, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, Bai Y, Bisanz JE, Bittinger K, Brejnrod A, Brislawn CJ, Brown CT, Callahan BJ, Caraballo-Rodriguez AM, Chase J, Cope EK, Da Silva R, Diener C, Dorrestein PC, Douglas GM, Durall DM, Duvallet C, Edwardson CF, Ernst M, Estaki M, Fouquier J, Gauglitz JM, Gibbons SM, Gibson DL, Gonzalez A, Gorlick K, Guo J, Hillmann B, Holmes S, Holste H, Huttenhower C, Huttley GA, Janssen S, Jarmusch AK, Jiang L, Kaehler BD, Bin Kang K, Keefe CR, Keim P, Kelley ST, Knights D, Koester I, Kosciolek T, Kreps J, Langille MGI, Lee J, Ley R, Liu Y-X, Loftfield E, Lozupone C, Maher M, Marotz C, Martin BD, McDonald D, McIver LJ, Melnik AV, Metcalf JL, Morgan SC, Morton JT, Naimey AT, Navas-Molina JA, Nothias LF, Orchanian SB, Pearson T, Peoples SL, Petras D, Preuss ML, Pruesse E, Rasmussen LB, Rivers A, Robeson MS II, Rosenthal P, Segata N, Shaffer M, Shiffer A, Sinha R, Song SJ, Spear JR, Swafford AD, Thompson LR, Torres PJ, Trinh P, Tripathi A, Turnbaugh PJ, Ul-Hasan S, vander Hooft JJJ, Vargas F, Vazquez-Baeza Y, Vogtmann E, von Hippel M, Walters W, Wan Y, Wang M, Warren J, Weber KC, Williamson CHD, Willis AD, Xu ZZ, Zaneveld JR, Zhang Y, Zhu Q, Knight R, Caporaso JG, (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat Biotechnol 37:852–857. https://doi.org/10.1038/s41587-019-0209-9
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13:581. https://doi.org/10.1038/nmeth.3869
Rognes T, Flouri T, Nichols B, Quince C, Mahe F (2016) VSEARCH: a versatile open source tool for metagenomics. Peerj 4. https://doi.org/10.7717/peerj.2584
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucl Acids Res 30:3059–3066. https://doi.org/10.1093/nar/gkf436
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072. https://doi.org/10.1128/aem.03006-05
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Gloeckner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucl Acids Res 41:D590–D596. https://doi.org/10.1093/nar/gks1219
Douglas GM, Maffei VJ, Zaneveld JR, Yurgel SN, Brown JR, Taylor CM, Huttenhower C, Langille MGI (2020) PICRUSt2 for prediction of metagenome functions. Nat Biotechnol 38:685–688. https://doi.org/10.1038/s41587-020-0548-6
Kanehisa M, Sato Y, Morishima K (2016) BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J Mol Biol 428:726–731. https://doi.org/10.1016/j.jmb.2015.11.006
Louca S, Parfrey LW, Doebeli M (2016) Decoupling function and taxonomy in the global ocean microbiome. Science 353:1272–1277. https://doi.org/10.1126/science.aaf4507
Vile MA, Wieder RK, Zivkovic T, Scott KD, Vitt DH, Hartsock JA, Iosue CL, Quinn JC, Petix M, Fillingim HM, Popma JMA, Dynarski KA, Jackman TR, Albright CM, Wykoff DD (2014) N-2-fixation by methanotrophs sustains carbon and nitrogen accumulation in pristine peatlands. Biogeochemistry 121:317–328. https://doi.org/10.1007/s10533-014-0019-6
Bulgarelli D, Rott M, Schlaeppi K, van Themaat EVL, Ahmadinejad N, Assenza F, Rauf P, Huettel B, Reinhardt R, Schmelzer E, Peplies J, Gloeckner FO, Amann R, Eickhorst T, Schulze-Lefert P (2012) Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488:91–95. https://doi.org/10.1038/nature11336
Lundberg DS, Lebeis SL, Paredes SH, Yourstone S, Gehring J, Malfatti S, Tremblay J, Engelbrektson A, Kunin V, del Rio TG, Edgar RC, Eickhorst T, Ley RE, Hugenholtz P, Tringe SG, Dangl JL (2012) Defining the core Arabidopsis thaliana root microbiome. Nature 488:86. https://doi.org/10.1038/nature11237
Van Cauwenberghe J, Michiels J, Honnay O (2015) Effects of local environmental variables and geographical location on the genetic diversity and composition of Rhizobium leguminosarum nodulating Vicia cracca populations. Soil Biol Biochem 90:71–79. https://doi.org/10.1016/j.soilbio.2015.08.001
Yu X, Yang J, Wang E, Li B, Yuan H (2015) Effects of growth stage and fulvic acid on the diversity and dynamics of endophytic bacterial community in Stevia rebaudianaBertoni leaves. Front Microbiol 6. https://doi.org/10.3389/fmicb.2015.00867
Horton MW, Bodenhausen N, Beilsmith K, Meng D, Muegge BD, Subramanian S, Vetter MM, Vilhjalmsson BJ, Nordborg M, Gordon JI, Bergelson J (2014) Genome-wide association study of Arabidopsis thaliana leaf microbial community. Nat Commun 5. https://doi.org/10.1038/ncomms6320
Ortega RA, Mahnert A, Berg C, Mueller H, Berg G (2016) The plant is crucial: specific composition and function of the phyllosphere microbiome of indoor ornamentals Fems. Microbiol Ecol 92. https://doi.org/10.1093/femsec/fiw173
Zhou J, Ning D (2017) Stochastic community assembly: does it matter in microbial ecology? MicrobiolMol Biol Rev 81. https://doi.org/10.1128/mmbr.00002-17
Warshan D, Espinoza JL, Stuart RK, Richter RA, Kim S-Y, Shapiro N, Woyke T, Kyrpides NC, Barry K, Singan V, Lindquist E, Ansong C, Purvine SO, Brewer HM, Weyman PD, Dupont CL, Rasmussen U (2017) Feathermoss and epiphytic Nostoc cooperate differently: expanding the spectrum of plant-cyanobacteria symbiosis. ISME J 11:2821–2833. https://doi.org/10.1038/ismej.2017.134
Stepniewska Z, Kuzniar A (2013) Endophytic microorganisms-promising applications in bioremediation of greenhouse gases. Appl Microbiol Biotechnol 97:9589–9596. https://doi.org/10.1007/s00253-013-5235-9
Coenye T, Vandamme P (2003) Diversity and significance of Burkholderia species occupying diverse ecological niches. Environ Microbiol 5:719–729. https://doi.org/10.1046/j.1462-2920.2003.00471.x
Belova SE, Pankratov TA, Dedysh SN (2006) Bacteria of the genus Burkholderia as a typical component of the microbial community of Sphagnum peat bogs. Microbiology 75:90–96. https://doi.org/10.1134/s0026261706010164
de Souza JT, Raaijmakers JM (2003) Polymorphisms within the prnD and pltC genes from pyrrolnitrin and pyoluteorin-producing Pseudomonas and Burkholderia spp. Fems Microbiol Ecol 43:21–34. https://doi.org/10.1016/s0168-6496(02)00414-2
Baldrian P (2006) Fungal laccases - occurrence and properties. Fems Microbiol Rev 30:215–242. https://doi.org/10.1111/j.1574-4976.2005.00010.x
Jassey VEJ, Chiapusio G, Gilbert D, Buttler A, Toussaint M-L, Binet P (2011) Experimental climate effect on seasonal variability of polyphenol/phenoloxidase interplay along a narrow fen-bog ecological gradient in Sphagnum fallax. Global Chang Biol 17:2945–2957. https://doi.org/10.1111/j.1365-2486.2011.02437.x
Toberman H, Evans CD, Freeman C, Fenner N, White M, Emmett BA, Artz RRE (2008) Summer drought effects upon soil and litter extracellular phenol oxidase activity and soluble carbon release in an upland Calluna heathland. Soil Biol Biochem 40:1519–1532. https://doi.org/10.1016/j.soilbio.2008.01.004
Schlaeppi K, Dombrowski N, Oter RG, van Themaat EVL, Schulze-Lefert P (2014) Quantitative divergence of the bacterial root microbiota in Arabidopsis thaliana relatives. Proc Nat Acad Sci United States Am 111:585–592. https://doi.org/10.1073/pnas.1321597111
Esson KC, Lin X, Kumaresan D, Chanton JP, Murrell JC, Kostka JE (2016) Alpha- and gammaproteobacterial methanotrophs codominate the active methane-oxidizing communities in an acidic boreal peat bog. Appl Environ Microbiol 82:2363–2371. https://doi.org/10.1128/aem.03640-15
Reed SC, Townsend AR, Cleveland CC, Nemergut DR (2010) Microbial community shifts influence patterns in tropical forest nitrogen fixation. Oecologia 164:521–531. https://doi.org/10.1007/s00442-010-1649-6
Renaudin M, Blasi C, Bradley RL, Bellenger J-P (2022) New insights into the drivers of moss-associated nitrogen fixation and cyanobacterial biomass in the eastern Canadian boreal forest. J Ecol 110:1403–1418. https://doi.org/10.1111/1365-2745.13881
A Putkinen T Larmola T Tuomivirta HMP Siljanen L Bodrossy E-S Tuittila H Fritze 2012 Water dispersal of methanotrophic bacteria maintains functional methane oxidation in Sphagnum mosses Front Microbiol 3 https://doi.org/10.3389/fmicb.2012.00015
Bragina A, Cardinale M, Berg C, Berg G (2013) Vertical transmission explains the specific Burkholderia pattern in Sphagnum mosses at multi-geographic scale. Front Microbiol 4:394. https://doi.org/10.3389/fmicb.2013.00394
Kox MAR, van den Elzen E, Lamers LPM, Jetten MSM, van Kessel M (2020) Microbial nitrogen fixation and methane oxidation are strongly enhanced by light in Sphagnum mosses. AMB Express 10:61. https://doi.org/10.1186/s13568-020-00994-9
Kox MAR, Kop LFM, Elzen van den E, Alen van TA, Lamers LPM, Kessel van MAHJ, Jetten MSM (2020) Functional redundancy of the methane-oxidising and nitrogen-fixing microbial community associated with Sphagnum fallax and Sphagnum palustre in two Dutch fens. Mires and Peat 26. https://doi.org/10.19189/MaP.2019.SNPG.StA.1885
Jourand P, Giraud E, Bena G, Sy A, Willems A, Gillis M, Dreyfus B, de Lajudie P (2004) Methylobacterium nodulans sp nov., for a group of aerobic, facultatively methylotrophic, legume root-nodule-forming and nitrogen-fixing bacteria. Int J Syst Evolut Microbiol 54:2269–2273. https://doi.org/10.1099/ijs.0.02902-0
Lin Y, Ye G, Liu D, Ledgard S, Luo J, Fan J, Yuan J, Chen Z, Ding W (2018) Long-term application of lime or pig manure rather than plant residues suppressed diazotroph abundance and diversity and altered community structure in an acidic Ultisol. Soil Biol Biochem 123:218–228. https://doi.org/10.1016/j.soilbio.2018.05.018
Kip N, Dutilh BE, Pan Y, Bodrossy L, Neveling K, Kwint MP, Jetten MSM, den Camp HJMO (2011) Ultra-deep pyrosequencing of pmoA amplicons confirms the prevalence of Methylomonas and Methylocystis in Sphagnum mosses from a Dutch peat bog. Environ Microbiol Rep 3:667–673. https://doi.org/10.1111/j.1758-2229.2011.00260.x
Kip N, Fritz C, Langelaan ES, Pan Y, Bodrossy L, Pancotto V, Jetten MSM, Smolders AJP, den Camp HJMO (2012) Methanotrophic activity and diversity in different Sphagnum magellanicum dominated habitats in the southernmost peat bogs of Patagonia. Biogeosciences 9:47–55. https://doi.org/10.5194/bg-9-47-2012
Kip N, Ouyang W, van Winden J, Raghoebarsing A, van Niftrik L, Pol A, Pan Y, Bodrossy L, van Donselaar EG, Reichart G-J, Jetten MSM, Damste JSS, den Camp HJMO (2011) Detection, isolation, and characterization of acidophilic methanotrophs from Sphagnum mosses. Appl Environ Microbiol 77:5643–5654. https://doi.org/10.1128/aem.05017-11
Kolton M, Weston DJ, Mayali X, Weber PK, McFarlane KJ, Pett-Ridge J, Somoza MM, Lietard J, Glass JB, Lilleskov EA, Shaw AJ, Tringe S, Hanson PJ, Kostka JE (2022) Defining the Sphagnum core microbiome across the North American continent reveals a central role for diazotrophicmethanotrophs in the nitrogen and carbon cycles of boreal peatland ecosystems. Mbio 13. https://doi.org/10.1128/mbio.03714-21
Kulichevskaya IS, Suzina NE, Liesack W, Dedysh SN (2010) Bryobacter aggregatus gen. nov., sp nov., a peat-inhabiting, aerobic chemo-organotroph from subdivision 3 of the Acidobacteria. Int J Syst Evolut Microbiol 60:301–306. https://doi.org/10.1099/ijs.0.013250-0
Pankratov TA, Dedysh SN (2010) Granulicella paludicola gen. nov., sp nov., Granulicella pectinivorans sp nov., Granulicella aggregans sp nov and Granulicella rosea sp nov., acidophilic, polymer-degrading acidobacteria from Sphagnum peat bogs. Int J Syst Evol Microbiol 60:2951–2959. https://doi.org/10.1099/ijs.0.021824-0
Lange BM, Rujan T, Martin W, Croteau R (2000) Isoprenoid biosynthesis: the evolution of two ancient and distinct pathways across genomes. Proc Nat Acad Sci United States Am 97:13172–13177. https://doi.org/10.1073/pnas.240454797
Fudyma JD, Lyon J, AminiTabrizi R, Gieschen H, Chu RK, Hoyt DW, Kyle JE, Toyoda J, Tolic N, Heyman HM, Hess NJ, Metz TO, Tfaily MM (2019) Untargeted metabolomic profiling of Sphagnum fallax reveals novel antimicrobial metabolites. Plant Direct 3. https://doi.org/10.1002/pld3.179
Hiruma K (2019) Roles of plant-derived secondary metabolites during interactions with pathogenic and beneficial microbes under conditions of environmental stress. Microorganisms 7. https://doi.org/10.3390/microorganisms7090362
Acknowledgements
This research was supported by the National Natural Science Foundation of China (42001093), the Second Tibetan Plateau Scientific Expedition (2019QZKK0304), the Youth Innovation Promotion Association of the Chinese Academy of Sciences (2022376), the Youth Science and Technology Innovation Team Program of Sichuan Province of China (2021JDTD011), and the Youth Innovation Program of CIB (QNTS202201). We wish to thank Prof. Meng Wang and Dr. Leming Ge of Northeast Normal University, Changchun, Jilin, China for leading us to the sample sites and helping us collect soil samples. We also thank to Prof. Lin Wu and the master of MouLi of Hubei Minzu University, Enshi, Hubei, China for assistance with the sample collection. We sincerely thank the editors and anonymous reviewers for their valuable comments on the manuscript.
Funding
This work was supported by the National Natural Science Foundation of China (42001093), the Second Tibetan Plateau Scientific Expedition (2019QZKK0304), the Youth Innovation Promotion Association of the Chinese Academy of Sciences (2022376), the Youth Science and Technology Innovation Team Program of Sichuan Province of China (2021JDTD011), and the Youth Innovation Program of CIB (QNTS202201).
Author information
Authors and Affiliations
Contributions
YW, DX, and HC conceptualized the study. DX and HC acquired the funding. YW, XC, and QQ investigated and collected samples. YW performed the experiments. YW and DX performed the data analysis. YW and DX wrote the original draft. All authors reviewed and contributed to the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Y., Xue, D., Chen, X. et al. Structure and Functions of Endophytic Bacterial Communities Associated with Sphagnum Mosses and Their Drivers in Two Different Nutrient Types of Peatlands. Microb Ecol 87, 47 (2024). https://doi.org/10.1007/s00248-024-02355-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-024-02355-6