Abstract
Various Lactobacillus paracasei strains are found in diverse environments, including dairy and plant materials and the intestinal tract of humans and animals, and are also used in the food industry or as probiotics. In this study, we have isolated a new strain L. paracasei subsp. paracasei IBB3423 from samples of raw cow milk collected in a citizen science project. IBB3423 showed some desired probiotic features such as high adhesion capacity and ability to metabolize inulin. Its complete genome sequence comprising the chromosome of 3,183,386 bp and two plasmids of 5986 bp and 51,211 bp was determined. In silico analysis revealed numerous genes encoding proteins involved in carbohydrate metabolism and of extracellular localization likely supporting interaction with host tissues. In vitro tests confirmed the high adhesion capacity of IBB3423 and showed that it even exceeds that of the highly adhesive L. rhamnosus GG. Curing of the larger plasmid indicated that the adhesive properties depend on the plasmid and thus could be determined by its pilus-encoding spaCBA genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lactobacillus is the largest genus of the lactic acid bacteria (LAB) group, which currently comprises 236 species listed in the List of Prokaryotic Names (October 2018; www.bacterio.net). Members of this genus represent an extremely diverse group of species with various physiological, biochemical, and genetic characteristics and are able to colonize diverse ecological niches, including the gastrointestinal tract (GIT) of humans and animals, plants, and dairy food environments (Makarova et al. 2006; Kandler 1983). Lactobacilli are rod-shaped, Gram-positive, non-spore-forming, lactic acid-producing, generally nonmotile bacteria that can survive mainly in anaerobic environments (Claesson et al. 2007; Kandler 1983). Due to their advantageous properties and a long history of safe use, lactobacilli are used as starter cultures in both traditional fermentation and various industrial bioprocesses (Stiles and Holzapfel 1997). Furthermore, having the generally recognized as safe (GRAS) status, many species are considered probiotics, i.e., live microorganisms intended to confer health benefits after consumption (Makarova et al. 2006; Fuller 1991; Ouwehand et al. 2002; Lebeer et al. 2008). The probiotic properties of microorganisms are determined by factors that include their adaptation to the GIT and those directly mediating the health effects to the host. The adaptation factors include the ability to survive in the harsh conditions of the GIT (low pH and high concentration of bile salts) and adherence to mucosal surfaces (mucus, extracellular matrix proteins (ECM), or epithelial cells) (Holzapfel et al. 2001; Marco et al. 2006; Lebeer et al. 2008). One of the most desirable traits for probiotic bacteria is good adherence capacity, as it is responsible for successful colonization of the mucosal surfaces (Ouwehand et al. 2000; Styriak et al. 2003). By adhering to the host gut, the probiotic microbiota provides beneficial effects through various mechanisms ensuring the maintenance of gut barrier function, enhancement of a balanced microbial flora, immunomodulation of gut mucosal defenses, and competitive exclusion of pathogens (Jankowska et al. 2008; Lim 2012). The process of adhesion is initially based on non-specific physical interactions between two surfaces (such as van der Waals forces, repulsive electrostatic double-layer forces, hydrophobic interactions, short-range forces, and ion bridging). Generally, hydrophobic cells adhere strongly to hydrophobic surfaces, and thus the phenomenon of cell surface hydrophobicity (CSH) is responsible for cell aggregation, and, as human body surfaces are mostly of a hydrophobic character, it promotes binding of hydrophobic bacteria to the intestine epithelial tissue (Krasowska and Sigler 2014). After the initial attachment to the epithelium surface, a more specific interplay between bacterial adhesion proteins and mucosal surfaces takes place, including binding to specific receptors (Pérez et al. 1998; Turroni et al. 2013). A large number of diverse factors mediating adhesion to different components of the mucosa have been identified largely owing to genome sequencing of various lactobacilli paired with integrated genomic techniques (Kant et al. 2011; Smokvina et al. 2013). However, in contrast to the often well-characterized adhesive determinants of pathogens and their specific host receptors, functional understanding of the adherence of commensal bacteria is only fragmentary. These bacterial adhesive factors include exopolysaccharides (EPSs), teichoic acids (TAs), surface layer (S-layer) proteins, and some other membrane- or cell wall-associated proteins protruding from the cell (Lebeer et al. 2008). Molecules involved in adhesion can be attached to the cell wall by LPXTG (Leu-Pro-X-Thr-Gly)-type anchor or other peptidoglycan-binding domains such as LysM, WxL, SH3, or PG (Kleerebezem et al. 2010; Visweswaran et al. 2014). Most adhesins of lactobacilli belong to a class of sortase-dependent proteins (Velez et al. 2007), which contain an N-terminal signal sequence for transport through the membrane and a C-terminal motif LPXTG for cell wall anchoring performed by the enzyme sortase (Navarre and Schneewind 1999). In Gram-positive pathogens, the SpaCBA pili employing an LPXTG-like motif are the major driver of adhesion (Mandlik et al. 2008). Similar structures have been also identified in many lactobacilli, for example, L. rhamnosus GG and L. paracasei LOCK 0919 (Kankainen et al. 2009; Aleksandrzak-Piekarczyk et al. 2016).
To interact with host receptors, bacterial adhesion proteins contain specific domains, among them are the following: mucin binding (MucBP), collagen binding (CnaB, collagenBindB, collagen_bind), fibronectin binding (FbpA), leucine-rich repeats (LRR), Ig-like (Big_3, BID_2), legume lectin-like (lectin L-type), and ZnuA (Barr et al. 2013; Dintilhac et al. 1997; Gabbianelli et al. 2011). Of those, the following have been studied in some detail: fibronectin-binding proteins in Lactobacillus casei BL23 (Muñoz-Provencio et al. 2010), collagen-binding surface proteins in L. reuteri NCIB11951 and L. acidophilus NCFM (Aleljung et al. 1994; Buck et al. 2005), mucus-adhesion-promoting protein (MapA) in L. reuteri (Miyoshi et al. 2006), and lectin-like mannose-specific adhesin in L. plantarum WCFS1 (Pretzer et al. 2005). A key role in adhesion to intestinal cells has also been established for so-called S-layer proteins in a number of lactobacilli such as L. brevis (Hynonen et al. 2002), L. crispatus (Toba et al. 1995; Antikainen et al. 2002), L. helveticus (Johnson-Henry et al. 2007), and L. acidophilus (Buck et al. 2005). Some probiotic strains devoid of the S-layer proteins encode an aggregation-promoting factor sharing several features with the S-layer proteins (Turpin et al. 2012). Additionally, cell surface–associated cell wall hydrolases Msp2/p40 and Msp1/p75 of L. casei have been shown to bind to mucin, collagen, and cultured epithelial cells (Bäuerl et al. 2010).
Comparative studies of lactobacilli have shown that their health efficacy is not only species-specific, but also varies between strains of a species. This observation justifies the need to characterize Lactobacillus strains individually, also at the genome level (Ventura et al. 2009; FAO/WHO 2002). In this context, the L. casei taxonomic group comprised of three related species (L. casei, L. paracasei, and L. rhamnosus) (Felis and Dellaglio 2007) is of great interest as numerous commonly utilized probiotic strains belong there (Kalliomäki et al. 2003; Sykora et al. 2005; Klaenhammer et al. 2008; Almeida et al. 2012; Kankainen et al. 2009; Douillard et al. 2013; Smokvina et al. 2013). Despite the groups’ importance and apparently comprehensive studies, some species affiliations of their members are turned out to be erroneous, e.g., a strain previously assigned as L. casei LOCK 0919 (Koryszewska-Baginska et al. 2013) basing on a re-examination of its genome sequence has now been renamed L. paracasei LOCK 0919.
In this study, we isolated a new strain L. paracasei subsp. paracasei IBB3423 from samples of raw cow milk from northeastern Poland. Functional tests revealed their high adhesion capacity and ability to metabolize inulin. Its DNA was sequenced and predicted to comprise two plasmids in addition to the chromosome. Bioinformatic analyses revealed the presence of a number of genes coding for potential adhesion proteins, including pilus-encoding spaCBA genes residing in one of the plasmids. The plasmid’s removal deprived the bacteria of their capacity to adhere, indicating that it most likely depends on the SpaCBA pili.
Materials and methods
Isolation of strains, growth conditions, and DNA extraction for genome sequencing
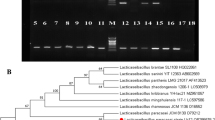
The raw cow milk samples were collected from northeastern Poland and L. paracasei subsp. paracasei IBB3423 strain was isolated from them by students (Supplementary Information S6) within the citizen science project (“Paths of Copernicus” MakeTogether program). Four other Lactobacillus strains (L. rhamnosus GG, L. rhamnosus LOCK 0900, L. rhamnosus LOCK 0908, and L. paracasei LOCK 0919) of human origin were obtained from commercial dietary supplements under the brand names of Dicoflor (Bayer, Germany; GG) and Latopic (Biomed, Poland; LOCK 0900, LOCK 0908 and LOCK 0919). Strain IBB3423 was isolated by serial dilution on MRS broth (De Man, Rogosa and Sharpe; Difco) solidified with 1% agar and incubating anaerobically (37 °C, 48 h). Individual colonies were inoculated into liquid MRS and propagated overnight anaerobically at 37 °C. Total DNA was extracted using a Genomic Mini kit (A&A Biotechnology, Poland) according to the manufacturer’s instruction with the following modifications. An overnight culture of L. paracasei subsp. paracasei IBB3423 in MRS medium was centrifuged (14,000 rpm, 1 min) and the pellet was suspended in TES (0.2 M Tris-HCl pH 8.0, 0.5 mM EDTA, 0.5 M sucrose) with lysozyme (10 mg/ml) and mutanolysin (10 U/ml) and incubated (37 °C, 1 h) with occasional mixing. The suspension was centrifuged (14,000 rpm, 3 min) and the pellet was gently suspended in the supplied buffer. The obtained DNA was characterized and quantified by measuring its OD at 230, 260, and 280 nm. Identification at species level was based on 16S rDNA sequencing following amplification with universal primers 27F (5′-AGAGTTTGATCCTGGCTCAG-3′) and 1492R (5′-GGTTACCTTGTTACGACTT-3′) (Lane 1991). The nucleotide sequence of the PCR product was compared with other sequences available using the Basic Local Alignment Search Tool (BLAST; https://blast.ncbi.nlm.nih.gov/Blast.cgi/; Altschul et al. 1990) and the ANI (average nucleotide identity) calculator software (https://www.ezbiocloud.net/tools/ani/; Yoon et al. 2017), which indicated that IBB3423 belongs to L. paracasei subsp. paracasei and has the highest similarity to L. paracasei subsp. paracasei TMW 1.1434.
Aggregation, hydrophobicity, and adherence tests
The tests for aggregation, hydrophobicity, and adherence to abiotic (glass and polystyrene) and biotic (collagen, gelatine, and mucus) surfaces were performed as described previously (Aleksandrzak-Piekarczyk et al. 2016). The following criteria were used to classify the adherence properties: A ≥ 3, strongly adherent; 3 > A > 2, moderately adherent; 2 > A > 1, weakly adherent; A ≤ 1, non-adherent.
For adherence tests to epithelial tissue, the colorectal carcinoma cell line Caco-2 (Cell Line Service GmbH, Germany) from 42nd passage was cultured in T75 Roux bottles (Greiner Bio-One GmbH, Germany) as described previously (Nowak et al. 2016) in DMEM (Dulbecco’s Modified Eagle’s Medium; Sigma-Aldrich, USA) supplemented with 10% FBS (fetal bovine serum; Thermo Fisher Scientific, USA), 25 mM HEPES (4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid; Sigma-Aldrich, USA), 4 mM GlutaMAX (Thermo Fisher Scientific, USA), a mixture of 100 μg/ml streptomycin, and 100 IU/ml penicillin (Sigma-Aldrich, USA). Cells were incubated (37 °C, 5% CO2, humidity > 95%) for 7 days to reach confluence and then were detached with TrypleTM Express (Thermo Fisher Scientific, USA) according to manufacturer’s instruction. The cell suspension was centrifuged (200×g, 5 min), the pellet was suspended in fresh DMEM, and the number and viability of cells were determined by trypan blue staining (Sigma-Aldrich, USA). For the adherence assay, Caco-2 cells were placed in a 24-well plate (2.5 × 105 cells/well) and left overnight, while bacteria were cultivated in MRS broth (24 h, 37 °C), centrifuged (9300×g, 10 min), washed with phosphate-buffered saline (PBS), and suspended in DMEM without supplements in the amount of 7–8 × 108 CFU/ml. Medium was aspirated from Caco-2 cells and 1 ml of bacteria in DMEM was added. The plate was incubated (2 h, 37 °C, 5% CO2, humidity > 95%) and non-adherent bacteria were removed by gentle washing with PBS. Caco-2 cells with adhered bacteria were detached with 200 μl 1% trypsin (Sigma-Aldrich, USA) for 10 min at 37 °C, scraped with a cell scraper (Greiner Bio-One GmbH, Germany), transferred into Eppendorf tubes, and centrifuged (9300×g, 10 min). To lyse the Caco-2 cells, the pellet was resuspended in 1 ml 0.1% Triton X-100 (Sigma-Aldrich, USA) and incubated for 5 min at RT. The released bacteria were counted by plating on MRS agar and incubating for 48 h at 37 °C. The adherence rate was calculated as follows: A[%] = (logA2/logA1) × 100, where A1 is the number of colony-forming units (CFU) of initial bacteria added to the well and A2 is the number (CFU) of adhered bacteria.
For microscopic visualization of adhering bacteria, the procedure described above was carried out in an 8-chamber Lab-TekTM microplate (Thermo Fisher Scientific, USA). After 2 h of incubation, non-adherent bacteria were removed; wells were washed with PBS, fixed with 70% methanol for 15 min, and then stained for 10 min with 0.1% crystal violet. After that, wells were washed with 70% ethanol and dried overnight. Samples were observed at 100× objective under a microscope (Nikon Eclipse Ci H600L, Japan) connected to a Nicon Digital Sight DS-U3 camera using NIS-elements BR 3.0 imaging software (Nikon, Japan).
Data were analyzed using two-way analysis of variance (ANOVA), and differences between values with normal distribution were evaluated by Student’s t test using OriginPro 6.1 software (OriginLab Corporation, Northampton, USA). The differences were deemed significant at P < 0.05. The results are presented as mean ± standard deviation (SD).
Genome sequencing and bioinformatic analyses
The total DNA of L. paracasei subsp. paracasei IBB3423 was subjected to whole genome sequencing (WGS) on a MiSeq system (Illumina, San Diego, CA) at the DNA Sequencing and Oligonucleotide Synthesis Laboratory, IBB PAS. DNA was sheared to the appropriate size and used for paired-end TruSeq (Illumina, USA) library construction following the manufacturer’s instructions. Additionally, the long-span mate-pair library was constructed from non-sheared DNA using Nextera Mate Pair Kit (Illumina, USA) following manufacturer’s instructions. Both libraries were sequenced in the paired-end mode using a v3 (600 cycles) chemistry kit (Illumina, USA). Obtained sequence reads were filtered by the quality and assembled using Newbler v3.0 software (Roche, USA). Gaps remaining in the genome assembly were closed by PCR amplification of DNA fragments followed by Sanger sequencing on an ABI3730xl Genetic Analyzer (Life Technologies, USA) using BigDye Terminator Mix v. 3.1 chemistry (Life Technologies, USA). To obtain complete nucleotide sequence of the genome errors, misassemblies were corrected using the Seqman software (DNAStar, USA). Annotation of open reading frames (orfs) and non-coding RNAs was done using the RAST server (http://rast.nmpdr.org/; Aziz et al. 2008) and checked by BLAST analysis when needed. RAST and BASYs (https://www.basys.ca/; Van Domselaar et al. 2005) software were applied for constructing Clusters of Orthologous Groups (COGs) of predicted proteins. Carbohydrate metabolic pathways were reconstructed using BlastKOALA tool from KEGG (Kyoto Encyclopedia of Genes and Genomes; https://www.kegg.jp/blastkoala/; Kanehisa et al. 2016). Carbohydrate-specific enzymes were annotated by the dbCAN prediction web server (http://csbl.bmb.uga.edu/dbCAN/; Yin et al. 2012). Transporters were predicted by searching the TCDB database (http://www.tcdb.org/; Saier et al. 2016) with the BLASTP program with the Expect value (E value) lower than e−05. Possible bacteriophage sequences were searched using PHAST (http://phast.wishartlab.com/; Zhou et al. 2011). CRISPR loci were identified using the CRISPRFinder tool (http://crispr.i2bc.paris-saclay.fr/; Grissa et al. 2007). Genome visualization was done using CGView server (http://cgview.ca/; Grant and Stothard 2008). The similarity of IBB3423 plasmids to other plasmids was evaluated using the standard BLASTN program at the NCBI site and best hits were used to show sequence similarity using Circoletto with the selected E value of 0.1 (http://tools.bat.infspire.org/circoletto/; Darzentas 2010). For comparative studies, the following 25 complete genome sequences of L. casei group were downloaded from NCBI: L. paracasei ATCC 334 (NC_008526.1), L. paracasei FAM18149 (NZ_CP017261.1), L. paracasei TK1501 (NZ_CP017716.1), L. paracasei IIA (NZ_CP014985.1), L. paracasei subsp. paracasei TMW 1.1434 (NZ_CP016355.1), L. paracasei subsp. paracasei 8700:2 (NC_022112.1), L. paracasei N1115 (NZ_CP007122.1), L. paracasei LOCK 0919 (NC_021721.1), L. paracasei subsp. paracasei JCM 8130 (NZ_AP012541.1), L. paracasei CAUH35 (NZ_CP012187.1), L. paracasei L9 (NZ_CP012148.1), L. paracasei KL1 (NZ_CP013921.1), L. paracasei HD1.7 (NZ_CP025582.1), L. paracasei EG9 (NZ_CP029546.1), L. paracasei Lpc10 (NZ_CP029686.1), L. paracasei LC355 (CP029536.1), L. paracasei Zhang (NC_014334.1), L. paracasei BD-II (NC_017474.1), L. paracasei LC2W (NC_017473.1), L. paracasei HDS-01 (NZ_CP026097.1), L. casei BL23 (NC_010999.1), L. casei 12A (NZ_CP006690.1), L. casei W56 (NC_018641.1), L. casei LC5 (NZ_CP017065.1), and L. casei ATCC 393 (NZ_AP012544.1). The average nucleotide identity between IBB3423 and respective Lactobacillus genomes was calculated with the ANI calculator software. Comparisons between chromosomes were performed using the LASTZ program with the step length of 20 and seed pattern of 12 of 19 (https://www.geneious.com/plugins/lastz-plugin/; Harris 2007).
Data access
The finished genome sequence for L. paracasei subsp. paracasei IBB3423 comprising the chromosome and two plasmids pLCAKO.1 and pLCAKO.2 has been deposited at the NCBI GenBank database with accession numbers CP022954, CP022955, and CP022956, respectively. The IBB3423 strain has been deposited at the publicly accessible Polish Collection of Microorganisms (PCM), culture no. 3007.
Sugar fermentation pattern
To determine the sugar fermentation profile of L. paracasei subsp. paracasei IBB3423 and other Lactobacillus strains, the API 50CH kit (BioMerieux, France) was used according to the manufacturer’s instructions. The fermentation patterns were recorded after 48 h of aerobic incubation at 37 °C. Fermentation of carbohydrate was detected by acid production demonstrated by a change in color of the pH indicator present in the medium.
Plasmid curing
The pLCAKO.2 plasmid was removed from strain IBB3423 by serial passages at a 10−3-fold dilution in MRS broth and culturing for 96 h at 37 °C under anaerobic condition. Appropriate dilutions of selected passages (after 4, 12, 20, 28, 32, 40, and 48 days of cultivation) were plated on MRS agar plates. After 48 h of incubation, visible colonies were randomly selected for further analysis. The absence of pLCAKO.2 plasmid was assessed by PCR with primers pLCAKO_3F (5′-CCTCCTTTAGACGCTGAACG-3′), pLCAKO_3R (5′-GGGCGGTACTTTATGGCAAC-3′), pLCAKO_7F (5′-CGCCTATCAAGTCGAAGGAG-3′), and pLCAKO_7R (5′-TCGAGCATCGCCTGCATACG-3′). A strain devoid of pLCAKO.2 was selected after 48 days of cultivation and named IBB3423 ΔpLCAKO.2.
Results
Sugar fermentation profile and preliminary studies of adhesion to bare PS plates of L. paracasei subsp. paracasei IBB3423
The ability of L. paracasei subsp. paracasei IBB3423 to utilize certain carbohydrates was compared with the fermentation profiles of well-characterized L. casei group bacteria with fully sequenced genomes as follows: L. rhamnosus GG, L. rhamnosus LOCK 0900, L. rhamnosus LOCK 0908, and L. paracasei LOCK 0919 (Kankainen et al. 2009; Aleksandrzak-Piekarczyk et al. 2013; Koryszewska-Baginska et al. 2013; Koryszewska-Baginska et al. 2014), originally isolated from the human gastrointestinal tract.
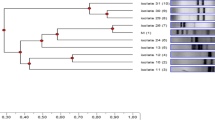
All the strains fermented numerous simple and complex carbohydrates (Fig. 1), but IBB3423 was unique among them being capable of metabolizing inulin, d-adonitol (also called d-ribitol), and l-sorbose but not l-rhamnose, dulcitol, inositol, and l-fucose. The inability to use those sugars is probably related to their low abundance in cow milk.
Fermentative profile of L. paracasei subsp. paracasei IBB3423. A positive result is indicated by black rectangles, partially by gray and negative by white rectangles. No strain could use glycerol, erythritol, d-xylose, l-xylose, methyl-β-d-xylopyranoside, methyl-α-d-mannopyranoside, d-melibiose, d-raffinose, starch, glycogen, d-lyxose, xylitol, d-fucose, l-arabitol, 2-ketogluconate potassium, or 5-ketogluconate potassium
In addition, L. rhamnosus GG and L. paracasei LOCK 0919 are strains of a considerable adhesion capacity (Aleksandrzak-Piekarczyk et al. 2016; Segers and Lebeer 2014) in contrast to poorly adherent L. paracasei LOCK 0900 and L. rhamnosus LOCK 0908 (Aleksandrzak-Piekarczyk et al. 2013; Koryszewska-Baginska et al. 2014). The properties of adhesion to microtiter PS plates of IBB3423, LOCK 0900, LOCK 0908, LOCK 0919, and GG were compared. L. rhamnosus GG isolate and L. paracasei LOCK 0919 showed high/moderate-level PS binding, while the L. paracasei LOCK 0900 and L. rhamnosus LOCK 0908 strains showed virtually no binding under the conditions tested. In comparison with them, L. paracasei IBB3423 presented the strongest adherences ability, even as much as 35% higher than the best adhesive L. rhamnosus GG strain (Figure S1).
Characteristics of L. paracasei subsp. paracasei IBB3423 genome
In order to identify the genetic determinants responsible for the strong adherence of L. paracasei subsp. paracasei IBB3423 and the wide range of metabolized carbon sources, we sequenced its entire genome. In addition to the circular chromosome of 3,183,386 bp, two circular plasmids designated pLCAKO.1 (5986 bp) and pLCAKO.2 (51,211 bp) were found. The GC content of the chromosome is 46.3% and, respectively, 42.7% and 43.8% for pLCAKO.1 and pLCAKO.2. These values agree well with the GC content in other lactobacilli (Sun et al. 2015); however, local GC content variation in the chromosome and a clear-cut GC skew at the origin of replication is evident (Fig. 2). A total of 3216 genes were identified in the chromosome, of which 3116 were annotated as protein-coding genes, resulting in the coding capacity of 84.5%. As with the GC content, the distribution of genes on the two DNA strands is substantially skewed around ori so that their transcription tends to agree with the direction of replicative fork migration. Fifty-nine tRNA genes representing all amino acids were found, and five complete rRNA operons, three on the forward strand, and two on the reverse strand. Most of the tRNAs genes are clustered in the close neighborhood of rRNA locus (Fig. 2). All the rRNA operons and nearly all tRNA genes are arranged in the direction of fork migration. Seventy-six genes were identified on pLCAKO.1 and pLCAKO.2 plasmids (Table 1).
A total of 2171 genes from entire genome (~ 67%) could be assigned a putative biological function, while the remaining ~ 33% genes were annotated as hypothetical or of unknown function. When their biological roles according to COG categories were analyzed, out of the total of 3189 predicted proteins, 1999 (62.7%) could be assigned to a COG functional category. About 10% of the proteins are involved in cellular processes and signaling (categories D, M, N, O, T, U, V), 16% in information storage and processing (categories J, K, L), almost 26% in metabolism (categories C, E, F, G, H, I, P, and Q), and ca. 11% are poorly characterized (categories R and S) (Supplementary Table S1). Proteins related to carbohydrate and amino acid transport and metabolism are particularly numerous, consistent with the wide fermentation profile of the strain and the inability of lactobacilli to synthetize most amino acids. Notably, no genes associated with pathogenesis were found, indicating that the IBB3423 strain should be safe to use as a probiotic.
A widespread abundance of prophages in the genomes of Lactobacillus spp. has been reported (Mercanti et al. 2016) and IBB3423 was typical in this respect. Four intact prophage regions ranging in length from 24.9 to 41.5 kb partially homologous to prophage sequences widely distributed among other lactobacilli were identified. All of them contain a phage attachment (ATT) site. Judging from the presence of genes involved in bacterial lysis (holin and phage lysin), two of those phages may follow the lysogenic pathway.
Clustered regularly interspaced short palindromic repeats (CRISPRs) are components of the type of bacterial immune system against mobile genetic elements, which are often found adjacent to the cas gene cluster encoding Cas protein complex (Haft et al. 2005). Type II-A/Lsal1 CRISPR-cas system is present in the IBB3423 genome. The CRISPR locus contains 26 perfect repeats of a 36-bp-long sequence (5′GCTCTTGAACTGATTGATTCGACATCTACCTGAGAC) and one imperfect repeat (5′ACTCTTGAACTGATTGATTCGACATCTACCTGAGAC). Four CRISPR-associated cas genes (LCAKO_2393, LCAKO_2394, LCAKO_2395, and LCAKO_2396) are present upstream of the DNA repeats.
The sequence assembly of the entire IBB3423 genome clearly indicated the presence of two plasmids: pLCAKO.1 and pLCAKO.2. Their organization is shown in Fig. 3. The smaller pLCAKO.1 plasmid contains nine genes, all transcribed in the clockwise orientation and mostly encoding proteins with unknown or hypothetical functions homologous to proteins from non-lactobacilli (Supplementary Table S2). The product of the repB gene harbors a Rep_trans superfamily domain (pfam02486), which is characteristic for rolling circle replicating (RC) plasmids (Balson and Shaw 1990).
Circular genetic maps of plasmids pLCAKO.1 and pLCAKO.2 of L. paracasei subsp. paracasei IBB3423. orfs are indicated by arrows showing the direction of transcription. The pilus-encoding spaCBA-srtC cluster is colored in red and lactose operon in green; an asterisk indicates pseudogenes. The two maps are drawn not to scale
The lager pLCAKO.2 harbors 67 putative genes, of which 64 were predicted to code for proteins. Most exhibit homology with genes located on chromosomes or plasmids of other lactobacilli. Putative biological functions could be assigned to 35 proteins. Fifteen transposase/insertion sequences and one gene encoding pin-related site recombinase/DNA invertase are present in pLCAKO.2 (Supplementary Table S2). The abundance of such elements likely reflects the occurrence of numerous rearrangements during plasmid evolution. The RepA protein encoded by pLCAKO.2 (gene LCAKO_2p1) contains RepA_N domain (pfam06970), which is characteristic for class F family pL32-type theta-replication proteins (Tanaka and Ogura 1998). That pLCAKO.2 is a theta-replicating plasmid is additionally indicated by its substantial size (~ 50 kb) as it has been shown that plasmids of this size found in natural isolates of Lactobacillus have been shown typically to replicate via the theta mechanism (Wang and Lee 1997). Plasmid replication and stability are closely related, and downstream of repA lies orf67 encoding a protein assigned to the ParA family of ATPases; they are involved in segregational stability preventing plasmid loss during cell division.
Among the genes encoding proteins of known function, pLCAKO.2 harbors genes encoding serine acetyltransferase, cystathionine gamma-lyase, and cystathionine beta-synthase enzymes, which are involved in amino acid metabolism and a lactose-specific PTS operon (Lac-PTS) responsible for lactose uptake and fermentation, comprising lacTEGF genes (LCAKO_2p26–LCAKO_2p29), and is absent in the chromosome. No such operon is present in the chromosome. Noteworthy is another operon with known function, the pilus- and sortase-encoding spaCBA-srtC gene cluster (LCAKO_2p51–LCAKO_2p54), which could play an important role in the adhesion to host cells. Its presence prompted us to investigate the dependence of the excellent adherence of IBB3423 on the pLCAKO.2 plasmid.
pLCAKO.1 shows no significant homologies with other plasmids deposited in the GenBank. In contrast, pLCAKO.2 exhibits conservation of almost all its sequence with other plasmids, albeit its gene layout is unique. This unique architecture confirms the earlier suggestion of numerous rearrangements based on the high content of mobile genetic elements in IBB3423. The most similar plasmid is pL11434-1 from L. paracasei subsp. paracasei TMW 1.1434 along 76% of the pLCAKO.2 length, including mobile elements and genes encoding Lac-PTS, serine acetyltransferase, cystathionine lyase, and several hypothetical proteins. Notably, a region homologous to the spaCBA-srtC cluster and downstream genes (orfs51–61) is present in two plasmids only—pLOCK 0919 (Aleksandrzak-Piekarczyk et al. 2016) and p1_LC355 (NZ_CP029537.1) (Fig. 4).
Sequence similarity between pLCAKO.2 plasmid and other lactobacilli plasmids. Plasmid sequences are placed at the circumference. Ribbons represent local alignments produced by BLAST, their width shows alignment length, and colors blue, green, orange, and red correspond to the alignment bit scores in the four quartiles, from the lowest (bit score below 25%) to the highest (bit score between 75 and 100%). Twisted ribbons connect sequences in opposite orientations (inverted). An asterisk indicates region homologous to the spaCBA-srtC cluster and downstream genes (orfs51–61) from the pLCAKO.2 plasmid. The following plasmid sequences were compared: pL11434-1 GenBank accession no. NZ_CP016356.1 from L. paracasei subsp. paracasei strain TMW 1.1434; pFAM18149.21 NZ_CP017262.1 from L. paracasei FAM18149; p3_IIA NZ_CP014988.1 from L. paracasei; p1_ATCC 334 NC_008502.1 from L. paracasei ATCC 334; p1_CAUH35 NZ_CP012188.1 from L. paracasei CAUH35; p2_8700:2 NC_022123.1 from L. paracasei subsp. paracasei 8700:2; p1_LC355 NZ_CP029537.1 from L. paracasei LC355; pEG9B NZ_CP029548.1 from L. paracasei EG9; pLOCK 0919 NC_021722.1 from L. paracasei LOCK 0919; pLC2W NC_017475.1 from L. paracasei LC2W; pLC1 NC_013200.1 from L. rhamnosus Lc 705; pBFE5264 NZ_CP014202.1 from L. rhamnosus BFE5264
Bearing in mind the unusually wide range of carbohydrates assimilated by IBB3423, we searched its genome factors involved in sugar metabolism. As expected, proteins assigned to the carbohydrate transport and metabolism category were many and constituted over 8% of all predicted proteins (Supplementary Table S1). The majority of carbohydrate transporters found in bacteria represents phosphoenolpyruvate (PEP)-dependent phosphotransferase systems (PTSs). Thirty-nine genes encoding PTS permeases from all seven recognized families, with various sugar substrate specificities are present in the IBB3423 genome. In addition, there are several transporters from the ATP-binding cassette (ABC) family and secondary sugar transporters from the major facilitator superfamily (MFS) (Supplementary Table S3). Many of these genes lie close genes encoding a glycosidase and transcriptional regulator, allowing hydrolysis of the incoming sugar and local transcriptional control. More than 40 glycosidases found include among others, 6-phospho-β-glucosidase (six genes), β-glucosidase (one gene), 6-phospho-β-galactosidase (one gene), β-galactosidase (two genes), α-glucosidase (four genes), α-galactosidase (three genes), α-fucosidase (two genes), and neopullulanase (two genes). Notably, the LCAKO_0451 glycosidase with a predicted β-fructosidase specificity contains a canonical LPXTG signal sequence, which suggests its extracytoplasmic localization. We posit that this enzyme could be important due to the strain’s ability to utilize the polyfructan inulin. Due to its size, inulin cannot enter the bacterial cell and has to be fragmented by an extracellular activity first to be used as a carbon source. Other genes encoding various carbohydrate-modifying enzymes from several CAZy “carbohydrate-active enzymes” families comprised numerous glycosyltransferases (39 genes), carbohydrate esterases (16 genes), carbohydrate-binding modules (5 genes), and one polysaccharide lyase. Additionally, members of the predicted carbon catabolite regulation network were identified, namely catabolite control protein A (CcpA; LCAKO_0880), phosphoryl transfer enzyme I (EI) (LCAKO_1965), HPr (LCAKO_1964), and HPr kinase/phosphorylase (LCAKO_1110).
The capacity to metabolize complex carbohydrates such as inulin is reflected in the IBB3423 genome. A putative operon involved in inulin degradation homologous to the fos cluster from other L. paracasei strains was identified (Goh et al. 2006). The fosRABCDXE operon of IBB3423 comprises genes encoding a transcriptional regulator FosR (LCAKO_0445), components of a putative fructose/mannose-specific PTS (LCAKO_0446–LCAKO_0450), and a β-fructosidase precursor FosE (LCAKO_0451). An intact glycogen metabolic pathway encoded by the glgBCDAP-amyB genes (LCAKO_2211 to LCAKO_2216) organized identically to the corresponding operons of other Lactobacillus species (Goh and Klaenhammer 2014) was identified in IBB3423.
In addition to the above, 15 common carbohydrate utilization pathways were predicted in the IBB3423 genome including glycolysis/gluconeogenesis, the citrate cycle, the pentose phosphate pathway, fructose, mannose, galactose, ascorbate, glucuronate, aldarate, sucrose, amino sugars and nucleotide sugars, pyruvate, glyoxylate and dicarboxylate, propanoate, butanoate metabolism, C5-branched dibasic acids, and inositol phosphates. In addition to the gene complement for glycolysis, six lactate dehydrogenase genes (both d- and l-) for the conversion of pyruvate into d- and l-lactate were identified.
The L. paracasei subsp. paracasei IBB3423 genome also contains a number of genes potentially involved in the biosynthesis of exopolysaccharides (Schmid et al. 2015) such as two of 28 kb gene clusters (LCAKO_2172 to LCAKO_2293) and 20 kb (LCAKO_2225 to LCAKO_2244).
Adhesion properties of strain IBB3423
To determine the adhesive properties of IBB3423, we performed a series assays using L. rhamnosus GG and L. paracasei LOCK 0919 as positive references (Aleksandrzak-Piekarczyk et al. 2016; Segers and Lebeer 2014) and L. rhamnosus LOCK 0908 as negative reference (Koryszewska-Baginska et al. 2014). To verify the role of the spaCBA-srtC gene cluster in adhesion, the IBB3423 strain was cured of the pLCAKO.2 plasmid and the adhesion properties of the resulting L. paracasei subsp. paracasei IBB3423 ΔpLCAKO.2 strain were compared with those of the parental IBB3423 and other control strains.
The hydrophobicity of the five strains testes varied significantly. IBB3423 showed the highest value (17.4% ± 6.8%), while the same strain after removing SpaCBA pili-encoding plasmid (L. paracasei subsp. paracasei IBB3423 ΔpLCAKO.2) was almost non-hydrophobic (0.1% ± 0.02%) (Table 2) (P < 0.05). The hydrophobicity of GG and LOCK 0908 was similar (3–4%) but significantly lower than that IBB3423 while, rather unexpectedly, strain 0919 was at least hydrophobic (ca. 1.5%) (Table 2). In contrast, the aggregation percentage after 24 h for all the strains except L. rhamnosus LOCK 0908, which was the least aggregative, was comparable (more than 80%) (Table 2).
Also the adherence of the bacteria tested to biotic and abiotic surfaces varied, and again, IBB3423 demonstrated the highest adherence ability to all surfaces (P < 0.05; Fig. 5). It adhered strongly to glass and gelatine and was medium adhesive to polystyrene, collagen, and mucus. The same strain (IBB3423 ΔpLCAKO.2) after removing its plasmid was significantly less adhesive demonstrating weak adhesion to all surfaces and non-adherence to gelatine. LOCK 0919 and GG showed strong, medium, or weak adhesive properties, depending on the surface, but always adhered better than the negative reference LOCK 0908 (Fig. 5).
To extend those characteristics to an in vivo situation more relevant to the bacteria–host interactions, we used epithelial Caco-2 cells as a model. L. paracasei subsp. paracasei IBB3423 adhered strongly to Caco-2 cells with an adherence rate of 97.5% ± 1.3%; LOCK 0919 and GG showed a similar lower rate, while L. rhamnosus LOCK 0908 displayed much weaker adherence (76.9% ± 10.9%) as shown in Fig. 6, which confirmed the earlier data (Aleksandrzak-Piekarczyk et al. 2016). Also, the elimination of pLCAKO.2 diminished adhesiveness to Caco-2 cells (adherence rate 92.5% ± 1.8%). The high standard deviation for that strain can be explained by the production and excretion of an amorphous, loose, slime-like substance strongly affecting the cell surface characteristics and probably disturbing its adherence (Nowak et al. 2016). Examples of microscopic pictures of adherence of Lactobacillus spp. strains to Caco-2 monolayer, confirming stronger adhesive properties of L. paracasei subsp. paracasei IBB3423 to Caco-2 cells than L. paracasei subsp. paracasei IBB3423 ΔpLCAKO.2, are presented in Fig. 7.
Adherence of various Lactobacillus strains to Caco-2 cells. aL. paracasei subsp. paracasei IBB3423, bL. paracasei subsp. paracasei IBB3423 ΔpLCAKO.2, cL. paracasei LOCK 0919, dL. rhamnosus LOCK 0908, and e Caco-2 cells (control). Representative microphotographs after staining with 0.1% crystal violet are shown
Genetic determinants of the adhesion capacity of IBB3423 strain
Cell surface proteins play a critical role in the molecular interactions between bacteria and their host. To understand the reasons for the high adhesive abilities of IBB3423, we searched its genome for factors that could be involved in adhesion to mucus, ECM, or epithelial cells. We identified 54 proteins encoded in the chromosome and five in the pLCAKO.2 plasmid potentially involved in specific adherence mechanisms (Supplementary Table S4), including 22 LPXTG-containing proteins, among which 20 also had a signal sequence for secretion. Among the LPXTG-containing proteins were pilin proteins SpaCBA (chromosomally encoded LCAKO_0529, LCAKO_0530 and LCAKO_0531, and plasmidically encoded pLCAKO.2_51, pLCAKO.2_52, and pLCAKO.2_53) and SpaDEF (LCAKO_2550, LCAKO_2551, and LCAKO_2552), all associated with a dedicated sortase. The SpaCBA proteins encoded by pLCAKO.2 plasmid shared 94%, 88%, and 95% amino acid identity, respectively, with their chromosomally located counterparts. We also detected eight proteins with diverse of collagen-binding domains such as Cna_B, collagenBindB, and collagen_bind and identified six proteins with one or three putative mucin-binding domains (MucBP). Other potential mucin-interacting proteins (LCAKO_0451, LCAKO_0596, and LCAKO_0985) contain BID_2 and Big_3 Ig-like domains. Furthermore, LCAKO_2606, LCAKO_2612, and LCAKO_2619 contain ECM-adhesive ZnuA domains, and LCAKO_1644, a fibronectin-binding protein A domain (FbpA).
We also identified a large surface-associated LCAKO_0110 protein of 3254 amino acids, with repeats of serine, alanine, and aspartic acid (Supplementary Table S4) encoded in a region carrying three glycosyltransferase genes. It has been suggested that glycosyltransferase carry out O-linked glycosylation of cell surface proteins on serine residues, thus creating a mucin-like structure (Tettelin 2001).
Comparative genome analysis of L. paracasei subsp. paracasei IBB3423
To further characterize IBB3423, we compared its genome with publicly available genomes of L. casei/paracasei strains. Complete genomes of five L. casei and 20 L. paracasei strains were found in the GenBank database, ranging in size between 2.9 and 3.2 Mbp, with a GC content of 46.2–47.9%, and predicted to encode 2600–3300 proteins (Supplementary Table S5). The L. paracasei subsp. paracasei IBB3423 genome falls within these values and shows > 98% identity with most of these species. The lowest degree of identity (< 80%) was with of L. casei LC5 and L. casei subsp. casei ATCC 393, while the highest (99.7%) with L. paracasei subsp. paracasei TMW 1.1434.
A detailed analysis revealed several local differences separating IBB3423 from all the other strains investigated. Thus, the genomes of four L. paracasei strains (IIA, KL1, N115, and Lpc10) contained inversions of the long, respective part of the IBB3423 chromosome (Fig. 8). In turn, IBB3423 differed from other strains in gene order in three separate regions (Fig. 8). Another unique feature of IBB3423 is the presence of multiple phage genes scattered across the genome. Several genes of IBB3423 are absent from the genomes from the L. casei group; they encode a cell surface protein containing the KxYKxGKxW signal peptide and MucBP and legume-lectin domains (LCAKO_0636), a transcription regulator (LCAKO_0637), an ATP-dependent endonuclease (LCAKO_1429), an AlwI family type II restriction endonuclease (LCAKO_1430), a methyl-directed repair DNA adenine methylase (LCAKO_1431), the bipolar DNA helicase HerA (LCAKO_2128), and a hypothetical protein (LCAKO_2129). Other IBB3423 genes are even not found in any other Lactobacillus spp., such as LCAKO_2843 and LCAKO_2938, LCAKO_3065 encoding hypothetical proteins, and LCAKO_3181 encoding an FRG domain-containing protein.
Discussion
Lactobacillus casei/paracasei strains can be found in various environments and this wide range explains the broad spectrum of their applications in dairy production, biotechnological, and health-related fields (Widyastuti and Rohmatussolihat 2014; Cai et al. 2009; Toh et al. 2013; Douillard et al. 2013; Smokvina et al. 2013). Owing to this potential, they are among the best-explored lactobacilli and new strains from the L. casei taxonomic group with novel functional properties are of great interest to both basic science and the biotech industry. In the present study, we isolated from raw cow milk and characterized a novel strain of attractive properties and named it IBB3423. It was recognized as L. paracasei based on its 16S rDNA partial sequence (Lane 1991) following current taxonomic guidelines (Judicial Commission of the International Committee on Systematics of Bacteria 2008). Whole genome sequencing confirmed the species identity and indicated the subspecies as paracasei. The strain showed desirable characteristics as a potential probiotic strain mainly due to its high adhesion capacity. Naturally, before this strain can get the probiotic status, there is a need to perform other proper tests confirming its probiotic properties and ability to survive in harsh conditions in the digestive tract of animals. According to the FAO/WHO guidelines (FAO/WHO 2002), these tests should include in vitro assays of, e.g., resistance to bile salts and gastric acidity, antimicrobial activity against pathogenic bacteria, the presence of transferable antibiotic resistance genes, and assessment of safety. It is also recommended to validate in vivo probiotic properties of the bacterial strain and to substantiate of health effects in the target host (FAO/WHO 2002).
To date, genomic sequences of 132 L. casei and L. paracasei strains are publicly available, 25 of which are complete (http://www.ncbi.nlm.nih.gov, last accessed in October 2018). The IBB3423 strain shows genomic features, not unlike those of other L. casei/paracasei strains as concerns the GC content (46.3%) and genome size. Notably, its chromosome of ca. 3 Mbp is the second largest among L. casei/paracasei. Also, the presence of two plasmids and their size (5986 bp and 51,211 bp) are typical for this group of bacteria. Previous studies have reported that strains of L. paracasei harbor up to four (Desmond et al. 2005) or perhaps even six plasmids (Smokvina et al. 2013) of different sizes. However, a recently published article claimed that the L. paracasei DPC2071 strain may harbor up to eleven plasmids (Stefanovic and McAuliffe 2018). These mobile replicons often carry genes conferring competitive advantage for the bacterium or critical for the industrial application such as those encoding production of bacteriocins, resistance to antibiotics, heavy metals, and phages or enabling utilization of lactose (Wang and Lee 1997). However, among the plasmid-borne genes of L. paracasei, only some have a known function (Smokvina et al. 2013). Also in the present study, many of the predicted proteins encoded by pLCAKO.1 and pLCAKO.2 plasmids were annotated as hypothetical (90% and 45%, respectively). Among those of known function are the pLCAKO.2 genes for lactose metabolism and pilus formation, both features relevant to the possible application of IBB3423. Interestingly, in contrast to pLCAKO.2, the smaller pLCAKO.1 plasmid encodes mainly proteins whose homologs are onlyfound in other genera, such as Staphylococcus, Pseudomonas, Enterococcus, and Streptococcus, indicating its possible horizontal transfer from distantly related bacteria.
The genomic relatedness between IBB3423 and other L. casei/paracasei strains is the strongest (99.7% identity) with the commercial strain L. paracasei subsp. paracasei TMW 1.1434. This strain is isogenic with L. paracasei subsp. paracasei F19 (Schott et al. 2016), a known probiotic strain able to bind gastric human and bovine mucin, collagens I and III, and fibronectin (Di Cerbo and Palmieri 2013). However, an alignment of the two genome sequences (Fig. 8) shows some unique regions in IBB3423. They contain mainly genes for phage-related proteins and hypothetical proteins, but also some of the known predicted functions such as cystathionine gamma-lyase, ABC-type amino acid transporter, type I restriction-modification system subunit R and M, or subunit M of type III R/M system. The overall high identity between IBB3423 and TMW 1.1434 explains to some extent their similar high adhesiveness, although they are of different origins: dairy and gastrointestinal, respectively. One should bear in mind, however, that the isolation source of a strain needs not to be the same as the original niche in which it has evolved, as strains can change habitats due to their adaptability (Ceapa et al. 2015).
The ability to utilize carbohydrates requires the presence of specific transporters for sugar uptake and functional metabolic pathways for its catabolism. In L. paracasei subsp. paracasei IBB3423, proteins from the functional category of carbohydrate transport and metabolism are the most abundant. This strain encodes enzymes of the Embden–Meyerhof–Parnas (EMP) and phosphoketolase pathways for homolactic and heterolactic lactic acid production from hexoses (Kandler 1983), as well as oxidative and non-oxidative branches of the pentose phosphate pathway (PP) (Tanaka et al. 2002) for utilization of pentoses. However, the non-oxidative PP pathway seems to lack a transaldolase-encoding gene. Regarding the conversion of pyruvate to lactate, at least six d- and l-lactate dehydrogenases are encoded in the IBB3423 genome. The complete Leloir pathway responsible for galactose utilization is also present. In bacteria, carbohydrate uptake is mediated by different classes of transporters, including ABC transporters, secondary transporters (permeases), and PTS transporters (Saier 2000). The PTS, involved in the uptake of mono- and oligosaccharides, is the primary sugar transport system in many lactobacilli (Lorca et al. 2007), but ABC transporters are also important for the transport of oligosaccharides (Monedero et al. 2007). The IBB3423 strain harbors a vast assortment of transporters—at least 39 PTS permeases, several ABC transporter family permeases, and other sugar transporters from the MFS superfamily. Besides these transporters for carbohydrate uptake, the IBB3423 genome encodes numerous carbohydrate-modifying enzymes, which reflects the strain’s sugar fermentation versatility including its unique ability to metabolize inulin, d-adonitol, and l-sorbose. Indeed, all the genes required for sorbose utilization, i.e., those encoding l-sorbose-phosphate-reductase and its transcriptional regulator, sorbitol-6-phosphate dehydrogenase, four components of a sorbose-specific PTS, and fructose-bisphosphate aldolase (Yebra et al. 2000) are present in the IBB3423 genome. Eleven genes for d-adonitol metabolism (LCAKO_2999 to LCAKO_3009) encode components of a mannose-type PTS, a transcriptional regulator, and six other enzymes (Bourand et al. 2013). Also, the fos operon involved in utilization of fructo-oligosaccharides (FOS), such as inulin, and the transport of free fructose (Goh et al. 2006) was identified in the IBB3423 genome. As inulin, one of the most studied prebiotics, is known to aid the development of desirable gastrointestinal microflora (Kolida et al. 2002), the ability to metabolize it is a very important property of probiotic bacteria. Among the bacteria that do metabolize prebiotic oligosaccharides are some strains of Lactobacillus and Bifidobacterium spp., and numerous in vitro and in vivo studies have shown that their growth is stimulated by FOS or other oligosaccharides. On the other hand, comparing with the control strains used by us, isolated from the human gastrointestinal tract, IBB3423 does not have the ability to ferment l-rhamnose, dulcitol, inositol, and l-fucose, probably due to their low abundance in cow milk. Interestingly, an in silico analysis has identified a complete myo-inositol (MI) utilization operon identical to that of the probiotic L. casei BL23 strain (Yebra et al. 2007), but it appears to be non-functional. Inositol, a sugar alcohol found, among other sources, in soil, is used for phosphate storage in plants, but is rarely used as an energy source by LAB (Yebra et al. 2007). So far, only some strains of L. casei have been found to metabolize MI with poor efficiency. One should note that even the presence of genes encoding enzymes for MI metabolism may not be sufficient to confer an ability to utilize MI (Zhang et al. 2010; Vinay-Lara et al. 2014). Moreover, we identified glgBCDAP-amyB genes connected with glycogen metabolism (LCAKO_2211 to LCAKO_2216) organized identically to operons of other Lactobacillus species (Goh and Klaenhammer 2014). It remains to be determined why IBB3423 cannot utilize glycogen despite having the relevant genes. Several reasons could be at play, including a lack of expression of these genes or their mutation.
Adherence to the intestinal surface is considered one of the most significant features of probiotic bacteria, as it facilitates colonization of the host and thus persistent protection against pathogens (Jankowska et al. 2008; Lim 2012). Bacteria of the genus Lactobacillus are able to adhere to various surfaces, but this feature is highly variable due to the variable presence of genes involved in adhesion. The adherence of L. paracasei subsp. paracasei IBB3423 to most surfaces is even higher than that of the best adhesive L. rhamnosus GG strain. Such high adhesiveness of a dairy strain is unexpected, as strains deriving from milk environments typically display weaker adhesion efficiency than those isolated from intestines/feces (Douillard et al. 2013). The very high adhesive capacity of IBB3423 appears to reflect the presence of as many as 59 genes encoding proteins containing putative adhesion domains, among them are SpaCBA and SpaDEF pilins encoded by chromosomal operons spaCBA and spaDEF and additional spaCBA operon on the pLCAKO.2 plasmid. All these operons had the same gene order and were associated with a gene for a pilin-specific sortase. To date, the SpaCBA proteins have been found mainly in the L. casei taxonomic group and much less frequently in other lactobacilli (Aleksandrzak-Piekarczyk et al. 2016). However, despite possessing complete pili operons, some L. casei strains do not form pili due to variations in the spaCBA sequence (Toh et al. 2013) or its transcriptional incompetence caused by a lack of functional − 35 and − 10 sequences (Aleksandrzak-Piekarczyk et al. 2016). In contrast to L. casei, in L. rhamnosus GG, the orthologous pili gene cluster is expressed and confers strong mucus-binding ability (Kankainen et al. 2009; Reunanen et al. 2012). Also the plasmidic spaCBA from L. paracasei LOCK 0919 seems to be functional and responsible for the strong adhesiveness of this strain (Aleksandrzak-Piekarczyk et al. 2016). It is postulated that insertion of an IS element in the spaC promoter region in both L. rhamnosus GG and L. paracasei LOCK 0919 allows the expression of the pili genes (Douillard et al. 2013; Aleksandrzak-Piekarczyk et al. 2016). Notably, the upstream region of the IBB3423 spaCBA operon harbored by pLCAKO.2 also contains an insertion element indicating that it likely is functional. Indeed, the removal of pLCAKO.2 led to a marked decrease of IBB3423 hydrophobicity and adhesiveness to biotic and abiotic surfaces. The presence of such a plasmid should therefore facilitate colonization and ensure longer persistence in the host’s gut thereby conferring a competitive advantage over other bacteria. It can also increase the range of inhabited environments. Plasmidic localization of the spaCBA operon, such as in IBB3423, is extremely rare albeit not unique since among all the plasmids deposited at GenBank only two, pLOCK 0919 from L. paracasei LOCK 0919 (Koryszewska-Baginska et al. 2013; Aleksandrzak-Piekarczyk et al. 2016) and p1_LC355 from L. paracasei LC355 (NZ_CP029537.1), harbor the same pilus gene cluster.
In conclusion, the newly identified dairy strain L. paracasei subsp. paracasei IBB3423 presents genomic and functional features making it a very promising candidate probiotic. Owing to its high adhesiveness comparable to that of the benchmark strain L. rhamnosus GG and the ability to use a wide range of saccharides including inulin, it should be able to compete with other commercially applied strains. However, additional functional studies in vitro and in vivo should be performed to determine whether IBB3423 can persist in the gastrointestinal tract and exhibits health-promoting action. Last but not least, the isolation of this novel bacterial strain of such interesting characteristics is proof of the productiveness of citizen science.
References
Aleksandrzak-Piekarczyk T, Koryszewska-Baginska A, Bardowski J (2013) Genome sequence of the probiotic strain Lactobacillus rhamnosus (formerly Lactobacillus casei) LOCK900. Genome Announc 1:e00640–e00613. https://doi.org/10.1128/genomeA.00640-13
Aleksandrzak-Piekarczyk T, Koryszewska-Bagińska A, Grynberg M, Nowak A, Cukrowska B, Kozakova H, Bardowski J (2016) Genomic and functional characterization of the unusual pLOCK 0919 plasmid harboring the spaCBA pili cluster in Lactobacillus casei LOCK 0919. Genome Biol Evol 8:202–217. https://doi.org/10.1093/gbe/evv247
Aleljung P, Shen W, Rozalska B, Hellman U, Ljungh A, Wadström T (1994) Purification of collagen-binding proteins of Lactobacillus reuteri NCIB 11951. Curr Microbiol 28:231–236
Almeida CC, Lorena SLS, Pavan CR, Akasaka HMI, Mesquita MA (2012) Beneficial effects of long-term consumption of a probiotic combination of Lactobacillus casei Shirota and Bifidobacterium breve Yakult may persist after suspension of therapy in lactose-intolerant patients. Nutr Clin Pract 27:247–251. https://doi.org/10.1177/0884533612440289
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Antikainen J, Anton L, Sillanpää J, Korhonen TK (2002) Domains in the S-layer protein CbsA of Lactobacillus crispatus involved in adherence to collagens, laminin and lipoteichoic acids and in self-assembly: adhesive domains in S-layer protein. Mol Microbiol 46:381–394. https://doi.org/10.1046/j.1365-2958.2002.03180.x
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST server: rapid annotations using subsystems technology. BMC Genomics 9:75. https://doi.org/10.1186/1471-2164-9-75
Balson DF, Shaw WV (1990) Nucleotide sequence of the rep gene of staphylococcal plasmid pCW7. Plasmid 24:74–80. https://doi.org/10.1016/0147-619X(90)90027-A
Barr JJ, Auro R, Furlan M, Whiteson KL, Erb ML, Pogliano J, Stotland A, Wolkowicz R, Cutting AS, Doran KS, Salamon P, Youle M, Rohwer F (2013) Bacteriophage adhering to mucus provide a non-host-derived immunity. Proc Natl Acad Sci 110:10771–10776. https://doi.org/10.1073/pnas.1305923110
Bäuerl C, Pérez-Martínez G, Yan F, Polk DB, Monedero V (2010) Functional analysis of the p40 and p75 proteins from Lactobacillus casei BL23. J Mol Microbiol Biotechnol 19:231–241. https://doi.org/10.1159/000322233
Bourand A, Yebra MJ, Boël G, Mazé A, Deutscher J (2013) Utilization of D-ribitol by Lactobacillus casei BL23 requires a mannose-type phosphotransferase system and three catabolic enzymes. J Bacteriol 195:2652–2661. https://doi.org/10.1128/JB.02276-12
Buck BL, Altermann E, Svingerud T, Klaenhammer TR (2005) Functional analysis of putative adhesion factors in Lactobacillus acidophilus NCFM. Appl Environ Microbiol 71:8344–8351. https://doi.org/10.1128/AEM.71.12.8344-8351.2005
Cai H, Thompson R, Budinich MF, Broadbent JR, Steele JL (2009) Genome sequence and comparative genome analysis of Lactobacillus casei: insights into their niche-associated evolution. Genome Biol Evol 1:239–257. https://doi.org/10.1093/gbe/evp019
Ceapa C, Lambert J, van Limpt K, Wels M, Smokvina T, Knol J, Kleerebezem M (2015) Correlation of Lactobacillus rhamnosus genotypes and carbohydrate utilization signatures determined by phenotype profiling. Appl Environ Microbiol 81:5458–5470. https://doi.org/10.1128/AEM.00851-15
Claesson MJ, van Sinderen D, O’Toole PW (2007) The genus Lactobacillus – a genomic basis for understanding its diversity. FEMS Microbiol Lett 269:22–28. https://doi.org/10.1111/j.1574-6968.2006.00596.x
Darzentas N (2010) Circoletto: visualizing sequence similarity with Circos. Bioinformatics 26:2620–2621. https://doi.org/10.1093/bioinformatics/btq484
Desmond C, Ross RP, Fitzgerald G, Stanton C (2005) Sequence analysis of the plasmid genome of the probiotic strain Lactobacillus paracasei NFBC338 which includes the plasmids pCD01 and pCD02. Plasmid 54:160–175. https://doi.org/10.1016/j.plasmid.2005.04.001
Di Cerbo A, Palmieri B (2013) Lactobacillus paracasei subsp. paracasei F19; a farmacogenomic and clinical update. Nutr Hosp 28:1842–1150. https://doi.org/10.3305/nh.2013.28.6.6831
Dintilhac A, Alloing G, Granadel C, Claverys JP (1997) Competence and virulence of Streptococcus pneumoniae: Adc and PsaA mutants exhibit a requirement for Zn and Mn resulting from inactivation of putative ABC metal permeases. Mol Microbiol 25:727–739
Douillard FP, Ribbera A, Järvinen HM, Kant R, Pietilä TE, Randazzo C, Paulin L, Laine PK, Caggia C, von Ossowski I, Reunanen J, Satokari R, Salminen S, Palva A, de Vos WM (2013) Comparative genomic and functional analysis of Lactobacillus casei and Lactobacillus rhamnosus strains marketed as probiotics. Appl Environ Microbiol 79:1923–1933. https://doi.org/10.1128/AEM.03467-12
FAO/WHO (2002) Guidelines for the evaluation of probiotics in food. Report of a joint FAO/WHO working group on drafting guidelines for the evaluation of probiotics in food. World Health Organization, London
Felis GE, Dellaglio F (2007) Taxonomy of lactobacilli and bifidobacteria. Curr Issues Intest Microbiol 8:44–61
Fuller R (1991) Probiotics in human medicine. Gut 32(4):439–442. https://doi.org/10.1136/gut.32.4.439
Gabbianelli R, Scotti R, Ammendola S, Petrarca P, Nicolini L, Battistoni A (2011) Role of ZnuABC and ZinT in Escherichia coli O157:H7 zinc acquisition and interaction with epithelial cells. BMC Microbiol 11:36. https://doi.org/10.1186/1471-2180-11-36
Goh YJ, Klaenhammer TR (2014) Insights into glycogen metabolism in Lactobacillus acidophilus: impact on carbohydrate metabolism, stress tolerance and gut retention. Microb Cell Factories 13:94. https://doi.org/10.1186/s12934-014-0094-3
Goh YJ, Zhang C, Benson AK, Schlegel V, Lee J-H, Hutkins RW (2006) Identification of a putative operon involved in fructooligosaccharide utilization by Lactobacillus paracasei. Appl Environ Microbiol 72:7518–7530. https://doi.org/10.1128/AEM.00877-06
Grant JR, Stothard P (2008) The CGView Server: a comparative genomics tool for circular genomes. Nucleic Acids Res 36:W181–W184. https://doi.org/10.1093/nar/gkn179
Grissa I, Vergnaud G, Pourcel C (2007) CRISPRFinder: a web tool to identify clustered regularly interspaced short palindromic repeats. Nucleic Acids Res 35:W52–W57. https://doi.org/10.1093/nar/gkm360
Haft DH, Selengut J, Mongodin EF, Nelson KE (2005) A guild of 45 CRISPR-associated (Cas) protein families and multiple CRISPR/Cas subtypes exist in prokaryotic genomes. PLoS Comput Biol 1:e60. https://doi.org/10.1371/journal.pcbi.0010060
Harris RS (2007) Improved pairwise alignment of genomic DNA. Ph.D. Thesis, The Pennsylvania State University.
Holzapfel WH, Haberer P, Geisen R, Björkroth J, Schillinger U (2001) Taxonomy and important features of probiotic microorganisms in food and nutrition. Am J Clin Nutr 73:365s–373s. https://doi.org/10.1093/ajcn/73.2.365s
Hynonen U, Westerlund-Wikstrom B, Palva A, Korhonen TK (2002) Identification by flagellum display of an epithelial cell- and fibronectin-binding function in the SlpA surface protein of Lactobacillus brevis. J Bacteriol 184:3360–3367. https://doi.org/10.1128/JB.184.12.3360-3367.2002
Jankowska A, Laubitz D, Antushevich H, Zabielski R, Grzesiuk E (2008) Competition of Lactobacillus paracasei with Salmonella enterica for adhesion to Caco-2 cells. J Biomed Biotechnol 2008:1–6. https://doi.org/10.1155/2008/357964
Johnson-Henry KC, Hagen KE, Gordonpour M, Tompkins TA, Sherman PM (2007) Surface-layer protein extracts from Lactobacillus helveticus inhibit enterohaemorrhagic Escherichia coli O157:H7 adhesion to epithelial cells. Cell Microbiol 9:356–367. https://doi.org/10.1111/j.1462-5822.2006.00791.x
Judicial Commission of the International Committee on Systematics of Bacteria (2008) The type strain of Lactobacillus casei is ATCC 393, ATCC 334 cannot serve as the type because it represents a different taxon, the name Lactobacillus paracasei and its subspecies names are not rejected and the revival of the name “Lactobacillus zeae” contravenes Rules 51b (1) and (2) of the International Code of Nomenclature of Bacteria. Opinion 82. Int J Syst Evol Microbiol 58:1764–1765. https://doi.org/10.1099/ijs.0.2008/005330-0
Kalliomäki M, Salminen S, Poussa T, Arvilommi H, Isolauri E (2003) Probiotics and prevention of atopic disease: 4-year follow-up of a randomised placebo-controlled trial. Lancet 361:1869–1871. https://doi.org/10.1016/S0140-6736(03)13490-3
Kandler O (1983) Carbohydrate metabolism in lactic acid bacteria. Antonie Van Leeuwenhoek 49:209–224. https://doi.org/10.1007/BF00399499
Kanehisa M, Sato Y, Morishima K (2016) BlastKOALA and GhostKOALA: KEGG tools for functional characterization of genome and metagenome sequences. J Mol Biol 428:726–731. https://doi.org/10.1016/j.jmb.2015.11.006
Kankainen M, Paulin L, Tynkkynen S, von Ossowski I, Reunanen J, Partanen P, Satokari R, Vesterlund S, Hendrickx APA, Lebeer S, De Keersmaecker SCJ, Vanderleyden J, Hamalainen T, Laukkanen S, Salovuori N, Ritari J, Alatalo E, Korpela R, Mattila-Sandholm T, Lassig A, Hatakka K, Kinnunen KT, Karjalainen H, Saxelin M, Laakso K, Surakka A, Palva A, Salusjarvi T, Auvinen P, de Vos WM (2009) Comparative genomic analysis of Lactobacillus rhamnosus GG reveals pili containing a human- mucus binding protein. Proc Natl Acad Sci 106:17193–17198. https://doi.org/10.1073/pnas.0908876106
Kant R, Blom J, Palva A, Siezen RJ, de Vos WM (2011) Comparative genomics of Lactobacillus: comparative analysis of 20 complete Lactobacillus genomes. Microb Biotechnol 4:323–332. https://doi.org/10.1111/j.1751-7915.2010.00215.x
Klaenhammer TR, Altermann E, Pfeiler E, Buck BL, Goh Y-J, O’Flaherty S, Barrangou R, Duong T (2008) Functional genomics of probiotic lactobacilli. J Clin Gastroenterol 42:S160–S162. https://doi.org/10.1097/MCG.0b013e31817da140
Kleerebezem M, Hols P, Bernard E, Rolain T, Zhou M, Siezen RJ, Bron PA (2010) The extracellular biology of the lactobacilli. FEMS Microbiol Rev 34:199–230. https://doi.org/10.1111/j.1574-6976.2009.00208.x
Kolida S, Tuohy K, Gibson GR (2002) Prebiotic effects of inulin and oligofructose. Br J Nutr 87:S193–S197. https://doi.org/10.1079/BJN/2002537
Koryszewska-Baginska A, Aleksandrzak-Piekarczyk T, Bardowski J (2013) Complete fenome sequence of the probiotic strain Lactobacillus casei (formerly Lactobacillus paracasei) LOCK919. Genome Announc 1:e00758–e00713. https://doi.org/10.1128/genomeA.00758-13
Koryszewska-Baginska A, Bardowski J, Aleksandrzak-Piekarczyk T (2014) Genome sequence of the probiotic strain Lactobacillus rhamnosus (formerly Lactobacillus casei) LOCK908. Genome Announc 2:e00120–e00114. https://doi.org/10.1128/genomeA.00120-14
Krasowska A, Sigler K (2014) How microorganisms use hydrophobicity and what does this mean for human needs? Front Cell Infect Microbiol 4:112. https://doi.org/10.3389/fcimb.2014.00112
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acids techniques in bacterial systematics. John Wiley & Sons, Chichester, pp 115–147
Lebeer S, Vanderleyden J, De Keersmaecker SCJ (2008) Genes and molecules of lactobacilli supporting probiotic action. Microbiol Mol Biol Rev 72:728–764. https://doi.org/10.1128/MMBR.00017-08
Lim S-M (2012) Factors affecting adhesion of lactic acid bacteria to Caco-2 cells and inhibitory effect on infection of Salmonella typhimurium. J Microbiol Biotechnol 22:1731–1739. https://doi.org/10.4014/jmb.1208.08049
Lorca GL, Barabote RD, Zlotopolski V, Tran C, Winnen B, Hvorup RN, Stonestrom AJ, Nguyen E, Huang L-W, Kim DS, Saier MH (2007) Transport capabilities of eleven Gram-positive bacteria: comparative genomic analyses. Biochim Biophys Acta Biomembr 1768:1342–1366. https://doi.org/10.1016/j.bbamem.2007.02.007
Makarova K, Slesarev A, Wolf Y, Sorokin A, Mirkin B, Koonin E, Pavlov A, Pavlova N, Karamychev V, Polouchine N, Shakhova V, Grigoriev I, Lou Y, Rohksar D, Lucas S, Huang K, Goodstein DM, Hawkins T, Plengvidhya V, Welker D, Hughes J, Goh Y, Benson A, Baldwin K, Lee J-H, Diaz-Muniz I, Dosti B, Smeianov V, Wechter W, Barabote R, Lorca G, Altermann E, Barrangou R, Ganesan B, Xie Y, Rawsthorne H, Tamir D, Parker C, Breidt F, Broadbent J, Hutkins R, O’Sullivan D, Steele J, Unlu G, Saier M, Klaenhammer T, Richardson P, Kozyavkin S, Weimer B, Mills D (2006) Comparative genomics of the lactic acid bacteria. Proc Natl Acad Sci 103:15611–15616. https://doi.org/10.1073/pnas.0607117103
Mandlik A, Swierczynski A, Das A, Ton-That H (2008) Pili in Gram-positive bacteria: assembly, involvement in colonization and biofilm development. Trends Microbiol 16:33–40. https://doi.org/10.1016/j.tim.2007.10.010
Marco ML, Pavan S, Kleerebezem M (2006) Towards understanding molecular modes of probiotic action. Curr Opin Biotechnol 17:204–210. https://doi.org/10.1016/j.copbio.2006.02.005
Mercanti DJ, Rousseau GM, Capra ML, Quiberoni A, Tremblay DM, Labrie SJ, Moineau S (2016) Genomic diversity of phages infecting probiotic strains of Lactobacillus paracasei. Appl Environ Microbiol 82:95–105. https://doi.org/10.1128/AEM.02723-15
Miyoshi Y, Okada S, Uchimura T, Satoh E (2006) A mucus adhesion promoting protein, MapA, mediates the adhesion of Lactobacillus reuteri to Caco-2 human intestinal epithelial cells. Biosci Biotechnol Biochem 70:1622–1628. https://doi.org/10.1271/bbb.50688
Monedero V, Mazé A, Boël G, Zúñiga M, Beaufils S, Hartke A, Deutscher J (2007) The phosphotransferase system of Lactobacillus casei: regulation of carbon metabolism and connection to cold shock response. J Mol Microbiol Biotechnol 12:20–32. https://doi.org/10.1159/000096456
Muñoz-Provencio D, Pérez-Martínez G, Monedero V (2010) Characterization of a fibronectin-binding protein from Lactobacillus casei BL23. J Appl Microbiol 108:1050–1059. https://doi.org/10.1111/j.1365-2672.2009.04508.x
Navarre WW, Schneewind O (1999) Surface proteins of Gram-positive bacteria and mechanisms of their targeting to the cell wall envelope. Microbiol Mol Biol Rev MMBR 63:174–229
Nowak A, Motyl I, Śliżewska K, Libudzisz Z, Klewicka E (2016) Adherence of probiotic bacteria to human colon epithelial cells and inhibitory effect against enteric pathogens. Int J Dairy Technol 69:532–539. https://doi.org/10.1111/1471-0307.12286
Ouwehand AC, Salminen S, Isolauri E (2002) Probiotics: an overview of beneficial effects. Antonie Van Leeuwenhoek 82:279–289
Ouwehand AC, Tölkkö S, Kulmala J, Salminen S, Salminen E (2000) Adhesion of inactivated probiotic strains to intestinal mucus: adhesion of inactivated probiotics. Lett Appl Microbiol 31:82–86. https://doi.org/10.1046/j.1472-765x.2000.00773.x
Pérez PF, Minnaard Y, Disalvo EA, De Antoni GL (1998) Surface properties of bifidobacterial strains of human origin. Appl Environ Microbiol 64:21–26
Pretzer G, Snel J, Molenaar D, Wiersma A, Bron PA, Lambert J, de Vos WM, van der Meer R, Smits MA, Kleerebezem M (2005) Biodiversity-based identification and functional characterization of the mannose-specific adhesin of Lactobacillus plantarum. J Bacteriol 187:6128–6136. https://doi.org/10.1128/JB.187.17.6128-6136.2005
Reunanen J, von Ossowski I, Hendrickx APA, Palva A, de Vos WM (2012) Characterization of the SpaCBA pilus fibers in the probiotic Lactobacillus rhamnosus GG. Appl Environ Microbiol 78:2337–2344. https://doi.org/10.1128/AEM.07047-11
Saier MH (2000) Families of transmembrane sugar transport proteins. Mol Microbiol 35:699–710. https://doi.org/10.1046/j.1365-2958.2000.01759.x
Saier MH, Reddy VS, Tsu BV, Ahmed MS, Li C, Moreno-Hagelsieb G (2016) The transporter classification database (TCDB): recent advances. Nucleic Acids Res 44:D372–D379. https://doi.org/10.1093/nar/gkv1103
Schott A-S, Behr J, Quinn J, Vogel RF (2016) MALDI-TOF mass spectrometry enables a comprehensive and fast analysis of dynamics and qualities of stress responses of Lactobacillus paracasei subsp. paracasei F19. PLoS One 11:e0165504. https://doi.org/10.1371/journal.pone.0165504
Segers ME, Lebeer S (2014) Towards a better understanding of Lactobacillus rhamnosus GG - host interactions. Microb Cell Factories 13:S7. https://doi.org/10.1186/1475-2859-13-S1-S7
Smokvina T, Wels M, Polka J, Chervaux C, Brisse S, Boekhorst J, van Hylckama Vlieg JET, Siezen RJ (2013) Lactobacillus paracasei comparative genomics: towards species pan-genome definition and exploitation of diversity. PLoS One 8:e68731. https://doi.org/10.1371/journal.pone.0068731
Stefanovic E, McAuliffe O (2018) Comparative genomic and metabolic analysis of three Lactobacillus paracasei cheese isolates reveals considerable genomic differences in strains from the same niche. BMC Genomics 19:205. https://doi.org/10.1186/s12864-018-4586-0
Stiles ME, Holzapfel WH (1997) Lactic acid bacteria of foods and their current taxonomy. Int J Food Microbiol 36:1–29. https://doi.org/10.1016/S0168-1605(96)01233-0
Styriak I, Nemcova R, Chang Y-H, Ljungh A (2003) Binding of extracellular matrix molecules by probiotic bacteria. Lett Appl Microbiol 37:329–333. https://doi.org/10.1046/j.1472-765X.2003.01402.x
Sun Z, Harris HMB, McCann A, Guo C, Argimón S, Zhang W, Yang X, Jeffery IB, Cooney JC, Kagawa TF, Liu W, Song Y, Salvetti E, Wrobel A, Rasinkangas P, Parkhill J, Rea MC, O’Sullivan O, Ritari J, Douillard FP, Paul Ross R, Yang R, Briner AE, Felis GE, de Vos WM, Barrangou R, Klaenhammer TR, Caufield PW, Cui Y, Zhang H, O’Toole PW (2015) Expanding the biotechnology potential of lactobacilli through comparative genomics of 213 strains and associated genera. Nat Commun 6:8322. https://doi.org/10.1038/ncomms9322
Sykora J, Valeckov AK, Amlerova J, Siala K, Dedek P, Watkins S, Varvarovska J, Stozicky F, Pazdiora P, Schwarz J (2005) Effects of a specially designed fermented milk product containing probiotic Lactobacillus casei DN-114 001 and the eradication of H. pylori in children: a prospective randomized double-blind study. J Clin Gastroenterol 39:692–698. https://doi.org/10.1097/01.mcg.0000173855.77191.44
Tanaka K, Komiyama A, Sonomoto K, Ishizaki A, Hall S, Stanbury P (2002) Two different pathways for D -xylose metabolism and the effect of xylose concentration on the yield coefficient of L -lactate in mixed-acid fermentation by the lactic acid bacterium Lactococcus lactis IO-1. Appl Microbiol Biotechnol 60:160–167. https://doi.org/10.1007/s00253-002-1078-5
Tanaka T, Ogura M (1998) A novel Bacillus natto plasmid pLS32 capable of replication in Bacillus subtilis. FEBS Lett 422:243–246. https://doi.org/10.1016/S0014-5793(98)00015-5
Tettelin H (2001) Complete genome sequence of a virulent isolate of Streptococcus pneumoniae. Science 293:498–506. https://doi.org/10.1126/science.1061217
Toba T, Virkola R, Westerlund B, Bjorkman Y, Sillanpaa J, Vartio T, Kalkkinen N, Korhonen TK (1995) A collagen-binding S-layer protein in Lactobacillus crispatus. Appl Environ Microbiol 61:2467–2471
Toh H, Oshima K, Nakano A, Takahata M, Murakami M, Takaki T, Nishiyama H, Igimi S, Hattori M, Morita H (2013) Genomic adaptation of the Lactobacillus casei group. PLoS One 8:e75073. https://doi.org/10.1371/journal.pone.0075073
Turpin W, Humblot C, Noordine M-L, Thomas M, Guyot J-P (2012) Lactobacillaceae and cell adhesion: genomic and functional screening. PLoS One 7:e38034. https://doi.org/10.1371/journal.pone.0038034
Turroni F, Serafini F, Foroni E, Duranti S, O’Connell Motherway M, Taverniti V, Mangifesta M, Milani C, Viappiani A, Roversi T, Sanchez B, Santoni A, Gioiosa L, Ferrarini A, Delledonne M, Margolles A, Piazza L, Palanza P, Bolchi A, Guglielmetti S, van Sinderen D, Ventura M (2013) Role of sortase-dependent pili of Bifidobacterium bifidum PRL2010 in modulating bacterium-host interactions. Proc Natl Acad Sci 110:11151–11156. https://doi.org/10.1073/pnas.1303897110
Van Domselaar GH, Stothard P, Shrivastava S, Cruz JA, Guo A, Dong X, Lu P, Szafron D, Greiner R, Wishart DS (2005) BASys: a web server for automated bacterial genome annotation. Nucleic Acids Res 33:W455–W459. https://doi.org/10.1093/nar/gki593
Velez MP, De Keersmaecker SCJ, Vanderleyden J (2007) Adherence factors of Lactobacillus in the human gastrointestinal tract. FEMS Microbiol Lett 276:140–148. https://doi.org/10.1111/j.1574-6968.2007.00908.x
Ventura M, O’Flaherty S, Claesson MJ, Turroni F, Klaenhammer TR, van Sinderen D, O’Toole PW (2009) Genome-scale analyses of health-promoting bacteria: probiogenomics. Nat Rev Microbiol 7:61–71. https://doi.org/10.1038/nrmicro2047
Vinay-Lara E, Hamilton JJ, Stahl B, Broadbent JR, Reed JL, Steele JL (2014) Genome –scale reconstruction of metabolic networks of Lactobacillus casei ATCC 334 and 12A. PLoS One 9:e110785. https://doi.org/10.1371/journal.pone.0110785
Visweswaran GRR, Leenhouts K, van Roosmalen M, Kok J, Buist G (2014) Exploiting the peptidoglycan-binding motif, LysM, for medical and industrial applications. Appl Microbiol Biotechnol. https://doi.org/10.1007/s00253-014-5633-7
Wang T-T, Lee BH (1997) Plasmids in Lactobacillus. Crit Rev Biotechnol 17:227–272. https://doi.org/10.3109/07388559709146615
Widyastuti Y, Rohmatussolihat FA (2014) The role of lactic acid bacteria in milk fermentation. Food Nutr Sci 05:435–442. https://doi.org/10.4236/fns.2014.54051
Yebra MJ, Veyrat A, Santos MA, Perez-Martinez G (2000) Genetics of L-sorbose transport and metabolism in Lactobacillus casei. J Bacteriol 182:155–163. https://doi.org/10.1128/JB.182.1.155-163.2000
Yebra MJ, Zuniga M, Beaufils S, Perez-Martinez G, Deutscher J, Monedero V (2007) Identification of a gene cluster enabling Lactobacillus casei BL23 to utilize myo-inositol. Appl Environ Microbiol 73:3850–3858. https://doi.org/10.1128/AEM.00243-07
Yin Y, Mao X, Yang J, Chen X, Mao F, Xu Y (2012) dbCAN: a web resource for automated carbohydrate-active enzyme annotation. Nucleic Acids Res 40:W445–W451. https://doi.org/10.1093/nar/gks479
Yoon S-H, Ha S, Lim J, Kwon S, Chun J (2017) A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Van Leeuwenhoek 110:1281–1286. https://doi.org/10.1007/s10482-017-0844-4
Zhang WY, Sun ZH, Yu DL, Airideng C, Chen W, Meng H, Zhang HP (2010) Comparative analysis of iol clusters in Lactobacillus casei strains. World J Microbiol Biotechnol 26:1949–1955. https://doi.org/10.1007/s11274-010-0375-x
Zhou Y, Liang Y, Lynch KH, Dennis JJ, Wishart DS (2011) PHAST: a fast phage search tool. Nucleic Acids Res 39:W347–W352. https://doi.org/10.1093/nar/gkr485
Funding
This study was funded by grant no. 2018/29/B/NZ9/02278 from the National Science Centre and MakeTogether program founded by the Ministry of Science and Higher Education “Paths of Copernicus” (Supplementary Information S6).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Marcin Grynberg, corresponding author only for citizen science matters.
Electronic supplementary material
ESM 1
(PDF 629 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Koryszewska-Bagińska, A., Gawor, J., Nowak, A. et al. Comparative genomics and functional analysis of a highly adhesive dairy Lactobacillus paracasei subsp. paracasei IBB3423 strain. Appl Microbiol Biotechnol 103, 7617–7634 (2019). https://doi.org/10.1007/s00253-019-10010-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-10010-1