Abstract
Efficient selection and production of antibody fragments in microbial systems remain to be a challenging process. To optimize microbial production of single-chain variable fragments (scFvs), we have chosen five model targets, 1) a hapten, Zearalenone (ZEN) mycotoxin, along with infectious agents 2) rabies virus, 3) Propionibacterium acnes, 4) Pseudomonas aeruginosa, and a cancer cell 5) acute myeloid leukemia cell line (HL-60). The scFv binders were affinity selected from a non-immunized human phage display scFv antibody library and genetically fused to the N-terminus of emerald green fluorescent protein (EmGFP). The scFv-EmGFP fusion constructs were subcloned into an expression vector, under the control of T7 promoter, C-terminally tagged with hexa-histidine and expressed in different Escherichia coli (E. coli) hosts. This enabled the detection of cells that expressed the correct scFv-EmGFP fusion, termed fluorobody, via bright fluorescent signal in the cytoplasm. Among the three E. coli hosts tested, an engineered E. coli B strain called SHuffle B that promotes disulfide bond formation in the cytoplasm appeared to be the most appropriate host. The recombinant fluorobodies were well expressed (2–8 mg/L), possessed the fluorescence property of EmGFP, and retained the ability to bind to their cognate targets. Their specific bindings were demonstrated by ELISA, fluorescence-linked immunosorbent assay (FLISA), flow cytometry, and fluorescent microscope imaging. The fluorobody expression platform in this study could be further adopted as a one-step immunostaining technique based on scFv, isolated from phage display library to numerous desired targets.
Key points
• E. coli SHuffle express T7 is a suitable expression host for scFv-EmGFP (fluorobody)
• Only the clones harboring scFv-EmGFP plasmid will show bright fluorescent signal
• This platform can be used to produce fluorobodies for numerous purposes
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Various types of antibody (Ab) fragments have been engineered for therapeutic and diagnostic purposes (Dübel 2007). The two common forms of small Ab fragments are single-chain variable-fragment (scFv) and single-domain Ab (sdAb). The scFv consists of light and heavy chain variable domains of immunoglobulin of mammals, linked with peptide linker (G4S)3. The average size of scFv is about 27 kDa and contains 2 predicted disulfide bonds (Ferrè and Clote 2005a). Phage display Ab technology is the most popular method for the generation of recombinant antibody (rAb) in the form of scFv or Fab. The key advantage of this technology relies on the simplicity but powerful affinity selection (bio-panning) procedure, and the direct linkage between the displayed Ab and its encoding gene within the phage genome (Kehoe and Kay 2005). Ab against desired target can be identified directly from diverse repertoires of Ab genes, generating high-affinity binding sites without the constraints imposed by classical method for generating either polyclonal or monoclonal Ab (McCafferty et al. 1990), such as high costs, stability of hybridoma cells, or batch-to-batch variation. Moreover, this method allows further engineering of the antibody into various formats to suit desired applications (Rangnoi et al. 2021), such as improved binding to FcγRI receptor (Robinson et al. 2015). Since the structure of the binding domain of the Ab is not glycosylated; contrary to therapeutic Ab which requires functional glycosylated constant domain (Fc), rAb can be expressed efficiently using expression systems which lack glycosylation machinery (Frenzel et al. 2013). This research focused on scFv format because it is small and can be obtained directly from biopanning of phage display scFv library. One of the common expression system for the production of scFvs especially for research purposes is Escherichia coli (E. coli), which is also a host for bacteriophage M13 (Spadiut et al. 2014). As expected, the expression level of various scFvs varied greatly, depending on different clones, despite a similar (65% identity) domain framework (Min and Yamabhai 2021).
For high sensitivity detection of biomolecules and the analysis of their interactions, Ab labeling with fluorescent probes, especially, fluorescein isothiocyanate (FITC) is one of the most widely used organic fluorophores in various applications (Hermanson 2008; Holmes and Lantz 2001). FITC can react with the free amino group of folded proteins, leading to stable thiourea bond and fluorescent Abs. However, the FITC may conjugate with the antigen-binding site of Abs, resulting in a partial or complete loss of reactivity of the Ab to its antigen (Sakamoto et al. 2012). To overcome these drawbacks, fusion of an Aequorea victoria green fluorescent protein (GFP) (Tsien 1998) with an scFv fragment, termed fluorobody, has been developed by genetic fusion of scFv to GFP. In this system, a direct conjugation of an scFv with a single GFP molecule (1:1 ratio) via a covalent bonding can improve the accuracy of quantitative analysis (Schwalbach et al. 2000). However, despite the use of fluorobodies as bio-probes in various detection formats, such as immunolabeling (Schwalbach et al. 2000), flow cytometry (Petrausch et al. 2007), fluorescent-linked immunosorbent assay (FLISA) (Oelschlaeger et al. 2002), the folding efficacy and the yield were generally poor (e.g., for fluorobody against s-triazine, 90% of the fusions were insoluble and only 25% could be refolded in E. coli BL21(DE3) pLysS) (Oelschlaeger et al. 2002). The yields of fluorobody against bacterial lipopolysaccharide (Griep et al. 1999) or hepatitis B surface antigen (HepBsAg) (Casey et al. 2000a) was ~ 100 to 200 µg/L in E. coli XL1-Blue-MRFX' Kan and Sure cells, respectively. This is likely because scFv has internal disulfide bonds (Ferrè and Clote 2005a), which require an environment containing enzymes that can promote the formation of correct bond formation between two cysteine molecules, such as the periplasm of E. coli. This requires the optimization of secretion of the scFv into the periplasm, where GFP is known not to fold (Ke et al. 2016). Further, the over-expression of protein in the periplasm to high yields in E. coli is known to be difficult to achieve (Karyolaimos and de Gier 2021; Olichon and Surrey 2007). To overcome these obstacles, E. coli SHuffle strains, which has been genetically engineered to promote correct disulfide bond formation in the cytoplasm (Lobstein et al. 2012), was selected as hosts for the expression of fluorobodies in this study. EmGFP (emerald GFP), which is a 27 kDa protein, derived from wildtype GFP (Ilagan et al. 2010) by introducing mutations (S65T, S72A, N149K, M153T, I167T), exhibits improved folding at 37 °C, is highly detectable, and has excitation maximum at 487 nm and emission maximum at 509 nm (Cubitt et al. 1999), was used as a fluorophore-fusion in this study. A suitable expression vector to produce scFv-EmGFP fusion or scFv-EmGFP fluorobody was also developed. The fluorobodies against diverse targets; namely, virus, mycotoxin, cancer cells and bacteria, were created and used as representative bio-detection probes with a myriad of applications in molecular biology. Finally, binding characteristic and their applications for quantitative and qualitative analysis in different assay formats were demonstrated.
Materials and methods
Materials, cell line, and bacterial strains
The pSERT/EmGFP vector containing EmGFP gene was purchased from Thermo Fisher Scientific (cat. #V35320, USA). Standard Zearalenone (ZEN) (cat. #CH-01S5) and ZEN conjugated with Bovine albumin serum; BSA (cat. #CJ-01-BSA) were prepared from Fusarium species (Aokin, Germany). Inactivated rabies virus (purified chick embryo cell vaccine, PCEC, LEP-Flury strain) was obtained from Rabipur (batch #2605, Chiron, India). Propionibacterium acnes DMST 14916 and Pseudomonas aeruginosa DMST 37186 were kindly provided by Dr. Griangsak Eumkeb, School of Sciences, Suranaree University of Technology, Thailand. E. coli SHuffle express T7 (E. coli SHuffle B, cat. #C3029, NEB, USA), SHuffle Express (E. coli SHuffle K-12, cat. #C3026, NEB, USA) and BL21 (DE3) (cat. #C2527, NEB, USA) strains were used to express fluorobodies. HL-60 cells were purchased from ATCC (cat. #CCL-240, Virginia, USA) and maintained with Iscove's Modified Dulbecco’s Medium (IMDM, cat. #12200036), supplemented with 20%FBS (cat. #10270106) and 1 × penicillin streptomycin (cat. #15140122). The cell culture media and supplements were purchased from Gibco (Eugene, OR, USA). Cells were cultured at 37 °C, 5% CO2.
Construction of an expression vector to produce fluorobodies
To construct a vector for the expression of scFv-EmGFP, the EmGFP gene from pSERT/EmGFP (cat. #V35320, Thermo Fisher Scientific, USA) was amplified by PCR using two primers; EmGFPNcoINotIFw: 5' CTG TGC CCA TGG GAA TTC AAG CTT GCG GCC GCA GGT GGC GGA GGG ATG GTG AGC AAG GGC GAG GAG-3' and EmGFPFlag6HisXhoIRv: 5'GCA CAG CTC GAG CTA GTG GTG GTG GTG GTG GTG CTT GTC GTC ATC GTC TTT GTA GTC CCC CTT GTA CAG CTC GTC CAT GCC-3', containing the NcoI and XhoI restriction sites (underlined), respectively. The amplified product was digested with NcoI and XhoI and ligated into the expression vector pET15b, which was digested using the same restriction enzymes. The map of the constructed plasmid, named pWS-Green is depicted in Fig. 1a. The integrity of the construct was confirmed by automated DNA sequencing (Macrogen, Korea).
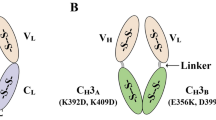
Construction of scFv-EmGFP fluorobodies. A Schematic representation and the DNA sequence of the regulatory 5′ UTR region of scFv-EmGFP constructs. The T7 promoter, lac operator, and ribosome binding site (RBS) along with the linker and EmGFP protein sequences are indicated. Amplified scFv DNA segments from phagemid vector (pMOD1) are digested and ligated into the NcoI and NotI restriction sites in the pWS-Green plasmid, resulting in fluorobody with 6xHis tag and FLAG tag at the C-terminus, to facilitate one-step affinity purification and detection. B 3D model of yPac1A8-EmGFP (anti-P. acnes) and yPgi3G4-EmGFP (anti-P. aeruginosa) generated by AlphaFold protein structure prediction and PyMOL molecular visualization system. The CDR regions of scFv, the VH-VL linker, scFv-EmGFP linker, EmGFP, 6xHis and FLAG tags are labeled
Cloning and expression of scFv-EmGFP fusions
To construct the scFv-EmGFP, the scFv genes, obtained from phagemid vectors (pMOD1) (Pansri et al. 2009), were digested with the NcoI and NotI restriction enzymes and ligated into corresponding digested pWS-Green (Fig. 1a). This resulted in the expression vector scFv-pWS-Green encoding scFv linked with N-terminus of EmGFP protein via G linker (GGGG) (Argos 1990), followed by C-terminal FLAG and 6xHis tags for one-step affinity purification and detection (Fig. 1a). The fused genes are under the control of T7 promoter and can be induced using isopropyl-β-D-thiogalactopyranoside (IPTG).
To produce the scFv-EmEGP fusions, the scFv-pWS-Green expression vectors were transformed into E. coli BL21 (DE3), SHuffle B and SHuffle K-12 strains to determine the suitable expression host. The fluorobodies were expressed according to previously published method (Lobstein et al. 2012) with some modifications. A single colony of each E. coli harboring the recombinant plasmid was inoculated into 5 mL of LB media containing 100 µg/mL of ampicillin and cultured at 30 °C with shaking 250 rpm, overnight. On the next day, 4 mL of overnight culture was inoculated into 400 mL of LB medium containing 100 µg/mL of ampicillin. Cells were cultured at 30 °C until OD600 reach 0.6, and expression was induced with 0.4 mM IPTG at 25 °C for 16 h before harvesting the cells. For negative control, the pET27b harboring gene encoding scFv fused with 6xHis was also expressed at the same time for comparison.
3D-modeling
Protein structure models of yPac1A8-EmGFP (anti-P. acnes) and yPgi3G4-EmGFP (anti-P. aeruginosa) were constructed by using AlphaFold Protein Structure Database (Jumper et al. 2021; Varadi et al. 2022) and PyMOL molecular visualization system from Schrödinger, LLC, USA (DeLano 2004).
Purification of scFv-EmGFP fusions
E. coli cell pellets were harvested by centrifugation at 8,000 rpm for 10 min, then re-suspended in the binding buffer (20 mM Tris–HCl, 300 mM NaCl and 20 mM imidazole, pH 7.9) with 1 mg/mL lysozyme. Cells were disrupted by intermittent sonication at 25% amplitude for 7 min on ice using 30 s pulse and 30 s break for cooling after adding 1 mM phenylmethylsulfonyl fluoride (PMSF). The cell debris was removed by centrifugation at 15,000 × g for 20 min at 4 °C, and the clear supernatant was applied directly onto Ni–NTA column, pre-equilibrated with the binding buffer. After the column was washed with binding buffer, the fusion protein was eluted with elution buffer (20 mM Tris–HCl, 300 mM NaCl and 250 mM imidazole, pH 7.9). Fractions containing scFv-EmGFP fusion were pooled and exchanged by dialysis with PBS buffer and stored at 4 °C for analysis. The soluble fraction and purity of the samples were assessed by denaturing in sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE). The concentration of purified proteins were determined by Bradford method (Bradford 1976), using BSA as a standard.
Gel electrophoresis and western blot analysis
The purified scFv-EmGFP fusions were analyzed by 12% SDS-PAGE. Protein bands were visualized by staining with Coomassie brilliant blue R-250 (Bio-Rad Laboratories, USA) (Laemmli 1970). All Blue Prestained Protein Standards (10–250 kDa) (cat. #1610373, Bio-Rad Laboratories, USA) was used as a molecular weight marker. Western blot was analyzed according to the manufacturer’s protocol (cat. #1703930, Bio-Rad Laboratories, USA). The proteins were electroblotted onto PVDF membrane at 100 V for 1.5 h, at 4 °C. Subsequently, the membrane was blocked for 1 h with 2% (w/v) skim milk in 1 × PBS (MPBS), at 4 °C for overnight. The membrane was then washed 3 times with 1 × PBS by rocking at room temperature for 1 min each time. After incubation with HisProbe-HRP conjugate (1:5000, cat. #15165, ThermoFisher Sceintific, USA,) for 1 h, the membrane was washed again. Finally, the protein target was visualized using Amersham ECL Prime Western Blotting Detection Reagent (cat. #RPN2232, GE Healcare Life Sciences, UK). Image was analyzed by ChemiDoc XRS Gel Documentation System (cat. #1708265, Bio-Rad, USA).
Conventional ELISA
ELISA experiments were performed as previously described for yrPCE1A7-EmGFP (Pruksametanan et al. 2012), and yZEN2A8-EmGFP (Sompunga et al. 2019).
Fluorescence-linked immunosorbent assay (FLISA)
A black Nunc-Immuno 96 well plate (cat. #437111, Thermo Fisher Scientific, USA) was coated with 1 μg of ZEN-BSA in 1 × PBS (for yZEN2A8-EmGFP) or 0.1 IU of rabies virus vaccine in 100 mM NaHCO3, pH 8.5 (for yrPCE1A7-EmGFP). For anti-bacterial fluorobodies, tenfold dilutions of heat-inactivated bacteria, ranging from 1.6 × 109 to 1.6 × 102 cells/mL in PBS were coated onto a black 96-well microtiter plate and incubated overnight at 37 °C, 1% (w/v) BSA or 2℅ skim milk was used as a negative control. After incubation at 4 °C overnight, the plates were blocked with 2% (w/v) MPBS at room temperature for 1 h, followed by washing 3 times with PBS. Then, 50 µL of 4% (w/v) MPBS and 100 μL of scFv-EmGFPs were added into each well and incubated at room temperature for 1 h. The wells were then washed three times with PBS containing 0.05% (v/v) Tween 20 (PBST) and twice with PBS. Finally, 100 µL of PBS were added. The fluorescence intensity was measured by a fluorescent microplate reader (Varioskan LUX multimode microplate reader (cat. #VL0000D0, ThermoScientific, USA). The excitation and emission wavelengths were 478–484 and 560–509 nm, respectively.
Competitive FLISA
To confirm the binding of anti-ZEN (hapten), yZEN2A8-EmGFP, competitive FLISA was performed as previously described for competitive ELISA (Rangnoi et al. 2011). The optimum concentration of different targets and scFv-EmGFP were determined by checkerboard titration as previously described (Sompunga et al. 2019). The IC50 values were estimated from the plot of inhibition curve as F/F0 vs concentration of ZEN at 50% F/F0. F and F0 were fluorescence intensities in wells with various amount of ZEN and zero, respectively.
Direct immunofluorescence staining of bacteria with fluorobodies
The immunofluorescence staining of P. acnes and P. aeruginosa using yPac1A8-EmGFP (anti-P. acnes) and yPgi3G4-EmGFP (anti-P. aeruginosa) was carried out as previously described for scFv-6xHis tagged (Min and Yamabhai 2021).
Flow cytometry analysis of HL-60 cancer cell using scFv-EmGFP
HL-60 cells were blocked with human IgG for 30 min at room temperature. Cells were then washed with 1xPBS containing 0.1% (w/v) BSA. After that, 0.1 μg/μL of anti-HL-60 cells fluorobody (y1HL63D6-EmGFP) was incubated with the cells for 1 h on ice. Then, the cells were washed by centrifugation at 330 × g for 5 min, 4 °C and resuspended with 1xPBS containing 0.1% (w/v) BSA. Propidium iodide (1 mg/mL) at 1:1000 dilution was added to exclude dead cells. Fluorescent intensity was detected by flow cytometer (cat. #A24858, Attune NxT Flow Cytometer, Thermo Fisher Scientific, USA). The events were acquired up to 10,000 events from Propidium iodide (PI) (cat. #P3566, Thermo Fisher Scientific, USA) negative population. For cells that were stained with y1HL63D6 scFv, the cells were stained with DyLight® 488 Anti-6X His tag® Ab (cat. #ab117512, Abcam, UK) for 1 h on ice prior to the washing step.
Cancer cell internalization assay
HL-60 cells were incubated with y1HL63D6-EmGFP at 37 °C, 5% CO2 for 2 h. Then, cells were striped with 0.05% (w/v) trypsin for 10 min before complete media was added and washed with PBS. Then, cells were carefully dropped onto a glass slide. After that, slow fade gold mountant (cat. #S36936, Invitrogen, USA) was added to mount the slide and prevent fluorescent signal fading. Cells were observed under fluorescence microscope (BX50 microscope (cat. #BX50-FL-PA, Olympus, Tokyo, Japan).
Results
Construction of an expression vector for fluorobody (scFv-EmGFP) production
Five scFv genes against ZEN (yZEN2A8) (Sompunga et al. 2019), rabies virus (yrPCE1A7: GenBank accession number OQ129413), P. acnes (yPac1A8) (Min and Yamabhai 2021), P. aeruginosa (yPgi3G4) (Min and Yamabhai 2021) and HL-60 AML cell line (y1HL63D6) (Sumphanapai et al. 2022), were obtained by bio-panning (affinity selection) of a compact non-immunized phage-display human scFv library named YamoI, which was constructed in our laboratory (Pansri et al. 2009). The scFv genes were cut from phagemid pMOD-1 vector that was used to construct the library, and subcloned into NcoI and NotI restriction sites of pWS-Green expression vector, such that scFv DNA fragment is fused with 5′ end of the EmGFP, resulting in the expression of scFv-EmGFP fluorobody. The construct is fused with FLAG tag, followed by 6xHistidine tag at the C-terminus for further detection and one-step purification (Fig. 1a). An example of the three-dimensional models of anti-bacterial scFv-EmGFP, namely, yPac1A8-EmGFP and yPgi3G4-EmGFP, is shown in Fig. 1b.
Expression of scFv-EmGFP fusions from different E. coli strains
In order to determine the suitable expression host for scFv-EmGFP expression, two scFv-EmGFP constructs, i.e., sAFH-3e3-EmGFP (Rangnoi et al. 2021) and yrPCE1A7-EmGFP were transformed into three different strains of E. coli, namely wild type (wt) DE3 and engineered SHuffle B and SHuffle K12 strains. The transformed E. coli appeared fluorescent green as shown in Fig. 2a. The level of fluorescence intensity was much higher for all E. coli strains expressing scFv-EmGFP when compared to EmGFP alone, as indicated by flow cytometry analysis (Fig. 2b). This is likely because for the empty vector, RBS is positioned distal from EmGFP start codon (43 bp) as illustrated in Fig. 1a. When scFv gene was subcloned between NcoI and NotI, the length between RBS and start codon is reduced (6 bp) to be optimal for translation, resulting in high level expression of the scFv-EmGFP fusion, which could be observed from cell lysate as bright fluorescent green as shown in Fig. 2a. Next, the binding of scFv-EmGFP fusion expressed from 3 different strains were determined by FLISA and ELISA. Two, fluorobodies (sAFH-3e3-EmGFP and yrPCE1A7-EmGFP) were used as examples of this study. As demonstrated in Fig. 2c and Figure S1, fluorobodies from engineered E. coli SHuffle B strain exhibited the highest binding activity among the three expression hosts when compared with engineered E. coli SHuffle K12 and DE3. Subsequently, all scFv-EmGFPs constructs were expressed in E. coli SHuffle B strain which has been genetically engineered to promote the formation for disulfide bonded proteins in the cytoplasm (Lobstein et al. 2012).
Properties of scFv-EmGFP generated from different E. coli expression hosts; wt E. coli BL21 (DE3), SHuffle K-12 and SHuffle B. a Tubes of cell lysate containing green fluorescence of scFv-EmGFP visualized with B-BOX Blue Light LED epi-illuminator (cat. #VE0100, SMOBiO, USA). b Flow cytometry analysis of three E. coli expression hosts (SH-B, SH-K and B) harboring 2 scFv-EmGFP constructs, yrPCE1A7- and sAFH-3e3-EmGFP. Empty pET15b vector was used as a negative control. c Schematic representation of how FLISA demonstrates the binding of scFv-EmGFP clones. EmGFP green beta barrel structure, blue antigen, scFv with linker, yellow hexagon FLAG tag and red oval 6xHis tag. sAFH-3e3- and yrPCE1A7-EmGFP obtained from crude cytoplasmic extract were checked for the binding activity against their targets by FLISA. BSA and skim milk were used as negative controls. The relative fluorescence units (RFU) values and standard errors from triplicate wells are shown. E. coli SHuffle B (SH-B); E. coli SHuffle K-12 (SH-K); wt E. coli BL21 (B)
Purification of scFv-EmGFP fusions
After IPTG induction, the fluorobodies were mainly expressed as soluble proteins in the cytosol of E. coli cells. To validate the correct active expression of fluorobodies, we purified them by one-step purification using Ni2+ affinity chromatography to determine the purity and size of the constructs by western blot and evaluate their binding activities. The purified products could be directly observed on a B-BOX Blue Light LED epi-illuminator as shown in Fig. 3a. The visualization of emerald-green fluorescence by fluorobody confirms the presence of the fusions in eluted fractions. SDS-PAGE and western blot analysis of partially purified products indicate that the anti-P. acnes yPac1A8-EmGFP (56 kDa) and anti-P. aeruginosa yPgi3G4-EmGFP (55 kDa) are soluble and expressed at the expected sizes (Fig. 3b). The parental anti-P. acnes yPac1A8 scFv (27.8 kDa) and anti-P. aeruginosa yPgi3G4 scFv (27 kDa) having 6xHis tags are used as controls. Western blot analysis using HisProbe-HRP revealed an extra band of nearly the same size as EmGFP fused with FLAG and 6xHis tags, in addition to the scFv-EmGFP band (Fig. 3b). Similar results were observed for other scFv-EmGFP as well (Figure S2 and S3). Besides, an extra band, which is likely to be an EmGFP dimer, was detected by western blot analysis of yrPCE1A7-EmGFP (Figure S3). These results suggested that proteolysis of the scFv-EmGFP occurred in E. coli (Casey et al. 2000a).
Purification of scFv-EmGFP fusion. a Fluorescence of eluted fractions of anti-P. acnes (yPac1A8-EmGFP) and anti-P. aeruginosa (yPgi3G4-EmGFP) are visualized by illuminating with B-BOX Blue Light LED epi-illuminator at 470 nm. The elution buffer alone was used as a negative control. b Reducing SDS-PAGE and western blot analysis of anti-bacterial fluorobodies. About 1.0 µg of anti-P. acnes yPac1A8 scFv (27.8 kDa), anti-P. acnes yPac1A8-EmGFP (56 kDa), anti-P. aeruginosa yPgi3G4 scFv (27 kDa, kept in 25.0 mg/mL BSA (arrowed); hence another big dark band), and anti-P. aeruginosa yPgi3G4-EmGFP (55 kDa) are loaded into each lane. All Blue Prestained Protein Standards is used as a marker. Asterisks are EmGFP fused with FLAG and 6xHis tags (28.8 kDa)
Detection of mycotoxins
To demonstrate the application of yZEN2A8-EmGFP for quantification of mycotoxin, competitive FLISA was performed. Firstly, optimal concentration of target and yZEN2A8-EmGFP protein were determined by checkerboard titration (Fig. 4a). Competitive FLISA was performed by using of 1:20 dilutions of yZEN2A8-EmGFP against 0.5 and 1.0 µg/well of ZEN-BSA. The IC50 of this scFv-EmGFP was approximaly 0.30–0.55 µg/ml, depending on coated ZEN-BSA per well. The best of IC50 value using this fluorobody was 0.30 µg/ml with a linear range from 0.1 to 5.0 µg/ml. (Fig. 4b). The limit of detection (LOD) of of this scFv-EmGFP Ab was 0.10 µg/ml. These results demonstrated that the yZEN2A8-EmGFP is applicable for the detection of contaminated mycotoxins as it can detect ZEN in range of the maximum allowance of contaminated in general food in Thailand (Anukul et al. 2013; Srianujata 2011) and EU (Wang et al. 2022), which were set at 30–1000 and 50–400 µg/kg, respectively.
Detection of mycotoxin based on binding activity of yZEN2A8-EmGFP to ZEN by competitive FLISA. a Checkerboard titration FLISA of purified scFv-EmGFP dilution from 1:20 to 1:1,280 was performed against 0.5 µg/well of ZEN-BSA (orange line) and 1 µg/well of ZEN-BSA (green line). The data is expressed as absorbance at excitation peak at 487 nm and an emission peak at 506 nm. b Competitive FLISA to detect free ZEN by using 1:20 dilution of yZEN2A8-EmGFP against 0.5 µg/well of ZEN-BSA (orange line) and 1 µg/well of ZEN-BSA (green line) are illustrated. Absorbance values (expressed as F/F0) were plotted against the logarithm of free toxins concentration. The IC50 of yZEN2A8-EmGFP with different coating target are 0.3–0.55 µg/mL
Detection of bacteria
To validate the application of fluorobody for the detection of bacteria, FLISA experiment was conducted on serial dilutions of P. acnes ranging from 1.6 × 109 to 1.6 × 102 cells/mL (Fig. 5a). P. acnes cells were incubated with various concentration of anti-P. acnes yPac1A8-EmGFP (0.1 to 10 μg/mL) and specific interaction was measured as fluorescence intensity in relative fluorescent units (RFU) by Varioskan LUX multimode microplate reader. FLISA result demonstrated that anti-P. acnes yPac1A8-EmGFP at 10 μg/mL could detect P. acnes at 1.6 × 107 cells/mL. However, P. acnes cells could not be detected by yPac1A8-EmGFP at concentrations below 2.5 μg/mL. In conclusion, the yPac1A8-EmGFP could also be used to observe whole bacterial cells as shown by direct immunofluorescence staining in Fig. 5b.
Detection of bacteria by scFv-EmGFP using a Utilizing FLISA, serial dilutions of P. acnes (from 1.6 × 109 to 1.6 × 102 cells/mL) are incubated with anti-P. acnes yPac1A8-EmGFP (from 10 to 0.1 μg/mL). 10 μg/mL of anti-P. acnes yPac1A8-EmGFP can detect 1.6 × 107 P. acnes/mL. The lines represent the average value of duplicate samples and error bars represent the standard error of the mean. b Direct immunofluorescence staining of planktonic cells of P. acnes DMST 14916 (upper panel) and P. aeruginosa DMST 37186 (lower panel) by incubating with anti-P. acnes yPac1A8-EmGFP and anti-P. aeruginosa yPgi3G4-EmGFP, respectively, and counterstained with DAPI. In DAPI panel, nucleoids of bacteria are blue. In GFP panel, P. acnes cells are stained green by anti-P. acnes yPac1A8-EmGFP and P. aeruginosa cells are stained green by anti-P. aeruginosa yPgi3G4-EmGFP. In merged panel, the overlapping of blue nucleoids and GFP signals are seen. The scale bars represent 10 μm
Detection of cancer cells
To investigate potential application of fluorobody for cancer research, scFv (designated y1HL63D6-EmGFP) against acute myeloid (AML) cell line (HL-60) which was selected by whole cell biopanning (Sumphanapai et al. 2022) was used as a model in this study. The binding ability of y1HL63D6-scFv-6xHis and y1HL63D6-EmGFP was compared by flow cytometry analysis. As demonstrated in Fig. 6a, the binding of y1HL63D6-EmGFP was slightly lower than that of y1HL63D6 scFv, with percentage of binding to HL-60 target cells at 95.0% and 99.5%, for y1HL63D6-EmGFP and y1HL63D6 scFv-6xHis, respectively. Next, potential application of y1HL63D6-EmGFP fluorobody for live cell imaging for in vitro internalization study was investigated. HL-60 cells were incubated with y1HL63D6-EmGFP at 37 °C for 2 h. The internalization of y1HL63D6-EmGFP was demonstrated by fluorescence microscope. Green-fluorescent spots appeared inside the cell outlines as shown in the bright field channel, indicating that after binding to the cell surface, the antibody could be internalized into the cells (Fig. 6b).
Application of scFv-EmGFP for cancer research. a Flow cytometry analysis of HL-60 cells against y1HL3D6 scFv and y1HL3D6-EmGFP. The fluorescent intensity was detected in BL-1 channel of the flow cytometer. Black peak: anti-His Dylight488 secondary Ab and PI, Green peak: y1HL3D6 scFv and y1HL3D6-EmGFP, respectively. b Image from fluorescent microscope showing in vitro endocytosis of y1HL3D6-EmGFP by HL-60 cells after incubation for 2 h. Cells were observed using GFP channel (left) and bright field channel (right). Endocytosed y1HL3D6-EmGFP are seen as green dots inside the cells
Discussion
In order to demonstrate the board diversity of our non-immunized human phage display antibody library (Pansri et al. 2009), we selected and identified scFv binders to various targets, ranging from small molecule hapten to virus, bacteria and human cells (Min and Yamabhai 2021; Pansri et al. 2009; Pranomphon et al. 2018; Pruksametanan et al. 2012; Rangnoi et al. 2011, 2021; Sompunga et al. 2019; Vu et al. 2017). Functionality of the selected scFv binders fused to GFP as an in-built feature were validating using FLISA, flow cytometry and immunostaining, to determine their potential uses for various downstream applications. In this study, we described a rapid and simple cloning method to construct a fluorobody comprising of scFv linked to the N-terminus of EmGFP and express the resulting scFv-EmGFP fusion in the cytoplasm of an engineered E. coli. The cloning procedure facilitates subcloning of insert obtained from phage display vector (pMOD), directly into this scFv-EmGFP fusion plasmid (pWS-Green), without PCR amplification. Positioning a start codon too far from the ribosomal binding site in an empty EmGFP (43 nt) vector is the advantage of this expression system because only the clones harboring plasmids containing scFv insert will bring the start codon closer to the ribosomal binding site (6 nt), which is optimal for initiation of translation, resulting in a bright fluorescent signal in the cytoplasm of E. coli.
To discover the optimal E. coli strain for expression of scFv-EmGFP constructs, wt E. coli BL21 (DE3), redox engineered SHuffle K and SHuffle B strains were used to express various fluorobodies. SHuffle B strains lacking the two reductive pathways (ΔtrxB, Δgor) and expressing cytoplasmic disulfide bond isomerase (dsbC) was the ideal expression host, due to the disulfide bonds in scFv (Cys24/Cys98 and Cys160-162/Cys227-229), which are identified by DiANNA 1.1 web server (Ferrè and Clote 2005a, b, 2006)), required for correct folding, stability, and binding (Fass 2012; Huang et al. 2006; Wörn and Plückthun 1998; Zhao et al. 2010). However, the redox state of the whole scFv-EmGFP fusion, having 6 cysteines, remains to be characterized. Since the functionality of various fluorobodies have been demonstrated in this study, it is fair to conclude that any aberrant disulfide bond that may have formed in the scFv-EmGFP fusion, does not interfere with its functionality.
The position of GFP fusion relative to scFv appears to be critical for the functionality of fluorobody, i.e., fluorescent intensity and antigen binding properties. Previous reports have shown that GFP-scFv format was superior to scFv-GFP format when expressed in non-redox engineered E. coli strains. However, the GFP-scFv must be secreted into culture media (Liu et al. 2019) or periplasm (Casey et al. 2000b), or produced as inclusion bodies (Sakamoto et al. 2012). In this study, we show that 5 functional scFv-GFP could be efficiently produced as soluble fusion protein in the cytoplasm using redox-engineered E. coli. Therefore, it can be stated that functional fluorobody both in the form of scFv-GFP or GFP-scFv can be produced if appropriate E. coli host is used for fluorobody expression.
The Ni2+ affinity chromatography was not able to purify the fluorobodies to homogeneity as observed by SDS-PAGE (Fig. 3b) and previous reports (Griep et al. 1999; Oelschlaeger et al. 2002). However, we also demonstrated that the crude and partial purified fluorobodies could be used for ELISA, FLISA, competitive FLISA, flow cytometry, cell internalization assay and immunofluorescence staining. To the best of our knowledge, this is the first report on using yZEN2A8-EmGFP for Zearalenone detection by competitive FLISA immunoassay. The IC50 value was approximately threefold lower (higher sensitivity) than that of competitive ELISA using scFv (Sompunga et al. 2019). In addition to reduction of assay time, since competitive FLISA signal is not amplified by conjugated secondary Ab and the enzyme reaction, the accuracy of quantitative analysis could be enhanced (Jeong et al. 2014; Sakamoto et al. 2012).
Even if the limit of detection (LOD) of fluorobody against bacteria is higher than that of scFv-6xHis, the key advantage of using scFv-EmGFP fusion is being a one-step detection probe. Further optimization of the scFv-EmGFP construct such as fusion of GFP unit at N-terminal of scFv, optimization of the length of scFv-GFP linker or replace the EmGFP with a wide variety of mutated chromophore (Heim and Tsien 1996) might lead to improved detection sensitivity and applicable for detection of multiple strains.
Application of fluorobody for flow cytometry analysis has been previously reported (Petrausch et al. 2007). In their study, unlike our fluorobody produced in engineered E. coli expression system, the fluorobody was produced by yeast expression. Live cell imaging is useful for tracking cellular mechanism. The use of fluorobody allows a direct visualization of cells, avoiding the staining steps required for scFv detection. Here, we demonstrated that the y1HL63D6-EmGFP could be endocytosed into HL-60 target cells (Fig. 6b). Similar application of scFv fusion to C-terminus of GFP for cell-based analysis of LDL uptake mechanisms using GFP-scFv against LDL has been reported (Cardinale et al. 2004; Faulin et al. 2014).
Currently, blue, cyan, green, yellow, orange, red, far-red, and infra-red fluorescent proteins are available for biomolecular applications (Day and Davidson 2009). Data presented in this study demonstrated that fluorobody platform is a suitable format for many diagnostic applications which can be extended to fusions to other GFP variants, for multiplex assays.
In conclusion, this research shows that our bacterial expression platform using engineered E. coli SHuffle B strain is suitable to produce fluorobodies for the detection of a wide variety of targets.
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
Code availability
Not applicable.
Change history
28 March 2023
A Correction to this paper has been published: https://doi.org/10.1007/s00253-023-12494-4
References
Anukul N, Vangnai K, Mahakarnchanakul W (2013) Significance of regulation limits in mycotoxin contamination in Asia and risk management programs at the national level. J Food Drug Anal 21(3):227–241. https://doi.org/10.1016/j.jfda.2013.07.009
Argos P (1990) An investigation of oligopeptides linking domains in protein tertiary structures and possible candidates for general gene fusion. J Mol Biol 211(4):943–958. https://doi.org/10.1016/0022-2836(90)90085-Z
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254. https://doi.org/10.1016/0003-2697(76)90527-3
Cardinale A, Filesi I, Mattei S, Biocca S (2004) Intracellular targeting and functional analysis of single-chain Fv fragments in mammalian cells. Methods 34(2):171–178. https://doi.org/10.1016/j.ymeth.2004.04.006
Casey JL, Coley AM, Tilley LM, Foley M (2000) Green fluorescent antibodies: novel in vitro tools. Annu Rev Biochem 13(6):445–452. https://doi.org/10.1016/j.ymeth.2004.04.00610.1093/protein/13.6.445
Casey JL, Coley AM, Tilley LM, Foley M (2000) Green fluorescent antibodies: novel in vitro tools. Protein Eng 13(6):445–452
Cubitt AB, Woollenweber LA, Heim R (1999) Chapter 2: understanding structure-function relationships in the aequorea victoria green fluorescent protein green fluorescent proteins. Methods Cell Biol 19–30. https://doi.org/10.1016/S0091-679X(08)61946-9
Day RN, Davidson MW (2009) The fluorescent protein palette: tools for cellular imaging. Chem Soc Rev 38(10):2887–2921. https://doi.org/10.1039/b901966a
Delano WL (2004) The PyMOL molecular graphics system. DeLano Scientific, San Carlos. http://pymol.sourceforge.net/
Dübel S (2007) Recombinant therapeutic antibodies. Appl Microbiol Biotechnol 74(4):723–729. https://doi.org/10.1007/s00253-006-0810-y
Fass D (2012) Disulfide bonding in protein biophysics. Annu Rev Biophys 41:63–79. https://doi.org/10.1146/annurev-biophys-050511-102321
Faulin TdES, Guilherme DF, Silva A, Abdalla DSP, Hering VR, Politi MJ, Maranhão AQ (2014) GFP-SCFV: expression and possible applications as a tool for experimental investigations of atherosclerosis. Biotechnol Progr 30(5):1206–1213. https://doi.org/10.1002/btpr.1935
Ferrè F, Clote P (2005) DiANNA: a web server for disulfide connectivity prediction. Nucleic Acids Res 33(Web Server issue):W230-2. https://doi.org/10.1093/nar/gki412
Ferrè F, Clote P (2005) Disulfide connectivity prediction using secondary structure information and diresidue frequencies. Bioinforma 21(10):2336–2346. https://doi.org/10.1093/bioinformatics/bti328
Ferrè F, Clote P (2006) DiANNA 1.1: an extension of the DiANNA web server for ternary cysteine classification. Nucleic Acids Res 34(Web Server issue):W182-5. https://doi.org/10.1093/nar/gkl189
Frenzel A, Hust M, Schirrmann T (2013) Expression of recombinant antibodies. Front Immunol 4:217. https://doi.org/10.3389/fimmu.2013.00217
Griep RA, van Twisk C, van der Wolf JM, Schots A (1999) Fluobodies: green fluorescent single-chain Fv fusion proteins. J Immunol Meth 230(1):121–130. https://doi.org/10.1016/S0022-1759(99)00131-3
Heim R, Tsien RY (1996) Engineering green fluorescent protein for improved brightness, longer wavelengths and fluorescence resonance energy transfer. Curr Biol 6(2):178–182. https://doi.org/10.1016/S0960-9822(02)00450-5
Hermanson GT (2008) Fluorescent probes Bioconjugate Techniques, 2nd edn. Academic Press, New York, pp 396–497
Holmes KL, Lantz LM (2001) Protein labeling with fluorescent probes Methods Cell Biol. vol 63, 2000/11/04 edn, pp 185–204. https://doi.org/10.1016/s0091-679x(01)63013-9
Huang BC, Davern S, Kennel SJ (2006) Mono and bivalent binding of a scFv and covalent diabody to murine laminin-1 using radioiodinated proteins and SPR measurements: effects on tissue retention in vivo. J Immunol Methods 313(1–2):149–160. https://doi.org/10.1016/j.jim.2006.04.006
Ilagan RP, Rhoades E, Gruber DF, Kao HT, Pieribone VA, Regan L (2010) A new bright green-emitting fluorescent protein–engineered monomeric and dimeric forms. FEBS J 277(8):1967–1978. https://doi.org/10.1111/j.1742-4658.2010.07618.x
Jeong GM, Kim YS, Jeong KJ (2014) A human kringle domain-based fluorescence-linked immunosorbent assay system. Anal Biochem 451:63–68. https://doi.org/10.1016/j.ab.2014.01.019
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Zidek A, Potapenko A, Bridgland A, Meyer C, Kohl SAA, Ballard AJ, Cowie A, Romera-Paredes B, Nikolov S, Jain R, Adler J, Back T, Petersen S, Reiman D, Clancy E, Zielinski M, Steinegger M, Pacholska M, Berghammer T, Bodenstein S, Silver D, Vinyals O, Senior AW, Kavukcuoglu K, Kohli P, Hassabis D (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596(7873):583–589. https://doi.org/10.1038/s41586-021-03819-2
Karyolaimos A, de Gier J-W (2021) Strategies to enhance periplasmic recombinant protein production yields in Escherichia coli. Front Bioeng Biotechnol 9:797334. https://doi.org/10.3389/fbioe.2021.797334
Ke N, Landgraf D, Paulsson J, Berkmen M (2016) Visualization of Periplasmic and Cytoplasmic Proteins with a Self-Labeling Protein Tag. J Bacteriol 198(7):1035–1043. https://doi.org/10.1128/JB.00864-15
Kehoe JW, Kay BK (2005) Filamentous Phage Display in the New Millennium. Chem Rev 105(11):4056–4072. https://doi.org/10.1021/cr000261r
Laemmli UK (1970) Cleavage of Structural Proteins during the Assembly of the Head of Bacteriophage T4. Nat 227(5259):680–685. https://doi.org/10.1038/227680a0
Liu M, Wang B, Wang F, Yang Z, Gao D, Zhang C, Ma L, Yu X (2019) Soluble expression of single-chain variable fragment (scFv) in Escherichia coli using superfolder green fluorescent protein as fusion partner. Appl Microbiol Biotechnol 103(15):6071–6079. https://doi.org/10.1007/s00253-019-09925-6
Lobstein J, Emrich CA, Jeans C, Faulkner M, Riggs P, Berkmen M (2012) SHuffle, a novel Escherichia coli protein expression strain capable of correctly folding disulfide bonded proteins in its cytoplasm. Microb Cell Fact 11:56. https://doi.org/10.1186/1475-2859-11-56
McCafferty J, Griffiths AD, Winter G, Chiswell DJ (1990) Phage antibodies: filamentous phage displaying antibody variable domains. Nat 348(6301):552–554. https://doi.org/10.1038/348552a0
Min TT, Yamabhai M (2021) Human hexa-histidine-tagged single-chain variable fragments for bioimaging of bacterial infections. ACS Omega 6(1):762–774. https://doi.org/10.1021/acsomega.0c05340
Oelschlaeger P, Srikant-Iyer S, Lange S, Schmitt J, Schmid RD (2002) Fluorophor-linked immunosorbent assay: a time- and cost-saving method for the characterization of antibody fragments using a fusion protein of a single-chain antibody fragment and enhanced green fluorescent protein. Anal Biochem 309(1):27–34. https://doi.org/10.1016/S0003-2697(02)00290-7
Olichon A, Surrey T (2007) Selection of genetically encoded fluorescent single domain antibodies engineered for efficient expression in Escherichia coli. J Biol Chem 282(50):36314–36320. https://doi.org/10.1074/jbc.M704908200
Pansri P, Jaruseranee N, Rangnoi K, Kristensen P, Yamabhai M (2009) A compact phage display human scFv library for selection of antibodies to a wide variety of antigens. BMC Biotechnol 9:6. https://doi.org/10.1186/1472-6750-9-6
Petrausch U, Dernedde J, Coelho V, Panjideh H, Frey D, Fuchs H, Thiel E, Deckert PM (2007) A33scFv–Green fluorescent protein, a recombinant single-chain fusion protein for tumor targeting. Protein Eng Des Sel 20(12):583–590. https://doi.org/10.1093/protein/gzm043
Pranomphon R, Srila W, Yamabhai M (2018) Generation of recombinant scFv antibody against Ochratoxin A (OTA). Indones J Biotechnol 22:107. https://doi.org/10.22146/ijbiotech.31121
Pruksametanan N, Yamabhai M, Khawplod P (2012) Selection of single chain human monoclonal antibody (scFv) against Rabies virus by phage display technology. In: 6th IEEE Intl Conf NANOMED, 4–7. p 78–81. https://doi.org/10.1109/NANOMED.2012.6509127
Rangnoi K, Jaruseranee N, O’Kennedy R, Pansri P, Yamabhai M (2011) One-step detection of aflatoxin-B1 using scFv-alkaline phosphatase-fusion selected from human phage display antibody library. Mol Biotechnol 49(3):240–249. https://doi.org/10.1007/s12033-011-9398-2
Rangnoi K, Rüker F, Wozniak-Knopp G, Cvak B, O’Kennedy R, Yamabhai M (2021) Binding Characteristic of Various Antibody Formats Against Aflatoxins. ACS Omega 6(39):25258–25268. https://doi.org/10.1021/acsomega.1c03044
Robinson M-P, Ke N, Lobstein J, Peterson C, Szkodny A, Mansell TJ, Tuckey C, Riggs PD, Colussi PA, Noren CJ, Taron CH, DeLisa MP, Berkmen M (2015) Efficient expression of full-length antibodies in the cytoplasm of engineered bacteria. Nat Commun 6(1):8072. https://doi.org/10.1038/ncomms9072
Sakamoto S, Pongkitwitoon B, Nakahara H, Shibata O, Shoyama Y, Tanaka H, Morimoto S (2012) Fluobodies against bioactive natural products and their application in fluorescence-linked immunosorbent assay. Antibodies 1(2):239–258. https://doi.org/10.3390/antib1020239
Schwalbach G, Sibler A-P, Choulier L, Deryckère F, Weiss E (2000) Production of fluorescent single-chain antibody fragments in Escherichia coli. Protein Expr Purif 18(2):121–132. https://doi.org/10.1006/prep.1999.1185
Sompunga P, Pruksametanan N, Rangnoi K, Choowongkomon K, Yamabhai M (2019) Generation of human and rabbit recombinant antibodies for the detection of Zearalenone by phage display antibody technology. Talanta 201:397–405. https://doi.org/10.1016/j.talanta.2019.04.034
Spadiut O, Capone S, Krainer F, Glieder A, Herwig C (2014) Microbials for the production of monoclonal antibodies and antibody fragments. Trends Biotechnol 32(1):54–60. https://doi.org/10.1016/j.tibtech.2013.10.002
Srianujata S (2004) Regulatory update and control measures for prevention and reduction of mycotoxins contamination in foods and feeds. 4th Asian conference on food and nutrition safety, Bali International Convention Center (BICC), Nusa Dua, Bali, Indonesia, 2–5 March.
Sumphanapai T, Chester K, Sawatnatee S, Yeung J, Yamabhai M (2022) Targeting acute myeloid cell surface using a recombinant antibody isolated from whole-cell biopanning of a phage display human scFv antibody library. Med Oncol 39(12):205. https://doi.org/10.1007/s12032-022-01806-9
Tsien RY (1998) The green fluorescent protein. Annu Rev Biochem 67:509–544. https://doi.org/10.1146/annurev.biochem.67.1.509
Varadi M, Anyango S, Deshpande M, Nair S, Natassia C, Yordanova G, Yuan D, Stroe O, Wood G, Laydon A, Zidek A, Green T, Tunyasuvunakool K, Petersen S, Jumper J, Clancy E, Green R, Vora A, Lutfi M, Figurnov M, Cowie A, Hobbs N, Kohli P, Kleywegt G, Birney E, Hassabis D, Velankar S (2022) AlphaFold protein structure database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res 50(D1):D439–D444. https://doi.org/10.1093/nar/gkab1061
Vu NX, Pruksametanan N, Srila W, Yuttavanichakul W, Teamtisong K, Teaumroong N, Boonkerd N, Tittabutr P, Yamabhai M (2017) Generation of a rabbit single-chain fragment variable (scFv) antibody for specific detection of Bradyrhizobium sp DOA9 in both free-living and bacteroid forms. PLoS One 12(6):e0179983. https://doi.org/10.1371/journal.pone.0179983
Wang Y, Zhang C, Wang J, Knopp D (2022) Recent Progress in Rapid Determination of Mycotoxins Based on Emerging Biorecognition Molecules: A Review. Toxins 14(2):73. https://doi.org/10.3390/toxins14020073
Wörn A, Plückthun A (1998) An intrinsically stable antibody scFv fragment can tolerate the loss of both disulfide bonds and fold correctly. FEBS Lett 427(3):357–361. https://doi.org/10.1016/s0014-5793(98)00463-3
Zhao JX, Yang L, Gu ZN, Chen HQ, Tian FW, Chen YQ, Zhang H, Chen W (2010) Stabilization of the single-chain fragment variable by an interdomain disulfide bond and its effect on antibody affinity. Int J Mol Sci 12(1):1–11. https://doi.org/10.3390/ijms12010001
Acknowledgements
The authors would like to acknowledge Natcha Pruksamethanan and Pensuda Sompunga for their technical assistants regarding the construction and characterization of fluorobody against rabies and ZEN, respectively.
Funding
This work was supported by Suranaree University of Technology (SUT), Science Research and Innovation (TSRI) [grant no. RTA6180012]. WS and TS were supported by Suranaree University of Technology and the Royal Golden Jubilee PhD program of Thailand, grant number PHD/0191/2557 and PHD/0098/2553, respectively. MY was also supported by the Distinguished Research Professor Grant (NRCT 808/2563) of the National Research Council of Thailand.
Author information
Authors and Affiliations
Contributions
WS constructed the scFv-EmGFP (fluorobody) expression vector, performed preliminary experiments, and wrote the first draft, TTM was responsible for the results of fluorobody against bacteria and 3D structure modeling, KR was responsible for the results of fluorobody against mycotoxin, TS was responsible for the results of fluorobody against cancer cell, MB supervised experiments on E. coli expression, 3D structure modelling and edited the manuscript. MY conceived of the study and edited the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original version of this article was revised due to a retrospective Open Access order.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Srila, W., Min, T.T., Sumphanapai, T. et al. Production and applications of fluorobody from redox-engineered Escherichia coli. Appl Microbiol Biotechnol 107, 1959–1970 (2023). https://doi.org/10.1007/s00253-023-12395-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-023-12395-6