Abstract
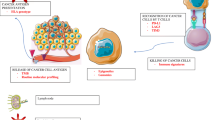
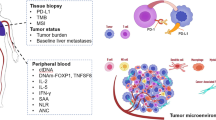
First-line immunotherapy in non-small-cell lung cancer largely improved patients’ survival. PD-L1 testing is required before immune checkpoint inhibitor initiation. However, this biomarker fails to accurately predict patients’ response. On the other hand, immunotherapy exposes patients to immune-related toxicity, the mechanisms of which are still unclear. Hence, there is an unmet need to develop clinically approved predictive biomarkers to better select patients who will benefit the most from immune checkpoint inhibitors and improve risk management. Single-cell technologies provide unprecedented insight into the tumor and its microenvironment, leading to the discovery of immune cells involved in immune checkpoint inhibitor response or toxicity. In this review, we will underscore the potential of the single-cell approach to identify candidate biomarkers improving non-small-cell lung cancer patients’ care.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Biomarker identification in advanced non-small-cell lung cancer (NSCLC): a crucial need
Lung cancer is the leading cause of cancer death worldwide [1]. In the last decade, the development of immune checkpoint inhibitors (ICI) has significantly improved clinical outcomes in advanced NSCLC [2,3,4,5]. Unfortunately, the pattern of ICI response is extremely heterogeneous, from hyper-progressors [6] to durable responders, [7] with the majority of patients not deriving significant benefits from ICI therapy [8]. Furthermore, ICI have a broad spectrum of toxicity and significant cost [9, 10]. Consequently, identifying robust predictive biomarkers is crucial but remains challenging despite numerous efforts.
Tumor programmed death ligand 1 (PD-L1) immunohistochemistry assay is used to determine the proportion of PD-L1 expressing tumor cells. PD-L1 expression was the first ICI biomarker to be published [2] and is currently the only one used in clinical routine. Multiple prospective trials have demonstrated a correlation between the level of tissue PD-L1 expression and clinical efficacy [11], notably KEYNOTE-001 [2], KEYNOTE 024 [12] and PACIFIC trial [13]. The Blueprint PD-L1 IHC Assay Comparison Project evaluated the diagnostic performances of 4 commercially available assays used in clinical trials, since each assay uses a specific clone for their monoclonal primary antibody: 22C3 for pembrolizumab, 28–8 for nivolumab, SP263 for nivolumab and SP142 for atezolizumab. 22C3, 28–8 and SP263 assays had similar performances but SP142 showed a decreased sensitivity [14]. However, the optimal nature of IHC staining (tumor cells, immune cells or both) and the cut-off value of PD-L1 staining positivity is still being debated. Moreover, some studies showed that PD-L1 expression failed to predict clinical outcomes under ICI therapy [15] and some patients with < 1% PD-L1 expression have substantial response rates [16]. For these reasons, PD-L1 expression is considered an imperfect biomarker that needs to be challenged. On the other hand, tumor mutational burden (TMB) quantifies the number of somatic mutations per coding area of a tumor genome using whole exome sequencing (WES) or FoundationOne CDx assay. TMB at the 10 mut/mb cut-point was approved by FDA in 2020 as a biomarker for pembrolizumab benefit across tumor types in the USA [17]. However, TMB faces several limitations in NSCLC such as the arbitrary threshold [18] or the absence of a consistent survival benefit of the TMB high phenotype [19, 20]. Notably, TMB failed to demonstrate its predictive value in NSCLC patients from the Checkmate 227 trial treated with nivolumab plus ipilimumab [21].
To date, PD-L1 expression and TMB are the only biomarkers used to stratify patients in prospective phase 3 clinical trials, whereas other biomarkers are still investigational. For instance, PD-L1 expression on tumor-infiltrating lymphocytes (TILS) was found to be strongly associated with outcome [22]. Despite being associated with objective response rate (ORR) and progression-free survival (PFS) in patients treated with pembrolizumab, lymphocyte infiltration did not substantially add to the predictive value of PD-L1 expression alone for overall survival (OS) [23]. In the blood, baseline Lung Immune Prognostic Index, combining derived neutrophil to lymphocyte ratio and lactate dehydrogenase level was associated with OS in NSCLC patients treated with ICI [24]. Finally, numerous potential predictive biomarkers of ICI efficacy are currently being studied: (i) T-effector and INF-γ-related gene signature [25]; (ii) peripheral blood markers such as proliferating PD-1+CD8 T cells [26] or serum interleukin-8 [27]; (iii) circulating tumor cells [28] or miRNAs/exosomes [29]. Most of these biomarkers still need a prospective validation to challenge PD-L1 expression in clinical routine.
Relevance of single-cell technologies in the identification of predictive biomarkers for immunotherapy in NSCLC
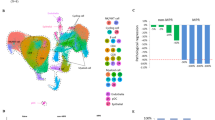
PD-L1 expression remains the only biomarker used to predict ICI response in clinical routine. However, the increasing knowledge on immune checkpoints shed the light on the complexity of their regulation [30], and accumulating evidence highlights the weakness of relying on one single molecule to predict treatment responsiveness. Thus, the ideal biomarker is still to be discovered and might result in a combinative approach [31] with more robustness to predict clinical benefits from immunotherapy. Over the years, the rising era of single-cell technologies [32, 33] unveiled the possibility to dissect a tumor and its microenvironment, including immune cells, with surgical precision by targeting multiple parameters at once. Multiple layers of information, spanning from surface protein expression to transcriptomic changes, can be integrated to have a comprehensive understanding of the mechanisms driving immunotherapy responses and, therefore, offering an opportunity to identify future biomarkers. Hereinafter, we will pinpoint the relevance of single-cell technologies in the identification of predictive biomarkers for immunotherapy in NSCLC (Table 1).
Mass cytometry
Deep immunophenotyping of a large number of cells at a single-cell resolution was made possible by mass cytometry. The samples are stained with antibodies conjugated to heavy-metal isotopes, which prevents spectral overlap observed with flow cytometry. Furthermore, the number of parameters that can be monitored considerably increased, up to 40–60 markers in daily practice [34], allowing for both identification and functional characterization of multiple cell subtypes at the protein level. These technological advances substantially improved our understanding of immune responses and provided the opportunity to uncover pathological alterations affecting the immune system in cancer and resistance mechanisms limiting the success of immunotherapies.
T cells expressing inhibitory immune checkpoints are hyporesponsive upon antigen stimulation and classified as exhausted [35]. Datar et al. [36] phenotyped the infiltrated leukocytes from 20 primary resected NSCLC to assess the biological implications of the expression of three negative immune checkpoints PD-1, LAG-3 and TIM-3. A panel of 35 markers enabled the monitoring of 9 different immune populations: CD8+ T cells, CD4+ T cells, regulatory T cells (Tregs), natural killer (NK) cells, NKT cells, B cells, granulocytes, macrophages and dendritic cells. PD-1 was mostly expressed on CD8+ T cells, CD25−FOXP3−CD4+ T helper, CD25+FOXP3+CD4+ Tregs and CD3+CD56+ NKT cells, LAG-3 on CD8+ T cells and TIM-3 was broadly expressed with macrophages, NK and NKT cells showing the highest expression. Regarding their coexpression, 5.4% of the CD3+ TILs were positive for the three markers, 9.1% were PD-1+LAG-3+, 21% were PD-1+TIM-3+ and 10.3% were LAG-3+TIM-3+. Interestingly, it was demonstrated that PD-1 and TIM-3 coexpression favoured the persistence of exhausted T-cells [37]. As the panel contained markers indicative of cell functions, they investigated the functional characteristics of the T cells expressing the above-mentioned immune checkpoints and observed that their coexpression was associated with higher expression of functional markers. PD-1+LAG-3+TIM-3+ T cells showed the highest expression of activation markers CD69 and 4-1BB, cytotoxicity marker granzyme GZMB, proliferation marker Ki67, apoptotic receptor FAS and pro-apoptotic protein BIM. Finally, they evaluated the impact of the three immune checkpoints on the survival of 90 patients treated with PD-1 inhibitors. Neither high PD-1 nor high TIM-3 T cell expression was predictive of an improved PFS; however, patients expressing high LAG-3 showed a significantly poorer outcome and patients with low expression of LAG-3 and high expression of PD-L1 had an improved PFS. These results suggest that LAG-3 could be involved in resistance to immunotherapy and could be targeted to improve ICI response. Accordingly, combined therapy of anti-LAG-3 relatlimab and anti-PD-1 nivolumab had greater results on PFS in treatment-naïve patients with advanced melanoma compared with nivolumab monotherapy [38].
The large panels used in mass cytometry give the possibility to map the relationships between different cell populations and shift the focus on other cell types that enhance anti-tumor response such as CD4+ T cells. Tay et al. [39] reviewed the multiple lines of evidence suggesting their implication in anti-tumor immunity. Blood samples from NSCLC patients treated with nivolumab revealed that Tregs were more present in the blood of non-responders [40], consistent with Kamada et al.’s [41] study that demonstrated the dampening effect of Tregs on anti-PD-1 response. Besides, responders had higher frequencies of CD62Llow CD8+ and CD4+ T cells [40], the latter containing effector and effector memory T cells crucial for the establishment of an effective and sustained anti-tumor response [42]. The authors performed mass cytometry to better characterize this population and identified CD62Llow CD4+ T cells as T-bet+CD27−FOXP3−CXCR3+ Th1 cells that correlated with cytotoxic CD8+ T cells and PD-1 expression on CD8+ T cells. CD62LlowCD8+ T cell subset was significantly increased in responders, although not as robustly as the CD62LlowCD4+ T cell population. The monitoring of the CD62LlowCD4+ T cell population before and after nivolumab treatment showed that long-term responders without disease progression in the 500 days after immunotherapy had higher frequencies of CD62LlowCD4+ T cells before treatment compared to short-term or non-responders. In addition, patients with an ongoing response between 12 to 92 weeks post-nivolumab had significantly higher frequencies of CD62LlowCD4+ T cells compared to patients who acquired treatment resistance. Notably, Liu et al. [43] also found evidence of Th1 expansion in responders after immunochemotherapy. The authors translated their findings into a predictive score, which could be used in a clinical setting. This predictive score was based on the percentages of CD62Llow and CD25+FOXP3+ among CD4+ T cells and significantly discriminated responders from non-responders after nivolumab treatment with a sensitivity of 92.9% and a specificity of 72.1% in a validation cohort of 86 patients. Hence, this study demonstrated the implication of CD4+ T cells in the maintenance of a sustained response after nivolumab. Immunomonitoring the CD62Llow CD4+ T cell subset in the clinic could be a useful tool to predict long-term responses.
It can be objected that mass cytometry is still expensive and time-consuming for routine use. Nevertheless, this technique can be used in exploratory studies before clinically validated methods in transitional research settings, as exemplified recently. Newel’s team demonstrated the heterogeneity of TILs, containing bystanders and tumor-specific CD8+ TILs characterized by high CD39 expression using mass cytometry. Notably, CD39+CD8+ T cells were predictive of response to immunotherapy in NSCLC patients [44, 45]. Therefore, they compared this technique to other clinically relevant methods for an accurate quantification of this population. Frequencies found with multiplex immunohistochemistry (mIHC) significantly correlated with mass cytometry data. Among patients treated with PD-1/PD-L1 inhibitors, responders had higher proportions of CD39+CD8+ T cells compared with non-responders whereas neither CD39+ cells nor CD8+ T cells retained significance.
scRNA-seq
scRNA-seq made possible the investigation of tumor heterogeneity in an unbiased manner as it does not require the ‘markers selection’ step of mass cytometry. With the identification of patterns of expression programs, not only do we access cell-type information but also cell-state across patients [46].
Cytotoxic CD8+ T cells are thought to be the main drivers of anti-tumor immunity and the cornerstone of cancer immunotherapy response [47]. Gueguen et al. [48] questioned the origins of CD8+ TILs in NSCLC and conducted a pseudotime analysis. Trajectory inference algorithms use single-cell data to predict cell fate over time, giving insights into a dynamic process that can hardly be witnessed [49]. They identified blood circulating memory-like precursor states GZMK+ and KLF2+ CD8+ T cells recruited at the tumor site and tissue-resident precursors XCL1+CD8+ T cells. Results suggested that these precursors converted into a transitional state GZMH+CD8+ T cells to become terminally differentiated CD8+ T cells with a dysfunctional/exhausted phenotype. The late CD8+ T cells were found to be the most cycling cells compared with precursors. PD-1+TIM-3+CD39+CD8+ T cells were Ki67+ and comprised a CD103+ fraction that expressed PD-L1 whereas PD-1−TIM-3− or PD-1+CD8+ T cells were Ki67low. These results were consistent with the TCR sequencing analysis where precursors shared many TCRs with GZMH+ and terminally differentiated CD8+ T cells. Besides, cycling cells TCRs preferentially overlapped with late CD8+ T cells, suggesting that the latter expand in response to tumor-antigen stimulation. Interestingly, they applied response signatures from Sade-Feldman et al. [50] that focused on melanoma patients treated with immunotherapy. KLF2+ and XCL1+ CD8+ T cells had an increased score for the good response signature; meanwhile, GZMH+ and terminally differentiated CD8+ T cells had an increased score for the poor response signature, consistent with the study of Guo et al. [51]. A similar model has since been proposed by Liu et al. [43] based on patients treated with immunochemotherapy. They developed the ‘clonal revival’ theory where pre-existing precursors locally expand and additional peripheral T cells are recruited at the tumor site to actively participate in the anti-tumor response following treatment as hypothesized in Wu et al.’s [52] paper, providing an explanatory mechanism of response to immunotherapy. Their results suggested that the tumor microenvironment promoted CXCL13 expression on CD8+ T cells whereas Banchereau et al. [53] described tissue-resident memory T cells (TRM) expressing high levels of CXCL13 transcript. TRM are characterized by CD103 surface protein expression and their accumulation is promoted by TGF-β1 in mice [54]. Their presence in tumor infiltrate is associated with improved prognosis [54]. Inflamed NSCLC were shown to overexpress CD103-encoding gene ITGAE compared with desert or immune-excluded tumors and its expression on CD8+ T cells was positively correlated with PD-L1 expression [53]. TRM expressed CXCL13, the immune checkpoints LAG3, PDCD1 (PD-1), HAVCR2 (TIM-3), TIGIT, CTLA4, a regulator of tissue-residency ZF683 and markers of tumor-specific response TOX [55] and ENTPD1 (CD39), consistent with Simoni et al.’s [44] study. These results strongly suggest that previously labelled exhausted CD8+ T cells were TRM. Additionally, single-cell TCR clonality analysis revealed that TRM shared most clonotypes with MKI67+ cluster, suggesting that it could result from a proliferation of TRM. Based on OAK trial data [56], they could demonstrate the predictive value of high ITGAE expression with improved OS for patients treated with atezolizumab compared with patients receiving docetaxel. Interestingly, CXCL13 is also involved in the formation of tertiary lymphoid structures (TLS). TLS arise around inflamed tissues and resemble secondary lymphoid organs [57]. They participate in anti-tumor immunity by increasing antigen presentation and cytokine-mediated signalling and improve ICI response in melanoma [58], which was also recently demonstrated in NSCLC [59]. Besides, Patil et al. [54] identified a plasma cell signature that could significantly predict increased OS in patients receiving immunotherapy. mIHC revealed that plasma cells were located at the vicinity of TLS, which presence was also associated to improved survival. Importantly, compared to known biomarkers such as PD-L1 or tissue TMB, the plasma cell signature still had the strongest impact on survival. It must be noted that MS4A1 (CD20) expression correlated with PDCD1 (PD-1) and CD274 (PD-L1) in the study of Chen et al. [60]. However, B cell signatures did not statistically improve the OS of atezolizumab-treated patients in the study of Patil et al. [54].
One major pitfall of scRNA-seq data analysis is the annotation of cell populations. Multiple datasets are often combined and transcriptomic findings are complemented by mass cytometry to validate protein expression levels on specific cell populations. However, it requires additional work and technical issues of combining two different approaches make it more complex. To circumvent this limit, multimodal single-cell technologies have been developed. Notably, CITE-seq allows simultaneous single-cell profiling of both transcripts and surface proteins by using sequenceable DNA oligonucleotides conjugated to antibodies that can bind to epitopes displayed on the cell surface [61]. Leader et al. [62] identified a ‘lung cancer immune activation module’ (LCAM) predictive of immunotherapy response. Based on the POPLAR trial results [25], they could determine that patients with increased frequencies of PD-1+CXCL13+ activated T cells, SPP1+ monocyte-derived macrophages, IgG+ plasma cells and a high TMB had an improved OS when treated with atezolizumab compared with docetaxel-treated patients. PD-1+CXCL13+ T cells could overlap with previously described subsets in studies by Banchereau et al. [53] and Gueguen et al. [48]. Notably, Chen et al. [60] found high expression of IgG encoding transcripts in plasma-like B cells which toxicity varied according to disease stage in culture with lung cancer A549 cell line. Activated T cells, including CD4+ and CD8+ T cells, were found to overexpress PD-1 and CD39 proteins and had an increased percentage of tumor-specific TCR clones. Interestingly, mIHC revealed that samples with high LCAM scores had plasma cells distributed around PD-1+ T cell aggregates or TLS made of PD-1+ T cells and CD20+ B cells, in line with other publications [59, 60]. It is possible to predict ligand-receptor interactions to better understand cell-to-cell communication within the tumor microenvironment. The study by Leader et al. [62] unveiled probable crosstalk between CXCL13+ T cells and CXCR5+ B cells, 41BBL+ B cells and 41BB+ T cells. Therefore, the presence of LCAM could translate an increased anti-tumor immune response that supports patients’ response to immunotherapy. CD274 (PD-L1) gene expression poorly correlated with LCAM score contrarily to TMB, underscoring the necessity to select better predictive tools that include both tumor intrinsic and immune cell infiltrate features.
Lau et al. [63] demonstrated that NSCLC tumor cells expressed human leucocyte antigen (HLA)-II transcripts, mainly HLA-DRB1. Besides, they identified a cluster of cytotoxic GZMB CD4+ T cells that expressed high levels of PDCD1 and CTLA4, suggesting that this population could be involved in the anti-tumor response after immunotherapy. PD-L1 and HLA-II expression in classic Hodgkin’s lymphoma (cHL) were predictive of an improved outcome after immunotherapy [64]. As cHL expresses little HLA-I molecules, it emphasizes the possibility of an anti-tumor immunity independent from HLA-I antigen presentation. In Lau et al.’s [63] study, mIHC analysis indicated that cytotoxic CD4+ T cells potentially interact with HLA-II-expressing tumor cells. The cytotoxic T cells had the greatest clonal expansion, again indicative of a tumor-specific immune response. Notably, high PD-L1 expression on the tumor was not associated with a longer time to progression. Based on a retrospective cohort of 123 individuals, the authors developed a predictive model, based on TMB and a cytotoxic score calculated by taking the mean of log-transformed expression of 25 cytotoxic-related gene signature (NKG7, CXCL13, GZMH, HAVCR2, CCL5, GZMK, CCL4, GZMA, CCL3, CST7, CCL4L2, ACP5, TNFRSF9, TIGIT, GZMB, PDCD1, PRF1, LYST, SIRPG, LAG3, CARD16, TUBA4A, PTMS, CD74, KLRD1) highly expressed in both CD4+ and CD8+ cytotoxic T cells, that could significantly differentiate patients with long and short time to progression after immunotherapy, regardless of HLA-I deficiency. Curiously, they also identified two cytotoxic CD8+ T cell clusters CD103+CD39+ that expressed negative immune checkpoint transcripts PDCD1, LAG3 and TIGIT that resemble and could potentially overlap with the CD103+ and the CD39+ CD8+ T cells previously mentioned [44, 45, 53]. Besides investigating immune populations involved in response to immunotherapy, scRNA-seq can also help to characterize immune cells that trigger treatment resistance. Serine threonine kinase LKB1 mutated tumors are known to poorly respond to immunotherapy [65]. Transcriptional profiling of STK11/LKB1 mutant mouse models revealed the presence of hypofunctional CD8+ T cells with a decrease in memory-like CD8+ T cells and an enrichment in M2-like macrophages [66, 67]. Furthermore, the level of lactate was increased in STK11/LKB1 mutant patients, which could explain the immunosuppressive environment [68]. The blockade of the lactate pathway through MCT4 knock-out or the inhibition of Axl in DCs sensitized LKB1 mutant tumors to immunotherapy, suggesting that combination therapies could circumvent treatment resistance in LKB1 mutant NSCLC patients.
High-dimensional imaging and spatial transcriptomics
Even though cell-to-cell interactions can be computationally predicted from scRNA-seq data, they are based on pairwise ligand-receptor expression; we therefore have no evidence of their actual behaviour in vivo. Imaging mass cytometry (IMC) can solve this problem and provide precious information on the NSCLC tumor architecture and immune cell crosstalk [69, 70]. Jia et al. [71] reported the first case of pseudoprogression after neoadjuvant immunochemotherapy. IMC enabled them to witness a shift from immune desert to infiltrated tumor that could have caused the enlarged lesions. The results showed cell-to-cell contact between CD8+ T cells and CD14+ and CD16+ monocytes, consistent with Leader et al. [62] predictions that involved IL-10, CCL3 and TNF receptors.
Another limit of scRNA-seq is the loss of spatial information regarding gene expression. Spatial transcriptomics enable us to analyse transcriptomic regulation in a tissue context while preserving the overall architecture of the sample [72]. The role of tumor-associated macrophages (TAMs) in promoting tumor growth and resistance to treatment is well known [73]. Larroquette et al. [74] demonstrated the predictive value of TAMs infiltration for NSCLC patients treated with immunotherapy. Indeed, patients with low CD163+ cell density within the tumor had improved clinical outcomes after immunotherapy. However, CD163+ cell infiltration in the stroma was not associated with the outcome, underscoring the importance of spatial information. The highly infiltrated tumors upregulated ITGAM (CD11b), CD27 and CCL5, a chemokine involved in TAM recruitment, whereas tumors with low CD163+ infiltrate upregulated HLA-E and BLC2. Among patients with high infiltrate of TAMs, some showed good treatment response, which was associated with an upregulation of genes associated with M1 phenotype and IFN-γ signalling pathway.
Other emerging technologies are being developed as screening tools for biomarkers in NSCLC, notably digital spatial profiling. Fluorescence images from formalin-fixed paraffin-embedded tissue sections stained with oligonucleotide-conjugated antibodies simultaneously identify proteins or RNA expressed in user-defined compartments such as tumor cells, stroma and immune cells [75]. Several studies demonstrated the feasibility and potential of such approach in both research and clinical settings [76,77,78]. However, this promising technology is still in its early stage and needs to be optimized before being used on a larger scale.
Contribution of single-cell technologies in the investigation of ICI-induced immune-related adverse events (irAEs) in NSCLC
The clinical benefits of ICI are limited by irAEs. Grade ≥ 3 irAEs occur in approximately 10% of NSCLC patients treated by ICI and warrant ICI suspension or permanent discontinuation [79, 80]. Chemotherapy combined with PD-1/PD-L1inhibitors is associated with a lower risk of severe irAEs than PD-1/PD-L1 inhibitors alone and the addition of CTLA-4 inhibitors increases the risk of grade ≥ 3 irAEs [81]. Although the development of irAEs correlates with improved efficacy of PD-1/PD-L1 inhibitors [82], ICI-related pneumonitis and myocarditis of any grade have been associated with decreased survival in NSCLC patients [83, 84]. IrAEs treatment mainly relies on corticosteroids, from low to high dosages according to severity. A study conducted on 2750 lung cancer patients treated by ICI showed that, of the 2% of patients with severe irAEs, 43% developed steroid-resistant irAEs [85]. Hence, the pathophysiology of these highly heterogenous toxicities requires further investigation to guide risk stratification of patients and management strategies and extend the clinical benefit of ICI [86]. Here, we will review the contribution of single-cell technologies to provide key insights into the mechanisms and potential biomarkers of ICI-related irAEs in NSCLC patients (Table 2).
Mechanisms underlying ICI-related irAEs
Although the pathophysiology of ICI-related irAEs resembles that of de novo autoimmune diseases, the precise underlying mechanisms have not been fully elucidated. Shared T cell clones were identified in matched ICI-related autoimmune skin lesions and tumor samples from NSCLC patients [87]. Besides, the discovery of tumor-associated self-antigens (Ag) combined with scRNA-seq further evidenced that autoreactive, napsin A–specific, CD8+ T cells are involved in both ICI efficacy and ICI-related inflammatory lung lesions [88]. This strongly suggests the existence of shared antigens and the capacity of ICI to simultaneously reinvigorate Ag-specific T cells in the two compartments. Besides reflecting treatment effectiveness, the identification of common pathways makes the occurrence of irAEs preventable.
Consistent with the previous notion, a study combining mass cytometry with scRNA-seq detected a predominant clonally expanded CD38hiCD127−CD8+ T cell population with effector and inflammatory phenotype driven by interferon in synovial fluid (SF) from ICI-related arthritis but not in SF from rheumatoid arthritis [89]. CD38hiCD127−CD8+ T cells from the peripheral blood of ICI-related arthritis were also increased, indicative of a systemic reaction to ICI rather than a local expansion [90]. Of note, CD38hi CD8+ T cells reinvigorated by ICI have been detected, using mass cytometry, in tumor and peripheral blood samples from NSCLC patients, where they correlated with improved responses to PD-1 inhibitors [90, 91].
Strikingly, the same mechanism has been suggested in Kim et al.’s [92] publication with CX3CR1hi and CXCR3hi CD8+ T cells, two subsets that could largely overlap with CD38hiCD127−CD8+ T cells from the study of Wang et al. [89]. Although scRNA-seq data show little expression of CD38 transcript by CX3CR1hi CD8+ T cells, the same was observed in CD38hiCD127−CD8+ T cells. Besides, CD38hiCD127−CD8+ T cells expressed high levels of CXR3 and CX3CR1. Kim et al. [92] took a step further with in silico cell-to-cell interaction analysis and in vitro migration assay suggesting that circulating CX3CR1hi CD8+ T cells are recruited into the joints by CXCL9/10/11/16-secreting myeloid cells. These data indicate that Ag-specific T cells activated by ICI and recruited from the periphery may contribute to the pathogenesis of ICI-related arthritis in NSCLC patients.
scRNA-seq and scTCR-seq analysis were also conducted on colon samples from healthy individuals and melanoma patients with or without ICI-related colitis [93]. As in ICI-related arthritis, crosstalk between CXCR3+ T cells and CXCL9/10+ myeloid cells was proposed to be involved in the recruitment of T cells. However, as opposed to ICI-related arthritis, a large proportion of clonally expanded TCRs from TRM were shared with cycling (MKI67+) effector (GZMB+ HLA-DRA+) CD8+ T cells in ICI-related colitis. The authors proposed that the differentiation of TRM into effector CD8+ T cells, further evidenced by pseudotime analysis, could explain the early onset of ICI-related colitis. Tregs play an important role in preventing autoimmune disease and can be depleted by CTLA-4 inhibitors. Tregs expressing a Th1 profile (STAT1+ IL12RB2+ IL10−) were enriched in CTLA-4 inhibitors (± PD-1 inhibitors)-related colitis, compared with healthy colon tissue, whereas no Tregs depletion was observed [93]. Besides, Tregs from ICI-related arthritis were enriched and displayed enhanced immunosuppressive functions [92]. These data indicate that the increased risk to develop grade ≥ 3 irAEs following administration of CTLA-4 inhibitors, alone or in combination, is not caused by depletion of CTLA-4+ Tregs [94], but is rather caused by alterations in expression programs of suppressive CD4+ T cells [95]. Importantly, CD4+ Th17 cells, but not Th1 cells, persisted in the peripheral blood of patients with ICI-related arthritis following steroid administration, suggesting a role of the former cell population in irAEs resistance to steroids [92].
Potential biomarker of ICI-related irAEs
Known risk factors of ICI-related irAEs in NSCLC patients include HLA-I homozygosity [96], BP180-specific immunoglobulin G [97], baseline levels of peripheral CD8+ T cells [98] and gut microbiome [99, 100]. The authors correlated the previously mentioned risk factors with improved responses to ICI. However, it is necessary to dissociate irAEs biomarkers from the efficacy of ICI, as the two events are closely related [101]. Mass cytometry analysis of peripheral blood samples from 46 NSCLC patients, prior to PD-1 inhibitor administration, identified 7 phenotypically distinct populations of regulatory B cells (Bregs) [102]. B-cell cytokine production was impaired in ICI-related irAEs patients, notably interleukine-10 (IL-10), a cytokine produced by Bregs with immunosuppressive properties. Consistent with this finding, the Bregs clusters from ICI-related irAEs patients were decreased and their defect participated in the emergence of ICI-related irAEs. Conversely, scRNA-seq analysis of peripheral blood samples from lung cancer patients revealed increased B cells, with substantially different B cell receptors repertoire, during ICI-related myocarditis remission vs onset [103]. Although changes in peripheral B cell populations were also associated with the development of severe ICI-related irAEs in melanoma patients, the incriminating phenotypes were different from those described in lung cancer patients [104]. This inter-tumor variability adds a layer of complexity to the study of ICI-related irAEs. Indeed, several studies have demonstrated that single-cell technologies are suitable tools to highlight rare events such as peripheral type-1 conventional dendritic cell [105], activated effector memory CD4+ T cell [106] and multimodal molecular states [107] as potential biomarkers of irAEs, dissociated from the response to ICI, in hepatocellular carcinoma, melanoma and metastatic thymic cancer patients, respectively.
Conclusion
Single-cell technologies have led to an unprecedented depth of analysis of the tumor microenvironment, paving the way toward an exhaustive comprehension of the mechanisms of action and resistance to ICI. Indeed, these multiparametric approaches provide highly resolutive data to monitor phenotypic and functional heterogeneity, and refine our knowledge on immune cell composition. New cell types supporting or dampening ICI response are being identified and changes occurring during ICI treatment help patients’ stratification.
This level of information cannot be captured by low-resolution techniques, which barely allow for the rough identification of immune populations and usually include a very limited number of functional markers. This major limitation applies to most techniques that are validated for clinical applications. Therefore, if high-throughput technologies are currently used for screening candidate biomarkers among thousands of immune subsets, we should see the premises of their use for clinical applications within the next coming years. Indeed, the high cost of these technologies, which currently prevents their use in clinical routine, is expected to decrease with barcoding and cell hashing. In addition, this cost must be weighed against the clinical benefit of a therapy better adapted to a patient’s immune profile, and against the cost of monoclonal antibody therapies whose efficacy is bound to be low or non-existent in certain subgroups of patients [9, 10]. This question is particularly relevant given the large number of ICI in phases II and III studies. Finally, the fact that the mechanisms of action and resistance are more complex than initially expected, as they involve a large number of immune subsets, makes it unlikely that a single parameter can accurately predict response to these drugs.
The major asset of immune biomarkers is their clinical actionnability, which opens perspectives of therapeutic intervention. Indeed, the identification of alternative immune checkpoints that are upregulated upon ligation of therapeutic monoclonal antibodies on their targets enables to highlight potential immunotherapy combinations. Upregulation of immune checkpoints such as LAG-3, TIM-3 and BTLA by immune cells has been described following PD-1, PD-L1 and CTLA4 inhibition and was linked with enhanced resistance to these drugs in NSCLC [108, 109]. Consistently, in the results of the phase II CITYSCAPE trial assessing the efficacy of tiragolumab in combination with atezolizumab, the simultaneous targeting of TIGIT and PD-L1 significantly enhanced survival in NSCLC patients, with a safety profile similar to that of atezolizumab alone [110]. Other applications are expected with the identification of immunosuppressive populations that interfere with T cell anti-tumor functions and involved in resistance to immunotherapy that may require depletion before treatment initiation.
The next hits are expected with spatially resolved multiplexed profiling approaches. Indeed, the spatial architecture of the immune microenvironment directly impacts cellular interactions, and has been linked with response to immunotherapy in renal cell carcinoma [111], melanoma [112, 113] and lung cancer [74]. Results of ongoing studies are eagerly awaited and should further enrich the overall picture of the determinants of responsiveness to ICI. The next challenges to overcome in the spatial analysis are yet to come, with needs for improvement of the RNA capture efficiency, as well as the resolution of spatial measurements, to better capture rare cell types while avoiding analysis of doublets.
To conclude, the accumulation of knowledge gained from these high-throughput technologies will help identify the next-generation predictive biomarkers of response to ICI and successfully meet the challenges in oncoimmunology.
References
Barta JA, Powell CA, Wisnivesky JP (2019) Global epidemiology of lung cancer. Ann Glob Health 85(1):8. https://doi.org/10.5334/aogh.2419
Garon EB, Rizvi NA, Hui R, Leighl N, Balmanoukian AS, Eder JP, Patnaik A, Aggarwal C, Gubens M, Horn L, Carcereny E, Ahn MJ, Felip E, Lee JS, Hellmann MD, Hamid O, Goldman JW, Soria JC, Dolled-Filhart M, Rutledge RZ, Zhang J, Lunceford JK, Rangwala R, Lubiniecki GM, Roach C, Emancipator K, Gandhi L, KEYNOTE-001 Investigators (2015) Pembrolizumab for the treatment of non-small-cell lung cancer. N Engl J Med 372(21):2018–28. https://doi.org/10.1056/NEJMoa1501824
Garon EB (2016) The race for combined checkpoint inhibition in NSCLC. Lancet Oncol 17(3):259–260. https://doi.org/10.1016/S1470-2045(15)00580-X
Ellis PM, Vella ET, Ung YC (2017) Immune checkpoint inhibitors for patients with advanced non–small-cell lung cancer: a systematic review. Clin Lung Cancer 18(5):444-459.e1. https://doi.org/10.1016/j.cllc.2017.02.001
Garon EB, Hellmann MD, Rizvi NA et al (2019) Five-year overall survival for patients with advanced non‒small-cell lung cancer treated with pembrolizumab: results from the phase I keynote-001 study. JCO JCO.19.00934. https://doi.org/10.1200/JCO.19.00934
Champiat S, Dercle L, Ammari S et al (2016) Hyperprogressive disease (HPD) is a new pattern of progression in cancer patients treated by anti-PD-1/PD-L1. Clin Cancer Res. https://doi.org/10.1158/1078-0432.CCR-16-1741
Qin A, Street L, Cease K et al (2017) Clinical determinants of durable clinical benefit of pembrolizumab in veterans with advanced non–small-cell lung cancer. Clin Lung Cancer 18(5):559–564. https://doi.org/10.1016/j.cllc.2017.01.012
Havel JJ, Chowell D, Chan TA (2019) The evolving landscape of biomarkers for checkpoint inhibitor immunotherapy. Nat Rev Cancer 19(3):133–150. https://doi.org/10.1038/s41568-019-0116-x
Wang Y, Zhou S, Yang F et al (2019) Treatment-related adverse events of PD-1 and PD-L1 inhibitors in clinical trials: a systematic review and meta-analysis. JAMA Oncol. https://doi.org/10.1001/jamaoncol.2019.0393
Georgieva M, da Silveira Nogueira Lima JP, Aguiar P, de Lima Lopes G, Haaland B (2018) Cost-effectiveness of pembrolizumab as first-line therapy for advanced non-small cell lung cancer. Lung Cancer 124:248–254. https://doi.org/10.1016/j.lungcan.2018.08.018
Bodor JN, Boumber Y, Borghaei H (2020) Biomarkers for immune checkpoint inhibition in non–small cell lung cancer (NSCLC). Cancer 126(2):260–270. https://doi.org/10.1002/cncr.32468
Reck M, Rodríguez-Abreu D, Robinson AG et al (2016) Pembrolizumab versus chemotherapy for PD-L1–positive non–small-cell lung cancer. N Engl J Med 375(19):1823–1833. https://doi.org/10.1056/NEJMoa1606774
Antonia SJ, Villegas A, Daniel D et al (2018) Overall survival with durvalumab after chemoradiotherapy in stage III NSCLC. N Engl J Med 379(24):2342–2350. https://doi.org/10.1056/NEJMoa1809697
Tsao MS, Kerr KM, Kockx M et al (2018) PD-L1 Immunohistochemistry comparability study in real-life clinical samples: results of blueprint phase 2 project. J Thorac Oncol 13(9):1302–1311. https://doi.org/10.1016/j.jtho.2018.05.013
Carbone DP, Reck M, Paz-Ares L et al (2017) First-line nivolumab in stage IV or recurrent non–small-cell lung cancer. N Engl J Med 376(25):2415–2426. https://doi.org/10.1056/NEJMoa1613493
Horn L, Spigel DR, Vokes EE et al (2017) Nivolumab versus docetaxel in previously treated patients with advanced non–small-cell lung cancer: two-year outcomes from two randomized, open-label, phase III trials (CheckMate 017 and CheckMate 057). J Clin Oncol 35(35):3924–3933. https://doi.org/10.1200/JCO.2017.74.3062
Marcus L, Fashoyin-Aje LA, Donoghue M et al (2021) FDA approval summary: pembrolizumab for the treatment of tumor mutational burden-high solid tumors. Clin Cancer Res 27(17):4685–4689. https://doi.org/10.1158/1078-0432.CCR-21-0327
Prasad V, Addeo A (2020) The FDA approval of pembrolizumab for patients with TMB >10 mut/Mb: was it a wise decision? No. Ann Oncol 31(9):1112–1114. https://doi.org/10.1016/j.annonc.2020.07.001
Hellmann MD, Callahan MK, Awad MM et al (2018) Tumor mutational burden and efficacy of nivolumab monotherapy and in combination with ipilimumab in small-cell lung cancer. Cancer Cell 33(5):853-861.e4. https://doi.org/10.1016/j.ccell.2018.04.001
Herbst RS, Giaccone G, de Marinis F et al (2020) Atezolizumab for first-line treatment of PD-L1–selected patients with NSCLC. N Engl J Med. https://doi.org/10.1056/NEJMoa1917346
Hellmann MD, Paz-Ares L, Bernabe Caro R et al (2019) Nivolumab plus ipilimumab in advanced non–small-cell lung cancer. N Engl J Med 381(21):2020–2031. https://doi.org/10.1056/NEJMoa1910231
Herbst RS, Soria J-C, Kowanetz M et al (2014) Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature 515(7528):563–567. https://doi.org/10.1038/nature14011
Hu-Lieskovan S, Lisberg A, Zaretsky JM et al (2019) Tumor characteristics associated with benefit from pembrolizumab in advanced non–small cell lung cancer. Clin Cancer Res 25(16):5061–5068. https://doi.org/10.1158/1078-0432.CCR-18-4275
Mezquita L, Auclin E, Ferrara R et al (2018) Association of the lung immune prognostic index with immune checkpoint inhibitor outcomes in patients with advanced non-small cell lung cancer. JAMA Oncol 4(3):351–357. https://doi.org/10.1001/jamaoncol.2017.4771
Fehrenbacher L, Spira A, Ballinger M et al (2016) Atezolizumab versus docetaxel for patients with previously treated non-small-cell lung cancer (POPLAR): a multicentre, open-label, phase 2 randomised controlled trial. The Lancet 387(10030):1837–1846. https://doi.org/10.1016/S0140-6736(16)00587-0
Kamphorst AO, Pillai RN, Yang S et al (2017) Proliferation of PD-1+ CD8 T cells in peripheral blood after PD-1–targeted therapy in lung cancer patients. PNAS 114(19):4993–4998. https://doi.org/10.1073/pnas.1705327114
Sanmamed MF, Carranza-Rua O, Alfaro C et al (2014) Serum interleukin-8 reflects tumor burden and treatment response across malignancies of multiple tissue origins. Clin Cancer Res 20(22):5697–5707. https://doi.org/10.1158/1078-0432.CCR-13-3203
Guibert N, Delaunay M, Lusque A et al (2018) PD-L1 expression in circulating tumor cells of advanced non-small cell lung cancer patients treated with nivolumab. Lung Cancer 120:108–112. https://doi.org/10.1016/j.lungcan.2018.04.001
Indini A, Rijavec E, Grossi F (2021) Circulating biomarkers of response and toxicity of immunotherapy in advanced non-small cell lung cancer (NSCLC): a comprehensive review. Cancers 13(8):1794. https://doi.org/10.3390/cancers13081794
He X, Xu C (2020) Immune checkpoint signaling and cancer immunotherapy. Cell Res 30(8):660–669. https://doi.org/10.1038/s41422-020-0343-4
Yarchoan M, Albacker LA, Hopkins AC et al (2019) PD-L1 expression and tumor mutational burden are independent biomarkers in most cancers. JCI Insight 4(6):126908. https://doi.org/10.1172/jci.insight.126908
Lee J, Hyeon DY, Hwang D (2020) Single-cell multiomics: technologies and data analysis methods. Exp Mol Med 52(9):1428–1442. https://doi.org/10.1038/s12276-020-0420-2
Stuart T, Satija R (2019) Integrative single-cell analysis. Nat Rev Genet 20(5):257–272. https://doi.org/10.1038/s41576-019-0093-7
Hartmann FJ, Bendall SC (2020) Immune monitoring using mass cytometry and related high-dimensional imaging approaches. Nat Rev Rheumatol 16(2):87–99. https://doi.org/10.1038/s41584-019-0338-z
Philip M, Schietinger A (2019) Heterogeneity and fate choice: T cell exhaustion in cancer and chronic infections. Curr Opin Immunol 58:98–103. https://doi.org/10.1016/j.coi.2019.04.014
Datar I, Sanmamed MF, Wang J et al (2019) Expression analysis and significance of PD-1, LAG-3 and TIM-3 in human non-small cell lung cancer using spatially-resolved and multiparametric single-cell analysis. Clin Cancer Res 25(15):4663–4673. https://doi.org/10.1158/1078-0432.CCR-18-4142
Yang R, Sun L, Li C-F et al (2021) Galectin-9 interacts with PD-1 and TIM-3 to regulate T cell death and is a target for cancer immunotherapy. Nat Commun 12(1):832. https://doi.org/10.1038/s41467-021-21099-2
Tawbi HA, Schadendorf D, Lipson EJ et al (2022) Relatlimab and nivolumab versus nivolumab in untreated advanced melanoma. N Engl J Med 386(1):24–34. https://doi.org/10.1056/NEJMoa2109970
Tay RE, Richardson EK, Toh HC (2021) Revisiting the role of CD4+ T cells in cancer immunotherapy—new insights into old paradigms. Cancer Gene Ther 28(1):5–17. https://doi.org/10.1038/s41417-020-0183-x
Kagamu H, Kitano S, Yamaguchi O et al (2020) CD4+ T-cell immunity in the peripheral blood correlates with response to anti-PD-1 therapy. Cancer Immunol Res 8(3):334–344. https://doi.org/10.1158/2326-6066.CIR-19-0574
Kamada T, Togashi Y, Tay C et al (2019) PD-1+ regulatory T cells amplified by PD-1 blockade promote hyperprogression of cancer. Proc Natl Acad Sci U S A 116(20):9999–10008. https://doi.org/10.1073/pnas.1822001116
Spitzer MH, Carmi Y, Reticker-Flynn NE et al (2017) Systemic immunity is required for effective cancer immunotherapy. Cell 168(3):487-502.e15. https://doi.org/10.1016/j.cell.2016.12.022
Liu B, Hu X, Feng K et al (2022) Temporal single-cell tracing reveals clonal revival and expansion of precursor exhausted T cells during anti-PD-1 therapy in lung cancer. Nat Cancer 3(1):108–121. https://doi.org/10.1038/s43018-021-00292-8
Simoni Y, Becht E, Fehlings M et al (2018) Bystander CD8+ T cells are abundant and phenotypically distinct in human tumour infiltrates. Nature 557(7706):575–579. https://doi.org/10.1038/s41586-018-0130-2
Yeong J, Suteja L, Simoni Y et al (2021) Intratumoral CD39+CD8+ T cells predict response to programmed cell death protein-1 or programmed death ligand-1 blockade in patients with NSCLC. J Thorac Oncol 16(8):1349–1358. https://doi.org/10.1016/j.jtho.2021.04.016
Suvà ML, Tirosh I (2019) Single-cell RNA sequencing in cancer: lessons learned and emerging challenges. Mol Cell 75(1):7–12. https://doi.org/10.1016/j.molcel.2019.05.003
Raskov H, Orhan A, Christensen JP, Gögenur I (2021) Cytotoxic CD8+ T cells in cancer and cancer immunotherapy. Br J Cancer 124(2):359–367. https://doi.org/10.1038/s41416-020-01048-4
Gueguen P, Metoikidou C, Dupic T et al (2021) Contribution of resident and circulating precursors to tumor-infiltrating CD8+ T cell populations in lung cancer. Sci Immunol 6(55):eabd5778. https://doi.org/10.1126/sciimmunol.abd5778
Saelens W, Cannoodt R, Todorov H, Saeys Y (2019) A comparison of single-cell trajectory inference methods. Nat Biotechnol 37(5):547–554. https://doi.org/10.1038/s41587-019-0071-9
Sade-Feldman M, Yizhak K, Bjorgaard SL et al (2018) Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175(4):998-1013.e20. https://doi.org/10.1016/j.cell.2018.10.038
Guo X, Zhang Y, Zheng L et al (2018) Global characterization of T cells in non-small-cell lung cancer by single-cell sequencing. Nat Med 24(7):978–985. https://doi.org/10.1038/s41591-018-0045-3
Wu TD, Madireddi S, de Almeida PE et al (2020) Peripheral T cell expansion predicts tumour infiltration and clinical response. Nature 579(7798):274–278. https://doi.org/10.1038/s41586-020-2056-8
Banchereau R, Chitre AS, Scherl A et al (2021) Intratumoral CD103+ CD8+ T cells predict response to PD-L1 blockade. J Immunother Cancer 9(4):e002231. https://doi.org/10.1136/jitc-2020-002231
Amsen D, van Gisbergen KPJM, Hombrink P, van Lier RAW (2018) Tissue-resident memory T cells at the center of immunity to solid tumors. Nat Immunol 19(6):538–546. https://doi.org/10.1038/s41590-018-0114-2
Scott AC, Dündar F, Zumbo P et al (2019) TOX is a critical regulator of tumour-specific T cell differentiation. Nature 571(7764):270–274. https://doi.org/10.1038/s41586-019-1324-y
Rittmeyer A, Barlesi F, Waterkamp D et al (2017) Atezolizumab versus docetaxel in patients with previously treated non-small-cell lung cancer (OAK): a phase 3, open-label, multicentre randomised controlled trial. The Lancet 389(10066):255–265. https://doi.org/10.1016/S0140-6736(16)32517-X
Sautès-Fridman C, Petitprez F, Calderaro J, Fridman WH (2019) Tertiary lymphoid structures in the era of cancer immunotherapy. Nat Rev Cancer 19(6):307. https://doi.org/10.1038/s41568-019-0144-6
Cabrita R, Lauss M, Sanna A et al (2020) Tertiary lymphoid structures improve immunotherapy and survival in melanoma. Nature 577(7791):561–565. https://doi.org/10.1038/s41586-019-1914-8
Patil NS, Nabet BY, Müller S et al (2022) Intratumoral plasma cells predict outcomes to PD-L1 blockade in non-small cell lung cancer. Cancer Cell 40(3):289-300.e4. https://doi.org/10.1016/j.ccell.2022.02.002
Chen J, Tan Y, Sun F et al (2020) Single-cell transcriptome and antigen-immunoglobin analysis reveals the diversity of B cells in non-small cell lung cancer. Genome Biol 21(1):152. https://doi.org/10.1186/s13059-020-02064-6
Stoeckius M, Hafemeister C, Stephenson W et al (2017) Large-scale simultaneous measurement of epitopes and transcriptomes in single cells. Nat Methods 14(9):865–868. https://doi.org/10.1038/nmeth.4380
Leader AM, Grout JA, Maier BB et al (2021) Single-cell analysis of human non-small cell lung cancer lesions refines tumor classification and patient stratification. Cancer Cell 39(12):1594-1609.e12. https://doi.org/10.1016/j.ccell.2021.10.009
Lau D, Khare S, Stein MM et al (2022) Integration of tumor extrinsic and intrinsic features associates with immunotherapy response in non-small cell lung cancer. Nat Commun 13:4053. https://doi.org/10.1038/s41467-022-31769-4
Roemer MGM, Redd RA, Cader FZ et al (2018) Major histocompatibility complex class II and programmed death ligand 1 expression predict outcome after programmed death 1 blockade in classic Hodgkin lymphoma. J Clin Oncol 36(10):942–950. https://doi.org/10.1200/JCO.2017.77.3994
Skoulidis F, Goldberg ME, Greenawalt DM et al (2018) STK11/LKB1 mutations and PD-1 inhibitor resistance in KRAS-mutant lung adenocarcinoma. Cancer Discov 8(7):822–835. https://doi.org/10.1158/2159-8290.CD-18-0099
Li H, Liu Z, Liu L et al (2022) AXL targeting restores PD-1 blockade sensitivity of STK11/LKB1 mutant NSCLC through expansion of TCF1+ CD8 T cells. Cell Reports Medicine 3(3):100554. https://doi.org/10.1016/j.xcrm.2022.100554
Qian Y, Guijarro I, Galan-Cobo A et al (2022) Abstract 2160: MCT4 blockade reverses lactate-mediated immunosuppression in LKB1-deficient NSCLC. Cancer Research 82(12_Supplement):2160. https://doi.org/10.1158/1538-7445.AM2022-2160
Hayes C, Donohoe CL, Davern M, Donlon NE (2021) The oncogenic and clinical implications of lactate induced immunosuppression in the tumour microenvironment. Cancer Lett 500:75–86. https://doi.org/10.1016/j.canlet.2020.12.021
Giesen C, Wang HAO, Schapiro D et al (2014) Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat Methods 11(4):417–422. https://doi.org/10.1038/nmeth.2869
Li R, Lin Y, Wang Y et al (2021) Characterization of the tumor immune microenvironment in lung squamous cell carcinoma using imaging mass cytometry. Front Oncol 11:620989. https://doi.org/10.3389/fonc.2021.620989
Jia W, Zhu H, Gao Q et al (2021) Case report: transformation from cold to hot tumor in a case of NSCLC neoadjuvant immunochemotherapy pseudoprogression. Front Immunol 12:633534. https://doi.org/10.3389/fimmu.2021.633534
Rao A, Barkley D, França GS, Yanai I (2021) Exploring tissue architecture using spatial transcriptomics. Nature 596(7871):211–220. https://doi.org/10.1038/s41586-021-03634-9
Chen Y, Song Y, Du W, Gong L, Chang H, Zou Z (2019) Tumor-associated macrophages: an accomplice in solid tumor progression. J Biomed Sci 26(1):78. https://doi.org/10.1186/s12929-019-0568-z
Larroquette M, Guegan J-P, Besse B et al (2022) Spatial transcriptomics of macrophage infiltration in non-small cell lung cancer reveals determinants of sensitivity and resistance to anti-PD1/PD-L1 antibodies. J Immunother Cancer 10(5):e003890. https://doi.org/10.1136/jitc-2021-003890
Merritt CR, Ong GT, Church SE et al (2020) Multiplex digital spatial profiling of proteins and RNA in fixed tissue. Nat Biotechnol 38(5):586–599. https://doi.org/10.1038/s41587-020-0472-9
Moutafi M, Martinez-Morilla S, Divakar P et al (2022) Discovery of biomarkers of resistance to immune checkpoint blockade in NSCLC using high-plex digital spatial profiling. J Thorac Oncol 17(8):991–1001. https://doi.org/10.1016/j.jtho.2022.04.009
Moutafi MK, Molero M, Martinez Morilla S et al (2022) Spatially resolved proteomic profiling identifies tumor cell CD44 as a biomarker associated with sensitivity to PD-1 axis blockade in advanced non-small-cell lung cancer. J Immunother Cancer 10(8):e004757. https://doi.org/10.1136/jitc-2022-004757
Zugazagoitia J, Gupta S, Liu Y et al (2020) Biomarkers associated with beneficial PD-1 checkpoint blockade in non-small-cell lung cancer (NSCLC) identified using high-plex digital spatial profiling. Clin Cancer Res 26(16):4360–4368. https://doi.org/10.1158/1078-0432.CCR-20-0175
Brahmer JR, Abu-Sbeih H, Ascierto PA et al (2021) Society for immunotherapy of cancer (SITC) clinical practice guideline on immune checkpoint inhibitor-related adverse events. J Immunother Cancer 9(6):e002435. https://doi.org/10.1136/jitc-2021-002435
Remon J, Reguart N, Auclin E, Besse B (2019) Immune-related adverse events and outcomes in patients with advanced non–small cell lung cancer: a predictive marker of efficacy? J Thorac Oncol 14(6):963–967. https://doi.org/10.1016/j.jtho.2019.02.031
Yan YD, Zhao Y, Zhang C, Fu J, Su YJ, Cui XL, Ma EL, Liu BL, Gu ZC, Lin HW (2022) Toxicity spectrum of immunotherapy in advanced lung cancer: a safety analysis from clinical trials and a pharmacovigilance system. EClinicalMedicine 50:101535. https://doi.org/10.1016/j.eclinm.2022.101535
Shankar B, Zhang J, Naqash AR et al (2020) Multisystem immune-related adverse events associated with immune checkpoint inhibitors for treatment of non-small cell lung cancer. JAMA Oncol 6(12):1952–1956. https://doi.org/10.1001/jamaoncol.2020.5012
Sławiński G, Wrona A, Dąbrowska-Kugacka A, Raczak G, Lewicka E (2020) Immune checkpoint inhibitors and cardiac toxicity in patients treated for non-small lung cancer: a review. Int J Mol Sci 21(19):7195. https://doi.org/10.3390/ijms21197195
Suresh K, Naidoo J (2020) Lower survival in patients who develop pneumonitis following immunotherapy for lung cancer. Clin Lung Cancer 21(3):e169–e170. https://doi.org/10.1016/j.cllc.2019.10.009
Luo J, Beattie JA, Fuentes P et al (2021) Beyond steroids: immunosuppressants in steroid-refractory or resistant immune-related adverse events. J Thorac Oncol 16(10):1759–1764. https://doi.org/10.1016/j.jtho.2021.06.024
Conroy M, Naidoo J (2022) Immune-related adverse events and the balancing act of immunotherapy. Nat Commun 13(1):392. https://doi.org/10.1038/s41467-022-27960-2
Berner F, Bomze D, Diem S et al (2019) Association of checkpoint inhibitor–induced toxic effects with shared cancer and tissue antigens in non–small cell lung cancer. JAMA Oncol 5(7):1–6. https://doi.org/10.1001/jamaoncol.2019.0402
Berner F, Bomze D, Lichtensteiger C et al (2022) Autoreactive napsin A-specific T cells are enriched in lung tumors and inflammatory lung lesions during immune checkpoint blockade. Sci Immunol 7(75):eabn9644. https://doi.org/10.1126/sciimmunol.abn9644
Wang R, Singaraju A, Marks KE, Shakib L, Dunlap G, Cunningham-Bussel A, Chen L, Tirpack A, Fein MR, Todd DJ, MacFarlane L, Goodman SM, DiCarlo EF, Massarotti EM, Sparks JA, Hamnvik OPR, Min L, Jonsson AH, Brenner MB, Chan KK, Bass AR, Donlin LT, Rao DA (2021) Clonally expanded CD38hi cytotoxic CD8 T cells define the T cell infiltrate in checkpoint inhibitor-associated arthritis. 2021.10.19.464961. https://doi.org/10.1101/2021.10.19.464961
Wu P, Zhao L, Chen Y et al (2021) CD38 identifies pre-activated CD8+ T cells which can be reinvigorated by anti-PD-1 blockade in human lung cancer. Cancer Immunol Immunother 70(12):3603–3616. https://doi.org/10.1007/s00262-021-02949-w
Kamphorst AO, Pillai RN, Yang S et al (2017) Proliferation of PD-1+ CD8 T cells in peripheral blood after PD-1–targeted therapy in lung cancer patients. Proc Natl Acad Sci 114(19):4993–4998. https://doi.org/10.1073/pnas.1705327114
Kim ST, Chu Y, Misoi M et al (2022) Distinct molecular and immune hallmarks of inflammatory arthritis induced by immune checkpoint inhibitors for cancer therapy. Nat Commun 13(1):1970. https://doi.org/10.1038/s41467-022-29539-3
Luoma AM, Suo S, Williams HL et al (2020) Molecular pathways of colon inflammation induced by cancer immunotherapy. Cell 182(3):655-671.e22. https://doi.org/10.1016/j.cell.2020.06.001
Arce Vargas F, Furness AJS, Litchfield K et al (2018) Fc effector function contributes to the activity of human anti-CTLA-4 antibodies. Cancer Cell 33(4):649-663.e4. https://doi.org/10.1016/j.ccell.2018.02.010
Grigoriou M, Banos A, Hatzioannou A et al (2021) Regulatory T-cell transcriptomic reprogramming characterizes adverse events by checkpoint inhibitors in solid tumors. Cancer Immunol Res 9(7):726–734. https://doi.org/10.1158/2326-6066.CIR-20-0969
Abed A, Law N, Calapre L et al (2022) Human leucocyte antigen genotype association with the development of immune-related adverse events in patients with non-small cell lung cancer treated with single agent immunotherapy. Eur J Cancer 172:98–106. https://doi.org/10.1016/j.ejca.2022.05.021
Hasan Ali O, Bomze D, Ring SS et al (2020) BP180-specific IgG is associated with skin adverse events, therapy response, and overall survival in non-small cell lung cancer patients treated with checkpoint inhibitors. J Am Acad Dermatol 82(4):854–861. https://doi.org/10.1016/j.jaad.2019.08.045
Wu K, Yang S, Li X, Xia B, Ma S, Chen X (2021) MA09.03 peripheral CD8+ T cells predicts immune-related adverse events and survival in advanced non-small cell lung cancer treated with immunotherapy. J Thorac Oncol 16(10):S911–S912. https://doi.org/10.1016/j.jtho.2021.08.154
Chau J, Yadav M, Liu B et al (2021) Prospective correlation between the patient microbiome with response to and development of immune-mediated adverse effects to immunotherapy in lung cancer. JCO 39(15_suppl):e21024–e21024. https://doi.org/10.1200/JCO.2021.39.15_suppl.e21024
Hakozaki T, Richard C, Elkrief A et al (2020) The gut microbiome associates with immune checkpoint inhibition outcomes in patients with advanced non-small cell lung cancer. Cancer Immunol Res 8(10):1243–1250. https://doi.org/10.1158/2326-6066.CIR-20-0196
Hu W, Wang G, Wang Y, Riese MJ, You M (2020) Uncoupling therapeutic efficacy from immune-related adverse events in immune checkpoint blockade. iScience 23(10):101580. https://doi.org/10.1016/j.isci.2020.101580
Patel AJ, Willsmore ZN, Khan N et al (2022) Regulatory B cell repertoire defects predispose lung cancer patients to immune-related toxicity following checkpoint blockade. Nat Commun 13(1):3148. https://doi.org/10.1038/s41467-022-30863-x
Lou B, Guo M, Chen F et al (2022) Single-cell RNA sequencing reveals the altered landscape of immune cells in immune checkpoint inhibitor related myocarditis. 2022.01.21.22269639. https://doi.org/10.1101/2022.01.21.22269639
Das R, Bar N, Ferreira M et al (2018) Early B cell changes predict autoimmunity following combination immune checkpoint blockade. J Clin Invest 128(2):715–720. https://doi.org/10.1172/JCI96798
Chuah S, Lee J, Song Y et al (2022) Uncoupling immune trajectories of response and adverse events from anti-PD-1 immunotherapy in hepatocellular carcinoma. J Hepatol 77(3):683–694. https://doi.org/10.1016/j.jhep.2022.03.039
Lozano AX, Chaudhuri AA, Nene A et al (2022) T cell characteristics associated with toxicity to immune checkpoint blockade in patients with melanoma. Nat Med 28(2):353–362. https://doi.org/10.1038/s41591-021-01623-z
Zhao C, Mulè MP, Martins AJ, Pinal-Fernandez I, Donahue RN, Chen J, Schlom J, Gulley JL, Mammen A, Tsang JS, Rajan A (2022) Contrasting autoimmune and treatment effects reveals baseline set points of immune toxicity following checkpoint inhibitor treatment. 2022.06.05.494592. https://doi.org/10.1101/2022.06.05.494592
Horvath L, Thienpont B, Zhao L, Wolf D, Pircher A (2020) Overcoming immunotherapy resistance in non-small cell lung cancer (NSCLC) - novel approaches and future outlook. Mol Cancer 19(1):141. https://doi.org/10.1186/s12943-020-01260-z
Thommen DS, Schreiner J, Müller P et al (2015) Progression of lung cancer is associated with increased dysfunction of T cells defined by coexpression of multiple inhibitory receptors. Cancer Immunol Res 3(12):1344–1355. https://doi.org/10.1158/2326-6066.CIR-15-0097
Cho BC, Abreu DR, Hussein M et al (2022) Tiragolumab plus atezolizumab versus placebo plus atezolizumab as a first-line treatment for PD-L1-selected non-small-cell lung cancer (CITYSCAPE): primary and follow-up analyses of a randomised, double-blind, phase 2 study. Lancet Oncol 23(6):781–792. https://doi.org/10.1016/S1470-2045(22)00226-1
Meylan M, Petitprez F, Becht E et al (2022) Tertiary lymphoid structures generate and propagate anti-tumor antibody-producing plasma cells in renal cell cancer. Immunity 55(3):527-541.e5. https://doi.org/10.1016/j.immuni.2022.02.001
Moldoveanu D, Ramsay L, Lajoie M et al (2022) Spatially mapping the immune landscape of melanoma using imaging mass cytometry. Sci Immunol 7(70):eabi5072. https://doi.org/10.1126/sciimmunol.abi5072
Martinez-Morilla S, Villarroel-Espindola F, Wong PF et al (2021) Biomarker discovery in patients with immunotherapy-treated melanoma with imaging mass cytometry. Clin Cancer Res 27(7):1987–1996. https://doi.org/10.1158/1078-0432.CCR-20-3340
Funding
This work has been financially supported by the Fondation ARC (grant SIGNIT202001327), INCa (grant 2012–064/2019–038 for D. O.), ITMO cancer (grant 2014-P032760 for A. S. C.) and the Fondation de France (grant 00076207 for A. S. C.). The team “Immunity and Cancer” was labeled “Equipe Fondation pour la Recherche Médicale (FRM) DEQ 20180339209” (for D.O.). S. F. is supported by a doctoral fellowship from Aix-Marseille University. D. O. is Senior Scholar of the Institut Universitaire de France.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
D. O. is a cofounder and shareholder of ImCheck Therapeutics, Alderaan Biotechnology and Emergence Therapeutics and has research funds from ImCheck Therapeutics, Alderaan Biotechnology, Cellectis and Emergence Therapeutics. The other authors declare no conflicts of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is a contribution to the special issue on: Single-cell and spatial multi-omics in clinical outcomes studies - Guest Editor: Brice Gaudillière
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wlosik, J., Fattori, S., Rochigneux, P. et al. Immune biology of NSCLC revealed by single-cell technologies: implications for the development of biomarkers in patients treated with immunotherapy. Semin Immunopathol 45, 29–41 (2023). https://doi.org/10.1007/s00281-022-00973-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00281-022-00973-1