Abstract
The RecA-family recombinase Rad51 is the central player in homologous recombination (HR), the faithful pathway for repairing DNA double-strand breaks (DSBs) during both mitosis and meiosis. The behavior of Rad51 protein in vivo is fine-tuned via posttranslational modifications conducted by multiple protein kinases in response to cell cycle cues and DNA lesions. Unrepaired DSBs and ssDNA also activate Mec1ATR and Tel1ATM family kinases to initiate the DNA damage response (DDR) that safeguards genomic integrity. Defects in HR and DDR trigger genome instability and result in cancer predisposition, infertility, developmental defects, neurological diseases or premature aging. Intriguingly, yeast Mec1ATR- and Tel1ATM-dependent phosphorylation promotes Rad51 protein stability during DDR, revealing how Mec1ATR can alleviate proteotoxic stress. Moreover, Mec1ATR- and Tel1ATM-dependent phosphorylation also occurs on DDR-unrelated proteins, suggesting that Mec1ATR and Tel1ATM have a DDR-independent function in protein homeostasis. In this minireview, we first describe how human and budding yeast Rad51 are phosphorylated by multiple protein kinases at different positions to promote homology-directed DNA repair and recombination (HDRR). Then, we discuss recent findings showing that intrinsic structural disorder and Mec1ATR/Tel1ATM-dependent phosphorylation are coordinated in yeast Rad51 to regulate both HR and protein homeostasis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
During homology-directed DNA repair and recombination (HDRR), DSBs are initially resected to generate single-stranded DNA (ssDNA). This ssDNA is rapidly protected by an ssDNA binding protein complex, RPA, which is subsequently replaced by Rad51 to form a right-handed nucleoprotein filament. This presynaptic filament is essential for homology search and strand invasion. A hallmark of the Rad51 family recombinases from yeast to mammals is that not only are they highly homologous in terms of amino acid sequences, but they also behave similarly in vitro. Surprisingly, heterologous expression of fission yeast Rad51 or human Rad51 fails to complement the HDRR defects of the budding yeast rad51 mutant (Shinohara et al. 1993). Further analyses unveiled that Rad51 recombinases do not act alone in vivo. Rad51 nucleoprotein filaments are regulated via the coordinated actions of diverse Rad51 mediators or interacting partners (Kowalczykowski 2015; Prakash et al. 2009, 2015; San Filippo et al. 2008). It is also noteworthy that Rad51 recombinases and their mediators often undergo a variety of post-translational modifications (e.g., phosphorylation, sumoylation or ubiquitination) in response to DNA damage agents, cell cycle cues or other signaling molecules (Burger et al. 2019; Cremona et al. 2012; Heyer 2015). Protein phosphorylation and dephosphorylation play key roles in many physiological processes and are often deregulated under pathological conditions. This reversible mechanism is mediated by various protein kinases and phosphatases through the addition or removal of a phosphate group (PO43−) of polar amino acids, including serine (S), threonine (T), tyrosine (Y) or histidine (H). In the budding yeast Saccharomyces cerevisiae, multiple kinases and their transducers are involved in coordinating different HR modules for repairing DNA lesions in mitosis and meiosis (Chuang et al. 2012; Crickard and Greene 2018). Although the strand exchange function of Rad51 is critical for repairing spontaneous DSBs during vegetative growth, it is repressed during meiosis by the meiosis-specific protein Hed1 (Busygina et al. 2008; Tsubouchi and Roeder 2006). Meiotic Rad51 plays a critical role in template choice for HDRR, supporting the strand exchange reaction carried out by the meiosis-specific RecA family protein Dmc1 to repair the programmed DSBs induced by Spo11 (Cloud et al. 2012). Here, we present an overview of Rad51 protein phosphorylation in human and budding yeast. We also discuss recent findings implying that the N-terminal domain (NTD) of yeast Rad51 has dual roles in regulating HDRR and Rad51 homeostasis via its intrinsic structural disorder and through Mec1ATR/Tel1ATM-dependent phosphorylation.
Human and yeast Rad51 recombinases are differentially phosphorylated by multiple protein kinases
Human Rad51 has been shown to be phosphorylated by three serine/threonine kinases [checkpoint kinase 1 (Chk1), polo-like kinase 1 (Plk1), casein kinase 2 (Ck2)] and two tyrosine kinases [Abelson tyrosine kinase (c-Abl) and mesenchymal-epithelial transition factor (c-Met)] (Chabot et al. 2019; Narayanaswamy et al. 2016; Popova et al. 2009; Sorensen et al. 2005; Subramanyam et al. 2016; Yata et al. 2012). The phosphorylation sites on human Rad51 and their biological functions in promoting HDRR are summarized in Table 1. The key function of Chk1 and its paralog Chk2 is to relay DNA damage response (DDR) signals from three DNA damage-sensing protein kinases, i.e., ATM (ataxia-telangiectasia mutated), ATR (ATM- and Rad3-Related), and DNA-dependent protein kinase (DNA-PKcs) (Blackford and Jackson 2017; Marechal and Zou 2013). DNA-PKcs are not present in the S. cerevisiae genome, whereas Mec1 and Tel1 are the S. cerevisiae orthologs of mammalian ATR and ATM, respectively (Craven et al. 2002). c-Abl is phosphorylated and activated by ATM (Wang et al. 2011), whereas c-Met signaling is wired into DDR pathways (Medova et al. 2013). Plk1 plays an important role in the initiation, maintenance, and completion of mitosis (Liu et al. 2017), and it is dephosphorylated and inactivated by protein phosphatase 2A (PP2A) through the ATM/Chk1 DDR pathway (Hyun et al. 2014; Lee et al. 2010). Intriguingly, Plk1 and Ck2 act synergistically during DDR. Plk1 phosphorylates human Rad51 at serine 14 (S14), which primes subsequent Ck2-mediated phosphorylation at Rad51 threonine 13 (T13) and triggers Rad51 binding to the forkhead-associated (FHA) domain of Nbs1 (Yata et al. 2012). Plk1 also phosphorylates Mre11, a component of the Mre11/Rad50/Nbs1 (MRN) complex, at S649 during DDR. Mre11-S649 phosphorylation enables subsequent Ck2-mediated phosphorylation at Mre11-S688 to impede loading of the MRN complex onto damaged DNA, thereby inhibiting HDRR and premature DNA damage checkpoint termination (Li et al. 2017). Further investigations are needed to reveal the relationship between the Rad51–NBS1 interaction and the formation of Rad51 foci during DDR.
Three protein kinases (Mec1ATR, Tel1ATM and Cdc28cdk) are known to phosphorylate budding yeast Rad51 at different target sites (Table 1) (Flott et al. 2011; Lim et al. 2020; Woo et al. 2020). Cdc28cdk, the catalytic subunit of cyclin-dependent protein kinase (CDK), is the master regulator of mitotic and meiotic cell cycles in S. cerevisiae. Using a monoclonal antibody specific for phospho-serines (S*) in PXS*P, PXS*PXR/K or S*PXR/K motifs, it was found that Cdc28cdk could phosphorylate S125 and S375 of an epitope-tagged Rad51 (HA-TEV-Rad51) both in vitro and in vivo (Table 1). Yeast mutant analyses further revealed that mutant non-phosphorylatable Rad51-2A (i.e., Rad51-S125A, S375A) and Rad51-2E (i.e., Rad51-S125E, S375E) impair the DNA binding affinity of Rad51 and the Rad51–Rad52 interaction (Lim et al. 2020). Although these results are important, it is noteworthy that the addition of an epitope or fusion protein tag(s) to native Rad51 often results in deleterious impacts to its normal cellular function and/or unexpected post-translational modification(s) (CNC and TFW, unpublished results). Further investigations are necessary to confirm phosphorylation of S125, S375 and S192 (see below) on native Rad51 protein in vivo using antisera targeted specifically against the corresponding phosphorylated peptides. It will also be important to determine if these modifications affect other biochemical or biological properties of native Rad51 in vivo, such as protein stability and nuclear import.
Mec1ATR and Tel1ATM preferentially phosphorylate S/T-Q motifs, i.e., S and T that are followed by glutamine (Q) (Traven and Heierhorst 2005). Mec1ATR alone can perform most of the consolidated functions of Tel1ATM and Mec1ATR in S. cerevisiae (Corcoles-Saez et al. 2018; Mallory and Petes 2000; Weinert et al. 1994). Yeast Rad51 contains four S/Q motifs, i.e., S2Q, S12Q, S30Q, and S192Q. Mec1ATR is likely responsible for S192 phosphorylation, with S192 being indispensable for Rad51 adenosine triphosphate (ATP) hydrolysis and DNA-binding activity in vitro as well as HDRR in vivo (Flott et al. 2011). Several lines of evidence indicate that the three S/Q motifs (S2Q, S12Q, S30Q) in yeast Rad51-NTD are authentically phosphorylated in a Mec1ATR/Tel1ATM-dependent manner (Woo et al. 2020). First, antisera specific to phosphorylated Rad51-S2Q, Rad51-S12Q and Rad51-S30Q peptides detect phosphorylated Rad51 during both vegetative growth and meiosis. Second, no or negligible signals are detected in corresponding antisera of three respective single-amino-acid substitution mutants (i.e., rad51-S2A, rad51-S12A or rad51-S30A), in the phosphorylation-defective mutant rad51-3A, and in the mec1-kd sml1Δ tel1Δ triple mutant. Third, phosphorylation of Rad51-NTD is only moderately diminished in the tel1Δ single mutant, indicating that Mec1ATR plays a more prominent role than Tel1ATM in Rad51-NTD phosphorylation. Interestingly, phosphorylation of Rad51-NTD also occurs during vegetative growth in the absence of genotoxin treatments or in sporulating spo11Δ diploid cells, but it is not detected in G1-arrested haploid cells or during early meiosis. The reduction of cellular DSB levels during meiosis of the spo11-hypomorphic strain (i.e., spo11-da-HA) leads to a corresponding reduction in phosphorylation levels of Rad51-NTD without apparent perturbation of steady-state Rad51 protein levels (Woo et al. 2020). Given that Spo11 is the catalytic center where meiotic recombination after the premeiotic S phase is initiated (Keeney 2008), we infer that the robustness of Rad51-NTD phosphorylation is tightly associated with different levels of DNA lesions. It is noteworthy that DSB levels are gradually increased as meiosis progresses in WT yeast (Joshi et al. 2015; Padmore et al. 1991). Accordingly, time-course immunoblots following cycloheximide-shutoff experiments (to establish for how long Rad51 is detectable upon inhibition of protein synthesis) show that Rad51 in early meiotic stages is hypophosphorylated and is indeed less stable than the hyperphosphorylated Rad51 in later meiosis (Woo et al. 2020). Together, these results suggest that spontaneous DSBs are responsible for inducing Mec1ATR/Tel1ATM-dependent Rad51-NTD phosphorylation during the vegetative S phase and the premeiotic S phase preceding meiotic Spo11-induced DSBs. It is crucial to further decipher if this low-level spontaneous phosphorylation of Rad51 primes for rapid and robust hyperphosphorylation in response to genotoxin treatments or meiotic DSBs in yeast.
Yeast Rad51 is a paradigm for how the ATR/ATM signaling network regulates both homologous recombination and protein homeostasis
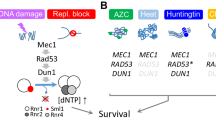
As displayed in Table 1, all known kinases that phosphorylate human Rad51 and yeast Rad51 at various target sites have positive roles in HDRR. Given our recent findings (Woo et al. 2020), summarized below from a mechanistic perspective, Rad51-NTD phosphorylation is unique because its primary role is to enhance Rad51 protein stability by preventing its degradation via the proteasomal pathway. Also noteworthy is that this function can be mimicked by replacing the wild-type (WT) RAD51 gene with the phosphomimetic mutant (rad51-3D) but not with the phosphorylation-defective mutant (rad51-3A). Overexpression of WT or even Rad51-3A proteins can also rescue the HDRR defects displayed by the rad51-3A and/or rad51 null (rad51Δ) mutants. Cycloheximide-shutoff experiments have further revealed that the half-lives of non-phosphorylated Rad51-3A proteins are ~ 30 min in vivo. In contrast, phosphorylated WT protein and Rad51-3D remain stable for > 180 min (Woo et al. 2020). These differing half-lives of Rad51 proteins readily explain why Rad51 phosphorylation has more profound impacts on Rad51-mediated DNA repair during meiosis than during vegetative growth (Woo et al. 2020), given that the mitotic S phase lasts for 20–30 min (Brewer et al. 1984; Slater et al. 1977) whereas the pre-meiotic S phase during synchronous meiosis of SK1 yeast lasts 65–80 min (Cha et al. 2000; Padmore et al. 1991). In addition, Spo11-induced DSBs take place 1.5–3.5 h after cells have been transferred to the meiosis medium, and the chromosomal foci of recombinases appear and disappear within a single peak (2.5–5 h), with maximum abundance at 3 h (Shinohara et al. 2000), implicating a long period (~ 5 h) when Rad51 is required to repair spontaneous DSBs in pre-meiotic S phase and the subsequent Spo11-induced DSBs (Padmore et al. 1991). The non-phosphorylated Rad51-3A is labile and fails to support DSB repair in dmc1Δ hed1Δ meiosis (Woo et al. 2020). Thus, higher Rad51 protein stability is required for meiotic DSB repair when Dmc1 is not available. Although the best-known functions of Mec1ATR and Tel1ATM are their roles in mediating DDR, they also have essential functions in regulating protein homeostasis or proteostasis (Corcoles-Saez et al. 2019, 2018). Along with the observations that hyperphosphorylated Rad51 is more stable than hypophosphorylated Rad51 during DDR, we suggest that Rad51 is a paradigm for Mec1ATR/Tel1ATM-dependent phosphorylation that couples Rad51 homeostasis to HDRR.
Rad51-NTD displays a nanny function in promoting protein expression
The NTD (1–66 amino acids) of budding yeast Rad51 is unique. Multiple sequence alignments of Rad51 proteins from a variety of model organisms (e.g., fission yeast, Neurospora crassa, Drosophila melanogaster, Caenorhabditis elegans, human, and mouse) have revealed that Rad51-NTD (1–66 amino acids) is specific to the genus Saccharomyces (Woo et al. 2020). Yeast mutants (rad51-ΔN) expressing NTD-truncated mutant proteins still possess the capability of promoting HDRR during both mitosis and meiosis but exhibit much lower efficiency in this function. Consistent with the HDRR-impaired phenotypes, steady-state levels of Rad51-ΔN protein in rad51-ΔN mutant cells were only ∼ 3% relative to those of WT Rad51. Further analyses demonstrated that Rad51-NTD can act autonomously to promote the expression of an exogenous protein, β-galactosidase (LacZ), with steady-state levels of Rad51-NTD-LacZ being ≥ 13.2-fold higher in vegetative cells than those of LacZ alone (Woo et al. 2020).
Although the highly abundant Rad51 proteins arising from efficient transcription or translation have been correlated with developmental or pathological conditions, such as respectively in mouse embryonic stem cells or irradiation-resistant tumor cells (Raderschall et al. 2002; Tichy et al. 2012), our understanding of how cells secure Rad51 protein stability in various physiological environments is limited (Ahmed et al. 2018; Ning et al. 2017; Woo et al. 2020). It will be critical to verify if other organisms also sustain such highly efficient protein turnover machineries to downregulate Rad51 and/or its nucleoprotein filaments during DDR and if such regulation is also susceptible to counteractions conferred by the yeast Rad51-NTD.
Intrinsic structural disorder is critical for the nanny function of Rad51-NTD
Many targets of Mec1ATR and Tel1ATM contain at least one S/T-Q cluster domain (SCD), which has been defined as the presence of at least three S/T-Q sites in a stretch of 50 amino acids in S. cerevisiae or 100 amino acids in mammals (Cheung et al. 2012; Traven and Heierhorst 2005). Yeast Rad51-NTD contains three SQ motifs that are phosphorylated dependently on Mec1ATR and Tel1ATM, so it can be ascribed as an SCD. The best-understood mechanism of SCD phosphorylation involves their association with binding partners harboring a forkhead-associated (FHA) domain (Durocher and Jackson 2002). For example, the human tumor suppressor protein CHK2 has an NH2-teminal SCD, followed by an FHA domain and a COOH-terminal catalytic kinase domain. ATR-dependent phosphorylation at CHK2-SCD induces CHK2 activation and phosphorylation-dependent oligomerization via the phospho-SCD/CHK2-FHA interaction (Xu et al. 2002). Moreover, the SCD1 domain (residues 1–29) of the S. cerevisiae Rad53 checkpoint kinase contains two adjacent TQ motifs (T5Q and T8Q) specifically required for recruitment and activation of the Dun1 kinase (Lee et al. 2008). Dun1 phosphorylates Sml1, a potent inhibitor of ribonucleotide reductase (Rnr1), at four serine residues (S56, S58, S60, S61), resulting in proteasomal degradation of Sml1 (Andreson et al. 2010). The sml1 null mutant was originally identified as a suppressor of mec1 viability (Zhao et al. 1998). Similarly, phosphorylation of the S. cerevisiae Hop1 SCD (residues 258–324) at T318Q promotes its interaction with the FHA domain of Mek1 protein kinase. Both Hop1 and Mek1 are meiosis-specific proteins essential for HDRR between homologous non-sister chromosomes (Carballo et al. 2008). Notably, it has been shown that there is low sequence complexity in SCDs enriched for S/T-Q motifs (Traven and Heierhorst 2005). Low sequence complexity and high content of S, T, Q, asparagine (N), proline (P), glycine (G) or charged amino acids is a common feature of many intrinsically disordered regions (IDRs) in proteins (Macossay-Castillo et al. 2019; Romero et al. 2001; Uversky 2019). IDRs are known to be involved in folding, proteasomal degradation, molecular recognition, and protein modifications (Tsvetkov et al. 2008; Uversky 2019; Wright and Dyson 1999). Highly-charged IDR protein sequences act as entropic bristles (EBs) that, when translationally fused to partner proteins, enhance water solubility (but not the overall quantity) of the partner proteins (Santner et al. 2012). However, assessments of the steady-state abundance of proteins with IDRs in cells are challenging because they are often proteolytically degraded, yet they sometimes form abnormal aggregates such as disease-causing prions that can persist in cells.
Inspired by these properties of IDRs, we recently reported that, like Rad51-NTD, the IDRs of several other yeast DDR proteins [e.g., Rad53-SCD1, Hop1-SCD and Sml1-NTD (residues 1–50)], as well as non-DDR proteins [e.g., the prion (nucleation) domains of three yeast prion-causing proteins (Sup35, Ure2 and New1) and the NTDs of Vps64 (Far9), Ssk2 and Kel1], possess autonomous and exchangeable activities to enhance high-level protein expression when they are artificially designed as N-terminal fusion tags of LacZ or GFP, among others. We have discovered an interesting correlation between relative LacZ activities and the overall S/T/Q/N percentages in the total amino acid content of these N-terminal IDRs (N-IDRs). Proteome-wide analyses also suggest that such high S/T/Q/N contents in N-IDRs confer a high predicted folding rate on the proteins that carry them. Intriguingly, all the above-mentioned N-IDRs in the non-DDR proteins also possess at least one S/T-Q motif that may be susceptible to phosphorylation by Mec1ATR and Tel1ATM. For instance, phosphorylation of Sup35-S17Q in response to DNA lesions has been assessed in immunoblots (Chuang et al. 2020). Therefore, we have proposed the “N-terminal intrinsic disorder facilitates abundance” (NIDFA) hypothesis that N-IDRs with high S/T/Q/N contents facilitate protein folding and some could be subject to proteostasis controlled by Mec1ATR and Tel1ATM due to the sporadic emergence of S/T-Q motifs (Chuang et al. 2020). Our NIDFA hypothesis could account for the functions of proteins that harbor an N-IDR but lack binding partners or that fold prior to protein–protein interaction, distinguishing it from two interesting hypothetical models that have been proposed previously. In the first of which, the IDRs in some proteins adopt distinct conformations upon binding to a partner protein, depending on the involvement of different binding partners, chaperones (to support protein folding or degradation) or post-translational modifications (Dyson and Wright 2002; Oldfield et al. 2008; Tompa et al. 2009; Wright and Dyson 1999). These interactions in turn protect IDRs from proteolytic degradation. Alternatively, the “N-terminal folding nucleation” (NFN) hypothesis illustrates that intramolecular interactions modulated by the structured N-terminal domains (SNTDs) that fold spontaneously during protein translation could serve as a nucleation point to organize the as yet unstructured amino acid chain and thus reduce the risk of degradation or aggregation of IDR-containing proteins (Simister et al. 2011). Further investigations must be carried out to delineate if the N-IDRs with high S/T/Q/N content in DDR and/or non-DDR proteins represent docking modules for Mec1ATR- and Tel1ATM-dependent regulation of protein homeostasis.
Conclusion and perspectives
Although our knowledge of the post-translational modifications of other DNA recombinase proteins, e.g., RadA, RecA, and Dmc1, is currently limited, findings regarding the post-translational modification of Rad51 have begun to reveal how cells fine-tune the activity of recombinases in HDRR. In addition to regulating the catalytic activity of Rad51, the unique Rad51-NTD in yeast has demonstrated another mechanism by which HDRR can be controlled. In conclusion, S. cerevisiae Rad51-NTD possesses dual functions to sustain sufficient levels of Rad51 protein under extreme physiological conditions, such as robust DNA lesions or long periods of DNA repair during vegetative growth and meiosis. As an IDR with high S/T/Q content, Rad51-NTD exhibits autonomous expression-enhancing activity for high-level production of native Rad51 and when fused to exogenous β-galactosidase in vivo. Furthermore, Rad51-NTD is an SCD harboring three putative Mec1/Tel1 target sites. Mec1ATR/Tel1ATM-dependent phosphorylation antagonizes the proteasomal degradation pathway and further extends the half-life of Rad51. Further investigations are needed to reveal the genetic determinants that control and/or regulate the protein-expression-enhancement function of SCDs in DDR proteins and even other IDRs of non-DDR proteins. Finally, given that > 1000 potential functional IDR segments have been identified in disease-related proteins (Anbo et al. 2019), in conjunction with the implications that deficiencies of ATM and ATR result in Ataxia-Telangiectasia and Seckel syndrome (Shiloh 2001), it is also important to understand whether IDRs or SCDs in human also exert similar functions in coordinating HDRR and protein homeostasis. A rather clear example of the involvement of IDRs in human disease is illustrated by the pathogenic mechanism of Huntington’s disease (HD) (DiFiglia et al. 1997), which is linked to the expansion of a polyglutamine (poly-Q) domain in the N-terminal part of huntingtin protein (Htt) leading to its aberrant aggregation. We propose that the enhanced IDR property conferred by the expanded poly-Q domain may be one of the reasons why Htt aggregates are highly stable, as shown in a yeast HD model (Meriin et al. 2002). Most intriguingly, Mec1ATR has been shown to be essential for relieving the cellular toxicity conferred by Htt aggregation in yeast (Corcoles-Saez et al. 2018), further implying a DDR-independent role of Mec1ATR in modulating the proteostasis of IDR-containing proteins.
Change history
14 March 2021
A Correction to this paper has been published: https://doi.org/10.1007/s00294-021-01161-8
References
Ahmed KM, Pandita RK, Singh DK, Hunt CR, Pandita TK (2018) Beta1-integrin impacts Rad51 stability and DNA double-strand break repair by homologous recombination. Mol Cell Biol. https://doi.org/10.1128/MCB.00672-17
Anbo H, Sato M, Okoshi A, Fukuchi S (2019) Functional segments on intrinsically disordered regions in disease-related proteins. Biomolecules. https://doi.org/10.3390/biom9030088
Andreson BL, Gupta A, Georgieva BP, Rothstein R (2010) The ribonucleotide reductase inhibitor, Sml1, is sequentially phosphorylated, ubiquitylated and degraded in response to DNA damage. Nucleic Acids Res 38:6490–6501. https://doi.org/10.1093/nar/gkq552
Blackford AN, Jackson SP (2017) ATM, ATR, and DNA-PK: the trinity at the heart of the DNA damage response. Mol Cell 66:801–817. https://doi.org/10.1016/j.molcel.2017.05.015
Brewer BJ, Chlebowicz-Sledziewska E, Fangman WL (1984) Cell cycle phases in the unequal mother/daughter cell cycles of Saccharomyces cerevisiae. Mol Cell Biol 4:2529–2531
Burger K, Ketley RF, Gullerova M (2019) Beyond the trinity of ATM, ATR, and DNA-PK: multiple kinases shape the DNA damage response in concert with RNA metabolism. Front Mol Biosci 6:61. https://doi.org/10.3389/fmolb.2019.00061
Busygina V, Sehorn MG, Shi IY, Tsubouchi H, Roeder GS, Sung P (2008) Hed1 regulates Rad51-mediated recombination via a novel mechanism. Genes Dev 22:786–795
Carballo JA, Johnson AL, Sedgwick SG, Cha RS (2008) Phosphorylation of the axial element protein Hop1 by Mec1/Tel1 ensures meiotic interhomolog recombination. Cell 132:758–770. https://doi.org/10.1016/j.cell.2008.01.035
Cha RS, Weiner BM, Keeney S, Dekker J, Kleckner N (2000) Progression of meiotic DNA replication is modulated by interchromosomal interaction proteins, negatively by Spo11p and positively by Rec8p. Genes Dev 14:493–503
Chabot T, Defontaine A, Marquis D, Renodon-Corniere A, Courtois E, Fleury F, Cheraud Y (2019) New phosphorylation sites of Rad51 by c-met modulates presynaptic filament stability. Cancers (Basel). https://doi.org/10.3390/cancers11030413
Cheung HC, San Lucas FA, Hicks S, Chang K, Bertuch AA, Ribes-Zamora A (2012) An S/T-Q cluster domain census unveils new putative targets under Tel1/Mec1 control. BMC Genom 13:664. https://doi.org/10.1186/1471-2164-13-664
Chuang CN, Cheng YH, Wang TF (2012) Mek1 stabilizes Hop1-Thr318 phosphorylation to promote interhomolog recombination and checkpoint responses during yeast meiosis. Nucleic Acids Res 40:11416–11427. https://doi.org/10.1093/nar/gks920
Chuang CN et al (2020) Intrinsic disorder codes for leaps of protein expression. BioRxiv (MS ID#: BIORXIV/2020/407247; submitted on December 9, 2020)
Cloud V, Chan YL, Grubb J, Budke B, Bishop DK (2012) Rad51 is an accessory factor for Dmc1-mediated joint molecule formation during meiosis. Science 337:1222–1225. https://doi.org/10.1126/science.1219379
Corcoles-Saez I, Dong K, Johnson AL, Waskiewicz E, Costanzo M, Boone C, Cha RS (2018) Essential function of Mec1, the budding yeast ATM/ATR checkpoint-response kinase, in protein homeostasis. Dev Cell 46(495–503):e492. https://doi.org/10.1016/j.devcel.2018.07.011
Corcoles-Saez I, Dong K, Cha RS (2019) Versatility of the Mec1(ATM/ATR) signaling network in mediating resistance to replication, genotoxic, and proteotoxic stresses. Curr Genet. https://doi.org/10.1007/s00294-018-0920-y
Craven RJ, Greenwell PW, Dominska M, Petes TD (2002) Regulation of genome stability by TEL1 and MEC1, yeast homologs of the mammalian ATM and ATR genes. Genetics 161:493–507
Cremona CA, Sarangi P, Zhao X (2012) Sumoylation and the DNA damage response. Biomolecules 2:376–388. https://doi.org/10.3390/biom2030376
Crickard JB, Greene EC (2018) The biochemistry of early meiotic recombination intermediates. Cell Cycle 17:2520–2530. https://doi.org/10.1080/15384101.2018.1553355
DiFiglia M, Sapp E, Chase KO, Davies SW, Bates GP, Vonsattel JP, Aronin N (1997) Aggregation of huntingtin in neuronal intranuclear inclusions and dystrophic neurites in brain. Science 277:1990–1993. https://doi.org/10.1126/science.277.5334.1990
Durocher D, Jackson SP (2002) The FHA domain. FEBS Lett 513:58–66. https://doi.org/10.1016/s0014-5793(01)03294-x
Dyson HJ, Wright PE (2002) Coupling of folding and binding for unstructured proteins. Curr Opin Struct Biol 12:54–60
Flott S, Kwon Y, Pigli YZ, Rice PA, Sung P, Jackson SP (2011) Regulation of Rad51 function by phosphorylation. EMBO Rep 12:833–839. https://doi.org/10.1038/embor.2011.127
Heyer WD (2015) Regulation of recombination and genomic maintenance. Cold Spring Harb Perspect Biol 7:a016501. https://doi.org/10.1101/cshperspect.a016501
Hyun SY, Hwang HI, Jang YJ (2014) Polo-like kinase-1 in DNA damage response. BMB Rep 47:249–255. https://doi.org/10.5483/bmbrep.2014.47.5.061
Joshi N, Brown MS, Bishop DK, Borner GV (2015) Gradual implementation of the meiotic recombination program via checkpoint pathways controlled by global DSB levels. Mol Cell 57:797–811. https://doi.org/10.1016/j.molcel.2014.12.027
Keeney S (2008) Spo11 and the formation of DNA double-strand breaks in meiosis. Genome Dyn Stab 2:81–123. https://doi.org/10.1007/7050_2007_026
Kowalczykowski SC (2015) An overview of the molecular mechanisms of recombinational DNA repair. Cold Spring Harb Perspect Biol. https://doi.org/10.1101/cshperspect.a016410
Lee H et al (2008) Diphosphothreonine-specific interaction between an SQ/TQ cluster and an FHA domain in the Rad53-Dun1 kinase. Cascade Mol Cell 30:767–778
Lee HJ, Hwang HI, Jang YJ (2010) Mitotic DNA damage response: polo-like kinase-1 is dephosphorylated through ATM-Chk1 pathway. Cell Cycle 9:2389–2398. https://doi.org/10.4161/cc.9.12.11904
Li Z, Li J, Kong Y, Yan S, Ahmad N, Liu X (2017) Plk1 phosphorylation of Mre11 antagonizes the DNA damage response. Cancer Res 77:3169–3180. https://doi.org/10.1158/0008-5472.CAN-16-2787
Lim G, Chang Y, Huh WK (2020) Phosphoregulation of Rad51/Rad52 by CDK1 functions as a molecular switch for cell cycle-specific activation of homologous recombination. Sci Adv 6:eaay669. https://doi.org/10.1126/sciadv.aay2669
Liu Z, Sun Q, Wang X (2017) PLK1, a potential target for cancer therapy. Transl Oncol 10:22–32. https://doi.org/10.1016/j.tranon.2016.10.003
Macossay-Castillo M, Marvelli G, Guharoy M, Jain A, Kihara D, Tompa P, Wodak SJ (2019) The balancing act of intrinsically disordered proteins: enabling functional diversity while minimizing promiscuity. J Mol Biol 431:1650–1670. https://doi.org/10.1016/j.jmb.2019.03.008
Mallory JC, Petes TD (2000) Protein kinase activity of Tel1p and Mec1p, two Saccharomyces cerevisiae proteins related to the human ATM protein kinase. Proc Natl Acad Sci USA 97:13749–13754. https://doi.org/10.1073/pnas.250475697
Marechal A, Zou L (2013) DNA damage sensing by the ATM and ATR kinases. Cold Spring Harb Perspect Biol. https://doi.org/10.1101/cshperspect.a012716
Medova M, Aebersold DM, Zimmer Y (2013) The molecular crosstalk between the MET receptor tyrosine kinase and the DNA damage response-biological and clinical aspects. Cancers (Basel) 6:1–27. https://doi.org/10.3390/cancers6010001
Meriin AB, Zhang X, He X, Newnam GP, Chernoff YO, Sherman MY (2002) Huntington toxicity in yeast model depends on polyglutamine aggregation mediated by a prion-like protein Rnq1. J Cell Biol 157:997–1004. https://doi.org/10.1083/jcb.200112104
Narayanaswamy PB, Tkachuk S, Haller H, Dumler I, Kiyan Y (2016) CHK1 and RAD51 activation after DNA damage is regulated via urokinase receptor/TLR4 signaling. Cell Death Dis 7:e2383. https://doi.org/10.1038/cddis.2016.291
Ning J, Wakimoto H, Peters C, Martuza RL, Rabkin SD (2017) Rad51 degradation: role in oncolytic virus-poly(ADP-Ribose) polymerase inhibitor combination therapy in glioblastoma. J Natl Cancer Inst 109:1–13. https://doi.org/10.1093/jnci/djw229
Oldfield CJ, Meng J, Yang JY, Yang MQ, Uversky VN, Dunker AK (2008) Flexible nets: disorder and induced fit in the associations of p53 and 14–3-3 with their partners. BMC Genom 9(Suppl 1):S1. https://doi.org/10.1186/1471-2164-9-S1-S1
Padmore R, Cao L, Kleckner N (1991) Temporal comparison of recombination and synaptonemal complex formation during meiosis in S. cerevisiae. Cell 66:1239–1256
Popova M, Shimizu H, Yamamoto K, Lebechec M, Takahashi M, Fleury F (2009) Detection of c-Abl kinase-promoted phosphorylation of Rad51 by specific antibodies reveals that Y54 phosphorylation is dependent on that of Y315. FEBS Lett 583:1867–1872. https://doi.org/10.1016/j.febslet.2009.04.044
Prakash R et al (2009) Yeast Mph1 helicase dissociates Rad51-made D-loops: implications for crossover control in mitotic recombination. Genes Dev 23:67–79. https://doi.org/10.1101/gad.1737809
Prakash R, Zhang Y, Feng W, Jasin M (2015) Homologous recombination and human health: the roles of BRCA1, BRCA2, and associated proteins. Cold Spring Harb Perspect Biol 7:a016600. https://doi.org/10.1101/cshperspect.a016600
Raderschall E, Stout K, Freier S, Suckow V, Schweiger S, Haaf T (2002) Elevated levels of Rad51 recombination protein in tumor cells. Cancer Res 62:219–225
Romero P, Obradovic Z, Li X, Garner EC, Brown CJ, Dunker AK (2001) Sequence complexity of disordered protein. Proteins 42:38–48
San Filippo J, Sung P, Klein H (2008) Mechanism of eukaryotic homologous recombination. Annu Rev Biochem 77:229–257. https://doi.org/10.1146/annurev.biochem.77.061306.125255
Santner AA, Croy CH, Vasanwala FH, Uversky VN, Van YY, Dunker AK (2012) Sweeping away protein aggregation with entropic bristles: intrinsically disordered protein fusions enhance soluble expression. Biochemistry 51:7250–7262. https://doi.org/10.1021/bi300653m
Shiloh Y (2001) ATM and ATR: networking cellular responses to DNA damage. Curr Opin Genet Dev 11:71–77. https://doi.org/10.1016/s0959-437x(00)00159-3
Shinohara A, Ogawa H, Matsuda Y, Ushio N, Ikeo K, Ogawa T (1993) Cloning of human, mouse and fission yeast recombination genes homologous to RAD51 and recA. Nat Genet 4:239–243. https://doi.org/10.1038/ng0793-239
Shinohara M, Gasior SL, Bishop DK, Shinohara A (2000) Tid1/Rdh54 promotes colocalization of rad51 and dmc1 during meiotic recombination. Proc Natl Acad Sci USA 97:10814–10819. https://doi.org/10.1073/pnas.97.20.10814
Simister PC, Schaper F, O’Reilly N, McGowan S, Feller SM (2011) Self-organization and regulation of intrinsically disordered proteins with folded N-termini. PLoS Biol 9:e1000591. https://doi.org/10.1371/journal.pbio.1000591
Slater ML, Sharrow SO, Gart JJ (1977) Cell cycle of Saccharomyces cerevisiae in populations growing at different rates. Proc Natl Acad Sci USA 74:3850–3854. https://doi.org/10.1073/pnas.74.9.3850
Sorensen CS, Hansen LT, Dziegielewski J, Syljuasen RG, Lundin C, Bartek J, Helleday T (2005) The cell-cycle checkpoint kinase Chk1 is required for mammalian homologous recombination repair. Nat Cell Biol 7:195–201. https://doi.org/10.1038/ncb1212
Subramanyam S, Ismail M, Bhattacharya I, Spies M (2016) Tyrosine phosphorylation stimulates activity of human RAD51 recombinase through altered nucleoprotein filament dynamics. Proc Natl Acad Sci USA 113:E6045–E6054. https://doi.org/10.1073/pnas.1604807113
Tichy ED, Pillai R, Deng L, Tischfield JA, Hexley P, Babcock GF, Stambrook PJ (2012) The abundance of Rad51 protein in mouse embryonic stem cells is regulated at multiple levels. Stem Cell Res 9:124–134. https://doi.org/10.1016/j.scr.2012.05.004
Tompa P, Fuxreiter M, Oldfield CJ, Simon I, Dunker AK, Uversky VN (2009) Close encounters of the third kind: disordered domains and the interactions of proteins. BioEssays 31:328–335. https://doi.org/10.1002/bies.200800151
Traven A, Heierhorst J (2005) SQ/TQ cluster domains: concentrated ATM/ATR kinase phosphorylation site regions in DNA-damage-response proteins. BioEssays 27:397–407. https://doi.org/10.1002/bies.20204
Tsubouchi H, Roeder GS (2006) Budding yeast Hed1 down-regulates the mitotic recombination machinery when meiotic recombination is impaired. Genes Dev 20:1766–1775
Tsvetkov P, Asher G, Paz A, Reuven N, Sussman JL, Silman I, Shaul Y (2008) Operational definition of intrinsically unstructured protein sequences based on susceptibility to the 20S proteasome. Proteins 70:1357–1366. https://doi.org/10.1002/prot.21614
Uversky VN (2019) Intrinsically disordered proteins and their “mysterious” (meta)physics. Front Phys. https://doi.org/10.3389/fphy.2019.00010
Wang X et al (2011) A positive role for c-Abl in Atm and Atr activation in DNA damage response. Cell Death Differ 18:5–15. https://doi.org/10.1038/cdd.2010.106
Weinert TA, Kiser GL, Hartwell LH (1994) Mitotic checkpoint genes in budding yeast and the dependence of mitosis on DNA replication and repair. Genes Dev 8:652–665
Woo TT, Chuang CN, Higashide M, Shinohara A, Wang TF (2020) Dual roles of yeast Rad51 N-terminal domain in repairing DNA double-strand breaks. Nucleic Acids Res 48:8474–8489. https://doi.org/10.1093/nar/gkaa587
Wright PE, Dyson HJ (1999) Intrinsically unstructured proteins: re-assessing the protein structure-function paradigm. J Mol Biol 293:321–331. https://doi.org/10.1006/jmbi.1999.3110
Xu X, Tsvetkov LM, Stern DF (2002) Chk2 activation and phosphorylation-dependent oligomerization. Mol Cell Biol 22:4419–4432. https://doi.org/10.1128/mcb.22.12.4419-4432.2002
Yata K, Lloyd J, Maslen S, Bleuyard JY, Skehel M, Smerdon SJ, Esashi F (2012) Plk1 and CK2 act in concert to regulate Rad51 during DNA double strand break repair. Mol Cell 45:371–383. https://doi.org/10.1016/j.molcel.2011.12.028
Zhao X, Muller EG, Rothstein R (1998) A suppressor of two essential checkpoint genes identifies a novel protein that negatively affects dNTP pools. Mol Cell 2:329–340
Acknowledgements
We thank John O’Brien for English editing and Rita Cha (Bangor University, UK) for sharing and discussing unpublished results on Sml1-regulated protein homeostasis. This work was supported by grants from Academia Sinica (AS-105-TP-B07 and AS108-TP-B07) and the Ministry of Science and Technology (MOST106-2311-B-001-016-MY3 and MOST109-2811-B-001-008-MY3), Taiwan, Republic of China. TTW was supported by a postdoctoral fellowship from Academia Sinica.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised due to a retrospective Open Access order.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Woo, TT., Chuang, CN. & Wang, TF. Budding yeast Rad51: a paradigm for how phosphorylation and intrinsic structural disorder regulate homologous recombination and protein homeostasis. Curr Genet 67, 389–396 (2021). https://doi.org/10.1007/s00294-020-01151-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-020-01151-2