Abstract
Objectives
This study investigates the influence of normal cohort (NC) size and the impact of different NCs on automated MRI-based brain atrophy estimation.
Methods
A pooled NC of 3945 subjects (NCpool) was retrospectively created from five publicly available cohorts. Voxel-wise gray matter volume atrophy maps were calculated for 48 Alzheimer’s disease (AD) patients (55–82 years) using veganbagel and dynamic normal templates with an increasing number of healthy subjects randomly drawn from NCpool (initially three, and finally 100 subjects). Over 100 repeats of the process, the mean over a voxel-wise standard deviation of gray matter z-scores was established and plotted against the number of subjects in the templates. The knee point of these curves was defined as the minimum number of subjects required for consistent brain atrophy estimation. Atrophy maps were calculated using each NC for AD patients and matched healthy controls (HC). Two readers rated the extent of mesiotemporal atrophy to discriminate AD/HC.
Results
The maximum knee point was at 15 subjects. For 21 AD/21 HC, a sufficient number of subjects were available in each NC for validation. Readers agreed on the AD diagnosis in all cases (Kappa for the extent of atrophy, 0.98). No differences in diagnoses between NCs were observed (intraclass correlation coefficient, 0.91; Cochran’s Q, p = 0.19).
Conclusion
At least 15 subjects should be included in age- and sex-specific normal templates for consistent brain atrophy estimation. In the study’s context, qualitative interpretation of regional atrophy allows reliable AD diagnosis with a high inter-reader agreement, irrespective of the NC used.
Clinical relevance statement
The influence of normal cohorts (NCs) on automated brain atrophy estimation, typically comparing individual scans to NCs, remains largely unexplored. Our study establishes the minimum number of NC-subjects needed and demonstrates minimal impact of different NCs on regional atrophy estimation.
Key Points
• Software-based brain atrophy estimation often relies on normal cohorts for comparisons.
• At least 15 subjects must be included in an age- and sex-specific normal cohort.
• Using different normal cohorts does not influence regional atrophy estimation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Brain atrophy plays a critical role in the progression and diagnosis of various neurodegenerative diseases, such as Alzheimer’s disease (AD) [1], and frontotemporal dementia [2], among others [3,4,5]. Moreover, brain volume changes are increasingly important for treatment monitoring, such as in multiple sclerosis [6]. However, detecting regional brain volume alterations on MRI, particularly subtle volume losses in the early stages of a disease, can be challenging and is subject to high inter-reader variation [7]. Software-augmented evaluations have demonstrated the potential to reduce this variation [8], which is desirable for accurate diagnosis and treatment monitoring.
Various software approaches are available to aid in detecting and quantifying brain volume changes [9, 10]. Exemplary software includes icobrain dm (icometrix), BIOMETRICA (jung diagnostics), NeuroQuant (Cortechs.ai), Quantib ND (Quantiv), and volBrain (free online tool, https://volbrain.upv.es/ [11]). These tools primarily provide volumes (e.g., in cm3) of larger-scale structures, such as the frontal lobe, often in the context of normal percentile curves. Approaches like VEOmorph (VEObrain) [8], VSRAD (Eisai) [12], and veganbagel (Open source, https://github.com/BrainImAccs/veganbagel) [13] derive voxel-wise z-score statistics based on (matched, in the case of veganbagel) normal cohorts and offer region-of-interest-based z-scores (VEOmorph and VSRAD) or color-coded overlays for interpretation (VEOmorph, VSRAD, and veganbagel). A key difference lies in the interpretability of the results, as color-coded atrophy maps allow for a more refined assessment of atrophy patterns.
One critical aspect of brain atrophy estimation is the use of normal cohorts for comparison. Depending on the approach, a patient may be evaluated in the context of the whole normal cohort, or may be matched to a subset of subjects in the normal cohort, considering factors such as age, sex, and potentially other factors like the scanner model [14,15,16,17,18]. The need for high-quality normal cohorts, ideally well-matched to the local setting, is widely recognized. However, the minimum required number of healthy subjects contributing to a normal cohort for consistent atrophy estimation and the effect of using different normal cohorts on diagnostic reliability have not been well-established in the literature.
Considering these research gaps, this study aims to:
-
1.
determine the minimum number of subjects needed for consistent brain atrophy estimation when using age- and sex-specific normal cohorts, and
-
2.
evaluate the effect of using different normal cohorts on detecting regional atrophy patterns using the mesiotemporal atrophy pattern in AD patients as an example.
By addressing these objectives, our study aims to contribute to a better understanding of the factors influencing the accuracy of automated brain atrophy estimation tools and provide insights into optimizing their use in a clinical setting.
Methods
The retrospective study has been approved by the local ethics committee (#2021-1424). The need for written informed consent was waived.
Software for atrophy estimation
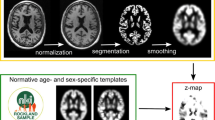
The open-source software veganbagel [13], an automated workflow for generating atrophy maps relative to age- and sex-specific normal templates, was adapted for the analysis. The workflow is depicted in Fig. 1. The Docker-based version of veganbagel was used (https://github.com/BrainImAccs/veganbagel, commit 6a2ac5f), which employs the standalone versions of CAT12.7 (r1713) [19] and SPM12 (version 7771) [20], eliminating the need for a MATLAB-license.
Visualization of the veganbagel workflow. Briefly, standardized preprocessing of structural T1-weighted imaging of subjects from a normal cohort is performed, comprising gray matter normalization, segmentation, modulation, and spatial smoothing using CAT12 for SPM12 with default settings. After preprocessing of healthy subjects, voxel-wise mean and standard deviation (SD) are computed for each sex and age (containing the actual age ±2 years), resulting in age- and sex-specific normal templates (green box). Voxel-wise z-score maps (= “atrophy maps”, red box) are then calculated for equally preprocessed subjects (yellow box), which express deviations from the age- and sex-specific normal templates. Atrophy maps may be inversely transformed into subject space and color-coded to generate overlays, with an example from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) for a male aged 63 years of age suffering from Alzheimer’s disease shown on the right
Consistent brain atrophy estimation
To establish the minimum number of healthy subjects needed for consistent atrophy detection, all healthy subjects from five different public cohorts were included into a pooled normal cohort (NCpool), if they met the following criteria: (a) age and sex were known; (b) a structural 3D T1-weighted dataset of the brain with a slice thickness of ≤ 1.5 mm was available; (c) the scan passed the cohort-internal quality control, if applicable; and (d) preprocessing with CAT12 was successful. The normal cohorts comprised the Lifespan Human Connectome Project Aging (HCP-A, started 2009, ongoing [21]), Information eXtraction from Images (IXI, 2005-2006), Nathan Kline Institute–Rockland Sample (Rockland, data sharing started 2010, ongoing [22]) as well as the healthy controls (HC) from the Alzheimer’s Disease Neuroimaging Initiative (ADNI, 2003, ongoing [23]) and the Open Access Series of Imaging Studies 3 (OASIS-3, published 2019, ongoing [24]). The ADNI was launched in 2003 as a public-private partnership, led by Principal Investigator Michael W. Weiner, MD (http://www.adni-info.org/). If there were multiple visits in the study, the first visit was used.
Patients with AD were retrieved from the ADNI database to serve as a surrogate for patients with brain atrophy. Patients with AD were included similar to the healthy subjects, but only if (a) structural imaging with a slice thickness ≤ 1 mm was available, and (b) based on the patients’ sex and age, there were ≥ 100 subjects of the same sex and age ± 2 years available in NCpool (see below).
To establish the minimum number of healthy subjects needed for consistent atrophy estimation, we performed an iterative process on the local High Performance Computing cluster. The process involved repeatedly calculating atrophy maps for each patient with AD using different normal templates, which were dynamically created using an increasingly larger number of randomly selected healthy subjects from NCpool. Healthy subjects were selected at random from NCpool to minimize effects of different scanners, sites, and cohorts. Furthermore, the whole process was repeated multiple times. A measure of the variance of the z-scores within the atrophy maps is then taken and plotted over the respective number of subjects contributing to the normal templates. We expected to see a considerable variance in z-scores with a smaller number of healthy subjects contributing to the normal templates, followed by a steady decrease and, finally, a plateau phase [25].
A detailed overview of the iterative process, which was applied to each patient with AD, can be found in Fig. 2. The process began with the random selection of three age- (± 2 years) and sex-matched healthy subjects from NCpool. Following the veganbagel methodology, these subjects were used to create mean and standard deviation (SD) normal templates, which were subsequently employed to compute z-score maps for the patient with AD. This entire procedure was carried out 100 times, with each iteration involving the random selection of three new eligible subjects from NCpool to create a fresh normal template. This resulted in 100 distinct z-score maps for each AD patient. The voxel-wise SD of z-score maps was calculated across the 100 repeats, and in a second step the spatial mean of SD was determined (\({\overline{SD} }_{{\text{spatial}}}\)), representing the consistency of brain atrophy estimates.
An overview of the process to establish the minimum number of healthy control subjects needed for consistent atrophy estimation. The process was repeated for every patient with AD included in this study. *Healthy subjects of the same sex and ± 2 years of age of the patient with AD were deemed eligible
For every AD patient, the process started with three healthy subjects forming the random dynamic normal templates, as described above. This number was incrementally increased by adding one healthy subject at a time (e.g., four subjects drawn, with the procedure above to be repeated 100 times) with up to 100 healthy subjects ultimately contributing to the random dynamic templates. Each subject was included in the normal templates only once, even if multiple scans were available (e.g., due to in-session repeat imaging within ADNI), but may be repetitively included during the 100 repeats. The number of repeats and the upper limit for subjects within the normal templates were informed by results from a prior veganbagel study [13], which utilized templates composed of 10 to 61 subjects. To ensure greater flexibility and comprehensiveness, our study broadened these parameters. The lower limit of three subjects contributing to the normal templates was established, since less than three subjects contributing to the normal templates was determined to yield an unrealistically high variance.
\({\overline{SD} }_{{\text{spatial}}}\) was plotted against the number of randomly selected subjects contributing to the normal templates. The “Kneedle” approach was used to determine the knee point of each curve, which involves fitting a smoothing spline to the data, normalizing, and finding the largest distance to a diagonal between the maximum and minimum of the data (https://github.com/etam4260/kneedle.) [26]. The maximum of the knee points across all patients, representing the point of diminishing returns when adding more normal subjects to the normal templates, was defined as the minimum number of subjects required for consistent results in brain atrophy evaluation.
Effect of different normal cohorts
To test the effect of using different normal cohorts for atrophy estimation on diagnostic reliability, we identified AD patients from the ADNI database for whom the previously established minimum number of age- and sex-specific subjects were available in each of the available non-ADNI normal cohorts (HCP-A, IXI, OASIS-3, and Rockland). HCs from the ADNI database were matched to the patients with AD based on age, sex, and scanner. We generated color-coded atrophy maps for each patient with AD and HC subject using veganbagel, separately using each normal cohort.
The atrophy maps were independently reviewed for the severity of mesiotemporal atrophy in a randomized order by two neuroradiologists with nine years of experience each (C.R. and J.C.), blinded to diagnosis and underlying normal cohort. Mesiotemporal atrophy is both a predictive and prognostic value in AD [27,28,29]. In the context of the study, it was rated for each hemisphere on a Likert scale, comprised of the following items: 0 = no atrophy, 1 = minimal to moderate atrophy (i.e., a few voxels of atrophy, as indicated by the atrophy map), 2 = marked atrophy (more prominent areas of volume loss, as noted in the atrophy map), 3 = severe atrophy (large areas of volume loss including voxels with z-scores ≥ 10). An AD diagnosis was assigned when the bihemispheric score was ≥ 2.
Inter-reader reliability was computed using Cohen’s Kappa, and sensitivity and specificity for the score-based AD diagnosis was determined for each normal cohort. To assess the agreement across the different normal cohorts, a two-way intraclass correlation coefficient was calculated. Cochran’s Q test and a pairwise McNemar test with Bonferroni correction were performed to compare the results. p < 0.05 was considered statistically significant. Statistical analysis was done using R v4.0.3 [30].
Data availability
All data used in the manuscript is either publicly available or available to qualified researchers from the respective cohort’s database (see “Acknowledgments”).
Results
Consistent brain atrophy estimation
The pooled normal cohort (NCpool) consisted of 3945 healthy subjects (55 ± 21 years, 57.9% female, Table 1, Figs. 3 and 4). A total of 48 patients with AD were included in the analysis (73 ± 7 years (range 55–82), 37.5% female, Fig. 3). Thirteen AD patients were scanned using a GE scanner, nine on Philips scanners, and 26 on Siemens scanners. A total of 27 different 3-T scanners were used for the AD patients.
Subjects included into the pooled normal cohort (to the left of the dotted line) and patients with Alzheimer’s disease from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) used for determining the minimum number of subjects to include into a normal cohort (to the right of the dotted line). *From the pooled normal cohort. HCP-A, Lifespan Human Connectome Project Aging; IXI, Information eXtraction from Images; OASIS-3, Open Access Series of Imaging Studies 3; Rockland, Nathan Kline Institute–Rockland Sample; QC, quality control
In all AD patients, a sharp drop of the \({\overline{SD} }_{{\text{spatial}}}\) was noted at a small number of healthy subjects included in the normal templates (Fig. 5). The knee points varied across patients, with the smallest at 9 subjects and the largest at 15 subjects (average 11.0 ± 1.2, median 11, inter quartile range 10 to 12). The minimum number of subjects required for consistent results in brain atrophy evaluation was therefore 15, with a corresponding \({\overline{SD} }_{{\text{spatial}}}\) of the z-scores of 0.34 ± 0.026 (range 0.297 to 0.432) across all patients.
Mean standard deviation of the voxel-wise z-scores over all repeats plotted over the number of randomly drawn subjects included in the dynamically generated normal. The maximum of all established knee points (= 15), representing the point of diminishing returns when adding more normal subjects to the normal templates, is denoted by the dashed black lines. Female patients are shown on the left and male patients on the right
Effect of different normal cohorts
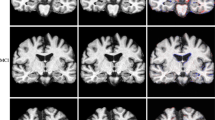
A subset of 21 patients with AD and 21 matched HCs had more than 15 healthy subjects available in the HCP-A, IXI, OASIS-3, and Rockland normal cohorts (Table 2). The inter-reader reliability between the two neuroradiologists was high, with an overall Cohen’s Kappa of 0.98 for the extent of the atrophy as determined on the visual rating scale (Table 3). For the individual cohorts, the Cohen’s Kappa was 1 for HCP-A and Rockland, 0.98 for IXI, and 0.93 for OASIS-3. Fig. 6 shows an example of each atrophy map derived.
Example of the color-coded z-score maps (= atrophy maps) calculated using veganbagel for a female patient with Alzheimer’s disease (AD), aged 67 years, derived using the four different normal cohorts. The color-coded z-score maps are overlaid onto the original 3D T1-weighted MRI acquisitions, shown in the axial plane
The readers agreed in the diagnosis of AD and HC in all cases. Table 4 lists the respective accuracy, sensitivities, and specificities, as well as the positive and negative predictive value for each reader and normal cohort. The intraclass correlation coefficient across the cohorts was 0.91. Cochran’s Q test did not show a significant difference across the different cohorts (p = 0.19). Likewise, no significant differences were found in the Bonferroni-corrected pairwise McNemar tests (HCP-A/IXI: p = 0.48; HCP-A/OASIS-3: p = 0.48; HCP-A/Rockland: p = 1; IXI/OASIS-3: p = 1; IXI/Rockland: p = 0.48; OASIS-3/Rockland: p = 0.48).
Discussion
Our study aimed to determine the minimum number of subjects required in a normal cohort for consistent software-based brain atrophy estimation and to evaluate the impact of using different normal cohorts on the qualitative assessment of mesiotemporal atrophy in Alzheimer’s disease. We found that at least 15 healthy subjects should be included in an age- and sex-specific normal cohort for consistent atrophy detection, and that using different normal cohorts does not significantly influence the qualitative evaluation of mesiotemporal atrophy or the imaging-based diagnosis of Alzheimer’s disease.
In our study, we evaluated the open-source software veganbagel [13], which implements an automated assessment of deviation in gray matter volume from a normal cohort using voxel-wise color-coded z-score maps. Veganbagel is based on proven methods, namely voxel-based morphometry (VBM) using CAT12 for SPM12. VBM reliably detects patterns in various contexts such as normal aging, neurodegenerative diseases, and psychiatric disorders [15, 31, 32]. More specifically, VBM-based approaches have proven valuable in detecting AD [1] and are routinely used in the clinical diagnosis of AD in Japan [33].
To conduct our analysis with a sufficiently large pool of healthy subjects, we created a pooled normal cohort consisting of subjects from five different normal cohorts with varying objectives, scanners, protocols, and quality. Previous studies have reported differences in quantitative brain atrophy estimation due to factors such as different scanners or protocols [16, 17], while others have demonstrated that volumes of subcortical structures may be interchangeable across different normal cohorts [34]. Given the objective of our analysis, we focused on sex and age as the main influences on brain volume and minimized the potential impact of other factors by not only randomly drawing subjects from the pooled normal cohort, but also by repeating the process 100 times, starting with three and ultimately including up to 100 healthy subjects contributing to the random dynamic normal templates in a computationally intensive approach.
Our study’s results may also enhance radiologists’ comprehension of the mechanisms underpinning automated brain atrophy estimation. Additionally, these findings can guide the decisions-making process when considering commercial solutions, especially in questioning undisclosed, inadequately sized, or insufficiently assessed normal cohorts.
Minimum number of subjects in a normal cohort
Previous findings in nuclear medicine have suggested a minimum number of 10–20 subjects for a normal cohort when evaluating brain glucose metabolism in the diagnostic workup of dementia [25, 35]. However, the minimum number of subjects in a normal cohort for consistent MRI-based brain atrophy estimation has not been established. Generally, it is assumed that a normal cohort must be as large as possible and as well adapted to a patient as possible with regard to sex, age, scanner, protocol, artifacts, and possibly other factors such as ethnicity or cultural background [14,15,16,17,18, 36].
It is important to recognize that no method for identifying a knee point of a curve is universally accepted, and all approaches rely on approximations dependent on various parameters. A definitive, objective threshold for \({\overline{SD} }_{{\text{spatial}}}\) would be ideal. However, it is important to recognize that any chosen cutoff might possess an element of arbitrariness. In our study, we observed that the variance in z-scores sharply diminishes and approaches 0 as more subjects are included in the normal templates. Given that the identified maximum knee point aligns visually with the knee point determined by the Kneedle method, we are confident that, within the scope of the study, a minimum 15 normal subjects is needed for consistent brain atrophy estimation.
Impact of different normal cohorts on atrophy detection
In the context of our study, we found that qualitative interpretation of regional mesiotemporal atrophy allowed for reliable AD diagnosis when using the different normal cohorts. Our current study outperforms the previously reported sensitivity and specificity for detection of AD in ADNI using veganbagel [13], likely due to evaluating a much smaller sample of patients in the current study. However, the current analysis is focused on the comparison of different normal cohorts, rather than diagnostic accuracy, which allowed for narrower inclusion/exclusion criteria. Nevertheless, the inter-reader agreement for the extent of mesiotemporal atrophy was excellent, which is notable since atrophy assessment on MRI without any software augmentation has been shown to have a low inter-reader agreement [7].
Limitations and future directions
The limitations of our study include the moderate sample size in the qualitative evaluation of the mesiotemporal atrophy and the evaluation of only one software approach (veganbagel). Other software for brain atrophy estimation, to our knowledge, either is not openly available or does not lend itself to the modifications needed for the conducted analyses. In the case of other open-source alternatives to CAT12/SPM12, such as the FSL or FreeSurfer, no fully integrated software packages for brain atrophy estimation are currently available.
As the number of subjects in the normal templates grows, there is a heighted probability that the same subjects may be repetitively included during the 100 iterations. However, considering the established minimum of 15 subjects and the study’s prerequisite for at least 100 age- and sex-matched subjects for every patient, these overlap likely do not distort our primary conclusions.
The current study leveraged five extensive normal cohorts, enriching the data variety. Yet, these cohorts predominately represent the population of the north-western regions of the world. It is paramount that subsequent studies address the applicability of our findings to the global demographic.
The structure of our experiment, especially its emphasis on numerous iterations, tends to mitigate outlier impacts—whether these outliers arise from atypical, presumed “normal” subjects or from subjects ill-matched to a given patient due to diverse imaging environments. There is an evident need for more focused studies on the resilience of normal templates, particularly those derived from a limited set of subjects.
Future studies should focus on the effects of combining different normal cohorts. A pooled normal cohort for clinical brain atrophy estimation may allow to recruit a sufficient number of healthy subjects for brain atrophy estimation at more extreme ages. Furthermore, a very large and heterogeneous normal cohort would enable more precise matching of patients with regard to factors such as scanners and protocols, enhancing the detection of subtle regional brain atrophy. Last, but not least, patients with other forms of neurodegenerative diseases should be evaluated to ensure that the findings are generalizable across different populations and clinical contexts.
Conclusion
In summary, our study indicates that a normal cohort should include at least 15 normal subjects, matched for age and sex, to consistently estimate brain atrophy using voxel-based morphometry. In the context of our study, using different normal cohorts did not significantly influence the qualitative assessment of regional mesiotemporal atrophy or the diagnosis of Alzheimer’s disease, and we observed a high inter-reader agreement. It is important to note, however, that these findings are influenced by our study’s particular design and parameters. Thus, caution is necessary when extrapolating these findings to other contexts without fully understanding the inherent assumptions and potential confounding factors.
Abbreviations
- AD:
-
Alzheimer’s disease
- ADNI:
-
Alzheimer’s Disease Neuroimaging Initiative
- HC:
-
Healthy control
- HCP-A:
-
Lifespan Human Connectome Project Aging
- IXI:
-
Information eXtraction from Images
- NCpool :
-
Pooled normal cohort
- OASIS-3:
-
Open Access Series of Imaging Studies 3
- Rockland:
-
Nathan Kline Institute–Rockland Sample
- SD:
-
Standard deviation
- VBM:
-
Voxel-based morphometry
References
Hedderich DM, Dieckmeyer M, Andrisan T et al (2020) Normative brain volume reports may improve differential diagnosis of dementing neurodegenerative diseases in clinical practice. Eur Radiol 30:2821–2829. https://doi.org/10.1007/s00330-019-06602-0
Fumagalli GG, Basilico P, Arighi A et al (2018) Distinct patterns of brain atrophy in Genetic Frontotemporal Dementia Initiative (GENFI) cohort revealed by visual rating scales. Alzheimers Res Ther 10:46. https://doi.org/10.1186/s13195-018-0376-9
Johnson EB, Gregory S (2019) Huntington’s disease: brain imaging in Huntington’s disease. Prog Mol Biol Transl Sci 165:321–369. https://doi.org/10.1016/bs.pmbts.2019.04.004
Reetz K, Gaser C, Klein C et al (2009) Structural findings in the basal ganglia in genetically determined and idiopathic Parkinson’s disease. Mov Disord 24:99–103. https://doi.org/10.1002/mds.22333
Boxer AL, Geschwind MD, Belfor N et al (2006) Patterns of brain atrophy that differentiate corticobasal degeneration syndrome from progressive supranuclear palsy. Arch Neurol 63:81–86. https://doi.org/10.1001/archneur.63.1.81
Sastre-Garriga J, Pareto D, Battaglini M et al (2020) MAGNIMS consensus recommendations on the use of brain and spinal cord atrophy measures in clinical practice. Nat Rev Neurol 16:171–182. https://doi.org/10.1038/s41582-020-0314-x
Scheltens P, Pasquier F, Weerts JG et al (1997) Qualitative assessment of cerebral atrophy on MRI: inter- and intra-observer reproducibility in dementia and normal aging. Eur Neurol 37:95–99. https://doi.org/10.1159/000117417
Kloppel S, Yang S, Kellner E et al (2018) Voxel-wise deviations from healthy aging for the detection of region-specific atrophy. NeuroImage Clin 20:851–860. https://doi.org/10.1016/j.nicl.2018.09.013
Scarpazza C, Ha M, Baecker L et al (2020) Translating research findings into clinical practice: a systematic and critical review of neuroimaging-based clinical tools for brain disorders. Transl Psychiatry 10:107. https://doi.org/10.1038/s41398-020-0798-6
Pemberton HG, Zaki LAM, Goodkin O et al (2021) Technical and clinical validation of commercial automated volumetric MRI tools for dementia diagnosis—a systematic review. Neuroradiology: 1–17. https://doi.org/10.1007/s00234-021-02746-3
Manjón JV, Coupé P (2016) volBrain: an online MRI Brain Volumetry System. Front Neuroinform 10:30. https://doi.org/10.3389/fninf.2016.00030
Matsuda H, Mizumura S, Nemoto K et al (2012) Automatic voxel-based morphometry of structural MRI by SPM8 plus diffeomorphic anatomic registration through exponentiated lie algebra improves the diagnosis of probable Alzheimer Disease. AJNR Am J Neuroradiol 33:1109–1114. https://doi.org/10.3174/ajnr.a2935
Caspers J, Heeger A, Turowski B, Rubbert C (2021) Automated age- and sex-specific volumetric estimation of regional brain atrophy: workflow and feasibility. Eur Radiol 31:1043–1048. https://doi.org/10.1007/s00330-020-07196-8
Kurth F, Thompson PM, Luders E (2018) Investigating the differential contributions of sex and brain size to gray matter asymmetry. Cortex 99:235–242. https://doi.org/10.1016/j.cortex.2017.11.017
Good CD, Johnsrude IS, Ashburner J et al (2001) A voxel-based morphometric study of ageing in 465 normal adult human brains. Neuroimage 14:21–36. https://doi.org/10.1006/nimg.2001.0786
Leung KK, Malone IM, Ourselin S et al (2015) Effects of changing from non-accelerated to accelerated MRI for follow-up in brain atrophy measurement. Neuroimage 107:46–53. https://doi.org/10.1016/j.neuroimage.2014.11.049
Dieckmeyer M, Roy AG, Senapati J et al (2021) Effect of MRI acquisition acceleration via compressed sensing and parallel imaging on brain volumetry. MAGMA. https://doi.org/10.1007/s10334-020-00906-9
Huang C-M, Doole R, Wu CW et al (2019) Culture-related and individual differences in regional brain volumes: a cross-cultural voxel-based morphometry study. Front Hum Neurosci 13:313. https://doi.org/10.3389/fnhum.2019.00313
Gaser C, Dahnke R, Thompson PM et al (2023) CAT – A computational anatomy toolbox for the analysis of structural MRI data. bioRxiv 2022.06.11.495736. https://doi.org/10.1101/2022.06.11.495736
Penny W, Friston K, Ashburner J et al (2006) Statistical parametric mapping : the analysis of functional brain images. Elsevier / Academic Press, Amsterdam, Boston
Bookheimer SY, Salat DH, Terpstra M et al (2019) The Lifespan Human Connectome Project in Aging: an overview. Neuroimage 185:335–348. https://doi.org/10.1016/j.neuroimage.2018.10.009
Nooner KB, Colcombe SJ, Tobe RH et al (2012) The NKI-Rockland Sample: a model for accelerating the pace of discovery science in psychiatry. Front Neurosci 6:152. https://doi.org/10.3389/fnins.2012.00152
Mueller SG, Weiner MW, Thal LJ et al (2005) The Alzheimer’s disease neuroimaging initiative. Neuroimaging Clin N Am 15:869-77-xi–xii. https://doi.org/10.1016/j.nic.2005.09.008
LaMontagne PJ, Benzinger TLS, Morris JC et al (2019) OASIS-3: longitudinal neuroimaging, clinical, and cognitive dataset for normal aging and Alzheimer disease. Medrxiv 2019.12.13.19014902. https://doi.org/10.1101/2019.12.13.19014902
Buchert R (2008) On the effect of sample size of the normal database on statistical power of single subject analysis. Nucl Med Commun 29:837. https://doi.org/10.1097/mnm.0b013e3283023f8d
Satopää V, Albrecht J, Irwin D, Raghavan B (2011) Finding a “Kneedle” in a haystack: detecting knee points in system behavior. 2011 31st Int Conf Distrib Comput Syst Work 166–171. https://doi.org/10.1109/icdcsw.2011.20
Jack CR, Knopman DS, Jagust WJ et al (2013) Tracking pathophysiological processes in Alzheimer’s disease: an updated hypothetical model of dynamic biomarkers. Lancet Neurol 12:207–216. https://doi.org/10.1016/s1474-4422(12)70291-0
Patel KP, Wymer DT, Bhatia VK et al (2020) Multimodality imaging of dementia: clinical importance and role of integrated anatomic and molecular imaging. Radiographics 40:200–222. https://doi.org/10.1148/rg.2020190070
Scheltens P, Leys D, Barkhof F et al (1992) Atrophy of medial temporal lobes on MRI in “probable” Alzheimer’s disease and normal ageing: diagnostic value and neuropsychological correlates. J Neurology Neurosurg Psychiatry 55:967. https://doi.org/10.1136/jnnp.55.10.967
R Core Team (2022) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Honea R, Crow TJ, Passingham D, Mackay CE (2005) Regional deficits in brain volume in schizophrenia: a meta-analysis of voxel-based morphometry studies. Am J Psychiatry 162:2233–2245. https://doi.org/10.1176/appi.ajp.162.12.2233
Minkova L, Habich A, Peter J et al (2017) Gray matter asymmetries in aging and neurodegeneration: a review and meta-analysis. Hum Brain Mapp 38:5890–5904. https://doi.org/10.1002/hbm.23772
Matsuda H (2016) MRI morphometry in Alzheimer’s disease. Ageing Res Rev 30:17–24. https://doi.org/10.1016/j.arr.2016.01.003
Vinke EJ, Huizinga W, Bergtholdt M et al (2019) Normative brain volumetry derived from different reference populations: impact on single-subject diagnostic assessment in dementia. Neurobiol Aging 84:9–16. https://doi.org/10.1016/j.neurobiolaging.2019.07.008
Chen W-P, Samuraki M, Yanase D et al (2008) Effect of sample size for normal database on diagnostic performance of brain FDG PET for the detection of Alzheimer’s disease using automated image analysis. Nucl Med Commun 29:270–276. https://doi.org/10.1097/mnm.0b013e3282f3fa76
Reuter M, Tisdall MD, Qureshi A et al (2015) Head motion during MRI acquisition reduces gray matter volume and thickness estimates. Neuroimage 107:107–115. https://doi.org/10.1016/j.neuroimage.2014.12.006
Acknowledgements
Computational infrastructure and support was provided by the Center for Information and Media Technology (ZIM) at the Heinrich Heine University of Duesseldorf (Germany).
ADNI: Collection and sharing of the Alzheimer’s Disease Neuroimaging Initiative (ADNI) data used for evaluation in this study was funded by the (National Institutes of Health Grant U01 AG024904) and DOD ADNI (Department of Defense award number W81XWH-12-2-0012). ADNI is funded by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: AbbVie, Alzheimer’s Association; Alzheimer’s Drug Discovery Foundation; Araclon Biotech; BioClinica, Inc.; Biogen; Bristol-Myers Squibb Company; CereSpir, Inc.; Cogstate; Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffmann-La Roche Ltd and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Alzheimer Immunotherapy Research & Development, LLC.; Johnson & Johnson Pharmaceutical Research & Development LLC.; Lumosity; Lundbeck; Merck & Co., Inc.; Meso Scale Diagnostics, LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Takeda Pharmaceutical Company; and Transition Therapeutics. The Canadian Institutes of Health Research is providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (https://fnih.org). The grantee organization is the Northern California Institute for Research and Education, and the study is coordinated by the Alzheimer’s Therapeutic Research Institute at the University of Southern California. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California. Data is available from http://adni.loni.usc.edu.
HCP-A: Research reported in this publication was supported by the National Institute On Aging of the National Institutes of Health under Award Number U01AG052564. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. Data is available from https://www.humanconnectome.org/study/hcp-lifespan-aging.
IXI: The Information eXtraction from Images (IXI) data is available under the CC BY-SA 3.0 license from https://brain-development.org/ixi-dataset/.
OASIS-3: Data were provided in part by OASIS-3: Principal Investigators: T. Benzinger, D. Marcus, J. Morris; NIH P50 AG00561, P30 NS09857781, P01 AG026276, P01 AG003991, R01 AG043434, UL1 TR000448, R01 EB009352. Data is available from https://www.oasis-brains.org.
Rockland: The Enhanced Nathan Kline Institute–Rockland Sample (NKI-RS) is available from http://fcon-1000.projects.nitrc.org/indi/enhanced/. See Nooner KB et al Front Neurosci. 2012;6:152. https://doi.org/10.3389/fnins.2012.00152.
Funding
Open Access funding enabled and organized by Projekt DEAL. Robert Dahnke was funded by the DFG project DA 2167/1-1.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Christian Rubbert.
Conflict of interest
The authors of this manuscript declare no relationships with any companies, whose products or services may be related to the subject matter of the article.
Statistics and biometry
Two of the authors have significant statistical expertise (CR and JC).
Informed consent
Written informed consent was waived by the Institutional Review Board.
Ethical approval
Institutional Review Board approval was obtained.
Study subjects or cohorts overlap
Our study is based on five large and well-known public datasets (the Alzheimer’s Disease Neuroimaging Initiative, the Lifespan Human Connectome Project Aging, the Information eXtraction from Images, the Nathan Kline Institute–Rockland Sample, and the Open Access Series of Imaging Studies); therefore, all subjects have extensively reported on before.
Methodology
• retrospective
• cross-sectional study
• multicenter study
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Data used in preparation of this article were obtained from the Alzheimer's Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at: http://adni.loni.usc.edu/wp-content/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rubbert, C., Wolf, L., Vach, M. et al. Normal cohorts in automated brain atrophy estimation: how many healthy subjects to include?. Eur Radiol 34, 5276–5286 (2024). https://doi.org/10.1007/s00330-023-10522-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-023-10522-5