Abstract
Objectives
Risk calculators (RCs) improve patient selection for prostate biopsy with clinical/demographic information, recently with prostate MRI using the prostate imaging reporting and data system (PI-RADS). Fully-automated deep learning (DL) analyzes MRI data independently, and has been shown to be on par with clinical radiologists, but has yet to be incorporated into RCs. The goal of this study is to re-assess the diagnostic quality of RCs, the impact of replacing PI-RADS with DL predictions, and potential performance gains by adding DL besides PI-RADS.
Material and methods
One thousand six hundred twenty-seven consecutive examinations from 2014 to 2021 were included in this retrospective single-center study, including 517 exams withheld for RC testing. Board-certified radiologists assessed PI-RADS during clinical routine, then systematic and MRI/Ultrasound-fusion biopsies provided histopathological ground truth for significant prostate cancer (sPC). nnUNet-based DL ensembles were trained on biparametric MRI predicting the presence of sPC lesions (UNet-probability) and a PI-RADS-analogous five-point scale (UNet-Likert). Previously published RCs were validated as is; with PI-RADS substituted by UNet-Likert (UNet-Likert-substituted RC); and with both UNet-probability and PI-RADS (UNet-probability-extended RC). Together with a newly fitted RC using clinical data, PI-RADS and UNet-probability, existing RCs were compared by receiver-operating characteristics, calibration, and decision-curve analysis.
Results
Diagnostic performance remained stable for UNet-Likert-substituted RCs. DL contained complementary diagnostic information to PI-RADS. The newly-fitted RC spared 49% [252/517] of biopsies while maintaining the negative predictive value (94%), compared to PI-RADS ≥ 4 cut-off which spared 37% [190/517] (p < 0.001).
Conclusions
Incorporating DL as an independent diagnostic marker for RCs can improve patient stratification before biopsy, as there is complementary information in DL features and clinical PI-RADS assessment.
Clinical relevance statement
For patients with positive prostate screening results, a comprehensive diagnostic workup, including prostate MRI, DL analysis, and individual classification using nomograms can identify patients with minimal prostate cancer risk, as they benefit less from the more invasive biopsy procedure.
Key Points
-
The current MRI-based nomograms result in many negative prostate biopsies. The addition of DL to nomograms with clinical data and PI-RADS improves patient stratification before biopsy.
-
Fully automatic DL can be substituted for PI-RADS without sacrificing the quality of nomogram predictions.
-
Prostate nomograms show cancer detection ability comparable to previous validation studies while being suitable for the addition of DL analysis.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Recently, multiparametric magnetic resonance imaging (mpMRI) has been established for identification and for targeted prostatic lesion biopsy through cognitive fusion [1], the stereotactic fusion of transrectal ultrasound (TRUS), and MRI [2], or in-bore MRI techniques [3]. With this approach, detection of significant prostate cancer (sPC) and enrollment into active surveillance have improved [4], while the approach promises to spare biopsies altogether in certain men [5, 6]. With growing evidence for the benefits of prostate MRI in prospective multi-center trials [7,8,9], guidelines increasingly recommend mpMRI prior to biopsy in biopsy-naïve or pre-biopsied men [10]. The decision whom to biopsy can be supported by risk calculators (RCs) incorporating demographic and clinical information such as age, digital rectal exam (DRE), prostate-specific antigen (PSA), and prostate volume, e.g., RCs from the European Randomized Study of Screening for Prostate Cancer (ERSPC) combine this information in logistic regression models [11] and visualize it using nomograms [12]. The positive predictive value (PPV) of clinical radiologist prostate MRI assessment using the prostate imaging reporting and data system (PI-RADS) [13] is limited, variable, and typically reported between 30% and 50% across different centers [14,15,16], resulting in many negative biopsies. RCs promise to improve patient selection before biopsy, which carries the risk of infection, bleeding, and hospitalization [17]. Recently, it has been shown that RCs benefit from the addition of PI-RADS [18]. Several such RCs have been proposed, none of which exhibits clear advantages over another at the current time [10], while there are concerns about calibration shifts over time and across institutions, reducing their benefits. Simultaneously, fully-automated analysis of prostate MRI using machine learning in the form of deep learning (DL) by convolutional neural networks has recently been demonstrated to provide sPC detection similar to clinical PI-RADS assessment by radiologists [19, 20]. Self-configuring network architectures for semantic image segmentation [21] and object detection [22] can adapt to a wide range of medical imaging modalities. The UNet DL architecture [23] has become especially popular in medical image segmentation [19,20,21]. Resulting networks indicate the spatial location of identified suspicious findings, allowing comparison to radiologist-identified regions [21, 22]. DL-based prostate MRI assessment carries the potential to make risk assessment tools more reproducible and to foster more widespread application of fully automatic image analysis. We hypothesized that risk estimation with logistic regression models based on demographic and clinical data but with the addition of fully-automated DL image assessment would be capable of performing similarly to previously established risk models using clinical PI-RADS assessment. The goal of our study was to re-evaluate established RCs for prostate cancer risk assessment using a large consecutive cohort from our institution to determine if clinical PI-RADS assessment and fully-automated DL prostate MRI assessment perform similarly in such models, and whether the additional benefit is obtained when both are combined.
Material and methods
Study sample
Multiparametric prostate MRI examinations between September 2014 and June 2021 were consecutively included in this retrospective single-center study if patients received combined extended systematic and targeted prostate biopsy and mpMRI at our institutions in Heidelberg, Germany. The institutional ethics committees approved the study and waived informed consent (S-164/2019). Exclusion criteria were (1) prior therapy for PCa (e.g., radiation therapy, ultrasound ablation); (2) previous or ongoing androgen-depriving treatment; (3) severe imaging artifacts; (4) previous prostate biopsy < 2 months ago or interval between MRI and subsequent biopsy > 6 months; and (5) unusually rare histopathology. Exams with any previous diagnosis of prostate cancer, including exams under Active Surveillance, were excluded from the risk calculator evaluation.
MRI protocol
Multiparametric MRI was performed using two 3.0 Tesla scanners (Megnetom Prisma, Biograph mMR; Siemens Healthineers) and one 1.5 Tesla MRI scanner (Magnetom Aera, Siemens Healthineers) based on PI-RADS recommendations [13, 24] and guidelines of the European Society of Urogenital Radiology [25, 26]. Examinations used the standard multichannel body coil and integrated spine phased-array coil. MRI acquisition parameters are detailed in Supplemental Table 1.
Biopsy scheme, PI-RADS assessment, and image segmentation
Radiologists assessed mpMRI according to PI-RADS v2.1 guidelines [13] during a clinical routine with access to previous reports, PSA levels, and dynamic contrast-enhanced T1-weighted images (DCE). After the multidisciplinary conference discussion, patients received extended systematic and targeted transperineal MRI/ultrasound-fusion biopsies matching the Ginsburg protocol [27]. For older cases, where only clinical PI-RADS v1 [25] assessments were available, previously-biopsied lesions from the original clinical report were reassessed with PI-RADS v2 by a board-certified radiologist without reviewing the biopsy result and with knowledge of the prior report, thus generating a consensus score using PI-RADS v2 criteria and preserving the relationship between MRI lesion and targeted biopsy result. Biopsies were guided by rigid or elastic software registration. Histopathology was assessed by a dedicated uropathologist with 18 years of experience (A.St.) and graded according to International Society of Urological Pathology (ISUP) standards [28]. sPC was defined as ISUP grade ≥ 2.
DL model configuration, training, calibration, and inference
Modified nnUNet models were trained for prostate and lesion segmentation with configuration details given in Supplemental Material S-1. Models use whole prostate bpMRI images for predictions and provide nnUNet softmax maps which provide voxel-based tumor probability and thus indicate both, lesion presence and localization. Activation maps were interpreted as the voxel-wise probability of finding sPC, with the maximum probability representing the patient-wise sPC probability score (UNet-probability). The continuous values of the UNet-probability were converted to a 5-point Likert scale (UNet-Likert). UNet-Likert thresholds were dynamically chosen [29] to target sensitivities or specificities similar to PI-RADS, as described in Supplemental Material S-2.
RCs and decision strategies
Patient age, DRE, PSA, fully automatic T2-weighted segmentation-based prostate volume, and previous biopsy results were available for analysis on each case so that data imputation was not necessary. RCs evaluated included Radtke et al [30], van Leeuwen et al [31], and Alberts et al [32] (MRI-ERSPC). For MRI-ERSPC and Radtke 2017, individual models for biopsy-naïve and previously-biopsied patients were considered. The PI-RADS/PSAD strategy defined low-risk exams as PI-RADS ≤ 2, or PI-RADS 3 with PSA density (PSAD) < 0.1 ng/mL [33] for comparison. To evaluate whether UNet-Likert provides comparable value to clinical PI-RADS assessment, PI-RADS was substituted by UNet-Likert in the RCs (UNet-Likert-substituted RCs), and for comparison in PI-RADS/PSAD (UNet-Likert/PSAD). As PI-RADS and DL may extract complementary information from image data, models combining DL with PI-RADS were calculated (UNet-probability-extended RCs), by estimating two-parameter logistic regression models including the RC score and UNet-probability, similar to the addition of age and PI-RADS to the ERSPC score [32]. Finally, to evaluate the most flexible parametrization, we fitted new multivariate logistic regression models not relying on fixed coefficients or parameter transforms (see Supplemental Material S-3 and Supplemental Table 2). We then used repeated 10-fold cross-validation on the training set to choose the candidate model with the highest cross-validation performance named the newly-fitted PI-RADS + UNet probability RC. Table 1 shows the parameters for each model.
Statistical analysis
Logistic regression models have been shown to be susceptible to calibration shifts when applied to datasets from new institutions [18, 34], thus, after establishing original and UNet-Likert-substituted model performance, adjustment of these models by intercept-only recalibration (a.k.a. recalibration in the large) and intercept/slope recalibration (logistic recalibration) on the training set was implemented [34,35,36]. In the UNet-probability-extended RCs, slope, and intercept were necessarily fitted as detailed in Supplemental Material S-4. As Radtke 2017 and MRI-ERSPC consist of separate models for biopsy-naïve or previously-biopsied patients, intercept-only recalibration, intercept/slope recalibration, and UNet-probability extension were done separately for each group. The newly-fitted PI-RADS + UNet probability RC is necessarily calibrated to the training data by its derivation. The exam-level predictive performance of Radtke 2017, Leeuwen 2017, and MRI-ERSPC-RCs was evaluated on the test set and compared to the UNet-Likert-substituted RCs, UNet-probability-extended RCs, and newly-fitted PI-RADS + UNet probability RC. Receiver operating characteristics (ROC) were used to evaluate calibration-independent sPC discrimination with area under the curve (AUC) comparisons. The Brier score was used for combined calibration-dependent evaluation of model calibration and discrimination. Calibration was further assessed using calibration plots and the ratio of average predicted risk to observed sPC rate (Exp/Obs-Ratio). Decision curve analysis (DCA), following the interpretation recommendations by Van Calster et al [37] for the opt-in decision to undergo targeted biopsy, was used to weigh the calibration-dependent benefit of correctly diagnosing a patient with sPC against the harm of over-diagnosing patients without sPC [38], with details given in Supplemental Material S-5. The p values were adjusted for multiple testing by Holm–Bonferroni correction. Statistical analysis was performed in R version 4.1.0.
Results
Study sample characteristics
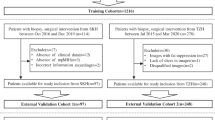
In total, 1627 MRI examinations were included which were temporally split in November 2018 into a training set of 1021 exams, used for DL training in 5-fold cross-validation, and 606 exams for independent testing. 834 exams in the training set and 517 exams in the test set had no previous prostate cancer diagnosis and were used for risk model estimation in cross-validation and subsequent testing. Figure 1 shows inclusion/exclusion criteria as a flowchart. Table 2 gives demographic and clinical characteristics. 1610 exams have been reported in previous publications on DL and radiomics [19, 20, 39], however, data have not been used for systematic clinical RC assessment or development. Regarding the PI-RADS/PSAD strategy, the test set of a previous study overlaps with 101 exams from biopsy-naïve patients with PI-RADS 3 lesions in the current study [33].
Diagram showing the cohort selection processing with inclusion and exclusion criteria according to STARD guidelines [53]. HIFU, high-intensity focused ultrasound; TULSA, transurethral ultrasound Ablation; STUMP, stromal tumor of uncertain malignant potential
Imaging-based performance (PI-RADS, DL) and PSA-heuristics
On the test set, clinical PI-RADS achieved 14% [44/317] specificity at 98% [197/200] sensitivity for PI-RADS ≥ 3, and 56% [178/317] specificity at 94% [188/200] sensitivity for PI-RADS ≥ 4. The PI-RADS/PSAD strategy achieved 39% [124/317] specificity at 97% [194/200] sensitivity.
UNet-probability alone achieved an AUC of 0.89 (95% CI: 0.86-0.92). UNet-Likert alone achieved 30% [94/317] specificity at 97% [194/200] sensitivity for UNet-Likert ≥ 3 and 57% [180/317] specificity at 92% [184/200] sensitivity for UNet-Likert ≥ 4. UNet-Likert/PSAD strategy achieved 45% [142/317] specificity at 96% [193/200] sensitivity. UNet-Likert ≥ 3 showed significantly higher specificity compared to PI-RADS ≥ 3 (p < 0.001) at similar sensitivities (p = 0.32). There was no significant difference in sensitivity or specificity between PI-RADS/PSAD and UNet-Likert/PSAD (p = 0.74 and p = 0.1, respectively), or between PI-RADS ≥ 4 and UNet-Likert ≥ 4 (p = 0.39 and p = 0.86, respectively).
Risk calculator performance (original, UNet-Likert-substituted)
Leeuwen 2017, Radtke 2017, and MRI-ERSPC original RCs achieved AUC of 0.90 (95% CI: 0.87–0.92), 0.89 (95% CI: 0.87–0.92), and 0.88 (95% CI: 0.85–0.91), while UNet-Likert-substituted RCs achieved 0.90 (95% CI: 0.88–0.93), 0.90 (95% CI: 0.87–0.93), and 0.90 (95% CI: 0.87–0.92), respectively (Fig. 2A, B). There was no significant difference in AUC between the three original RCs (p = 0.20) or UNet-Likert-substituted RCs (p = 0.26) in global testing, so neither Leeuwen 2017, Radtke 2017 nor MRI-ERSPC showed superior AUC. Also, the original and UNet-Likert-substituted versions of each RC showed no significant difference in AUC (p = 1.00).
ROCs indicating discrimination ability of RCs. Classification models with no skill would lie on the identity bisector (gray). A Original RCs (Leeuwen 2017 (red), Radtke 2017 (black), and MRI-ERSPC (green), compared to UNet-probability (yellow)); B UNet-Likert-substituted RCs compared to UNet-probability (yellow); C UNet-probability-extended RCs compared to UNet-probability (yellow); D, F differences between original (black), UNet-Likert-substituted (red), and UNet-probability-extended (green) variants of Radtke 2017 (D), Leeuwen 2017 (E), and MRI-ERSPC (F) compared to the newly-fitted PI-RADS + UNet probability RC (purple)
Risk calculator calibration (original, UNet-Likert-substituted)
Original Leeuwen 2017, Radtke 2017, and MRI-ERSPC RCs had Brier scores of 0.14 (95% CI: 0.12–0.16), 0.22 (95% CI: 0.19–0.24), and 0.20 (95% CI: 0.17–0.22), respectively, with Leeuwen 2017 significantly better calibrated than MRI-ERSPC (p < 0.001) and Radtke 2017 (p = 0.001) (Fig. 3, top, lower is better). After intercept-only calibration, Brier scores of all RCs improved to 0.12 (95% CI: 0.10–0.14, p < 0.001), 0.13 (95% CI: 0.11–0.15, p < 0.001), and 0.13 (95% CI: 0.11–0.15, p < 0.001), respectively (Fig. 3, middle). Intercept/slope recalibration led to only minor statistically insignificant improvements over the intercept-only calibration, at 0.12 (95% CI: 0.11–0.14, p = 0.90), 0.13 (95% CI: 0.11–0.15, p = 1.00), and 0.13 (95% CI: 0.11–0.15, p = 0.61), respectively (Fig. 3, bottom). The Exp./Obs.-Ratio the ratios for the original RCs were 1.29, 1.69, and 0.48, respectively, indicating Radtke 2017 and Leeuwen 2017 overestimated sPC risk while MRI-ERSPC was underestimated. After intercept-only or intercept/slope calibration, Exp./Obs.-Ratio improved to 1.04, 0.99, and 1.02, respectively, with diminishing differences in over- or underestimation between the models. UNet-Likert-substituted RCs improved Brier scores for all RCs, however, only the improvement for MRI-ERSPC was statistically significant (p < 0.001) (filled triangles in Fig. 3, top). Supplemental Fig. 1 shows calibration plots before and after intercept-only calibration.
Brier scores, measuring both discrimination and calibration, for original versions w/o recalibration (top), after intercept-only recalibration (middle), and after intercept and slope refitting (bottom). Lower Brier scores indicate better fit. RCs were analyzed with PI-RADS (hollow circle) or UNet-Likert (filled triangle) as their MRI assessment method. As models show no significant difference in discrimination, Brier scores vary mostly due to calibration. Leeuwen 2017 shows the best out-of-the-box calibration, followed by MRI-ERSPC and Radtke 2017. Substitution with UNet-Likert worsens calibration for Leeuwen 2017 and Radtke 2017 slightly but results in an improvement for MRI-ERSPC. After intercept-only calibration, the Brier scores of all models improve drastically, as the worst re-calibrated model has lower scores than the best-uncalibrated model. The effect of UNet-Likert substitution on Brier scores is minimal after intercept-only recalibration. Fitting the model’s slope in addition to the intercept does not lead to improved Brier scores overall and does not decrease Brier score differences between original and substituted models
Performance of combined DL and PI-RADS (UNet-probability-extended RCs, newly-fitted PI-RADS + UNet probability RC)
UNet-probability-extended RCs for Leeuwen 2017, Radtke 2017, and MRI-ERSPC achieved AUC of 0.92 (95% CI: 0.89–0.94), 0.92 (95% CI: 0.90–0.95), and 0.92 (95% CI: 0.90–0.94) (Fig. 2C), respectively, and resulted in a significant improvement to the original Radtke 2017 (p < 0.001), Leeuwen 2017 (p < 0.01), and MRI-ERSPC (p < 0.01). The coefficients for UNet-probability-extended RCs are given in Supplemental Table 3.
The highest test-set cross-validation performance of the candidate models was provided by candidate model #3 which thus was selected as the newly-fitted PI-RADS + UNet probability RC (see Supplemental Table 2). Optimal parameters in this model included age, DRE, reciprocal square root of prostate volume, natural logarithm of PSA, biopsy status, PI-RADS score, and UNet-probability resulting in AUC of 0.93 (95% CI: 0.90–0.95) (Fig. 2D–F) and Brier score of 0.10 (95% CI: 0.09–0.12). While the AUC improvements of the newly-fitted PI-RADS + UNet probability RC compared to the UNet-probability-extended RCs were not statistically significant (p > 0.12), there was a minor improvement. The nomogram for the newly-fitted PI-RADS + UNet probability RC is given in Fig. 4, with model parameter significance and odds ratios given in Table 3 indicating the independent contribution of PI-RADS and UNet-probability.
Nomogram for calculating the probability of finding sPC from demographic data, clinical data, radiologist’s mpMRI assessment (PI-RADS), and the predictions of a fully-automated DL system (UNet-probability). Each parameter is projected orthogonally onto the points scale, then points for all parameters are summed. The predicted probability is obtained by orthogonal projection of the total points scale onto the sPC probability scale
Benefit of avoiding biopsies through risk stratification
Net benefit curves from DCA are given in Fig. 5. Leeuwen 2017 was on par with the PI-RADS/PSAD strategy for thresholds below 20% and showed improved net benefit above that threshold. Radtke 2017 showed benefits against the default strategies in the relevant range while MRI-ERSPC appeared harmful for thresholds below 20% (Fig. 5A). With prevalence adjustment by intercept-only recalibration, Radtke 2017 and MRI-ERSPC compensate for their miscalibration and consequently closely approximate Leeuwen 2017 such that there is no longer a clear benefit for a single RC. UNet-Likert-substituted RCs provide higher net benefits than their respective original RCs. UNet-probability-extended RCs improve over the original RCs with intercept/slope recalibration, UNet-Likert-substituted RC counterparts, and the PI-RADS/PSAD strategy for risk thresholds above 10%. The newly-fitted PI-RADS + UNet probability RC shows further minimal improvement over the UNet-probability-extended RCs, but not over the entire range of relevant thresholds.
Decision curve analysis comparing existing RCs (Leeuwen 2017 (red), Radtke 2017 (black), and MRI-ERSPC (green)), and the newly-fitted PI-RADS + UNet probability RC (purple). The fixed biopsy decision strategy, where the threshold for biopsy is set at PI-RADS 3 with PSAD of ≥ 0.1 ng/mL2, is shown in comparison (blue). A Shows the RCs performance as is (solid) or with intercept-only recalibration (dotted), B shows each strategy as is (solid) and its UNet-Likert-substituted variant (dotted). Leeuwen 2017 shows achieves a net benefit comparable to the PI-RADS/PSAD strategy and can even exceed it for risk thresholds above 20%, even if manual PI-RADS is substituted with DL image assessment. C Shows models after intercept/slope recalibration (solid) and their UNet-probability-extended variants (dotted line) against the newly-fitted PI-RADS + UNet probability RC (purple)
Comparison of the absolute number of examinations receiving a recommendation to avoid biopsy by different decision strategies demonstrates that PI-RADS ≥ 4 cut-off spares 37% [190/517] of biopsy sessions at the cost of missing sPC in 12 of 190 spared sessions, which corresponds to a negative predictive value (NPV) of 94%. By adding PSAD through the PI-RADS/PSAD decision strategy, biopsy avoidance is reduced to 25% [130/517] while only 6 sPC are missed in 130 spared biopsies (NPV 95%). The newly-fitted PI-RADS + UNet probability RC at the 15% threshold can spare 49% [252/517] of biopsies, 12% more than the PI-RADS ≥ 4 cut-offs (p < 0.001) and 24% more than the PI-RADS/PSAD strategy (p < 0.001), while missing 16 sPC out of 252 spared biopsies, maintaining the NPV of the PI-RADS ≥ 4 cut-off at 94% (p = 0.98) and staying comparable to 95% NPV of PI-RADS/PSAD (p = 0.24). These improvements indicate the contributory effect of the UNet-derived information to PI-RADS and clinical information for patient stratification. Table 4 compares the number of biopsies spared by each decision strategy, with the finer histopathological stratification given in Supplemental Table 4. An exemplary case for the clinical benefit of the newly-fitted PI-RADS + UNet probability RC at the 15% risk threshold is shown in Fig. 6.
Risk stratification example from the test cohort: a 66-year-old male presents with negative DRE, PSA elevation to 6.2 ng/mL, prostate volume of 82.5 mL, and PSAD of 0.075 ng/mL. The previous prostate biopsy did not find any prostate cancer but MRI uncovers a lesion in the posterolateral peripheral zone with apparent diffusion coefficient restriction, categorized as PI-RADS 4. nnUNet co-detects the lesion and assigns a UNet-probability of 31%, corresponding to UNet-Likert 4. While Leeuwen 2017 and Radtke 2017 RCs follow the PI-RADS recommendation for biopsy with 20% and 45% sPC risk, respectively, the newly-fitted PI-RADS + UNet probability RC sets the sPC probability at only 10.5%, below the 15% threshold discussed for biopsy recommendation. MRI-TRUS fusion biopsy did not find any prostate cancer in 30 systematic and nine targeted cores, confirming the newly fitted RCs recommendation to spare the prostate biopsy in this case
Discussion
We find that fully-automated DL biparametric MRI assessment by UNet-Likert scores can substitute for clinical PI-RADS assessment without performance deterioration. After recalibration by adjusting the models’ intercept, all RC models exhibited similar net benefits. Substitution of PI-RADS by UNet-Likert scores demonstrated tendencies for improvement but combining PI-RADS with UNet-probability demonstrated improved discrimination and net benefit, suggesting extraction of complementary information from imaging. Diagnostically important information appears to be present in discordant MRI readings of radiologists and DL. However, DL systems trained with radiologist PI-RADS assessment instead of histopathologically proven sPC as ground truth may not provide similar complementary information. Our results suggest that DL may be able to support diagnostic assessment in settings with limited experience in prostate MRI, as it provided on par performance with experienced radiologists in the current study.
We demonstrated that nearly half of biopsies may spared by the newly-fitted PI-RADS + UNet probability RC while providing an NPV of 94%, which lies above the expected NPV of 86% (95% CI: 0.79–0.91) for mpMRI at this prevalence [16]. Almost doubling the number of spared biopsies comes at the cost of missing 16 sPC cases when using the 15% threshold, compared to six missed sPCs for PI-RADS/PSAD or 12 missed sPCs for PI-RADS ≥ 4. While this increase in false negative cases did not result in a significantly lower NPV, clinicians should critically weigh the decreased morbidity of spared prostate biopsies against the possibility of missing a small number of sPC. For risk-averse patients, establishing a follow-up plan to delay the biopsy instead of avoiding it has the potential to mitigate the consequences of missing sPC and should be investigated further. Being able to quantify and visualize the risk before undergoing an invasive procedure is an additional tool for shared decision-making with the patients about the benefits and harms of the procedure.
The models showed varying degrees of miscalibration when applied to our consecutive test set, but improved substantially by adjusting only the model’s intercept, shown by Brier scores and calibration curves. Leeuwen 2017, the uncalibrated RC from van Leeuwen et al [31], showed the best calibration overall. Leeuwen 2017 and Radtke 2017 slightly overestimated sPC risk while MRI-ERSPC consistently underestimated it before recalibration, which was already observed in previous studies [18, 34, 35, 40, 41] and can reduce their net benefit in DCA. The net benefit of an overestimating model always remains higher than the biopsy-all strategy if the risk threshold remains lower than the cohort prevalence, meaning an overestimating model cannot be harmful at low thresholds. The net benefit of underestimating models approaches the biopsy-none strategy with increasing miscalibration, so they can be harmful to risk thresholds below the cohort prevalence. As current practice favors biopsy for most patients [10], we assume that reasonable risk thresholds lie below the cohort prevalence, so overestimating models (Leeuwen 2017 und Radtke 2017) have an advantage benefit over underestimating MRI-ERSPC ones (MRI-ERSPC), which are potentially harmful in DCA. Deniffel et al [34] raised concerns over the potential miscalibration of RCs and argued that MRI-ERSPC and Radtke 2017 were not clinically useful through DCA. Without calibration, Radtke 2017 showed better clinical utility compared to the default strategies for thresholds over 10% in this study, and Leeuwen 2017 was on par with PI-RADS/PSAD and can even surpass it. With recalibration, all RCs have a higher net benefit than PI-RADS/PSAD.
Our study showed that DL image analysis, PI-RADS, and clinical and demographic parameters have complementary risk prediction values for sPC before biopsy. Predictive performance is also expected to increase with the addition of more risk factors, e.g., free-to-total PSA ratio [42], family history [43], body-mass index [44], or genomic markers [45,46,47,48]. DL should be further investigated for clinical decisions after biopsy, as MRI assessments are also a significant predictor for biochemical recurrence and prostatectomy outcomes [48,49,50].
There are limitations to this study. The RCs shown here are applicable for biopsy-naïve or previously negatively biopsied patients, as these represent the typical screening population. In active surveillance, the use of DL to predict tumor progression risk remains to be investigated [46]. Transperineal MRI/TRUS fusion biopsy provided the reference standard, while prostatectomy specimens would allow for more detailed lesion localization and sPC diagnosis, but prostatectomy cohorts are biased toward sPC-positive cases. As RCs are used in patient stratification before any intervention, the cohort examined in this study much more closely models a typical screening cohort in which an RC would be used, while a prostatectomy cohort would exclude most of the patients who would benefit most from the RC models due to their low-risk status and potential decision to avoid biopsy. The biopsy scheme used in this study has been shown to detect 97% of sPC found at radical prostatectomy [2], providing high-quality ground truth. DL image analysis was performed on bi-parametric MRI, while radiologists interpreted mpMRI including DCE. As the quality of predictions did not decline after PI-RADS was substituted by DL, our data suggest that bi-parametric MRI provides sufficient information. Previous studies showed only minor contributions of DCE to MRI assessment as well [51]. PSA showed a significant contribution to predictions, while PI-RADS and DL image analysis contributed similarly, suggesting that radiologists base their PI-RADS assessment primarily on the mpMRI imaging appearance, although they have access to PSA and PSAD, as intended by PI-RADS [24]. Radiologist-delineated lesions informed the reference standard through targeted biopsy cores while suspicious regions from UNet predictions could not be probed in a retrospective analysis, potentially underestimating the diagnostic potential of DL. However, at our institution, prostate MRI was read by experienced radiologists familiar with subtle manifestations of sPC and frequent review of cases in interdisciplinary boards, and providing high sensitivity at the PI-RADS ≥ 3 threshold of up to 98% compared to extended systematic and targeted biopsies [19]. The ground truth provided by sensitive clinical imaging assessment and a high-quality biopsy scheme leads to an exceptional targeted mapping quality of the prostate while maintaining a clinical workflow. For some of the examinations in the training set, only PI-RADS v1 was available. In these cases, post-hoc reassessment by a board-certified radiologist was performed without knowledge of the biopsy result. By considering the original PI-RADS v1 score, the result was a consensus score, which assured that a representative PI-RADS v2 score was used for training, e.g., a score for which agreement of image characteristics and score were further confirmed. For DL training, this reassessment had only a minor effect as training was based on histopathology from a systematic and targeted prostate biopsy and not PI-RADS, affecting only the ground truth if a lesion was not biopsied by using PI-RADS v1 criteria, which would qualify for biopsy by PI-RADS v2 criteria. As a sensitive approach to prostate assessment was chosen, the probability of this effect with regard to sPC was further minimized. For examinations in the test set, reassessment was not necessary and did therefore not affect performance comparisons. Logistic regression models, as used in the analyzed RCs, enable explainable nomograms but other classifiers should be investigated further, such as support vector machines or random forest classifiers, which have been successfully applied to radiomics [39] and to assess biochemical recurrence [52]. This study retrospectively evaluated cases from a single high-volume tertiary-care center. The benefit of DL-based and updated RCs should be further validated in multi-centric studies.
In conclusion, fully-automated DL prostate MRI assessment not only confirms its similar performance to clinical PI-RADS assessment but also demonstrates complementarity to the latter, with the effect of increased predictive performance of logistic regression risk estimation models utilizing both parameters, thus suggesting an attractive approach for improvement of diagnostic quality for patient stratification before biopsy.
Abbreviations
- AUC:
-
Area under the receiver operating characteristics curve
- DL:
-
Deep learning
- DRE:
-
Digital rectal examination
- ISUP:
-
International Society of Urological Pathology
- mpMRI:
-
Multiparametric magnetic resonance imaging
- PI-RADS:
-
Prostate imaging reporting and data system
- PPV/NPV:
-
Positive predictive value/negative predictive value
- PSA(D):
-
Prostate-specific antigen (density)
- RC:
-
Risk calculator
- ROC:
-
Receiver operating characteristics
- sPC:
-
Clinically significant prostate cancer
- TRUS:
-
Transrectal ultrasound
References
Panebianco V, Valerio MC, Giuliani A et al (2018) Clinical utility of multiparametric magnetic resonance imaging as the first-line tool for men with high clinical suspicion of prostate cancer. Eur Urol Oncol 1:208–214. https://doi.org/10.1016/j.euo.2018.03.008
Radtke JP, Schwab C, Wolf MB et al (2016) Multiparametric magnetic resonance imaging (MRI) and MRI-transrectal ultrasound fusion biopsy for index tumor detection: correlation with radical prostatectomy specimen. Eur Urol 70:846–853. https://doi.org/10.1016/j.eururo.2015.12.052
Schimmöller L, Blondin D, Arsov C et al (2016) MRI-guided in-bore biopsy: differences between prostate cancer detection and localization in primary and secondary biopsy settings. AJR Am J Roentgenol 206:92–99. https://doi.org/10.2214/AJR.15.14579
Radtke JP, Kuru TH, Bonekamp D et al (2016) Further reduction of disqualification rates by additional MRI-targeted biopsy with transperineal saturation biopsy compared with standard 12-core systematic biopsies for the selection of prostate cancer patients for active surveillance. Prostate Cancer Prostatic Dis 19:283–291. https://doi.org/10.1038/pcan.2016.16
Ahmed HU, El-Shater Bosaily A, Brown LC et al (2017) Diagnostic accuracy of multi-parametric MRI and TRUS biopsy in prostate cancer (PROMIS): a paired validating confirmatory study. Lancet 389:815–822. https://doi.org/10.1016/S0140-6736(16)32401-1
van der Leest M, Cornel E, Israël B et al (2019) Head-to-head comparison of transrectal ultrasound-guided prostate biopsy versus multiparametric prostate resonance imaging with subsequent magnetic resonance-guided biopsy in biopsy-naive men with elevated prostate-specific antigen: a large prospective multicenter clinical study. Eur Urol 75:570–578. https://doi.org/10.1016/j.eururo.2018.11.023
Kasivisvanathan V, Rannikko AS, Borghi M et al (2018) MRI-targeted or standard biopsy for prostate-cancer diagnosis. N Engl J Med 378:1767–1777. https://doi.org/10.1056/NEJMoa1801993
Rouvière O, Puech P, Renard-Penna R et al (2019) Use of prostate systematic and targeted biopsy on the basis of multiparametric MRI in biopsy-naive patients (MRI-FIRST): a prospective, multicentre, paired diagnostic study. Lancet Oncol 20:100–109. https://doi.org/10.1016/S1470-2045(18)30569-2
Wegelin O, Exterkate L, van der Leest M et al (2019) The FUTURE trial: a multicenter randomised controlled trial on target biopsy techniques based on magnetic resonance imaging in the diagnosis of prostate cancer in patients with prior negative biopsies. Eur Urol 75:582–590. https://doi.org/10.1016/j.eururo.2018.11.040
Mottet N, van den Bergh RCN, Briers E et al (2021) EAU-EANM-ESTRO-ESUR-SIOG guidelines on prostate cancer—2020 update. Part 1: screening, diagnosis, and local treatment with curative intent. Eur Urol 79:243–262. https://doi.org/10.1016/j.eururo.2020.09.042
Steyerberg E, Roobol M, Kattan M, Van der Kwast T, De Koning H, Schröder F (2007) Prediction of indolent prostate cancer: validation and updating of a prognostic nomogram. J Urol 177:107–112. https://doi.org/10.1016/j.juro.2006.08.068
Kranse R, Roobol M, Schröder FH (2008) A graphical device to represent the outcomes of a logistic regression analysis. Prostate 68:1674–1680. https://doi.org/10.1002/pros.20840
Turkbey B, Rosenkrantz AB, Haider MA et al (2019) Prostate imaging reporting and data system version 2.1: 2019 update of prostate imaging reporting and data system version 2. Eur Urol 76:340–351. https://doi.org/10.1016/j.eururo.2019.02.033
Venderink W, van Luijtelaar A, van der Leest M et al (2019) Multiparametric magnetic resonance imaging and follow-up to avoid prostate biopsy in 4259 men. BJU Int 124:775–784. https://doi.org/10.1111/bju.14853
Westphalen AC, McCulloch CE, Anaokar JM et al (2020) Variability of the positive predictive value of PI-RADS for prostate MRI across 26 centers: experience of the society of abdominal radiology prostate cancer disease-focused panel. Radiology 296:76–84. https://doi.org/10.1148/radiol.2020190646
Drost FH, Osses D, Nieboer D et al (2020) Prostate magnetic resonance imaging, with or without magnetic resonance imaging-targeted biopsy, and systematic biopsy for detecting prostate cancer: a cochrane systematic review and meta-analysis. Eur Urol 77:78–94. https://doi.org/10.1016/j.eururo.2019.06.023
Loeb S, Vellekoop A, Ahmed HU et al (2013) Systematic review of complications of prostate biopsy. Eur Urol 64:876–892. https://doi.org/10.1016/j.eururo.2013.05.049
Radtke JP, Giganti F, Wiesenfarth M et al (2019) Prediction of significant prostate cancer in biopsy-naïve men: validation of a novel risk model combining MRI and clinical parameters and comparison to an ERSPC risk calculator and PI-RADS. PLoS One 14:e0221350. https://doi.org/10.1371/journal.pone.0221350
Netzer N, Weißer C, Schelb P et al (2021) Fully automatic deep learning in bi-institutional prostate magnetic resonance imaging: effects of cohort size and heterogeneity. Invest Radiol 56:799–808. https://doi.org/10.1097/rli.0000000000000791
Schelb P, Kohl S, Radtke JP et al (2019) Classification of cancer at prostate mri: deep learning versus clinical PI-RADS assessment. Radiology 293:607–617. https://doi.org/10.1148/radiol.2019190938
Isensee F, Jaeger PF, Kohl SAA, Petersen J, Maier-Hein KH (2021) nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods 18:203–211. https://doi.org/10.1038/s41592-020-01008-z
Baumgartner M, Jäger PF, Isensee F, Maier-Hein KH (2021) nnDetection: a self-configuring method for medical object detection. In: de Bruijne M, Cattin PC, Cotin S et al (eds) Medical image computing and computer assisted intervention—MICCAI 2021. Springer International Publishing, Cham, pp 530–539. https://doi.org/10.1007/978-3-030-87240-3_51
Ronneberger O, Fischer P, Brox T (2015) U-Net: convolutional networks for biomedical image segmentation. In: Navab N, Hornegger J, Wells WM, Frangi AF (eds) Medical image computing and computer-assisted intervention—MICCAI 2015. Springer International Publishing, Cham, pp 234–241. https://doi.org/10.1007/978-3-319-24574-4_28
Weinreb JC, Barentsz JO, Choyke PL et al (2016) PI-RADS prostate imaging—reporting and data system: 2015, version 2. Eur Urol 69:16–40. https://doi.org/10.1016/j.eururo.2015.08.052
JO Barentsz J Richenberg R Clements et al (2012) ESUR prostate MR guidelines 2012 Eur Radiol 22 746–757. https://doi.org/10.1007/s00330-011-2377-y.
Dickinson L, Ahmed HU, Allen C et al (2011) Magnetic resonance imaging for the detection, localisation, and characterisation of prostate cancer: recommendations from a European Consensus Meeting. Eur Urol 59:477–494. https://doi.org/10.1016/j.eururo.2010.12.009
Kuru TH, Wadhwa K, Chang RT et al (2013) Definitions of terms, processes and a minimum dataset for transperineal prostate biopsies: a standardization approach of the Ginsburg Study Group for Enhanced Prostate Diagnostics. BJU Int 112:568–577. https://doi.org/10.1111/bju.12132
Egevad L, Delahunt B, Srigley JR, Samaratunga H (2016) International Society of Urological Pathology (ISUP) grading of prostate cancer—an ISUP consensus on contemporary grading. APMIS 124:433–435. https://doi.org/10.1111/apm.12533
Schelb P, Wang X, Radtke JP et al (2021) Simulated clinical deployment of fully automatic deep learning for clinical prostate MRI assessment. Eur Radiol 31:302–313. https://doi.org/10.1007/s00330-020-07086-z
Radtke JP, Wiesenfarth M, Kesch C et al (2017) Combined clinical parameters and multiparametric magnetic resonance imaging for advanced risk modeling of prostate cancer—patient-tailored risk stratification can reduce unnecessary biopsies. Eur Urol 72:888–896. https://doi.org/10.1016/j.eururo.2017.03.039
van Leeuwen PJ, Hayen A, Thompson JE et al (2017) A multiparametric magnetic resonance imaging-based risk model to determine the risk of significant prostate cancer prior to biopsy. BJU Int 120:774–781. https://doi.org/10.1111/bju.13814
Alberts AR, Roobol MJ, Verbeek JFM et al (2019) Prediction of high-grade prostate cancer following multiparametric magnetic resonance imaging: improving the Rotterdam European Randomized Study of Screening for Prostate cancer Risk Calculators. Eur Urol 75:310–318. https://doi.org/10.1016/j.eururo.2018.07.031
Görtz M, Radtke JP, Hatiboglu G et al (2021) The value of prostate-specific antigen density for prostate imaging-reporting and data system 3 lesions on multiparametric magnetic resonance imaging: a strategy to avoid unnecessary prostate biopsies. Eur Urol Focus 7:325–331. https://doi.org/10.1016/j.euf.2019.11.012
Deniffel D, Healy GM, Dong X et al (2021) Avoiding unnecessary biopsy: MRI-based risk models versus a PI-RADS and PSA density strategy for clinically significant prostate cancer. Radiology 300:369–379. https://doi.org/10.1148/radiol.2021204112
Püllen L, Radtke JP, Wiesenfarth M et al (2020) External validation of novel magnetic resonance imaging-based models for prostate cancer prediction. BJU Int 125:407–416. https://doi.org/10.1111/bju.14958
Remmers S, Kasivisvanathan V, Verbeek JFM, Moore CM, Roobol MJ (2022) Reducing biopsies and magnetic resonance imaging scans during the diagnostic pathway of prostate cancer: applying the rotterdam prostate cancer risk calculator to the PRECISION trial data. Eur Urol Open Science 36:1–8. https://doi.org/10.1016/j.euros.2021.11.002
Van Calster B, Wynants L, Verbeek JFM et al (2018) Reporting and interpreting decision curve analysis: a guide for investigators. Eur Urol 74:796–804. https://doi.org/10.1016/j.eururo.2018.08.038
Vickers AJ, Elkin EB (2006) Decision curve analysis: a novel method for evaluating prediction models. Med Decis Making 26:565–574. https://doi.org/10.1177/0272989x06295361
Zhang KS, Schelb P, Kohl S et al (2021) Improvement of PI-RADS-dependent prostate cancer classification by quantitative image assessment using radiomics or mean ADC. Magn Reson Imaging 82:9–17. https://doi.org/10.1016/j.mri.2021.06.013
Petersmann A-L, Remmers S, Klein T et al (2021) External validation of two MRI-based risk calculators in prostate cancer diagnosis. World J Urol 39:4109–4116. https://doi.org/10.1007/s00345-021-03770-x
Doan P, Graham P, Lahoud J et al (2021) A comparison of prostate cancer prediction models in men undergoing both magnetic resonance imaging and transperineal biopsy: Are the models still relevant? BJU Int 128:36–44. https://doi.org/10.1111/bju.15554
Nan L, Guo K, Li M, Wu Q, Huo S (2022) Development and validation of a multi-parameter nomogram for predicting prostate cancer: a retrospective analysis from Handan Central Hospital in China. PeerJ 10:e12912. https://doi.org/10.7717/peerj.12912
Patel HD, Koehne EL, Shea SM et al (2022) Risk of prostate cancer for men with prior negative biopsies undergoing magnetic resonance imaging compared with biopsy-naive men: a prospective evaluation of the PLUM cohort. Cancer 128:75–84. https://doi.org/10.1002/cncr.33875
Nasri J, Barthe F, Parekh S et al (2022) Nomogram predicting adverse pathology outcome on radical prostatectomy in low-risk prostate cancer men. Urology 166:189–195. https://doi.org/10.1016/j.urology.2022.02.019
Hu D, Cao Q, Tong M et al (2022) A novel defined risk signature based on pyroptosis-related genes can predict the prognosis of prostate cancer. BMC Med Genomics 15:24. https://doi.org/10.1186/s12920-022-01172-5
Beksac AT, Ratnani P, Dovey Z et al (2021) Unified model involving genomics, magnetic resonance imaging and prostate-specific antigen density outperforms individual co-variables at predicting biopsy upgrading in patients on active surveillance for low risk prostate cancer. Cancer Rep 5:e1492. https://doi.org/10.1002/cnr2.1492
Wu C, Zhu J, King A et al (2021) Novel strategy for disease risk prediction incorporating predicted gene expression and DNA methylation data: a multi-phased study of prostate cancer. Cancer Commun (Lond) 41:1387–1397. https://doi.org/10.1002/cac2.12205
Huang W, Randhawa R, Jain P et al (2022) A novel artificial intelligence-powered method for prediction of early recurrence of prostate cancer after prostatectomy and cancer drivers. JCO Clin Cancer Inform 6:e2100131. https://doi.org/10.1200/CCI.21.00131
Mazzone E, Gandaglia G, Ploussard G et al (2022) Risk stratification of patients candidate to radical prostatectomy based on clinical and multiparametric magnetic resonance imaging parameters: development and external validation of novel risk groups. Eur Urol 81:193–203. https://doi.org/10.1016/j.eururo.2021.07.027
van Dijk-de Haan MC, Boellaard TN, Tissier R et al (2022) Value of different magnetic resonance imaging-based measurements of anatomical structures on preoperative prostate imaging in predicting urinary continence after radical prostatectomy in men with prostate cancer: a systematic review and meta-analysis. Eur Urol Focus 8:1211–1225. https://doi.org/10.1016/j.euf.2022.01.015
Tavakoli AA, Hielscher T, Badura P et al (2022) Contribution of dynamic contrast-enhanced and Diffusion MRI to PI-RADS for detecting clinically significant prostate cancer. Radiology 306:186–199. https://doi.org/10.1148/radiol.212692
Tan YG, Fang AHS, Lim JKS et al (2022) Incorporating artificial intelligence in urology: Supervised machine learning algorithms demonstrate comparative advantage over nomograms in predicting biochemical recurrence after prostatectomy. Prostate 82:298–305. https://doi.org/10.1002/pros.24272
Bossuyt PM, Reitsma JB, Bruns DE et al (2015) STARD 2015: an updated list of essential items for reporting diagnostic accuracy studies. Radiology 277:826–832. https://doi.org/10.1148/radiol.2015151516
Funding
This research received research support from Bundesministerium für Wirtschaft und Klimaschutz (BMWK): grant no. 01MT21004B. Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is David Bonekamp.
Conflict of interest
Albrecht Stenzinger reports speakers honoraria for Aignostics, Amgen, Astra Zeneca, AGCT, Bayer, Bristol-Myers Squibb, Eli Lilly, Illumina, Incyte, Janssen, MSD, Novartis, Qlucore, Pfizer, Roche, Seagen, Seattle Genetics, Takeda, Thermo Fisher and grants from Bayer, Bristol-Myers Squibb, Chugai, and Incyte. Heinz-Peter Schlemmer declares consulting fees or honoraria from Bayer, Bracco; travel support from Siemens, Bayer, Bracco; consultancy for Bayer; grants/grants pending from EU, BMBF, Deutsche Krebshilfe; payment for lectures from Bayer, Bracco. Heinz-Peter Schlemmer is member of the Advisory Editorial Board of European Radiology. He has not taken part in the review or selection process of this article. David Bonekamp received lecture payments from Bayer Vital. Heinz-Peter Schlemmer is a member of the European Radiology Editorial Board. He has not taken part in the review or selection process of this article. The remaining authors declare no relationships with any companies, whose products or services may be related to the subject matter of the article.
Statistics and biometry
Thomas Hielscher, a co-author of this study, is a biostatistician at the German Cancer Research Center (DKFZ), Heidelberg, Germany, and contributed to the statistical analysis for this paper.
Informed consent
Written informed consent was waived by the Institutional Review Board (S-164/2019) in Heidelberg.
Ethical approval
Institutional Review Board approval was obtained with ethics review in Heidelberg (S-164/2019).
Study subjects or cohorts overlap
One thousand six hundred ten out of the 1627 MRI examinations included in this study have been previously reported. The previous publications focused on DL and radiomics [19, 20, 39], however, data have not been used for systematic clinical RC assessment or development. Regarding the PI-RADS/PSAD biopsy strategy, the test set of a previous study overlaps with 101 exams from biopsy-naïve patients with PI-RADS 3 lesions in the current study [33].
Methodology
-
Retrospective
-
Diagnostic and prognostic study
-
Performed at one institution
Additional information
Publisher’s Note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schrader, A., Netzer, N., Hielscher, T. et al. Prostate cancer risk assessment and avoidance of prostate biopsies using fully automatic deep learning in prostate MRI: comparison to PI-RADS and integration with clinical data in nomograms. Eur Radiol (2024). https://doi.org/10.1007/s00330-024-10818-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00330-024-10818-0