Abstract
Purpose
We aimed to develop and externally validate a nomogram based on MRI volumetric parameters and clinical information for deciding when SBx should be performed in addition to TBx in man with suspicious prostate MRI.
Materials and methods
Retrospective analyses of single (IMPROD, NCT01864135) and multi-institution (MULTI-IMPROD, NCT02241122) clinical trials. All men underwent a unique rapid biparametric magnetic resonance imaging (IMPROD bpMRI) consisting of T2-weighted imaging and three separate DWI acquisitions. Men with IMPROD bpMRI Likert scores of 3–5 were included. Logistic regression models were developed using IMPROD trial (n = 122) and validated using MULTI-IMPROD trial (n = 262) data. The model’s performance was evaluated in the terms of PCa detection with Gleason Grade Group 1 (clinically insignificant prostate cancer, iPCa) and > 1 (clinically significant prostate cancer, csPCa). Net benefits and decision curve analyses (DCA) were compared. Combined biopsies were used for reference.
Results
The developed nomogram included age, PSA, prostate volume, MRI suspicion score (IMPROD bpMRI Likert or PIRADsv2.1 score), MRI-suspicion lesion volume percentage, and lesion location. All these variables were significant predictors of csPCa in SBx in multivariable analysis. In the validation cohort (n = 262) using different nomogram cutoffs, 19–43% of men would have avoided SBx while missing 1–4% of csPCa and avoiding detection of 9–20% of iPCa. Similar performance was found for nomograms using IMPROD bpMRI Likert score or v2.1.
Conclusions
The developed nomogram demonstrated potential to select men with a clinical suspicion of PCa who would benefit from performing SBx in addition to TBx. Public access to the nomogram is provided at:https://petiv.utu.fi/multiimprod/.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
In the recent years, Magnetic Resonance Imaging (MRI) is increasingly used in the diagnostic pathways for prostate cancer (PCa). Simplified MRI protocols with shorter acquisition time and with no use of contrast agents (biparametric MRI) hold the promise to further reduce costs of prostate imaging and promote its use in any patient at risk of PCa [1, 2]. Furthermore, compared to other tools such as biomarkers, prostate MRI allows to characterize and localize PCa lesions and to eventually perform targeted biopsy of any suspicious lesions. Given the reported superiority of the MRI-targeted biopsy (TBx) compared to systematic biopsy (SBx) in recent prospective clinical trials (PROMIS [3], PRECISION [4], and MRI-FIRST [5]), there is a growing interest to evaluate if SBx is needed in addition to TBx in men with a clinical suspicion of PCa. Avoiding SBx would reduce the risks of infection, bleeding and pain associated with additional cores sampling [6, 7]. Additionally, the incidental detection of clinically insignificant PCa (iPCa) is more frequent in SBx than in TBx [8]. Finally, the workload for pathologist evaluating biopsy cores would be reduced from reviewing 12–18 cores to 2–6 cores. However, the risk of missing clinically significant PCa (csPCa) must be addressed. A recent Cochrane meta-analysis of the results of studies evaluating MRI and MRI-targeted biopsy suggests that omitting systematic biopsy would miss approximately 16% of csPCa in biopsy-naive patients and 10% in the repeat-biopsy setting.
Several multivariable models to predict negative SBx have been developed to avoid systematic cores at the time of target biopsy [10, 11]. None of these models showed excellent accuracy, and their clinical benefit is questionable. There are at least three reasons behind TBx failure: (1) Misdiagnosis of the lesion by the radiologist either due to the low quality of prostate MRI or misinterpretation; (2) Presence of csPCa that is not visible on MRI; (3) Targeting error.
Since smaller lesions in big prostates are more likely to be missed by both the reader or by the person performing the TBx, we aimed to develop and externally validate a nomogram based on MRI volumetric parameters and clinical information for deciding when SBx should be performed in addition to TBx in man with suspicious prostate MRI.
Materials and methods
Study design and population
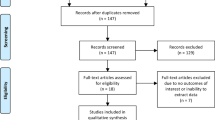
We retrospectively analyzed data from two prospective clinical trials involving a total of 499 patients (Supporting Material Figure S1). All patients underwent biparametric MRI according to IMPROD protocol (IMPROD bpMRI). Patients with negative MRI (IMPROD bpMRI Likert score/PIRADsv2.1 score of 1–2) were excluded from the analysis (n = 112). Additionally, four men were excluded: three having a PIRADsv2.1 score > 2 while IMPROD bpMRI was Likert 2 and no TBx was performed; one having PIRADsv2.1 score of 1 while IMPROD bpMRI was Likert 3 (Supporting Material Figure S1). Imaging findings for these four men are presented in the Supporting Material (Figure S2–S4) and the following 15 tweet series on twitter https://twitter.com/jambor_ivan/status/1185272940671180800. The final population consisted of two cohort: a single institution development cohort, IMPROD trial (NCT01864135) n = 122 and a multi-institution validation cohort, MULTI-IMPROD trial (NCT02241122) n = 262. All the study biopsies were taken during 4/2013—05/2017. The patient study flow is shown in Figure S1.
IMPROD bpMRI protocol and MRI reporting
As described before, the MRI examination was performed using surface array body coils at 3 T in Turku (Verio, Siemens), Helsinki (Skyra, Siemens), and Tampere (Skyra, Siemens), while 1.5 T (Aera, Siemens) MRI scanner was used in Pori [12]. All imaging data sets were prospectively reported by a local radiologist (at least one year of prostate MRI experience at the beginning of the MULTI-IMPROD trial) and confirmed or re-reported centrally by one designated central reader (IJ, 5 years of prostate MRI experience at the beginning of the MULTI-IMPROD trial) to guarantee reporting integrity using a dedicated IMPROD bpMRI reporting system developed before initialization of the trial (See details at https://mrc.utu.fi/mri/improd). Following the completion of the trial, Prostate Imaging Reporting and Data System version 2.1 (PI-RADSv2.1) scores [12] were assigned by the central reader (IJ). Volumetric analysis was performed using manually delineating whole prostate volumes and all MRI suspicious lesions on axial T2-weigthed images.
Volumes were calculated as the sum of prostate/lesion area on each slice multiplied by slice thicknesses (3 mm). Cancer volume on MRI was calculated as the sum of all the lesion volumes in each patient. Total percentage of cancer on MRI was calculated as the ratio of cancer volume on MRI divided the whole prostate volume. The MRI protocols, all MRI data and MRI reports are freely available at the following address: https://petiv.utu.fi/improd/ and https://petiv.utu.fi/multiimprod/.
Biopsy procedure and histopathological analysis
In IMPROD trial, only the dominant lesion was targeted (two targeted biopsy cores), whereas in MULTI-IMPROD trial, up to two lesions were targeted (two targeted biopsy cores per each lesion). A cognitive targeting method was used with the exception of one center (Helsinki) where a MRI-TRUS fusion platform was used (n = 45). Every patient in the present study underwent the first TBx of every MRI suspicious lesion. The same surgeon subsequently performed 12 cores SBx using a sextant template. The location of each SBx core was random and thus may overlap in some degree with MRI lesions. Histopathology of all biopsies was reported separately at each center by a uropathologist, each with at least 5 years of experience in genitourinary pathology at the beginning of the trial, using the 2014 International Society of Urological Pathology modified Gleason grading system [13]. The Gleason score for each patient was assigned using the overall Gleason score from all SBx and TBx cores. Findings from MRI and clinical data were not available for the pathologists.
Outcome measurements and statistical analysis
Outcomes of this study were the rates of csPCa, Gleason Grade Group (GGG) > 1 and iPCa, GGG = 1, in SBx, TBx and overall (SBx + TBx).
Descriptive characteristics of the development and validation cohorts were calculated. Continuous variables are reported as medians ± standard deviation (when appropriate) and compared by the Mann Whitney test for independent groups. Differences in rates were tested by the chi-square test.
To graphically assess how prostate and lesion dimensions impact the diagnostic accuracy of TBx and SBx, we plotted the actual probability of diagnosing csPCa and iPCa in SBx and TBx according to the percentage of prostate involved by tumor.
Univariable and Multivariable analyses were performed in the development cohort to evaluate predictors of csPCa in SBx, TBx and overall (SBx + TBx).
Two different logistic regression models were developed for each of the three outcomes (csPCa in SBx, TBx, SBx + TBx) including either PI-RADSv2.1 or IMPROD bpMRI Likert score in addition to the other variables. The models with the best area under the curve (AUC) in the development cohort were selected and the linear prediction of the logistic function was used to perform the decision curve analysis in the validation cohort.
Finally, to provide physicians and patients with an outline for deciding when to perform SBx in addition to TBx, we performed the systematic analysis of model-derived cutoffs, considering the predicted probability of the models in the validation cohort. Statistical analyses were performed using Stata 14 (StataCorp LP, College Station, TX, USA). All tests were 2-sided with a significance level set at p < 0.05.
Results
Patients’ characteristic for the included 122 and 262 men in the developmental (IMPROD trial) and validation (MULTI-IMPROD) cohorts, respectively, are presented in Table 1. PSA density (0.20 vs 0.16, p 0.004), index lesion volume (0.98 vs 0.73, p 0.015) and csPCa detection rate in TBx (60% vs 47%, p 0.01) were slightly higher in the developmental cohort but no difference were found in overall csPCa detection rate ( csPCa detection rate in TBx + SBx, p 0.051). CsPCa detection rate according to Likert 1–2, 3 and 4–5 lesion were respectively 4.5%, 47.6% and 79.7% in the development cohort, 10.8%, 30.5% and 70.3% in the validation cohort. No difference was found in csPCa detection rate according to PIRADsv2.1 score.
Univariate analysis in the developmental cohort
The results of univariate analysis evaluating predictors of csPCa in SBx, TBx and all cores (SBx + TBx) are shown in Table 2. In contrast to TBx, MRI suspicious lesion location and prostate volume were significant predictors of csPCa in SBx with p values of 0.039 and 0.02, respectively. The remaining significant predictors of csPCa in SBx were similar to TBx. The total volume of the suspicious lesion on MRI, index lesion volume and the percentage of prostate involved by cancer emerged as important predictors csPCa in SBx, TBx and all cores (SBx + TBx).
Figure 1 graphically shows that the actual probability of diagnosing csPCa in SBx and TBx increase with the increase of the percentage of prostate involved by a suspicious lesion on MRI. However, in patients with low percentages, the probability of finding csPCa in SBx only is very low and actually lower than the probability of finding iPCa.
Multivariable analysis in the developmental cohort
The results of multivariable analysis evaluating predictors of csPCa in SBx using IMPROD bpMRI Likert score and PI-RADsv2.1 are shown in Table 3 and Supporting Material Table S1, respectively. The corresponding results for predictors of csPCa in TBx are shown in Table 4 and Supporting Material Table S2, respectively. Finally, the results of multivariable analysis evaluating predictors of csPCa in SBx + TBx using IMPROD bpMRI Likert score and PI-RADsv2.1 are shown in Supporting Material Table S3 and S4, respectively. In contrast to predictors of csPCa in TBx and SBx + TBx, PSA and MRI-suspicion lesion location were a significant predictor of csPCa only in SBx (Table 3 and Supporting Material Table S1). Decision curve analysis for SBx, TBx, and SBx + TBx are shown in Supporting Material Figure S6, S7, and S8, respectively.
Risk calculator
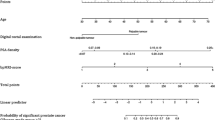
A risk calculator for deciding when SBx should be performed in addition to TBx in men with suspicious MRI findings was developed using the developmental cohort (IMPROD, n = 122) while the results using the validation cohort (MULTI-IMPROD, n = 262) are shown in Table 5. The developed nomogram included age, PSA, prostate volume, MRI suspicion score (IMPROD bpMRI Likert or PIRADsv2.1 score), MRI-suspicion lesion volume percentage, and lesion location. Beta coefficients of the logit function are shown in Supporting Material Table S5-S6.
The external validity of the risk calculator was performed in the validation cohort (n = 262) using DCA and evaluating the clinical utility of the risk calculator at different cut-offs (Table 5). DCA showed a net clinical benefit for every value of predicted probability above 10%.
The systematic analysis of the model derived cutoffs revealed that 19–43% of men would have avoided SBx while missing 1–4% of csPCa and avoiding detection of 9–20% of iPCa (Table 5). Similar performance was found for nomograms using IMPROD bpMRI Likert score or PIRADsv2.1.
Discussion
In this study, using data of two prospective trials, we developed a novel instrument based onto readily-available clinical parameters (age, PSA, prostate volume, IMPROD bpMRI Likert or PIRADsv2.1 score, total percentage of cancer volume on MRI and lesion location) that can help the physician to understand the added value of SBx in patients who underwent TBx of MRI suspicious lesion.
Avoiding SBx in patients who have a suspicious MRI lesion represents a great challenge for a urologist.
In an ideal world, where MRI and targeting systems show perfect performance, avoiding SBx would result in less complications and a lower rate of diagnosis of iPCa.
Recently Wegelin et al., in a post hoc analysis of a randomized controlled trial, compared adverse events among three mpMRI-based TBx techniques of the prostate in men with prior negative SBx and a persisting suspicion of PCa [6, 14]. No difference were found in PCa detection rates between in-bore MRI TBx, MRI-TRUS fusion TBx, and cognitive registration TRUS TBx [14], however, the number of biopsy cores taken was different between the various techniques and was significantly associated with the occurrence of adverse events (OR 1.11 [95% CI 1.06–1.17, p < 0.001]) [6].
To the best of our knowledge no level 1 evidence is available in the biopsy naïve setting. Indeed, when mpMRI is positive (i.e. PIRADsv2.1 > 3), EAU guidelines suggest performing 12-cores SBx in addition to TBx of every suspicious lesion in biopsy naïve patients (Level of evidence: 2A; strength rating: strong) and TBx only in patients with prior negative biopsy (Level of evidence: 2A; strength rating: weak) [15].
Although this might not be feasible in every center at the moment, the aim of this study was to find a subset of biopsy naïve men with a clinical suspicion of PCa that can safely avoid 12-core SBx without compromising the detection of csPCa.
Unlike prior studies, we considered MRI volumetric parameters and we found that adding the prostate volume and percentage of prostate involved by tumor to a model based on age, PSA, lesion location and lesion suspicious score, increased the model predictive accuracy for csPCa on SBx from 0.82 to 0.86 for the model using IMPROD bpMRI Likert and from 0.80 to 0.85 for the model using PIRADsv2.1. Our rationale for this decision was that smaller lesions in big prostates are more likely to be missed by a reader interpreted prostate MRI or by the person performing TBx.
Turning findings into clinical practice, in a 56 years old man with PSA 4 ng/ml and one PIRADsv2.1 4 lesion of 1 cm (0.52 mL), the estimated probability of finding csPCa in SBx is 10.4% in case of prostate volume 60 mL (0.87% of prostate involved by tumor), as opposed to 5.7% in case of prostate volume 80 mL (0.65% of prostate involved by tumor).
Among the models for prediction of sPC in SBx, the nomogram developed by Sathianathen et al. considered the clinical setting (biopsy naive, previous negative biopsy and Active surveillance patients), number of MRI lesions and the highest PIRADS score [11]. Such model showed clinical benefit in a range of probability between 10 and 30%. The lack of external validation limits the generalization of these findings. Moreover, the model was developed to predict csPCa in systematic cores only (with negative TBx). Indeed, our outcome was intentionally csPCa in SBx irrespective of the results of TBx. We believe that information on every positive core (from TBx or SBx) are still necessary when planning focal therapies and to assess the oncologic outcome of the patient since the current risk stratification tools take into account the number of positive cores (e.g. NCCN risk categories that stratifies patients into favorable or unfavorable intermediate-risk based on the number of cores with GGG 2 [16]).
Similar to our risk calculator, the model developed by Stabile et al. predicts csPCa in SBx. Their model ultimately included age, PSA, Prostate volume, number of MRI lesions, PIRADS score and the biopsy setting (biopsy naive vs previous negative biopsy patients) [10]. However, the DCA showed modest clinical benefit in the external validation cohort and the authors concluded that the number of systematic biopsies spared compared with the number of aggressive PCa missed is negligible. Moreover, no useful clinical model can be developed to safely identify those patients who could avoid SBx in addition to TBx. Conversely, our risk calculator showed a clinical benefit in the external validation cohort for the low threshold of probabilities. The addition of volumetric parameters derived from MRI seems to be the key to develop nomograms that help in this scenario.
Biomarkers such us a 4Kscore, selectMDx or the Stockholm3 test could also help to predict the risk of csPCa in SBx in patients with a positive MRI. However, the evidence in this regard is very limited and while MRI is becoming widely available, molecular tests are not yet used in routine clinical practice [17,18,19,20].
The present study is not devoid of limitations. First, we acknowledge that we used a cohort of patients enrolled in a prospective clinical trial that received MRI and biopsies according to the highest standard of care with a central prospective review of MRI images for prostate cancer detection. Although, this is a reflective of the daily clinical practice in referral academic centers it may be limit the validity of our findings to nonacademic centers where radiologists and urologists are still in the initial phase of their learning curve. Moreover, we used the combination of SBx + TBx rather than the Radical prostatectomy specimen or template transperineal saturation biopsy as a reference standard. Even if the previous studies showed a certain rate of discordance between Bx GGG and final pathology GGG [21,22,23], this still represents the standard of care and none of the two other options would have been feasible.
Conclusions
In conclusion, patients with a small lesion on MRI in big prostate benefit less from standard cores at the time of MRI-targeted biopsy. We developed and externally validated in a multi-institutional cohort a risk calculator to predict the added values of SBx to TBx that could help urologist avoid unnecessary biopsy sampling, reduce detection of iPCa while maintaining detection of csPCa.
Availability of data and material
Free public access to IMPROD bpMRI protocol, all IMPROD and MULTI-IMPROD data sets, MRI reports, clinical data with pathology and follow up data is provided on the trials’ servers: https://petiv.utu.fi/improd/ and https://petiv.utu.fi/multiimprod/. Free public access to the developed model is provided at https://petiv.utu.fi/multiimprod/.
Abbreviations
- csPCa:
-
Clinically significant PCa
- iPCa:
-
Clinically insignificant PCa iPCa
- SBx:
-
Magnetic Resonance Imaging-targeted biopsy
- TBx:
-
Systematic biopsy
- MRI:
-
Magnetic Resonance Imaging
References
Perez IM, Jambor I, Kauko T et al (2020) Qualitative and quantitative reporting of a unique biparametric MRI: towards biparametric MRI-based nomograms for prediction of prostate biopsy outcome in men with a clinical suspicion of prostate cancer (IMPROD and MULTI-IMPROD Trials). J Magn Reson Imaging 51:1556–1567
Jambor I, Bostrom PJ, Taimen P et al (2017) Novel biparametric MRI and targeted biopsy improves risk stratification in men with a clinical suspicion of prostate cancer (IMPROD Trial). J Magn Reson Imaging 46:1089–1095
Ahmed HU, El-Shater Bosaily A, Brown LC et al (2017) Diagnostic accuracy of multi-parametric MRI and TRUS biopsy in prostate cancer (PROMIS): a paired validating confirmatory study. Lancet 25(389):815–822
Kasivisvanathan V, Rannikko AS, Borghi M et al (2018) MRI-targeted or standard biopsy for prostate-cancer diagnosis. N Engl J Med 10(378):1767–1777
Rouvière O, Puech P, Renard-Penna R et al (2019) Use of prostate systematic and targeted biopsy on the basis of multiparametric MRI in biopsy-naive patients (MRI-FIRST): a prospective, multicentre, paired diagnostic study. Lancet Oncol 20:100–109
Wegelin O, Exterkate L, van der Leest M et al (2019) Complications and adverse events of three magnetic resonance imaging-based target biopsy techniques in the diagnosis of prostate cancer among men with prior negative biopsies: results from the FUTURE trial, a multicentre randomised controlled trial. Eur Urol Oncol 2(6):617–624
Eineluoto JT, Jarvinen P, Kilpelainen T et al (2018) Patient experience of systematic versus fusion prostate biopsies. Eur Urol Oncol 1:202–207
Wajswol E, Winoker JS, Anastos H et al (2019) A cohort of transperineal electromagnetically tracked magnetic resonance imaging/ultrasonography fusion-guided biopsy: assessing the impact of inter-reader variability on cancer detection. BJU Int 125:531–540
Drost FJH, Osses DF, Nieboer D et al (2019) Prostate MRI, with or without MRI-targeted biopsy, and systematic biopsy for detecting prostate cancer. Cochrane Database Syst Rev 4(4):CD012663
Dell'Oglio P, Stabile A, Soligo M et al (2019) There is no way to avoid systematic prostate biopsies in addition to multiparametric magnetic resonance imaging targeted biopsies. Eur Urol Oncol 3(1):112–118
Sathianathen NJ, Warlick CA, Weight CJ et al (2019) A clinical prediction tool to determine the need for concurrent systematic sampling at the time of magnetic resonance imaging-guided biopsy. BJU Int 123:612–617
Jambor I, Verho J, Ettala O et al (2019) Validation of IMPROD biparametric MRI in men with clinically suspected prostate cancer: a prospective multi-institutional trial. PLoS Med 16:e1002813
Epstein JI, Egevad L, Amin MB, et al (2016) The 2014 International Society of Urological Pathology (ISUP) Consensus Conference on Gleason Grading of Prostatic Carcinoma: Definition of Grading Patterns and Proposal for a New Grading System. Am J Surg Pathol 40:244–52
Wegelin O, Exterkate L, van der Leest M et al (2019) The FUTURE Trial: a multicenter randomised controlled trial on target biopsy techniques based on magnetic resonance imaging in the diagnosis of prostate cancer in patients with prior negative biopsies. Eur Urol 75:582–590
Mottet N, van den Bergh RCN, Briers E, et al. EAU - ESTRO - ESUR - SIOG Guidelines on Prostate Cancer 2018. European Association of Urology Guidelines 2018 Edition, Vol. presented at the EAU Annual Congress Copenhagen 2018. Arnhem, The Netherlands European Association of Urology Guidelines Office, 2018
Carroll PH, Mohler JL (2018) NCCN Guidelines Updates: Prostate Cancer and Prostate Cancer Early Detection. J Natl Compr Canc Netw 16:620–623
Falagario UG, Martini A, Wajswol E et al (2019) Avoiding Unnecessary Magnetic Resonance Imaging (MRI) and biopsies: negative and positive predictive value of MRI according to prostate-specific antigen density, 4Kscore and Risk Calculators. Eur Urol Oncol
Van Neste L, Hendriks RJ, Dijkstra S et al (2016) Detection of high-grade prostate cancer using a urinary molecular biomarker-based risk score. Eur Urol 70:740–748
Gronberg H, Eklund M, Picker W et al (2018) Prostate cancer diagnostics using a combination of the Stockholm3 blood test and multiparametric magnetic resonance imaging. Eur Urol 74:722–728
Vickers A, Cronin A, Roobol M et al (2010) Reducing unnecessary biopsy during prostate cancer screening using a four-kallikrein panel: an independent replication. J Clin Oncol 20(28):2493–2498
Beksac AT, Sobotka S, Xu P et al (2019) Downgrading of grade group after radical prostatectomy: comparison of multiparametric magnetic resonance imaging guided fusion biopsy and standard 12-core biopsy. Urology 127:80–85
Sanguedolce F, Falagario UG, Castellan P et al (2020) Bioptic intraprostatic chronic inflammation predicts adverse pathology at radical prostatectomy in patients with low-grade prostate cancer. Urol Oncol
Falagario UG, Beksac AT, Martini A, et al (2019) Defining prostate cancer at favorable intermediate risk: the potential utility of magnetic resonance imaging and genomic Tests. J Urol. 10.1097JU0000000000000134
Acknowledgements
Open access funding provided by Università di Foggia within the CRUI-CARE Agreement.
Funding
IMPROD trial, MULTI-IMPROD trial and the corresponding developmental work were financially supported by grants from the Instrumentarium Research Foundation, Sigrid Jusélius Foundation, Turku University Hospital, TYKS-SAPA research fund, Finnish Cancer Society, Finnish Cultural Foundation, and Orion Research Foundation. Pekka Taimen was supported by Clinical Researcher Funding from the Academy of Finland. No funding bodies had any role in the study design, data collection and analysis, decision to publish, or preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
UF: Project development, Data analysis, Methodology, Validation, Manuscript writing. IJ: Project development, Data collection or management, Data analysis, Protocol development, Funding acquisition, Methodology, Validation, Manuscript writing. PT: Data collection or management, Validation, Manuscript editing. KTS: Data collection or management, Manuscript editing. EK: Data collection or management, Manuscript editing. HM: Data analysis, Methodology, Manuscript editing. IMP: Data analysis, Methodology, Manuscript editing. JK: Data collection or management, Validation, Manuscript editing. AK: Data collection or management, Manuscript editing. JV: Data collection or management, Manuscript editing. AT: Project development, Funding acquisition, Supervision, Manuscript editing. HJA: Protocol development, Funding acquisition, Supervision, Manuscript editing. GC: Project development, Funding acquisition, Supervision, Manuscript editing. PB: Data collection or management, Protocol development, Funding acquisition, Manuscript editing. OE: Data collection or management, Supervision, Validation, Manuscript editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Research involving human participants, their data or biological material
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Ethics Committee of Turku University hospital. All patients signed informed consent. IMPROD: ClinicalTrials.gov Identifier: NCT01864135. MULTI-IMPROD: ClinicalTrials.gov Identifier: NCT02241122.
Informed consent
All patients included in this study signed an informed consent before enrollment (IMPROD trial, ClinicalTrials.gov Identifier: NCT01864135; Multi-IMPROD trial, ClinicalTrials.gov Identifier: NCT02241122).
Code availability
STATA Codes used in the statistical analyses in this manuscript are available upon request to the corresponding author. Free public access to the calculator presented in this manuscript is provided on the MULTI-IMPROD trial’s server: https://petiv.utu.fi/multiimprod/.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Falagario, U., Jambor, I., Taimen, P. et al. Added value of systematic biopsy in men with a clinical suspicion of prostate cancer undergoing biparametric MRI-targeted biopsy: multi-institutional external validation study. World J Urol 39, 1879–1887 (2021). https://doi.org/10.1007/s00345-020-03393-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00345-020-03393-8