Abstract
Treatment response and resistance in major depressive disorder (MDD) show a significant genetic component, but previous studies had limited power also due to MDD heterogeneity. This literature review focuses on the genetic factors associated with treatment outcomes in MDD, exploring their overlap with those associated with clinically relevant symptom dimensions. We searched PubMed for: (1) genome-wide association studies (GWASs) or whole exome sequencing studies (WESs) that investigated efficacy outcomes in MDD; (2) studies examining the association between MDD treatment outcomes and specific depressive symptom dimensions; and (3) GWASs of the identified symptom dimensions. We identified 13 GWASs and one WES of treatment outcomes in MDD, reporting several significant loci, genes, and gene sets involved in gene expression, immune system regulation, synaptic transmission and plasticity, neurogenesis and differentiation. Nine symptom dimensions were associated with poor treatment outcomes and studied by previous GWASs (anxiety, neuroticism, anhedonia, cognitive functioning, melancholia, suicide attempt, psychosis, sleep, sociability). Four genes were associated with both treatment outcomes and these symptom dimensions: CGREF1 (anxiety); MCHR1 (neuroticism); FTO and NRXN3 (sleep). Other overlapping signals were found when considering genes suggestively associated with treatment outcomes. Genetic studies of treatment outcomes showed convergence at the level of biological processes, despite no replication at gene or variant level. The genetic signals overlapping with symptom dimensions of interest may point to shared biological mechanisms and potential targets for new treatments tailored to the individual patient’s clinical profile.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Major depression disorder (MDD) is a leading cause of disability worldwide and ranks as the fourth leading cause of morbidity, according to the World Health Organization [1]. Estimates of the lifetime prevalence of depression range from 7 to 20% [2] and the 12-month prevalence can be as high as 5% [1].
Major guidelines recommend pharmacotherapy as first-line treatment for moderate to severe MDD [3, 4]. Antidepressants are among the most commonly prescribed medications worldwide [5] and have proven efficacy in reducing depressive symptoms. Currently, clinicians have access to several treatment options belonging to different classes of antidepressants, but none of them has clear evidence of superiority over the others and the remission rates with antidepressant therapy are still concerningly low [6]. Indeed, only ~ 30% of patients achieves remission after the first treatment, and even after four trials of treatment the percentage of remission does not reach 70% [7, 8]. After each treatment failure, the chance of response to the next antidepressant decreases and the risk of treatment-resistant depression (TRD), which is commonly defined as the lack of response to at least two treatments, increases [9, 10]. The trial-and-error process of finding an effective antidepressant can be prolonged and demoralizing, leading to delayed recovery and potentially contributing to the chronicity of the disease. Additionally, lack of treatment response exposes patients to a range of distressing and debilitating side effects [11, 12], which can undermine the therapeutic alliance.
Therefore, understanding the individual factors that influence treatment response in MDD is of primary importance to improve outcomes related not only to depressive symptoms, but also quality of life and overall functioning. Treatment response has a hereditary basis—for example, a concordance of antidepressant response- has been demonstrated in affected members of the same family [13] and a significant single nucleotide polymorphism (SNP)-based heritability (h2SNP) was found by genome-wide association studies (GWASs) [14]. Clinical and socio-demographic variables also contribute to treatment outcomes [15, 16]. Certain risk factors, such as suicidality and comorbid anxiety, may have a genetic basis overlapping with that of treatment response; conversely, socio-demographic variables and clinical factors such as duration and severity of the depressive episode, may exert effects independent from the genetics involved in treatment outcomes [17].
Although MDD is conceptualized as a single disorder, its diagnosis is formulated from a combination of symptoms that present with considerable variability, as over 1000 unique symptom combinations can be observed [18]. This phenotypic variability likely reflects the biological and environmental heterogeneity among patients, and may be partly linked to the heterogeneity observed in treatment response. Strong evidence shows that certain symptoms tend to co-occur, identifying subtypes of depression [19]. This is the case, for example, of atypical depression, which is characterized by the presence of mood reactivity and reversed neurovegetative symptoms (e.g. increased appetite/weight and hypersomnia). These symptoms likely have similar pathophysiological correlates and share common polygenic liabilities [20, 21]. On the other hand, distinct symptom domains are expressions of the heterogeneous genetic architecture of depression [22]. Consistently, specific symptom profiles tend to exhibit greater responsiveness to particular medications because of their pharmacodynamic profiles, in line with recommendations by the Canadian Network for Mood and Anxiety Treatments (CANMAT) guidelines [4]. Furthermore, certain depressive symptom dimensions such as anxiety, hypersomnia, anhedonia, suicidality, are associated with poorer treatment response, suggesting shared genetics between specific depressive symptom profiles and treatment outcomes [18].
Therefore, the aims of this narrative review are to (1) summarize the genetic factors associated with treatment efficacy outcomes in MDD; and (2) to explore the overlap between these genetic factors and those involved in specific symptom dimensions previously associated with treatment outcomes in MDD. This approach can provide information useful to identify specific biomarkers for treatment personalization, with implications for future research and clinical practice. To reach the second aim, we first reviewed the literature to identify the genetic signals associated with MDD treatment outcomes, focusing on GWASs and whole exome sequencing studies (WESs), given the instability of results from candidate gene studies [23, 24]. Then, we investigated whether these genetic associations overlap with those of MDD treatment outcomes. The rationale of this approach is to contribute to interpreting the existing literature in terms of genetic factors associated with treatment outcomes that may also be linked to specific symptoms, thereby helping to dissect the biological and clinical heterogeneity of poor treatment response and suggesting new treatment targets specific to certain symptom dimensions rather than MDD overall.
Methods
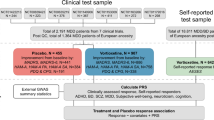
We searched PubMed, GWAS catalog (https://www.ebi.ac.uk/gwas/), GWAS atlas (https://atlas.ctglab.nl/), and medRxiv (https://www.medrxiv.org/) for original research articles on the following topics: (1) GWASs or WESs that investigated efficacy outcomes in MDD (response, remission, symptom improvement, or TRD, as defined in each original study); (2) the association between MDD treatment outcomes and specific depressive symptoms/dimensions; (3) GWASs that investigated the genetics of the symptom dimensions identified in (2).
When more GWASs were available in (3) for the same trait, we focused on the largest study demonstrating a significant h2SNP. We considered association signals at locus, gene, and gene set level. As the main topic of this work was to review the genetics of treatment outcomes in MDD, for studies in (1) we decided to present both genetic associations surviving multiple testing correction and suggestive association signals (p-value < 5 × 10–8 and 5 × 10–8 ≤ p-value < 5 × 10–6 at locus level, respectively). Although this was not a systematic review, we aimed to provide a comprehensive consideration of GWASs and WESs relevant to MDD treatment outcomes.
Finally, based on the overlap between genes identified (either significant or suggestive associations) from the SNP- and gene-level analyses of treatment outcomes and those significatively associated with the clinical dimensions in (3), we conducted a gene-drug interaction analysis via www.DGIdb.org [25].
Results
Genetic associations with treatment outcomes
A total of 13 GWASs and one WESs were included in this review, and their main characteristics are summarized in Table 1. Most studies had relatively small sample size (i.e., < 10 K), with only three exceptions, including one large study with 154,433 participants [26]; only 6 studies demonstrated a significant h2SNP of the outcome(s) of interest, with values generally around 0.8–0.10.
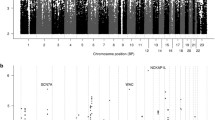
At locus level, 11 genome-wide significant associations were identified, spanning across multiple genes, including ITGA9, NRXN3, UST, MECOM, FTO, and MCHR1 (Table 2A). Additional suggestive signals were identified, including for example LINGO2, CACNA1C, PRG3, ITGA1, EPHB1 and SLC27A1 genes (Supplementary Table 1). However, we underline that these non-significant results have unclear relevance and were not replicated across studies.
A total of 25 genes were associated with the outcomes in the gene-level analyses, including LZTS3, PRNP, OR4K2, PPFIBP1, and GPHA2 (see Table 2B for all results). Suggestive associations were identified, including ADGRG5, MAP3K2, the solute carrier genes SLC17A4 and SLCO3A1, the glutamate receptor gene GRM3,, and genes encoding for many zinc finger proteins (Supplementary Table 2). However, none of these were replicated across studies.
Noteworthy, both suggestive and significant association findings had some consistency in terms of biological processes involved at the pathway level, showing an involvement of the immune system (e.g.: NCR3, LST1, and LCN2), synaptic transmission and dendritic spine formation (e.g.: PPFIBP1, PRNP, LZTS3, and NRXN3), neurogenesis and differentiation (e.g.: NRXN3, MECOM, CGREF1 and MAP1A), and regulation of transcription and gene expression (e.g.: DHX8, MECOM, ETV4, MEPCE, and PFAS). These observations are supported by the findings from gene set enrichment analyses (GSEAs). In particular, while no enrichment for specific gene sets was replicated across studies, GSEAs showed that many gene-sets are involved in or regulate similar biological processes. For example, signal transduction (Rhodopsine-like receptors A/1 R-HSA-373076 and Calcium-activated potassium channel activity GO:0015269); gene expression and nuclear functions (Chromosomal part GO:0044427 and Chromosome pathway GO:0005694), neurotransmission and synapse activity (Neuronal action potential GO:0019228, Transmission of nerve impulse GO:0019226, Long term potentiation hsa04720), immune function (Lymphocyte mediated immunity GO:0002449) ( Table 2C and Supplementary Table 3).
Genetic associations with symptom dimensions
Nine symptom dimensions were identified as associated with poor treatment outcomes in MDD and studied by previous GWASs showing significant h2SNP, namely: anxiety symptoms, neuroticism (including symptoms of apathy, worthlessness, guilt, loneliness, and excessive worry), anhedonia, cognitive functioning, melancholia, suicide attempt, psychotic symptoms, sleep symptoms, and sociability. No GWASs showed significant h2SNP for other symptoms also associated with poor therapeutic outcomes, such as irritable mood, inner tension, dissociative symptoms, and reverse or typical neurovegetative symptoms (with the exception of sleep-related symptoms) [15, 16, 27, 28].
The main characteristics of the included studies are summarised in Table 1, whilst full details and results are reported in the Supplementary Tables 1, 2, and 3. The selected GWASs were performed on large samples, in the range of 500 K participants, and all reported multiple significant genetic associations with the traits of interest (Table 1 and Supplementary Tables 1 and 2).
Given the focus of this review, a comprehensive description of these results is beyond our aims, while we were interested in describing the potential overlap with the genetic associations found for MDD treatment outcomes. Among the genes significatively associated with treatment outcomes, four overlapped with those associated with symptom dimensions, more precisely with anxiety (CGREF1), neuroticism (MCHR1) and sleep (FTO, NRXN3) (Table 3, in bold).
When considering also genes suggestively associated with treatment outcomes, additional signals were found in common with anxiety, anhedonia, executive functions, and sociability, as well as more overlapping genes for neuroticism and sleep (Table 3).
At gene set level, exact matches were found only with sleep, involving gene sets related to synaptic activity (hsa04730, hsa04730), G alpha signalling (R-HSA-418594), taurine/hypotaurine metabolism (hsa00430), immune response (BIOCARTA_VEGF_PATHWAY, hsa04612), and Alzheimer’s disease (hsa05010). However, even if no exact gene set overlaps were found between MDD treatment outcomes and other symptom dimensions, we identified patterns of possible overlap in some biological mechanisms involved. For example, gene sets associated with neuroticism, executive functions, and suicide attempts are predominantly related to neurogenesis and neurotransmission. Similarly, also other biological processes are involved both in MDD treatment outcome and some of the clinical dimensions, like synaptic plasticity (executive functions), immune response (suicide attempt, sleep), and nucleic acid and gene expression (suicide attempt) (Supplementary Table 3).
New therapeutic targets for treating symptom dimensions associated with poor response?
Based on the overlap discussed in the previous paragraph, we conducted a gene-drug interaction analysis. Of the four significant overlapping genes, no compounds were found to interact with either NRXN3 or CGREF1, while interactions were found only for the melanin concentrating hormone receptor 1 (MCHR1) and the fat mass and obesity associated gene (FTO, also known as ALKBH9). MCHR1, also known as SLC1, is a G-protein coupled receptor (GPCR) which binds the melanin-concentrating hormone (MCH, or PMCH) and inhibits cAMP accumulation while stimulating intracellular calcium influx. MCH is likely involved in the regulation of feeding behaviour, mood, sleep–wake cycle and energy balance [29]. Four still non-approved compounds resulted to interact with MCHR1, of which SNAP-7941 is the lead compound of MCHR1-inhibitors and displayed promising anxiolytic, antidepressant, and anorectic effects, even though not replicated in clinical trials [30]. Another MCHR1 antagonist, BMS-830216, is currently in phase 2 for the treatment of obesity [31].
Concerning FTO, eight compounds were found, all already approved as antineoplastics (INFɑ-2A, INFɑ-2B, mercaptopurine, and bisantrene), antiviral (ribavirin), antiarrhythmic (atenolol), antihypertensive (atenolol, hydrochlorothiazide), and disease-modifying antirheumatic drugs (azathioprine). FTO’s exact physiological function is yet to be uncovered; however, it is a non-heme iron enzyme located in the nucleus and likely related to growth, development, BMI, obesity, and type 2 diabetes mellitus [32].
When considering the overlap with genes associated with poor treatment outcome at a suggestive level, interesting gene-drug interactions were found for GRM3 with risperidone and two selective mGluR2/3 agonists (LY2969822 and LY404039, and the corresponding prodrug of the latter LY2140023); and CACNA1C with haloperidol, citalopram, valproate, and gabapentinoids. For all gene-drug interactions, see Table 3.
Discussion
The identification of genetic factors modulating MDD treatment outcomes has been challenging and led to few clinical applications, limited to genes involved in drug metabolism [33]. In the 50 years after the first evidence of a substantial heritability coming from family genetic studies [33], many studies focused on candidate genes, with minor and mainly unreplicated findings. In the last 10 years, GWASs produced more interesting findings, due to a more extensive coverage of the genome and larger samples, as discussed in this review. To date, it has been estimated that genetics may account for up to 60% of the variance in treatment resistance according to pedigree-based heritability [34].
However, the polygenic nature of MDD treatment outcomes and the relatively limited size of most samples resulted in scattered results which do not generally overlap, at least at SNP or gene level. The redundancies among the pathways modulating antidepressant outcomes and the heterogeneity of depressive symptoms are likely involved in the discrepancy of results at SNP and gene level [35,36,37,38]. The approach used in the present review aimed to partially overcome these issues, by integrating the genetic signals associated with antidepressant outcomes and specific symptom dimensions of clinical relevance, and by extending the analysis to pathways, pointing out potential mechanisms involved in treatment resistance and possible treatment targets.
Both at variant/gene level and gene set level, treatment outcomes were linked to gene expression regulation, central nervous system (CNS) development, synaptic plasticity, and immune system activity. One speculative interpretation of these results is that anomalies in the regulation of gene expression concur with abnormal CNS development (from tissue differentiation to synapse formation) and with an aberrant brain-immunity interplay, resulting in increased risk of developing more severe and less treatment-responsive MDD.
We identified four genes that were significantly associated with both treatment outcomes and the clinical dimensions of interest, namely CGREF1 (anxiety), MCHR1 (neuroticism), FTO and NRXN3 (sleep). NRXN3 (neurexin 3) encodes for a surface protein acting as cell adhesion molecule-receptor and it is likely involved in synaptic plasticity [39]. Other than with treatment response, it was also associated with sleep health (Supplementary Tables 1 and 2), suggesting a protective function on brain physiological activity. Polymorphisms in this gene have been linked to obesity and substance use disorders [40, 41], suggesting an influence on impulse control. This increases the interest towards this gene and the possibility to target/modulate it by future therapies, despite currently there are no known compounds/molecules targeting NRXN3.
FTO (fat mass and obesity-associated protein) and MCHR1 (melanin concentrating hormone receptor 1) were both significantly associated with TRD and with sleep and neuroticism, respectively. Both are involved in energy homeostasis, but FTO acts more on a cellular level regulating adipogenesis and fat mass [32, 42,43,44], while MCHR1 is involved in the regulation of feeding behaviour and energy balance and has known effects on mood and sleep–wake cycle [45]. Variants in MCHR1 have been linked to a reduced expression in the dorsolateral prefrontal cortex [46] and are in linkage disequilibrium with another variant recently linked to an increased risk of bipolar disorder [47]. Several drugs interact with FTO but, as far as we know, no specific compound has been developed or studied specifically. Interestingly, FTO also maps to a genomic region showing significant local genetic correlation between MDD and type 2 diabetes mellitus and obesity, suggesting that FTO could be implicated in the shared etiopathogenesis between MDD and insulin resistance-related conditions [48]. We found FTO to interact with antihypertensive drugs, i.e. hydrochlorothiazide (a diuretic) and atenolol (a β-blocker). Angiotensin agents, calcium-channel blockers, and β-blockers (but not diuretics) have been recently associated with decreased rates of depression [49]. On the other hand, MCHR1 has been investigated and promising results were found for some compounds [50, 51], but results were not replicated [30].
CGREF1 (cell growth regulator with EF-hand domain 1) inhibits cell proliferation, despite its function has not been fully elucidated [52]. We found that this gene was associated with remission to antidepressants and anxiety (Supplementary Table 2), but it was implicated in other traits as well by previous GWASs, such as brain measures, risk-taking behaviours, wellbeing, and educational attainment [53,54,55,56,57]. Therefore, CGREF1 may be involved in the modulation of multiple but likely connected phenotypes, which are relevant for antidepressant effects. Currently, there are no known compounds that target CGREF1.
Other genes were not significantly associated with antidepressant outcomes, however suggestive findings were reported (Supplementary Tables 1–3). Among these genes, we outline GRM3, [58], SLCO3A1, LINGO1 and LINGO2. EPHB1 (Supplementary Table 1) regulates chemotaxis and proliferation of neural progenitors in the hippocampus [58], a well known region for mood disorders physiopathology and AD efficacy [59, 60]. It was first identified as potentially involved in antidepressant response by one of the first GWASs in this field, which reported a suggestive association with a SNP located downstream of this gene [61]; however, this GWAS was not formally included in the present review, as it included both unipolar (MDD) and bipolar depression, while this work was focused on MDD. This gene was also associated with several symptom dimensions of interest, including neuroticism, anhedonia, and executive functions. EPHB1 has a key role in axon guidance and, with other ephrin-B receptors, is involved in the development and maturation of dendritic spine and synapse formation [58]. To date, no specific drug exists to selectively target EPHB1, however it interacts with progesterone [25], which has been proved to modulate the expression of γ-aminobutyric acid type-A receptors (GABAAR) via its metabolite allopregnanolone [62]. Noteworthy, brexanolone, a synthetic allopregnanolone analogous, is approved for the treatment of postpartum depression [63].
GRM3 represents an interesting finding. Unlike NMDA (N-Methyl-D-aspartic acid) and AMPA (α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid) receptors, glutamate metabotropic receptors are G-protein coupled receptors (GPCR) with more complex effects, involved in synapse plasticity, for example long-term depression (LTD) at excitatory synapses [64]. GRM3 was associated with neuroticism and executive functions (Supplementary Table 2). Thus, compounds acting at this site may hypothetically be beneficial for patients experiencing a depressive episode with a sense of guilt and worthlessness, interpersonal sensibility, and/or cognitive symptoms. To date, the only drug that seems to interact with GRM3 is risperidone, a second-generation antipsychotic (SGA) used as adjunctive therapy in psychotic depression or as augmentation to antidepressants in TRD [65]. Other compounds targeting glutamate receptors are under study for the treatment of schizophrenia [66] and depressive disorders, such as MGS-0210 [67], which is a selective metabotropic glutamate receptor antagonist with antidepressant-like activity [68].
Even if only suggestive, the association of the organic anion transporter SLCO3A1 with remission after treatment may suggest a role of endogenous organic anions, vasopressin and prostaglandins in MDD outcome [69, 70]. Noteworthy, SLCO3A1 is significatively expressed in the CNS, especially in oligodendrocytes [70, 71] but also in neurons and grey matter glial cells [69, 72]. However, SLCO3A1 activity is not yet fully understood and new interactions have been found with other exogenous compounds [73], such as modulation of its expression levels by valproic acid [74].
The role of energy metabolism in mood disorders is well known, as discussed above for FTO and MCHR1, and there is an increasing consensus on the involvement, for example, of glucidic metabolism in brain disorders [75, 76]. We observed significant association with many genes involved in feeding behaviour and sleep–wake cycle. LINGO1 and LINGO2 were associated with symptom remission and neuroticism, and are likely involved in synapse assembly [77], other than being associated with BMI [78]. No drug exists to date targeting these genes, but they are modulated by resveratrol and vitamin D [79] and they may represent putative targets for future complementary treatments, for example in patients with higher levels of neuroticism and BMI [80, 81].
This review provides a comprehensive overview of GWASs and WESs on treatment outcomes in MDD, also leveraging an innovative, clinically-oriented approach to explore the complex genetics of MDD treatment outcomes. By integrating genetic signals associated with MDD treatment outcomes and specific depressive symptom dimensions, our approach may pave the way for developing targeted treatments for non-responsive patients exhibiting specific symptom profiles. However, several limitations should be acknowledged. Firstly, we did not perform statistical analyses to test the genetic overlap between treatment outcomes and the symptom dimensions of interest; however, this was beyond the aims of this paper, being this work intended as a review of the literature. A common limitation of the included GWASs on MDD treatment outcomes was the relatively small sample size, and the resulting limited power to detect genome-wide significant associations and to replicate findings across studies. Last but not least, pharmacogenetic findings and targets identified by gene-drug interactions need functional validation to assess their potential clinical relevance and applicability. This consideration outlines the importance of using complementary and integrated research approaches, for example in vitro/in vivo models evaluating compound properties and activity.
In conclusion, this review presents significant insights into the genomics of treatment outcomes in MDD, highlighting the existence of genetic factors overlapping with specific clinical dimensions that are in turn associated with poor treatment outcomes. We prioritised four genes, including CGREF1, MCHR1, FTO, and NRXN3, which are linked to both MDD treatment outcomes and relevant clinical dimensions. These findings highlight the potential for developing new treatments that target specific depressive symptom dimensions, contributing to the advancement of precision psychiatry.
References
W. World Health Organization (2023) Depression. https://www.who.int/health-topics/depression#tab=tab_1. Accessed 31 May 2024
Sadock BJ, Sadock VA, Ruiz P, Kaplan HI (2017) Kaplan and Sadock’s Comprehensive Textbook of Psychiatry. Wolters Kluwer Health/Lippincott Williams & Wilkins, Philadelphia
Taylor DM, Barnes TR, Young AH (2021) In Maudsley Prescribing Guidelines in Psychiatry, 14th edn. Wiley Blackwell, Hoboken
Yatham LN et al (2018) Canadian network for mood and anxiety treatments (CANMAT) and international society for bipolar disorders (ISBD) 2018 guidelines for the management of patients with bipolar disorder. Bipolar Disord 20(2):97–170. https://doi.org/10.1111/bdi.12609
Fuentes AV, Pineda MD, Venkata KCN (2018) Comprehension of top 200 prescribed drugs in the US as a resource for pharmacy teaching, training and practice. Pharm (Basel) 6(2):43. https://doi.org/10.3390/pharmacy6020043
Sinyor M, Schaffer A, Levitt A (2010) The sequenced treatment alternatives to relieve depression (STAR*D) trial: a review. Can J Psychiatry 55(3):126–135. https://doi.org/10.1177/070674371005500303
Rush AJ et al (2006) Acute and longer-term outcomes in depressed outpatients requiring one or several treatment steps: a STAR*D report. AJP 163(11):1905–1917. https://doi.org/10.1176/ajp.2006.163.11.1905
Warden D, Rush AJ, Trivedi MH, Fava M, Wisniewski SR (2007) The STAR* D project results: a comprehensive review of findings. Curr Psychiatry Rep 9(6):449–459
McIntyre RS et al (2023) Treatment-resistant depression: definition, prevalence, detection, management, and investigational interventions. World Psychiatry 22(3):394–412. https://doi.org/10.1002/wps.21120
Sforzini L et al (2022) A Delphi-method-based consensus guideline for definition of treatment-resistant depression for clinical trials. Mol Psychiatry 27(3):1286–1299. https://doi.org/10.1038/s41380-021-01381-x
Oliva V et al (2021) Gastrointestinal side effects associated with antidepressant treatments in patients with major depressive disorder: a systematic review and meta-analysis. Prog Neuropsychopharmacol Biol Psychiatry 109:110266
Serretti A, Mandelli L, Laura M (2010) Antidepressants and body weight: a comprehensive review and meta-analysis. J Clin Psychiatry 71(10):979
Franchini L, Serretti A, Gasperini M, Smeraldi E (1998) Familial concordance of fluvoxamine response as a tool for differentiating mood disorder pedigrees. J Psychiatr Res 32(5):255–259
Pain O et al (2021) Identifying the common genetic basis of antidepressant response. Biol Psychiatry Glob Open Sci 2(2):115–126. https://doi.org/10.1016/j.bpsgos.2021.07.008
De Carlo V, Calati R, Serretti A (2016) Socio-demographic and clinical predictors of non-response/non-remission in treatment resistant depressed patients: a systematic review. Psychiatry Res 240:421–430. https://doi.org/10.1016/j.psychres.2016.04.034
Kautzky A et al (2017) Refining prediction in treatment-resistant depression: results of machine learning analyses in the TRD III sample. J Clin Psychiatry 79(1):14989
Fabbri C et al (2019) The genetics of treatment-resistant depression: a critical review and future perspectives. Int J Neuropsychopharmacol 22(2):93–104
Fried EI, Nesse RM (2015) Depression is not a consistent syndrome: an investigation of unique symptom patterns in the STAR*D study. J Affect Disord 172:96–102. https://doi.org/10.1016/j.jad.2014.10.010
Lamers F et al (2010) Identifying depressive subtypes in a large cohort study: results from the Netherlands study of depression and anxiety (NESDA). J Clin Psychiatry 71(12):8450
Badini I et al (2022) Depression with atypical neurovegetative symptoms shares genetic predisposition with immuno-metabolic traits and alcohol consumption. Psychol Med 52(4):726–736. https://doi.org/10.1017/S0033291720002342
Milaneschi Y et al (2016) Polygenic dissection of major depression clinical heterogeneity. Mol Psychiatry 21(4):516–522
Thorp JG, Marees AT, Ong J-S, An J, MacGregor S, Derks EM (2020) Genetic heterogeneity in self-reported depressive symptoms identified through genetic analyses of the PHQ-9. Psychol Med 50(14):2385–2396
Border R et al (2019) No support for historical candidate gene or candidate gene-by-interaction hypotheses for major depression across multiple large samples. Am J Psychiatry 176(5):376–387. https://doi.org/10.1176/appi.ajp.2018.18070881
Fabbri C et al (2017) Consensus paper of the WFSBP task force on genetics: genetics, epigenetics and gene expression markers of major depressive disorder and antidepressant response. World J Biol Psychiatry 18(1):5–28. https://doi.org/10.1080/15622975.2016.1208843
Cannon M et al (2024) DGIdb 5.0: rebuilding the drug-gene interaction database for precision medicine and drug discovery platforms. Nucl Acids Res 52(D1):D1227–D1235. https://doi.org/10.1093/nar/gkad1040
Kang J et al (2023) Genome-wide association study of treatment resistant depression highlights shared biology with metabolic traits. MedRxiv. https://doi.org/10.1101/2022.08.10.22278630
Huang Y et al (2023) Comparison on the clinical features in patients with or without treatment-resistant depression: a national survey on symptomatology of depression report. Psychiatry Res 319:114972. https://doi.org/10.1016/j.psychres.2022.114972
Uher R et al (2012) Depression symptom dimensions as predictors of antidepressant treatment outcome: replicable evidence for interest-activity symptoms. Psychol Med 42(5):967–980. https://doi.org/10.1017/S0033291711001905
Verret L et al (2003) A role of melanin-concentrating hormone producing neurons in the central regulation of paradoxical sleep. BMC Neurosci 4:19. https://doi.org/10.1186/1471-2202-4-19
Basso AM et al (2006) Lack of efficacy of melanin-concentrating hormone-1 receptor antagonists in models of depression and anxiety. Eur J Pharmacol 540(1):115–120. https://doi.org/10.1016/j.ejphar.2006.04.043
Squibb B-M (2015) Randomized, double-blind, placebo-controlled, ascending multiple-dose and parallel arm study to evaluate the safety, pharmacokinetics and pharmacodynamics of BMS-830216 (prodrug of BMS-819881) in obese subjects’, clinicaltrials.gov, clinical trial registration NCT00909766. https://clinicaltrials.gov/study/NCT00909766. Accessed 01 Jan 2024
PubChem FTO - FTO alpha-ketoglutarate dependent dioxygenase (human). https://pubchem.ncbi.nlm.nih.gov/gene/FTO/human. Accessed 20 May 2024
Bousman CA et al (2021) Review and consensus on pharmacogenomic testing in psychiatry. Pharmacopsychiatry 54(1):5–17. https://doi.org/10.1055/a-1288-1061
Wigmore EM et al (2020) Genome-wide association study of antidepressant treatment resistance in a population-based cohort using health service prescription data and meta-analysis with GENDEP. Pharmacogenomics J 20(2):329–341. https://doi.org/10.1038/s41397-019-0067-3
Adams MJ, Lewis CM, McIntosh AM, the P. G. C. M. D. D. W. Group (2024) Genome-wide study of major depression in 685,808 diverse individuals identifies 697 independent associations, infers causal neuronal subtypes and biological targets for novel pharmacotherapies. MedRxiv. https://doi.org/10.1101/2024.04.29.24306535
Ferrari F, Villa RF (2017) The neurobiology of depression: an integrated overview from biological theories to clinical evidence. Mol Neurobiol 54(7):4847–4865. https://doi.org/10.1007/s12035-016-0032-y
Krishnan V, Nestler EJ (2008) The molecular neurobiology of depression. Nature 455(7215):894–902. https://doi.org/10.1038/nature07455
Rantamäki T, Yalcin I (2016) Antidepressant drug action–from rapid changes on network function to network rewiring. Prog Neuropsychopharmacol Biol Psychiatry 64:285–292. https://doi.org/10.1016/j.pnpbp.2015.06.001
Zhang R, Jiang H, Liu Y, He G (2022) Structure, function, and pathology of neurexin-3. Genes Dis 10(5):1908–1919. https://doi.org/10.1016/j.gendis.2022.04.008
Heard-Costa NL et al (2009) NRXN3 is a novel locus for waist circumference: a genome-wide association study from the CHARGE consortium. PLoS Genet 5(6):e1000539. https://doi.org/10.1371/journal.pgen.1000539
Hishimoto A et al (2007) Neurexin 3 polymorphisms are associated with alcohol dependence and altered expression of specific isoforms. Hum Mol Genet 16(23):2880–2891. https://doi.org/10.1093/hmg/ddm247
Huang Y et al (2015) Meclofenamic acid selectively inhibits FTO demethylation of m6A over ALKBH5. Nucleic Acids Res 43(1):373–384. https://doi.org/10.1093/nar/gku1276
Jia G et al (2011) N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat Chem Biol 7(12):885–887. https://doi.org/10.1038/nchembio.687
Wei J et al (2018) Differential m6A, m6Am, and m1A demethylation mediated by FTO in the cell nucleus and cytoplasm. Mol Cell 71(6):973-985.e5. https://doi.org/10.1016/j.molcel.2018.08.011
MCHR1 melanin concentrating hormone receptor 1 [Homo sapiens (human)]-Gene-NCBI. https://www.ncbi.nlm.nih.gov/gene/2847. Accessed 25 May 2024
Ochoa D et al (2021) Open targets platform: supporting systematic drug-target identification and prioritisation. Nucleic Acids Res 49(D1):D1302–D1310. https://doi.org/10.1093/nar/gkaa1027
Mullins N et al (2021) Genome-wide association study of over 40,000 bipolar disorder cases provides new insights into the underlying biology. Nat Genet 53(6):817. https://doi.org/10.1038/s41588-021-00857-4
Fanelli G et al (2024) Local patterns of genetic sharing challenge the boundaries between neuropsychiatric and insulin resistance-related conditions. MedRxiv. https://doi.org/10.1101/2024.03.07.24303921
Kessing LV, Rytgaard HC, Ekstrøm CT, Torp-Pedersen C, Berk M, Gerds TA (2020) Antihypertensive drugs and risk of depression: a nationwide population-based study. Hypertension 76(4):1263–1279. https://doi.org/10.1161/HYPERTENSIONAHA.120.15605
ATC0175 | MCH1R antagonist | MedChemExpress, MedchemExpress.com. https://www.medchemexpress.com/atc0175.html. Accessed 25 May 2024
Chaki S et al (2005) ATC0175: an orally active melanin-concentrating hormone receptor 1 antagonist for the potential treatment of depression and anxiety. CNS Drug Rev 11(4):341–352. https://doi.org/10.1111/j.1527-3458.2005.tb00052.x
Xiang M, Gao Y, Zhou Y, Wang M, Yao X (2023) A novel nomogram based on cell cycle-related genes for predicting overall survival in early-onset colorectal cancer. BMC Cancer 23(1):595. https://doi.org/10.1186/s12885-023-11075-y
Clifton EAD et al (2018) Genome-wide association study for risk taking propensity indicates shared pathways with body mass index. Commun Biol 1:36. https://doi.org/10.1038/s42003-018-0042-6
Lee J et al (2023) Quantifying the causal impact of biological risk factors on healthcare costs. Nat Commun 14(1):5672. https://doi.org/10.1038/s41467-023-41394-4
Okbay A et al (2022) Polygenic prediction of educational attainment within and between families from genome-wide association analyses in 3 million individuals. Nat Genet 54(4):437–449. https://doi.org/10.1038/s41588-022-01016-z
Sha Z et al (2021) The genetic architecture of structural left-right asymmetry of the human brain. Nat Hum Behav 5(9):1226–1239. https://doi.org/10.1038/s41562-021-01069-w
Wainberg M et al (2024) Genetic architecture of the structural connectome. Nat Commun 15(1):1962. https://doi.org/10.1038/s41467-024-46023-2
EPHB1-ephrin type-B receptor 1-Homo sapiens (Human) | UniProtKB | UniProt. https://www.uniprot.org/uniprotkb/P54762/entry#function. Accessed 23 May 2024
Dusi N, Barlati S, Vita A, Brambilla P (2015) Brain structural effects of antidepressant treatment in major depression. Curr Neuropharmacol 13(4):458–465. https://doi.org/10.2174/1570159X1304150831121909
Perlman K et al (2019) A systematic meta-review of predictors of antidepressant treatment outcome in major depressive disorder. J Affect Disord 243:503–515. https://doi.org/10.1016/j.jad.2018.09.067
Ising M et al (2009) A genomewide association study points to multiple loci that predict antidepressant drug treatment outcome in depression. Arch Gen Psychiatry 66(9):966–975. https://doi.org/10.1001/archgenpsychiatry.2009.95
Kapur J, Joshi S (2021) Progesterone modulates neuronal excitability bidirectionally. Neurosci Lett 744:135619. https://doi.org/10.1016/j.neulet.2020.135619
Walton N, Maguire J (2019) Allopregnanolone-based treatments for postpartum depression: why/how do they work? Neurobiol Stress 11:100198. https://doi.org/10.1016/j.ynstr.2019.100198
Kang SJ, Kaang B-K (2016) Metabotropic glutamate receptor dependent long-term depression in the cortex. Korean J Physiol Pharmacol 20(6):557–564. https://doi.org/10.4196/kjpp.2016.20.6.557
Kennedy SH et al (2016) Canadian network for mood and anxiety treatments (CANMAT) 2016 clinical guidelines for the management of adults with major depressive disorder:section 3. Pharmacological treatments. Can J Psychiatry 61(9):540–560. https://doi.org/10.1177/0706743716659417
Kryszkowski W, Boczek T (2021) The G protein-coupled glutamate receptors as novel molecular targets in schizophrenia treatment—a narrative review. J Clin Med 10(7):1475. https://doi.org/10.3390/jcm10071475
NCATS inxight drugs — MGS-0210. https://drugs.ncats.io/substance/12XH8EKL2A. Accessed 24 May 2024
Chaki S et al (2004) MGS0039: a potent and selective group II metabotropic glutamate receptor antagonist with antidepressant-like activity. Neuropharmacology 46(4):457–467. https://doi.org/10.1016/j.neuropharm.2003.10.009
Hagenbuch B, Stieger B (2013) The SLCO (former SLC21) superfamily of transporters. Mol Aspects Med 34(2):396–412. https://doi.org/10.1016/j.mam.2012.10.009
SLCO3A1 protein expression summary-the human protein atlas. https://www.proteinatlas.org/ENSG00000176463-SLCO3A1. Accessed 22 May 2024
SLCO3A1 solute carrier organic anion transporter family member 3A1 [Homo sapiens (human)]-gene-NCBI. https://www.ncbi.nlm.nih.gov/gene/28232. Accessed 22 May 2024
Huber RD et al (2007) Characterization of two splice variants of human organic anion transporting polypeptide 3A1 isolated from human brain. Am J Physiol Cell Physiol 292(2):C795–C806. https://doi.org/10.1152/ajpcell.00597.2005
Li T et al (2023) Organic anion transporting polypeptide 3a1 is a novel influx pump for perfluorooctane sulfonate in sertoli cells and contributes to its reproductive toxicity. Chemosphere 345:140428. https://doi.org/10.1016/j.chemosphere.2023.140428
Pharmaco-transcriptomics | DrugBank online. https://go.drugbank.com/pharmaco/transcriptomics?page=3699. Accessed 29 May 2024
Fanelli G et al (2022) Insulinopathies of the brain? genetic overlap between somatic insulin-related and neuropsychiatric disorders. Transl Psychiatry 12(1):59. https://doi.org/10.1038/s41398-022-01817-0
Possidente C, Fanelli G, Serretti A, Fabbri C (2023) Clinical insights into the cross-link between mood disorders and type 2 diabetes: a review of longitudinal studies and Mendelian randomisation analyses. Neurosci Biobehav Rev 152:105298. https://doi.org/10.1016/j.neubiorev.2023.105298
LINGO2 leucine rich repeat and Ig domain containing 2 [Homo sapiens (human)]-Gene-NCBI. https://www.ncbi.nlm.nih.gov/gene/158038. Accessed 23 May 2024
Speliotes EK et al (2010) Association analyses of 249,796 individuals reveal 18 new loci associated with body mass index. Nat Genet 42(11):937–948. https://doi.org/10.1038/ng.686
Pirmoradi Z et al (2024) Resveratrol and 1,25-dihydroxyvitamin D decrease Lingo-1 levels, and improve behavior in harmaline-induced essential tremor, suggesting potential therapeutic benefits. Sci Rep 14(1):9864. https://doi.org/10.1038/s41598-024-60518-4
Moore A, Beidler J, Hong MY (2018) Resveratrol and depression in animal models: a systematic review of the biological mechanisms. Molecules 23(9):2197. https://doi.org/10.3390/molecules23092197
Wei R-M et al (2023) Resveratrol ameliorates maternal separation-induced anxiety- and depression-like behaviors and reduces Sirt1-NF-kB signaling-mediated neuroinflammation. Front Behav Neurosci 17:1172091. https://doi.org/10.3389/fnbeh.2023.1172091
Huang S-S et al (2023) Investigating genetic variants for treatment response to selective serotonin reuptake inhibitors in syndromal factors and side effects among patients with depression in Taiwanese Han population. Pharmacogenomics J 23(2):50–59. https://doi.org/10.1038/s41397-023-00298-8
Fabbri C et al (2021) Genetic and clinical characteristics of treatment-resistant depression using primary care records in two UK cohorts. Mol Psychiatry 26(7):3363–3373. https://doi.org/10.1038/s41380-021-01062-9
Kang H-J et al (2020) Genetic markers for later remission in response to early improvement of antidepressants. Int J Mol Sci. https://doi.org/10.3390/ijms21144884
Li QS, Tian C, Hinds D (2020) Genome-wide association studies of antidepressant class response and treatment-resistant depression. Transl Psychiatry 10(1):1–12. https://doi.org/10.1038/s41398-020-01035-6
Fabbri C et al (2019) Genome-wide association study of treatment-resistance in depression and meta-analysis of three independent samples. Br J Psychiatry 214(1):36–41. https://doi.org/10.1192/bjp.2018.256
Fabbri C et al (2018) New insights into the pharmacogenomics of antidepressant response from the GENDEP and STAR*D studies: rare variant analysis and high-density imputation. Pharmacogenomics J 18(3):413–421. https://doi.org/10.1038/tpj.2017.44
Li QS, Tian C, Seabrook GR, Drevets WC, Narayan VA (2016) Analysis of 23andMe antidepressant efficacy survey data: implication of circadian rhythm and neuroplasticity in bupropion response. Transl Psychiatry 6(9):e889–e889. https://doi.org/10.1038/tp.2016.171
Hunter AM et al (2013) A genome-wide association study of a sustained pattern of antidepressant response. J Psychiatr Res 47(9):1157–1165. https://doi.org/10.1016/j.jpsychires.2013.05.002
Tansey KE et al (2012) Genetic predictors of response to serotonergic and noradrenergic antidepressants in major depressive disorder: a genome-wide analysis of individual-level data and a meta-analysis. PLoS Med 9(10):e1001326. https://doi.org/10.1371/journal.pmed.1001326
Uher R et al (2010) Genome-wide pharmacogenetics of antidepressant response in the GENDEP project. AJP 167(5):555–564. https://doi.org/10.1176/appi.ajp.2009.09070932
Friligkou E et al (2024) Gene discovery and biological insights into anxiety disorders from a multi-ancestry genome-wide association study of > 1.2 million participants. MedRxiv. https://doi.org/10.1101/2024.02.14.24302836
Nagel M, Watanabe K, Stringer S, Posthuma D, van der Sluis S (2018) Item-level analyses reveal genetic heterogeneity in neuroticism. Nat Commun 9(1):905. https://doi.org/10.1038/s41467-018-03242-8
Ward J et al (2019) Novel genome-wide associations for anhedonia, genetic correlation with psychiatric disorders, and polygenic association with brain structure. Transl Psychiatry 9(1):327. https://doi.org/10.1038/s41398-019-0635-y
Hatoum AS et al (2023) Genome-wide association study shows that executive functioning is influenced by GABAergic processes and is a neurocognitive genetic correlate of psychiatric disorders. Biol Psychiatry 93(1):59–70. https://doi.org/10.1016/j.biopsych.2022.06.034
Cai N et al (2015) Sparse whole-genome sequencing identifies two loci for major depressive disorder. Nature 523(7562):588–591. https://doi.org/10.1038/nature14659
Docherty AR et al (2020) Genome-wide association study of suicide death and polygenic prediction of clinical antecedents. Am J Psychiatry 177(10):917–927. https://doi.org/10.1176/appi.ajp.2020.19101025
Barkhuizen W, Pain O, Dudbridge F, Ronald A (2020) Genetic overlap between psychotic experiences in the community across age and with psychiatric disorders. Transl Psychiatry 10(1):1–12. https://doi.org/10.1038/s41398-020-0765-2
Bralten J et al (2021) Genetic underpinnings of sociability in the general population. Neuropsychopharmacol 46(9):1627–1634. https://doi.org/10.1038/s41386-021-01044-z
Goodman MO et al (2024) Genome-wide association analysis of composite sleep health scores in 413,904 individuals. MedRxiv. https://doi.org/10.1101/2024.02.02.24302211
Uher R et al (2013) Common genetic variation and antidepressant efficacy in major depressive disorder: a meta-analysis of three genome-wide pharmacogenetic studies. AJP 170(2):207–217. https://doi.org/10.1176/appi.ajp.2012.12020237
Funding
Open access funding provided by Alma Mater Studiorum - Università di Bologna within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
Alessandro Serretti is or has been a consultant/speaker for Abbott, Abbvie, Angelini, AstraZeneca, Clinical Data, Boehringer Ingelheim, Bristol-Myers Squibb, Eli Lilly, GlaxoSmithKline, Innovapharma, Italfarmaco, Janssen, Lundbeck, Naurex, Pfizer, Polifarma, Sanofi, Servier, and Taliaz. Chiara Fabbri was a speaker for Janssen. The other authors declare no conflicts of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Martone, A., Possidente, C., Fanelli, G. et al. Genetic factors and symptom dimensions associated with antidepressant treatment outcomes: clues for new potential therapeutic targets?. Eur Arch Psychiatry Clin Neurosci (2024). https://doi.org/10.1007/s00406-024-01873-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00406-024-01873-1