Abstract
Serine incorporator 5 (SERINC5 or SER5) is a multipass transmembrane protein with ill-defined cellular activities. SER5 was recently described as a human immunodeficiency virus 1 (HIV-1) restriction factor capable of inhibiting HIV-1 that does not express its accessory protein Nef (Δ Nef). SER5 incorporated into the viral membrane impairs the entry of HIV-1 by disrupting the fusion between the viral and the plasma membrane after envelope receptor interaction induced the first steps of the fusion process. The mechanisms of how SER5 prevents membrane fusion are not fully understood and viral envelope proteins were identified that escape the SER5-mediated restriction. Primate lentiviruses, such as HIV-1 and simian immunodeficiency viruses (SIVs), use their accessory protein Nef to downregulate SER5 from the plasma membrane by inducing an endocytic pathway. In addition to being directly antiviral, recent data suggest that SER5 is an important adapter protein in innate signaling pathways leading to the induction of inflammatory cytokines. This review discusses the current knowledge about HIV-1 restriction by SER5.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Discovery of the SERINC5 anti-HIV-1 activity and Nef counteraction
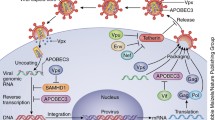
Some host cellular proteins act as the first line of defense against infection of human immunodeficiency virus 1 (HIV-1) and other viruses by blocking directly different cellular steps of the viral replication cycle [1]. These host proteins also called restriction factors are part of the cellular innate immune system [2]. Well-described inhibitory proteins of HIV-1 include APOBEC3 cytidine deaminases [3,4,5,6], Tetherin/BST2 [7,8,9,10], SAMHD1 [11,12,13], and TRIM5α [14, 15]. Most recently, MX2 [16,17,18,19], SER3/5 [20,21,22], IFITM3 [23,24,25], SLNF11 [26] and MARCH2/8 [27, 28] have been reported. This review discusses the current knowledge about HIV-1 restriction by SER5. Studies focused on Nef function allowed the discovery of SER3 and 5 (SER3, SER5) as antiviral factors. Initially, it was observed that Nef from HIV-1 [29] and the glycoGag from the murine leukemia virus (MLV) [30] increase the infectivity of HIV-1 particles due to an unknown function in the virus producer cell. To understand these observations, in 2015, two independent studies with different approaches were able to identify these new restriction factors that are antagonized by the Nef. These experiments were based on proteomic analyzes [21] and transcriptional profiles [20] of cells permissive or resistant to HIV-1 not expressing Nef (∆ Nef) that allowed the identification of SER3 and SER5 proteins incorporated in virions generated from viral DNAs not encoding functional Nef [20, 21]. Besides HIV-1 Nef and MLV glycoGag, the S2 protein of the equine infectious anemia virus (EIAV) and also the glycoGag of the feline leukemia virus (FeLV) have been shown to counteract SER5 [20, 21, 31,32,33].
SER genes and evolution
The serine incorporator (SERINC or SER) proteins are a family of eukaryote multi-transmembrane proteins highly conserved among different species [34], with recent evidence increasingly supporting their role in intrinsic immunity against retroviruses [22]. S. cerevisiae and D. melanogaster have only one gene of this family named TMS1. C. elegans, C. intestinalis, and C. savignyi encode likely two homologous genes to SER/TMS1: Y57E12AL.1 and R11H6.2 in C. elegans; F6YGN3 and F7A8J9 in C. intestinalis and, H2YQ96 and H2Z2X8 in C. savignyi [35, 36]. In mammals, the SER protein family comprises five members (SER1-5). Previous topologies of the SER encoding genes have shown that the genes amplified by two duplication events, one before and one after the vertebrate-invertebrate split [37] which gave rise to two clusters of genes encoding SER1/2/3 and SER4/5 [22, 37]. In humans, there are five alternatively spliced isoforms of SER5, SER5-001, -004, -005, -008a, and -008b, which differ in the region coding for the C-terminal end and the transmembrane domains [38]. SER5 (here we discuss isoform 001) is found in the plasma membrane and is composed of 10 transmembrane domains, five extracellular loops, and four intracellular loops (ICL) [39]. SER5-001 is the longest isoform (461 amino acids) and encodes for an additional transmembrane domain with an essential role in restricting HIV-1 replication [38]. In addition, human SER1/3/4 [20, 21, 40, 41], coelacanth SER2 [35], rodent and lagomorph SER3/5 [42], and feline SER5 [33] do have antiviral activity.
Expression of SERINC proteins
In contrast to many other restriction factors, whose expression is regulated by interferons, it is interesting to note that the expression of SER5 is independent of interferon-induced signaling [20, 21, 43]. The immunodetection of SER proteins is still difficult due to the lack of specific antibodies to probe the endogenous level of these proteins [43]. SER5 is N-glycosylated at residue N294, which is critical for its steady-state level, and probably the non-glycosylated SER5 is degraded by proteasome as a quality control mechanism [44]. In mice, SER1 and SER3 are expressed in the central nervous system and SER3 mRNA is also found in the kidney and testis [45]. In humans, SER5 is ubiquitously expressed across tissues with a specific expression cluster in the liver (https://www.proteinatlas.org/ENSG00000164300-SERINC5/tissue), and the expression of SER5 is induced during the differentiation of monocytes to the myeloid lineage [43] as well as it is upregulated in oligodendrocytes during myelination [46]. In patients with HIV-1 compared to the uninfected individuals, the level of SER5 mRNA was found to be downregulated [47].
Cellular functions of SERINC5
Initially, SER proteins were characterized as carrier proteins, to incorporate serines into membrane lipids, such as phosphatidylserine and sphingolipids [48]. However, the knock-out of SER1 in some immune cells did not alter the serine lipid composition and function [49]. Moreover, SER5 did not alter the lipid composition in the HIV-1 particles [34]. Recently SER3 and SER5 were reported to have an intracellular role in innate sensing and signaling. Both SER3 and SER5 enhance the expression of type I IFN and the nuclear factor κB (NF- κB) contributing to the antiviral activity [50]. After infection with Sendai virus or stimulation with poly (I:C) or lipopolysaccharides (LPS), SER5 internalizes into the mitochondria membrane and associates with the mitochondrial membrane protein MAVS (mitochondrial antiviral signaling protein). As a result, MAVS aggregates and complexes with tumor necrosis factor receptor-associated factor 6 (TRAF6) through K63-mediated ubiquitylation. The complex MAVS-SER5-TRAF6 induces the phosphorylation of IRF3 and IκBα, which activates NF-κB (including the non-canonical pathway) and the expression of genes encoding type I IFNs and cytokines by activating the transcription factors interferon regulatory factor 3 (IRF3), IRF7, and NF-κB. Thus, SER5 functions here between MAVS and TBK1 [50] (Fig. 1). Interestingly, it was also reported that IFN-I stimulation enhances the surface levels of endogenous SER5 in a process that relocates the intracellular SER5 and stabilizes it on the plasma membrane. This was confirmed after observing the opposite effect by blocking the Jak/STAT signaling pathway with ruxolitinib [51] (Fig. 2). Those findings place SER5 as a factor involved in intracellular immune signaling pathways beyond its initially antiviral role associated with the plasma membrane.
Model of IFN-1 and inflammatory pathways induced by SER5 according Zeng et al. [50]. After the viral infection by Sendai virus or the stimulation with poly (I:C) or LPS, SER5 relocates from the plasma membrane to the mitochondrial membrane. SER5 associates with the transmembrane domains of the mitochondrial membrane protein MAVS and with the E3 ubiquitin ligase TRAF6. TRAF6 associates also with MAVS. This stable complex is the result of the MAVS aggregation and the polyubiquitination of TRAF6. This complex could have three effects (1) the activation of the canonical NF-κB pathway: after the phosphorylation of the IκBα repressor, the complex p50/p56 (p65 also named RelA) is transported to the nucleus and induces the expression of pro-inflammatory genes like ISG15, TNF-α IL-6; (2) the activation of the non-canonical NF-κB pathway; and (3) the phosphorylation of IRF3 and the translocation of phosphorylated dimers to the nucleus to induce the activation expression of IFN-I
Model of the effect of IFN-I on SER5. Type I interferon increases the amount of SER5 in the plasma membrane by reducing its recycling or endocytosis. This effect can be antagonized by ruxolitinib, which inhibits the Jak/STAT signalization pathway. This IFN effect is not dependent on the modulation of mRNA or protein levels of SER5 [51]
SERINC5 inhibits HIV-1
SER5 restricts HIV-1 replication when the virus does not express Nef (HIV-1 ∆Nef) [20, 21]. In contrast, human SER1 and SER2 display very limited or no activity against HIV-1 [40, 52]. However, SER2 from other species as coelacanth and Xenopus can inhibit HIV-1 [35]. These findings may indicate that the activity of SER2 was initially directed against retroviruses found in these species and it was lost during evolution. Further support for an ancient antiviral function of SER proteins comes from the finding that the envelope glycoprotein of the prototype foamy virus (a primate retrovirus that belongs to spumaviruses) counteracts the HIV-1 inhibition of coelacanth SER2 [35]. SER3 proteins from different species (human, primate, mouse, frog, and zebrafish) also moderately inhibit HIV-1 [21, 40, 42]. Despite the rapid turnover and low expression level of human SER4 (at least 250-fold lower than that of human SER5), this protein can also restrict HIV-1 as strongly as SER5 when the protein level is restored by transfecting 250-fold more SER4 than SER5 plasmids [40, 41].
In general, the antiviral activity of SERs is exerted on HIV-1 ∆ Nef viruses, only when SER proteins are packaged into viral particles [20, 21]. Various SER proteins have different abilities to incorporate into the budding virus membrane, where, SER5 is very efficiently associated in its glycosylated form [40, 44]. SER5 is localized in the detergent-resistant domains in the plasma membrane, where the budding of HIV takes place [40]. This indicates a passive mechanism of the SER5 association with the virus [22].
The restriction activity of SER occurs predominantly during the entry of the virus at the stage of fusion of viral and plasma membranes. Exactly how SER proteins block HIV infection of target cells is not fully understood, however, some hypotheses have been postulated [53,54,55,56]. It is possible that the infection of target cells is reduced by virion incorporated SER5 due to an alteration of the formation and/or enlargement of the fusion pore affecting the envelope-mediated fusion process [21, 57, 58]. In addition, it was suggested that SER5 proteins alter the conformation of the viral envelope and prevent the entry of the virus into the host cell [39, 52]. However, the fusion process between the cell and the HIV-1 WT or ∆ Nef has been found to have the same fusion capabilities independently of the SER5 expression status of the producer cells and thus, the inhibitory effect of SER5 may reduce further steps during the viral entry [51]. Regardless of Nef expression, several reports demonstrate that the antiviral activity of SER5 depends on an ill-defined sensitivity of viral Env proteins [58,59,60]. The HIV-1 trimeric envelope spike is formed by the Env surface subunit (gp120) with its five variable regions (V1 to V5) and the transmembrane subunit (gp41) with its cytoplasmic tail. The open conformation of the envelope protein seems to be essential for the SER5 restriction [59, 61] and SER5 appears to inactivate sensitive but not resistant Env glycoproteins [58, 59, 62, 63] that include some wildtype HIV-1 envelopes and variants with truncations of the cytoplasmic tail [64, 65].
Whether a direct interaction between SER and HIV-1 envelope is relevant is unclear [64]. SER5 could interact with Env in the plasma membrane and the extent of the binding correlates with the sensitivity of Env to SER5 [59]. It was suggested that Env-SER5 interaction is mediated by the extracellular loop 3 and 5 of SER5 which have a spacing that matches that between the MPER regions (membrane-proximal external region) within the HIV-1 Env trimer [39]. In contrast, it was also discussed that SER5 disrupts Env clusters without a direct binding and no co-distribution of Env-SER in virions was detected [66].
Envelopes from other viruses have been also demonstrated to be sensitive, such as amphotropic MLV and influenza A virus (IAV). In contrast, diverse viral surface glycoproteins e.g. derived from vesicular stomatitis virus (VSV-G), Ebola virus (EBOV), equine infectious anemia virus (EIAV), avian leukosis virus A (ALV-A), and feline leukemia virus subgroup B (FeLV-B) and the ecotropic MLV are resistant to the antiviral SER5 activity [31, 33, 65, 67]. In principle, the reason why some envelopes are sensitive to SER5 or insensitive is not clear [65]. Initially, it was postulated that envelopes with a low pH-dependent endocytic entry pathway were SER5 resistant [20, 21], however, this was not demonstrated with other glycoproteins [65]. In SER5 resistant HIV-1 envelopes, the resistance correlates to a closed conformation of Env where the variable loops V1–V3 interact with each other [52, 59, 66, 68]. Additionally, the viral core also seems to be an important factor that determines the sensitivity of some envelope proteins. For instance, the mason-pfizer monkey virus (M-PMV) glycoprotein is sensitive to SER5 when it is used to pseudotype MLV or M-PMV cores, but shows resistance when it is used on HIV-1 cores [65]. With other viral glycoproteins similar observations were made, where the SER5 resistance was viral core-dependent [65].
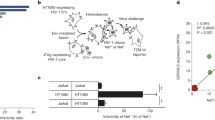
Another interesting antiviral function of SER5 that is also counteracted by Nef is an enhanced recognition of such HIV-1 particles by primary myeloid cells, which leads to the production of proinflammatory cytokines like granulocyte–macrophage colony-stimulating factor (GM-CSF), GRO alpha, IL-6, IL-8, and TNF-α. The mechanism of this virus recognition is not completely clear and may involve sensing of viral RNAs [69].
Nef counteracts SERINC5
HIV-1 Nef prevents the incorporation of SER5 into budding virions by inducing the down-regulation of SER5 from the plasma membrane [22]. Nef is an accessory protein of 27 to 32 kDa, encoded only by primate lentiviruses (HIV-1, HIV-2, and SIV) [70]. Nef was found to reduce the expression of several surface proteins in the infected cells, modulate signaling pathways in T cells and increase the infectivity of viral particles [71]. While most studies on ectopic expressed SER5 concluded that Nef is inducing cellular degradation of SER5, a study on the mRNA and protein levels of endogenous SER5 observed that Nef modulates only the SER5 surface localization without altering the steady-state levels of SER5 [51] (Fig. 3). Uniformly in all studies, HIV-1 Nef reduced the amount of SER5 on the cell surface by 70–90% [33, 51, 54, 72]. Myristoylated Nef associates with the plasma membrane as homodimers [54] where it interacts with SER5. In a second step, Cyclin K (CCNK) and cyclin-dependent-kinase 13 (CDK13) are recruited and the serine 360 in SER5 on the intracellular loop 4 (ICL4) is phosphorylated. This phosphorylation induces a conformational change in ICL4 that facilitates the interaction of SER5 with Nef by increasing the accessibility of residues L350/I352 to Nef which potentiates the binding [55]. When the conformational change is maintained, an unknown endocytic motif of ICL4 is exposed and the adaptor protein 2 (AP-2) is recruited [55]. The Nef C-terminal dileucine motif ExxxLL interacts with the AP-2 and the Nef N-terminal (amino acids 32–39) bind with SER5 [20, 21, 54, 73]. Consequently, Nef induces polyubiquitination of SER5 via a K48- and K63-linkage as a prerequisite for lysosomal degradation [74, 75]. This assembly, SER5, Nef, cyclin K/CDK13, and AP-2, causes endocytosis and lysosome-mediated degradation of SER5 from the membrane [55, 75, 76]. SER5 is relocated especially to Rab7 + late endosomes [20, 54, 74, 75], however, SER5 has been also found in Rab5 + early and Rab11 + recycling endosomes [75].
HIV-1 counteracts the antiviral activity of SER5 by its Nef protein.. When Nef is absent, the surface expression of SER5 in the plasma membrane is not altered and SER5 is incorporated into the newly formed viral particle. The infectivity of the virus will depend on its envelope. Virions with a SER5-sensitive envelope have an impaired infectivity while a resistant envelope allows the infection of the target cell even in the presence of SER5. If Nef is present, SER5 associates with the AP-2 complex and Nef with low affinity. Afterwards, Cyclin K/CDK13 is recruited and phosphorylates SER5 in the intracellular loop 4 (ICL4). In consequence, the AP-2 complex is recruited which induces the endogenization of SER5 to Rab7 + late endosomes, SER5 is downregulated from the plasma membrane and thus is not incorporated into the budding virion keeping its infectivity capacity [55]
The activity of Nef proteins from the HIV-1 group M subtypes A, B, C, and D was found to be not identical and especially the subtype B Nef clones showed the highest function in SER5 internalization, followed by the Nef from subtype D, while Nef proteins of subtypes A and C displayed the lowest capacity to induce SER5 internalization [77]. The ability of Nef to counteract SER3 and SER5 varies among patients living with HIV-1. While most of the Nef proteins were capable of strongly antagonizing SER3 and SER5, some mutations were associated with a better counteraction of SER3 (N51T, H116N, and 188S) and SER5 (S163C) by increasing the HIV-1 infectivity [78]. The counteraction of SER5 by Nef was also associated with a lower viremia in HIV-1 infected patients [79]. Some natural polymorphisms in HIV-1 Nef from elite controllers (e.g., K94E and H116N) were described to display lower infectivity and replication capacity in the presence of SER5 [80]. Other Nef polymorphisms, here in the dileucine motif, were found to be associated with the CD4 and SER5 downregulation. This finding supports a relation between Nef downregulation of SER5 and the rate of plasma CD4+ T cell decline [81]. However, a different study with the Nef of the simian immunodeficiency virus (SIV) showed that the activity against SER5 can be separated from the general downregulating functions of Nef. After mutating some important residues for counteracting SER5, Nef still reduced the amount of CD3, CD4, and MHC-I from the plasma membrane [82].
Concluding remarks
In summary, the cellular functions of the SER proteins appear to keep some secrets, and solving these puzzles may be the solution to understand their antiviral activity. Different retroviruses evolved independent pathways to remove SER5 from the plasma membrane to prevent SER5 incorporation. HIV Nef, EIAV S2, and MLV/FeLV glycoGag are unrelated retroviral proteins that have in common that evolution independently directed them to SER5 counteraction. In addition, viral envelopes have been shown to have an important role in conferring resistance against SER5 inhibition. It is unclear why retroviruses use two strategies to escape SER5, but possibly suggesting a strong in vivo restriction by SER5 [67, 83].
References
Boso G, Kozak CA (2020) Retroviral restriction factors and their viral targets: restriction strategies and evolutionary adaptations. Microorganisms 8:1–34. https://doi.org/10.3390/microorganisms8121965
Colomer-Lluch M, Ruiz A, Moris A, Prado JG (2018) Restriction factors: from intrinsic viral restriction to shaping cellular immunity against HIV-1. Front Immunol 9:2876. https://doi.org/10.3389/fimmu.2018.02876
Aydin H, Taylor MW, Lee JE (2014) Structure-guided analysis of the human APOBEC3-HIV restrictome. Structure 22:668–684. https://doi.org/10.1016/j.str.2014.02.011
Desimmie BA, Delviks-Frankenberrry KA, Burdick RC et al (2014) Multiple APOBEC3 restriction factors for HIV-1 and one vif to rule them all. J Mol Biol 426:1220–1245. https://doi.org/10.1016/j.jmb.2013.10.033
Arias JF, Koyama T, Kinomoto M, Tokunaga K (2012) Retroelements versus APOBEC3 family members: no great escape from the magnificent seven. Front Microbiol 3:1–12. https://doi.org/10.3389/fmicb.2012.00275
Delviks-frankenberry KA, Desimmie BA, Pathak VK (2020) Structural insights into APOBEC3-mediated. Viruses. https://doi.org/10.3390/v12060587
Arias JF, Iwabu Y, Tokunaga K (2011) Structural basis for the antiviral activity of BST-2/tetherin and its viral antagonism. Front Microbiol 2:1–9. https://doi.org/10.3389/fmicb.2011.00250
Evans DT, Serra-Moreno R, Singh RK, Guatelli JC (2011) BST-2/tetherin: a new component of the innate immune response to enveloped viruses. Trends Microbiol 18:388–396. https://doi.org/10.1016/j.tim.2010.06.010
Li SX, Barretta BS, Guoa K, Santiago ML (2016) Tetherin/BST-2: restriction factor or immunomodulator? Curr HIV Res 14:235–246. https://doi.org/10.2174/1570162x14999160224102752
Nomaguchi M, Fujita M, Adachi A (2008) Role of HIV-1 Vpu protein for virus spread and pathogenesis. Microbes Infect 10:960–967. https://doi.org/10.1016/j.micinf.2008.07.006
Wu L (2013) Cellular and biochemical mechanisms of the retroviral restriction factor SAMHD1. ISRN Biochem 728392:1–11. https://doi.org/10.1155/2013/728392
Goldstone DC, Ennis-Adeniran V, Hedden JJ et al (2011) HIV-1 restriction factor SAMHD1 is a deoxynucleoside triphosphate triphosphohydrolase. Nature 480:379–382. https://doi.org/10.1038/nature10623
Deutschmann J, Gramberg T (2021) SAMHD1 … and viral ways around it. Viruses 13:1–16. https://doi.org/10.3390/v13030395
Murrell B, Vollbrecht T, Guatelli J, Wertheim JO (2016) The evolutionary histories of antiretroviral proteins SERINC3 and SERINC5 do not support an evolutionary arms race in primates. J Virol 90:8085–8089. https://doi.org/10.1128/jvi.00972-16
Cloherty APM, Rader AG, Compeer B, Ribeiro CMS (2021) Human TRIM5α: autophagy connects cell-intrinsic HIV-1 restriction and innate immune sensor functioning. Viruses 13:1–32. https://doi.org/10.3390/v13020320
Betancor G, Dicks MDJ, Jimenez-Guardeño JM et al (2019) The GTPase domain of MX2 interacts with the HIV-1 capsid, enabling its short isoform to moderate antiviral restriction. Cell Rep 29:1923-1933.e3. https://doi.org/10.1016/j.celrep.2019.10.009
Staeheli P, Haller O (2018) Human MX2/MxB: a potent interferon-induced postentry inhibitor of herpesviruses and HIV-1. J Virol. https://doi.org/10.1128/JVI.00709-18
Fackler OT, Keppler OT (2013) MxB/Mx2: the latest piece in HIV’s interferon puzzle. EMBO Rep 14:1028–1029. https://doi.org/10.1038/embor.2013.172
Haller O, Staeheli P, Schwemmle M, Kochs G (2015) Mx GTPases: dynamin-like antiviral machines of innate immunity. Trends Microbiol 23:154–163. https://doi.org/10.1016/j.tim.2014.12.003
Rosa A, Chande A, Ziglio S et al (2015) HIV-1 Nef promotes infection by excluding SERINC5 from virion incorporation. Nature 526:212–217. https://doi.org/10.1038/nature15399
Usami Y, Wu Y, Göttlinge HG (2015) SERINC3 and SERINC5 restrict HIV-1 infectivity and are counteracted by Nef. Nature 526:218–223. https://doi.org/10.1016/j.physbeh.2017.03.040
Firrito C, Bertelli C, Vanzo T et al (2018) SERINC5 as a new restriction factor for human immunodeficiency virus and murine leukemia virus. Annu Rev Virol 5:323–340. https://doi.org/10.1146/annurev-virology-092917-043308
Lee WYJ, Fu RM, Liang C, Sloan RD (2018) IFITM proteins inhibit HIV-1 protein synthesis. Sci Rep 8:1–15. https://doi.org/10.1038/s41598-018-32785-5
Foster TL, Wilson H, Iyer SS et al (2016) Resistance of transmitted founder HIV-1 to IFITM-mediated restriction. Cell Host Microbe 20:429–442. https://doi.org/10.1016/j.chom.2016.08.006
Perreira JM, Chin CR, Feeley EM, Brass AL (2013) IFITMs restrict the replication of multiple pathogenic viruses. J Mol Biol 425:4937–4955. https://doi.org/10.1016/j.jmb.2013.09.024
Li M, Kao E, Gao X et al (2012) Codon-usage-based inhibition of HIV protein synthesis by human schlafen 11. Nature 491:125–128. https://doi.org/10.1038/nature11433
Zhang Y, Lu J, Liu X (2018) MARCH2 is upregulated in HIV-1 infection and inhibits HIV-1 production through envelope protein translocation or degradation. Virology 518:293–300. https://doi.org/10.1016/j.virol.2018.02.003
Zhang Y, Tada T, Ozono S et al (2020) MARCH8 inhibits viral infection by two different mechanisms. Elife 9:1–14. https://doi.org/10.7554/ELIFE.57763
Münch J, Rajan D, Schindler M et al (2007) Nef-mediated enhancement of virion infectivity and stimulation of viral replication are fundamental properties of primate lentiviruses. J Virol 81:13852–13864. https://doi.org/10.1128/jvi.00904-07
Pizzato M (2010) MLV glycosylated-gag is an infectivity factor that rescues Nef-deficient HIV-1. Proc Natl Acad Sci USA 107:9364–9369. https://doi.org/10.1073/pnas.1001554107
Chande A, Cuccurullo EC, Rosa A et al (2016) S2 from equine infectious anemia virus is an infectivity factor which counteracts the retroviral inhibitors SERINC5 and SERINC3. Proc Natl Acad Sci USA 113:13197–13202. https://doi.org/10.1073/pnas.1612044113
Usami Y, Popov S, Gottlinger HG (2014) The Nef-Like effect of murine leukemia virus glycosylated gag on HIV-1 infectivity is mediated by its cytoplasmic domain and depends on the AP-2 adaptor complex. J Virol 88:3443–3454. https://doi.org/10.1128/jvi.01933-13
Cano-Ortiz L, Gu Q, de Sousa-Pereira P et al (2021) Feline leukemia virus-B envelope together with its GlycoGag and human immunodeficiency virus-1 Nef mediate resistance to feline SERINC5. J Mol Biol. https://doi.org/10.1016/j.jmb.2021.167421
Trautz B, Wiedemann H, Lüchtenborg C et al (2017) The host-cell restriction factor SERINC5 restricts HIV-1 infectivity without altering the lipid composition and organization of viral particles. J Biol Chem 292:13702–13713. https://doi.org/10.1074/jbc.M117.797332
Ramdas P, Bhardwaj V, Singh A et al (2021) Coelacanth SERINC2 inhibits HIV-1 infectivity and is counteracted by envelope glycoprotein from foamy virus. J Virol 95:e00229-e321. https://doi.org/10.1128/jvi.00229-21
Howe KL, Achuthan P, Allen J et al (2020) Ensembl 2021. Nucleic Acids Res 49:539–547. https://doi.org/10.1093/nar/gkaa942
Haq F, Saeed U, Khalid R et al (2019) Phylogenetic analyses of human 1/2/8/20 paralogons suggest segmental duplications during animal evolution. 3 Biotech 9:1–8. https://doi.org/10.1007/s13205-019-1768-7
Zhang X, Zhou T, Yang J et al (2017) Identification of SERINC5-001 as the predominant spliced isoform for HIV-1 restriction. J Virol. https://doi.org/10.1128/jvi.00137-17
Pye VE, Rosa A, Bertelli C et al (2020) A bipartite structural organization defines the SERINC family of HIV-1 restriction factors. Nat Struct Mol Biol 27:78–83. https://doi.org/10.1038/s41594-019-0357-0
Schulte B, Selyutina A, Opp S et al (2017) Localization to detergent-resistant membranes and HIV-1 core entry inhibition correlate with HIV-1 restriction by SERINC. Physiol Behav 176:139–148. https://doi.org/10.1016/j.virol.2017.12.005
Qiu X, Eke IE, Johnson SF et al (2020) Proteasomal degradation of human SERINC4: a potent host antiHIV-1 factor that is antagonized by nef. Curr Res Virol Sci. https://doi.org/10.1016/j.crviro.2020.100002
de Sousa-Pereira P, Abrantes J, Bauernfried S et al (2019) The antiviral activity of rodent and lagomorph SERINC3 and SERINC5 is counteracted by known viral antagonists. J Gen Virol 100:278–288. https://doi.org/10.1099/jgv.0.001201
Zutz A, Schölz C, Schneider S et al (2020) SERINC5 is an unconventional HIV restriction factor that is upregulated during myeloid cell differentiation. J Innate Immun 12:399–409. https://doi.org/10.1159/000504888
Sharma S, Lewinski MK, Guatellia J (2018) An N-glycosylated form of SERINC5 is specifically incorporated into HIV-1 virions. J Virol 92:1–15. https://doi.org/10.1128/JVI.00753-18
Grossman TR, Luque JM, Nelson N (2000) Identification of a ubiquitous family of membrane proteins and their expression in mouse brain. J Exp Biol 203:447–457. https://doi.org/10.1242/jeb.203.3.447
Krueger WHH, Gonye GE, Madison DL et al (1997) TPO1, a member of a novel protein family, is developmentally regulated in cultured oligodendrocytes. J Neurochem 69:1343–1355. https://doi.org/10.1046/j.1471-4159.1997.69041343.x
Hernández-López EG, González-Enríquez GV, Torres-Mendoza BM et al (2021) Downregulation of SERINC5 expression in buffy coats of HIV-1-infected patients with detectable or undetectable viral load. Mol Biol Rep 48:4247–4252. https://doi.org/10.1007/s11033-021-06438-2
Inuzuka M, Hayakawa M, Ingi T (2005) Serine, an activity-regulated protein family, incorporates serine into membrane lipid synthesis. J Biol Chem 280:35776–35783. https://doi.org/10.1074/jbc.M505712200
Chu EPF, Elso CM, Pollock AH et al (2017) Disruption of Serinc1, which facilitates serine-derived lipid synthesis, fails to alter macrophage function, lymphocyte proliferation or autoimmune disease susceptibility. Mol Immunol 82:19–33. https://doi.org/10.1016/j.molimm.2016.12.007
Zeng C, Waheed AA, Li T et al (2021) SERINC proteins potentiate antiviral type I IFN production and proinflammatory signaling pathways. Sci Signal 14:1–14. https://doi.org/10.1126/scisignal.abc7611
Passos V, Zillinger T, Casartelli N et al (2019) Characterization of endogenous SERINC5 protein as anti-HIV-1 factor. J Virol 93:e01221-e1319. https://doi.org/10.1128/JVI.01221-19
Beitari S, Ding S, Pan Q, Finzi A, Lianga C (2017) Effect of HIV-1 Env on SERINC5 antagonism. J Virol 91:1–13. https://doi.org/10.1128/JVI.02214-16
Gonzalez-Enriquez GV, Escoto-Delgadillo M, Vazquez-Valls E, Torres-Mendoza BM (2017) SERINC as a restriction factor to inhibit viral infectivity and the interaction with HIV. J Immunol Res. https://doi.org/10.1155/2017/1548905
Staudt RP, Smithgall TE (2020) Nef homodimers down-regulate SERINC5 by AP-2-mediated endocytosis to promote HIV-1 infectivity. J Biol Chem 295:15540–15552. https://doi.org/10.1074/jbc.RA120.014668
Chai Q, Li S, Collins MK et al (2021) HIV-1 Nef interacts with the cyclin K/CDK13 complex to antagonize SERINC5 for optimal viral infectivity. Cell Rep. https://doi.org/10.1016/j.celrep.2021.109514
Staudt RP, Alvarado JJ, Emert-Sedlak LA et al (2020) Structure, function, and inhibitor targeting of HIV-1 Nef-effector kinase complexes. J Biol Chem 295:15158–15171. https://doi.org/10.1074/jbc.REV120.012317
Cohen FS, Melikyan GB (2004) The energetics of membrane fusion from binding, through hemifusion, pore formation, and pore enlargement. J Membr Biol 199:1–14. https://doi.org/10.1007/s00232-004-0669-8
Sood C, Marin M, Chande A et al (2017) SERINC5 protein inhibits HIV-1 fusion pore formation by promoting functional inactivation of envelope glycoproteins. J Biol Chem 292:6014–6026. https://doi.org/10.1074/jbc.M117.777714
Zhang X, Shi J, Qiu X et al (2019) CD4 expression and Env conformation are critical for HIV-1 restriction by SERINC5. J Virol. https://doi.org/10.1128/jvi.00544-19
Liu Y, Wang H, Zhang J et al (2020) SERINC5 inhibits the secretion of complete and genome-free hepatitis B virions through interfering with the glycosylation of the HBV envelope. Front Microbiol 11:1–15. https://doi.org/10.3389/fmicb.2020.00697
Featherstone A, Aiken C (2020) SERINC5 inhibits HIV-1 infectivity by altering the conformation of gp120 on HIV-1 particles. J Virol 94:e00594-e620
Ahmad I, Li S, Li R et al (2019) The retroviral accessory proteins S2, Nef, and glycoMA use similar mechanisms for antagonizing the host restriction factor SERINC5. J Biol Chem 294:7013–7024. https://doi.org/10.1074/jbc.RA119.007662
Seaman MS, Janes H, Hawkins N et al (2010) Tiered categorization of a diverse panel of HIV-1 Env pseudoviruses for assessment of neutralizing antibodies. J Virol 84:1439–1452. https://doi.org/10.1128/jvi.02108-09
Haider T, Snetkov X, Jolly C (2021) HIV envelope tail truncation confers resistance to SERINC5 restriction. Proc Natl Acad Sci USA 118:e2101450118. https://doi.org/10.1073/pnas.2101450118
Diehl WE, Guney MH, Vanzo T et al (2021) Influence of different glycoproteins and of the virion core on SERINC5 antiviral activity. Viruses 13:1279. https://doi.org/10.3390/v13071279
Chen YC, Sood C, Marin M et al (2020) Super-resolution fluorescence imaging reveals that serine incorporator protein 5 inhibits human immunodeficiency virus fusion by disrupting envelope glycoprotein clusters. ACS Nano 14:10929–10943. https://doi.org/10.1021/acsnano.0c02699
Timilsina U, Umthong S, Lynch B, Stablewski A (2020) Serinc5 potently restricts retrovirus infection in vivo. MBio 11:1–18. https://doi.org/10.1128/mBio.00588-20
Usami Y, Göttlinger H (2013) HIV-1 nef responsiveness is determined by env variable regions involved in trimer association and correlates with neutralization sensitivity. Cell Rep 5:802–812. https://doi.org/10.1016/j.celrep.2013.09.028
Pierini V, Gallucci L, Stürzel CM et al (2021) SERINC5 can enhance proinflammatory cytokine production by primary human myeloid cells in response to challenge with HIV-1 particles. J Virol 95:1–19. https://doi.org/10.1128/JVI.02372-20
Hirsch VM, Dapolito G, Goeken R, Campbell BJ (1995) Phylogeny and natural history of the primate lentiviruses, SIV and HIV. Curr Opin Genet Dev 5:798–806. https://doi.org/10.1016/0959-437X(95)80014-V
Basmaciogullari S, Pizzato M (2014) The activity of Nef on HIV-1 infectivity. Front Microbiol 5:1–12. https://doi.org/10.3389/fmicb.2014.00232
Trautz B, Pierini V, Wombacher R et al (2016) The antagonism of HIV-1 Nef to SERINC5 particle infectivity restriction involves the counteraction of virion-associated pools of the restriction factor. J Virol 90:10915–10927. https://doi.org/10.1128/JVI.01246-16
Ananth S, Morath K, Trautz B et al (2019) Multifunctional roles of the N-terminal region of HIV-1 SF2 Nef are mediated by three independent protein interaction sites. J Virol. https://doi.org/10.1128/jvi.01398-19
Li S, Ahmad I, Shi J et al (2018) Murine leukemia virus glycosylated Gag reduces murine SERINC5 protein expression at steady-state levels via the endosome/lysosome pathway to counteract SERINC5 antiretroviral activity. J Virol 93:1–15. https://doi.org/10.1128/jvi.01651-18
Shi J, Xiong R, Zhou T et al (2018) HIV-1 Nef antagonizes SERINC5 restriction by downregulation of SERINC5 via the endosome/lysosome system. J Virol 92:1–16. https://doi.org/10.1128/jvi.00196-18
daSilva LLP, Sougrat R, Burgos PV et al (2009) Human immunodeficiency virus type 1 Nef protein targets CD4 to the multivesicular body pathway. J Virol 83:6578–6590. https://doi.org/10.1128/jvi.00548-09
Jin SW, Mwimanzi FM, Mann JK et al (2020) Variation in HIV-1 nef function within and among viral subtypes reveals genetically separable antagonism of serinc3 and serinc5. PLoS Pathog 16:1–22. https://doi.org/10.1371/journal.ppat.1008813
Kruize Z, van Nuenen AC, van Wijk SW et al (2021) Nef obtained from individuals with hiv-1 vary in their ability to antagonize serinc3-and serinc5-mediated hiv-1 restriction. Viruses 13:1–16. https://doi.org/10.3390/v13030423
Toyoda M, Kamori D, Tan TS et al (2020) Impaired ability of Nef to counteract SERINC5 is associated with reduced plasma viremia in HIV-infected individuals. Sci Rep 10:1–12. https://doi.org/10.1038/s41598-020-76375-w
Jin SW, Alsahafi N, Kuang XT et al (2019) Natural HIV-1 Nef polymorphisms impair SERINC5 downregulation activity. Cell Rep 29:1449-1457.e5. https://doi.org/10.1016/j.celrep.2019.10.007
Mumby MJ, Johnson AL, Trothen SM et al (2021) An amino acid polymorphism within the HIV-1 Nef dileucine motif functionally uncouples cell surface CD4 and SERINC5 downregulation. J Virol 95:e00588-e621. https://doi.org/10.1128/JVI.00588-21
Janaka SK, Palumbo AV, Tavakoli-Tameh A, Evans DT (2021) Selective disruption of SERINC5 antagonism by Nef impairs simian immunodeficiency virus replication in primary CD4 + T cells. J Virol. https://doi.org/10.1128/jvi.01911-20
Heigele A, Kmiec D, Regensburger K et al (2016) The potency of Nef-mediated SERINC5 antagonism correlates with the prevalence of primate lentiviruses in the wild. Cell Host Microbe 20:381–391. https://doi.org/10.1016/j.chom.2016.08.004
Acknowledgements
We thank Ananda Ayyappan Jaguva Vasudevan for critical reading of the manuscript. We thank Wioletta Hörschken for excellent technical assistance. LCO is supported by DAAD scholarship, CM is supported by the Heinz-Ansmann foundation for AIDS research. We thank Dr. Friedhelm Haak (Hannover) for his generous support.
Funding
Open Access funding enabled and organized by Projekt DEAL. Funding was provided by Deutscher Akademischer Austauschdienst.
Author information
Authors and Affiliations
Contributions
LCO: literature review, writing, original draft preparation, figures. TL: supervision. CM: Conceptualization, writing, review and editing, supervision.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Edited by: Hanna-Mari Baldauf.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is published as part of the Special Issue on Immunobiology of Viral Infections.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cano-Ortiz, L., Luedde, T. & Münk, C. HIV-1 restriction by SERINC5. Med Microbiol Immunol 212, 133–140 (2023). https://doi.org/10.1007/s00430-022-00732-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00430-022-00732-x