Abstract
Purpose
Loss of expression of DLG2 has been identified in a number of cancers to contribute to the disease by resulting in increased tumor cell proliferation and poor survival. In light of the previous evidence that DLG2 alters the cell cycle and affects proliferation, combined with indications that DLG2 is involved in NLRP3 inflammasome axis we speculated that DLG2 has an immune function. So far, there is no data that clearly elucidates this role, and this study was designed to investigate DLG2 in inflammatory colon disease and in colon cancer as well as its impact on inflammasome induction.
Methods
The DLG2 expression levels were established in publicly available inflammation, colon cancer and mouse model datasets. The overexpression and silencing of DLG2 in colon cancer cells were used to determine the effect of DLG2 expression on the activation of the inflammasome and subsequent cytokine release.
Results
The expression of DLG2 is repressed in inflammatory colon diseases IBD and Ulcerative colitis as well as colorectal cancer tissue compared to healthy individuals. We subsequently show that induction with inflammatory agents in cell and animal models results in a biphasic alteration of DLG2 with an initial increase followed by an ensuing decrease. DLG2 overexpression leads to a significant increase in expression of IL1B, IκBζ and BAX, components that result in inflammasome formation. DLG2 silencing in THP1 cells resulted in increased release of IL-6 into the microenvironment which once used to treat bystander COLO205 cells resulted in an increase in STAT3 phosphorylation and an increase proliferating cells and more cells in the G2/M phase. Restoration of DLG2 to the colon resulted in reduced AKT and S6 signaling.
Conclusion
DLG2 expression is altered in response to inflammation in the gut as well as colon cancer, resulting in altered ability to form inflammasomes.
Trial registration
NCT03072641.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colon cancer is one of the leading causes of cancer related deaths worldwide. In the western world, poor diets are increasing the incidences of obesity and altering the diversity of commensal bacteria (Singh et al. 2017). The increasing rates of obesity within global populations are increasing the total number of individuals at an elevated risk for developing colorectal cancer (Lund et al. 2011). Another factor leading to the increased incidences of colorectal cancer in the general population is advancing age (Kolligs 2016). Additionally, inflammatory bowel diseases such as Crohn’s disease and ulcerative colitis are also known factors that increase the risk of colorectal cancer development and subsequent mortality (Lund et al. 2011). It is important to note that there are generally considered to be two major pathways for development of colon cancer development, spontaneous (SCC) (Shi et al. 2020) also referred to as adenoma–carcinoma-sequence (Ozawa et al. 2021) and Colitis associated colon cancer (CAC) (Wang et al. 2020; Zhang et al. 2020a, b) referred to as inflammation–dysplasia–carcinoma-sequence (Ozawa et al. 2021). SCC results from otherwise healthy mucosa transforming into precancerous polyps, then progressing to adenocarcinomas and then cancerous lesions with a number of specific genetic alterations that occur at each transition step (Jones et al. 2008). One of the first alterations is the loss or mutation of APC followed by KRAS mutation and then loss of p53, with IL-27 loss compounding the effect of p53 loss in other cancer forms (Dibra et al. 2016a, b). With CAC the order of alterations differs, resulting in flatter lesions with varying degrees of dysplasia. In CAC the loss of p53 occurs earlier in this process and APC later (Jones et al. 2008). Common to both pathways is the activation of COX-2 in response to inflammation (Sharma et al. 2001; Chu et al. 2004; Janakiram and Rao 2009). Underpinning the transitions is a constitutive activation of NFκβ signaling which in this context results in tumor cell survival, proliferation and cell cycle progression (Curtin et al. 2010; Shi et al. 2019).
The colon is home to the largest population of microbes in the human body and encounters the highest concentration of pathogens, representing a large risk of infection if there is a disturbance in the microbiota (Jahani-Sherafat et al. 2018). Therefore, the colon has a large number of immune cells present. The innate immune system is triggered through pattern recognition receptors (PRRs), which include Toll like receptors (TLR) and retinoic acid inducible gene-I (RIG-I) (Ostvik et al. 2020). PRRs, TLRs and RIGs activate the inflammatory pathways in response to Pathogen Associated Molecular Patterns (PAMPs) (Santiago et al. 2020) or Damage Associated Molecular Patterns (DAMPs) (Matzinger 1994; Zhang et al. 2010). The upregulation of NFκβ in response to a DAMP or PAMP initiates the upregulation of proinflammatory factors required for priming of the NLRP3 inflammasome, by inducing the upregulation of IL-1β (Bank et al. 2014; Hai Ping et al. 2016). The activation of the inflammasome requires a second signal such as ATP (Jang et al. 2021). Mutations in the components of the inflammasomes often result in an increased susceptibility to cancer. The Q705K SNP in NLRP3 is associated with higher inflammasome activation (Verma et al. 2012) and poor patient survival in advanced stage colorectal cancer (Ungerback et al. 2012). It has been proposed that this SNP combined with external stimuli can result in increased IL-1β and IL-18 production (Verma et al. 2012), and IL-1β has been shown to downregulate pTEN by activation of NFκβ in colon cancer (Hai Ping et al. 2016), whilst IL-18 contributes to inflammasome mediated protection against tumorigenesis in colitis patients (Zaki et al. 2010). IL-18 has also been shown to be activated downstream of the retinoic acid receptor (RAR) as well as promote T- Helper 17 (Th 17) responses. The role of NLRP3 in colon cancer has been controversial with studies showing that high expression of NLRP3 drives epithelial-mesenchymal transition (EMT) (Shao et al. 2020) and results in poor survival (Shi et al. 2021). However, in direct contradiction another study has shown that NLRP3 mediates inhibition of metastatic growth (Dupaul-Chicoine et al. 2015). It is however important to note that the expression of NLRP3 and activation of the inflammasome differs with the type of cell mediating the inflammasome effect (Jang et al. 2021). Previous studies have shown that inflammasome activation requires NFKBIZ (IκBζ), a lesser known nuclear Factor kappa B (NFκβ) inhibitor, that regulates transcription of NFκβ targets by binding p50 or p52 subunits of NFκβ (Yamazaki et al. 2001). Furthermore, it can directly bind and negatively regulate STAT3 and induce apoptosis (Willems et al. 2016). Loss or altered NFKBIZ results in chronic inflammation by inhibiting the production of IL-10 (Horber et al. 2016) as well as diminished inflammasome priming (Kim et al. 2020). Chronic and/or overactivation of the inflammasome and resulting increase in IL-6 expression result in poor outcomes in colon cancer patients by activating STAT3 (Corvinus et al. 2005; Slattery et al. 2007; Xiong et al. 2008). When taken together this highlights the dual nature and the importance of concise control of the inflammasome and inflammatory responses in human colorectal cancers.
Recently, low DLG2 expression in osteosarcoma (Shao et al. 2019), ovarian cancer (Zhuang et al. 2019) and neuroblastoma (Keane et al. 2020, 2021; Siaw et al. 2020) has been identified as contributing to disease etiology, with low DLG2 expression in neuroblastoma showing increased cell proliferation and poor survival (Keane et al. 2020). Additionally, preliminary data show that DLG2 induces cell death by modulating BAX/BCL2 in response to DNA damage (Keane et al. 2022), colocalizing in the same pathways as the NLRP3 inflammasome. It has been shown that DLG2 can be induced by treatment with 13´cis retinoic acid (Siaw et al. 2020) with retinoic acid response element 1 (RARRES1) directly regulating DLG2 (Sahab et al. 2010). Additionally, DLG2 induces p53 mediated cell death in response to UVC irradiation (Keane et al. 2022).
In light of the building body of evidence that DLG2 is an important tumor suppressor gene we here investigate DLG2 and NFKBIZ and their impact on the inflammasome induction in inflammatory colon disease and in colon cancer. We detail the importance of DLG2 in activating the inflammasome complex and the effect of inflammation on DLG2.
Methods
Gene expression analysis
Data for analyses and comparison of DLG2 expression between the different patient subgroups was imported from the R2 platform (http://r2.amc.nl). The six independent colon cohorts; (Galamb et al. 2008) (GSE4183), (Haberman et al. 2019) (GSE109142), (Vancamelbeke et al. 2017) (GSE75214), (Jiang et al. 2008) (GSE10950), (Sabates-Beliver et al. 2007) (GSE8671) and (Agesen et al. 2012) (GSE24551). The inflammation time series mouse models, DSS (Fang et al. 2012) (GSE22307) and Colon T cell transfer (Fang et al. 2011) (GSE27302). The microarray data was downloaded as the centered log2 fold change.
Cell Lines and cell culture
Human colon adenocarcinoma cell lines SW480 and COLO205 were obtained from ATCC Cell Line Collection and THP 1-ASC-GFP monocytes were obtained from Invovogen. The SW480 and COLO205 cell lines were maintained in RPMI 1640 (ThermoFisher Scientific) supplemented with 10% FBS, 1% L-Glutamine (ThermoFisher Scientific), 10 mM HEPES solution (ThermoFisher Scientific) and 1 mM sodium pyruvate (ThermoFisher Scientific). THP 1-ASC-GFP cells were cultured in RPMI-1640 with L-glutamine (Merck), 10% heat inactivated premium grade FBS (Biowest), 10 mM HEPES, 1 mM sodium pyruvate (Merck), 0,45% glucose (Merck) and 100U/ml penicillin–streptomycin (Merck) at 37 °C and 5% CO2. Zeocin (200 µg/ml) (Invivogen) was added to the culture medium as per the manufacturer’s instructions. Cell density was maintained between 5 × 105 and 1.5 × 106 cells/ml and cells were used up to passage number ten. Differentiation was conducted with 100 ng/ml PMA (Merck) for 72 h followed by 72 h of rest in fresh media. Priming of undifferentiated or differentiated cells was conducted with 500 ng/ml ultrapure LPS (Invivogen) for up to 24 h as indicated. Activation of inflammasome complex formation was conducted with 5 mM ATP (Merck) for 30 min.
Plasmids, siRNAs and transfections
DLG2 (NM_001351274.2) overexpression plasmids on a backbone of pCMV6-AC-GFP (catalogue # PS100010) vector were purchased from Origene Technologies. siRNA targeting DLG2 (s4122) or Silencer™ Select Negative control No. 1 siRNA (4,390,843) was purchased from Ambion (ThermoFischer Scientific). SW480 cells were grown to 80% confluence and subsequently transfected with; DLG2 plasmid, empty vector “mock” (pCMV6-AC-GFP), si-DLG2 or scrambled control “mock”. 100 ng plasmid-DNA or 10 pmol siRNA was complexed with 0.3 µl of Lipofectamine 2000 according to the Lipofectamine 2000 reagent forward transfection protocol (Invitrogen; ThermoFisher Scientific).
Inclusion of study subjects
Study subjects who underwent colonoscopy at the Sahlgrenska University Hospital, Gothenburg, Sweden were consecutively included in the study. Reasons for referral to colonoscopy for each participant are presented in Additional file 1. Forty controls and twenty patients who were diagnosed with colon cancer were included. The prerequisite for inclusion into the control group was ≥ 18 years of age, and a normal-appearing mucosa in the entire colon, e.g. patients with any significant pathology such as colonic polyps or adenomas, inflammatory bowel disease, malignancy, ischemic colitis etc. were excluded. Possibility of microscopic colitis was ruled out by light microscopic examination of biopsy specimens obtained from the mid-portion of the ascending colon as well as from the sigmoid. Presence of colonic diverticula was accepted provided there were no signs of acute diverticulitis and/or diverticulosis-associated colitis. The prerequisite for inclusion into the colon cancer group was the presence of at least one malignant tumor in the colon and ≥ 18 years of age. Tumors were classified according to the Tumor–Node–Metastasis (TNM) staging system (Compton et al. 2000).
Collection of tissue samples
During the diagnosis colonoscopy, mucosa samples were obtained from the mid-portion of the ascending colon (right side samples) as well from the sigmoid (left side samples) using a regular biopsy forceps. If applicable, a tissue sample was also collected from the tumor. The distance between the tumor and the matching cancer mucosa that was sampled on the same side as the tumor was approximately 10 cm. Tissue samples were frozen immediately in liquid nitrogen, and stored at − 80 °C until used.
Bacterial treatment in flies
The strain white (w-1118) (Bloomington Drosophila Stock Center) were used in the study. Newly laid eggs (within 3 h) were transferred to apple-agar plates with either control food (15% dry yeast, 17% mashed potato powder, 14% apple juice, 3% sugar, 1 grinded Probion placebo tablet per 10 g of food) or food supplemented with Probion Clinica (15% dry yeast, 17% mashed potato powder, 14% apple juice, 3% sugar, 1 grinded Probion Clinica tablet per 10 g of food). Larvae raised on control food were transferred to food supplemented with Probion Clinica after 1, 2, 3 or 4 days. After 5 days the larvae gut from 5 five larvae from each experiment were dissected and used for RNA extraction.
One Probion Clinica (Wasa Medicals AB, Halmstad, Sweden) tablet includes 7 × 109 CFUs Bifidobacterium lactis Bl-04 (ATCC SD5219), 3,5 × 109 CFUs Lactobacillus acidophilus NCFM (ATCC 700396) and 0.32 g inulin/xanthan mix.
DNA and RNA isolation
RNA was isolated from tissue samples using Qiagen AllPrep DNA/RNA/Protein Kit according to the manufacturer’s instructions. The samples were kept at – 20 °C until analysis. cDNA was synthesized from total RNA using the High Capacity cDNA Reverse Transcription Kit (no. 4368814, ThermoFisher Scientific) and run on a Bio-Rad T100 Thermal Cycler (Bio-Rad laboratories). RNA from SW480 and THP-1 cells were extracted with RNeasy Kit (Qiagen) according to manufacturer’s protocol. RNA was quantified by NanoDrop (NanoDrop Technologies) and 2 µg of RNA was reverse-transcribed into double stranded cDNA on a T-professional Basic Gradient thermal cycler (Biometra) using the High Capacity cDNA Reverse Transcription kit (Applied Biosystems). cDNA corresponding to 20 ng of RNA was used for each qPCR reaction.
Quantitative PCR analysis
The relative gene expression of selected genes was quantified using TaqMan® Assays labelled with FAM-MGB (ThermoFisher Scientific) (Table 1) or by SYBR green (Table 2). Samples were run as duplicates in 96-well plates. Polymerase chain reactions were carried out in 5 μl reactions with 1 × TaqMan™ Gene Expression Master Mix (no. 4369016, ThermoFisher Scientific), 1 × gene-specific assay and 2.5 μl cDNA. The plates were run and analyzed using the Pikoreal qPCR System (ThermoFisher Scientific) according to the manufacturer’s protocol. Thresholds and baselines were set manually and Ct values were extracted. All Ct values were normalized to the mean of the reference genes; ACTb, GAPDH, GUSB and PPIA (ΔCt) for each sample.
Protein analysis by Western blot
Protein was extracted from the transfected cells in 96 well plates (1 × 104 cells/well), by aspirating the media and incubating on ice for 5 min then adding ice cold RIPA buffer (ThermoFisher Scientific, 89901). Western blot analysis was performed using a Mini-PROTEAN® TGX™ 8–20% gradient gel (Bio-Rad), protein was blotted onto LF-PVDF membrane (8 min, 25 V and 2.5A) using a Trans-Blot® Turbo™ Transfer System (Bio-Rad). Blots were subsequently blocked for 1 h in 5% milk in TBST buffer (0.1% Tween-20 and 150 mM NaCl in 10 mM Tris–HCl, pH 7.4) as per the manufacturer’s recommendations. Primary antibodies (Table 3) were diluted in PBST (0.1% Tween-20 in PBS). After incubation with the primary antibodies the membranes were washed 3 × 10 min in TBST 0.1% (0.1% Tween-20 in tris-buffered saline). Secondary antibodies; Starbright B520 goat anti-rabbit (12,005,870, 1:5000, BioRad) and Starbright B700 goat anti-mouse (12,004,159, 1:5000, BioRad) were incubated for 1 h at RT. Transient transfection and Western blot analyses were performed in triplicate as three independent experiments. Image detection was performed on ChemiDoc MP (BioRad) and band intensity was quantified using Image lab™ (v. 6.1, BioRad). Protein loading from the different experiments and gels were normalized against total loaded protein from stain free images. The p-STAT, p-AKT, p-FOXO3 and p-S6 levels were calculated relative to the total STAT (STAT), total AKT (AKT), total FOXO3 (FOXO3) and total S6 (S6) protein quantities respectively. GAPDH was included as loading control.
Quantification of inflammasome formation by microscopy
5 µM DRAQ5™ nuclear stain (ThermoFisher Scientific) was added to the cells 5 min before imaging. Live cell imaging was performed at 37 °C, 5% CO2 using the Olympus scanR High-Content Screening Station and CellR software, UPLSAPO 20 × objective and Hamamatsu C8484 CCD camera. FITC and Cy5 fluorophore channels were used to visualize PYCARD-GFP and DRAQ5™ nuclear stain respectively. Each stimulation condition was imaged at 72 locations. Replicate experiments were performed on fresh aliquots of cells cultivated, treated and imaged at separate times.
Image analysis (FIJI)
Inflammasomes were identified using WEKA Trainable Segmentation plugin. The plugin was trained using 30 images of varying conditions and verified under all conditions. Identified specks were counted using the particle analyzer plugin. Cell nuclei were counted and used to determine total number of cells. Briefly, Gaussian blur (sigma = 2) was applied to images followed by an auto threshold step. Images were then converted to a binary image followed by 2-D watershed and the particle analyzer plugin was used to count cell nuclei.
Statistical analysis
All data presented are plotted as Tukey’s box and whisker plots showing IQR, line at the median, + at the mean with whiskers ± 1.5-fold of interquartile range from at least three independent experiments, or as a Kaplan–Meier to determine event free survival over time. For all multi-group analyses, differences were determined by one-way ANOVA test followed by Holm–Sidak’s multiple comparison test. For comparisons between two groups a Mann–Whitney U test was used: *p < 0.05, **p < 0.01, ***p < 0.001. All analyses were conducted using GraphPad Prism version 8.0.1 for Windows, (GraphPad Software, http://www.graphpad.com).
Results
DLG2 expression was low in inflamed tissues and in colorectal tumors, whereas NLRP3 and NFKBIZ expressions were high in inflamed tissues
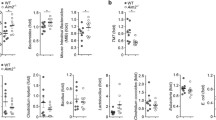
We evaluated the expression of DLG2, NLRP3 and NFKBIZ genes using publicly available microarray data for the independent colon cohorts (GSE4183; Fig. 1a, e, i) (Galamb et al. 2008), (GSE109142; Fig. 1b, f, j) (Haberman et al. 2019), (GSE75214; Fig. 1c, g, k) (Vancamelbeke et al. 2017), (GSE10950; Fig. 1d, h, l) (Jiang et al. 2008), obtained from the R2 Genomics Analysis and Visualization Platform (http://r2.amc.nl). In the different datasets gene expression was investigated in samples from patients with inflammatory bowel disease (IBD), adenoma or colon cancer compared to healthy controls (Fig. 1a); ulcerative colitis (UC) patients compared to controls (Fig. 1b); or ulcerative colitis (UC) patients with active vs. inactive disease state (Fig. 1c) and case controlled colorectal tumor samples compared to paired healthy mucosa (Fig. 1d). DLG2 was downregulated in IBD (log2 FC = 0.81, p < 0.05), adenoma (log2 FC = 1.1, p < 0.1) and colon cancer (log2 FC = 1.3, p < 0.01; Fig. 1a). DLG2 also showed a decrease in expression in UC compared to the control (log2 FC = 0.14, p < 0.01; Fig. 1b) and further decrease in expression when UC was active (log2 FC = 0.22, p < 0.001; Fig. 1c). A large downregulation in DLG2 was seen in the paired healthy-tumor colon tissue from colorectal cancer patients (log2 FC = 12.6, p < 0.001; Fig. 1d). There was no difference in NLRP3 expression in samples from patients with IBD, adenoma or colon cancer compared to healthy controls (Fig. 1e). An increased NLRP3 expression was detected in UC compared to control samples (log2 FC = 0.59, p < 0.001; Fig. 1f), and a further increase in NLRP3 expression when the UC was active (log2 FC = 0.74, p < 0.001; Fig. 1g). A lower NLRP3 expression was seen in the colon tissue compared to paired healthy mucosa in colorectal cancer patients (log2 FC = 1.1, p < 0.01; Fig. 1h). The NFKBIZ expression was higher in IBD samples compared to controls (log2 FC = 0.59, p < 0.001; Fig. 1i), however, no difference from controls was detected in adenoma or colon cancer samples (Fig. 1i). There was also an increased NFKBIZ expression in UC samples compared to the control (log2 FC = 1.8, p < 0.001; Fig. 1j) and a further increase in NFKBIZ expression when UC was active (log2 FC = 0.78, p < 0.001; Fig. 1k). However, a downregulation in NFKBIZ was seen in the paired tumor tissue compared to healthy mucosa from colorectal cancer patients (log2 FC = 0.52, p < 0.001; Fig. 1l). Additional cytokines were investigated with IL1RN showing increased expression in IBD, adenoma and CRC compared to healthy controls, however no significant difference between groups in TGFB1, IL4, IL10, IL13, IL27 or IL37 expression (Supplementary Fig. 1).
The alteration of DLG2, NLRP3 and NFKBIZ gene expression in colon inflammation and cancer. a DLG2, e NLRP3 and i NFKBIZ gene expression by sample type including; 8 Healthy patients, 15 IBD patients, 15 adenoma patients and 15 colon cancer patients (cohort GSE4183). b DLG2, f NLRP3, and j NFKBIZ expression in 20 healthy individuals compared to 206 with Ulcerative colitis (UC) (cohort GSE109142). c DLG2, g NLRP3 and k NFKBIZ expression in 73 patients UC patients with active disease compared to 23 patients with inactive disease (cohort GSE75214). d DLG2, h NLRP3 and l NFKBIZ expression of 24 case-controlled tumor mucosa samples (cohort GSE10950). The expression data are presented as median centered log2 fold change and plotted as Tukeys box and whisker plots showing IQR, line at the median, + at the mean with whiskers ± 1.5-fold of interquartile range. Data outside the whiskers are shown as outliers. The paired data are shown as an individual symbol with a connecting line. *p < 0.05, **p < 0.01, ***p < 0.001
DLG2 expression was initially upregulated followed by downregulation over time in response to inflammatory signals
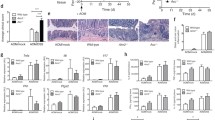
We evaluated the expression of DLG2, NLRP3 and NFKBIZ genes using publicly available microarray data in mouse colon from mice treated with Dextran Sulfate Sodium (DSS) to induce a colitis like phenotype (Fang et al. 2012) (GSE22307) and T cell transfer (Fang et al. 2011) (GSE27302) to model chronic colitis, obtained from the R2 Genomics Analysis and Visualization Platform (http://r2.amc.nl). DLG2 was upregulated in the colitis mouse model 4 days after DSS treatment with no difference between 0 and 6 days of DSS treatment (log2 FC = 0.47, p < 0.001) (Fig. 2a). When given a T-cell transfer, DLG2 expression in mice was decreased after 4 and 6 weeks (log2 FC = 0.57, p < 0.05 and log2 FC = 0.52, p < 0.05, respectively; Fig. 2b). When THP-1 monocytes were treated with Lipopolysaccharides (LPS) to induce immune responses, there was an initial increase in DLG2 expression 12 h after exposure (log2 FC = 0.879, p < 0.001; Fig. 2c) then a decrease in DLG2 was detected 24 h post exposure (log2 FC = 1.63, p < 0.001; Fig. 2c). The expression of the Drosophila melanogaster DLG2 ortholog dmDLG increased in fly larvae gut cells in response to Bifidobacterium lactis Bl-04 and Lactobacillus acidophilus NCFM, 24 h post treatment (log2 FC = 0.98, p < 0.01; Fig. 2d), with a progressive and gradual decrease in dmDLG over time until four days post treatment (log2 FC = − 0.64, p < 0.05; Fig. 2d). NLRP3 expression increased after six days of DSS treatment (log2 FC = 1.9, p < 0.001; Fig. 2e) and after T cell transfer by increasing expression between 4 and 6 weeks after treatment (log2 FC = 1.0, p < 0.01 and log2 FC = 1.6, p < 0.01, respectively; Fig. 2f). When THP-1 cells were treated with LPS there was no alteration in NLRP3 expression over time (Fig. 2g). NFKBIZ expression responded to DSS treatment after six days (log2 FC = 0.52, p < 0.01; Fig. 2h) and to T cell transfer by increasing its expression across all time points against the control up to 6 weeks after treatment (log2 FC = 0.80, p < 0.05, log2 FC = 1.5, p < 0.001 and log2 FC = 1.4, p < 0.01, respectively; Fig. 2i). When THP-1 cells were treated with LPS there was an initial increase in NFKBIZ expression 12 h after exposure (log2 FC = 0.83, p < 0.001; Fig. 2j) with the increase sustained 24 h post exposure (log2 FC = 0.84, p < 0.001; Fig. 2j).
The response of DLG2, NLRP3 and NFKBIZ gene expression to inflammation in mouse, cell and fly models. a DLG2, e NLRP3 and h NFKBIZ gene expression in the colon tissue of 5 mice for each time point in response to DSS treatment at 0, 2-, 4- and 6-days post treatment (cohort GSE22307). b DLG2, f NLRP3, and i NFKBIZ expression in the colon tissue of 4 mice in response to T-cell transfer at 0, 2-, 4- and 6- weeks post transfer (cohort GSE27302). c DLG2, g NLRP3 and j NFKBIZ expression in THP1 cells in response to LPS treatment at 0, 12 and 24 h. d The expression of dmDLG in Drosophila melanogaster gut in response to lactate bacteria treatment at 0, 1-, 2-, 3-, 4- days post treatment. The expression data are presented as median centered log2 fold change and plotted as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001
DLG2 overexpression results in increased NFKB components
Using differentiated THP-1 monocytes we compared mock transfection to DLG2 overexpressed and subsequent activation with either growth media, LPS or LPS with ATP. Gene expression analysis of NFKB1, NFKBIZ, RELA and RELB (Fig. 3 a-d) was performed in response to the growth conditions. We determined that there was no difference in RELA expression between the control and the DLG2 transfected cells (Fig. 3a). RELB showed a consistent upregulation in response to DLG2 overexpression, a stronger effect than both LPS and ATP treatments had (log2FC = 3.55, p < 0.01, log2FC = 4.46, p < 0.01 and log2FC = 4.17, p < 0.01; Fig. 3b). We investigated the expression of NFKB1 and showed that like RELB, the expression was consistently upregulated in the DLG2 expressed cells, with no additional effect by addition of LPS or ATP (log2FC = 3.51, p < 0.01, log2FC = 3.27, p < 0.01 and log2FC = 3.65, p < 0.01; Fig. 3c). Finally, we investigated the expression of NFKBIZ which was upregulated across all of the activations compared to the control with DLG2 overexpressed cells showing higher expression (log2FC = 3.77, p < 0.001, log2FC = 4.18, p < 0.001 and log2FC = 2.667, p < 0.001; Fig. 3d). We subsequently confirmed that the effects of DLG2 overexpression seen on gene expression level, also affected the protein expression, visualized by immunoblot for; DLG2, p-P65 Ser536, RELA, RELB, p-P105 Ser932, NFκB (P105), and IκBζ, using GAPDH as loading control (Fig. 3e). With the p-P65 Ser536 immunoblot indicating a low level of P65-phosphorylation across all activations. Despite the p-P105 Ser932 immunoblot detected higher amounts of phosphorylated NFκB (P105) in the DLG2 transfected cells, these transfections maintained lower relative phosphorylation compared to total amount of NFκB (P105) (Fig. 3e). DLG2 stimulated inflammasome formation and increased apoptosis in macrophage like cells.
The response of NFκβ members to increased DLG2 expression and activation of the inflammasome in THP-1 cells. The expression of a RELA, b RELB, c NFKB1 and d NFKBIZ in response to control conditions (circle symbol), LPS priming (square symbol) and LPS + ATP treatment (triangle symbol) with or without DLG2 overexpression. e Representative immunoblot showing the expression of; DLG2, p-P65 (Ser 536), RELA, RELB, p-P105 (Ser 932), NFκB1, IκBζ and GAPDH. Each experiment was performed in triplicate. The gene expression data are presented as log2 fold change and plotted as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001
DLG2 overexpression results in inflammasome formation in THP1 cells
Using differentiated THP-1 monocytes we compared mock transfection to DLG2 overexpressed cells and treated the cells with either growth medium, LPS or LPS with ATP. We determined the gene expression level of IL1B, IL6, BAX and BCL2 (Fig. 4a–d) and NLRP3 (supplementary 2). IL1B showed higher expression in the DLG2 transfected cells regardless of activation when compared to the equivalent activation (log2FC = 5.45, p < 0.001, log2FC = 3.22, p < 0.01 and log2FC = 3.03, p < 0.01, for the control, LPS and LPS + ATP respectively; Fig. 4a). DLG2 attenuated IL6 expression after activation with LPS and LPS and ATP (log2FC = 2.78, p < 0.001 and log2FC = 3.29, p < 0.001), with no difference in non-activated cells (Fig. 4b). DLG2 overexpression also resulted in consistently higher BAX expression across all activations (log2FC = 1.45, p < 0.01, log2FC = 1.90, p < 0.001 and log2FC = 1.57, p < 0.01; Fig. 4c) and consistently lower BCL2 expression across all activations (log2FC = 0.99, p < 0.01, log2FC = 1.01, p < 0.01 and log2FC = 1.33, p < 0.01; Fig. 4d). We subsequently determined the protein expression by immunoblot for DLG2, TIG1, BAX, BCL2, ser727 p-STAT3, total STAT3, ser235/236 p-S6, total S6 and GAPDH (Fig. 4e). DLG2 overexpression resulted in increased BAX expression in non-activated, LPS and LPS + ATP stimulated cells, and protein expression of BCL2 was decreased across all activations, which agreed with the gene expression data. STAT3 phosphorylation increased stepwise in the mock transfected cells with the LPS + ATP treatment showing the highest phosphorylation. The overexpression of DLG2 resulted in an increase of STAT3 phosphorylation during the LPS only treatment and a subsequent decrease during the LPS + ATP treatment (Fig. 4e). Furthermore, p-S6 was also decreased in all DLG2 transfections while total S6 expression remained unaffected (Fig. 4e). The expression of TIG1 is increased in the control activations compared to the controls. As we previously determined overexpression of DLG2 resulted in an increase in IL1B gene expression across all conditions. Finally, we investigated PYCARD/ASC speck formation in THP1 cells. We detected that DLG2 overexpression resulted in an increase in PYCARD speck formation (8.3% more, p < 0.01; Fig. 4f) and DLG2 silencing inhibited PYCARD speck formation (6.99% less, p < 0.01; Fig. 4f) in THP1 cells with stably transfected GFP tagged PYCARD compared to the control. To confirm the transfection efficiency, we determined DLG2 protein by immunoblot (Fig. 4g).
The response of cytokines and apoptotic proteins to increased DLG2 level and formation of the inflammasome in THP-1 cells. The gene expression of a IL1B, b IL6, c BAX and d BCL2 in response to control conditions (circle symbol), LPS priming (square symbol) and LPS + ATP treatment (triangle symbol) with or without DLG2 overexpression. e Representative immunoblot showing the expression of; BAX, BCL2, p-STAT3, STAT3, p-S6, S6, TIG1 and GAPDH. f Activation of the inflammasome in DLG2 silenced, control and DLG2 overexpressed THP1-ASC-GFP cells, in response to LPS and ATP treatment, as determined by PYCARD/ASC speck formation, presented as a percentage of observed cells. g The transfection efficiency of DLG2 in THP1 cells for silenced, control and DLG2 overexpression. Each experiment was performed in triplicate. The expression data are presented as log2 fold change and plotted as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001
DLG2 activated inflammasomes result in decreased colon cell proliferation
To model the effect of inflammasome formation on adjacent colon cancer cells we first quantified the amount of IL-1β and IL-6 in the supernatant taken from the transfected and activated THP-1 cells by immunoblot (Fig. 5a). We could show that silencing of DLG2 expression resulted in a slight decrease of IL-1β and a strong increase in IL-6, while overexpression of DLG2 had the opposite effect (Fig. 5a). We subsequently tested if the altered expression in IL-6 and IL-1β would affect the tumor microenvironment and modify signaling in colon cancer cells by treating COLO205 cells with the supernatant from THP1 transfected cells combined with regular growth media (1:1) followed by cell growth for 72 h. We detected that the DLG2 knockdown THP1 cell media increased the proliferation of COLO205 (22.0% more cells/ml, p < 0.05; Fig. 5b) as well as increasing the proportion of cells in G2/M phase (62.8% more G2/M cells, p < 0.05; Fig. 5c) when compared to the control cells. DLG2 overexpression resulted in the opposite of this, decreasing the cell proliferation (7.3% less cells, p < 0.01; Fig. 5b), and the number of cells in G2/M (34,9% less G2/M cells, p < 0.01; Fig. 5c). To show that NFκB and apoptosis signaling pathways were affected in response to these treatments we visualized protein expression of DLG2, RELA, RELB, IκBζ, NFKB1, BCL2, BAX, p-STAT3 Ser727, total STAT3 and GAPDH by immunoblot. These results showed that media from THP-1 DLG2-silenced cells decreased protein expression of RELB and BAX, and increased ser727 phosphorylation of STAT3 in COLO205 cells (Fig. 5d, e). Media from THP 1 DLG2 overexpressed cells resulted in increased protein level of RELB, NFκB1 and decreased level of phosphorylation of STAT3 (Fig. 5d, e).
The response of THP-1 cytokine production in colon cancer COLO205 cells. a Representative western blot showing the level of IL-1β and IL-6 production and release in the cell medium in DLG2 silenced, control and DLG2 overexpressed THP-1 cells in response to LPS and ATP, normalized to total protein level. b The effect of cell medium from DLG2 silenced (siDLG2), control and DLG2 overexpressed (DLG2) THP-1 cells on COLO205 cell proliferation, and c percentage of COLO205 cells in G2/M phase. Representative immunoblot showing the effect of THP-1 inflammasome growth media on the expression of d RELA, RELB, IκBζ, NFκB1, e BAX, BCL2, p-STAT3, STAT3, normalized to GAPDH. Each experiment was performed in triplicate. The expression data are presented as log2 fold change and plotted as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001
DLG2 expression was low in colon tumors and controlled signaling pathways
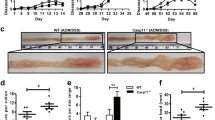
We could show that DLG2 gene expression was not significantly different between the ascending and descending colon in healthy controls (Log2 fold change = 0.04, p > 0.05; Fig. 6a). DLG2 expression in the tumor tissue was lower than the paired mucosa sample (Log2 fold change = 1.89, p < 0.001), as well as the paired ascending and descending colon mucosa from the distal healthy colon tissue (Log2 fold change = 1.36, p < 0.05, Log2 fold change = 1.32, p < 0.05, respectively; Fig. 6a). Using publicly available microarray colon adenoma data (Sabates-Beliver et al. 2007) (GSE8671) we determined the expression of DLG2 relative to adenoma size. We could also show that DLG2 expression decreased as colon adenoma size increased to 1.1–1.5 cm and larger than 1.5 cm when compared to tumors under 1 cm in diameter (Log2 fold change = 1.32, p < 0.01 and Log2 fold change = 1.32, p < 0.01; Fig. 6b). To confirm these results, we determined proliferation in the colon cancer cells SW480 after DLG2 silencing or overexpression, DLG2 silencing resulted in an increase in SW480 proliferation (29.4% more cells, p < 0.001; Fig. 6c) and overexpression resulted in a decrease in proliferation (19.6% less cells, p < 0.001; Fig. 6c) compared to the control, 48 h after transfection. Using the dataset (Agesen et al. 2012) (GSE24551) to determine the if the expression of DLG2 resulted in altered survivability in colorectal cancer patients we performed a Kaplan–Meier survival analysis, with high DLG2 expression increasing the probability of 5-year patient survival.
The expression of DLG2 in colon cancer and the effect of low DLG2 level in colon cancer cells. a The expression of DLG2 in 20 paired samples from the healthy ascending colon (Healthy A), healthy descending colon (Healthy D), mucosa from colon cancer patients 10 cm adjacent to the tumor (Mucosa) and colon cancer tumor (Tumor). b TheDLG2 expression in colon adenomas comparing 32 samples < 1 cm, 16 samples between 1 and 1.5 cm and 15 samples larger than 1.5 cm. c The effect of DLG2 silencing (siDLG2) and overexpression (DLG2) on SW480 cell proliferation 48 h post transfection. d Kaplan–Meier survival curve showing the 5-year event free survival probability of colon cancer patients. e–g The gene expression of e NLRP3, f NFKBIZ, and g FOXO3 in response to DLG2 overexpression in SW480 cells. Normalized level of protein phosphorylation of h AKT1, i FOXO3 and j S6 in response to DLG2 overexpression in SW480 cells. k Representative immunoblot showing the effect of DLG2 overexpression on SW480 cells showing the expression of DLG2, p-AKT (s473), AKT1, p-FOXO3a (s318/321), FOXO3, p-S6 (s235/236) and S6, normalized to GAPDH. Each experiment was performed in triplicate. The expression data are presented as log2 fold change and plotted as either Tukey’s box and whisker plots showing IQR, line at the median, + at the mean with whiskers ± 1.5-fold of interquartile range or as mean. *p < 0.05, **p < 0.01, ***p < 0.001
We detected lower level of NLRP3 (Log2 fold change = − 1.1, p < 0.05; Fig. 6e), and higher levels of NFKBIZ (Log2 fold change = 1.05, p < 0.05; Fig. 6f) and FOXO3 (Log2 fold change = 0.53, p < 0.05; Fig. 6g) gene expressions after DLG2 overexpression and compared to the mock transfection. At the protein level we subsequently determined that the percentage of phosphorylation of AKT (15.0%, p < 0.05; Fig. 6h), FOXO3 (26.4%, p < 0.05; Fig. 6i), and S6 (51.2%, p < 0.05; Fig. 6j) was lower in Sw480 cells overexpressing DLG2, as visualized in a representative immunoblot (Fig. 6k).
Discussion
The digestive system is a complex series of organs that contains a high percentage of immune cells in order to protect the body from pathogens in the event of disruption of the physical mucosal and epithelial barrier. If the barrier is broken, invading pathogens trigger a PRR immune response and can induce inflammasome formation, the purpose of which is to induce an immune cascade to prepare the innate immune system as well as initiate repair of the physical barrier. Autoimmune diseases with chronic inflammation often separate these functions with chronic immune cascade signaling and minimal repair.
In this study we showed that DLG2 was downregulated in human colon tumor tissue with the lowest DLG2 level seen in larger size adenomas, and that silencing of DLG2 caused an increase in colon cancer cell proliferation in vitro. Previously, it has been established that DLG2 directly binds FASL which has been shown to mediate NLRP3 inflammasome mediated apoptosis in liver metastases (Dupaul-Chicoine al. 2015). Here we show that overexpression of DLG2 resulted in increased NFKBIZ expression, which is required for inflammasome activation (Horber et al. 2016). Increased expression of NFKBIZ is also known to curtail STAT3 activity and inhibit proliferation (Wu et al. 2009). Previous reports identified mutations in NFKBIZ in colon cancer which disrupt a stop codon, producing an abnormally long C-terminal region. This mutation is thought to affect the interactions with nuclear factor-κB complexes that bind to that region, altering the transcriptional regulation of its target genes and leading to cancer predisposition (Esteban-Jurado et al. 2015). Furthermore, NFKBIZ has been shown to be downregulated in bladder cancer and to affect the PI3K/AKT/mTOR pathway to inhibit proliferation (Xu et al. 2021). To test if DLG2 influenced these pathways we investigated the phosphorylation levels of AKT, FOXO3 and S6 in colon cancer cells. We could show that DLG2 overexpression reduced AKT phosphorylation at S473, the phosphorylation level of FOXO3 was maintained but the total protein was increased resulting in a decrease in the percentage of phosphorylation. The largest effect in response to DLG2 overexpression was observed in the decrease in phosphorylated S6, highlighting that mTORC1 signaling seems to be altered with DLG2 expression. High levels of phosphorylated S6 in colon cancer have been shown to be related to high nodal metastasis and high tumor histologic grade (Lai et al. 2014). Previous results have shown that the colon of colitis animal models have a high degree of phosphorylation of S6 suggesting that mTOR is involved in the disease, with inhibition of mTOR attenuating DSS induced colitis (Hu et al. 2016). In the ulcerative colitis (UC) patient data, in accordance with previous literature, NFKBIZ was upregulated in UC patients and even more in active UC cases, whereas we could show that DLG2 was downregulated.
To investigate if DLG2 downregulation is a direct result of inflammation, we investigated the effect of inflammation on DLG2 gene expression in mouse, cell and fly models. We could show that acute inflammation initially resulted in an increase in DLG2 expression across all models with a subsequent decrease over time, whereas NLRP3 and NFKBIZ trended to increase over time. The initial increase and subsequent decrease in DLG2 expression in response to inflammation implies that DLG2 could respond to an inflammation feedback loop. To validate if DLG2 expression affected the inflammatory cell response we investigated the formation of inflammasomes in THP1 cells after DLG2 silencing, which resulted in decreased inflammasome formation after LPS and ATP stimulation (supplementary 3). Additionally, we were able to show that overexpression of DLG2 resulted in increased NFKBIZ, RELB, NFKB1 and IL-1β expression but a decrease in IL-6. It has previously been shown that DLG2 increases p53 expression (Keane et al. 2022), which can inhibit IL-6 expression (Zhang et al. 2016) and cause IL-1β mediated cell cycle arrest (Guadagno et al. 2015). The homodimerization of p50 combined with IκBζ results in the transcription of anti-inflammatory genes and functions as a tumor suppressor (Cartwright et al. 2016). Previously, it has been shown that NFKBIZ prepares the priming stage of inflammasome formation by controlling NLRP3 and pro IL-1β expression. The second signal required for inflammasome formation in the colon is the release of ADP by injured cells (Zhang et al. 2020a, b). Additionally, ATP is commonly thought of as an activator and extracellular levels of ATP are also commonly elevated in the tumor microenvironment (Di Virgilio and Adinolfi 2017; Alvarez et al. 2021). It has been noted that this signal results in an increase of IL-1β with marginal effect on TNF-α and IL-6. We show that p-P105 Ser932, which is activated by TNF-α, remains constant in the control activations and the DLG2 transfected activations, implying that TNF-α levels are also stable.
This work builds on the growing body of evidence that DLG2 functions as a tumor suppressor. Recently, low DLG2 expression has been found in osteosarcoma (Shao et al. 2019), ovarian cancer (Zhuang et al. 2019) and neuroblastoma (Keane et al. 2020, 2021; Siaw et al. 2020). The limitation of these studies has been that the mechanism and function of DLG2 has not been directly shown, but generally inferred from bioinformatic analysis. Here we show that DLG2 is downregulated in inflammatory bowel diseases such as UC as well as colon cancers, indicating that DLG2 alteration occurs early in the tumorigenesis process. We also show that the downregulation is directly dependent on inflammation. The effect of DLG2 loss is lowered expression of NFKB1 and NFKBIZ, both of which are protective against carcinogens that cause genotoxic damage, providing a molecular mechanism for previous results showing that DLG2 maintains genome integrity (Keane et al. 2022). Furthermore, previous bioinformatic results suggested that DLG2 is involved in DNA replication (Keane et al. 2020), the cell cycle (Keane et al. 2020), apoptosis (Shao et al. 2019) and chemokine signaling (Shao et al. 2019). We have been able to confirm in colon cancer cells that increased DLG2 results in an increase in BAX and a decrease in BCL2 resulting in lower cell proliferation. Additionally, DLG2 overexpression in THP-1 cells resulted in an altered cytokine and growth factor profile which was subsequently used to treat colon cancer cells decreasing the number in G2/M cell cycle phase. Which may be due to cell death caused by pyroptosis, however this would need to be further evaluated. Furthermore, a decrease in the phosphorylation of S6, consistent with control of proliferation was detected after DLG2 overexpression. Decreased S6 phosphorylation has been shown to inhibit the synthesis of the chemokine IL-8 (Ang et al. 2019). Finally, DLG2 loss has been shown to increase cyclin A2 and result in S phase progression and DNA replication (Keane et al. 2020), a similar function to the integral inflammasome component GSDMD (Wang et al. 2018).
Modulation of the immune system to treat cancers has been increasing over time, however, due to the complex nature and function of both the digestive system and immune system there has been some difficulty in developing immunotherapies for colon cancer. In this study we have shown that DLG2 in macrophage like cells can activate the formation of the inflammasome. The suppression of DLG2 seen in chronic inflammatory disease patients is one of the early changes that occurs and facilitates the formation of tumors. Restoration of DLG2 in the colon may provide a mechanism for improved immunotherapy function as well as attenuating inflammatory bowel diseases. However, further work will be needed to evaluate the therapeutic potential of DLG2 modulation.
Data availability
The datasets generated during and/or analysed during the current study are available in the R2: genomics analysis visualization platform, http://r2.amc.n listed in the methods by GSE identification number.
Abbreviations
- CAC:
-
Colitis associated colorectal cancer
- SCC:
-
Spontaneous colorectal cancer
- TLR:
-
Toll like receptors
- PRR:
-
Pattern recognition receptor
- RIG-I:
-
Retinoic acid inducible gene-I
- EMT:
-
Epithelial-mesenchymal transition
- RARRES1 :
-
Retinoic acid response element 1
- IBD:
-
Inflammatory bowel disease
- UC:
-
Ulcerative colitis
- DSS:
-
Dextran sulfate sodium
References
Agesen TH, Sveen A, Merok MA, Lind GE, Nesbakken A, Skotheim RI, Lothe RA (2012) ColoGuideEx: a robust gene classifier specific for stage II colorectal cancer prognosis. Gut 61(11):1560–1567
Alvarez CL, Troncoso MF, Espelt MV (2021) Extracellular ATP and adenosine in tumor microenvironment: Roles in epithelial-mesenchymal transition, cell migration, and invasion. J Cell Physiol
Ang Z, Koean RAG, Er JZ, Lee LT, Tam JKC, Guo H, Ding JL (2019) Novel AU-rich proximal UTR sequences (APS) enhance CXCL8 synthesis upon the induction of rpS6 phosphorylation. PLoS Genet 15(4):e1008077
Bank S, Andersen PS, Burisch J, Pedersen N, Roug S, Galsgaard J, Turino SY, Brodersen JB, Rashid S, Rasmussen BK, Avlund S, Olesen TB, Hoffmann HJ, Thomsen MK, Thomsen VO, Frydenberg M, Nexo BA, Sode J, Vogel U, Andersen V (2014) Associations between functional polymorphisms in the NFkappaB signaling pathway and response to anti-TNF treatment in Danish patients with inflammatory bowel disease. Pharmacogenomics J 14(6):526–534
Cartwright T, Perkins ND, L. W. C, (2016) NFKB1: a suppressor of inflammation, ageing and cancer. FEBS J 283(10):1812–1822
Chu EC, Chai J, Tarnawski AS (2004) NSAIDs activate PTEN and other phosphatases in human colon cancer cells: novel mechanism for chemopreventive action of NSAIDs. Biochem Biophys Res Commun 320(3):875–879
Compton C, Fenoglio-Preiser CM, Pettigrew N, Fielding LP (2000) American Joint Committee on cancer prognostic factors consensus conference: colorectal working group. Cancer 88(7):1739–1757
Corvinus FM, Orth C, Moriggl R, Tsareva SA, Wagner S, Pfitzner EB, Baus D, Kaufmann R, Huber LA, Zatloukal K, Beug H, Ohlschlager P, Schutz A, Halbhuber KJ, Friedrich K (2005) Persistent STAT3 activation in colon cancer is associated with enhanced cell proliferation and tumor growth. Neoplasia 7(6):545–555
Curtin K, Wolff RK, Herrick JS, Abo R, Slattery ML (2010) Exploring multilocus associations of inflammation genes and colorectal cancer risk using hapConstructor. BMC Med Genet 11:170
Di Virgilio F, Adinolfi E (2017) Extracellular purines, purinergic receptors and tumor growth. Oncogene 36(3):293–303
Dibra D, Mitra A, Newman M, Xia X, Cutrera JJ, Gagea M, Kleinerman ES, Lozano G, Li S (2016a) Lack of immunomodulatory interleukin-27 enhances oncogenic properties of mutant p53 In Vivo. Clin CancerRes 22(15):3876–3883
Dibra D, Xia X, Mitra A, Cutrera JJ, Lozano G, Li S (2016b) Mutant p53 inconcert with an interleukin-27 receptor alpha deficiency causes spontaneous liver inflammation, fibrosis, andsteatosis in mice. Hepatol 63(3):1000–1012
Dupaul-Chicoine J, Arabzadeh A, Dagenais M, Douglas T, Champagne C, Morizot A, Rodrigue-Gervais IG, Breton V, Colpitts SL, Beauchemin N, Saleh M (2015) The Nlrp3 inflammasome suppresses colorectal cancer metastatic growth in the liver by promoting natural killer cell tumoricidal activity. Immunity 43(4):751–763
Esteban-Jurado C, Vila-Casadesus M, Garre P, Lozano JJ, Pristoupilova A, Beltran S, Munoz J, Ocana T, Balaguer F, Lopez-Ceron M, Cuatrecasas M, Franch-Exposito S, Pique JM, Castells A, Carracedo A, Ruiz-Ponte C, Abuli A, Bessa X, Andreu M, Bujanda L, Caldes T, Castellvi-Bel S (2015) Whole-exome sequencing identifies rare pathogenic variants in new predisposition genes for familial colorectal cancer. Genet Med 17(2):131–142
Fang K, Bruce M, Pattillo CB, Zhang S, Stone R 2nd, Clifford J, Kevil CG (2011) Temporal genomewide expression profiling of DSS colitis reveals novel inflammatory and angiogenesis genes similar to ulcerative colitis. Physiol Genomics 43(1):43–56
Fang K, Zhang S, Glawe J, Grisham MB, Kevil CG (2012) Temporal genome expression profile analysis during t-cell-mediated colitis: identification of novel targets and pathways. Inflamm Bowel Dis 18(8):1411–1423
Galamb O, Gyorffy B, Sipos F, Spisak S, Nemeth AM, Miheller P, Tulassay Z, Dinya E, Molnar B (2008) Inflammation, adenoma and cancer: objective classification of colon biopsy specimens with gene expression signature. Dis Markers 25(1):1–16
Guadagno J, Swan P, Shaikh R, Cregan SP (2015) Microglia-derived IL-1beta triggers p53-mediated cell cycle arrest and apoptosis in neural precursor cells. Cell Death Dis 6:e1779
Haberman Y, Karns R, Dexheimer PJ, Schirmer M, Somekh J, Jurickova I, Braun T, Novak E, Bauman L, Collins MH, Mo A, Rosen MJ, Bonkowski E, Gotman N, Marquis A, Nistel M, Rufo PA, Baker SS, Sauer CG, Markowitz J, Pfefferkorn MD, Rosh JR, Boyle BM, Mack DR, Baldassano RN, Shah S, Leleiko NS, Heyman MB, Grifiths AM, Patel AS, Noe JD, Aronow BJ, Kugathasan S, Walters TD, Gibson G, Thomas SD, Mollen K, Shen-Orr S, Huttenhower C, Xavier RJ, Hyams JS, Denson LA (2019) Ulcerative colitis mucosal transcriptomes reveal mitochondriopathy and personalized mechanisms underlying disease severity and treatment response. Nat Commun 10(1):38
Hai Ping P, Feng Bo T, Li L, Nan Hui Y, Hong Z (2016) IL-1beta/NF-kb signaling promotes colorectal cancer cell growth through miR-181a/PTEN axis. Arch Biochem Biophys 604:20–26
Horber S, Hildebrand DG, Lieb WS, Lorscheid S, Hailfinger S, Schulze-Osthoff K, Essmann F (2016) The atypical inhibitor of NF-kappaB, IkappaBzeta, controls macrophage interleukin-10 expression. J Biol Chem 291(24):12851–12861
Hu S, Chen M, Wang Y, Wang Z, Pei Y, Fan R, Liu X, Wang L, Zhou J, Zheng S, Zhang T, Lin Y, Zhang M, Tao R, Zhong J (2016) mTOR inhibition attenuates dextran sulfate sodium-induced colitis by suppressing T cell proliferation and balancing TH1/TH17/Treg profile. PLoS ONE 11(4):e0154564
Jahani-Sherafat S, Alebouyeh M, Moghim S, Ahmadi Amoli H, Ghasemian-Safaei H (2018) Role of gut microbiota in the pathogenesis of colorectal cancer; a review article. Gastroenterol Hepatol Bed Bench 11(2):101–109
Janakiram NB, Rao CV (2009) Role of lipoxins and resolvins as anti-inflammatory and proresolving mediators in colon cancer. Curr Mol Med 9(5):565–579
Jang JH, Kim DH, Surh YJ (2021) Dynamic roles of inflammasomes in inflammatory tumor microenvironment. NPJ Precis Oncol 5(1):18
Jiang X, Tan J, Li J, Kivimae S, Yang X, Zhuang L, Lee PL, Chan MT, Stanton LW, Liu ET, Cheyette BN, Yu Q (2008) DACT3 is an epigenetic regulator of Wnt/beta-catenin signaling in colorectal cancer and is a therapeutic target of histone modifications. Cancer Cell 13(6):529–541
Jones S, Chen WD, Parmigiani G, Diehl F, Beerenwinkel N, Antal T, Traulsen A, Nowak MA, Siegel C, Velculescu VE, Kinzler KW, Vogelstein B, Willis J, Markowitz SD (2008) Comparative lesion sequencing provides insights into tumor evolution. Proc Natl Acad Sci U S A 105(11):4283–4288
Keane S, Ameen S, Lindlof A, Ejeskar K (2020) Low DLG2 gene expression, a link between 11q-deleted and MYCN-amplified neuroblastoma, causes forced cell cycle progression, and predicts poor patient survival. Cell Commun Signal 18(1):65
Keane S, Martinsson T, Kogner P, Ejeskar K (2021) The loss of DLG2 isoform 7/8, but not isoform 2, is critical in advanced staged neuroblastoma. Cancer Cell Int 21(1):170
Keane S, de Weerd HA, Ejeskar K (2022) DLG2 impairs dsDNA break repair and maintains genome integrity in neuroblastoma. DNA Repair (amst) 112:103302
Kim J, Ahn H, Yu S, Ahn JH, Ko HJ, Kweon MN, Hong EJ, An BS, Lee E, Lee GS (2020) IkappaBzeta controls NLRP3 inflammasome activation via upregulation of the Nlrp3 gene. Cytokine 127:154983
Kolligs FT (2016) Diagnostics and epidemiology of colorectal cancer. Visceral Med 32(3):158–164
Lai EY, Chen ZG, Zhou X, Fan XR, Wang H, Lai PL, Su YC, Zhang BY, Bai XC, Li YF (2014) DEPTOR expression negatively correlates with mTORC1 activity and tumor progression in colorectal cancer. Asian Pac J Cancer Prev 15(11):4589–4594
Lund EK, Belshaw NJ, Elliott GO, Johnson IT (2011) Recent advances in understanding the role of diet and obesity in the development of colorectal cancer. Proc Nutr Soc 70(2):194–204
Matzinger P (1994) Tolerance, danger, and the extended family. Annu Rev Immunol 12:991–1045
Ostvik AE, Svendsen TD, Granlund AVB, Doseth B, Skovdahl HK, Bakke I, Thorsvik S, Afroz W, Walaas GA, Mollnes TE, Gustafsson BI, Sandvik AK, Bruland T (2020) Intestinal epithelial cells express immunomodulatory ISG15 during active ulcerative colitis and Crohn’s disease. J Crohns Colitis 14(7):920–934
Ozawa N, Yokobori T, Osone K, Katayama C, Suga K, Komine C, Shibasaki Y, Shiraishi T, Okada T, Kato R, Ogawa H, Sano A, Sakai M, Sohda M, Ojima H, Miyazaki T, Motegi Y, Ide M, Yao T, Kuwano H, Shirabe K, Saeki H (2021) PD-L1 upregulation is associated with activation of the DNA double-strand break repair pathway in patients with colitic cancer. Sci Rep 11(1):13077
Sabates-Beliver J, Van der Flier LG, de Palo M, Cattaneo E, Maake C, Rehrauer H, Laczko E, Kurowski MA, Bujnicki JM, Menigatti M, Luz J, Ranalli TV, Gornes V, Pastorelli A, Faggiani R, Anti M, Jiricny J, Clevers H, Marra G (2007) Transcriptome profile of human colorectal adenomas. Mol Cancer Res 5(12):1263–1275
Sahab ZJ, Hall MD, Zhang L, Cheema AK, Byers SW (2010) Tumor suppressor RARRES1 regulates DLG2, PP2A, VCP, EB1, and Ankrd26. J Cancer 1:14–22
Santiago L, Castro M, Sanz-Pamplona R, Garzon M, Ramirez-Labrada A, Tapia E, Moreno V, Layunta E, Gil-Gomez G, Garrido M, Pena R, Lanuza PM, Comas L, Jaime-Sanchez P, Uranga-Murillo I, Del Campo R, Pelegrin P, Camerer E, Martinez-Lostao L, Munoz G, Uranga JA, Alcalde A, Galvez EM, Ferrandez A, Bird PI, Metkar S, Arias MA, Pardo J (2020) Extracellular granzyme a promotes colorectal cancer development by enhancing gut inflammation. Cell Rep 32(1):107847
Shao YW, Wood GA, Lu J, Tang QL, Liu J, Molyneux S, Chen Y, Fang H, Adissu H, McKee T, Waterhouse P, Khokha R (2019) Cross-species genomics identifies DLG2 as a tumor suppressor in osteosarcoma. Oncogene 38(2):291–298
Shao X, Lei Z, Zhou C (2020) NLRP3 promotes colorectal cancer cell proliferation and metastasis via regulating epithelial mesenchymal transformation. Anticancer Agents Med Chem 20(7):820–827
Sharma RA, Gescher A, Plastaras JP, Leuratti C, Singh R, Gallacher-Horley B, Offord E, Marnett LJ, Steward WP, Plummer SM (2001) Cyclooxygenase-2, malondialdehyde and pyrimidopurinone adducts of deoxyguanosine in human colon cells. Carcinogenesis 22(9):1557–1560
Shi W, Zou R, Yang M, Mai L, Ren J, Wen J, Liu Z, Lai R (2019) Analysis of genes involved in ulcerative colitis activity and tumorigenesis through systematic mining of gene co-expression networks. Front Physiol 10:662
Shi YJ, Zhao QQ, Liu XS, Dong SH, Li X, Liu C, Wang H (2020) Toll-like receptor 4 regulates spontaneous intestinal tumorigenesis by up-regulating IL-6 and GM-CSF. J Cell Mol Med 24(1):385–397
Shi F, Wei B, Lan T, Xiao Y, Quan X, Chen J, Zhao C, Gao J (2021) Low NLRP3 expression predicts a better prognosis of colorectal cancer. Biosci Rep 41(4)
Siaw JT, Javanmardi N, Eynden JVD, Lind DE, Fransson S, Martinez-Monleon A, Djos A, Sjoberg RM, Ostensson M, Caren H, Troen G, Beiske K, Berbegall AP, Noguera I, Lai WY, Kogner P, Palmer RH, Hallberg B, Martinsson T (2020) 11q deletion or ALK activity curbs DLG2 expression to maintain an undifferentiated state in neuroblastoma. Cell Rep 32(12):108171
Singh RK, Chang HW, Yan D, Lee KM, Ucmak D, Wong K, Abrouk M, Farahnik B, Nakamura M, Zhu TH, Bhutani T, Liao W (2017) Influence of diet on the gut microbiome and implications for human health. J Transl Med 15(1):73
Slattery ML, Wolff RK, Herrick JS, Caan BJ, Potter JD (2007) IL6 genotypes and colon and rectal cancer. Cancer Causes Control 18(10):1095–1105
Ungerback J, Belenki D, Jawad ul-Hassan A, Fredrikson M, Fransen K, Elander N, Verma D, Soderkvist P (2012) Genetic variation and alterations of genes involved in NFkappaB/TNFAIP3- and NLRP3-inflammasome signaling affect susceptibility and outcome of colorectal cancer. Carcinogenesis 33(11):2126–2134
Vancamelbeke M, Vanuytsel T, Farre R, Verstockt S, Ferrante M, Van Assche G, Rutgeerts P, Schuit F, Vermeire S, Arijs I, Cleynen I (2017) Genetic and transcriptomic bases of intestinal epithelial barrier dysfunction in inflammatory bowel disease. Inflamm Bowel Dis 23(10):1718–1729
Verma D, Sarndahl E, Andersson H, Eriksson P, Fredrikson M, Jonsson JI, Lerm M, Soderkvist P (2012) The Q705K polymorphism in NLRP3 is a gain-of-function alteration leading to excessive interleukin-1beta and IL-18 production. PLoS ONE 7(4):e34977
Wang WJ, Chen D, Jiang MZ, Xu B, Li XW, Chu Y, Zhang YJ, Mao R, Liang J, Fan DM (2018) Downregulation of gasdermin D promotes gastric cancer proliferation by regulating cell cycle-related proteins. J Dig Dis 19(2):74–83
Wang A, Deng S, Chen X, Yu C, Du Q, Wu Y, Chen G, Hu L, Hu C, Li Y (2020) miR-29a-5p/STAT3 positive feedback loop regulates TETs in colitis-associated colorectal cancer. Inflamm Bowel Dis 26(4):524–533
Willems M, Dubois N, Musumeci L, Bours V, Robe PA (2016) IkappaBzeta: an emerging player in cancer. Oncotarget 7(40):66310–66322
Wu Z, Zhang X, Yang J, Wu G, Zhang Y, Yuan Y, Jin C, Chang Z, Wang J, Yang X, He F (2009) Nuclear protein IkappaB-zeta inhibits the activity of STAT3. Biochem Biophys Res Commun 387(2):348–352
Xiong H, Zhang ZG, Tian XQ, Sun DF, Liang QC, Zhang YJ, Lu R, Chen YX, Fang JY (2008) Inhibition of JAK1, 2/STAT3 signaling induces apoptosis, cell cycle arrest, and reduces tumor cell invasion in colorectal cancer cells. Neoplasia 10(3):287–297
Xu T, Rao T, Yu WM, Ning JZ, Yu X, Zhu SM, Yang K, Bai T, Cheng F (2021) Upregulation of NFKBIZ affects bladder cancer progression via the PTEN/PI3K/Akt signaling pathway. Int J Mol Med 47(6)
Yamazaki S, Muta T, Takeshige K (2001) A novel IkappaB protein, IkappaB-zeta, induced by proinflammatory stimuli, negatively regulates nuclear factor-kappaB in the nuclei. J Biol Chem 276(29):27657–27662
Zaki MH, Vogel P, Body-Malapel M, Lamkanfi M, Kanneganti TD (2010) IL-18 production downstream of the Nlrp3 inflammasome confers protection against colorectal tumor formation. J Immunol 185(8):4912–4920
Zhang Q, Raoof M, Chen Y, Sumi Y, Sursal T, Junger W, Brohi K, Itagaki K, Hauser CJ (2010) Circulating mitochondrial DAMPs cause inflammatory responses to injury. Nature 464(7285):104–107
Zhang T, Li H, Shi J, Li S, Li M, Zhang L, Zheng L, Zheng D, Tang F, Zhang X, Zhang F, You X (2016) p53 predominantly regulates IL-6 production and suppresses synovial inflammation in fibroblast-like synoviocytes and adjuvant-induced arthritis. Arthritis Res Ther 18(1):271
Zhang C, Qin J, Zhang S, Zhang N, Tan B, Siwko S, Zhang Y, Wang Q, Chen J, Qian M, Liu M, Du B (2020a) ADP/P2Y1 aggravates inflammatory bowel disease through ERK5-mediated NLRP3 inflammasome activation. Mucosal Immunol 13(6):931–945
Zhang Y, Pu W, Bousquenaud M, Cattin S, Zaric J, Sun LK, Ruegg C (2020b) Emodin inhibits inflammation, carcinogenesis, and cancer progression in the AOM/DSS model of colitis-associated intestinal tumorigenesis. Front Oncol 10:564674
Zhuang RJ, Bai XX, Liu W (2019) MicroRNA-23a depletion promotes apoptosis of ovarian cancer stem cell and inhibits cell migration by targeting DLG2. Cancer Biol Ther 20(6):897–911
Acknowledgements
We thank the Jane and Dan Olsson foundation, Assar Gabrielssons Foundation, Royal Physiographic Society of Lund and University of Skövde for financial support. We thank the staff at the Centre for Cellular Imaging at the University of Gothenburg for their advice and assistance in fluorescence imaging. A-L. Helminen and H. Björkqvist are acknowledged for help with implementation of the NCT03072641 study and collection of patient samples. We thank L. Munro for clinical database assistance, and J. Flach and M. Åkerström for technical assistance.
Funding
Open access funding provided by University of Skövde. Jane and Dan Olsson foundation (2020–2029), Assar Gabrielssons Foundation (FB21-06), Royal Physiographic Society of Lund (NA) and University of Skövde.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation and acquisition was performed by PR and YW. Data collection and analysis were performed by SK and MH. Funding was acquired by KE and SK. The first draft of the manuscript was written by SK and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose. The authors declare that they have no conflicts of interest.
Ethics approval
The study was approved by the Regional Ethical Review Board in Gothenburg (no. 233-10) and written informed consent was obtained from all participants. The procedures used in this study adhere to the tenets of the Declaration of Helsinki.
Consent to participate
For all research subjects, freely-given, informed consent to participate in the study was obtained.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
432_2022_4029_MOESM1_ESM.tif
The gene expression of (a) IL1N and TGFB1, and (b) IL4, IL10, IL13, IL27 or IL37 by sample type including; 8 Healthy patients, 15 IBD patients, 15 adenoma patients and 15 colon cancer patients (cohort GSE4183). IL1RN showing increased expression in IBD, adenoma and CRC compared to healthy controls. Data is plotted as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001 (TIF 1939 KB)
432_2022_4029_MOESM2_ESM.tif
The gene expression of NLRP3 under mock or DLG2 transfection combined with the three activation stages; basal, activation by LPS and activation by LPS + ATP. Each experiment was performed in triplicate. The gene expression data are presented as log2 fold change and plotted as mean ± SD (TIF 1129 KB)
432_2022_4029_MOESM3_ESM.tif
Representative images of inflammasome formation as determined by PYCARD/ASC speck formation, observed in THP1 cells activated with LPS + ATP for; (a) DLG2 silenced, (b) Mock transfection or (c) DLG2 overexpression (TIF 3378 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Keane, S., Herring, M., Rolny, P. et al. Inflammation suppresses DLG2 expression decreasing inflammasome formation. J Cancer Res Clin Oncol 148, 2295–2311 (2022). https://doi.org/10.1007/s00432-022-04029-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-022-04029-7