Abstract
Introduction
Low back pain is the leading contributor to disability burden globally. It is commonly due to degeneration of the lumbar intervertebral discs (LDD). Magnetic resonance imaging (MRI) is the current best tool to visualize and diagnose LDD, but places high time demands on clinical radiologists. Automated reading of spine MRIs could improve speed, accuracy, reliability and cost effectiveness in radiology departments. The aim of this review and meta-analysis was to determine if current machine learning algorithms perform well identifying disc degeneration, herniation, bulge and Modic change compared to radiologists.
Methods
A PRISMA systematic review protocol was developed and four electronic databases and reference lists were searched. Strict inclusion and exclusion criteria were defined. A PROBAST risk of bias and applicability analysis was performed.
Results
1350 articles were extracted. Duplicates were removed and title and abstract searching identified original research articles that used machine learning (ML) algorithms to identify disc degeneration, herniation, bulge and Modic change from MRIs. 27 studies were included in the review; 25 and 14 studies were included multi-variate and bivariate meta-analysis, respectively. Studies used machine learning algorithms to assess LDD, disc herniation, bulge and Modic change. Models using deep learning, support vector machine, k-nearest neighbors, random forest and naïve Bayes algorithms were included. Meta-analyses found no differences in algorithm or classification performance. When algorithms were tested in replication or external validation studies, they did not perform as well as when assessed in developmental studies. Data augmentation improved algorithm performance when compared to models used with smaller datasets, there were no performance differences between augmented data and large datasets.
Discussion
This review highlights several shortcomings of current approaches, including few validation attempts or use of large sample sizes. To the best of the authors' knowledge, this is the first systematic review to explore this topic. We suggest the utilization of deep learning coupled with semi- or unsupervised learning approaches. Use of all information contained in MRI data will improve accuracy. Clear and complete reporting of study design, statistics and results will improve the reliability and quality of published literature.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Low back pain (LBP) is the leading contributor to disability burden globally and work disability [1]. LBP has a multifactorial aetiology, however lumbar disc degeneration (LDD) identified by magnetic resonance imaging (MRI) increases the risk of self-reported LBP up to 3.6-fold [2]. Although MRI should not be used in LBP diagnosis unless serious pathology is suspected [3], it is the current best available imaging tool to view soft tissue disorders, playing a role in LBP management and surgical treatment planning. MRI use throughout medicine is increasing, due to clinical benefits and improved patient safety. Demands on radiology departments have grown considerably with required radiological work hours outstripping those available [4]. MRIs take longer to read than other radiographic studies due to complexity and volume [4], with clinical spine MRIs, averaging 14–19 min to assess by an experienced radiologist [5]. The COVID-19 pandemic has exacerbated this pressure, hampering health care service provision including LBP diagnosis and treatment [6].
Disc pathology identification and classification using MR imaging
Several grading systems are used to evaluate LDD [7,8,9]. It is commonly assessed with the 1–5 grade Pfirrmann scale [10]. Grade 1 healthy discs appear hyperintense, or bright on T2 weighted (T2W) MRI, due to their hydration. As discs dehydrate and degrade, image signal intensity is lost and at grade 5 badly degenerate discs appear black on T2W MRI, as depicted in Fig. 1 [11]. Shape changes are indicative: healthy discs are elliptical, whereas degenerated discs are flatter [12]. Modic change (MC) describes a bone marrow lesion in the vertebra adjacent to the bony endplate. MC type 1, associated with inflammation or increased water content in the endplate, is darker on T1 weighted (T1W) scans and brighter on T2W; types 2 and 3 show hyperintensity and hypointensity, respectively, on both T1W and T2W scans, as depicted in Fig. 2 [13,14,15]. Research to date has focused upon type 1, associated with advanced LDD, pain severity and worse prognosis [16]. The combination of LDD and endplate signal change is strongly associated with LBP [17, 18]. Capturing an accurate description of LDD on MRI presents several challenges. Distinguishing between the expected, age-related disc change in spine structures and abnormal or rapid degeneration that might lead to pain symptoms is difficult [19]. One study reported degenerate discs in 96% MRIs from 80 + year olds, who were not experiencing back pain [20]. There is high inter-rater variation between intervertebral disc pathology diagnoses and gradings [21,22,23]. The Pfirrmann scale only has moderate inter-rater agreement [24, 25] and can be difficult to use, with failures to distinguish early signs of degeneration [26]. The term disc bulge—whilst commonly used, lacks standardization and can be confusing, leading to poor communication between medical professionals [25].
Pfirrmann grades on MRI. Image courtesy of Pfirrmann, C. W. A., Metzdorf, A., Zanetti, M., Hodler, J., & Boos, N. (2001). Magnetic resonance classification of lumbar intervertebral disc degeneration. Spine, 26(17), 1873–1878. https://doi.org/10.1097/00007632-200109010-00011
Modic change on MRI. Typical Modic changes. a Type I changes; b type II changes; c type III changes. Image courtesy of Chen Y, Bao J, Yan Q, Wu C, Yang H, Zou J. Distribution of Modic changes in patients with low back pain and its related factors. Eur J Med Res. 2019 Oct 9;24(1):34. https://doi.org/10.1186/s40001-019-0393-6. PMID: 31,597,571, PMCID: PMC6784341
Machine learning
Machine learning (ML) developments offer a standardized approach and may detect patterns that have so far evaded human radiological enquiry. Further artificial intelligence (AI)-enabled solutions could improve the efficiency of reading scans [27] and accurately depict degeneration and other disc pathologies, thereby assisting clinicians and radiologists with correct diagnoses. Considering the shortfalls described above, simply to match human grading performance would not be optimal—ML may one day extend beyond current grading schemas. Several different ML approaches are currently used for automated reading of MRI scans [28] and inevitably their effectiveness needs to be evaluated. Deep learning (DL), algorithms use multiple layers of interconnected processing units able to extract and compile complex features from an input. Advantages to spine imaging such as standardized coding of defined phenotypes, improved accuracy of reading will improve research and patient care. Automated object recognition and classification for a broad range of spinal pathologies include successfully segmenting vertebral bodies on MRI [29,30,31,32,33,34,35]. Progress has been made in detection and classification of spinal deformity [36, 37] and high-performance ML models predict success or failure of spinal operations and postoperative complications [38, 39]. Early detection of pathological compositional rather than structural disc changes—has been demonstrated by comparing T1ro and T2 MRI relaxation times, important to note when there is a dearth of early process diagnostic tools [26]. Columbo software has recently been granted a world-first FDA clearance for their spine MRI-reading software [40]. While there are many encouraging reports of high accuracy models compared to human radiologists, there has been few replication studies which formally test algorithm validity. Along with replication trials, systematic, robust comparisons and evaluation of performance metrics that classify disc degeneration, herniation, budge or Modic change are needed. The aim of this review is to determine the ability of current ML technology for the classification of degenerate, herniated and bulged discs and Modic change. Successful ML models offer the exciting potential of real change for spine radiology, yet this promise is tempered by practical obstacles. Radiology departments will need to purchase special processing hardware (such as graphics processing units) to utilize algorithms yet may lack negotiating power to purchase at best price [41]. Rapidly improving technology presents the obvious threat of new ML assets being quickly out-of-date. Data and concept drifts need to be monitored as they can significantly undermine model performance in real-world settings [41, 42]. Models need to be not only accurate and reliable to be implemented clinically, but they need also regulatory approval. The beneficial, cost-effective implementation of ML technology in routine clinical practice goes well beyond the development and validation of software.
What this review will add
AI-enabled applications are increasing in use throughout medicine. ML models reading MRI could save radiologists time and potentially surpass human diagnostic or prognostic accuracy. In clinical settings, an algorithm must be cost- and time-efficient, reproducible, offer standardized outcomes that are user-friendly and easy to integrate into approved picture archiving communication systems (PACS) software. ML is new in medicine and the contribution to spinal MRI reading depends upon how effectively and reliably detect it can classify and grade disc degenerative conditions including herniation, bulge and Modic change. The primary aims of this review and meta-analysis are to identify (1) if one model or software approach performs consistently well in identifying lumbar disc degeneration, herniation, bulge and Modic change, (2) if any MRI diagnostic tool is more amenable to ML and (3) document limitations of current ML applications.
Methods
A systematic review protocol was developed in accordance with PRISMA guidelines. The review was registered with PROSPERO (CRD42021261659) on 13th of July 2021 and is accessible (https://www.crd.york.ac.uk/prospero/). Four electronic databases were searched: Cihal, Embase, PubMed, Web of Science (including Medline) on 18th June 2021. Search terms, commands and outputs for databases are shown in Supplementary 1 (S1). Grey literature search details in S2. Details of article search results shown in PRISMA flow diagram (Fig. 3).
Studies were included if they met the following criteria: (1) original research articles (2) used ML algorithms to diagnose or measure LDD, herniation, bulge or Modic change and (3) used MRIs performed in adult humans. Abstracts, comparison trials or observational studies, detailing relevant findings or validation research for established algorithms were included. No date or language limits were set. Exclusion criteria specified case reports, editorials, letters, other meta-analyses or reviews. Only papers specifically using ML algorithms to identify or grade disc degeneration, herniation or bulge or MC were included. ML performance was to be compared to at least one human radiologist. Studies that focused on related conditions, for example spinal stenosis (only), were excluded. Studies using an imaging modality other than MRI were excluded. Algorithms that focused on image parameters necessary for but not sufficient to analyze LDD, for example segmentation (only) were excluded. This search was corroborated, and inclusion agreed by consensus of co-authors including a consultant radiologist (RC, IGS and AI). Eligible studies (n = 27) were identified and included in the review (Table 1). Study details such as prospective/retrospective design, algorithm development/validation and use of pre-processing or data augmentation were documented.
Articles were tabled according to the PROBAST tool [43] to summarize the design of each study, assess risk of bias and determine the applicability of included models (S10). Studies that did not use standard disease definitions or did not report standard participant details or recruitment numbers or those failing to report variance statistics were ranked either unclear or high risk of bias.
Meta-analysis
Studies were grouped according to classification measures. Groups included Pfirrmann and MC grades and binary or numerical LDD, herniation and bulge classifications. Performance metrics for correctly classifying LDD such as accuracy, sensitivity, specificity, area under the curve (AUC) and F1 were recorded, with the primary aim of the analysis to identify if one algorithm consistently outperformed others. Authors of studies who did not report accuracy or variance statistics were contacted by email for these details. Pan et al. (2021) employed an unconventional accuracy definition, incompatible with standard metrics reported by other articles. These authors did not respond to our request for standard accuracy measurements and this study was omitted from the meta-analysis [44]. Zheng and colleagues developed a DL algorithm for segmentation with additional disc measurements to diagnose LDD without ML so this study was excluded from the meta-analysis [45].
MRI acquisition parameters recorded included Tesla (T) strength and plane (axial/sagittal/transverse). MRI sequences T1W and T2W were recorded, quantitative sequences such as T1rho, T2 mapping, DIXON, spectral fat suppression were not. Standard deviation (SD) and 95% confidence intervals (CI) of performance metrics were extracted. When CI was not reported, Z scores (1.96) were used to transform SD to CI. In studies reporting train-test split validation without variability, SD was inferred using the variability mean of other included studies.
Sensitivity and specificity bivariate mixed effects regression was performed on studies reporting both measures (Table 2) [46]. Subsequently, a multivariate mixed effects regression of accuracy, sensitivity, specificity, AUC and F1 was performed for studies included in the meta-analysis (Table 3). Regression was fitted using rma.mv function from R package Metafor (version 3.4–0). Validity of the regression was assessed using the restricted log-likelihood plots. Logit transformation was applied to both analyses. Algorithm, LDD classification, data augmentation and internal/external validation and scaled year of publication were used as predictors. Structure of the random effects correlation between measures of each study was defined as unstructured as described before [46]. ANOVA (Wald tests) were used to group categorical variables. Post-hoc Tukey tests for significant pairwise comparisons following ANOVA were run with false discovery rate (FDR)-adjusted p-values. Python (3.9.12) software was used for the analysis (scipy 1.8.0, statsmodels 0.13.2). Sensitivity and specificity bivariate mixed effects regression was performed on studies reporting both measures (Table 2) [46]. Subsequently, a multivariate mixed effects regression of accuracy, sensitivity, specificity, AUC and F1 was performed for studies included in the meta-analysis (Table 3). Regression was fitted using rma.mv function from R package Metafor (version 3.4–0). Validity of the regression was assessed using the restricted log-likelihood plots. Logit transformation was applied to both analyses. Algorithm, LDD classification, data augmentation and internal/external validation were used as predictors. Structure of the random effects correlation between measures of each study was defined as unstructured as described before [46]. ANOVA (Wald tests) were used to group categorical variables. Post-hoc Tukey tests for significant pairwise comparisons following ANOVA were run with FDR-adjusted p-values. Python (3.9.12) software was used for the analysis Python (3.9.12) software was used for the analysis (scipy 1.8.0, statsmodels 0.13.2).
Results
Types of studies
Of the 27 studies included, 22 were aimed at algorithm development, 2/27 reported development and external validation [45, 47] and 3/27 solely focused on external validation [48,49,50]. Most studies (24/27) used retrospective, pre-existing datasets from hospital or university collections; however, 4/27 studies prospectively examined patient scans [51,52,53,54]. Two distinct themes emerged from included articles: several had a clear focus on the underlying algorithm development and were written from a technical perspective while others, including the external validation studies, were written by and for a clinical audience. In three cases author groups published two different studies which used the same dataset [51, 55,56,57,58,59].
Magnetic resonance imaging specifications
Most studies used sagittal plane MRI (21/27), but one study examined disc herniation and two studies examined disc bulge using the axial plane, while 3/27 studies used both planes. All studies reported using T2W sequences and 7/27 of them additionally used T1W sequences. From studies reporting MR field strength (22/27), most used images acquired on a 1.5 Tesla (T) scanner, some in combination with 3 T scanners. One study used a 0.4 T [52] while 3/27 studies did not report the magnetic field strength. Due to inconsistencies of MRI sequences and planes, along with several studies failing to report MR scanner field strength, MRI acquisition parameters were not included as a predictors or variables in the meta-regression. Lewandrowski et al. (2020) used T2 fast spin echo in their software that graded herniation to generate radiology reports [60].
LDD classifications
14/27 studies used standard disease classifications including 6/27 investigating Pfirrmann grading and 3/27 MC, with one study examining both [50]. One study used numerical grading for herniation and 2/27 used descriptive grading for LDD while the remaining studies gave binary classifications for disc herniation, degeneration, or bulge. Most studies (14/27) used a single radiologist’s grading to establish ground truth. 5/27 studies used two raters, 2/27 studies used three raters and 1/27 study used four to validate ground truth. 3/27 studies did not report how MRIs were rated and 2/27 relied upon previous ratings (from medical reports). Of the studies using more than one human rater to establish ground truth, k-values for inter-rater agreement ranged from 0.54 [61] to 0.93 [47].
Performance metrics and algorithms
Studies used either tenfold [51, 53, 55,56,57, 61, 62] five-fold [54, 59, 63,64,65,66] or random sample split validation [47, 51, 52, 55, 56, 60, 67,68,69,70]. The bivariate model of 14 studies showed differences in performances between types of classifications. Specifically, studies examining herniation had higher performance metrics than those examining disc bulge. In the bivariate model, external validation studies performed on a par with developmental studies and studies using data augmentation showed superior performance (Table 2). In the multivariate analysis of 25 studies, external validation papers did not perform as well as development studies (Table 3). Studies using data augmentation had higher performance metrics than others. However, Table 4 shows this effect is lost when studies using data augmentation are compared only to studies using large sample sizes.
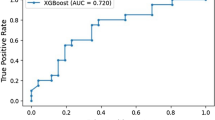
Sensitivity (Fig. 4) and specificity (Fig. 5) forest plots were produced. Performance receiver operating curve (ROC) is shown in Fig. 6. Forest plots depicting accuracy (S4), AUC (S5), F-1 (S6) and precision (S7) were made. These plots show the extreme heterogeneity between the included studies.
Forest plot of sensitivity. Forest plots depicting sensitivity of all algorithms examined in included studies. Grey shadow lines correspond to the DerSimonian and Laird adjusted variation. Reference marker sizes correspond to participants numbers of each study. * indicates algorithm performance with data augmentation, # indicates external validation studies. Confidence interval (CI), deep learning (DL), k nearest neighbor (kNN), random forest (RF), support vector machine (SVM)
Forest plot of specificity. Forest plots depicting specificity of all algorithms examined in included studies. Grey shadow lines correspond to the DerSimonian and Laird adjusted variation. Reference marker sizes correspond to participants numbers of each study. * indicates algorithm performance with data augmentation, # indicates external validation studies. Confidence interval (CI), deep learning (DL), k nearest neighbor (kNN), random forest (RF), support vector machine (SVM)
Sixteen studies with image datasets used DL algorithms, DL studies generally used large datasets, the average number of DL disc images was 4,211 (min 169, max 17,800), whereas non-DL mean disc images were 613 (min 93, max 2,500). DL models averaged an accuracy of 87.0% (SD 7.0%), specificity of 90.4% (SD 6.3%) and sensitivity 88.2% (SD 7.2%). 8/25 studies compared performance of several algorithms, including DL, SVM, kNN, NB and RF. Other studies used nonspecific algorithms which were listed as custom models (2/25).
Risk of bias and quality assessment
There were large amounts of missing information: many studies did not report ethical approval, participant consent or waiving of consent for retrospectively designed studies. Basic participant details were missing; participants’ mean age, or recruitment site were often not reported. The PROBAST tool, completed and agreed by RC and IGS ranked 7/27 studies as low, 16/27 unclear and 4/27 high risk of bias (S10). For many studies performance measures and CI information were missing. We contacted authors of 11/27 studies asking for the variance of reported statistics; of these two responded—but only one provided the requested information. Only 2/27 studies provided statements of data availability [45, 47]—this is now standard in fields like genetic epidemiology—and just one study provided a link to their algorithm code [45]. PROBAST judgments of the applicability of included studies was generally poor—however most included studies were developmental rather than models that could immediately translate to useable clinical or research tools. For several questions assessing study quality and applicability many papers did not include any or appropriate detail. Information was mainly missing for the following four questions, prohibiting judgments quality and applicability:
-
3.1
Was the outcome determined appropriately?
-
4.4
Were participants with missing data handled appropriately?
-
4.3
Were all enrolled participants included in the analysis?
-
4.7
Were relevant model performance measures evaluated appropriately?
Discussion
Most studies included in this systematic review used DL models for LDD, bulge or herniation classification or grading by Pfirrmann or MC criteria. DL models, made up of complex layered input networks, might be expected to surpass other algorithms for reading spine MRI. This expectation was echoed by the choice of many authors of studies in this review to utilize DL models. However, our meta-analysis results do not show differences in DL performance compared to other ML approaches. DL algorithms tend to have a high false positive error rate and are difficult to calibrate [71]. Development and validation of SpineNet predominated DL studies. While used for research purposes, this model may be the most suitable candidate to develop for clinical practice. The software can detect six spine abnormalities including MC with ~ 87% accuracy, [72] Pfirrmann grade, disc narrowing, spondylolisthesis and central canal stenosis [73]. External validation of SpineNet demonstrated a balanced accuracy of 76% for Pfirrmann and 78% for MC classification in a large population dataset [50] and class average accuracy of 55% for Pfirrmann grade classification [48]. An external validation of an open-source version of SpineNet (V2) [73] is currently under review.
Supervised models predominate medical research; however performance improvements may come from incorporating other training techniques such as semi-supervised learning which labels a small portion of a large training dataset. Subsequent unlabelled data can improve classification results. Lewandroski and colleagues (2020) successfully combined supervised, semi-supervised and unsupervised training of a natural language processing (NLP) algorithm to generate radiologist reports. A 3D model was fitted to MRIs (axial, sagittal and transverse) in a large dataset, with high performance metrics reported for separate DL models that graded disc bulge and herniation and stenosis. Detailed radiological reports with only trivial differences from human reports were generated [60]. This innovative study design demonstrates the effective use of semi- and unsupervised learning with NLP, an addition which could be adopted to DL reading of MRIs.
Two DL studies used extended Pfirrmann grading systems—which allow for uncertainties between exact categories [61], and reported high agreement with ground truth [45, 61]. Niemeyer et al., (2021) compared analyses that treated Pfirrmann grades as categorical, numerical and continuous variables, and found a linear regression of a continuous variable had higher numbers of slight errors but gave best results overall, with fewer large deviations from ground truth [61]. The extended scale had good correlation with disc hydration and other markers of degeneration [45]. Zheng et al., (2022) went on to externally validate their model in a small dataset and have sought intellectual property rights for the software, proposing their model can easily integrate into existing MRI systems [45].
Pan et al. (2021) successfully integrated three CNN models for location, definition and classification of discs and used ResNet modification to reduce overfitting [44]. Gao et al., (2021) reported similarly high accuracy to Pan et al. (2021), adding a regularization method to their two CNN models to enhance separation of differential features. Classification according to Pfirrmann categories is a challenge for DL and CNN, with many samples “between grades”—training samples are often at the margins of a grade, so can easily be misclassified in testing, reducing model performance [53]. A more complex architecture including a memory component to reduce overfitting and include information about neighboring tissue was used by Han et al.’s (2018) CNN model, resulting in very high performance. This study demonstrates architectural modifications can incorporate information from a range of spine tissues captured in MRI rather than limiting to a single structure [63]. In a similar fashion Gong et al. (2021) used several features of surrounding spine tissues in axial MRIs, positing axial images may better capture multiple structures [54].
Castro-Mateos et al.’s. (2016) small, prospective study compared four different algorithm classifiers and found NN-2 performed best. Authors note an effective system must pick up on features not detected by human radiologists; while not included in their design, this implies that a component of unsupervised training may have benefitted their model [52]. While these CNN models perform well, a specific constraint to this study and an over-arching limitation of the review is the incompatibility of Pfirrmann grading with LDD progression which presents a challenge to algorithm performance.
Linear approaches commonly used include support vector machines (SVM) that are ‘noise-tolerant’ classifiers with reduced propensity to overfitting, however these algorithms tend not to handle complex or extensive data [71]. SVM are typically used for classification problems, and rely on linearly separating data, based on pattern differences. ‘Support vectors’ are datapoints on the periphery of a category—thus they support the plane of separation or decision boundary. SVM are most successful in datasets where large margins between categories exist. As described, the Pfirrmann grading system may be more precise when used as a continuous than categorical measure, which more accurately reflects the progression of degenerating discs [45, 61]. Stand-out SVM results include those from Oktay and colleagues (2014) which achieved high accuracy identifying disc herniation by extracting several image features (intensity, texture, whole shape, context) and focusing on the difference image created by comparing T1W and T2W MRIs [74]. They state that using the difference image helps to disregard artifacts of either T1W or T2W images [74]. SVM is computationally inexpensive, therefore attractive for model development. For these models to perform well, images must be simplified with pre-processing techniques such as grey scale reduction [51, 57, 70, 74], window cropping [56], and thresholding [55]. Conceptually SVM appears less capable of handling the complexities of identifying disc pathology in MRIs than DL models.
Herniation classification was tested with kNN, SVM, NB and two types of dimensionality reduction by Ghosh et al. (2011) who found concatenating different texture, shape and intensity features improved performance [64]. Using a RF classifier to diagnose disc budge and desiccation, Sundarsingh and Kesavan (2020) also found combining one texture and one shape feature improved accuracy [68]. These small studies collectively report increased success with the incorporation of multiple features.
It is common to use small datasets for algorithm development, big data are available but often require extensive cleaning and preparation [75]. The use of data augmentation is prevalent among AI-developers, the creation of additional training data can improve validation performance [76]. Six studies in our review used varying extents of augmentation. Athertya and colleagues identified MC1 and MC2 cases in images from 100 participants [55, 56]. Ten MC1 cases were augmented to 160 using synthetic minority over sampling technique and MC2 cases were doubled from 88 to 176 [55, 56]. It is difficult to acquire ‘case’ scans for training, however a curated dataset of original MRIs may provide a better training tool than one so aggressively augmented [77, 78]. Other studies have also employed extreme augmentation, for example Tsai and colleagues (2021) used rotation and contrast and brightness augmentation ran several trials augmenting between 50 and 550 images, with the model maintaining high accuracy [67]. Niemeyer and colleagues (2021) used elaborate pre-processing [61], not undertaken in usual care which does reduce applicability, however this model outperformed most others in the review, and it will be interesting to see if training with such manicured data translates to good performance in any subsequent external validation studies. Jamaludin and colleagues (2016) produced a texture feature extraction guided algorithm to assess MC from images focused on features of vertebral endplates. They used data augmentation, in a similar, but less extreme fashion to Niemeyer et al. (2021), in both training and testing datasets posited to better reflect real-world circumstances [66]. Su et al. (2022) also employed a lighter touch, randomly cropping and rotating training images to enhance real-world applicability of their model [47]. MRI images may be less amenable to augmentation than non-medical, natural image classification tasks—augmenting scans may distort MRI datasets, creating unrealistic images [78]. Augmentation specifically designed for fMRI showed a 26% algorithm performance improvement over more modest traditional augmentation results [78, 79].
AI promises exciting developments such as the prediction of patient response to treatment—and differentiating patients to optimal treatments. The onus to publish novel findings is not limited to computer science. The performance of any algorithm purported to assist medicine needs to be well- and repeatedly validated, however we only found three replication studies [48,49,50]. CoLumbo software appears to have been marketed to radiology departments based on one published external validation study [49]. In this study, authors note the software could benefit from future improvement, yet it is unclear if such improvements have been undertaken. We contacted the company for publications validating the software and received documentation reporting a trial in three hospitals conducted to receive EU Medical Devices Directive approval (available from CoLumbo marketing). External validation of successful algorithms remains an essential part of real-world applicability. Another recent publication testing a “widely available algorithm” used for LDD grading reported it failed significantly on unseen data [79]. The algorithm being tested was not made public and their work has not been included in this review, but findings present a cautionary tale against blind acceptance of model outputs [79]. It also demonstrates the need for new ground truth labels, repeated validation studies and close surveillance of software performance integrity.
Strengths and limitations
This systematic review has some limitations. Firstly, several studies had very small sample sizes. Nikravan, Ebrahimzadeh and colleagues developed binary herniation classifiers. Using SVM and NN [59] followed by a kNN algorithm, which used grey thresholding intensity features to identify disc rupture and leaking annulus pulposus [58]. Good performance was reported in both studies with the same 30 participants, but only limited generalizability from samples of this size may be inferred [52, 58, 59, 62, 64].
Included studies had either a strong computer science approach or a clinical focus. A second limitation was that computer science-focused reports often omitted participant details. Reported performance metrics differed between studies and many lacked variance statistics, posing challenges to meta-analysis; frequently ethical committee statements, basic participant details, missing data contingencies, basic model variance statistics or reference to disc degeneration grading systems were missing—and none of the development studies reported a contingency table. Incomplete reporting has been documented by other reviewers, who have called for “international standards for study protocols and reporting that recognize challenges” inherent to DL and ML [80]. A recent large systematic review and meta-analysis of ML for medical diagnosis found that while speciality DL algorithms were generally clinically accurate, there was unacceptably high variation in reporting of metrics, methodology and reference standards [81]. Parsimonious reporting is an unnecessary waste in biomedical research [82] and robust clinical peer review is lacking, with a dearth of randomized trials, prospective or external validation studies; estimated as low as 6% of all published AI studies [83, 84].
The third significant limitation was that studies performed poorly in the PROBAST risk of bias and quality assessment. This is in part due to the lack of availability of an AI-specific assessment tool. We and others are frustrated by the lack of an AI-specific tool and anticipate the Delphi group’s publication of tools like TRIPOD-AI [41, 80].
Conclusion
MRI reading may be most suited to DL processing, with the presentation of greater quantities and more varied data. Semi- and unsupervised components will improve the chance of detecting patterns that currently elude human radiologists. Successful models will likely benefit from using greater numbers of features and the incorporation of information from tissue surrounding the disc. The use of multiple MRI planes and all clinical sequencies, along with incorporating the T1W and T2W difference image, may provide richer data for ML algorithms to process. Employing continuous grading classifications, more sympathetic to the progression of LDD may also be useful.
We encourage the use and sharing of large datasets for developing and validating models. While data augmentation may seem an attractive bypass, susceptibility to model overfitting threatens practical performance. Progression of this field is hampered by lack of external validation studies, although such work is the backbone of any robust model in any field.
The current scientific and reporting quality of ML studies to identify LDD is overall insufficient, and none have been reliably implemented as a clinical decision-making tool. Widely acceptable methodological and reporting guidelines for ML in LDD research are warranted yet remain unavailable. We attempted to pursue missing data to improve the strength of our conclusions, with limited success. Future research should aim to bridge the gap between biomedical engineering literature and clinical value of the software. This review highlights the need to move beyond simply matching radiologists' interpretation to extracting quantitative LDD representations that effectively utilize the full complexity of data contained in spine MRI sequencing.
Availability of data and material
Supplementary documents provided detailing systematic search strategy.
Code availability
Supplementary document.
References
Maher C, Underwood M, Buchbinder R (2017) Non-specific low back pain. Lancet 389(10070):736–747. https://doi.org/10.1016/S0140-6736(16)30970-9
MacGregor AJ, Andrew T, Sambrook PN, Spector TD (2004) Structural, psychological, and genetic influences on low back and neck pain: A study of adult female twins. Arthritis Care Res (Hoboken) 51(2):160–167. https://doi.org/10.1002/art.20236
National Institute for Health and Care Excellence Low back pain and sciatica in over 16s: assessment and management (NG59). http://www.nice.org.uk/guidance/ng59
McDonald RJ, Schwartz KM, Eckel LJ, Diehn FE, Hunt CH, Bartholmai BJ, Erickson BJ, Kallmes DF (2015) The effects of changes in utilization and technological advancements of cross-sectional imaging on radiologist workload. Acad Radiol 22(9):1191–1198. https://doi.org/10.1016/j.acra.2015.05.007
Hirschmann A, Cyriac J, Stieltjes B, Kober T, Richiardi J, Omoumi P (2019) Artificial intelligence in musculoskeletal imaging: Review of current literature, challenges, and trends. Semin Musculoskelet Radiol 23(3):304–311. https://doi.org/10.1055/s-0039-1684024
dos Santos IN, Pernambuco ML, da Silva AMB, Ruela G de A, de Oliveira AS (2021) Association between musculoskeletal pain and telework in the context of the COVID 19 pandemic: an integrative review. Rev Bras Med Trab 19(3):342–350. https://doi.org/10.47626/1679-4435-2021-812
Mysliwiec LW, Cholewicki J, Winkelpleck MD, Eis GP (2010) MSU Classification for herniated lumbar discs on MRI: Toward developing objective criteria for surgical selection. Eur Spine J 19(7):1087–1093. https://doi.org/10.1007/s00586-009-1274-4
Riesenburger RI, Safain MG, Ogbuji R, Hayes J, Hwang SW (2015) A novel classification system of lumbar disc degeneration. J Clin Neurosci 22(2):346–351. https://doi.org/10.1016/j.jocn.2014.05.052
Guellil N, Argawal N, Krieghoff M, Kaden I, Hohaus C, Meisel HJ, Schenk P (2022) Novel methods to measure height and volume in healthy and degenerated lumbar discs in MRIs: A reliability assessment study. Diagnostics 12(6). https://doi.org/10.3390/diagnostics12061437
Pfirrmann CWA, Dora C, Schmid MR, Zanetti M, Hodler J, Boos N (2004) MR image-based grading of lumbar nerve root compromise due to disk herniation: Reliability study with surgical correlation. Radiology 230(2):583–588. https://doi.org/10.1148/radiol.2302021289
Harada GK, Siyaji ZK, Younis S, Louie PK, Samartzis D, An HS (2020) Imaging in spine surgery: current concepts and future directions. Spine Surg Relat Res 4(2):99–110. https://doi.org/10.22603/SSRR.2020-0011
Videman T, Battié MC, Gibbons LE, Maravilla K, Manninen H, Kaprio J (2003) Associations between back pain history and lumbar MRI findings. Spine (Phila Pa 1976) 28(6):582–588. https://doi.org/10.1097/00007632-200303150-00013
Modic MT, Steinberg PM, Ross JS, Masaryk TJ, Carter JR (1988) Degenerative disk disease: Assessment of changes in vertebral body marrow with MR imaging. Radiology 166(1 I):193–199. https://doi.org/10.1148/radiology.166.1.3336678
Modic MT, Masaryk TJ, Ross JS, Carter JR (1988) Imaging of degenerative disk disease. Radiology 168(1):177–186. https://doi.org/10.1148/radiology.168.1.3289089
Kjaer P, Korsholm L, Bendix T, Sorensen JS, Leboeuf-Yde C (2006) Modic changes and their associations with clinical findings. Eur Spine J 15(9):1312–1319. https://doi.org/10.1007/s00586-006-0185-x
Dudli S, Fields AJ, Samartzis D, Karppinen J, Lotz JC (2016) Pathobiology of Modic changes. Eur Spine J 25(11):3723–3734. https://doi.org/10.1007/s00586-016-4459-7
Määttä JH, Wadge S, MacGregor A, Karppinen J, Williams FMK (2015) ISSLS PRIZE WINNER: Vertebral endplate (Modic) change is an independent risk factor for episodes of severe and disabling low back pain. Spine (Phila Pa 1976) 40(15). https://journals.lww.com/spinejournal/Fulltext/2015/08010/ISSLS_Prize_Winner__Vertebral_Endplate__Modic_.10.aspx
Teraguchi M, Yoshimura N, Hashizume H, Muraki S, Yamada H, Oka H, Minamide A, Nakagawa H, Ishimoto Y, Nagata K, Kagotani R, Tanaka S, Kawaguchi H, Nakamura K, Akune T, Yoshida M (2015) The association of combination of disc degeneration, end plate signal change, and Schmorl node with low back pain in a large population study: the Wakayama Spine Study. Spine J 15(4):622–628. https://doi.org/10.1016/j.spinee.2014.11.012
Watanabe T, Otani K, Sekiguchi M, Konno S-I (2022) Relationship between lumbar disc degeneration on MRI and low back pain: A cross-sectional community study. Fukushima J Med Sci 68(2):97–107. https://doi.org/10.5387/fms.2022-17
Brinjikji W, Luetmer PH, Comstock B, Bresnahan BW, Chen LE, Deyo RA, Halabi S, Turner JA, Avins AL, James K, Wald JT, Kallmes DF, Jarvik JG (2015) Systematic literature review of imaging features of spinal degeneration in asymptomatic populations. Am J Neuroradiol 36(4):811–816. https://doi.org/10.3174/ajnr.A4173
van Rijn JC, Klemetsö N, Reitsma JB, Majoie CBLM, Hulsmans FJ, Peul WC, Stam J, Bossuyt PM, den Heeten GJ (2005) Observer variation in MRI evaluation of patients suspected of lumbar disk herniation. Am J Roentgenol 184(1):299–303. https://doi.org/10.2214/ajr.184.1.01840299
Herzog R, Elgort DR, Flanders AE, Moley PJ (2017) Variability in diagnostic error rates of 10 MRI centers performing lumbar spine MRI examinations on the same patient within a 3-week period. Spine J 17(4):554–561. https://doi.org/10.1016/j.spinee.2016.11.009
Fu MC, Webb ML, Buerba RA, Neway WE, Brown JE, Trivedi M, Lischuk AW, Haims AH, Grauer JN (2016) Comparison of agreement of cervical spine degenerative pathology findings in magnetic resonance imaging studies. Spine J 16(1):42–48. https://doi.org/10.1016/j.spinee.2015.08.026
Pfirrmann CW, Metzdorf A, Zanetti M, Hodler J, Boos N (2001) Magnetic resonance classification of lumbar intervertebral disc degeneration. Spine (Phila Pa 1976) 26(17):1873–1878. https://doi.org/10.1097/00007632-200109010-00011
Li Y, Fredrickson V, Resnick DK (2015) How should we grade lumbar disc herniation and nerve root compression? A systematic review. Clin Orthop Relat Res 473(6):1896–1902. https://doi.org/10.1007/s11999-014-3674-y
Pandit P, Talbott JF, Pedoia V, Dillon W, Majumdar S (2016) T1ρ and T2-based characterization of regional variations in intervertebral discs to detect early degenerative changes. J Orthop Res 34(8):1373–1381. https://doi.org/10.1002/jor.23311
Gao KT, Tibrewala R, Hess M, Bharadwaj UU, Inamdar G, Link TM, Chin CT, Pedoia V, Majumdar S (2022) Automatic detection and voxel-wise mapping of lumbar spine Modic changes with deep learning. JOR Spine 5(2):e1204. https://doi.org/10.1002/jsp2.1204
Castiglioni I, Rundo L, Codari M, di Leo G, Salvatore C, Interlenghi M, Gallivanone F, Cozzi A, D’Amico NC, Sardanelli F (2021) AI applications to medical images: From machine learning to deep learning. Physica Medica, pp 839–24. https://doi.org/10.1016/j.ejmp.2021.02.006
Zhang L, Wang H (2021) A novel segmentation method for cervical vertebrae based on PointNet++ and converge segmentation. Comput Methods Programs Biomed 200105798. https://doi.org/10.1016/j.cmpb.2020.105798
Pang S, Pang C, Su Z, Lin L, Zhao L, Chen Y, Zhou Y, Lu H, Feng Q (2022) DGMSNet: Spine segmentation for MR image by a detection-guided mixed-supervised segmentation network. Med Image Anal 75102261. https://doi.org/10.1016/j.media.2021.102261
Li H, Luo H, Liu Y (2019) Paraspinal muscle segmentation based on deep neural network. Sensors (Switzerland) 19(12): https://doi.org/10.3390/s19122650
Hwang J, Hwang S (2021) Exploiting global structure information to improve medical image segmentation. Sensors 21(9):3249. https://doi.org/10.3390/s21093249
Silvoster L, Kumar RMS (2020) Efficient segmentation of lumbar intervertebral disc from MR images. IET Image Process 14(13):3243–3253. https://doi.org/10.1049/iet-ipr.2019.0971
Rehman F, Ali Shah SI, Riaz N, Gilani SO (2019) A Robust Scheme of Vertebrae Segmentation for Medical Diagnosis. IEEE Access 7120387–120398. https://doi.org/10.1109/ACCESS.2019.2936492
Kim Y, Kim D (2009) A fully automatic vertebra segmentation method using 3D deformable fences. Comput Med Imaging Graph 33(5):343–352. https://doi.org/10.1016/j.compmedimag.2009.02.006
Lee S, Choe EK, Kang HY, Yoon JW, Kim HS (2020) The exploration of feature extraction and machine learning for predicting bone density from simple spine X-ray images in a Korean population. Skeletal Radiol 49(4):613–618. https://doi.org/10.1007/s00256-019-03342-6
Lin H (2008) Identification of spinal deformity classification with total curvature analysis and artificial neural network. IEEE Trans Biomed Eng 55(1):376–382. https://doi.org/10.1109/TBME.2007.894831
Scheer JK, Oh T, Smith JS, Shaffrey CI, Daniels AH, Sciubba DM, Hamilton DK, Protopsaltis TS, Passias PG, Hart RA, Burton DC, Bess S, Lafage R, Lafage V, Schwab F, Klineberg EO, Ames CP (2018) Development of a validated computer-based preoperative predictive model for pseudarthrosis with 91% accuracy in 336 adult spinal deformity patients. Neurosurg Focus 45(5):E11. https://doi.org/10.3171/2018.8.FOCUS18246
Arvind V, Kim JS, Oermann EK, Kaji D, Cho SK (2018) Predicting surgical complications in adult patients undergoing anterior cervical discectomy and fusion using machine learning. Neurospine 15(4):329–337. https://doi.org/10.14245/ns.1836248.124
CoLumbo AI spine assistant receives an FDA 510(K) clearance. https://columbo.me/columbo-ai-spine-assistant-receives-an-fda-510k-clearance/ (Jan. 21, 2023).
Goergen SK, Frazer HML, Reddy S (2022) Quality use of artificial intelligence in medical imaging: What do radiologists need to know? J Med Imaging Radiat Oncol 66(2):225–232. https://doi.org/10.1111/1754-9485.13379
Soin A, Merkow J, Long J, Cohen JP, Saligrama S, Kaiser S, Borg S, Tarapov I, Lungren MP (2022) CheXstray: Real-time multi-modal data concordance for drift detection in medical imaging AI. https://doi.org/10.48550/arxiv.2202.02833
Wolff RF, Moons KGM, Riley RD, Whiting PF, Westwood M, Collins GS, Reitsma JB, Kleijnen J, Mallett S (2019) PROBAST: a tool to assess the risk of bias and applicability of prediction model studies. Ann Intern Med 170(1):51–58. https://doi.org/10.7326/M18-1376
Pan Q, Zhang K, He L, Dong Z, Zhang L, Wu X, Wu Y, Gao Y (2021) Automatically diagnosing disk bulge and disk herniation with lumbar magnetic resonance images by using deep convolutional neural networks: Method development study. JMIR Med Inform 9(5). https://doi.org/10.2196/14755
Zheng HD, Sun YL, Kong DW, Yin MC, Chen J, Lin YP, Ma XF, Wang HS, Yuan GJ, Yao M, Cui XJ, Tian YZ, Wang YJ (2022) Deep learning-based high-accuracy quantitation for lumbar intervertebral disc degeneration from MRI. Nat Commun 13(1):841. https://doi.org/10.1038/s41467-022-28387-5
Reitsma JB, Glas AS, Rutjes AWS, Scholten RJPM, Bossuyt PM, Zwinderman AH (2005) Bivariate analysis of sensitivity and specificity produces informative summary measures in diagnostic reviews. J Clin Epidemiol 58(10):982–990. https://doi.org/10.1016/j.jclinepi.2005.02.022
Su ZH, Liu J, Yang MS, Chen ZY, You K, Shen J, Huang CJ, Zhao QH, Liu EQ, Zhao L, Feng QJ, Pang SM, Li SL, Lu H (2022) Automatic grading of disc herniation, central canal stenosis and nerve roots compression in lumbar magnetic resonance image diagnosis. Front Endocrinol (Lausanne) 13. https://doi.org/10.3389/fendo.2022.890371
Grob A, Loibl M, Jamaludin A, Winklhofer S, Fairbank JCT, Fekete T, Porchet F, Mannion AF (2022) External validation of the deep learning system ‘SpineNet’ for grading radiological features of degeneration on MRIs of the lumbar spine. Eur Spine J 31(8):2137–2148. https://doi.org/10.1007/s00586-022-07311-x
Lehnen NC, Haase R, Faber J, Rüber T, Vatter H, Radbruch A, Schmeel FC (2021) Detection of degenerative changes on MR images of the lumbar spine with a convolutional neural network: A feasibility study. Diagnostics 11(5):902. https://doi.org/10.3390/diagnostics11050902
McSweeney TP, Saarakkala S, Tiulpin A, Jamaludin A, Kadir T, Niinimäki J, Karppinen J, Määttä J (2022) External validation of Spinenet, a deep learning model for automated grading of lumbar disc degeneration MRI features, using the Northern Finland Birth Cohort. Osteoarthritis Cartilage 30(Supplementary 1):S86. https://doi.org/10.1016/j.joca.2022.02.106
Beulah A, Sharmila TS, Pramod VK (2018) Disc bulge diagnostic model in axial lumbar MR images using Intervertebral disc Descriptor (IdD). Multimed Tools Appl 77(20):27215–27230. https://doi.org/10.1007/s11042-018-5914-8
Castro-Mateos I, Hua R, Pozo JM, Lazary A, Frangi AF (2016) Intervertebral disc classification by its degree of degeneration from T2-weighted magnetic resonance images. Eur Spine J 25(9):2721–2727. https://doi.org/10.1007/s00586-016-4654-6
Gao F, Liu S, Zhang X, Wang X, Zhang J (2021) Automated grading of lumbar disc degeneration using a push-pull regularization network based on MRI. J Magn Reson Imaging 53(3):799–806. https://doi.org/10.1002/jmri.27400
Gong H, Liu J, Li S, Chen B (2021) Axial-SpineGAN: Simultaneous segmentation and diagnosis of multiple spinal structures on axial magnetic resonance imaging images. Phys Med Biol 66(11):115014. https://doi.org/10.1088/1361-6560/abfad9
Athertya JS, Saravana Kumar G, Govindaraj J (2019) Detection of Modic changes in MR images of spine using local binary patterns. Biocybern Biomed Eng 39(1):17–29. https://doi.org/10.1016/j.bbe.2018.09.003
Athertya JS, Kumar GS (2021) Classification of certain vertebral degenerations using MRI image features. Biomed Phys Eng Express 7(4):45013. https://doi.org/10.1088/2057-1976/ac00d2
Beulah A, Sharmila TS, Pramod VK (2022) Degenerative disc disease diagnosis from lumbar MR images using hybrid features. Visual Comp 38(8):2771–2783. https://doi.org/10.1007/s00371-021-02154-x
Ebrahimzadeh E, Fayaz F, Nikravan M, Ahmadi F, Dolatabad MR (2018) Towards an automatic diagnosis system for lumbar disc herniation: The significance of local subset feature selection. Biomed Eng (Singapore) 30(6):1850044. https://doi.org/10.4015/S1016237218500448
Nikravan M, Ebrahimzadeh E, Izadi MR, Mikaeili M (2016) Toward a computer aided diagnosis system for lumbar disc herniation disease based on MR images analysis. Biomed Eng (Singapore) 28(6):1650042. https://doi.org/10.4015/S1016237216500423
Lewandrowski KU, Muraleedharan N, Eddy SA, Sobti V, Reece BD, León JFR, Shah S (2020) Feasibility of deep learning algorithms for reporting in routine spine magnetic resonance imaging. Int J Spine Surg 14S86–S97. https://doi.org/10.14444/7131
Niemeyer F, Galbusera F, Tao Y, Kienle A, Beer M, Wilke HJ (2021) A deep learning model for the accurate and reliable classification of disc degeneration based on MRI data. Invest Radiol 56(2):78–85. https://doi.org/10.1097/RLI.0000000000000709
Koh J, Chaudhary V, Dhillon G (2012) Disc herniation diagnosis in MRI using a CAD framework and a two-level classifier. Int J Comput Assist Radiol Surg 7(6):861–869. https://doi.org/10.1007/s11548-012-0674-9
Han Z, Wei B, Mercado A, Leung S, Li S (2018) Spine-GAN: Semantic segmentation of multiple spinal structures. Med Image Anal, pp 5023–5035. https://doi.org/10.1016/j.media.2018.08.005
Ghosh S, Alomari RS, Chaudhary V, Dhillon G (2011) Composite features for automatic diagnosis of intervertebral disc herniation from lumbar MRI. In: Proceedings of the Annual International Conference of the IEEE Engineering in Medicine and Biology Society, EMBS, pp 5068–5071. https://doi.org/10.1109/IEMBS.2011.6091255
He X, Landis M, Leung S, Warrington J, Shmuilovich O, Li S (2017) Automated grading of lumbar disc degeneration via supervised distance metric learning. Medical Imaging 2017: Computer-Aided Diagnosis 101341013443. https://doi.org/10.1117/12.2253688
Jamaludin A, Kadir T, Zisserman A (2016) Automatic Modic changes classification in spinal MRI. In: Lecture Notes in Computer Science (including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics) 940214–26. https://doi.org/10.1007/978-3-319-41827-8_2
Tsai JY, Hung IYJ, Guo YL, Jan YK, Lin CY, Shih TTF, Chen B bin, Lung CW (2021) Lumbar disc herniation automatic detection in magnetic resonance imaging based on deep learning. Front Bioeng Biotechnol, 9. https://doi.org/10.3389/fbioe.2021.708137
Sundarsingh S, Kesavan R (2020) Diagnosis of disc bulge and disc desiccation in lumbar MRI using concatenated shape and texture features with random forest classifier. Int J Imaging Syst Technol 30(2):340–347. https://doi.org/10.1002/ima.22374
Jamaludin A, Lootus M, Kadir T, Zisserman A, Urban J, Battié MC, Fairbank J, McCall I (2017) ISSLS PRIZE IN BIOENGINEERING SCIENCE 2017: Automation of reading of radiological features from magnetic resonance images (MRIs) of the lumbar spine without human intervention is comparable with an expert radiologist. Eur Spine J 26(5):1374–1383. https://doi.org/10.1007/s00586-017-4956-3
Hashia B, Mir AH (2020) Texture features’ based classification of MR images of normal and herniated intervertebral discs. Multimed Tools Appl 79(21–22):15171–15190. https://doi.org/10.1007/s11042-018-7011-4
Guo C, Pleiss G, Sun Y, Weinberger KQ (2017) On calibration of modern neural networks. 34th International Conference on Machine Learning, ICML 2017, pp 32130–2143..
Jamaludin A, Kadir T, Zisserman A (2017) SpineNet: Automated classification and evidence visualization in spinal MRIs. Med Image Anal, pp 4163–4173. https://doi.org/10.1016/j.media.2017.07.002
Windsor R, Jamaludin A, Kadir T, Zisserman A SpineNetV2: Automated Detection, Labelling and Radiological Grading Of Clinical MR Scans. arXiv e-prints. arXiv:2205.01683, May 01, 2022. http://arxiv.org/abs/2205.01683
Oktay AB, Albayrak NB, Akgul YS (2014) Computer aided diagnosis of degenerative intervertebral disc diseases from lumbar MR images. Comput Med Imaging Graph 38(7):613–619. https://doi.org/10.1016/j.compmedimag.2014.04.006
Galbusera F, Casaroli G, Bassani T (2019) Artificial intelligence and machine learning in spine research. JOR Spine 2(1):e1044. https://doi.org/10.1002/jsp2.1044
Chlap P, Min H, Vandenberg N, Dowling J, Holloway L, Haworth A (2021) A review of medical image data augmentation techniques for deep learning applications. J Med Imaging Radiat Oncol 65(5):545–563. https://doi.org/10.1111/1754-9485.13261
Shorten C, Khoshgoftaar TM (2019) A survey on image data augmentation for deep learning. J Big Data 6(1). https://doi.org/10.1186/s40537-019-0197-0
Nguyen KP, Fatt CC, Treacher A, Mellema C, Trivedi MH (2021) Anatomically informed data augmentation for functional MRI with applications to deep learning, pp 1–11. https://doi.org/10.1117/12.2548630.
Hornung AL, Hornung CM, Mallow GM, Barajas JN, Espinoza Orías AA, Galbusera F, Wilke H-J, Colman M, Phillips FM, An HS, Samartzis D (2022) Artificial intelligence and spine imaging: limitations, regulatory issues and future direction. Eur Spine J 1–15. https://doi.org/10.1007/s00586-021-07108-4
Liu X et al (2019) A comparison of deep learning performance against health-care professionals in detecting diseases from medical imaging: a systematic review and meta-analysis. Lancet Digital Health 1(6):e271–e297
Aggarwal R, Sounderajah V, Martin G, Ting DSW, Karthikesalingam A, King D et al (2021) Diagnostic accuracy of deep learning in medical imaging: a systematic review and meta-analysis. Npj Digital Medicine. Nat Res. https://doi.org/10.1038/s41746-021-00438-z
Cohen JF, Korevaar DA, Altman DG, et al (2016) STARD 2015 guidelines for reporting diagnostic accuracy studies: explanation and elaboration. BMJ Open 6:e012799. https://doi.org/10.1136/bmjopen-2016-012799
Kim DW, Jang HY, Kim KW, Shin Y, Park SH (2019) Design characteristics of studies reporting the performance of artificial intelligence algorithms for diagnostic analysis of medical images: Results from recently published papers. Korean J Radiol 20(3):405–410. https://doi.org/10.3348/kjr.2019.0025
Steffens D, Hancock MJ, Pereira LS, Kent PM, Latimer J, Maher CG (2016) Do MRI findings identify patients with low back pain or sciatica who respond better to particular interventions? A systematic review. Eur Spine J 25:1170–1187
Funding
This work has been supported by funding from Versus Arthritis (Grant Number 22467). R.C & T.M.S are supported by the Disc4All Marie Curie Consortium project. (H2020-MSCA-ITN-ETN-2020 GA: 9557).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose. The authors have no conflicts of interest to declare that are relevant to the content of this article. All authors certify that they have no affiliations with or involvement in any organization or entity with any financial interest or non-financial interest in the subject matter or materials discussed in this manuscript. The authors have no financial or proprietary interests in any material discussed in this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Warmest thanks to Jaro Karppinen and Francesco Gualdi for proof-reading, comments and encouragement.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Compte, R., Granville Smith, I., Isaac, A. et al. Are current machine learning applications comparable to radiologist classification of degenerate and herniated discs and Modic change? A systematic review and meta-analysis. Eur Spine J 32, 3764–3787 (2023). https://doi.org/10.1007/s00586-023-07718-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00586-023-07718-0