Abstract
Biological nanochannels or nanopores play a crucial role in basic biochemical processes in cells. Artificial nanopores possessing dimensions comparable to the size of biological molecules and mimicking the function of biological ion channels are of particular interest with respect to the design of biosensors with a sensitivity that can go down to the fM level and even to single molecule detection. Nanopore-based analysis (NPA) is currently a new research field with fascinating prospects. This review (with 118 refs.) summarizes the progress made in this field in the recent 10 years. Following an introduction into the fundamentals of NPA, we demonstrate its potential by describing selected methods for sensing (a) proteins such as streptavidin, certain antibodies, or thrombin via aptamers; (b) oligomers, larger nucleic acids, or micro-RNA; (c) small molecules, (d) ions such as K(I) which is vital to the maintenance of life, or Hg(II) which is dangerous to health. We summarize the results and discuss the merits and limitations of the various methods at last.

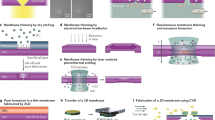
Schematic of a signal-off system and a signal-on system in nanopore analysis. The effective diameter of nanopores decreases when targets undergo certain interactions with receptors attached on the inner surface of the nanopore. Correspondingly, the current will drop on appearance of the analyte. This is referred to as a “signal-off” system. Conversely, it is called a “signal-on” system.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biological ion channels embed in biological cell to communicate the matter and energy with the extracellular world [1]. They play a crucial role in various significant physiological activities [2–4]. The function of biological ion channels has allured a lot of attention from both scientists and engineers. They fabricate solid-state artificial nanopores to mimic functions of biological ion channels [5–9]. Except for many similar functions of the biological ion channels, synthetic nanopores such as carbon nanotubes [10–13], silicon-based nanopores [14–20], graphene nanopores [21–24], and polymeric nanopores [25–27] possess ascendances, multi-functions and stability. Benefit by these characteristics, many applications including sensing [28–32], energy conversion [33–35], nanofluidic circuits [36, 37] and filtration [38–40] are possible to achieve. Among all these potential applications, we focus mainly on the nanopore-based analysis (NPA) for their applications in detection of the biochemical species in last 10 years.
The principle of NPA can be described briefly: molecules access in or attach on the surface of a pore, thereby leading the ionic current changes can be detected [41]. The nanopore membrane is located in the middle of two electrochemical chambers separated into cis- and trans- compartments, each containing conducting buffers. Under an applied voltage, electrolyte ions flow through the nanopore, which is measured as current in the electrical instrument. The effective diameter of the nanopore will be decreased when targets have certain interaction with receptors which are attached on the inner surface of the nanopore. Correspondingly, the current signal will drop along with the appearance of the target; we call it signal off system. Conversely, the effective diameter of nanopores will be increased when targets change the structure of receptor or come out of the nanopore. Correspondingly, the current signal will rise along with the appearance of the target; we call it signal on system (Fig. 1). Using nanopores in sensing of biomolecules has distinct advantages; for example, one can detect analytes via their size [42], shape [43] or charge [44]. The range of analytes that can be detected with nanopores now spans peptides, proteins, bimolecular complexes, enzymes, organic polymers and small molecules [45]. In this review, we discuss the application of the NPA in detection of nucleic acids, proteins, small molecules and ions. We end with a brief conclusion of the advantages and challenges of the NPA.
Schematic of the signal off system and signal on system illustrates the principle of the NPA. The effective diameter of nanopores will be decreased when targets have certain interaction with receptors which are attached on the inner surface of the nanopore. Correspondingly, the current signal will drop along with the appearance of the target; we call it signal off system. Conversely, we call it signal on system
The NPA for detection of biochemical species
Detection of proteins
The NPA technology develops in an application for detection of protein [46–48]. Siwy et al. fabricate a single conical gold nanopore with biochemical molecular-recognition agent (MRA) as a protein biosensor [49]. They investigate three MRA/analyte systems including the biotin/streptavidin, protein-G/immunoglobulin (IgG), and an antibody to the protein ricin as the MRA and ricin as the analyte. For example the diameter of the streptavidin (SA) molecule (~5 nm), and the final diameter after the MRA modified is ~5 nm for SA sensor. Because the size of protein molecule is comparable with the nanopore diameters, when the protein is recognized, the effective diameter of nanopore decreases. This is the signal off system. Current–voltage (I–V) curves for the biotinylated nanopore after exposure to two negative control proteins 100 nM lysozyme and BSA, the signal show little change, indicating that the sensor does not respond to proteins that do not recognize by the biotin MRA. In contrast, the ionic current drop substantially after immersed to a solution with 180 pM streptavidin (Fig. 2a). They use the time required for blockage, τ b, to determine the analyte concentration. The IgG concentrations in 100–10 nM range can be detected. Chen group also use the MRA to detect a variety of biotin binding proteins by modified an OmpG nanopore with a biotinylated PEG molecule [50].
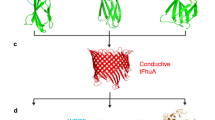
a Current–voltage curves for the streptavidin sensor are presented in the presence of no protein (×), 100 nM lysozyme (♦), and 180 pM SA (▲) [49]. Copyright © 2005, American Chemical Society b A single asymmetric nanopore functionalized with biorecognition elements (biotin-PEO3-amine) is used for streptavidin analytics [51]. Copyright © 2008, American Chemical Society c Artificial nanopores are applied to analyze the antibodies. His6-tagged protein A is immobilized on the surface of trisNTA-modified nanopores. IgG antibodies are identified because of their specific interaction times with protein A receptors. The species and subclass the antibodies belong to can also be detected in the system [56]. d An α-hemolysin pore functioned by a DNA aptamer detects thrombin [57]. Copyright © 2012, Rights Managed by Nature Publishing Group
Ali et al. report a facile strategy based on the electrostatically assemble biorecognition with the target into conical nanopores for constructing a nanobiosensor. In this system the target SA (∼5 nm) and the dimension of the nanopore (∼8 nm) are comparable in size and the detection limit of SA is 1 pM (Fig. 2b) [51–53]. Maglia et al. evolve Cytolysin A nanopore, which can be isolated into three nanopore types. The three ClyA nanopores with different diameter (33, 37, 42 Å) allowed the selective entry of proteins inside the nanopore [54, 55].
Recently, Wei et al. [56] detect single protein by using solid-state artificial nanopores modified with biological receptors (Fig. 2c). In their design, gold-coated silicon nitride nanopores functionalized with adequately few multivalent nitrilotriacetic acid (NTA) groups. By adjusting the ratio of NTA and the fabrication of ethylene glycol which can couple to NTA, only a single NTA tag can be controlled to fabricate on the nanopore. They use the NTA receptor, which act as binding sites for His-tagged proteins. When nanopores are immersed in the solution with His-tagged protein A, the current across it changes from an empty pore to a pore transiently blocked by one protein A molecule, or back-up process. The authors design another nanopore platform that is used to detect IgG. In this system, His6-tagged protein A is stably immobilized within a trisNTA-modified pore. IgG antibodies (=ligands) interact with the His6-tagged protein A, are detected by the resistive pulse technique. This nanopore sensor which based on how long the pore is blocked could detect single molecules of IgG and distinguish between various IgG subtypes from different organisms. Unbinding of His-tagged protein A was not detected. To control the concentration and geometry of receptors, Rotem et al. [57] also achieve a sensitivity detection of protein by using a α-hemolysin pore (Fig. 2d). Other groups use nanopores to study protein translation [58, 59], DNA-protein interactions [60, 61], and proteins folding [62, 63].
The NPA for detection of proteins has traits in common. (1) The nanopore mouth dimension and the target are comparable in size. (2) By choosing or tuning the receptors which can recognize or bind the specific proteins leads to the detection with high specificity. (3) When the proteins enter into the nanopores, combine with the receptor, lead to the decrease of the effective diameter, and the ion current decreases. These are signal off systems. NPA application in detection of protein can achieve single-molecule sensitivity because the diameter of nanopore can be controlled accurately. Compared with other certain protein detection technologies, the NPA offers a simple detection method relying on the electrical read-out to downstream signal processing, which avoiding complex operations.
Detection of nucleic acids
The NPA is firstly applied possibility to rapidly sequencing DNA [64–66]. In 1996 year, a group of scientists made a discovery that the ion channel could in principle provide direct, high-speed detection of the sequence of bases in single molecules of DNA or RNA [41]. They use an electric field to drive single-stranded DNA and RNA molecules through a pore-forming protein and detect the signal of ionic current in nanopores [67, 68] (Fig. 3a). This system uses the Staphylococcus aureus toxin, α-hemolysin (α-HL) (Fig. 3b), the use of which as a biosensor is pioneered by Bayley and his coworkers [69–73]. Both Ghadiri group and Akeson group show that polymerase enzymes can be used to move DNA across the α-HL nanopores [74–76]. Gundlach group introduced the MspA pore and showed the convincing sequencing data [77, 78]. Maglia et al. use modified ClyA nanopore to recognize and chaperone DNA [79]. Long group modified the protein nanopores as biomolecular sensors [80–83]. These experiments dawn some conclusions. For example, Contrast on polyC, polyA can block the nanopore a greater degree; the nanopore has not a clear distinct between purine and pyrimidine ribonucleotides. The order of the nucleobases in a polynucleotide can be detected by the signal changes of ion current though nanopores. Both kilo-base length polymers (single-stranded genomic DNA or RNA) and small molecules (e.g., nucleosides) can be identified and characterized without amplification or labeling. The NPA offers a unique analytical capability that makes inexpensive, rapid DNA sequencing possible [84, 85].
a The nanopore is used in strand-sequencing detection. The ionic current amplitude through a α-hemolysin nanopore changes in the two statuses, an open pore and a blocked one by a single-strand DNA. The red bracket means this system cannot distinguish the ~12 nucleotides [67]. Copyright © 2008, Rights Managed by Nature Publishing Group b Side view of α-hemolysin pore from Staphylococcus aureus is presented [69]. Copyright © 1996, American Association for the Advancement of Science c The specific miRNA is detected using solid-state 3-nm-diameter nanopore in a 7-nm-thick membrane [82]. Copyright © 2010, Nature Publishing Group
Artificial nanopores synthesized by materials such as silicon nitride have superiority to be detected of molecules. The capabilities of these sensors are influenced by both the diameter of the nanopore and the thickness of the membrane. Wanunu et al. show a sub-micrometer area of a SiN membrane with thicknesses as small as 6 nm provides a detection platform for the RNA molecules, which without the time-consuming labelling or amplification methods [86]. Their experiments show that reducing the thickness of the membrane can increase signal amplitudes from biomolecules. Nucleic acids with as few as ten base pairs can be detected by using 3-nm-diameter nanopores in sub-10-nm thick membranes. The various short nucleic acids with similar molecular weights are also discriminated on differences in their physical dimensions. They detect the miR122a which from 1 mg of rat liver total RNA using by a 3-nm-diameter nanopore in 7-nm-thick membrane. Target microRNA is first hybridized to a probe. This probe/target duplex is then enriched through binding to the viral protein. From the measurement data, the concentration of miR122a dilute 20-fold in sample is 0.7 fmol/μL, translate 78 ± 2 pg miR122a/mg liver RNA in rat liver cells. Gu et al. use one nanopore to detect of multiple miRNA simultaneously [87]. When it comes to the thickness of nanopore, the graphene nanopores have to mentioned because of it has one atom thickness. Graphene nanopores show great potential for the detection of DNA sequencing rather than other solid-state artificial nanopores [88–98]. The nanopore provides a highly confined space which made the nucleic acids analyzed at high throughput [99–101]. However, such thin membranes significantly limit the surface of the pore available to interact with DNA, leading to the translocation speeds of DNA in nanopores is too fast. Thus, electrical signal cannot be resolved with sufficient accuracy in the detection system [102, 103].
To regulate the translocation speeds of nucleic acid might be one of the most important factors in detection of DNA or RNA sequencing use solid-state artificial nanopores. During detection of nucleic acids sequencing both signals off system (nucleic acids enter into nanopores) and signal on system (nucleic acids come out of nanopores) are used together. Compared with the proteins, the size of nucleic acids are smaller which requires smaller and special character of nanopores, such as protein nanopores and graphene nanopores. Furthermore, taking advantage of nucleic acids are designable and modified, the NPA can combined with conventional characterization techniques such as gel electrophoresis, atomic force microscopy, transmission electron microscopy and laser scanning confocal microscopy. The amplification method such as the interaction with protein can improve the detection limit of nucleic acids by the NPA technology.
Detection of small molecules
The NPA technology can be used to detect and select small molecules by binding them in nanopores. Bayley H et al. equipped α- hemolysin nanopore with cyclodextrins to detect of organic molecules [104]. Kasianowicz and Bezrukov group studied the interaction between polymer molecules and protein nanopores [105–107]. Both of them examine the blockages of ion current when analysts through nanopores, which belongs to signal off system. Till now, small molecules and nucleic acids cannot be analyzed simultaneously in a nanopore sensor. Xia et al. have found that a more complex DNA nanostructure can be introduced to the nanopore [31]. The complex DNA nanostructure contains multiple target-binding sites on each of its long concatamers and provides a built-in amplification mechanism (Fig. 4a). This nanopore is prepared from poly(ethyleneterephthalate) membranes. It has a diameter of 79 ± 7 nm. When target nucleic acids exist, the DNA supersandwich structures are assembled to decrease the effective diameter of the nanopore; while ATP exists, the DNA supersandwich structures are disassembled to increase the effective diameter of the nanopore, that means both the signal off and signal on system in this nanopore detection platform. This nanopore sensor has many advantages, such as enhancing signal intensity, a better detection limit (the detection limit of DNA is 10 fM and ATP is 1 nM), and anti-interference capability. More importantly, it can be used to analysis single-base mismatch resolution and discrimination among different types of nucleoside triphosphates. It could also be used in complex matrices when the interfering substances concentration is high in the buffer solution. It even can be directly used in serum.
a The scheme of two-way nanopore sensor. A more complex DNA nanostructure (supersandwich structures) can be introduced to the nanopore. It shows high sensitive to DNA or ATP. The detection limt of DNA is 10 fM and ATP is 1 nM. More importantly, it could also be used in complex matrices when the interfer substances concentration is high in the buffer solution. It even can be directly used in serum [31]. Copyright © 2013 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim b The scheme of β-CD modified nanopore system. It is responsive to L-His and induce a large difference in the enantiomeric ionic currents which makes it can be used in practical application [108]. Copyright © 2011, American Chemical Society
Li et al. report a simple enantioselective device. They modify a single artificial β-cyclodextrin (β-CD) to the single conical nanopore system (Fig. 4b) [108]. Putting this β-CD-modified nanopore into a solution of L-His, they find a decrease of the transmembrane ionic current due to the selective binding of L-His to the nanopore wall that is occurs inside the confined geometry. While, immersing this β-CD-modified nanopore in solutions of D-His or other aromatic amino acids, no significant changes of ionic current are found. It shows that this β-CD-modified nanopore has high selective recognition of histidine enantiomers by monitoring of ionic current signatures. They also find that the ionic currents decrease gradually with increasing L-His concentration from 0 to 1 mM.
There are some characteristics of the NPA using in the detection of small molecules. (1) With the size of small molecules down, the detection by binding event direct which require the smaller diameter of nanopores (most of protein nanopores); (2) Detecting of small molecules using the NPA, the signal changes are inconspicuous without the amplified technology. (3) Statistical analysis of each blockage should be investigated in nanopore experiments [109]. (4) The NPA detection platform is expected to develop into a live assay for disease related molecular targets, and with many practical applications in biotechnology and life science.
Detection of ions
Jiang et al. report a potassium-responsive nanopore (Fig. 5a) [110]. It is mainly rely on the conformational changes of the G4 DNA chains in the presence of potassium ions (K+). The reason is that the structure transition of the G4 DNA chains from loose packing to the i-motif structure after binding with K+. Thus the effective pore sizes decrease which leading to the ionic current drop. As shown in Fig. 5b, this nanopore/DNA hybrid system has an ion concentration effect that provides a nonlinear response to K+ at the concentration ranging from 0 to 1500 μM. They also construct a biomimetic zinc activated ion channel by introducing the zinc fingers into the nanopores (Fig. 5c) [111]. This nanopore is responsive to zinc ions. In the presence of zinc ions, the zinc fingers fold into finger like conformations, thus the effective diameter of the channel increase. In turn, the biomimetic ion channel is activated. However, in the absence of zinc, they find a low ion conductance. This nanopore has high specificity. As shown in Fig. 5d, it is not responsive to other metal ions. If introduce a T-rich ssDNA to the nanopores, it can construct a biomimetic mercury (II)-gated nanopore by forming a stable T-Hg2+-T complex [112]. Cu2+ also is detected by peptide conformational changes in nanopores [113].
a A single nanopore is fabricated by conformational transition of G4 DNA which can response to K+ [110]. Copyright © 2009 American Chemical Society b The current data of the NPA system. The blue line is the absence of G4 DNA modification. The red line is the presence of G4 DNA modification. The green line is the addition of the complementary DNA strands. c The scheme of biomimetic zinc activated ion channel. d The histogram of the nanopore. It shows the specificity of the system [111]. Copyright © 2010 Royal Society of Chemistry
The dimension of ions is the smallest among targets, which means that they can only be detected by indirect analysis, such as the interactions with nanopores, the conformation transition between ions and the nucleic acids or polypeptide et al. The conformational change induces the change in the effective size of nanopores and these are either signal on or signal off system. The key features of the NPA using for ions detection is closely imitate these molecular interactions happening in living organisms.
Summary
The advantages of the NPA for detection of biochemical species are as follows:
-
(1)
The NPA has high sensitivity; even the single molecule can be detected.
-
(2)
The NPA has high specificity; the target can be detected from the analogues or in the presence of interfering substance.
-
(3)
The NPA for target detection requires very low sample volumes and without sample complicated preparation.
-
(4)
Study the dynamics of interaction between receptors and targets in nanopores by the patch clamp technique can promote the understanding of the molecular mechanism.
-
(5)
The NPA has the broad analytical range. In addition to the biomolecules, small molecules and ions, other molecules such as nanoparticles [114], inorganic molecules [115–117] and organic molecules [118] can also be detected.
There are also many challenges in application the NPA for detection of biochemical species.
-
(1)
The limit of materials and dimensions cause that the NPA for detection platform can only be used in vitro.
-
(2)
How to further improve the NPA technology for detection of targets specificity. Especially for proteins which have complex spacial structures, the specificity is even more important in bimolecular analysis.
Considered that with the development of nanotechnique, the fabrication of nanopores will be more diverse and accurate. The NPA as a new kind of detection methods arouses widespread interest for its incredibly merits and wide application. It presents improved capabilities for the area of single molecule detection, discriminating molecules with different configuration and mimicking the transmembrane protein features. The NPA is expected that the simple electronic device fabricated with high sensitivity and specificity, which can be used in practical life, such as clinic diagnostics, routine laboratory detection, food safety, and environmental monitoring.
References
Heginbotham L, Abramson T, MacKinnon R (1992) A functional connection between the pores of distantly related ion channels as revealed by mutant K+ channels. Science 258:1152–1155
Eisenberg B (1998) Ionic channels in biological membranes: natural nanotubes. Acc Chem Res 31:117–123
Perozo E, Cortes DM, Sompornpisut P, Kloda A, Martinac B (2002) Open channel structure of MscL and the gating mechanism of mechanosensitive channels. Nature 418:942–948
Beckstein O, Biggin PC, Bond P, Bright JN, Domene C, Grottesi A, Holyoake J, Sansom M (2003) Ion channel gating: insights via molecular simulations. FEBS Lett 555:85–90
Bayley H, Martin CR (2000) Resistive-pulse sensings from microbes to molecules. Chem Rev 100:2575–2594
Gyurcsányi RE (2008) Chemically-modified nanopores for sensing. TrAC Trends Anal Chem 27:627–639
Reiner JE, Balijepalli A, Robertson JW, Campbell J, Suehle J, Kasianowicz JJ (2012) Disease detection and management via single nanopore-based sensors. Chem Rev 112:6431–6451
Hernandez-Ainsa S, Keyser UF (2014) DNA origami nanopores: developments, challenges and perspectives. Nanoscale 6:14121–32
Taniguchi M (2015) Selective multi-detection using nanopores. Anal Chem 87:188–199
Bachtold A, Hadley P, Nakanishi T, Dekker C (2001) Logic circuits with carbon nanotube transistors. Science 294:1317–1320
Siwy ZS, Davenport M (2010) Biosensors: making nanopores from nanotubes. Nat Nanotechnol 5:174–175
Wang MS, Chen Q, Peng LM (2008) Field-emission characteristics of individual carbon nanotubes with a conical Tip: the validity of the fowler–Nordheim theory and maximum emission current. Small 4:1907–1912
Harrell CC, Lee SB, Martin CR (2003) Synthetic single-nanopore and nanotube membranes. Anal Chem 75:6861–6867
Ho C, Qiao R, Heng JB, Chatterjee A, Timp RJ, Aluru NR, Timp G (2005) Electrolytic transport through a synthetic nanometer-diameter pore. Proc Natl Acad Sci U S A 102:10445–10450
Li J, Stein D, McMullan C, Branton D, Aziz MJ, Golovchenko JA (2001) Ion-beam sculpting at nanometre length scales. Nature 412:166–169
Yossifon G, Chang YC, Chang HC (2009) Rectification, gating voltage, and interchannel communication of nanoslot arrays due to asymmetric entrance space charge polarization. Phys Rev Lett 103:154502
Storm AJ, Chen JH, Ling XS, Zandbergen HW, Dekker C (2003) Fabrication of solid-state nanopores with single-nanometre precision. Nat Mater 2:537–540
Wei C, Bard AJ, Feldberg SW (1997) Current rectification at quartz nanopipet electrodes. Anal Chem 69:4627–4633
Wu MY, Smeets RM, Zandbergen M, Ziese U, Krapf D, Batson PE, Dekker NH, Dekker C, Zandbergen HW (2009) Control of shape and material composition of solid-state nanopores. Nano Lett 9:479–484
Zhang B, Zhang YH, White HS (2004) The nanopore electrode. Anal Chem 76:6229–6238
Siwy ZS, Davenport M (2010) Nanopores: graphene opens up to DNA. Nat Nanotechnol 5:697–698
Garaj S, Hubbard W, Reina A, Kong J, Branton D, Golovchenko JA (2010) Graphene as a subnanometre trans-electrode membrane. Nature 467:190–193
Merchant CA, Healy K, Wanunu M, Ray V, Peterman N, Bartel J, Fischbein MD, Venta K, Luo Z, Johnson AT, Drndic M (2010) DNA translocation through graphene nanopores. Nano Lett 10:2915–2921
Schneider GF, Kowalczyk SW, Calado VE, Pandraud G, Zandbergen HW, Vandersypen LM, Dekker C (2010) DNA translocation through graphene nanopores. Nano Lett 10:3163–3167
Apel P (2001) Track etching technique in membrane technology. Radiat Meas 34:559–566
Siwy ZS (2003) Preparation of synthetic nanopores with transport properties analogous to biological channels. Surf Sci 532–535:1061–1066
Chantiwas R, Park S, Soper SA, Kim BC, Takayama S, Sunkara V, Hwang H, Cho YK (2011) Flexible fabrication and applications of polymer nanochannels and nanoslits. Chem Soc Rev 40:3677–3702
Howorka S, Siwy ZS (2009) Nanopore analytics: sensing of single molecules. Chem Soc Rev 38:2360–2384
Tian Y, Wen L, Hou X, Hou G, Jiang L (2012) Bioinspired ion-transport properties of solidstate single nanochannels and their applications in sensing. ChemPhysChem 13:2455–2470
Sexton LT, Horne LP, Martin CR (2007) Developing synthetic conical nanopores for biosensing applications. Mol Bio Syst 3:667–685
Liu NN, Jiang YN, Zhou YH, Xia F, Guo W, Jiang L (2013) Two-Way nanopore sensing of sequence-specific oligonucleotides and small-molecule targets in complex matrices using integrated DNA supersandwich structures. Angew Chem Int Ed 52:1842–2007
Xia F, Guo W, Mao YD, Hou X, Xue J, Xia H, Wang L, Song Y, Ji H, Ouyang Q, Wang Y, Jiang L (2008) Gating of single synthetic nanopores by proton-driven DNA molecular motors. J Am Chem Soc 130:8345–8350
Xie YB, Wang XW, Xue JM, Jin K, Chen L, Wang YG (2008) Electric energy generation in single track-etched nanopores. Appl Phys Lett 93:163116–163119
Wen L, Hou X, Tian Y, Zhai J, Jiang L (2010) Bio-inspired photoelectric conversion based on smart-gating nanochannels. Adv Funct Mater 20:2636–2642
Guo W, Cao LX, Xia JC et al (2010) Energy harvesting with single-Ion-selective nanopores: a concentration gradient driven nanofluidic power source. Adv Funct Mater 20:1339–1344
Jiang YN, Liu NN, Guo W, Xia F, Jiang L (2013) Highly-efficient gating of solid-state nanochannels by DNA supersandwich structure containing ATP aptamers: a nanofluidic IMPLICATION logic device. J Am Chem Soc 134:15395–15401
Vlassiouk I, Kozel TR, Siwy ZS (2009) Biosensing with nanofluidic diodes. J Am Chem Soc 131:8211–8220
Martin CR, Siwy ZS (2004) Molecular filters—pores within pores. Nat Materm 3:284–285
Calvo A, Yameen B, Williams FJ, Soler-Illia GJ, Azzaroniet O (2009) Mesoporous films and polymerbrushes helping each other to modulate ionic transport in nanoconfined environments: an interesting example of synergism in functional hybrid assemblies. J Am Chem Soc 131:10866–10868
Vlassiouk I, Smirnov S, Siwy ZS (2008) Ionic selectivity of single nanochannels. Nano Lett 8:1978–1985
Kasianowicz J, Brandin E, Branton D, Deamer DW (1996) Characterization of individual polynucleotide molecules using a membrane channel. Proc Natl Acad Sci U S A 93:13770–13773
Heng JB, Ho C, Kim T, Timp R, Aksimentiev A, Grinkova YV, Sligar S, Schulten K, Timp G (2004) Sizing DNA using a nanometer-diameter pore. Biophys J 87:2905–2911
Wang H, Dunning JE, Huang AP, Nyamwanda JA, Branton D (2004) DNA heterogeneity and phosphorylation unveiled by single-molecule electrophoresis. Proc Natl Acad Sci U S A 101:13472–13477
Cervera J, Schiedt B, Neumann R, Mafé S, Ramírez P (2006) Ionic conduction, rectification, and selectivity in single conical nanopores. J Chem Phys 124:104706
Zhao Q, Wang D, Jayawardhana DA, Guan X (2008) Stochastic sensing of biomolecules in a nanopore sensor array. Nanotechnology 19:505504
Howorka S, Siwy ZS (2012) Nanopores as protein sensors. Nat Biotechnol 30:506–507
Oukhaled A, Bacri L, Pastoriza-Gallego M, Betton JM, Pelta J (2012) Sensing proteins through nanopores: fundamental to applications. ACS Chem Biol 7:1935–1949
Li W, Bell NA, Hernández-Ainsa S, Thacker VV, Thackray AM, Bujdoso R, Keyser UF (2013) Single protein molecule detection by glass nanopores. ACS Nano 7:4129–4134
Siwy Z, Trofin L, Kohli P, Baker LA, Trautmann C, Martin CR (2005) Protein biosensors based on biofunctionalized conical gold nanotubes. J Am Chem Soc 127:5000–5001
Fahie M, Chisholm C, Chen M (2015) Resolved single-molecule detection of individual species within a mixture of anti-biotin antibodies using an engineered monomeric nanopore. ACS Nano 9:1089–1098
Ali M, Yameen B, Neumann R, Ensinger W, Knoll W, Azzaroni O (2008) Biosensing and supramolecular bioconjugation in single conical polymer nanochannels. Facile incorporation of biorecognition elements into nanoconfined geometries. J Am Chem Soc 130:16351–16357
Ali M, Schiedt B, Neumann R, Ensinger W (2010) Biosensing with functionalized single asymmetric polymer. Nanochannels Macromol Biosci 10:28–32
Ali M, Nasir S, Ensinger W (2015) Bioconjugation-induced ionic current rectification in aptamer-modified single cylindrical nanopores. Chem Commun 51:3454–3457
Soskine M, Biesemans A, De Maeyer M, Maglia G (2013) Tuning the size and properties of ClyA nanopores assisted by directed evolution. J Am Chem Soc 135:13456–13463
Van Meervelt V, Soskine M, Maglia G (2014) Detection of two isomeric binding configurations in a protein-aptamer complex with a biological nanopore. ACS Nano 8:12826–12835
Wei R, Gatterdam V, Wieneke R, Tampe R, Rant U (2012) Stochastic sensing of proteins with receptor-modified solid-state nanopores. Nat Nanotechnol 7:257–263
Rotem D, Jayasinghe L, Salichou M, Bayley H (2012) Protein detection by nanopores equipped with aptamers. J Am Chem Soc 134:2781–2787
Yusko EC, Johnson JM, Majd S, Prangkio P, Rollings RC, Li J, Yang J, Mayer M (2011) Controlling protein translocation through nanopores with bio-inspired fluid walls. Nat Nanotechnol 6:253–260
Nivala J, Marks DB, Akeson M (2013) Unfoldase-mediated protein translocation through an α-hemolysin nanopore. Nat Biotechnol 31:247–250
Hornblower B, Coombs A, Whitaker RD, Kolomeisky A, Picone SJ, Meller A, Akeson M (2007) Single-molecule analysis of DNA-protein complexes using nanopores. Nat Methods 4:315–317
Liu NN, Yang ZK, Lou XD, Wei BM, Zhang JT, Gao PC, Hou RZ, Xia F (2015) Nanopore-based DNA-probe sequence-evolution method unveiling characteristics of protein–DNA binding phenomena in a nanoscale confined space. Anal Chem. doi:10.1021/acs.analchem.5b00375
Talaga DS, Li J (2009) Single-molecule protein unfolding in solid state nanopores. J Am Chem Soc 131:9287–9297
Rodriguez-Larrea D, Bayley H (2013) Multistep protein unfolding during nanopore translocation. Nat Nanotechnol 8:288–295
Rostovtseva TK, Bezrukov SM (1998) ATP transport through a single mitochondrial channel, VDAC, studied by current fluctuation analysis. Biophys J 74:2365–2373
Venkatesan B, Bashir R (2011) Nanopore sensors for nucleic acid analysis. Nat Nanotechnol 6:615–624
Cherf GM, Lieberman KR, Lam CE, Karplus K, Akeson M (2012) Automated forward and reverse ratcheting of DNA in a nanopore at 5-Å precision. Nat Biotechnol 30:344–348
Branton D, Deamer DW, Marziali A, Bayley H, Benner SA, Butler T, Di Ventra M, Garaj S, Hibbs A, Huang X, Jovanovich SB, Krstic PS, Lindsay S, Ling XS, Mastrangelo CH, Meller A, Oliver JS, Pershin YV, Ramsey JM, Riehn R, Soni GV, Tabard-Cossa V, Wanunu M, Wiggin M, Schloss JA (2008) The potential and challenges of nanopore sequencing. Nat Biotechnol 26:1146–1153
Deamer DW, Akeson M (2000) Nanopores and nucleic acids: prospects for ultrarapid sequencing. Trends Biotechnol 18:147–151
Song L, Hobaugh M, Shustak C, Cheley S, Bayley H, Gouaux JE (1996) Structure of staphylococcal α-hemolysin, a heptameric transmembrane pore. Science 274:1859–1865
Howorka S, Movileanu L, Braha O, Bayley H (2001) Kinetics of duplex formation for individual DNA strands within a single protein nanopore. Proc Natl Acad Sci U S A 98:12996–13001
Stoddart D, Heron AJ, Mikhailova E, Maglia G, Bayley H (2009) Single-nucleotide discrimination in immobilized DNA oligonucleotides with a biological nanopore. Proc Natl Acad Sci U S A 106:7702–7707
Clarke J, Wu HC, Jayasinghe L, Patel A, Reid S, Bayley H (2009) Continuous base identification for single-molecule nanopore DNA sequencing. Nat Nanotechnol 4:265–270
Bayley H, Cremer PS (2001) Stochastic sensors inspired by biology. Nature 413:226–230
Ashkenasy N, Sanchez-Quesada J, Bayley H, Ghadiri MR (2005) Recognizing a single base in an individual DNA strand: a step toward DNA sequencing in nanopores. Angew Chem Int Ed 44:1401–1404
Cockroft SL, Chu J, Amorin M, Ghadiri MR (2008) A single-molecule nanopore device detects DNA polymerase activity with single-nucleotide resolution. J Am Chem Soc 130:818–820
Lieberman KR, Cherf GM, Doody MJ, Olasagasti F, Kolodji Y, Akeson M (2010) Processive replication of single DNA molecules in a nanopore catalyzed by phi29 DNA polymerase. J Am Chem Soc 132:17961–17972
Butler TZ, Pavlenok M, Derrington IM, Niederweis M, Gundlach JH (2008) Single-molecule DNA detection with an engineered MspA protein nanopore. Proc Natl Acad Sci U S A 105:20647–20652
Manrao EA, Derrington IM, Laszlo AH, Langford KW, Hopper MK, Gillgren N, Pavlenok M, Niederweis M, Gundlach JH (2012) Reading DNA at single-nucleotide resolution with a mutant MspA nanopore and phi29 DNA polymerase. Nat Biotechnol 30:349–353
Franceschini L, Soskine M, Biesemans A, Maglia G (2013) A nanopore machine promotes the vectorial transport of DNA across membranes. Nat Commun 4:2415–2422
Wang HY, Gu Z, Cao C, Wang J, Long YT (2013) Analysis of a single alpha-synuclein fibrillation by the interaction with a protein nanopore. Anal Chem 85:8254–8261
Ying YL, Cao C, Long YT (2014) Single molecule analysis by biological nanopore sensors. Analyst 139:3826–3835
Ying YL, Zhang J, Gao R, Long YT (2013) Nanopore-based sequencing and detection of nucleic acids. Angew Chem Int Ed 52:13154–13161
Ying YL, Li DW, Liu Y, Dey SK, Kraatz HB, Long YT (2012) Recognizing the translocation signals of individual peptide-oligonucleotide conjugates using an alpha-hemolysin nanopore. Chem Commun 48:8784–8786
Braha O, Walker B, Cheley S (1997) Designed protein pores as components for biosensors. Chem Biol 4:497–505
Meller A, Nivon L, Brandin E, Golovchenko J, Branton D (2000) Rapid nanopore discrimination between single oligonucleotide molecules. Proc Natl Acad Sci USA 97:1079–1084
Wanunu M, Dadosh T, Ray V, Jin J, McReynolds L, Drndic M (2010) Rapid electronic detection of probe-specific microRNAs using thin nanopore sensors. Nat Nanotechnol 5:807–817
Zhang XY, Wang Y, Fricke BL, Gu LQ (2014) Programming nanopore Ion flow for encoded multiplex MicroRNA detection. ACS Nano 8:3444–3450
Postma HW (2010) Rapid sequencing of individual DNA molecules in graphene nanogaps. Nano Lett 10:420–425
Nelson T, Zhang B, Prezhdo OV (2010) Detection of nucleic acids with graphene nanopores: ab initio characterization of a novel sequencing device. Nano Lett 10:3237–3242
Avdoshenko SM, Nozaki D, Gomes da Rocha C, Gonzalez JW, Lee MH, Gutierrez R, Cuniberti G (2013) Dynamic and electronic transport properties of DNA translocation through graphene nanopores. Nano Lett 13:1969–1976
Wells DB, Belkin M, Comer J, Aksimentiev A (2012) Assessing graphene nanopores for sequencing DNA. Nano Lett 12:4117–4123
Sathe C, Zou X, Leburton JP, Schulten K (2011) Computational investigation of DNA detection using graphene nanopores. ACS Nano 5:8842–8851
Traversi F, Raillon C, Benameur SM, Liu K, Khlybov S, Tosun M, Krasnozhon D, Kis A, Radenovic A (2013) Detecting the translocation of DNA through a nanopore using graphene nanoribbons. Nat Nanotechnol 8:939–945
Larkin J, Henley R, Bell DC, Cohen-Karni T, Rosenstein JK, Wanunu M (2013) Slow DNA transport through nanopores in hafnium oxide membranes. ACS Nano 7:10121–10128
Garaj S, Liu S, Golovchenko JA, Branton D (2013) Molecule-hugging graphene nanopores. Proc Natl Acad Sci U S A 110:12192–12196
Lv W, Liu S, Li X, Wu R (2014) Spatial blockage of ionic current for electrophoretic translocation of DNA through a graphene nanopore. Electrophoresis 35:1144–1151
Banerjee S, Wilson J, Shim J, Shankla M, Corbin EA, Aksimentiev A, Bashir R (2015) Slowing DNA transport using graphene-DNA interactions. Adv Funct Mater 25:936–946
Lv W, Chen M, Wu RA (2013) The impact of the number of layers of a graphene nanopore on DNA translocation. Soft Matter 9:960–966
Bissels U, Wild S, Tomiuk S, Holste A, Hafner M, Tuschl T, Bosio A (2009) Absolute quantification of microRNAs by using a universal reference. RNA 15:2375–2384
Chen P, Mitsui T, Farmer DB, Golovchenko J, Gordon RG, Branton D (2004) Atomic layer deposition to fine-tune the surface properties and diameters of fabricated nanopores. Nano Lett 4:1333–1337
Lqbal SM, Akin D, Bashir R (2007) Solid-state nanopore channels with DNA selectivity. Nat Nanotechnol 2:243–248
Smeets RMM, Keyser UF, Dekker NH, Dekker C (2008) Noise in solid-state nanopores. Proc Natl Acad Sci U S A 105:417–421
Squires AH, Hersey JS, Grinstaff MW, Meller A (2013) A nanopore-nanofiber mesh biosensor to control DNA translocation. J Am Chem Soc 135:16304–16307
Gu LQ, Braha O, Conlan S, Cheley S, Bayley H (1999) Stochastic sensing of organic analytes by a pore-forming protein containing a molecular adapter. Nature 398:686–690
Reiner JE, Kasianowicz JJ, Nablo BJ, Robertson JWF (2010) Theory for polymer analysis using nanopore-based single-molecule mass spectrometry. Proc Natl Acad Sci U S A 107:12080–12085
Balijepalli A, Robertson JW, Reiner JE, Kasianowicz JJ, Pastor RW (2013) Theory of polymer-nanopore interactions refined using molecular dynamics simulations. J Am Chem Soc 135:7064–7072
Krasilnikov O, Rodrigues C, Bezrukov S (2006) Single polymer molecules in a protein nanopore in the limit of a strong polymer-pore attraction. Phys Rev Lett 97:018301–018304
Han CP, Hou X, Zhang HC, Guo W, Li H, Jiang L (2011) Enantioselective recognition in biomimetic single artificial nanochannels. J Am Chem Soc 133:7644–7647
Gu Z, Ying YL, Cao C, He P, Long YT (2015) Accurate data process for nanopore analysis. Anal Chem 87:907–913
Hou X, Guo W, Xia F, Nie FQ, Dong H, Tian Y, Wen L, Wang L, Cao L, Yang Y, Xue J, Song Y, Wang Y, Liu D, Jiang L (2009) A biomimetic potassium responsive nanochannel: G-quadruplex DNA conformational switching in a synthetic nanopore. J Am Chem Soc 131:7800–7805
Tian Y, Hou X, Wen LP, Guo W, Song Y, Sun H, Wang Y, Jiang L, Zhu D (2010) A biomimetic zinc activated ion channel. Chem Commun 46:1682–1684
Tian Y, Zhang Z, Wen LP, Ma J, Zhang Y, Liu W, Zhai J, Jiang L (2013) A biomimetic mercury (II)-gated single nanochannel. Chem Commun 49:10679–10681
Mereuta L, Schiopu I, Asandei A, Park Y, Hahm KS, Luchian T (2012) Protein nanopore-based, single-molecule exploration of copper binding to an antimicrobial-derived, histidine-containing chimera peptide. Langmuir 28:17079–17091
Davenport M, Healy K, Pevarnik M, Ni T, Cabrini S, Morrison AP, Siwy ZS, Létant SE (2012) The role of pore geometry in single nanoparticle detection. ACS Nano 6:8366–8380
Ali M, Tahir M, Siwy Z, Neumann R, Tremel W, Ensinger W (2011) Hydrogen peroxide sensing with horseradish peroxidase-modified polymer single conical nanochannels. Anal Chem 83:1673–1680
Cortez ML, Pallarola D, Ceolin M, Azzaroni O, Battaglini F (2013) Electron transfer properties of dual self-assembled architectures based on specific recognition and electrostatic driving forces: its application to control substrate inhibition in horseradish peroxidase-based sensors. Anal Chem 85:2414–2422
Pallarola D, Bildering C, Pietrasanta LI, Queralto N, Knoll W, Battaglini F, Azzaroni O (2012) Recognitiondriven layer-by-layer construction of multiprotein assemblies on surfaces: a biomolecular toolkit for building up chemoresponsive bioelectrochemical interfaces. Phys Chem Chem Phys 14:11027–11039
Heins EA, Siwy ZS, Baker LA, Martin CR (2005) Detecting single porphyrin molecules in a conically shaped synthetic nanopore. Nano Lett 5:1824–1829
Acknowledgments
This research is supported by National Basic Research Program of China (21375042), National Basic Research Program of China (973 program, 2013CB933000), 1000 Young Talent (to Fan Xia) and initiatory financial support from HUST.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Liu, N., Yang, Z., Ou, X. et al. Nanopore-based analysis of biochemical species. Microchim Acta 183, 955–963 (2016). https://doi.org/10.1007/s00604-015-1560-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00604-015-1560-2