Abstract
Background
Pseudomonas aeruginosa is a nosocomial bacterium responsible for variety of infections. Inappropriate use of antibiotics could lead to emergence of multidrug-resistant (MDR) P. aeruginosa strains. Herein, a virulent phage; vB_PaeM_PS3 was isolated and tested for its application as alternative to antibiotics for controlling P. aeruginosa infections.
Methods
Phage morphology was observed using transmission electron microscopy (TEM). The phage host range and efficiency of plating (EOP) in addition to phage stability were analyzed. One-step growth curve was performed to detect phage growth kinetics. The impact of isolated phage on planktonic cells and biofilms was assessed. The phage genome was sequenced. Finally, the therapeutic potential of vB_PaeM_PS3 was determined in vivo.
Results
Isolated phage has an icosahedral head and a contractile tail and was assigned to the family Myoviridae. The phage vB_PaeM_PS3 displayed a broad host range, strong bacteriolytic ability, and higher environmental stability. Isolated phage showed a short latent period and large burst size. Importantly, the phage vB_PaeM_PS3 effectively eradicated bacterial biofilms. The genome of vB_PaeM_PS3 consists of 93,922 bp of dsDNA with 49.39% G + C content. It contains 171 predicted open reading frames (ORFs) and 14 genes as tRNA. Interestingly, the phage vB_PaeM_PS3 significantly attenuated P. aeruginosa virulence in host where the survival of bacteria-infected mice was markedly enhanced following phage treatment. Moreover, the colonizing capability of P. aeruginosa was markedly impaired in phage-treated mice as compared to untreated infected mice.
Conclusion
Based on these findings, isolated phage vB_PaeM_PS3 could be potentially considered for treating of P. aeruginosa infections.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
Pseudomonas aeruginosa is a widely distributed opportunistic pathogen that could survive in water, soil, animals and humans [1]. P. aeruginosa causes a variety of acute and chronic infections especially in immunocompromised patients [2, 3]. P. aeruginosa infections are very difficult to eradicate owing to higher resistance to antibiotics [4]. P. aeruginosa has been recently listed as a multidrug resistant (MDR) pathogen by the World Health Organization (WHO) and represents one of the biggest threats to public health [5]. This resistance could be related to bacterial inherited antibiotic resistance or to the acquisition of resistance genes through mobile genetic elements [4]. Additionally, it has been suggested that biofilm formation markedly contributes to bacterial resistance to various antibiotics, including quinolones, aminoglycosides and β-lactams leading to long term persistence [6, 7]. Biofilms confer higher antibiotic resistance than planktonic cells due to diminished antibiotic penetration through extracellular polymeric matrix in addition to the lower metabolic activity of persister cells [8, 9]. Therefore, there is a critical demand to develop new approaches to inhibit P. aeruginosa biofilms and keep this pathogen under control. Latterly, researchers have indicated that phage therapy could be considered as an effective approach to eradicate P. aeruginosa biofilms [10, 11].
Bacteriophages also known informally as phages are viruses that selectively infect and replicate within bacterial cells and considered the most prevalent entities on the earth [12]. Phage therapy refers to the use of bacteriophages in treatment of bacterial infections as potential alternatives to overcome limitations of antibiotics [13]. Phages possess remarkable advantages over antibiotics and are extremely specific to their host bacteria as well as they do not cause harmful side effects to normal microbiota [14]. In addition, phages are easily isolated and highly effective at destroying bacteria in biofilms [15, 16].
Unlike chemical antibiotics, phages have different strategies for eliminating resistant bacteria and have low environmental impact due to their natural origin [17, 18]. An important feature of phages is their specificity of action, exhibiting a narrow spectrum of activity. Furthermore, many reports have shown that phages are highly effective against various MDR pathogens in humans and animals [19, 20]. Hence, phage therapy possesses a great potential to replace antibiotic treatment in treatment of various bacterial infections [21].
While many previous studies have reported isolation of a huge number of phages infecting P. aeruginosa, there is still a tremendous lack of knowledge regarding either genomic characterization [22, 23], the antibiofilm activity or host safety of isolated phages targeting P. aeruginosa [24,25,26,27,28]. Hence, the present research is directed to introduce and widely characterize a virulent phage infecting P. aeruginosa. The phage genome, physical characteristics, environmental stability as well as antibiofilm activity will be uncovered in the present study. Finally, the capacity of isolated phage to limit P. aeruginosa virulence potential in host will be evaluated in vivo. The outcomes of current study are expected to be valuable for establishing effective treatment and controlling infectious diseases caused by P. aeruginosa.
Material and methods
Bacterial strains
Fifteen P. aeruginosa strains isolated from various clinical sources were provided by laboratories at the University Hospital, Zagazig, Egypt, with no direct involvement of patients in the study. In addition, P. aeruginosa reference strains, PAO1, ATCC 9027, and ATCC 27853, were involved in present study (Table 1).
Bacteriophage isolation, purification, and propagation
Wastewater samples were collected from Zagazig city, Egypt, for phage isolation by enrichment procedure as described previously [29]. Briefly, wastewater samples were centrifugated (6000 × g, 10 min), then the supernatant was filtrated through a 0.4-μm sterilized syringe filter. P. aeruginosa was grown to exponential phase, infected with the obtained filtrate in double concentrated tryptone soya (TS) broth then incubated with shaking for 24 h at 37°C. Afterwards, the enriched culture was centrifuged, filtered using 0.22 µm filter, then spot assay was performed to detect presence of phages in the filtrate as described [30]. Phage purification was performed using double agar layer method as described before [31]. Phage propagation was carried out according to Gencay et al. [32] by adding 5 mL of phage SM buffer to plates with confluent lysis, left overnight at 4°C and filtered by a 0.22 filter. Finally, the phages were kept at higher titer in a refrigerator at 4°C [32].
Transmission electron microscopy (TEM)
The morphology of phage particles was determined using transmission electron microscope (Hitachi H600A, Japan) exactly as previously described following staining with 2% phosphotungstic acid [33].
Host range determination and efficiency of plating (EOP) analysis
Host range specificity of isolated phage was determined using standard spot assay [34]. Bacteriophage efficiency of plating was determined by double agar layer technique. Briefly, tenfold serially diluted phage (100 µL) was mixed with exponential phase bacterial culture (100 µL) in soft agar layer and poured over the surface of TS agar plates followed by incubation at 37°C. The plaque forming units (PFUs) were counted for susceptible strains. The relative EOP values were determined by calculating the ratio of phage titer for a given bacteria to phage titer of the relevant host bacteria [35].
Phage stability to environmental conditions
Thermal stability of isolated phage was assessed by incubating phage particles at various temperatures (4–100°C) for 1 h. Similarly, phage pH stability was evaluated by incubating phage for 1 h in SM buffer adjusted at wide pH range (3–12) adjusted by 1 M HCl or 1 M NaOH. The number of surviving phages was determined by the double agar layer technique from triplicate assays [36].
Phage growth kinetics analysis
One-step growth curve for isolated phage was carried out to detect phage burst size and latent period as described [37]. Initially, bacterial host strain was cultured to reach exponential phase [108 colony forming units (CFU)/mL] and mixed with isolated phage at multiplicity of infection (MOI) of 0.1. Then, the mixture was centrifuged for 10 min at 10000 × g, pellets were resuspended in fresh TS broth and incubated at 37°C. Simultaneously, phage titer was determined by collecting samples of 100 µL at 5 min intervals then plated by double agar layer method.
In vitro killing assay and Biofilm inhibition assay
In vitro bacteriolytic activity of isolated phage was determined by measuring optical density (OD600) as mentioned before [38]. The phage was added to exponential phase bacterial cultures at different MOIs (0.1, 1, and 10). The phage/bacteria mixture was incubated at 37°C for 24 h. Bacterial culture without phage was used as a control. The phage-induced bacterial lysis (bacterial growth inhibition) was observed by measuring change in OD600. The antibiofilm activity of isolated phage was determined as exactly as previously described [39]. The phage was diluted in sterile TS broth to reach different MOIs (0.1, 1 and 10), then added to wells containing preformed bacterial biofilms followed by incubation at 37°C for 24 h. Control wells received an equivalent amount of TS broth only. The biomass of formed biofilms was quantified spectrophotometrically by the crystal violet technique using a microplate reader (Biotek, USA).
Determination of the frequency of bacteriophage insensitive mutants (BIMs)
The frequency of occurrence of bacteriophage insensitive mutants (BIMs) was determined as previously described [40]. Briefly, bacterial host culture was mixed with the phage suspension at MOI of 100 and incubated for 10 min at 37°C. Then, 100 µL of phage-bacterium mixture was serially diluted, plated on TS agar and incubated overnight at 37°C. The number of resulting colonies was counted from triplicate assays and the BIM frequency was estimated by dividing surviving bacterial colonies by initial viable counts and results were expressed as means ± standard deviation.
Phage-antibiotic synergy assay
The synergetic effect between isolated phage and two antibiotics commonly used for pseudomonas infections and represent different antibiotic classes, gentamicin (aminoglycoside) and ciprofloxacin (quinolone), was determined using checkerboard microdilution assay by calculating the fractional inhibitory concentration index (FICI) [41]. Briefly, the antibiotic was two-fold serially diluted and the vB_PaeM_PS3 phage was tenfold serially diluted followed by addition of bacterial suspension (105 CFU/mL) to each well of 96-well plate. The plates were incubated overnight at 37°C. The FICI was calculated using the following equation: FICI = FIC antibiotic + FIC phage; FIC antibiotic = Cantibiotic / MICantibiotic; FIC phage = Cphage / MICphage, where MICantibiotic and MICphage are the respective minimum inhibitory concentration (MICs) of the antibiotic and the phage alone, and Cantibiotic and Cphage are the respective concentrations of the antibiotic and phage in combination. The results were interpreted as the follow; synergy if FICI was ≤ 0.5; indifferent if 0.5 ˂ FICI ≤ 1; additive if 1 ˂ FIC ≤ 2 and antagonistic if FICI > 2.
Bacteriophage genome sequencing and data analysis
The genome of isolated phage was extracted using QIAamp1 DNA Mini kit (QIAGEN, Germany) following the manufacturer guidelines. The Nextera XT DNA Library Preparation Kit (Illumina, USA) was used to prepare the DNA library. The phage genome was sequenced at Genomics and Epigenomics Program, Cairo, Egypt, using Illumina Miseq next-generation sequencing. The raw sequence within the phage genome was checked for quality with FastQC and reads trimming was performed using Trimmomatic v0.36 [42]. The trimmed reads were assembled de novo into a single contig using Unicycler v0.4.8 [43]. The assembly quality was checked with Quality Assessment Tool for Genome Assemblies (QUAST—v4.4) [44]. Sequence analysis and annotation of resulting open reading frames (ORFs) were predicted using Prokka v1.14 [45] and Rapid Annotation Subsystems Technology (RAST) [46]. The virus circular genome map was created using the CGView [47]. Transfer RNA was detected using the online tool tRNAscan-SE 1.21 [48]. The phylogenetic analysis of phage whole genome was performed using the MEGA X program v10.2.2. [49]. Additionally, the nucleotide sequences of both the phage major capsid protein and terminase large subunit genes were compared with their corresponding sequences of reference bacteriophages deposited in the NCBI database to determine the phylogenetic position of the recently isolated phage [50]. Comparison of phage genome with similar phages was accomplished using the EasyFig program [51]. A dot plot was constructed using the Gepard-2.1 [52]. The annotated genome sequence of isolated phage was submitted to the National Center for Biotechnology Information (NCBI) nucleotide database and deposited in the GenBank under accession number (GenBank Acc. No OQ411628).
In vivo mice infection assay
The impact of phage on Pseudomonas infectivity was investigated in vivo using mice [26]. Briefly, five mice groups; each one contains 15 mice were included in the experiment. In the 1st group, mice were infected intraperitoneally (IP) with P. aeruginosa (2.5 × 107 CFU/mL), while in the 2nd group, mice were infected with P. aeruginosa and treated IP with phage at MOI of 100 (2.5 × 109 PFU/mL). The 3rd group represents mice without bacterial infection but inoculated only with phage. As negative controls, mice injected with PBS only and non-injected mice were included in the experiment. Mice survival was monitored and statistically analysed using Log-rank test. Additionally, three mice from each group were subjected to bacterial load and phage titer determination. Mice spleen and liver were collected, then bacterial burden as well as phage titer were quantified and statistically analyzed with P value < 0.05 is considered significant.
Results
Isolation and characterization of phage
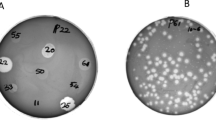
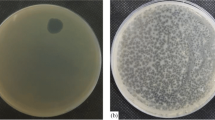
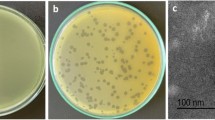
The phage was isolated from wastewater sample using P. aeruginosa PS3 as a host. Isolated phage produced clear circular plaques surrounded by halos with a diameter of 4–5 mm (Fig. 1a). Phage particles morphology were observed using TEM revealing that isolated phage has an icosahedral head of about 70 nm in diameter and a contractile tail of 100 nm in length (Fig. 1b). The phage was classified according guidelines of International Committee Taxonomy of Viruses (ICTV) and found to belong to the order Caudovirales and the family Myoviridae. The phage isolated herein was named as vB_PaeM_PS3 following the phage nomenclature recommendation [53].
Determination of host range and efficiency of plating (EOP)
The host range specificity of vB_PaeM_PS3 lysate was examined using the spot assay. Results show that isolated phage has a unique lytic profile and was able to infect 10 out of 18 of tested P. aeruginosa strains (55.5%) indicating a higher lytic efficiency of isolated phage (Table 1). To further assess the phage vB_PaeM_PS3 infection capability, EOP analysis was performed for each susceptible strain. vB_PaeM_PS3 phage was able to infect and form plaques (produce new progeny) on all susceptible P. aeruginosa strains with different infectivity patterns. Results show that only one P. aeruginosa isolate; PS15 exhibits efficient production higher than host strain (Table 1).
Environmental stability and one-step growth curve
Sensitivity of vB_PaeM_PS3 to different environmental conditions was characterized and phage survival under various temperatures and pH values was assessed. As shown in Fig. 2a, the phage vB_PaeM_PS3 was considerably stable at temperatures range from 40 to 60°C. However, the activity of vB_PaeM_PS3 was entirely diminished when incubated at 70–100°C. Additionally, the vB_PaeM_PS3 titer remained relatively unchanged when incubated at pH values ranging from 5 to 10. However, the phage infectivity slightly decreased at pH 11 and completely lost at pH 3 & 12 as shown in Fig. 2b. The one-step growth curve was used to characterize vB_PaeM_PS3 growth characters. The phage vB_PaeM_PS3 exhibited a short latent period of 10 min with host-cell lysis releasing about of 132 new virions per infected cell (Fig. 3).
In vitro killing and biofilm degradation assay
The bacteriolytic activity of vB_PaeM_PS3 phage was determined in broth culture medium. P. aeruginosa host strain (PS3) was infected with vB_PaeM_PS3 phage at various MOIs (0.1, 1 and 10). As shown in Fig. 4, growth of vB_PaeM_PS3-infected P. aeruginosa significantly decreased compared to the control P. aeruginosa culture without phage treatment. Remarkably, the cell lysis capacity of the phage was MOI dependent and bacterial inhibition was more obvious at higher MOIs. Furthermore, the ability of vB_PaeM_PS3 to eradicate biofilms of five clinical P. aeruginosa isolates recovered from various sources as well as the reference strains; ATCC 27853 and 9027 was evaluated. The phage vB_PaeM_PS3 showed a remarkable biofilm degrading efficiency both at higher and lower MOIs (MOI of 10 and 0.1, respectively) (Fig. 5). In addition, there was a considerable decrease in bacterial biomass compared to the control phage-untreated bacteria.
Effect of vB_PaeM_PS3 phage treatment on P. aeruginosa biofilms. Bacterial isolates were allowed to form biofilms for 24 h then treated with phage at various MOIs (0.1, 1 & 10) for 24 h. Biofilm biomass was evaluated by crystal violet (CV) staining. Data was expressed as means ± SE from three independent replicates with P < 0.05 was considered significant
Frequency of BIMs
The frequency of emergence of BIMs is considered as one of the major important characteristics for phage therapy. It is worth noting that the mutation frequency parameter in the host strain was [1.8 ± 0.1] × 10–3. This result indicates low ability to develop phage-resistance mutants and long-term effectiveness using phage therapy confirming that vB_PaeM_PS3 phage would be a promising candidate for phage therapy in the future.
The phage vB_PaeM_PS3 exhibits a synergistic effect on commonly used antibiotics
To establish the synergistic effect of vB_PaeM_PS3 phage in combination with conventional used antibiotics; gentamicin and ciprofloxacin, the FICIs values were determined. Interestingly, the MICs values for gentamicin and ciprofloxacin were markedly reduced when combined with vB_PaeM_PS3 phage giving FIC values of 0.16 and 0.26, respectively which were lower than 0.5 indicating a synergistic effect. MICs values of gentamicin and ciprofloxacin dropped from 32 and 16 to 2 and 4 µg ̸ mL, respectively. These results clearly suggest that isolated phage could be used in combination with antibiotics to effectively control P. aeruginosa infections.
Phage genome analysis
The genome of vB_PaeM_PS3 comprises 93,922 bp of dsDNA with 49.39% G + C content. The isolated phage was found to be a member of the Myoviridae family and the genus Pakpunavirus. As shown in Table 2 and Fig. 6a, a total of 171 ORFs were identified, of which 27 were assigned as functional proteins whereas hypothetical proteins were encoded by 144 ORFs. The functional proteins were divided into four major classes; structure proteins (ORF 17, ORF 20, ORF 31, ORF 35, ORF 39, and ORF 41); DNA metabolism, repair and replication which involves components that help in viral replication and enzymes responsible for modification of infected cell surface polysaccharides (ORF 51, ORF 61, ORF 64, ORF 66, ORF 67, ORF 68, ORF 75, ORF 86, ORF 88, ORF 90, ORF 91, ORF 157, ORF 159, ORF 161, ORF 165, ORF 170, and ORF 171); packaging and assembly proteins (ORF 15) and finally host cell lysis (ORF 10, ORF 42, ORF 167 and ORF 168). Interestingly, a cluster of 14 tRNA genes was predicted in the phage vB_PaeM_PS3 genome and listed in Supplementary Table S1. Of note that neither lysogenic genes nor host related sequences were identified in vB_PaeM_PS3 genome, confirming the lytic nature of vB_PaeM_PS3. In addition, the genes related to P. aeruginosa antibiotic resistance as well as toxins and virulence proteins production were absent in vB_PaeM_PS3 genome. Phylogenetic analysis based on the overall similarity with completely sequenced phage genomes in database shows that vB_PaeM_PS3 shares greatest nucleotide similarity with Pseudomonas phage vB_PaeM_SCUT-S2 (GenBank Acc. No MK340761.1), Pseudomonas phage vB_PaM_EPA1 (GenBank Acc. No MN013356.1), Pseudomonas phage PaYy-2 (GenBank Acc. No MH725810.1) and Pseudomonas phage SRT6 (GenBank Acc. No MH370478.1) representing percent identities of 96%, 95.2%, 95%, and 94.9%, respectively (Table 3, Fig. 6b, c and Supplementary Fig. S1). Moreover, neighbor-joining (NJ) phylogenetic trees were constructed for the phage major capsid protein and terminase large subunit to better illustrate the evolutionary relationships of vB_PaeM_PS3 with other bacteriophages. As shown in Fig. 7a and b, vB_PaeM_PS3 phage displayed close relation to other Caudovirales related pseudomonas phages, specifically, Pseudomonas phage PAK P1 and Pseudomonas phage PaYy-2, respectively. These results support and are in accordance with the whole genome phylogeny.
General features of vB_PaeM_PS3 genome. (a) Schematic genomic map of vB_PaeM_PS3 phage. The inner rings represent genome location, GC skew + (green) and GC skew (purple) and GC content (black). ORFs are represented by colored arrows. Functional ORFs were classified into four groups: yellow; DNA packaging, red; DNA metabolism, repair and replication, blue; structural proteins and purple; lysis proteins. Hypothetical proteins are indicated in grey. The figure was generated using CGView program. (b) Phylogenetic tree of vB_PaeM_PS3 using whole genome sequence and other closely related sequences. Phylogenetic tree was generated using MEGA-X computer program. (c) Comparative genomic analysis of the phage vB_PaeM_PS3 with homologous phages. Similarity level among phage sequences is represented by the colored scale bar from 20 to 100%. The coding sequences are represented by directional arrows. Predicted ORFs in vB_PaeM_PS3 genome are listed below. Genomic comparison was performed and plotted using Easyfig program
The phage vB_PaeM_PS3 exhibits a potent in vivo antibacterial activity against P. aeruginosa
The impact of vB_PaeM_PS3 on P. aeruginosa virulence was assessed in vivo. The mice infected with P. aeruginosa showed 100% mortality rate at 24 h. On contrast, the survival rate of P. aeruginosa infected mice was significantly improved (66.7%) following treatment with vB_PaeM_PS3 (Fig. 8a). Importantly, no death was recorded either in mice injected with vB_PaeM_PS3 alone or negative control mice and they remained healthy over the course of experiment. Additionally, both bacterial burden and vB_PaeM_PS3 titer were determined in isolated mice organs. Interestingly, treatment of P. aeruginosa-infected mice with vB_PaeM_PS3 effectively lowered Pseudomonas colonization in mice organs. The results revealed that bacterial loads in liver and spleen isolated from P. aeruginosa mice were markedly higher (27 × 104 ± 28, 52 × 104 ± 21 CFUs/g, respectively) compared to those of phage-untreated mice (4837 ± 16, 9210 ± 17 CFUs/g, respectively) (Fig. 8b, c). It is worth mentioning that the phage vB_PaeM_PS3 was completely eliminated and not detected in organs isolated from mice at 72 h post-inoculation.
Impact of phage vB_PaeM_PS3 on P. aeruginosa pathogenesis in vivo. a) Survival rate of mice infected with P. aeruginosa and treated with isolated phage. Mice survival was plotted using Kaplan–Meier survival curve and statically analyzed using Log-rank test. Bacterial count and phage titer in the liver (b) and spleen (c) isolated from mice at 24, 48, and 72 h after infection. Bacterial and phage load were represented on left and right y axis, respectively. Data are presented as means ± SE of three independent experiments
Discussion
P. aeruginosa is a serious nosocomial pathogen which causes a variety of health illnesses due to increased prevalence of resistant strains [54]. Therefore, phage therapy has received a significant interest as an alternative to antibiotics as well as a new strategy to alleviate antimicrobial resistance [55]. One of the major benefits of phage therapy over antibiotics is host specificity without influencing human microbial flora [56]. Consequently, isolation and characterization of new lytic phages targeting antibiotic resistant P. aeruginosa would be helpful to combat these infections that threaten human healthcare.
Lytic phages are more preferred for phage therapy as compared with temperate ones that could lysogenize bacterial cells. In addition, toxin genes should be absent in the genomes of phages applied for phage therapy [57]. In this context, a novel virulent phage vB_PaeM_PS3 targeting P. aeruginosa was isolated herein and fully characterized. The morphological features coupled with genomic analysis revealed that vB_PaeM_PS3 belonging to the family Myoviridae and order Caudovirales that represents approximately 95% of isolated phages [58]. The phage vB_PaeM_PS3 exhibited a relatively strong lytic potential on tested P. aeruginosa isolates. These findings were further confirmed by EOP analysis which is considered a rigorous test for evaluating phage infectivity indicated the productive infectivity of vB_PaeM_PS3 and releasing of new progeny rather than by non-productive infection or lysis from without [59]. Additionally, the isolated phage demonstrated a reasonable environmental stability and optimal growth over a broad range of temperature and pH ranges. Phage tolerance to various environmental conditions is very relevant if phage would be involved in therapeutic applications as well as during storage and production process [60]. This finding agrees with earlier studies that reported that the majority of phages survive well at pH range of 5 to 9 [61]. The vB_PaeM_PS3 phage has a latent period of 10 min and burst size of approximately 132 virions per infected cell with a higher likelihood of previously isolated phage Ps12-on-D with latent period of 10 min and an average size of 115 PFU/infected cell [62]. Phages had short latent period and large burst size are more effective and preferable for clinical use [63]. Consequently, vB_PaeM_PS3 could be considered as a good candidate for biocontrol and phage therapy.
The combination of phage and antibiotics is of great importance to avoid the development of antibiotic resistant bacterial strains [64]. Several studies have demonstrated that phages could produce synergistic antimicrobial effects when combined with conventional antibiotics [65]. Current results show a synergetic effect of vB_PaeM_PS3 when combined with traditionally used antibiotics in the treatment of pseudomonas infections, hence supporting promising application of isolated phage for therapy.
The most critical aspect associated with P. aeruginosa infections is biofilm formation [66, 67]. Fortunately, phages have been found to be highly effective in treatment of recalcitrant infections and biofilm removal [68]. The most common mechanism of phage antibiofilm activity could be due to production of different enzymes such as depolymerases and endolysins [69]. Depolymerase can specifically degrade the extracellular polysaccharides in bacterial matrix facilitating phage adsorption, penetration and lysis of bacterial cell [70]. Furthermore, endolysins can degrade bacterial peptidoglycan and assist in release of new phage progeny out of host cell [71]. Current results show that vB_PaeM_PS3 possesses a potent antibiofilm activity and therefore is an attractive biological agent for therapeutic application.
In order to consider phages for in vivo therapeutic applications, it is important to ascertain that phage genome devoid of any lysogenic, antibiotic resistance and virulence-related genes [72]. Importantly, the analysis of vB_PaeM_PS3 genome revealed the absence of both virulence-encoding and lysogeny genes indicating the true lytic nature of bacteriophage. Furthermore, phage genome analysis allowed the detection of 14 tRNAs. The presence of tRNAs is often found in myoviruses with large genomes and is common in strictly virulent or lytic phages [73, 74]. The phage-encoded tRNA genes are generally present in clusters and promote an efficient and more rapid translation rate [75]. In addition, it is thought that tRNAs are responsible for phages have short latent period and large burst size because. It has been reported that tRNAs enable phages propagation and enhance viral replication kinetics [76].
The above findings become more convincing in light of previous reports that documented the effectiveness of local and systemic application of phages in controlling infections by P. aeruginosa [77]. For instance, Jeon and Yong found that phage nasal inhalation effectively reduced P. aeruginosa count in mice lungs with pneumonia [78]. Similarly, other successful clinical trials revealed the efficacy of phage therapy against ear and burn infections caused by P. aeruginosa [79, 80]. Intriguingly, the phage vB_PaeM_PS3 has been found to be highly efficient in attenuation of P. aeruginosa virulence in mice, which further encourages its future application in phage therapy. Not only vB_PaeM_PS3 enhanced survival of P. aeruginosa-infected mice, it markedly reduced bacterial proliferation and colonization in tissues obtained from infected mice. It is worth mentioning that no detectable side effects or harmful signs were observed in mice upon phage treatment over the experiment course. Given the promising results of this study, it may be useful to incorporate vB_PaeM_PS3 phage in treatment of human infections with P. aeruginosa.
In conclusion, vB_PaeM_PS3 phage is a novel lytic phage, member of the family Myoviridae and the genus Pakpunavirus. The phage vB_PaeM_PS3 exhibited an optimum environmental stability added to its effective antimicrobial activity and the ability to disrupt bacterial biofilms. Genomic analysis revealed that phage vB_PaeM_PS3 does not contain any toxic genes or integrases and the detected tRNAs could further enhance phage protein translation that make the phage independent on the host. Furthermore, in vivo results proved the safety and efficiency of isolated phage against P. aeruginosa as an alternative to traditionally used antibiotics. Based on these findings, the phage vB_PaeM_PS3 fulfills all requirements of effective phages for further application as an innovative candidate against P. aeruginosa infections. However, further investigations about phage formulations are required to be involved in clinical trials.
Data availability
The authors confirm that the data supporting the findings of this study are available within the article.
Abbreviations
- MDR:
-
Multidrug resistant
- ORFs:
-
Open reading frames
- WHO:
-
World Health Organization
- TS:
-
Tryptone soya
- TEM:
-
Transmission electron microscopy
- EOP:
-
Efficiency of plating
- PFUs:
-
Plaque forming units
- CFUs:
-
Colony-forming units
- MOI:
-
Multiplicity of infection
- OD:
-
Optical density
- BIMs:
-
Bacteriophage insensitive mutants
- FICI:
-
Fractional inhibitory concentration index
- MIC:
-
Minimum inhibitory concentration
- QUAST:
-
Quality Assessment Tool for Genome Assemblies
- RAST:
-
Rapid Annotation Subsystems Technology
- NCBI:
-
National Center for Biotechnology Information
- IP:
-
Intraperitoneally
- ICTV:
-
International Committee Taxonomy of Viruses
- NJ:
-
Neighbor-joining
References
Crone S, Vives-Flórez M, Kvich L, Saunders AM, Malone M, Nicolaisen MH et al (2020) The environmental occurrence of Pseudomonas aeruginosa. APMIS 128(3):220–231. https://doi.org/10.1111/apm.13010
Rossi E, La Rosa R, Bartell JA, Marvig RL, Haagensen JA, Sommer LM et al (2021) Pseudomonas aeruginosa adaptation and evolution in patients with cystic fibrosis. Nat Rev Microbiol 19(5):331–342. https://doi.org/10.1038/s41579-020-00477-5
Jurado-Martín I, Sainz-Mejías M, McClean S (2021) Pseudomonas aeruginosa: An audacious pathogen with an adaptable arsenal of virulence factors. Int J Mol Sci 22(6):3128. https://doi.org/10.3390/ijms22063128
Breidenstein EB, de la Fuente-Núñez C, Hancock RE (2011) Pseudomonas aeruginosa: all roads lead to resistance. Trends Microbiol 19(8):419–426. https://doi.org/10.1016/j.tim.2011.04.005
Daikos GL, da Cunha CA, Rossolini GM, Stone GG, Baillon-Plot N, Tawadrous M et al (2021) Review of ceftazidime-avibactam for the treatment of infections caused by Pseudomonas aeruginosa. Antibiotics 10(9):1126. https://doi.org/10.3390/antibiotics10091126
Pang Z, Raudonis R, Glick BR, Lin T-J, Cheng Z (2019) Antibiotic resistance in Pseudomonas aeruginosa: mechanisms and alternative therapeutic strategies. Biotechnol Adv 37(1):177–192. https://doi.org/10.1016/j.biotechadv.2018.11.013
Hancock RE, Speert DP (2000) Antibiotic resistance in Pseudomonas aeruginosa: mechanisms and impact on treatment. Drug Resist Updat 3(4):247–255. https://doi.org/10.1054/drup.2000.0152
Ma L, Conover M, Lu H, Parsek MR, Bayles K, Wozniak DJ (2009) Assembly and development of the Pseudomonas aeruginosa biofilm matrix. PLoS Pathog 5(3):e1000354. https://doi.org/10.1371/journal.ppat.1000354
Spoering AL, Lewis K (2001) Biofilms and planktonic cells of Pseudomonas aeruginosa have similar resistance to killing by antimicrobials. J Bacteriol 183(23):6746–6751. https://doi.org/10.1128/JB.183.23.6746-6751.2001
Forti F, Roach DR, Cafora M, Pasini ME, Horner DS, Fiscarelli EV et al (2018) Design of a broad-range bacteriophage cocktail that reduces Pseudomonas aeruginosa biofilms and treats acute infections in two animal models. Antimicrob Agents Chemother 62(6):e02573-e2617. https://doi.org/10.1128/AAC.02573-17
Waters EM, Neill DR, Kaman B, Sahota JS, Clokie MR, Winstanley C et al (2017) Phage therapy is highly effective against chronic lung infections with Pseudomonas aeruginosa. Thorax 72(7):666–667. https://doi.org/10.1136/thoraxjnl-2016-209265
Clokie MR, Millard AD, Letarov AV, Heaphy S (2011) Phages in nature. Bacteriophage 1(1):31–45. https://doi.org/10.4161/bact.1.1.14942
Golkar Z, Bagasra O, Pace DG (2014) Bacteriophage therapy: a potential solution for the antibiotic resistance crisis. J Infect Dev Ctries 8(02):129–136. https://doi.org/10.3855/jidc.3573
Moelling K, Broecker F, Willy C (2018) A wake-up call: we need phage therapy now. Viruses 10(12):688. https://doi.org/10.3390/v10120688
Khalifa L, Shlezinger M, Beyth S, Houri-Haddad Y, Coppenhagen-Glazer S, Beyth N et al (2016) Phage therapy against Enterococcus faecalis in dental root canals. J Oral Microbiol 8:32157. https://doi.org/10.3402/jom.v8.32157
Chang H-C, Chen C-R, Lin J-W, Shen G-H, Chang K-M, Tseng Y-H et al (2005) Isolation and characterization of novel giant Stenotrophomonas maltophilia phage φSMA5. App Environ Microbiol 71(3):1387–1393. https://doi.org/10.1128/AEM.71.3.1387-1393.2005
Abedon ST, García P, Mullany P, Aminov R (2017) Editorial: Phage Therapy: Past, Present and Future. Front Microbiol 8:981. https://doi.org/10.3389/fmicb.2017.00981
Wittebole X, De Roock S, Opal SM (2014) A historical overview of bacteriophage therapy as an alternative to antibiotics for the treatment of bacterial pathogens. Virulence 5(1):226–235. https://doi.org/10.4161/viru.25991
Schooley RT, Biswas B, Gill JJ, Hernandez-Morales A, Lancaster J, Lessor L et al (2017) Development and use of personalized bacteriophage-based therapeutic cocktails to treat a patient with a disseminated resistant Acinetobacter baumannii infection. Antimicrob Agents Chemother 61(10):e00954-e1017. https://doi.org/10.1128/AAC.00954-17
Dedrick RM, Guerrero-Bustamante CA, Garlena RA, Russell DA, Ford K, Harris K et al (2019) Engineered bacteriophages for treatment of a patient with a disseminated drug-resistant Mycobacterium abscessus. Nat Med 25(5):730–733. https://doi.org/10.1038/s41591-019-0437-z
Casey E, Van Sinderen D, Mahony J (2018) In vitro characteristics of phages to guide ‘real life’phage therapy suitability. Viruses 10(4):163. https://doi.org/10.3390/v10040163
Tiwari BR, Kim S, Rahman M, Kim J (2011) Antibacterial efficacy of lytic Pseudomonas bacteriophage in normal and neutropenic mice models. J Microbiol 49:994–999. https://doi.org/10.1007/s12275-011-1512-4
Wroe JA, Johnson CT, García AJ (2020) Bacteriophage delivering hydrogels reduce biofilm formation in vitro and infection in vivo. J Biomed Mater Res A 108(1):39–49. https://doi.org/10.1002/jbm.a.36790
Garbe J, Wesche A, Bunk B, Kazmierczak M, Selezska K, Rohde C et al (2010) Characterization of JG024, a Pseudomonas aeruginosa PB1-like broad host range phage under simulated infection conditions. BMC Microbiol 10(1):1–10. https://doi.org/10.1186/1471-2180-10-301
Oliveira VC, Bim FL, Monteiro RM, Macedo AP, Santos ES, Silva-Lovato CH et al (2020) Identification and characterization of new bacteriophages to control multidrug-resistant Pseudomonas aeruginosa biofilm on endotracheal tubes. Front Microbiol 11:580779. https://doi.org/10.3389/fmicb.2020.580779
Alvi IA, Asif M, Tabassum R, Aslam R, Abbas Z, Rehman SU (2020) RLP, a bacteriophage of the family Podoviridae, rescues mice from bacteremia caused by multi-drug-resistant Pseudomonas aeruginosa. Arch Virol 165:1289–1297. https://doi.org/10.1007/s00705-020-04601-x
Yang Y, Shen W, Zhong Q, Chen Q, He X, Baker JL et al (2020) Development of a bacteriophage cocktail to constrain the emergence of phage-resistant Pseudomonas aeruginosa. Front Microbiol 11:327. https://doi.org/10.3389/fmicb.2020.00327
Ceyssens P-J, Lavigne R, Mattheus W, Chibeu A, Hertveldt K, Mast J et al (2006) Genomic analysis of Pseudomonas aeruginosa phages LKD16 and LKA1: Establishment of the φKMV subgroup within the T7 supergroup. J Bacteriol 188(19):6924–6931. https://doi.org/10.1128/JB.00831-06
Lerdsittikul V, Thongdee M, Chaiwattanarungruengpaisan S, Atithep T, Apiratwarrasakul S, Withatanung P et al (2022) A novel virulent Litunavirus phage possesses therapeutic value against multidrug resistant Pseudomonas aeruginosa. Sci Rep 12(1):21193. https://doi.org/10.1038/s41598-022-25576-6
Tan D, Gram L, Middelboe M (2014) Vibriophages and their interactions with the fish pathogen Vibrio anguillarum. Appl Environ Microbiol 80(10):3128–3140. https://doi.org/10.1128/AEM.03544-13
Kropinski AM, Mazzocco A, Waddell TE, Lingohr E, Johnson RP (2009) Enumeration of Bacteriophages by Double Agar Overlay Plaque Assay. In: Clokie, M.R., Kropinski, A.M. (eds) Bacteriophages. Methods Mol Biol™, vol 501. Humana Press; 69–76 https://doi.org/10.1007/978-1-60327-164-6_7
Gencay YE, Birk T, Sørensen MCH, Brøndsted L (2017) Methods for Isolation, Purification, and Propagation of Bacteriophages of Campylobacter jejuni . In: Butcher, J., Stintzi, A. (eds) Campylobacter jejuni. Methods Mol Biol, vol 1512. Humana Press, New York, NY; 19–28. https://doi.org/10.1007/978-1-4939-6536-6_3
Essoh C, Blouin Y, Loukou G, Cablanmian A, Lathro S, Kutter E et al (2013) The susceptibility of Pseudomonas aeruginosa strains from cystic fibrosis patients to bacteriophages. PLoS One 8(4):e60575. https://doi.org/10.1371/journal.pone.0060575
González-Villalobos E, Ribas-Aparicio RM, Montealegre GER, Belmont-Monroy L, Ortega-García Y, Aparicio-Ozores G et al (2021) Isolation and characterization of novel bacteriophages as a potential therapeutic option for Escherichia coli urinary tract infections. Appl Microbiol Biotechnol 105:5617–5629. https://doi.org/10.1007/s00253-021-11432-6
Khan Mirzaei M, Nilsson AS (2015) Isolation of phages for phage therapy: a comparison of spot tests and efficiency of plating analyses for determination of host range and efficacy. PLoS one 10(3):e0118557. https://doi.org/10.1371/journal.pone.0118557
Shahin K, Bouzari M, Wang R (2018) Isolation, characterization and genomic analysis of a novel lytic bacteriophage vB_SsoS-ISF002 infecting Shigella sonnei and Shigella flexneri. J Med Microbiol 67(3):376–386. https://doi.org/10.1099/jmm.0.000683
Kropinski AM (2018) Practical Advice on the One-Step Growth Curve. In: Clokie, M., Kropinski, A., Lavigne, R. (eds) Bacteriophages. Methods Mol Biol, vol 1681. Humana Press, New York, NY; 41–47. https://doi.org/10.1007/978-1-4939-7343-9_3
Peng S-Y, Chen L-K, Wu W-J, Paramita P, Yang P-W, Li Y-Z et al (2020) Isolation and characterization of a new phage infecting Elizabethkingia anophelis and evaluation of its therapeutic efficacy in vitro and in vivo. Front Microbiol 11:728. https://doi.org/10.3389/fmicb.2020.00728
Horváth M, Kovács T, Koderivalappil S, Ábrahám H, Rákhely G, Schneider G (2020) Identification of a newly isolated lytic bacteriophage against K24 capsular type, carbapenem resistant Klebsiella pneumoniae isolates. Sci Rep 10(1):5891. https://doi.org/10.1038/s41598-020-62691-8
Wittmann J, Dreiseikelmann B, Rohde C, Rohde M, Sikorski J (2014) Isolation and characterization of numerous novel phages targeting diverse strains of the ubiquitous and opportunistic pathogen Achromobacterxylosoxidans. PLoS One 9(1):e86935. https://doi.org/10.1371/journal.pone.0086935
Jo A, Ding T, Ahn J (2016) Synergistic antimicrobial activity of bacteriophages and antibiotics against Staphylococcus aureus. Food Sci Biotechnol 25:935–940. https://doi.org/10.1007/s10068-016-0153-0
Brown J, Pirrung M, Dashboard MLAFQC (2017) integrates FastQC results into a web-based, interactive, and extensible FASTQ quality control tool. Bioinformatics 33(19):3137–3139. https://doi.org/10.1093/bioinformatics/btx373
Wick RR, Judd LM, Gorrie CL, Holt KE (2017) Completing bacterial genome assemblies with multiplex MinION sequencing. Microb Genom 3(10):e000132. https://doi.org/10.1099/mgen.0.000132
Gurevich A, Saveliev V, Vyahhi N, Tesler G (2013) QUAST: quality assessment tool for genome assemblies. Bioinformatics 29(8):1072–1075. https://doi.org/10.1093/bioinformatics/btt086
Prokka ST (2014) rapid prokaryotic genome annotation. Bioinformatics 30(14):2068–2069. https://doi.org/10.1093/bioinformatics/btu153
McNair K, Aziz RK, Pusch GD, Overbeek R, Dutilh BE, Edwards R (2018) Phage Genome Annotation Using the RAST Pipeline. In: Clokie, M., Kropinski, A., Lavigne, R. (eds) Bacteriophages. Methods Mol Biol vol 1681. Humana Press, New York, NY; https://doi.org/10.1007/978-1-4939-7343-9_17.
Stothard P, Wishart DS (2005) Circular genome visualization and exploration using CGView. Bioinformatics 21(4):537–539. https://doi.org/10.1093/bioinformatics/bti054
Chan PP, Lowe TM (2019) tRNAscan-SE: Searching for tRNA Genes in Genomic Sequences. In: Kollmar, M. (eds) Gene Prediction. Methods Mol Biol vol 1962. Humana, New York, NY; https://doi.org/10.1007/978-1-4939-9173-0_1
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35(6):1547. https://doi.org/10.1093/molbev/msy096
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Sullivan MJ, Petty NK, Beatson SA (2011) Easyfig: a genome comparison visualizer. Bioinformatics 27(7):1009–1010. https://doi.org/10.1093/bioinformatics/btr039
Krumsiek J, Arnold R, Rattei T (2007) Gepard: a rapid and sensitive tool for creating dotplots on genome scale. Bioinformatics 23(8):1026–1028. https://doi.org/10.1093/bioinformatics/btm039
Kropinski AM, Prangishvili D, Lavigne R (2009) Position paper: the creation of a rational scheme for the nomenclature of viruses of Bacteria and Archaea. Environ Microbiol 11(11):2775–2777. https://doi.org/10.1111/j.1462-2920.2009.01970.x
Gellatly S, Hancock R (2013) Pseudomonas aeruginosa: new insights into pathogenesis and host defenses. Pathog Dis 67(3):159–173. https://doi.org/10.1111/2049-632X.12033
Pires DP, Vilas Boas D, Sillankorva S, Azeredo J (2015) Phage therapy: a step forward in the treatment of Pseudomonas aeruginosa infections. J Virol 89(15):7449–7456. https://doi.org/10.1128/JVI.00385-15
Drulis-Kawa Z, Majkowska-Skrobek G, Maciejewska B, Delattre A-S, Lavigne R (2012) Learning from bacteriophages-advantages and limitations of phage and phage-encoded protein applications. Curr Protein Pept Sci 13(8):699–722. https://doi.org/10.2174/138920312804871193
Fernández L, Gutiérrez D, García P, Rodríguez A (2019) The perfect bacteriophage for therapeutic applications—a quick guide. Antibiotics 8(3):126. https://doi.org/10.3390/antibiotics8030126
Swanson MM, Reavy B, Makarova KS, Cock PJ, Hopkins DW, Torrance L et al (2012) Novel bacteriophages containing a genome of another bacteriophage within their genomes. PloS one 7(7):e40683. https://doi.org/10.1371/journal.pone.0040683
Turner D, Hezwani M, Nelson S, Salisbury V, Reynolds D (2012) Characterization of the Salmonella bacteriophage vB_SenS-Ent1. J Gen Virol 93(9):2046–2056. https://doi.org/10.1099/vir.0.043331-0
Capra M, Quiberoni A, Reinheimer J (2006) Phages of Lactobacillus casei/paracasei: response to environmental factors and interaction with collection and commercial strains. J Appl Microbiol 100(2):334–342. https://doi.org/10.1111/j.1365-2672.2005.02767.x
Jamalludeen N, Johnson RP, Friendship R, Kropinski AM, Lingohr EJ, Gyles CL (2007) Isolation and characterization of nine bacteriophages that lyse O149 enterotoxigenic Escherichia coli. Vet Microbiol 124(1–2):47–57. https://doi.org/10.1016/j.vetmic.2007.03.028
Akremi I, Merabishvili M, Jlidi M, Haj Brahim A, Ben Ali M, Karoui A et al (2022) Isolation and Characterization of Lytic Pseudomonas aeruginosa Bacteriophages Isolated from Sewage Samples from Tunisia. Viruses 14(11):2339. https://doi.org/10.3390/v14112339
Abedon ST, Herschler TD, Stopar D (2001) Bacteriophage latent-period evolution as a response to resource availability. Appl Environ Microbiol 67(9):4233–4241. https://doi.org/10.1128/AEM.67.9.4233-4241.2001
Coulon C, Vinogradov E, Filloux A, Sadovskaya I (2010) Chemical analysis of cellular and extracellular carbohydrates of a biofilm-forming strain Pseudomonas aeruginosa PA14. PLoS One 5(12):e14220. https://doi.org/10.1371/journal.pone.0014220
Wei Q, Ma LZ (2013) Biofilm matrix and its regulation in Pseudomonas aeruginosa. Int J Mol Sci 14(10):20983–21005. https://doi.org/10.3390/ijms141020983
Manohar P, Madurantakam Royam M, Loh B, Bozdogan B, Nachimuthu R, Leptihn S (2022) Synergistic effects of phage–antibiotic combinations against Citrobacter amalonaticus. ACS Infect Dis 8(1):59–65. https://doi.org/10.1021/acsinfecdis.1c00117
Lin Y, Chang RYK, Britton WJ, Morales S, Kutter E, Chan H-K (2018) Synergy of nebulized phage PEV20 and ciprofloxacin combination against Pseudomonas aeruginosa. Int J Pharm 551(1–2):158–165. https://doi.org/10.1016/j.ijpharm.2018.09.024
Motlagh AM, Bhattacharjee AS, Goel R (2016) Biofilm control with natural and genetically-modified phages. World J Microbiol Biotechnol 32:1–10. https://doi.org/10.1007/s11274-016-2009-4
Tian F, Li J, Nazir A, Tong Y (2021) Bacteriophage–a promising alternative measure for bacterial biofilm control. Infect Drug Resist 14:205–217
Yan J, Mao J, Xie J (2014) Bacteriophage polysaccharide depolymerases and biomedical applications. BioDrugs 28(3):265–274. https://doi.org/10.1007/s40259-013-0081-y
Schmelcher M, Donovan DM, Loessner MJ (2012) Bacteriophage endolysins as novel antimicrobials. Future Microbiol 7(10):1147–1171. https://doi.org/10.2217/fmb.12.97
Endersen L, Guinane CM, Johnston C, Neve H, Coffey A, Ross RP et al (2015) Genome analysis of Cronobacter phage vB_CsaP_Ss1 reveals an endolysin with potential for biocontrol of Gram-negative bacterial pathogens. J Gen Virol 96(2):463–477. https://doi.org/10.1099/vir.0.068494-0
Santos SB, Kropinski AM, Ceyssens P-J, Ackermann H-W, Villegas A, Lavigne R et al (2011) Genomic and proteomic characterization of the broad-host-range Salmonella phage PVP-SE1: creation of a new phage genus. J Virol 85(21):11265–11273. https://doi.org/10.1128/JVI.01769-10
Bailly-Bechet M, Vergassola M, Rocha E (2007) Causes for the intriguing presence of tRNAs in phages. Genome Res 17(10):1486–1495
Lu S, Le S, Tan Y, Zhu J, Li M, Rao X et al (2013) Genomic and proteomic analyses of the terminally redundant genome of the Pseudomonas aeruginosa phage PaP1: establishment of genus PaP1-like phages. PloS one 8(5):e62933. https://doi.org/10.1371/journal.pone.0062933
Jun JW, Yun SK, Kim HJ, Chai JY, Park SC (2014) Characterization and complete genome sequence of a novel N4-like bacteriophage, pSb-1 infecting Shigella boydii. Res Microbiol 165(8):671–678. https://doi.org/10.1016/j.resmic.2014.09.006
Hagens S, Habel A, Von Ahsen U, Von Gabain A, Bläsi U (2004) Therapy of experimental Pseudomonas infections with a nonreplicating genetically modified phage. Antimicrob Agents Chemother 48(10):3817–3822. https://doi.org/10.1128/AAC.48.10.3817-3822.2004
Jeon J, Yong D (2019) Two novel bacteriophages improve survival in Galleria mellonella infection and mouse acute pneumonia models infected with extensively drug-resistant Pseudomonas aeruginosa. Appl Environ Microbiol 85(9):e02900-e2918. https://doi.org/10.1128/AEM.02900-18
Wright A, Hawkins C, Änggård E, Harper D (2009) A controlled clinical trial of a therapeutic bacteriophage preparation in chronic otitis due to antibiotic-resistant Pseudomonas aeruginosa; a preliminary report of efficacy. Clin otolaryngol 34(4):349–357. https://doi.org/10.1111/j.1749-4486.2009.01973.x
Jault P, Leclerc T, Jennes S, Pirnay JP, Que Y-A, Resch G et al (2019) Efficacy and tolerability of a cocktail of bacteriophages to treat burn wounds infected by Pseudomonas aeruginosa (PhagoBurn): a randomised, controlled, double-blind phase 1/2 trial. Lancet Infect Dis 19(1):35–45. https://doi.org/10.1016/S1473-3099(18)30482-1
Acknowledgements
The authors would like to acknowledge the medical staff at clinical laboratories of Zagazig University Hospitals in Zagazig, Egypt, for providing clinical Pseudomonas aeruginosa isolates for this study with no direct involvement of patients in the study.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
Conceptualization, A.E., G.S., and M.A; methodology, A.A., and M.A; validation, A.A., and M.A; investigation, A.E., G.S., and M.A; data curation, A.A., and M.A; writing-original draft preparation, A.A., and M.A; writing-review and editing, A.A., and M.A; visualization, A.A., A.E., G.S., and M.A; supervision, G.S., A.E., and M.A.; all authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The study was carried out according to the Declaration of Helsinki and approved by Zagazig University Institutional Review Board (ZU-IRB) under the number (ZU-IRB #10219). The animal procedures in this study were approved by Zagazig University Institutional Animal Care and Use Committee (ZU-IACUC) with approval number (ZU-IACUC/3/F/72/2022) and were carried out according to Guide for the Care and Use of Laboratory Animals (8th edition, National Academies Press).
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abdelghafar, A., El-Ganiny, A., Shaker, G. et al. A novel lytic phage exhibiting a remarkable in vivo therapeutic potential and higher antibiofilm activity against Pseudomonas aeruginosa. Eur J Clin Microbiol Infect Dis 42, 1207–1234 (2023). https://doi.org/10.1007/s10096-023-04649-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-023-04649-y