Abstract
Each year, hundreds of thousands of farmed Atlantic salmon escape from fish farms into the wild. Some of these escapees enter freshwater, and manage to interbreed with native populations. To hinder further genetic introgression in native populations, the use of sterile triploid salmon within commercial aquaculture is being examined. However, if triploid escapees migrate into freshwater, they may still have ecological impacts on local populations. In the present study, we used microsatellite DNA genotyping to determine the ploidy of 3794 farmed escapees captured in 17 Norwegian rivers in the period 2007–2014. Although a previous study has reported an average of 2 % triploids in Norwegian fish farms during this exact period, here, we only observed 7 (0.18 %) triploids among the escapees captured in freshwater. In addition, we identified three trisomic escapees. For the triploids where the within-river capture location was determined, they were only observed in the lower reaches and not on the spawning grounds. It is concluded that propensity for triploid Atlantic salmon to migrate into freshwater following escape from a fish farm is significantly lower than for normal diploid salmon escapees. Therefore, commercial production of triploids should not only be seen as an effective way of stopping genetic introgression, it will also significantly reduce the numbers of escapees entering rivers, which in turn limits ecological interactions and potential disease transmission.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Atlantic salmon (Salmo salar L.) aquaculture industry has a number of environmental challenges, of which farmed escapees and their potential genetic interaction with wild conspecifics represents one of the most significant (Taranger et al. 2015). Each year, hundreds of thousands of farmed salmon escape into the wild. While many of these disappear never to be seen again (Skilbrei et al. 2015a), some enter freshwater and spawn with wild salmon (Lura and Saegrov 1991; Saegrov et al. 1997). As a result, genetic changes, caused by introgression of non-native farmed salmon, have been documented in native Atlantic salmon populations in Ireland, Canada and Norway (Clifford et al. 1998b; Crozier 1993; Glover et al. 2013, 2012; Skaala et al. 2006).
As one of the measures to mitigate the potential effects of genetic interactions with wild populations, commercial production of triploid salmon, that are sterile and therefore unable to produce viable offspring, is being explored. The procedures to induce triploidy are relatively straightforward and have been used in a number of experiments (e.g., Frenzl et al. 2014; Leclercq et al. 2011; Taylor et al. 2014). However, while triploid salmon are sterile, they can still develop secondary sexual characteristics, and it has been demonstrated that farmed triploid males can successfully coax normal diploid wild females to mate with them (Fjelldal et al. 2014). This could lead to many of the eggs being unfertilized depending upon the numbers of wild males also present during spawning. Therefore, if there are large numbers of triploid farmed salmon on the spawning grounds, genetic interactions could be replaced by ecological interactions instead.
As Atlantic salmon aquaculture is primarily based upon freshwater production in tanks on land, and marine production in cages, the majority of fish escaping from aquaculture installations occur during the marine phase of the production cycle. Therefore, in order for triploid escapees to disturb spawning in rivers (Fjelldal et al. 2014), they first need to migrate into freshwater. Immature diploid farmed Atlantic salmon escapees have been documented to occasionally enter freshwater soon after escape from marine cages (Madhun et al. 2015; Skilbrei et al. 2015a). However, freshwater migration in salmon is primarily linked with adult maturation, and it is therefore possible that sterile triploid escapees will display reduced motivation to enter freshwater. An experimental study to evaluate this was conducted in Ireland by releasing groups of diploid and triploid salmon smolts into the sea (Cotter et al. 2000). The results of that study demonstrated that adult return rates to freshwater were significantly lower (approximately 4 times) in the triploid experimental group. In a recent study of spontaneous triploidy (i.e., triploids that arise from a random event during development as opposed to a deliberate protocol), it was demonstrated that the Norwegian aquaculture industry had produced approximately 2 % spontaneous triploids in the period 2007–2014, and for the first time, demonstrated that triploid salmon, originating from a commercial fish farm entered a river (Glover et al. 2015). However, the authors concluded that it would be essential to analyse a larger number of escapees in rivers in order to determine the relative frequency of this behavior.
Scales are often sampled from salmon that have been captured by angling in rivers. These can be read to differentiate between farmed escapees and wild fish (Fiske et al. 2006; Lund and Hansen 1991). Thereafter, these scales can be used to isolate DNA in order to address a wide variety of evolutionary and conservation questions (Glover et al. 2012; Karlsson et al. 2011; Nielsen and Hansen 2008; Nielsen et al. 1997). Recently, identification of triploid Atlantic salmon, using microsatellite DNA genotyping was validated against other triploid identification methods that require fresh tissues (Glover et al. 2015). Consequently, it is now possible to go back into historical scale archives in order to identify and investigate the frequency of triploid Atlantic salmon escapees in rivers.
The overall aim of the present study was to determine the frequency of triploid farmed salmon escapees in Norwegian rivers in the period 2007–2014, and, compare this to the observed frequency of triploid salmon in Norwegian farms in the same period (2 %) (Glover et al. 2015). In order to achieve this, we used microsatellite genotyping to determine the ploidy of 3794 farmed Atlantic salmon escapees captured in 17 Norwegian rivers.
Materials and methods
Samples
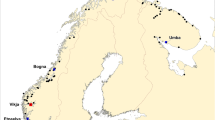
The samples of farmed escaped salmon, upon which the present study is based, originate from 17 rivers in the western part of Norway in the period 2007–2014 (Fig. 1). A description of the number of samples by year is available (Supplementary information 1).
The first and primary source of the samples is the historical scale archives owned by the Norwegian company Rådgivende Biologer AS. This company has conducted an extensive sampling of farmed and wild salmon in a variety of rivers in the west of Norway since the early 1990s. These samples are donated to the company by anglers who take the samples themselves once they have killed the fish for personal consumption. Thus, the samples were collected on a voluntary basis from dead fish killed as part of a licensed recreational activity (angling). The authors of this work have therefore not played any part in this specific sampling.
The second and minor source of samples for this study originate from farmed salmon captured and killed in a new trapping facility located in the river Etne in the west of Norway. The upstream trap is in place as part of a conservation plan to protect the local salmon population in the river from further genetic interactions with farmed escapees. The putatively identified farmed salmon (based upon morphological characteristics) are humanly killed by a sharp blow to the head. Thereafter, their identification is confirmed by scale reading (Lund and Hansen 1991).
Genotyping
DNA was isolated from 1 to 3 fish scales in 96 well format using either of the following two DNA isolation methods. The DNeasy blood and tissue kit from Qiagen, or the HotSHOT method (Truett et al. 2000). In both cases, either the manufacturers’ instructions or the publication´s standard description of the method were implemented. Each DNA extraction plate contained at least 2 blank cells as negative controls to enable their unique identification.
Eighteen microsatellite loci were amplified for each fish. These included the following loci amplified in three multiplex reactions (for specific PCR conditions see Supplementary information 2): SSsp3016 (Genbank no. AY372820), SSsp2210, SSspG7, SSsp2201, SSsp1605, SSsp2216 (Paterson et al. 2004), Ssa197, Ssa171, Ssa202 (O´Reilly et al. 1996), SsaD157, SsaD486, SsaD144 (King et al. 2005), Ssa289, Ssa14 (McConnell et al. 1995), SsaF43 (Sanchez et al. 1996), SsaOsl85 (Slettan et al. 1995), MHC I (Grimholt et al. 2002) MHC II (Stet et al. 2002). PCR products were analysed on an ABI 3730 Genetic Analyser and sized by a 500LIZ™ size-standard. Automatically binned alleles were manually checked by two researchers. All putatively identified triploid salmon were genotyped twice for validation. Individual salmon that were not successfully genotyped ≥10 of the 18 loci were removed from the data set (with the exception of one clearly triploid salmon genotyped for 9 loci, see Supplementary information 1).
Identified triploids (see below) were sexed genetically using the sdY gene (Eisbrenner et al. 2014; Yano et al. 2012). Individuals displaying amplicons of exon 2 and 4 were designated as males. Amplicons were identified on the ABI 3730 Genetic Analyser using identical conditions to amplify microsatellites described above.
Protocol for triploid identification using microsatellites
Microsatellite genotyping to identify triploid Atlantic salmon has recently been validated against flow cytometry and red blood cell diameter analyses (Glover et al. 2015). We have therefore used an identical protocol to identify the triploid salmon here. In short, we reported an individual as triploid where it displayed three alleles at two or more of the loci genotyped. Fish displaying more than 3 alleles were regarded as potentially polluted samples, and fish that displayed 3 alleles at just one of the loci were regarded as trisomic. Examples of how the genotyping data look in a normal diploid, a triploid and a trisomic salmon are presented in Supplementary information 3. The described approach to identify triploid salmon is also identical to a study of spontaneous triploids in cultured Atlantic salmon in a Baltic fish hatchery (Ozerov et al. 2010), and is similar to approaches used for identification of triploids in other organisms in the wild (Darvill et al. 2012; Garner et al. 2008; Hernandez-Urcera et al. 2012; Liebert et al. 2004).
Results
DNA was isolated from a total of 3941 samples collected from 17 rivers in the period 2007–2014. Of these, 3794 (96 %) individual samples were genotyped at 10 or more of the 18 microsatellite loci. The raw genotypic data for all of these 3794 individuals are provided (Supplementary information 1). Only these individuals are included in the present analyses. Sample drop-out is normal in all genetic data sets, and based upon previous studies where we have genotyped triploid salmon both in a pedigree based material (Solberg et al. 2013), and samples that were cross-validated against non-genetic ploidy determination methods (Glover et al. 2015), it is concluded that sample drop-out is random and not connected to ploidy.
A total of 7 (0.18 %) triploid, and 3 (0.08 %) trisomic farmed salmon escapees were observed among the samples captured in rivers (Tables 1, 2). Concentrating on the triploid escapees, these were only observed in four of the studied rivers, and only in the last years of the sampling period (i.e., 2010–2014). Both males (5) and females (2) were identified. Four of the escapees were captured in the river Oselva in 2014. The triploid escapees captured in the river Oselva in 2014 were fish of a similar size, and it is not unlikely that may have originated from the same farm source.
The exact within-river location of capture of the triploid escapees was available for five of the triploids observed in freshwater (Table 2). All five of these triploids were captured in the lower stretches of their respective rivers. In addition, all of the triploids with available data were less than 2 kg.
Discussion
This is the first study to investigate the relative frequency of diploid and triploid farmed Atlantic salmon escapees in rivers. Based upon genetic-ploidy determination of 3794 escapees re-captured in 17 rivers in the period 2007–2014, the following main results are presented: (1) Only 7 (0.18 %) of the escapees analysed were triploid, which is approximately 10-fold lower than the frequency observed in Norwegian farms in this time-period (2 %) (Glover et al. 2015). Working on the premise that ploidy does not influence likelihood of escape from a net pen, this study clearly demonstrates significantly reduced motivation of triploids to migrate into freshwater following escape from a farm when compared with diploid escapees. (2) Where the within-river location of capture was recorded, triploids were only observed in the lower sections or estuarine parts of the rivers. This further suggests reduced motivation to ascend rivers. (3) Triploids were only observed in three of the 8 sampling years, and only in 4 of the 17 rivers investigated. This suggests that entry of triploid escapees to freshwater is unpredictable, and that it is potentially triggered by specific conditions. Based upon these observations, we conclude that commercial production of triploid Atlantic salmon will not only function as an effective barrier to stop further gene flow to wild populations, it would reduce the numbers of farmed escapees entering rivers. In turn, this would reduce other environmental impacts on wild populations in freshwater, such as ecological interactions and possible disease transmission.
Genetic interactions between farmed escaped Atlantic salmon and wild conspecifics is a topic that has generated an entire field of research (Ferguson et al. 2007). Farmed salmon have probably been escaping from aquaculture installations since the very start of the industry in the early 1970s, and long-term studies have observed escapees in rivers over many years (Fiske et al. 2006). Both the pioneering studies conducted in Ireland (Clifford et al. 1998a, b; Crozier 1993), as well as more recent and extensive studies conducted in Norway (Glover et al. 2013, 2012; Skaala et al. 2006), have demonstrated genetic changes in native populations as a result of farmed escapees interbreeding. Furthermore, studies conducted in the wild have clearly demonstrated reduced fitness of the offspring of farmed escapees (Fleming et al. 2000; McGinnity et al. 2003; Skaala et al. 2012), and the underlying genomic causes of this are starting to be identified (Besnier et al. 2015). Clearly, while a switch to triploid production will not be able to reverse introgression that has already occurred, hindering further erosion of genetic integrity in native populations is important. This is especially so given the fact that domestication of farmed Atlantic salmon is ongoing, and the genetic differences between farmed and wild salmon in important fitness related traits, for example growth, continues to diverge with successive generations (Glover et al. 2009; Solberg et al. 2012).
The results of the present study, together with an earlier study involving release of triploid and diploid smolt groups in Ireland (Cotter et al. 2000), strongly suggest that commercial production of triploid salmon will not only hinder further genetic interactions with wild populations, it is likely that the frequency of escapees in rivers will drop significantly due to their apparent reduced motivation for freshwater migration. Thus, triploid production should also be viewed as a strategy to limit other interactions with both juvenile and adult wild salmon in rivers, such as spawning competition and potential disease transmissions. For example, a recent study has demonstrated that virus-infected farmed escaped salmon enter rivers, and thus represent a potential disease transmission threat to wild salmon populations (Madhun et al. 2015). If the salmon aquaculture industry was based upon production of triploid salmon, then the opportunity for transmission of diseases to juvenile and young fish in rivers may be significantly reduced as a result of their limited motivation for migration into freshwater.
The low number of triploid escapees observed in freshwater in this study precluded the ability to determine trends of this occurrence. However, despite the fact that secondary sexual characteristics were not recorded for these triploid fish, their size and capture location within the rivers (lower stretches) is typical of immature farmed Atlantic salmon that enter freshwater soon after escape from a farm (Madhun et al. 2015; Skilbrei et al. 2015a, b). Thus, while triploid male salmon that have developed secondary sexual characteristics can partake in spawning (Fjelldal et al. 2014), the evidence presented here indicates that where triploids enter freshwater, their distribution in the river is typical of immature diploid escapees, and it is unlikely that they would attempt to partake in and therefore disrupt wild fish spawning. We cannot exclude the possibility that some individuals would nevertheless migrate further upstream and attempt to spawn.
In the period in which the present study was conducted, almost no deliberate commercial production of triploid Atlantic salmon occurred. However, recent developments in production techniques have solved some of the early challenges experienced with commercial production of triploids (Fjelldal and Hansen 2010; Fraser et al. 2012; Frenzl et al. 2014; Leclercq et al. 2011; Taylor et al. 2015). In Norway, which is the world´s largest Atlantic salmon producing country, recent government licensing schemes have encouraged companies to initiate triploid production at a commercial level. It is therefore likely that the frequency of triploid Atlantic salmon production in Norway will expand in the coming years. The results of this study support this development in order to hinder further genetic introgression as well as potential ecological and disease interactions with wild populations in rivers.
References
Besnier F, Glover KA, Lien S, Kent M, Hansen MM, Shen X, Skaala O (2015) Identification of quantitative genetic components of fitness variation in farmed, hybrid and native salmon in the wild. Heredity 115:47–55. doi:10.1038/hdy.2015.15
Clifford SL, McGinnity P, Ferguson A (1998a) Genetic changes in an Atlantic salmon population resulting from escaped juvenile farm salmon. J Fish Biol 52:118–127
Clifford SL, McGinnity P, Ferguson A (1998b) Genetic changes in Atlantic salmon (Salmo salar) populations of northwest Irish rivers resulting from escapes of adult farm salmon. Can J Fish Aquat Sci 55:358–363
Cotter D, O’Donovan V, O’Maoileidigh N, Rogan G, Roche N, Wilkins NP (2000) An evaluation of the use of triploid Atlantic salmon (Salmo salar L.) in minimising the impact of escaped farmed salmon on wild populations. Aquaculture 186:61–75. doi:10.1016/s0044-8486(99)00367-1
Crozier WW (1993) Evidence of genetic interaction between escaped farmed salmon and wild Atlantic salmon (Salmo salar L) in a Northern Irish river. Aquaculture 113:19–29
Darvill B, Lepais O, Woodall LC, Goulson D (2012) Triploid bumblebees indicate a direct cost of inbreeding in fragmented populations. Mol Ecol 21:3988–3995. doi:10.1111/j.1365-294X.2012.05679.x
Eisbrenner WS et al (2014) Evidence for multiple sex-determining loci in Tasmanian Atlantic salmon (Salmo salar). Heredity 113:86–92. doi:10.1038/hdy.2013.55
Ferguson A, Fleming IA, Hindar K, Skaala Ø, McGinnity P, Cross T, Prodohl P (2007) Farm escapees. In: Verspoor E, Stradmeyer L, Nielsen JL (eds) The Atlantic salmon. Genetics, conservation and management. Blackwell, Oxford, pp 357–398
Fiske P, Lund RA, Hansen LP (2006) Relationships between the frequency of farmed Atlantic salmon, Salmo salar L., in wild salmon populations and fish farming activity in Norway, 1989–2004. ICES J Mar Sci 63:1182–1189. doi:10.1016/j.icesjms.2006.04.006
Fjelldal PG, Hansen T (2010) Vertebral deformities in triploid Atlantic salmon (Salmo salar L.) underyearling smolts. Aquaculture 309:131–136. doi:10.1016/j.aquaculture.2010.09.027
Fjelldal PG, Wennevik V, Fleming IA, Hansen T, Glover KA (2014) Triploid (sterile) farmed Atlantic salmon males attempt to spawn with wild females Aquaculture Environment. Interactions 5:155–162. doi:10.3354/aei00102
Fleming IA, Hindar K, Mjolnerod IB, Jonsson B, Balstad T, Lamberg A (2000) Lifetime success and interactions of farm salmon invading a native population. Proc R Soc Lond Ser B-Biol Sci 267:1517–1523
Fraser TWK, Fjelldal PG, Hansen T, Mayer I (2012) Welfare considerations of triploid fish. Rev Fish Sci 20:192–211. doi:10.1080/10641262.2012.704598
Frenzl B et al (2014) Triploid and diploid Atlantic salmon show similar susceptibility to infection with salmon lice Lepeophtheirus salmonis. Pest Manag Sci 70:982–988. doi:10.1002/ps.3639
Garner SR, Madison BN, Bernier NJ, Neff BD (2008) Juvenile growth and aggression in diploid and triploid Chinook salmon Oncorhynchus tshawytscha (Walbaum). J Fish Biol 73:169–185. doi:10.1111/j.1095-8649.2008.01923.x
Glover KA et al (2015) The frequency of spontaneous triploidy in farmed Atlantic salmon produced in Norway during the period 2007–2014. BMC Genet. doi:10.1186/s12863-015-0193-0
Glover KA, Ottera H, Olsen RE, Slinde E, Taranger GL, Skaala O (2009) A comparison of farmed, wild and hybrid Atlantic salmon (Salmo salar L.) reared under farming conditions. Aquaculture 286:203–210. doi:10.1016/j.aquaculture.2008.09.023
Glover KA, Quintela M, Wennevik V, Besnier F, Sørvik AGE, Skaala O (2012) Three decades of farmed escapees in the wild: a spatio-temporal analysis of population genetic structure throughout Norway. PLoS ONE 7(8):e43129
Glover KA, Pertoldi C, Besnier F, Wennevik V, Kent M, Skaala Ø (2013) Atlantic salmon populations invaded by farmed escapees: quantifying genetic introgression with a Bayesian approach and SNPs. BMC Genet 14:4. doi:10.1186/1471-2156-14-74
Grimholt U, Drablos F, Jorgensen SM, Hoyheim B, Stet RJM (2002) The major histocompatibility class I locus in Atlantic salmon (Salmo salar L.): polymorphism, linkage analysis and protein modelling. Immunogenetics 54:570–581. doi:10.1007/s00251-002-0499-8
Hernandez-Urcera J, Vera M, Magadan S, Pino-Querido A, Cal RM, Martinez P (2012) Development and validation of a molecular tool for assessing triploidy in turbot (Scophthalmus maximus). Aquaculture 330:179–184. doi:10.1016/j.aquaculture.2011.11.039
Karlsson S, Moen T, Lien S, Glover KA, Hindar K (2011) Generic genetic differences between farmed and wild Atlantic salmon identified from a 7 K SNP-chip. Mol Ecol Resour 11:247–253. doi:10.1111/j.1755-0998.2010.02959.x
King TL, Eackles MS, Letcher BH (2005) Microsatellite DNA markers for the study of Atlantic salmon (Salmo salar) kinship, population structure, and mixed-fishery analyses. Mol Ecol Notes 5:130–132. doi:10.1111/j.1471-8286.2005.00860.x
Leclercq E, Taylor JF, Fison D, Fjelldal PG, Diez-Padrisa M, Hansen T, Migaud H (2011) Comparative seawater performance and deformity prevalence in out-of-season diploid and triploid Atlantic salmon (Salmo salar) post-smolts. Comp Biochem Physiol A-Mol Integr Physiol 158:116–125. doi:10.1016/j.cbpa.2010.09.018
Liebert AE, Johnson RN, Switz GT, Starks PT (2004) Triploid females and diploid males: underreported phenomena in Polistes wasps? Insect Soc 51:205–211. doi:10.1007/s00040-004-0754-0
Lund RA, Hansen LP (1991) Identification of wild and reared Atlantic salmon, Salmo salaar L., using scale characters. Aquac Fish Manage 22:499–508
Lura H, Saegrov H (1991) Documentation of successful spawning of escaped farmed female Atlantic salmon, Salmo salar, in Norwegian rivers. Aquaculture 98:151–159
Madhun AS et al (2015) Potential disease interaction reinforced: double-virus infected escaped farmed Atlantic salmon, Salmo salar L., recaptured in a nearby river. J Fish Dis 38:209–219
McConnell SK, Oreilly P, Hamilton L, Wright JN, Bentzen P (1995) Polymorphic microsatellite loci from Atlantic salmon (Salmo salar)—genetic differentiation of North-American and European populations. Can J Fish Aquat Sci 52:1863–1872
McGinnity P et al (2003) Fitness reduction and potential extinction of wild populations of Atlantic salmon, Salmo salar, as a result of interactions with escaped farm salmon. Proc R Soc Lond Ser B-Biol Sci 270:2443–2450. doi:10.1097/rspb.2003.2520
Nielsen EE, Hansen MM (2008) Waking the dead: the value of population genetic analyses of historical samples. Fish Fish 9:450–461. doi:10.1111/j.1467-2979.2008.00304.x
Nielsen EE, Hansen MM, Loeschcke V (1997) Analysis of microsatellite DNA from old scale samples of Atlantic salmon Salmo salar: a comparison of genetic composition over 60 years. Mol Ecol 6:487–492
O’Reilly PT, Hamilton LC, McConnell SK, Wright JM (1996) Rapid analysis of genetic variation in Atlantic salmon (Salmo salar) by PCR multiplexing of dinucleotide and tetranucleotide microsatellites. Can J Fish Aquat Sci 53:2292–2298
Ozerov MY et al (2010) High Gyrodactylus salaris infection rate in triploid Atlantic salmon Salmo salar. Dis Aquat Organ 91:129–136. doi:10.3354/dao02242
Paterson S, Piertney SB, Knox D, Gilbey J, Verspoor E (2004) Characterization and PCR multiplexing of novel highly variable tetranucleotide Atlantic salmon (Salmo salar L.) microsatellites. Mol Ecol Notes 4:160–162. doi:10.1111/j.1471-8286.2004.00598.x
Saegrov H, Hindar K, Kalas S, Lura H (1997) Escaped farmed Atlantic salmon replace the original salmon stock in the River Vosso, western Norway. ICES J Mar Sci 54:1166–1172
Sanchez JA, Clabby C, Ramos D, Blanco G, Flavin F, Vazquez E, Powell R (1996) Protein and microsatellite single locus variability in Salmo salar L (Atlantic salmon). Heredity 77:423–432
Skaala O, Wennevik V, Glover KA (2006) Evidence of temporal genetic change in wild Atlantic salmon, Salmo salar L., populations affected by farm escapees. ICES J Mar Sci 63:1224–1233. doi:10.1016/j.icesjms.2006.04.005
Skaala Ø, Glover Kevin A, Barlaup Bjørn T, Svåsand T, Besnier F, Hansen Michael M, Borgstrøm R (2012) Performance of farmed, hybrid, and wild Atlantic salmon (Salmo salar) families in a natural river environment. Can J Fish Aquat Sci 69:1994–2006. doi:10.1139/f2012-118
Skilbrei OT, Heino M, Svåsand T (2015a) Using simulated escape events to assess the annual numbers and destinies of escaped farmed Atlantic salmon of different life stages, from farms sites in Norway. ICES J Mar Sci 72:670–685
Skilbrei OT, Normann ES, Meier S, Olsen RE (2015b) Use of fatty acid profiles to monitor the escape history of farmed Atlantic salmon. Aquac Environ Interact 7:1–13
Slettan A, Olsaker I, Lie O (1995) Atlantic salmon, Salmo salar, microsatellites at the SsOSL25, SsOSL85, SsOSL311, SsOSL417 loci. Anim Genet 26:281–282
Solberg MF, Kvamme BO, Nilsen F, Glover KA (2012) Effects of environmental stress on mRNA expression levels of seven genes related to oxidative stress and growth in Atlantic salmon Salmo salar L. of farmed, hybrid and wild origin. Bmc Res Notes 5:672
Solberg MF, Glover KA, Nilsen F, Skaala Ø (2013) Does domestication cause changes in growth reaction norms? A study of farmed, wild and hybrid atlantic salmon families exposed to environmental stress. Plos One 8(1):e54469
Stet RJM, de Vries B, Mudde K, Hermsen T, van Heerwaarden J, Shum BP, Grimholt U (2002) Unique haplotypes of co-segregating major histocompatibility class II A and class II B alleles in Atlantic salmon (Salmo salar) give rise to diverse class II genotypes. Immunogenetics 54:320–331. doi:10.1007/s00251-002-0477-1
Taranger GL et al (2015) Risk assessment of the environmental impact of Norwegian Atlantic salmon farming Ices. J Marine Sci 72:997–1021. doi:10.1093/icesjms/fsu132
Taylor JF et al (2015) Adult triploid Atlantic salmon (Salmo salar) have higher dietary histidine requirements to prevent cataract development in seawater. Aquac Nutr 21:18–32. doi:10.1111/anu.12130
Taylor JF, Bozzolla P, Frenzl B, Matthew C, Hunter D, Migaud H (2014) Triploid Atlantic salmon growth is negatively affected by communal ploidy rearing during seawater grow-out in tanks. Aquaculture 432:163–174. doi:10.1016/j.aquaculture.2014.05.014
Truett GE, Heeger P, Mynatt RL, Truett AA, Walker JA, Warman ML (2000) Preparation of PCR-quality mouse genomic DNA with hot sodium hydroxide and tris (HotSHOT) Biotechniques 29:52-+
Yano A et al (2012) An immune-related gene evolved into the master sex-determining gene in rainbow trout. Oncorhynchus Mykiss Curr Biol 22:1423–1428. doi:10.1016/j.cub.2012.05.045
Acknowledgments
This study was financed using resources from the Norwegian Department of Industry and Fisheries and the Norwegian Research Council funded project INTERACT. Jim B. Bos was partially financed by an Erasmus mobility grant and Yongkai Tang was jointly-financed by the Chinese Scholarship Council and a NFR mobility grant between China and Norway. We would like to thank the anglers who have taken the vast majority of samples upon which the present study was made possible.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Glover, K.A., Bos, J.B., Urdal, K. et al. Genetic screening of farmed Atlantic salmon escapees demonstrates that triploid fish display reduced migration to freshwater. Biol Invasions 18, 1287–1294 (2016). https://doi.org/10.1007/s10530-016-1066-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-016-1066-9