Abstract
A longstanding assumption of glucose tracing experiments is that all glucose is microbially utilized during short incubations of ≤2 days to become microbial biomass or carbon dioxide. Carbon use efficiency (CUE) estimates have consequently ignored the formation of residues (non-living microbial products) although such materials could represent an important sink of glucose that is prone to stabilization as soil organic matter. We examined the dynamics of microbial residue formation from a short tracer experiment with frequent samplings over 72 h, and conducted a meta-analysis of previously published glucose tracing studies to assess the generality of these experimental results. Both our experiment and meta-analysis indicated 30–34% of amended glucose-C (13C or 14C) was in the form of residues within the first 6 h of substrate addition. We expand the conventional efficiency calculation to include residues in both the numerator and denominator of efficiency, thereby deriving a novel metric of the potential persistence of glucose-C in soil as living microbial biomass plus residues (‘carbon stabilization efficiency’). This new metric indicates nearly 40% of amended glucose-C persists in soil 180 days after amendment, the majority as non-biomass residues. Starting microbial biomass and clay content emerge as critical factors that positively promote such long term stabilization of labile C. Rapid residue production supports the conclusion that non-growth maintenance activity can illicit high demands for C in soil, perhaps equaling that directed towards growth, and that residues may have an underestimated role in the cycling and sequestration potential of C in soil.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
As the largest pool of carbon (C) in the biosphere, soil organic matter (SOM) and its transformation represent a critical component of global C cycling. Carbon entering the soil is primarily plant-derived but subsequent processing relies heavily on microorganisms. Microbial decomposition and metabolism convert organic C into biosynthetic molecules that may be actively recycled, become stable SOM resistant to decay, or lost from soil as respiration. Understanding the conditions which drive this partitioning can help to forecast depletion of soil C stocks and carbon dioxide (CO2) efflux, particularly as climate stressors such as warming amplify rates of belowground C transformation (Melillo et al. 2017; Kuzyakov et al. 2019).
Substantial insights into the processing of C in soil have been gained using isotopic tracer studies. Labeled substrates, particularly glucose, are used in tracer experiments to determine how quickly microbial uptake occurs, whether the added substrate is used anabolically or mineralized for energy generation, and what the dynamics of SOM stabilization in soil are like. The mean residence time of glucose in the soil solution is typically seconds to hours (Boddy et al. 2007; Hill et al. 2008; Fischer et al. 2010; Leitner et al. 2012; Gunina and Kuzyakov 2015). Glucose itself appears unlikely to become stabilized on mineral or organic surfaces via abiotic reactions, given rapid microbial uptake and the uncharged nature of the molecule (Gunina and Kuzyakov 2015). Mean residence time of tracer C in microbial biomass depends on the turnover time of the microbial community, typically 1–10 days for active bacteria and 100–300 days for fungi, although the range is large (Rousk and Baath 2011; Koch et al. 2018). Rates of tracer C loss from soil will peak within days of amendment because of microbial respiratory demands and thereafter decline as stabilization of microbial residues occurs (e.g., up to 20% of the original added C stabilized in SOM after 3 years; Cheshire 1979).
Glucose C was originally assumed to be partitioned only between microbial biomass and respiration during short (≤2 day) incubations (Dictor et al. 1998). However, it was eventually observed that a substantial portion of the isotope remained unextractable from soil by liquid extraction. This missing C was attributed to microbial biomass unextractable after chloroform fumigation. Extraction coefficients (KEC) were thus developed from short glucose tracing experiments (i.e., the in situ labeling method using 13C or 14C) (Sparling and West 1988; Joergensen 1996) to correct fumigation-based estimates of microbial biomass. In situ labeling as well as other calibration techniques have produced KEC values of 0.35–0.45, suggesting that 55–65% of isotopic label becomes “stuck” in unextractable forms of living biomass like bacterial cell walls and spores (Lynch and Panting 1980). Although this assumption that all unextractable C is living microbial biomass is “probably wrong” (Joergensen 1996; Dictor et al. 1998; Jenkinson et al. 2004), no attempt has been made to clarify the issue or assess its impacts on soil biogeochemistry such as microbial biomass determination and substrate utilization.
Microbial residues are an alternative explanation for unextractable tracer C. We operationally define residues as any non-living organic material of microbial origin including extracellular enzymes, antibiotics, exopolysaccharides, and necromass (Gregorich et al. 1991; Mueller et al. 1998; Costa et al. 2018). Such materials may be bound to the cell surface (e.g., lipopolysaccharides), exist as dissolved organic C, or be associated with soil particles via organo-mineral complexes. Given their varied forms, residues are often quantified indirectly as the amount of label in soil minus that in microbial biomass (Gregorich et al. 1991; Gunina and Kuzyakov 2015; Moreno-Cornejo et al. 2015; Creamer et al. 2016):
As SOM precursors, residues effectively link the dynamics of microbial activity on hourly time scales with the capacity of soil to build C stocks of much longer residence time (Allison et al. 2010; Kallenbach et al. 2015).
Ignoring residue production, particularly over short time scales, has two major consequences. The first is overestimation of living microbial biomass. MBC is overestimated by over two times when a KEC factor of 0.45 is used (e.g., = 1/0.45 = 2.22) if unextractable C is non-living residues rather than biomass. Second, microbial carbon use efficiency estimation (CUE; the proportion of total C uptake that microbes partition towards growth) is also predicated on an assumption of no residue production during short (≤2 day) incubation. A conventional calculation of CUE is as follows:
The denominator of Eq. (2) would be an unreliable estimate of total uptake if additional tracer sinks like non-biomass residues exist beyond MBC and CO2. Whether residue production occurs during short incubations remains unknown; the few studies to have examined the temporal dynamics of residue formation have been over much longer time scales and none have assessed the potential impact on microbial CUE.
Here we quantitatively assess glucose partitioning in soil to address whether residue formation may occur during short incubations, and the impacts that residue formation would have for estimation of CUE in soil. We conducted a laboratory experiment to measure glucose partitioning among major pools, including residues, and a subsequent meta-analysis to assess the generality of our experimental results.
Methods
Glucose tracing experiment
A tracing experiment was conducted to examine the partitioning of glucose into major soil pools within hours of amendment. A representative temperate forest soil was collected from the Harvard Forest Long Term Ecological Research (LTER) site in Petersham, MA, USA. Mineral soil (0–15 cm depth) below the organic horizon of the Gloucester series (fine loamy, mixed, mesic, typic Dystrochrepts) was collected, sieved (<2 mm), and stored at 4 °C until experimental use. Gravimetric water content (0.47 g g−1) was determined by drying at 105 °C for 24 h. Field capacity (0.90 g g−1) was determined from the gravimetric water content of saturated soil after gravity draining for 48 h. Soil organic C (5.7% of dry soil), microbial biomass concentrations (1% of SOC), pH (4.2) and C:N ratio (24) were typical for these soils (Frey et al. 2014). All soil was pre-incubated for 48 h at 25 °C before glucose addition.
Labeled glucose addition to soil allows for the determination of substrate-C uptake and mineralization, where microbial growth is inferred from 13C incorporation into microbial biomass. Five replicate soil samples (40 g each) and one soil control (nutrient solution without glucose) were prepared for each target harvest time of 6, 12, 18, 24, 32, 40, 48, 60, 72 h. Unlabeled glucose was mixed with 99 at% U-13C glucose to achieve total enrichment of 5 at%. The amendment solution delivered 0.05 mg glucose-C g−1 dry soil, along with trace nutrients (0.1% MgCl2, 0.2% KH2PO4, and 0.1% K2SO4) and sufficient sodium nitrate to achieve a molar C:N ratio = 40 (Wadso 2009). This experimental design was replicated for a second amendment concentration of 2.0 mg glucose-C g−1 dry soil and additional nitrogen to maintain a C:N ratio = 40 of solution. Amendments were briefly mixed by spatula into the soil, increasing soil moisture to 65% of field capacity. The lower glucose amendment was chosen to represent a typical substrate amendment rate used in other tracer experiments (Frey et al. 2013); the higher glucose amendment was chosen to induce logistic growth (Reischke et al. 2014).
Specimen cups containing the soil were parafilm covered and incubated at 25 °C until harvesting. Ninety minutes prior to harvest, cups were uncovered and sealed within 1 L jars fitted with rubber septa. Jars were flushed for 15 min with CO2-free air. At the time of harvest, 15 mL of headspace was sampled via syringe and injected into evacuated exetainers (12 mL). Jars were then opened and the soil immediately extracted for microbial biomass by chloroform fumigation extraction (Vance et al. 1987). Briefly, 15 g of each sample was fumigated with ethanol-free chloroform for 24 h, after which the sample was extracted in 0.05 M K2SO4. A paired 15 g sample was immediately extracted without fumigation. Total dissolved organic C (DOC) extracts were frozen at −20 °C and shipped, along with headspace samples, to the University of California (Davis) Stable Isotope Facility (Davis, CA, USA) for total C and 13C quantification. The difference in total DOC concentration between fumigated and non-fumigated samples was used to calculate microbial biomass C. Remaining soil was frozen at −20 °C and later dried and finely ground for determination of total soil C and N, and 13C quantification at the University of New Hampshire Stable Isotope Facility (Durham, NH, USA).
All estimates of growth, respiration, and CUE were calculated using published mixing models and equations (Brant et al. 2006). Total microbial growth (13MBC; μg C g−1 soil) was calculated as the product of total microbial biomass (F DOC–NF DOC; μg C g−1 soil) and the percent of total microbial biomass labeled (%13MBC):
where at% F DOC, F DOC, at% NF DOC, and NF DOC represent the atom % and total C concentrations (μg C g−1 soil) of fumigated (F) and non-fumigated (NF) K2SO4 extracts, respectively. at% MBCt and at% MBCc are the atom % of sample treatments and natural abundance controls, and at% sol is the atom % of amendment solution (5 at%). A typical extraction efficiency of 0.45 was applied to account for unextractable biomass (Joergensen 1996; Jenkison et al. 2004). Finally, CUE was calculated using Eq. 2 where 13CO2 is the cumulative respiration derived from added glucose (μg 13CO2–C g−1 soil).
Meta-analysis
We conducted a meta-analysis of previously published data to assess the generality of our experimental results across soil types and study-specific treatments. For our purposes, we included only those studies that provided sufficient detail to construct a budget that could account for ~100% of the amended glucose tracer.
The search for relevant studies was conducted using the online ISI Web of Science Core Collection in September 2018. Search criteria were “C-13 AND glucose AND soil” (264 results) and “C-14 AND glucose AND soil” (378 results). Studies from this list also needed to meet the following criteria to be included in the analysis: (1) at least two of the three major tracer sinks (CO2, MBC, bulk soil) were reported so that a budget could be reconstructed and residue amounts calculated using Eq. 1, (2) MBC estimates were made using a fumigation-extraction technique and a KEC value <1 applied to account for unextractable biomass, (3) incubation times did not exceed 180 days. This time frame was chosen as net movement of tracer plateaued within 50 days of amendment. Search criteria resulted in a final list of 315 observations from 18 studies. Data presented in graphical form were gleaned using PlotDigitizer X (v. 2.0.1). Tracer remaining in the DOC pool was assumed to be negligible unless otherwise stated since the majority of studies reported undetectable concentrations at harvest. The Satterthwaite method was used to generate pooled variance estimates when tracer concentrations were estimated from other data (e.g., % tracer in CO2 = 100% − % tracer in soil) (Satterthwaite 1946).
The primary response variables in the meta-analysis dataset (percent of tracer in MBC, soil, CO2, and residue pools and CUE) were asymptotic over time. Non-linear least squares modeling was conducted using a self-starting asymptotic R function (SSasymp) to assess model fit using the entire dataset of 0–180 incubation days. The data was further subset into observations ≤2 days (the length of a typical CUE assay) and >24 days (when response variables had generally reached an asymptote). Subsetting in this manner allowed us to control for incubation length during some analyses.
The denominator of CUE is intended to capture all C uptake by microbes. This term may be underestimated if some C is used for non-biomass residue formation (e.g., Eq. 1). We developed a second CUE metric (“CUEr”) to additionally account for any residues as a portion of microbial C uptake:
A metric of carbon stabilization efficiency (“CSE”) was also formulated to express the potential of a soil to stabilize all microbial products, whether biomass or residue:
We assume that all tracer C is rapidly utilized such that a term for the original, unprocessed substrate is unnecessary. Similar to the ‘carbon sequestration efficiency’ of Gleixner (2013) or CUETP (total microbial products) of Joergensen and Wichern (2018), CSE provides a meaningful link between microbial metabolism and SOM formation at ecosystem scales.
Three independent variables were examined as predictors of glucose partitioning and efficiency: starting MBC at the onset of experimentation, glucose amendment concentration, and soil clay content. Linear mixed effects, partial regression models were fit to the subsetted data (≤2 and >24 incubation days) and full 180-day dataset with random terms to account for non-independence of results by publication source. Partial regression allowed the removal of (1) clay and starting MBC effects from one another, as a positive relationship was observed between these predictors and (2) the effect of time from the full 180-day dataset. Normality of residuals and homogeneity of variance were tested and transformations applied if failed. Multiple linear regression was not used as only a small number of observations (n = 29) had all three drivers reported.
In addition to the mixed effects modeling, meta-analysis results were analyzed following the recommendations outlined by Viechtbauer (2010) for the ‘metafor’ R package. The effect size and variance of mean value responses were estimated using the ‘escalc’ function. These data were fitted to multivariate meta-analytic models with the rma.mv function using a restricted maximum likelihood method and a random term to account for clustering of the data by publication source. Normality of model residuals was assessed with the Shapiro test and, if failed, a transformation applied to the response variable. Publication bias was also assessed using the ‘ranktest’ and ‘weightfunct’ functions. In most cases, fewer observations existed for meta-analysis than with comparable mixed effects modeling because variance estimates also had to be reported from the literature. As the results from mixed effects and meta-analytical approaches were similar, only the former are presented. All stats were performed in R (v. 3.6.0).
Results
Glucose tracing experiment
Quantification of 13C in the pools of CO2, MBC, DOC, and bulk soil C indicates relatively stable tracer concentrations throughout the 72-h incubation following 0.05 mg glucose-C g−1 dry soil addition (Fig. 1). No tracer was found in the DOC pool even after the shortest incubation interval of 6 h, suggesting rapid microbial uptake of added substrate. Steady concentrations in microbial biomass (~45% of tracer applied) throughout the incubation suggest that any accumulation of tracer C in the MBC pool was being offset by losses from respiration, biomass turnover, or exudation. Even after an extraction coefficient of 0.45 was applied to MBC estimates, the residue pool (13Soil C – 13MBC; gray shaded area Fig. 1) represented ~40% of tracer C remaining in the soil within 6 h of incubation, or about 30% of the total glucose added. Carbon use efficiency (Eq. 2) significantly declined during incubation from 0.86 to 0.74 (p < 0.001).
Glucose-derived C in the pools of respiration (CO2), soil C, microbial biomass C (MBC), and dissolved organic C (DOC) over 72 h after amendment with 0.05 or 2.0 mg glucose-C g−1 dry soil. Measurements were made after 6, 12, 18, 24, 32, 40, 48, 60, and 72 h of incubation with tracer. Error bars are one standard deviation. 13C respiration is cumulative 13CO2 emission, depicted here as the excess beyond 13C remaining in soil (pink shaded region). Microbial residues are calculated as the difference between 13C in soil and the sum of (13MBC + 13DOC) (gray shaded region). (Color figure online)
Higher glucose amendment (2.0 mg glucose-C g−1 dry soil) substantially altered tracer dynamics. A logistic pattern of growth occurred as evident from (1) a lag phase of ~24 h with insignificant tracer found as MBC or CO2 followed by (2) a depletion of tracer from DOC and appearance in MBC and CO2 as microbial growth was initiated (Fig. 1). Evidence for logistic growth is supported by 18O-water tracing that was conducted simultaneously (Geyer et al. 2019). All tracer C in soil was in the form of DOC (extractable by K2SO4) during the lag phase suggesting glucose was not microbially utilized or abiotically stabilized via reaction with native SOM or other means. A residue pool was not observed during the lag phase but developed rapidly once microbial growth was initiated, reaching ~50% of the initial glucose amendment after 72 h of incubation. This indicates that residue formation is linked to microbial activity.
Meta-analysis
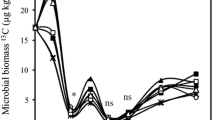
Meta-analysis of 18 glucose tracing studies supports the generality of our laboratory experiment. In the initial hours of incubation, 42% (±2.12 S.E.) of tracer was in MBC, 34% (±1.07 S.E.) was found as residue, and 13% (±1.91 S.E.) had been mineralized to CO2 (Table 1, y-intercepts). The proportion of tracer in soil, MBC, and residues significantly declined during the first days of incubation, whereas the proportion in CO2 increased, all significantly fit by asymptotic functions (p < 0.05). Changes in tracer concentrations among pools largely stopped within 24 days of incubation although the percent in residues continued to slowly decline through the 180 days of incubation surveyed. The majority of tracer was ultimately lost from soil as CO2 (58% ± 0.78 S.E.), with the remaining left in soil as MBC (11% ± 1.11 S.E.) or residues (19% ± 9.06 S.E.) after 180 days (Fig. 2).
The partitioning of glucose-derived C (percent of C added) over 180 incubation days between the pools of respiration (CO2), soil C, microbial biomass C (MBC), and residue C (=Soil C – MBC). Data represents 315 observations from 18 published studies (Supp. Table 1). Statistics generated from non-linear least squares modeling of responses over time. *p < 0.05; **p < 0.01; ***p < 0.001
Similar to our experimental results, the meta-analysis indicates accumulation of tracer as residues within hours of glucose addition. This occurred despite the application of extraction coefficients in all studies to account for unextractable C as microbial biomass. Extraction coefficients ranged from 0.29 to 0.45 (mean = 0.41) among the 18 studies we examined. Of the 40 observations where incubations lasted ≤2 days, only two used an extraction coefficient that fully accounted for unextractable C as biomass such that no reside was found [i.e., Eq. (1) = 0]. Extraction coefficients would need to be smaller to adequately fully account for all unextractable C as living biomass in soil, or a significant percent of tracer accumulated as non-biomass residues.
CUE estimates were impacted by the inclusion of residues in the calculation (Fig. 3). Average CUE using a conventional ratio ignoring residues [Eq. (2)] was 0.78 (±0.03 S.E.) during the shortest incubations (Table 1, y-intercept). CUEr, where residues have been added to the denominator to account for all microbial uptake, had significantly lower efficiency of 0.46 (±0.02 S.E.) at the onset of incubation. Incorporating residues into the CUE denominator and numerator as CSE resulted in significantly higher efficiency of 0.83 (±0.02 S.E.) during the same time period. All three CUE metrics declined over time and reached significantly different asymptotes (Fig. 3). While CUE and CUEr declined to ~0.15, CSE remained relatively high at 0.40 (±0.01 S.E.). Thus, 40% of tracer appears to remain in soil as microbial products (living or dead) after 180 days, presumably as precursors to stable SOM.
Trends in carbon use efficiency (CUE) over 180 days after glucose tracer addition. Data represents 315 observations from 18 published studies (Supp. Table 1). Solid line fit to green circles: CUE (13 or 14MBC/(13 or 14MBC + 13 or 14CO2)); dashed line fit to red squares: CUEr (13 or 14MBC/(13 or 14MBC + 13 or 14CO2 + 13 or 14Residue C)); dash-dot line fit to blue triangles: CSE ((13 or 14MBC + 13 or 14Residue C)/(13 or 14MBC + 13 or 14CO2 + 13 or 14Residue C)). Inset expands detail from the first 50 incubation days. (Color figure online)
The amount of starting microbial biomass and soil clay content were particularly influential predictors of tracer partitioning according to regression analysis (Table 1). Starting microbial biomass (757.7 μg C g−1 ± 32.3 S.E., n = 192) had significant positive effects on tracer accumulation in the microbial biomass and soil pools, but negative effects on accumulation as residues or CO2. Clay content (18.9% ± 1.25 S.E., n = 135) had significant positive effects on tracer presence in both soil and residue pools, but a negative effect on accumulation in microbial biomass and C mineralization. Glucose amendment level (899.6 μg C g−1 ± 56.1 S.E., n = 315) had weak but significant positive effects on tracer mineralization at the expense of presence in soil or residues. These effects on tracer partitioning resulted in starting MBC being the strongest predictor of CUE, while glucose amendment concentration had a weaker, negative effect. Starting MBC was also the strongest predictor of CUEr and CSE, although clay content was a significant positive predictor of the latter as well. Approximately 70% and 76% of the variability in CUE and CUEr was explained by starting MBC alone, respectively, while 41% of the variability in CSE was due to starting MBC (26%) and clay content (15%).
Discussion
Glucose tracing experiments have traditionally assumed that tracer C is partitioned only between microbial biomass and CO2 pools during short incubations ≤2 days (Joergensen 1996; Dictor et al. 1998; Jenkinson et al. 2004). Microbial residues, operationally defined as non-living organic C of microbial origin, have been suggested to accumulate only later in time as necromass accumulates or cells exude products to favorably modify their environment for growth. As a result, CUE and extraction coefficients used in the estimation of microbial biomass do not account for residues. Given recent evidence for the largely microbial provenance of SOM (Schmidt et al. 2011; Miltner et al. 2012; Kallenbach et al. 2016), residues should no longer be overlooked.
We found from our lab experiment that 30% of amended C is present as microbial residues within 6 h of tracer amendment, much sooner than is generally assumed. Meta-analysis of 18 previously published glucose tracing studies corroborates this result: ~34% of tracer C was found in the form of residues within minutes of amendment, nearly as much as in microbial biomass. Contrary to expectation, residues also appear to decline over time after substrate addition rather than accrue. These results suggest that soil microorganisms have the potential to rapidly synthesize and exude substrate C and/or that biomass turnover generates necromass-derived residues almost immediately after tracer addition. Given that the estimated lifespan of soil bacteria may average roughly two days (Rousk and Baath 2011; Koch et al. 2018), it should perhaps be expected that residues could form this quickly. Rapid residue formation is also consistent with the prioritization of maintenance activities before growth; many residue products like exoenzymes and capsular polysaccharides for biofilm formation may be constitutively produced before growth can occur (Joergensen and Wichern 2018; Marchus et al. 2018). The slow decline and asymptote in residue mass over time likely represents a slowing of microbial recycling until less biologically available, stabilized materials remain in soil.
The low standard error of the mean for glucose partitioning was surprising given the differences among the 18 surveyed studies in soil biology (e.g., starting MBC; range 37–1654 µg C g−1 dry), texture (e.g., percent clay; range 1.7–50%), and experimental conditions (e.g., glucose amendment; range 0.0009–5000 µg C g−1 dry). For example, ~42% (±2.1 S.E.) of glucose accumulated as MBC and 13% (±1.91 S.E.) as CO2 in the hours immediately after amendment. CUE was relatively fixed at ~75% [=42%/(42% + 13%)] for the studies we surveyed as a result of the low standard error. Since this has been reported frequently by others (Dijkstra et al. 2011, 2015; Frey et al. 2013; Jones et al. 2018; Mehnaz et al. 2019), glucose metabolism appears to be a conserved process in soil. This could limit the range of insights gained from isotopic glucose tracing if this method is insensitive to other drivers of CUE, such as differences in trait-based features of microbial communities (Fierer et al. 2007; Morrissey et al. 2017; Malik et al. 2019).
In contrast to short incubations, glucose partitioning over longer (>24 day) incubations was responsive to other soil variables. For instance, high starting MBC led to a greater percent of tracer accumulating in MBC and less mineralized to CO2 or present as residues. The net effect of these drivers was tracer retention in soil when initial MBC was higher. High microbial biomass is likely accompanied by biofilm production and denser cell consortia that enhance the exchange and recycling of residues, effectively promoting C return to the biomass pool. Additionally, tracer retention in soil over longer periods of time may be under the control of certain master variables (e.g., SOM content) that also positively affect microbial biomass.
Clay content and glucose amendment levels appear to be secondarily influential drivers (after MBC) on tracer partitioning. Others have suggested that clay enhances microbial biosynthesis by serving as a reservoir of exchangeable organic substrates (van Veen et al. 1985; Gregorich et al. 1991). Although we also observed a positive relationship between clay content and starting MBC, clay content negatively affected tracer acquisition by microbes while strongly promoting residue mass. From this it appears that clay content competes with microbial biomass by scavenging exudates and necromass or physically protecting it (Jilling et al. 2018). This clay effect may be transient; for example, the exchangeability of residues with organo-mineral complexes can increase with soil pH (Kleber et al. 2015) or elevated root exudation (Keiluweit et al. 2015). We caution that use of high glucose addition rates, although less consistent and weaker in effect than other predictors, can inflate glucose mineralization (Schneckenberger et al. 2008) and thereby reduce CUE estimates below in-situ levels.
Certain conditions are associated with maximal CUE in soil and thus the potential to build SOM (Sauvadet et al. 2018; Kallenbach et al. 2019). For example, CUE appears to be highest when temperatures do not exceed species-specific thresholds (Steinweg et al. 2008; Schipper et al. 2014; Min et al. 2016) and when resource stoichiometry is in balance with metabolic requirements (Keiblinger et al. 2010; Sinsabaugh et al. 2013). From our present work, the initial MBC amount emerges as another critical factor that enhances CUE [Eq. (2)] by promoting accumulation of tracer C in microbial biomass and dampening its mineralization. In contrast, clay content was not an important predictor of CUE as its effects on tracer accumulation in MBC and CO2 pools offset one another.
Focusing on a two-pool definition of CUE [i.e., MBC and CO2; Eq. (2)] appears shortsighted given the evidence that residues can form rapidly, have substantial mass, and are important SOM precursors. Tracer C is nearly as likely to be in the form of residues as MBC after short incubation (34% vs. 42% of added tracer-C, respectively), and almost twice as likely after longer incubation (19% vs. 11%, respectively). An ecosystem perspective on CUE (sensu Geyer et al. 2016) that accounts for substrate recycling and matrix stabilization in the form of residues is merited. CUEr adds residues into the efficiency denominator [Eq. (6)] to fully account for substrate uptake, reducing the average efficiency observed 48 h after glucose addition from 0.65 to 0.40 for the studies surveyed here (Fig. 4). Adding residue mass into the CUE denominator and numerator (CSE), yields an efficiency of 0.75; 75% of added glucose was retained in soil as MBC or any non-biomass microbial products which may collectively be SOM precursors. By factoring in residues, CSE may directly link microbial efficiency with ecosystem-scale SOM formation more than a conventional CUE metric could.
Conceptual diagram summarizing the trends in glucose-derived C found by meta-analysis of 315 observations in 18 studies (Supp. Table 1) over time after glucose addition (bottom panel). While the total amount of glucose-C remaining in soil declines over time, the relative proportion found as residues (“Res”) increases compared to microbial biomass C (“MBC”). All pools, including respiration (“CO2”), reach an asymptote within ~24 incubation days. Three equations of carbon use efficiency are derived from this data: CUE (MBC/(MBC + CO2)); CUEr (MBC/(MBC + CO2 + Residue C)); CSE ((MBC + Residue C)/(MBC + CO2 + Residue C)). Solid rectangles enclose ‘efficient’ pools positively related to the efficiency metric; dashed rectangles enclose ‘inefficient’ pools negatively related to the metric. The traditional 2-pool formulation of CUE does not account for the substantial residue C pool. We expand on this definition by including residue C as a source of ‘inefficiency’ within CUEr and by including residue C as a source of ‘efficiency’ within CSE (carbon sequestration efficiency)
Our discussion to this point has assumed that tracer C in soil that is not MBC, after applying an extraction coefficient to account for unextractable biomass, is microbial residues. Two further hypotheses warrant mention: this tracer is (1) additional unextractable biomass that needs to be accounted for with lower KEC values, or (2) glucose that has been directly stabilized on organic or mineral surfaces without microbial processing. The first hypothesis would require lowering the average KEC from 0.41 to 0.22 for the studies we surveyed with the shortest incubations of ≤2 days (n = 40 observations) (Fig. 5). A coefficient this low is unlikely given the range of coefficient values that have been found via direct and indirect calibration (0.21–0.59, mean = 0.46; Joergensen 1996). Direct stabilization of glucose in the second hypothesis is also unlikely, given that glucose is an uncharged molecule that undergoes rapid microbial uptake (Fischer et al. 2010; Glanville et al. 2016). After sterilization to prevent microbial uptake, glucose is largely re-extractable from soil (Bremer and van Kessel 1990; Kuzyakov and Jones 2006; Hill et al. 2008). Our laboratory experiment also shows that, when microbial uptake is delayed by high glucose concentrations, glucose extractability from soil is unaffected and non-biomass ‘residues’ do not form (Fig. 1). We find only one reference reporting substantial sorption of glucose in soil (~7% of the initial amendment after 5 h) that may serve as an upper bound on such processes (Fischer et al. 2010).
The relationship between extraction coefficient (KEC) used in estimation of microbial biomass C and the resultant proportion of glucose-derived C that would be attributed to residue C (=Soil C – MBC). Smaller KEC values result in larger MBC estimates, thereby accounting for a greater proportion of glucose-derived C remaining in soil such that residue C declines. Bottom panel is a frequency histogram of KEC values used in the 18 studies we examined by meta-analysis (mean KEC = 0.41). Top panel depicts the mean residue C estimated immediately after tracer addition for the 18 studies we examined (black dot = 34%; see Table 1). Solid line indicates hypothetical changes in this quantity of residue C with use of other KEC values, including full accounting of all glucose-derived C in soil as MBC (i.e., no residue C) with a KEC value of 0.22
Direct quantification of residues would support our conclusions but are difficult to make. Culture-based approaches can permit physical isolation of biomass from exuded products, although natural growth conditions are sacrificed. Isotopic tracing combined with nuclear magnetic resonance imaging or mass spectrometry are adept at characterizing metabolites in soils, but discriminating between intracellular and exuded, non-living compounds remains a challenge (Derrien et al. 2007; Simpson et al. 2018) alongside many of other technical difficulties with working in soil.
We conclude that residues likely have an underestimated role in soil C cycling given the evidence that microbes can generate non-biomass residues as rapidly as living biomass. This is consistent with the wide-ranging challenges microorganisms must overcome before growth in soil can presumably occur: preventing desiccation with capsular sugars, acquiring C and nutrients via exogenous metabolism (Joergensen and Wichern 2018), and combating neighbors (Maynard et al. 2017) among others. Increasing severity of environmental stresses caused by climate change or other anthropogenic activity may induce greater shifts towards such investments at the expense of growth. Understanding when, why, and to what extent microbial residues form will inform fundamental aspects of soil science like microbial biomass estimation, but also provide continuity between spatially and temporally narrow concepts like microbial activity with those like SOM formation at much broader scales of relevance.
References
Allison SD, Wallenstein MD, Bradford MA (2010) Soil-carbon response to warming dependent on microbial physiology. Nat Geosci 3:336–340
Bardgett R, Saggar S (1994) Effects of heavy-metal contamination on the short-term decomposition of labeled [C-14] glucose in a pasture soil. Soil Biol Biochem 26:727–733
Basler A, Dippold M, Helfrich M, Dyckmans J (2015) Microbial carbon recycling—an underestimated process controlling soil carbon dynamics—part 1: a long-term laboratory incubation experiment. Biogeosciences 12:5929–5940
Boddy E, Hill PW, Farrar J, Jones DL (2007) Fast turnover of low molecular weight components of the dissolved organic carbon pool of temperate grassland field soils. Soil Biol Biochem 39:827–835
Bore EK, Apostel C, Halicki S et al (2017) Soil microorganisms can overcome respiration inhibition by coupling intra- and extracellular metabolism: C-13 metabolic tracing reveals the mechanisms. Isme J 11:1423–1433
Brant JB, Sulzman EW, Myrold DD (2006) Microbial community utilization of added carbon substrates in response to long-term carbon input manipulation. Soil Biol Biochem 38:2219–2232
Bremer E, van Kessel C (1990) Extractability of microbial C-14 and N-15 following addition of variable rates of labeled glucose and (NH4)2SO4 to soil. Soil Biol Biochem 22:707–713
Bremer E, Kuikman P (1994) Microbial utilization of C-14 U glucose in soil is affected by the amount and timing of glucose additions. Soil Biol Biochem 26:511–517
Chander K, Brookes P (1991) Microbial biomass dynamics during the decomposition of glucose and maize. Soil Biol Biochem 23:917–925
Chander K, Joergensen RG (2001) Decomposition of C-14 glucose in two soils with different amounts of heavy metal contamination. Soil Biol Biochem 33:1811–1816
Cheshire MV (1979) Nature and origin of carbohydrates in soil. Academic Press, Waltham, p 216
Costa OYA, Raaijmakers JM, Kuramae EE (2018) Microbial extracellular polymeric substances: ecological function and impact on soil aggregation. Front Microbiol 9:14
Creamer CA, Jones DL, Baldock JA, Farrell M (2014) Stoichiometric controls upon low molecular weight carbon decomposition. Soil Biol Biochem 79:50–56
Creamer CA, Jones DL, Baldock JA, Rui YC, Murphy DV, Hoyle FC, Farrell M (2016) Is the fate of glucose-derived carbon more strongly driven by nutrient availability, soil texture, or microbial biomass size?. Soil Biol Biochem 103:201–212
Degens B, Sparling G (1996) Changes in aggregation do not correspond with changes in labile organic C fractions in soil amended with C-14-glucose. Soil Biol Biochem 28:453–462
Derrien D, Marol C, Balesdent J (2007) Microbial biosyntheses of individual neutral sugars among sets of substrates and soils. Geoderma 139:190–198
Dictor MC, Tessier L, Soulas G (1998) Reassessement of the K-ec coefficient of the fumigation-extraction method in a soil profile. Soil Biol Biochem 30:119–127
Dijkstra P, Salpas E, Fairbanks D, Miller EB, Hagerty SB, van Groenigen KJ, Hungate BA, Marks JC, Koch GW, Schwartz E (2015) High carbon use efficiency in soil microbial communities is related to balanced growth, not storage compound synthesis. Soil Biol Biochem 89:35–43
Dijkstra P, Thomas SC, Heinrich PL, Koch GW, Schwartz E, Hungate BA (2011) Effect of temperature on metabolic activity of intact microbial communities: evidence for altered metabolic pathway activity but not for increased maintenance respiration and reduced carbon use efficiency. Soil Biol Biochem 43:2023–2031
Fierer N, Bradford MA, Jackson RB (2007) Toward an ecological classification of soil bacteria. Ecology (Washington D C) 88:1354–1364
Fischer H, Ingwersen J, Kuzyakov Y (2010) Microbial uptake of low-molecular-weight organic substances out-competes sorption in soil. Eur J Soil Sci 61:504–513
Frey SD, Lee J, Melillo JM, Six J (2013) The temperature response of soil microbial efficiency and its feedback to climate. Nat Clim Change 3:395–398
Frey SD, Ollinger S, Nadelhoffer K, Bowden R, Brzostek E, Burton A, Caldwell BA, Crow S, Goodale CL, Grandy AS, Finzi A, Kramer MG, Lajtha K, LeMoine J, Martin M, McDowell WH, Minocha R, Sadowsky JJ, Templer PH, Wickings K (2014) Chronic nitrogen additions suppress decomposition and sequester soil carbon in temperate forests. Biogeochemistry 121:305–316
Geyer KM, Dijkstra P, Sinsabaugh R, Frey SD (2019) Clarifying the interpretation of carbon use efficiency in soil through methods comparison. Soil Biol Biochem 128:79–88
Geyer KM, Kyker-Snowman E, Grandy AS, Frey SD (2016) Microbial carbon use efficiency: accounting for population, community, and ecosystem-scale controls over the fate of metabolized organic matter. Biogeochemistry 127:173–188
Glanville HC, Hill PW, Schnepf A, Oburger E, Jones DL (2016) Combined use of empirical data and mathematical modelling to better estimate the microbial turnover of isotopically labelled carbon substrates in soil. Soil Biol Biochem 94:154–168
Gregorich EG, Voroney RP, Kachanoski RG (1991) Turnover of carbon through the microbial biomass in soils with different textures. Soil Biol Biochem 23:799–805
Gunina A, Kuzyakov Y (2015) Sugars in soil and sweets for microorganisms: review of origin, content, composition and fate. Soil Biol Biochem 90:87–100
Hill PW, Farrar JF, Jones DL (2008) Decoupling of microbial glucose uptake and mineralization in soil. Soil Biol Biochem 40:616–624
Jenkinson DS, Brookes PC, Powlson DS (2004) Measuring soil microbial biomass. Soil Biol Biochem 36:5–7
Jilling A, Keiluweit M, Contosta AR et al (2018) Minerals in the rhizosphere: overlooked mediators of soil nitrogen availability to plants and microbes. Biogeochemistry 139:103–122
Joergensen RG (1996) The fumigation-extraction method to estimate soil microbial biomass: calibration of the k(EC) value. Soil Biol Biochem 28:25–31
Joergensen RG, Wichern F (2018) Alive and kicking: why dormant soil microorganisms matter. Soil Biol Biochem 116:419–430
Jones DL, Hill PW, Smith AR, Farrell M, Ge T, Banning NC, Murphy DV (2018) Role of substrate supply on microbial carbon use efficiency and its role in interpreting soil microbial community-level physiological profiles (CLPP). Soil Biol Biochem 123:1–6
Kallenbach CM, Frey SD, Grandy AS (2016) Direct evidence for microbial-derived soil organic matter formation and its ecophysiological controls. Nat Commun 7:10
Kallenbach CM, Grandy AS, Frey SD, Diefendorf AF (2015) Microbial physiology and necromass regulate agricultural soil carbon accumulation. Soil Biol Biochem 91:279–290
Kallenbach CM, Wallenstein MD, Schipanksi ME, Grandy AS (2019) Managing agroecosystems for soil microbial carbon use efficiency: ecological unknowns, potential outcomes, and a path forward. Front Microbiol 10:8
Keiblinger KM, Hall EK, Wanek W, Szukics U, Hammerle I, Ellersdorfer G, Bock S, Strauss J, Sterflinger K, Richter A, Zechmeister-Boltenstern S (2010) The effect of resource quantity and resource stoichiometry on microbial carbon-use-efficiency. FEMS Microbiol Ecol 73:430–440
Keiluweit M, Bougoure JJ, Nico PS, Pett-Ridge J, Weber PK, Kleber M (2015) Mineral protection of soil carbon counteracted by root exudates. Nat Clim Change 5:588–595
Kleber M, Eusterhues K, Keiluweit M, Mikutta C, Mikutta R, Nico PS (2015) Mineral-organic associations: formation, properties, and relevance in soil environments. In: Sparks DL (ed) Advances in agronomy, vol 130. Elsevier Academic Press Inc, San Diego, pp 1–140
Koch BJ, McHugh TA, Hayer M, Schwartz E, Blazewicz SJ, Dijkstra P, van Gestel N, Marks JC, Mau RL, Morrissey EM, Pett-Ridge J, Hungate BA (2018) Estimating taxon-specific population dynamics in diverse microbial communities. Ecosphere 9:15
Kuzyakov Y, Jones DL (2006) Glucose uptake by maize roots and its transformation in the rhizosphere. Soil Biol Biochem 38:851–860
Kuzyakov Y, Horwath WR, Dorodnikov M, Blagodatskaya E (2019) Review and synthesis of the effects of elevated atmospheric CO2 on soil processes: no changes in pools, but increased fluxes and accelerated cycles. Soil Biol Biochem 128:66–78
Ladd J, Jocteurmonrozier L, Amato M (1992) Carbon turnover and nitrogen transformations in an Alfisol and Vertisol amended with [u-C-14] glucose and [n-15] ammonium sulfate. Soil Biol Biochem 24:359–371
Leitner S, Wanek W, Wild B, Haemmerle I, Kohl L, Keiblinger K, Zechmeister-Boltenstern S, Richter A (2012) Influence of litter chemistry and stoichiometry on glucan depolymerization during decomposition of beech (Fagus sylvatica L.) litter. Soil Biol Biochem 50:174–187
Lynch JM, Panting LM (1980) Cultivation and the soil biomass. Soil Biol Biochem 12:29–33
Malik AA, Puissant J, Goodall T, Allison SD, Griffiths RI (2019) Soil microbial communities with greater investment in resource acquisition have lower growth yield. Soil Biol Biochem 132:36–39
Marchus KA, Blankinship JC, Schimel JP (2018) Environmental controls on extracellular polysaccharide accumulation in a California grassland soil. Soil Biol Biochem 125:86–92
Maynard DS, Crowther TW, Bradford MA (2017) Fungal interactions reduce carbon use efficiency. Ecol Lett 20:1034–1042
Mehnaz KR, Corneo PE, Keitel C, Dijkstra FA (2019) Carbon and phosphorus addition effects on microbial carbon use efficiency, soil organic matter priming, gross nitrogen mineralization and nitrous oxide emission from soil. Soil Biol Biochem 134:175–186
Melillo JM, Frey SD, DeAngelis KM, Werner WJ, Bernard MJ, Bowles FP, Pold G, Knorr MA, Grandy AS (2017) Long-term pattern and magnitude of soil carbon feedback to the climate system in a warming world. Science 358:101–104
Miltner A, Bombach P, Schmidt-Brucken B, Kastner M (2012) SOM genesis: microbial biomass as a significant source. Biogeochemistry 111:41–55
Min K, Lehmeier CA, Ballantyne F, Billings SA (2016) Carbon availability modifies temperature responses of heterotrophic microbial respiration, carbon uptake affinity, and stable carbon isotope discrimination. Front Microbiol 7:10
Moreno-Cornejo J, Zornoza R, Doane TA, Faz A, Horwath WR (2015) Influence of cropping system management and crop residue addition on soil carbon turnover through the microbial biomass. Biol Fertil Soils 51:839–845
Morrissey EM, Mau RL, Schwartz E, McHugh TA, Dijkstra P, Koch BJ, Marks JC, Hungate BA (2017) Bacterial carbon use plasticity, phylogenetic diversity and the priming of soil organic matter. ISME J 11:1890–1899
Mueller T, Magid J, Jensen LS, Svendsen H, Nielsen NE (1998) Soil C and N turnover after incorporation of chopped maize, barley straw and blue grass in the field: evaluation of the DAISY soil-organic-matter submodel. Ecol Model 111:1–15
Reischke S, Rousk J, Baath E (2014) The effects of glucose loading rates on bacterial and fungal growth in soil. Soil Biol Biochem 70:88–95
Rousk J, Baath E (2011) Growth of saprotrophic fungi and bacteria in soil. FEMS Microbiol Ecol 78:17–30
Saggar S, Hedley CB, Giddens KM, Salt GJ (2000) Influence of soil phosphorus status and nitrogen addition on carbon mineralization from C-14-labelled glucose in pasture soils. Biol Fertil Soils 32:209–216
Saggar S, Tate K, Feltham C et al (1994) Carbon turnover in a range of allophanic soils amended with C-14-labeled glucose. Soil Biol Biochem 26:1263–1271
Satterthwaite FE (1946) An approximate distribution of estimates of variance components. Biom Bull 2:110–114
Sauvadet M, Lashermes G, Alavoine G, Recous S, Chauvat M, Maron PA, Bertrand I (2018) High carbon use efficiency and low priming effect promote soil C stabilization under reduced tillage. Soil Biol Biochem 123:64–73
Schipper LA, Hobbs JK, Rutledge S, Arcus VL (2014) Thermodynamic theory explains the temperature optima of soil microbial processes and high Q(10) values at low temperatures. Glob Change Biol 20:3578–3586
Schmidt MWI, Torn MS, Abiven S, Dittmar T, Guggenberger G, Janssens IA, Kleber M, Koegel-Knabner I, Lehmann J, Manning DAC, Nannipieri P, Rasse DP, Weiner S, Trumbore SE (2011) Persistence of soil organic matter as an ecosystem property. Nature (London) 478:49–56
Schneckenberger K, Demin D, Stahr K, Kuzyakov Y (2008) Microbial utilization and mineralization of C-14 glucose added in six orders of concentration to soil. Soil Biol Biochem 40:1981–1988
Simpson AJ, Simpson MJ, Soong R (2018) Environmental nuclear magnetic resonance spectroscopy: an overview and a primer. Anal Chem 90:628–639
Sinsabaugh RL, Manzoni S, Moorhead DL, Richter A (2013) Carbon use efficiency of microbial communities: stoichiometry, methodology and modelling. Ecol Lett 16:930–939
Smith JL, Bell JM, Bolton H, Bailey VL (2007) The initial rate of C substrate utilization and longer-term soil C storage. Biol Fertil Soils 44:315–320
Sparling GP, West AW (1988) A direct extraction method to estimate soil microbial C—calibration in situ using microbial respiration and C-14-labeled cells. Soil Biol Biochem 20:337–343
Steinweg JM, Plante AF, Conant RT, Paul EA, Tanaka DL (2008) Patterns of substrate utilization during long-term incubations at different temperatures. Soil Biol Biochem 40:2722–2728
van Veen JA, Ladd JN, Amato M (1985) Turnover of carbon and nitrogen through the microbial biomass in a sandy loam and a clay soil with C-14 U-glucose and N-15 (NH4)2SO4 under different moisture regimes. Soil Biol Biochem 17:747–756
Vance ED, Brookes PC, Jenkinson DS (1987) An extraction method for measuring soil microbial biomass-C. Soil Biol Biochem 19:703–707
Viechtbauer W (2010) Conducting meta-analyses in R with the metafor package. J Stat Softw 36:1–48
Wadso I (2009) Characterization of microbial activity in soil by use of isothermal microcalorimetry. J Therm Anal Calorim 95:843–850
Witter E, Dahlin S (1995) Microbial utilization of [U-C-14]-labelled straw and [U-C-13]-labelled glucose in soils of contrasting pH and metal status. Soil Biol Biochem 27:1507–1516
Wu J, Brookes P, Jenkinson D (1993) Formation and destruction of microbial biomass during the decomposition. Soil Biol Biochem 25:1435–1441
Acknowledgements
This work was supported by grants to SDF from the U.S. Department of Energy Genomic Science Program (DE-S0016590) and the U.S. Department of Agriculture National Institute of Food and Agriculture through the New Hampshire Agricultural Experiment Station (NHAES; Hatch 1003421). This is NHAES Scientific Contribution Number 2848.
Author information
Authors and Affiliations
Contributions
KG designed the study, performed the research, and wrote the manuscript. All authors contributed substantially to revisions.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Data accessibility statement
Data has been archived in Dryad with DOI: https://doi.org/10.5061/dryad.jm63xsj8p.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Responsible Editor: Susan E. Crow.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Geyer, K., Schnecker, J., Grandy, A.S. et al. Assessing microbial residues in soil as a potential carbon sink and moderator of carbon use efficiency. Biogeochemistry 151, 237–249 (2020). https://doi.org/10.1007/s10533-020-00720-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10533-020-00720-4