Abstract
Melanoma is a highly immunogenic malignancy with an elevated mutational burden, diffuse lymphocytic infiltration, and one of the highest response rates to immune checkpoint inhibitors (ICIs). However, over half of all late-stage patients treated with ICIs will either not respond or develop progressive disease. Spatial imaging technologies are being increasingly used to study the melanoma tumor microenvironment (TME). The goal of such studies is to understand the complex interplay between the stroma, melanoma cells, and immune cell-types as well as their association with treatment response. Investigators seeking a better understanding of the role of cell location within the TME and the importance of spatial expression of biomarkers are increasingly turning to highly multiplexed imaging approaches to more accurately measure immune infiltration as well as to quantify receptor-ligand interactions (such as PD-1 and PD-L1) and cell-cell contacts. CyTOF-IMC (Cytometry by Time of Flight - Imaging Mass Cytometry) has enabled high-dimensional profiling of melanomas, allowing researchers to identify complex cellular subpopulations and immune cell interactions with unprecedented resolution. Other spatial imaging technologies, such as multiplexed immunofluorescence and spatial transcriptomics, have revealed distinct patterns of immune cell infiltration, highlighting the importance of spatial relationships, and their impact in modulating immunotherapy responses. Overall, spatial imaging technologies are just beginning to transform our understanding of melanoma biology, providing new avenues for biomarker discovery and therapeutic development. These technologies hold great promise for advancing personalized medicine to improve patient outcomes in melanoma and other solid malignancies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Melanoma is an immunogenic malignancy with an elevated tumor mutational burden (TMB) and a highly active immune tumor microenvironment (TME) [1]. While historically associated with poor clinical outcomes, late-stage metastatic melanoma patients have seen a significant improvement in prognosis with the advent and clinical implementation of immune checkpoint inhibitors (ICIs) [2,3,4]. The favorable response to these therapies has been linked to a number of factors, such as abundance of neoantigens and the degree of infiltrating lymphocytes within the melanoma TME [5,6,7]. In recent years, spatial imaging technologies have been increasingly used to study the immune infiltration of melanomas, with the goal of understanding the complex interplay between stromal and tumor cells, as well as distinct immune and tumor-infiltrating lymphocyte subsets (TILs), and their effect on ICI response.
Improved approaches to characterize the melanoma TME carries a degree of urgency with clear clinical significance as more than 50% of late-stage metastatic melanoma patients treated with ICIs will either not respond or develop progressive disease [8]. With the recent approval of anti-LAG-3 ICIs for the treatment of metastatic melanoma, a better understanding of the role of the melanoma TME and mechanisms mediating patient response to anti-PD-1 monotherapy compared to combination therapy with anti-CTLA-4 or LAG-3, is clearly needed [9]. Furthermore, results from recent melanoma neoadjuvant clinical trials, SWOG S1801 (NCT03698019) and NADINA (NCT04949113), are poised to redefine management of high-risk resectable stage III melanomas [10]. Molecular profiling studies of pre- and post-treated melanomas in the neoadjuvant setting provides an efficient platform to understand mechanisms of therapy response with measured pathological response available in a relatively short period of time. Such melanoma study designs provide a critical opportunity to identify biomarkers to personalize adjuvant therapy and improve patient outcomes [11]. In this review, we will provide a brief survey of current multiplexed imaging technologies and highlight recent studies that have employed such approaches to characterize the spatial immune landscape of melanomas to better understand ICI responses.
Multiplexed imaging technologies
Limitations in conventional multiplexed tissue staining and antibody-based imaging, such as hematoxylin and eosin (H&E), immunohistochemistry (IHC) and immunofluorescence (IF) approaches has previously limited the scope and scale of melanoma TME research. However, studies have shown the improved predictive power of biomarkers of ICI response when incorporating spatial TME information, such as cell-type specific expression in specific regions of melanomas [5, 12]. Such studies have reinforced the need to employ more advanced highly multiplexed approaches for translational research.
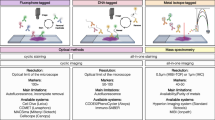
To address these limitations, various multiplexed spatial imaging methods have been developed to study the melanoma TME in search of new clinically predictive biomarkers over the past decade (Table 1). Multiplexed IF and cyclical multiplexed immunofluorescence techniques such as OPAL-7 [13], tyramide signal amplification (TSA) [14], t-CyCIF (Tissue-based Cyclic Immunofluorescence) [15], or MILAN (Multiple Iterative Labeling by Antibody Neodeposition) [16] can image from 6 to 60 markers using repeated cycles of antibody deposition and imaging. Among these, the more highly-multiplexed technologies have enabled researchers to extract more cell-type specific spatial information, such as t-CyCIF, which makes use of repeated cycles of low-plex fluorescence staining and imaging for up to 60 markers [15]. T-CyCIF has recently been applied to the melanoma TME to evaluate sequential melanoma biopsies obtained from the same patient, revealing co-evolution of differences in immune composition among different clonal lineages, while another study on metastatic melanoma lymph nodes used a 45-plexed t-CyCIF antibody panel to classify prognostically relevant immune-cell neighborhoods [17, 18]. T-CyCIF has also been combined with other multiplexed technologies including PickSeq and NanoString to spatially characterize melanoma cellular neighborhoods, their transition from precursor states to melanoma in situ and invasive tumor, and the immune-suppressive environment along the tumor-stromal boundary [19].
Recent advancements in lower-cost spatial immune profiling technology include bright-field multiplexed imaging, which enables imaging of triple-plexed bright-field colors for clinical and histopathological diagnostic assays using standard pathology-lab equipment [22]. Another advancement is nine-color multiplexed immunofluorescence imaging that has been applied to characterize cell phenotype diversity and immunosuppression patterns within the TME of malignant pleural mesothelioma [33].
In contrast, mass-spectroscopy-based imaging and analysis methods, such as MALDI (Matrix Assisted Laser Desorption/Ionization), DESI (Desorption Electrospray Ionization), SIMS (Secondary Ion Mass Spectrometry), and MIBI-TOF (Multiplexed Ion Beam Imaging by Time-of-Flight MS), can detect up to thousands of analytes including metabolites on a sample without using repetitive imaging cycles [29, 34, 35]. For example, MIBI-TOF, which can image up to 36-plex markers (or in theory, up to 100 markers) and uses a tunable ion-beam that can be adjusted to various sample depths and image resolutions, has been applied to image cellular phenotypes and tissue structure, revealing regional variability in tumor cell phenotypes in several different breast cancer subtypes including triple-negative breast cancer [35, 36]. CyTOF-IMC, another recent addition to the spatial imaging toolkit, can detect up to 40 proteins on a single sample of formalin-fixed paraffin-embedded (FFPE) tissue and has recently been applied to characterize the melanoma TME [23,24,25,26].
Exponential development in the spatial transcriptomics imaging field has enabled researchers to spatially image transcripts from up to 20,000 different protein-coding genes at mid to high-resolution, using technologies including NanoString [37], Visium [38], and MER-FISH [39]. Generally, spatial transcriptomics technologies fall into two categories. First, those that spatially segment tissue into a limited number of relevant cell types and interrogate large numbers of transcripts across those selected groups of cells, albeit without achieving single-cell resolution, such as NanoString [37, 40]. Second, those that provide single-cell, or close-to single-cell resolution for a more limited number of transcripts, such as MER-FISH [40]. In the intermediary zone are technologies that include Visium, which allow for lower-resolution whole transcriptome sequencing of ‘patches’ of 5–50 cells [38, 41]. These technologies have provided detailed insights into the TME in the context of ICI treatment for melanoma liver metastases. A comprehensive analysis of cutaneous and uveal melanoma liver metastases identified differences in the TME between these melanoma subtypes, such as PD-L1 expression and the ratio of exhausted CD8 T cells to several other T cell subsets [30]. High-plex spatial RNA profiling has been instrumental in revealing cell type-specific biomarker expression in the TME during early melanoma evolution, including prominent expression of S100A8 and S100A9 in melanoma-adjacent keratinocytes [42]. Furthermore, these technologies have been used to examine the immune landscape of tumor-associated lymph nodes in melanoma, leading to the identification of potential biomarkers of CD11c activation that could be clinically relevant for predicting survival outcomes [37].
Spatial transcriptomics platforms, including NanoString and Visium, have greatly advanced our understanding of gene transcription by facilitating the detailed analysis of mRNA levels at a high spatial resolution [37,38,39,40,41]. Of note, transcriptomic-based technologies do possess inherent limitations as some isoforms may not be accurately differentiated by these platforms and mRNA levels are not always congruent with protein expression. Thus, protein-based methodologies cannot be wholly substituted by spatial transcriptomics alone. Integration of spatial proteomic and transcriptomic profiling allowed by currently available platforms has already shown promise in offering a more holistic view of the cellular landscape [23, 24]. However, the field still faces significant challenges in merging these data streams, largely due to technological limitations in comprehensively measuring both the transcriptome and proteome simultaneously.
CyTOF studies in cutaneous melanoma
Flow cytometry has long been a cornerstone in the field of cellular analytics, providing a means to measure, quantify and analyze characteristics of single cells in suspension in a multiplexed workflow. The Helios CyTOF platform introduced the use of rare earth metal isotopes for antibody labeling and integrated mass spectroscopy into flow cytometry as an analysis technique, thus allowing for separation of cells based on their mass-to-charge ratio and eliminating issues with spectral overlap from traditional flow cytometry platforms [43, 44]. This significantly expanded the capabilities of flow cytometry technology and allowed for the simultaneous analysis of up to 60 parameters using metal-tagged antibodies, albeit without preserving spatial orientation of cells within tissues [44,45,46]. The technology used to develop the Helios CyTOF platform paved the way for the development of the Hyperion CyTOF-IMC platform that can multiplex image up to 40 markers using rare earth metal isotope-conjugated antibodies [47].
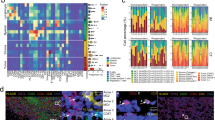
CyTOF-IMC can detect antibodies at sub-cellular resolution simultaneously on the same tissue slide using high-frequency laser ablation of antibody-tagged metal isotopes in a low-dispersion laser ablation chamber, for either FFPE or fresh tissue [47]. In brief, tissues are labeled with metal-tagged antibodies using standard immunohistochemistry methods, air dried, laser-ablated pixel by pixel, and transported by a mixed argon and helium stream to the CyTOF mass cytometer for measurement (Fig. 1) [47]. Cellular images generated by CyTOF-IMC undergo a two-step analytical process. Images are first segmented into individual cells using a variety of software. Next, cells are phenotyped using methods based either on unsupervised clustering (e.g. Phenograph) [48], FlowSOM [49], or probabilistic classification models (e.g. Astir) [50] (Table 2 provides a non-exhaustive overview of some of the more commonly-used tools available for these tasks). Advanced spatial analysis, including the calculation of median intercellular distances, and contact enrichment, can be performed using tools such as imcRtools [51] (see [52] for a more comprehensive review of analytical pipelines).
Illustration of the data acquisition workflow used for CyTOF-IMC: (1) One ~ 5 μm-thick section of a tissue microarray (TMA), usually composed of 1 mm2 melanoma cores (2) is stained with a cocktail of antibodies labeled with metal isotopes (colored asterisks). (3) Samples are ablated with a high energy laser in a rastered pattern, and the resulting ionized isotope plumes are analyzed by a mass cytometer, which returns the number and type of metal isotopes per pixel (one pixel = 1micron x 1micron). (4) Each antibody results in a single image per sample, and together are layered to construct a multi-image stack. Since 2014, CyTOF-IMC has been utilized to spatially analyze a wide array of tissues and tumor-types, including breast, lung, and melanoma, as well as to identify biomarkers of response to treatment in breast cancer (including HR and HER2) or immunotherapy (anti PD-1 or anti-CTLA4) in melanoma [24, 26, 47, 60, 61]
Two of the earliest publications to use CyTOF to characterize melanoma patient samples utilized the Helios platform to characterize peripheral blood mononuclear cells (PBMCs) or immune profiles of tumor biopsies from patients treated with anti-PD-1 monotherapy, or combined anti-PD-1 and anti-CTLA-4 [62, 63]. One early study conducted on stage IV melanoma patients receiving anti-PD-1 immunotherapy found that the frequency of CD14 + CD16 − HLA-DRhi monocytes in PBMCs prior to therapy initiation was a strong indicator of progression-free and overall survival, suggesting their potential use in informing treatment decisions [62]. Another study identified activated T-cell signatures and T-cell populations in responders to both treatment modalities (anti-PD-1 and anti-CTLA-4) using a panel of multiplexed antibodies to characterize immune cell populations [63]. Further analysis revealed an EOMES + CD69 + CD45RO + effector memory T-cell phenotype that was significantly more abundant in responders to combined immunotherapy than non-responders. The gene expression profile of this population was associated with longer progression-free survival in patients treated with single-agent therapy and showed greater tumor shrinkage in both treatments, revealing insights into response and resistance mechanisms to ICIs [63]. These early CyTOF flow cytometry studies provided valuable information regarding abundance of immune cell populations present in the melanoma TME. However, the advent of the Hyperion IMC has facilitated the multiplexed characterization of spatial relationships of cells and biomarkers within the melanoma TME using the CyTOF platform.
Four recent studies have utilized CyTOF-IMC to directly characterize the melanoma TME (Table 3). Published in 2021, CyTOF-IMC was employed to analyze the TME of patients with metastatic melanoma who received ICI, and to identify indicative factors of treatment response [25]. Rather than segmenting cells, the authors used a newly designed version of the AQUA software to measure marker intensity in molecularly defined compartments. Multivariable analyses revealed significant associations of 12 markers with progression-free survival, and 7 markers with overall survival, which included b2-microglobulin [25].
In our study, published in 2022, we utilized CyTOF-IMC to profile more than 230,000 individual cells from 5 benign nevi and 67 melanomas and identified melanoma, lymphocyte, macrophage/monocyte, and stromal cell populations, allowing for in-depth spatial quantification of the melanoma microenvironment [26]. While prior studies have shown that the abundance of TILs in the TME is associated with better prognosis and response to ICIs [64,65,66], we demonstrated that within the pre-treatment melanoma TME, it was the abundance of proliferating antigen-experienced cytotoxic T-cells (CD8 + CD45RO + Ki67+) and their proximity to melanoma cells that best informed on ICI responses in our analyzed cohort [26]. This indicated that a shorter distance between melanoma and nearby antigen-experienced cytotoxic T-cells within the TME was linked to a favourable response, highlighting the potential of CyTOF-IMC to quantify spatial cell-cell interactions for ICI biomarker studies.
Two other studies were published in 2022 that utilized CyTOF-IMC but combined this approach with other modalities to investigate the immune environment of melanoma. Specifically, researchers investigated the role of regulatory and pro-inflammatory cytokine-expressing B cells in patients with melanoma, utilizing flow cytometry, CyTOF-IMC, single-cell RNAseq, immunofluorescence staining and transcriptomic analysis [23]. The group found that patients had enhanced circulating regulatory B cell populations and reduced pro-inflammatory B cell populations compared to healthy volunteers [23]. They also found that cytokine-expressing B cells in the melanoma TME assembled in clusters and interacted with T-cells and T-regulatory cells via various signaling pathways [23]. Patient-derived B cells were found to promote T-regulatory cell differentiation and T-helper cell proliferation in a TGF-β-dependent manner, an effect further enhanced with anti-PD-1 checkpoint blockade [23]. These findings highlight the bidirectional crosstalk between B and T cell subsets with immunosuppressive attributes in the context of melanoma. In another CyTOF IMC study, researchers reported the use of multiplexed mass cytometry–based imaging of protein markers and RNA transcripts to investigate the chemokine landscape and immune infiltration in metastatic melanoma samples [24]. The study showed that tumors lacking immune infiltration had low levels of antigen presentation and markers of inflammation, and were devoid of most of the profiled chemokines [24]. In contrast, infiltrated tumors expressed high cytokine levels, with CXCL9 and CXCL10 being localized in patches associated with dysfunctional T-cells expressing the B lymphocyte chemoattractant CXCL13 [24]. This study also found that T-cells play a role in B cell recruitment and potentially in B cell follicle formation, and that the formation of tertiary lymphoid structures may be accompanied by naïve and naïve-like T-cell recruitment, which can contribute to antitumor activity. These two CyTOF-IMC articles demonstrate the potential of combining multiple spatial ‘omic’ technologies to make important TME discoveries.
Future studies combining multiple spatial ‘omic’ technologies (e.g. CyTOF-IMC, NanoString, Visium, ATAC-Seq, microbiome analysis, and plasma metabolite analysis) will undoubtedly lead to a more deeper understanding of TME biology. One challenge with integrating multiple profiling approaches is that often these technologies require different biological material inputs (i.e. plasma, tissue sections, or tissue lysate). Already, studies have expanded IMC technologies to detect mRNA and proteins with single-cell and spatial resolution in melanoma and breast tumours, capturing three mRNAs along with a panel of proteins to reveal correlations between mRNA and proteins at both the single-cell and cell population levels [24, 67]. Furthermore, we recently interrogated microbiome composition, plasma metabolite makeup, and employed CyTOF-IMC to analyze the immune landscape of melanoma tumors prior to fecal microbiota transplantation (FMT) in order to more comprehensively characterize the immune infiltrates in responsive and non-responsive patients to combination FMT and ICI therapy [68].
Limitations
Spatial imaging proteomic and transcriptomics technologies, while powerful, are newer techniques that often grapple with limitations and compromises. These include balancing single-cell resolution with high multiplexing capabilities, limitations in availability of required reagents such as high-quality antibodies and probes, as well as challenges associated with high costs and slow data acquisition times coupled with the complexity of data analysis. For example, analyzing and interpreting the large and complex datasets produced by CyTOF-IMC requires sophisticated tools and computational expertise. Recent advances in machine learning have allowed for improved cell segmentation and classification in the analysis and interpretation of large and complex datasets produced by CyTOF-IMC. For example, a recent study utilized CyTOF-IMC and deep learning to analyze the TME of lung adenocarcinoma samples from 416 patients, profiling 1.6 million cells using a combination of classical and modern machine learning based computer vision algorithms in order to reveal distinct immune lineages and activation states with clinical correlates, and demonstrating the potential of artificial intelligence in predicting patient progression using a single 1-mm2 tumor core [57, 61]. Finally, as the technology for spatial imaging and transcriptomics continue to advance, analytical tools, imaging analysis methods and data processing pipelines will need to keep pace. These rapid advances provide both great opportunities and present unique challenges to researchers, as consolidation of best-practices for application and data analysis are continually evolving to meet the needs of the ever-advancing armamentarium of spatial imaging technologies.
Conclusion
The future of single-cell spatial profiling will be driven by the development of novel technologies that can overcome current limitations and provide more nuanced insights into single-cell dynamics. These advancements will enable multiplexed, cell-type specific, and tumor location-specific analysis of biomarkers, which could significantly enhance their predictive power. Furthermore, the integration of 3D spatial imaging using technologies such as CyTOF-IMC may provide a more comprehensive view of the cellular landscape. Ultimately, the integration of 3D biomarker imaging across multi-omic platforms would significantly improve our understanding of the TME [69].
Within the melanoma TME research field, these single cell imaging advancements are poised to have an immediate and profound impact. With the recent approval of anti-LAG-3 for the treatment of late-stage metastatic melanoma, identification of biomarkers that can inform on prescription of monotherapy anti-PD-1, versus combinations that include anti-CTLA-4 and LAG-3, are urgently needed. Moreover, single cell technologies amenable to predicting response in the neoadjuvant setting could significantly advance personalized medicine by tailoring adjuvant treatments to individual patient profiles [11].
As our review has underscored, recent studies utilizing single cell imaging approaches are just beginning to illuminate the cellular composition and functional dynamics of the melanoma TME. This area of investigation has the potential to uncover novel biomarkers for diagnosis and prognosis, particularly in the context of ICI response, and to identify new targets for therapeutic intervention.
Search methodology
In this review, a search of peer-reviewed scientific articles and literature reviews was completed using PubMed with the following search terms: (“Melanoma”) AND (“Tumor micro-environment” OR “Tumour micro-environment” OR “Tumor microenvironment” OR “Tumour microenvironment” OR “Cellular Microenvironment” OR “Cellular Micro-environment” OR “Tumour immune microenvironment” OR “Tumor immune microenvironment” OR “Tumour immune micro-environment” OR “Tumor immune micro-environment”) AND (“spatial landscape” OR “spatial”), within the date range of 2014–2023 (the earliest CyTOF-IMC paper being released in 2014), retrieving 82 results. After full-text review, we included 3 papers on melanoma spatial imaging using CyTOF-IMC, and captured an additional 6 papers on other melanoma spatial imaging technologies. We also applied a snowballing search methodology using the references cited in the articles identified in the literature search, locating an additional 1 pertinent paper (see Table 3). For the snowballing search, articles were limited to those including CyTOF-IMC or other relevant spatial imaging technologies, melanoma or another relevant cancer, or other pertinent articles. Each identified item was assessed for relevance by a member of the study team and final calls were made by the senior author. This review is intended to summarize high-impact, relevant and recent literature on spatial imaging technologies, specifically CyTOF-IMC, for the tumor microenvironment and immune landscape in melanoma, and is not meant to be exhaustive.
References
Alkallas R, Lajoie M, Moldoveanu D, Hoang KV, Lefrançois P, Lingrand M, Ahanfeshar-Adams M, Watters K, Spatz A, Zippin JH, Najafabadi HS, Watson IR (2020) Multi-omic analysis reveals significantly mutated genes and DDX3X as a sex-specific Tumor suppressor in cutaneous Melanoma. Nat Cancer 6(1):635–652. https://doi.org/10.1038/s43018-020-0077-8
Chamoto K, Yaguchi T, Tajima M, Honjo T (2023) Insights from a 30-year journey: function, regulation and therapeutic modulation of PD1. Nat Rev Immunol 23:682–695. https://doi.org/10.1038/s41577-023-00867-9
Wei SC, Duffy CR, Allison JP (2018) Fundamental mechanisms of immune checkpoint blockade therapy. Cancer Discov 9(8):1069–1086. https://doi.org/10.1158/2159-8290.CD-18-0367
Luke JJ, Flaherty KT, Ribas A, Long GV (2017) Targeted agents and immunotherapies: optimizing outcomes in Melanoma. Nat Rev Clin Oncol 8(14):463–482. https://doi.org/10.1038/nrclinonc.2017.43
Tumeh PC, Harview CL, Yearley JH, Shintaku IP, Taylor EJM, Robert L, Chmielowski B, Spasic M, Henry G, Ciobanu V, West AN, Carmona M, Kivork C, Seja E, Cherry G, Gutierrez AJ, Grogan TR, Mateus C, Tomasic G, Glaspy JA, Emerson RO, Robins H, Pierce RH, Elashoff DA, Robert C, Ribas A (2014) PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 7528(515):568–571. https://doi.org/10.1038/nature13954
Snyder A, Makarov V, Merghoub T, Yuan J, Zaretsky JM, Desrichard A, Walsh LA, Postow MA, Wong P, Ho TS, Hollmann TJ, Bruggeman C, Kannan K, Li Y, Elipenahli C, Liu C, Harbison CT, Wang L, Ribas A, Wolchok JD, Chan TA (2014) Genetic basis for clinical response to CTLA-4 blockade in Melanoma. N Engl J Med 23(371):2189–2199. https://doi.org/10.1056/NEJMoa1406498
Van Allen EM, Miao D, Schilling B, Shukla SA, Blank C, Zimmer L, Sucker A, Hillen U, Geukes Foppen MH, Goldinger SM, Utikal J, Hassel JC, Weide B, Kaehler KC, Loquai C, Mohr P, Gutzmer R, Dummer R, Gabriel S, Wu CJ, Schadendorf D, Garraway LA (2015) Genomic correlates of response to CTLA-4 blockade in metastatic Melanoma. Science (350)6257:207–211. https://doi.org/10.1126/science.aad0095
Satya D, Douglas BJ (2019) Immune-related adverse events and anti-tumor efficacy of immune checkpoint inhibitors. JITC 1(7):306. http://jitc.bmj.com/content/7/1/306.abstract
Tawbi HA, Schadendorf D, Lipson EJ, Ascierto PA, Matamala L, Castillo Gutiérrez E, Rutkowski P, Gogas HJ, Lao CD, De Menezes JJ, Dalle S, Arance A, Grob JJ, Srivastava S, Abaskharoun M, Hamilton M, Keidel S, Simonsen KL, Sobiesk AM, Li B, Hodi FS, Long GV (2022) Relatlimab and Nivolumab versus Nivolumab in untreated advanced Melanoma. N Engl J Med 1(386):24–34. https://doi.org/10.1056/NEJMoa2109970
Long GV, Menzies AM, Scolyer RA (2023) Neoadjuvant Checkpoint Immunotherapy and Melanoma: The Time Is Now. J Clin Oncol 17(41):3236–3248. https://doi.org/10.1200/JCO.22.02575
Lucas MW, Versluis JM, Rozeman EA, Blank CU (2023) Personalizing neoadjuvant immune-checkpoint inhibition in patients with Melanoma. Nat Rev Clin Oncol 6(20):408–422. https://doi.org/10.1038/s41571-023-00760-3
Taube JM, Klein A, Brahmer JR, Xu H, Pan X, Kim JH, Chen L, Pardoll DM, Topalian SL, Anders RA (2014) Association of PD-1, PD-1 ligands, and other features of the tumor immune microenvironment with response to anti–PD-1 therapy. Clin Cancer Res 19(20):5064–5074. https://doi.org/10.1158/1078-0432.CCR-13-3271
Conway JW, Rawson RV, Lo S, Ahmed T, Vergara IA, Gide TN, Attrill GH, Carlino MS, Saw RPM, Thompson JF (2022) Unveiling the Tumor immune microenvironment of organ-specific Melanoma metastatic sites. JITC 9(10):e004884. https://jitc.bmj.com/content/10/9/e004884
Berry S, Giraldo NA, Green BF, Cottrell TR, Stein JE, Engle EL, Xu H, Ogurtsova A, Roberts C, Wang D, Nguyen P, Zhu QF, Soto-Diaz S, Loyola J, Sander IB, Wong PF, Jessel S, Doyle J, Signer D, Wilton R, Roskes JS, Eminizer M, Park S, Sunshine JC, Jaffee EM, Baras A, De Marzo AM, Topalian SL, Kluger H, Cope L, Lipson EJ, Danilova L, Anders RA, Rimm DL, Pardoll DM, Szalay AS, Taube JM (2021) Analysis of multispectral imaging with the AstroPath platform informs efficacy of PD-1 blockade. Science 6547(372):eaba2609. https://doi.org/10.1126/science.aba2609
Lin JR, Izar B, Wang S, Yapp C, Mei S, Shah P, Santagata S, Sorger PK (2018) Highly multiplexed immunofluorescence imaging of human tissues and tumors using t-CyCIF and conventional optical microscopes. eLife (7):e31657. https://doi.org/10.7554/eLife.31657
Bosisio FM, Antoranz A, van Herck Y, Bolognesi MM, Marcelis L, Chinello C, Wouters J, Magni F, Alexopoulos L, Stas M, Boecxstaens V, Bechter O, Cattoretti G, van den Oord J (2020) Functional heterogeneity of lymphocytic patterns in primary Melanoma dissected through single-cell multiplexing. eLife (9):e53008. https://doi.org/10.7554/eLife.53008
Liu D, Lin JR, Robitschek EJ, Kasumova GG, Heyde A, Shi A, Kraya A, Zhang G, Moll T, Frederick DT, Chen YA, Wang S, Schapiro D, Ho LL, Bi K, Sahu A, Mei S, Miao B, Sharova T, Alvarez-Breckenridge C, Stocking JH, Kim T, Fadden R, Lawrence D, Hoang MP, Cahill DP, Malehmir M, Nowak M, Brastianos PK, Lian CG, Ruppin E, Izar B, Herlyn M, Van Allen EM, Nathanson K, Flaherty KT, Sullivan RJ, Kellis M, Sorger PK, Boland GM (2021) Evolution of delayed resistance to immunotherapy in a Melanoma responder. Nat Med 6(27):985–992. https://doi.org/10.1038/s41591-021-01331-8
Maus RLG, Leontovich AA, Moore RM, Fogarty Z, Guo R, Davidson TM, Tekin B, Atherton C, Schimke JM, Dicke BA, Chen BJ, Markovic SN (2022) Quantitative spatial evaluation of tumor-immune interactions in the immunotherapy setting of metastatic Melanoma lymph nodes. Front Immunol (13):1024039. https://www.frontiersin.org/articles/10.3389/fimmu.2022.1024039
Nirmal AJ, Maliga Z, Vallius T, Quattrochi B, Chen AA, Jacobson CA, Pelletier RJ, Yapp C, Arias-Camison R, Chen YA, Lian CG, Murphy GF, Santagata S, Sorger PK (2022) The spatial Landscape of Progression and Immunoediting in primary Melanoma at single-cell resolution. Cancer Discov 6(12):1518–1541. https://doi.org/10.1158/2159-8290.CD-21-1357
Karras P, Bordeu I, Pozniak J, Nowosad A, Pazzi C, Van Raemdonck N, Landeloos E, Van Herck Y, Pedri D, Bervoets G, Makhzami S, Khoo JH, Pavie B, Lamote J, Marin-Bejar O, Dewaele M, Liang H, Zhang X, Hua Y, Wouters J, Browaeys R, Bergers G, Saeys Y, Bosisio F, van den Oord J, Lambrechts D, Rustgi AK, Bechter O, Blanpain C, Simons BD, Rambow F, Marine JC (2022) A cellular hierarchy in Melanoma uncouples growth and Metastasis. Nature 7930(610):190–198. https://doi.org/10.1038/s41586-022-05242-7
Nguyen T, Kocovski N, Macdonald S, Yeang HXA, Wang M, Neeson PJ (2021) Multiplex Immunohistochemistry Analysis of Melanoma Tumor-Infiltrating Lymphocytes. In: Hargadon Kristian M (ed) Melanoma: methods and protocols, vol 2265. Springer US, New York, NY, p 557–572
Ugolini F, Pasqualini E, Simi S, Baroni G, Massi D (2022) Bright-field multiplex immunohistochemistry assay for tumor microenvironment evaluation in Melanoma tissues. Cancers 15(14):3682. https://doi.org/10.3390/cancers14153682
Harris RJ, Willsmore Z, Laddach R, Crescioli S, Chauhan J, Cheung A, Black A, Geh JLC, MacKenzie Ross AD, Healy C, Tsoka S, Spicer J, Lacy KE, Karagiannis SN (2022) Enriched circulating and tumor-resident TGF-β + regulatory B cells in patients with melanoma promote FOXP3 + Tregs. OncoImmunology 1(11):2104426. https://doi.org/10.1080/2162402X.2022.2104426
Hoch T, Schulz D, Eling N, Gómez JM, Levesque MP, Bodenmiller B (2022) Multiplexed imaging mass cytometry of the chemokine milieus in Melanoma characterizes features of the response to immunotherapy. Sci Immunol 70(7):eabk1692. https://doi.org/10.1126/sciimmunol.abk1692
Martinez-Morilla S, Villarroel-Espindola F, Wong PF, Toki MI, Aung TN, Pelekanou V, Bourke-Martin B, Schalper KA, Kluger HM, Rimm DL (2021) Biomarker Discovery in patients with immunotherapy-treated Melanoma with Imaging Mass Cytometry. Clin Cancer Res 7(27):1987–1996. https://doi.org/10.1158/1078-0432.CCR-20-3340
Moldoveanu D, Ramsay L, Lajoie M, Anderson-Trocme L, Lingrand M, Berry D, Perus LJM, Wei Y, Moraes C, Alkallas R, Rajkumar S, Zuo D, Dankner M, Xu EH, Bertos NR, Najafabadi HS, Gravel S, Costantino S, Richer MJ, Lund AW, Del Rincon SV, Spatz A, Miller WH, Jamal R, Lapointe R, Mes-Masson AM, Turcotte S, Petrecca K, Dumitra S, Meguerditchian AN, Richardson K, Tremblay F, Wang B, Chergui M, Guiot MC, Watters K, Stagg J, Quail DF, Mihalcioiu C, Meterissian S Watson IR spatially mapping the immune landscape of Melanoma using imaging mass cytometry. Sci Immunol 70(7):eabi5072. https://doi.org/10.1126/sciimmunol.abi5072
Casadonte R, Caprioli RM (2011) Proteomic analysis of formalin-fixed paraffin-embedded tissue by MALDI imaging mass spectrometry. Nat Protoc 11(6):1695–1709. https://doi.org/10.1038/nprot.2011.388
Casadonte R, Kriegsmann M, Kriegsmann K, Hauk I, Meliß RR, Müller CSL, Kriegsmann J (2021) Imaging mass spectrometry-based proteomic analysis to differentiate Melanocytic Nevi and Malignant Melanoma. Cancers 13(13):3197. https://doi.org/10.3390/cancers13133197
Taylor MJ, Lukowski JK, Anderton CR (2021) Spatially resolved Mass Spectrometry at the single cell: recent innovations in Proteomics and Metabolomics. JASMS 4(32):872–894. https://doi.org/10.1021/jasms.0c00439
Hoefsmit EP, Rozeman EA, Van TM, Dimitriadis P, Krijgsman O, Conway JW, da Silva IP, van der Wal JE, Ketelaars SLC, Bresser K (2020) Comprehensive analysis of cutaneous and uveal Melanoma liver metastases. JITC 2(8):e001501. https://doi.org/10.1136/jitc-2020-001501
Thrane K, Eriksson H, Maaskola J, Hansson J, Lundeberg J (2018) Spatially resolved Transcriptomics enables dissection of genetic heterogeneity in stage III cutaneous malignant Melanoma. Cancer Res 20(78):5970–5979. https://doi.org/10.1158/0008-5472.CAN-18-0747
Quek C, Bai X, Long GV, Scolyer RA, Wilmott JS (2021) High-Dimensional Single-Cell Transcriptomics in Melanoma and Cancer Immunotherapy. Genes 10(12):1629. https://doi.org/10.3390/genes12101629
Parra ER, Zhai J, Tamegnon A, Zhou N, Pandurengan RK, Barreto C, Jiang M, Rice DC, Creasy C, Vaporciyan AA, Hofstetter WL, Tsao AS, Wistuba II, Sepesi B, Haymaker C (2021) Identification of distinct immune landscapes using an automated nine-color multiplex immunofluorescence staining panel and image analysis in paraffin Tumor tissues. Sci Rep 1(11):4530. https://doi.org/10.1038/s41598-021-83858-x
Buchberger AR, DeLaney K, Johnson J, Li L (2018) Mass Spectrometry Imaging: a review of emerging advancements and future insights. Anal Chem (90)1:240–265. https://doi.org/10.1021/acs.analchem.7b04733
Keren L, Bosse M, Thompson S, Risom T, Vijayaragavan K, McCaffrey E, Marquez D, Angoshtari R, Greenwald NF, Fienberg H, Wang J, Kambham N, Kirkwood D, Nolan G, Montine TJ, Galli SJ, West R, Bendall SC Angelo M MIBI-TOF: a multiplexed imaging platform relates cellular phenotypes and tissue structure. Sci Adv 10(5):eaax5851. https://doi.org/10.1126/sciadv.aax5851
Angelo M, Bendall SC, Finck R, Hale MB, Hitzman C, Borowsky AD, Levenson RM, Lowe JB, Liu SD, Zhao S, Natkunam Y, Nolan GP (2014) Multiplexed ion beam imaging of human breast tumors. Nat Med 4(20):436–442. https://doi.org/10.1038/nm.3488
Beasley GM, Therien AD, Holl EK, Al-Rohil R, Selim MA, Farrow NE, Pan, Haynes P, Liang Y, Tyler DS, Hanks BA, Nair SK (2021) Dissecting the immune landscape of Tumor draining lymph nodes in Melanoma with high-plex spatially resolved protein detection. CII 70(2):475–483. https://doi.org/10.1007/s00262-020-02698-2
Ståhl PL, Salmén F, Vickovic S, Lundmark A, Navarro JF, Magnusson J, Giacomello S, Asp M, Westholm JO, Huss M, Mollbrink A, Linnarsson S, Codeluppi S, Borg Å, Pontén F, Costea PI, Sahlén P, Mulder J, Bergmann O, Lundeberg J, Frisén J (2016) Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 6294(353):78–82. https://doi.org/10.1126/science.aaf2403
Chen KH, Boettiger AN, Moffitt JR, Wang S, Zhuang X (2015) Spatially resolved, highly multiplexed RNA profiling in single cells. Science 6233(348):aaa6090. https://doi.org/10.1126/science.aaa6090
Hernandez S, Lazcano R, Serrano A, Powell S, Kostousov L, Mehta J, Khan K, Lu W, Solis LM (2022) Challenges and opportunities for Immunoprofiling using a spatial high-Plex Technology: the NanoString GeoMx® Digital spatial profiler. Front Oncol (12):890410. https://www.frontiersin.org/articles/10.3389/fonc.2022.890410
Piñeiro AJ, Houser AE, Ji AL (2022) Research Techniques Made Simple: Spatial Transcriptomics. JID 4(142):993–1001.e1001. https://doi.org/10.1016/j.jid.2021.12.014
Kiuru M, Kriner MA, Wong S, Zhu G, Terrell JR, Li Q, Hoang M, Beechem J, McPherson JD (2022) High-plex spatial RNA profiling reveals cell type–specific biomarker expression during Melanoma development. JID 5(142):1401–1412.e20. https://doi.org/10.1016/j.jid.2021.06.041
Bandura DR, Baranov VI, Ornatsky OI, Antonov A, Kinach R, Lou X, Pavlov S, Vorobiev S, Dick JE, Tanner SD (2009) Mass Cytometry: technique for real time single cell multitarget immunoassay based on inductively coupled plasma time-of-flight mass spectrometry. Anal Chem 16(81):6813–6822. https://doi.org/10.1021/ac901049w
Iyer A, Hamers AAJ, Pillai AB (2022) CyTOF® for the masses. Front Immunol (13):815828. https://www.frontiersin.org/articles/10.3389/fimmu.2022.815828
Gadalla R, Noamani B, MacLeod BL, Dickson RJ, Guo M, Xu W, Lukhele S, Elsaesser HJ, Razak ARA, Hirano N (2019) Validation of CyTOF against flow cytometry for immunological studies and monitoring of human cancer clinical trials. Front Oncol (9):415. https://doi.org/10.3389/fonc.2019.00415
Lee BH, Kelly G, Bradford S, Davila M, Guo XV, Amir EAD, Thrash EM, Solga MD, Lannigan J, Sellers B, Candia J, Tsang J, Montgomery RR, Tamaki SJ, Sigdel TK, Sarwal MM, Lanier LL, Tian Y, Kim C, Hinz D, Peters B, Sette A, Rahman AH (2019) A modified injector and sample acquisition protocol can improve data quality and reduce inter-instrument variability of the Helios mass cytometer. Cytometry A 9(95):1019–1030. https://doi.org/10.1002/cyto.a.23866
Giesen C, Wang HAO, Schapiro D, Zivanovic N, Jacobs A, Hattendorf B, Schüffler PJ, Grolimund D, Buhmann JM, Brandt S, Varga Z, Wild PJ, Günther D, Bodenmiller B (2014) Highly multiplexed imaging of Tumor tissues with subcellular resolution by mass cytometry. Nat Methods 4(11):417–422. https://doi.org/10.1038/nmeth.2869
Levine JH, Simonds EF, Bendall SC, Davis KL, Amir ED, Tadmor MD, Litvin O, Fienberg HG, Jager A, Zunder ER, Finck R, Gedman AL, Radtke I, Downing JR, Pe’er D, Nolan GP (2015) Data-driven phenotypic dissection of AML reveals progenitor-like cells that correlate with prognosis. Cell 1(162):184–197. https://doi.org/10.1016/j.cell.2015.05.047
Van Gassen S, Callebaut B, Van Helden MJ, Lambrecht BN, Demeester P, Dhaene T, Saeys Y (2015) FlowSOM: using self-organizing maps for visualization and interpretation of cytometry data. Cytometry A 7(87):636–645. https://doi.org/10.1002/cyto.a.22625
Geuenich MJ, Hou J, Lee S, Ayub S, Jackson HW, Campbell KR (2021) Automated assignment of cell identity from single-cell multiplexed imaging and proteomic data. Cell Syst 12(12):1173–1186.e1175. https://doi.org/10.1016/j.cels.2021.08.012
Jonas W, Bernd B, Nils E (2021) An end-to-end workflow for multiplexed image processing and analysis. bioRxiv:2021.2011.2012.468357. http://biorxiv.org/content/early/2021/11/13/2021.11.12.468357.abstract
Milosevic V (2023) Different approaches to Imaging Mass Cytometry data analysis. Bioinform Adv 1(3):vbad046. https://doi.org/10.1093/bioadv/vbad046.
Stringer C, Wang T, Michaelos M, Pachitariu M (2021) Cellpose: a generalist algorithm for cellular segmentation. Nat Methods 1(18):100–106. https://doi.org/10.1038/s41592-020-01018-x
Schmidt U, Weigert M, Broaddus C, Myers G (2018) Cell detection with Star-Convex polygons. In: Frangi AF, Schnabel JA, Davatzikos C, Alberola-López C, Fichtinger G (eds) Medical Image Computing and Computer assisted intervention – MICCAI 2018, vol 11071. Springer International Publishing, Cham, p 265–273
Greenwald NF, Miller G, Moen E, Kong A, Kagel A, Dougherty T, Fullaway CC, McIntosh BJ, Leow KX, Schwartz MS, Pavelchek C, Cui S, Camplisson I, Bar-Tal O, Singh J, Fong M, Chaudhry G, Abraham Z, Moseley J, Warshawsky S, Soon E, Greenbaum S, Risom T, Hollmann T, Bendall SC, Keren L, Graf W, Angelo M, Van Valen D (2022) Whole-cell segmentation of tissue images with human-level performance using large-scale data annotation and deep learning. Nat Biotechnol 4(40):555–565. https://doi.org/10.1038/s41587-021-01094-0
Zanotelli VRT, Bodenmiller B (2022) ImcSegmentationPipeline: A pixel-classification based multiplexed image segmentation pipeline. Zenodo. https://doi.org/10.5281/zenodo.3841961. Accessed 30 August 2023
Karimi E, Rezanejad M, Fiset B, Perus L, McDowell SAC, Arabzadeh A, Beugnot G, Siegel P, Guiot MC, Quail DF (2022) Machine learning meets classical computer vision for accurate cell identification. bioRxiv:2022.2002.2027.482183. http://biorxiv.org/content/early/2022/02/28/2022.02.27.482183.abstract
Aghaeepour N, Nikolic R, Hoos HH, Brinkman RR (2011) Rapid cell population identification in flow cytometry data. Cytometry A 1(79A):6–13. https://doi.org/10.1002/cyto.a.21007
Zhang W, Li I, Reticker-Flynn NE, Good Z, Chang S, Samusik N, Saumyaa S, Li Y, Zhou X, Liang R, Kong CS, Le QT, Gentles AJ, Sunwoo JB, Nolan GP, Engleman EG, Plevritis SK (2022) Identification of cell types in multiplexed in situ images by combining protein expression and spatial information using CELESTA. Nat Methods 6(19):759–769. https://doi.org/10.1038/s41592-022-01498-z
Jackson HW, Fischer JR, Zanotelli VRT, Ali HR, Mechera R, Soysal SD, Moch H, Muenst S, Varga Z, Weber WP, Bodenmiller B (2020) The single-cell pathology landscape of Breast cancer. Nature 7796(578):615–620. https://doi.org/10.1038/s41586-019-1876-x
Sorin M, Rezanejad M, Karimi E, Fiset B, Desharnais L, Perus LJM, Milette S, Yu MW, Maritan SM, Doré S, Pichette É, Enlow W, Gagné A, Wei Y, Orain M, Manem VSK, Rayes R, Siegel PM, Camilleri-Broët S, Fiset Pierre O, Desmeules P, Spicer JD, Quail DF, Joubert P, Walsh LA (2023) Single-cell spatial landscapes of the lung tumour immune microenvironment. Nature 7948(614):548–554. https://doi.org/10.1038/s41586-022-05672-3
Krieg C, Nowicka M, Guglietta S, Schindler S, Hartmann FJ, Weber LM, Dummer R, Robinson MD, Levesque MP, Becher B (2018) High-dimensional single-cell analysis predicts response to anti-PD-1 immunotherapy. Nat Med 2(24):144–153. https://doi.org/10.1038/nm.4466
Gide TN, Quek C, Menzies AM, Tasker AT, Shang P, Holst J, Madore J, Lim SY, Velickovic R, Wongchenko M, Yan Y, Lo S, Carlino MS, Guminski A, Saw RPM, Pang A, McGuire HM, Palendira U, Thompson JF, Rizos H, Silva IP, Batten M, Scolyer RA, Long GV, Wilmott JS (2019) Distinct Immune cell populations define response to Anti-PD-1 monotherapy and Anti-PD-1/Anti-CTLA-4 combined Therapy. Cancer Cell 2(35):238–255.e236. https://doi.org/10.1016/j.ccell.2019.01.003
Azimi F, Scolyer RA, Rumcheva P, Moncrieff M, Murali R, McCarthy SW, Saw RP, Thompson JF (2012) Tumor-infiltrating lymphocyte grade is an independent predictor of sentinel lymph node status and survival in patients with cutaneous Melanoma. J Clin Oncol 21(30):2678–2683. https://doi.org/10.1200/JCO.2011.37.8539
The Cancer Genome Atlas Network (2015) Genomic classification of cutaneous Melanoma. Cell 7(161):1681–1696. https://doi.org/10.1016/j.cell.2015.05.044
Hamid O, Schmidt H, Nissan A, Ridolfi L, Aamdal S, Hansson J, Guida M, Hyams DM, Gómez H, Bastholt L, Chasalow SD, Berman D (2011) A prospective phase II trial exploring the association between Tumor microenvironment biomarkers and clinical activity of ipilimumab in advanced Melanoma. J Transl Med 1(9):204. https://doi.org/10.1186/1479-5876-9-204
Schulz D, Zanotelli VRT, Fischer JR, Schapiro D, Engler S, Lun XK, Jackson HW, Bodenmiller B (2018) Simultaneous multiplexed imaging of mRNA and proteins with subcellular resolution in Breast Cancer tissue samples by Mass Cytometry. Cell Syst 1(6):25–36.e25. https://doi.org/10.1016/j.cels.2017.12.001
Routy B, Lenehan JG, Miller WH, Jamal R, Messaoudene M, Daisley BA, Hes C, Al KF, Martinez-Gili L, Punčochář M, Ernst S, Logan D, Belanger K, Esfahani K, Richard C, Ninkov M, Piccinno G, Armanini F, Pinto F, Krishnamoorthy M, Figueredo R, Thebault P, Takis P, Magrill J, Ramsay L, Derosa L, Marchesi JR, Parvathy SN, Elkrief A, Watson IR, Lapointe R, Segata N, Mansour Haeryfar SM, Mullish BH, Silverman MS, Burton JP, Maleki Vareki S (2023) Fecal microbiota transplantation plus anti-PD-1 immunotherapy in advanced Melanoma: a phase I trial. Nat Med 8(29):2121–2132. https://doi.org/10.1038/s41591-023-02453-x
Kuett L, Catena R, Özcan A, Plüss A, Schraml P, Moch H, de Souza N, Bodenmiller B, Consortium Cancer Grand Challenges Imaxt (2022) Three-dimensional imaging mass cytometry for highly multiplexed molecular and cellular mapping of tissues and the Tumor microenvironment. Nat Cancer 1(3):122–133. https://doi.org/10.1038/s43018-021-00301-w
Acknowledgements
Ian R. Watson is supported by funding from the Canada Research Chairs Program; Canadian Institute of Health Research (CIHR – Grant # PJT-178341, PJT-180496, PJT 186002); Omics Data Against Cancer (ODAC) grant funded by Genome Quebec, Oncopole and IVADO; the Victor Liu McGill Interdisciplinary Initiative in Infection and Immunity (MI4); generous donations from Dr. K. Jean Baggs; and generous donations from the Rachel and Jason Schwartz Family Foundation. Jamie Magrill is supported by a McKeown Scholarship from the McGill MD-PhD Program and a Thérèse & David Bohbot Scholarship from JCF Montreal, as well as generous funding from the St. John’s Legacy Foundation.
Author information
Authors and Affiliations
Contributions
JM drafted the initial manuscript, created figures, and carried out substantial revisions and editing. DM, JG, and ML provided intellectual input and carried out revisions and editing on the manuscript. IRW provided intellectual input and carried out substantial revisions and editing on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Presented at the 9th International Congress on Cancer Metastasis through the Lymphovascular System, May 4–6, 2023, in San Francisco, CA. To be published in a Special Issue of Clinical and Experimental Metastasis: Molecular Mechanisms of Cancer Metastasis.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Magrill, J., Moldoveanu, D., Gu, J. et al. Mapping the single cell spatial immune landscapes of the melanoma microenvironment. Clin Exp Metastasis 41, 301–312 (2024). https://doi.org/10.1007/s10585-023-10252-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10585-023-10252-4