Abstract
Immunotherapy has been established as a promising therapy for different cancer types. However, many patients experience primary or secondary resistance to treatment. Immune cells and anti-inflammatory factors are regulated by long noncoding RNAs (lncRNAs). In addition, lncRNAs have a role in immune resistance through antigen presentation loss or attenuation, PD-L1 upregulation, loss of T-cell activities, and activation of G-MDSCs and Tregs in the tumor environment. LncRNAs can also influence the interaction between cancer stem cells and immune cells in the tumor microenvironment, potentially resulting in cancer stem cell resistance to immunotherapy. Immunological-related lncRNAs can influence immune responses either directly by affecting neighboring protein-coding genes or indirectly by sponging miRNAs through various mechanisms. We have emphasized the role and levels of expression of lncRNAs that have been linked to immune cell formation, differentiation, and activation, which may have an influence on immunotherapy efficacy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer and the immune system are bonded strongly together. Immune cells switch from a passive sentinel to an active responder when they detect foreign threats or endogenous alterations in the microenvironment [1]. Cancer immunotherapy (CI) is one of the developing therapy forms which can supplement conventional cancer treatments like chemotherapy, radiation and surgery [2]. Conventional cancer treatments can be destructive to all dividing cells leading to severe side effects like anemia, exhaustion, and emotional stress [3]. However, CI aims to activate the immune system directly to identify and destroy tumor cells and is therefore potentially more specific. The primary characteristics of CI are the broadness of response, specificity and memory against tumor antigens, which can result in better clinical outcomes and consequently improve quality of life, especially in patients with metastatic disease [4]. CI encompasses many different approaches like the use of monoclonal antibodies (mAbs), checkpoint inhibitors (CPIs), cytokines, vaccines, and a chimeric antigen receptor-T cell (CAR-T cell) therapy (Fig. 1) [5]. These approaches can be either active or passive immunotherapies.
Types of cancer immunotherapies. The figure depicts different cancer immunotherapies, including monoclonal antibodies (mAbs), checkpoint inhibitors, cytokines, vaccines, and chimeric antigen receptor-T cells (CAR-T cells). The figure was created using Biorender (https://biorender.com/)
In passive immunotherapies, immune checkpoint inhibitors (ICIs) are different monoclonal antibodies which deactivate checkpoint receptors and activate T cells to kill cancerous cells. Anti-PDL1 and anti-CTLA-4 ICIs are passive immunotherapy drugs which have recently been used [6]. Active immunotherapies mainly aim to produce long-lasting memory responses, via the exploitation of immune cells such as dendritic cells which are potent immunomodulatory cytokines and antigen-presenting cells (APC) [7]. However, there are various obstacles to CI which make active and passive immunotherapies highly challenging. The most common challenge is that many individuals fail to respond to CI due to high mutation rates. This suggests that CI must be individualized by carefully comprehending and identifying the rate-limiting step operating on a specific patient [8]. Several cellular mechanisms can affect the functions of the immune system, which could influence the patient’s response to immunotherapy. LncRNAs are one of these strategies. LncRNAs are transcripts that are longer than 200 nucleotides and lack the ability of protein-encoding [9], due to the presence of 7-methyl guanosine (m7G) at their 5′ ends. LncRNAs are not appropriately spliced at the 3’ ends compared to mRNA, which has full splicing at the 3’ end [10]. Although more than 90% of lncRNAs have no protein-coding potential, certain transcripts have been shown to produce short open reading frames (ORFs) of 300 nt or less, which could be translated into small peptides [11].
LncRNAs have a critical role in determining immune responses, therefore they form a logical class of target in cancer immunotherapy studies. Because of their condition-specific expression pattern, lncRNAs are an appealing target for prospective biomarkers and treatments [12]. LncRNAs play a role in immune cells` regulation system; for example; in many particular types of immune cells, the expression of lncRNAs is stimulated by the activation of Toll-like receptor (TLR) when bind with intracellular signalling pathways (such as NF-κB). Also, it can be suppressed by activation of cytokines receptors such as IL-6 or TGF-β, consequently, mediating immune responses [13]. In addition, lncRNAs control cytokines and immune checkpoints and enhance the development of the immunosuppressive environment, which either promotes the progression or suppression of tumor and drug resistance [14]. In this review, we try to summarize the critical roles and functions of these lncRNA-mediated approaches, which have considerable potential in immunotherapies to facilitate cancer treatment and diagnosis.
The role of lncRNAs in gene expression
LncRNAs have a crucial role in different biological processes, including transcriptional activation and interference, chromatin remodeling, mRNA translation and RNA processing [15] (Fig. 2). Most lncRNAs’ biological activities and molecular mechanisms are still substantially unknown, with only a few being partially understood. Existing evidence suggests that these metabolites serve key roles in the epigenetic, transcriptional, and post-transcriptional control of various cellular activities, including protein-coding gene expression [16].
Mechanisms of lncRNAs in the cell. The figure depicts the mechanisms of lncRNAs in the nucleus via (1) enhancing DNA transcription; (2) targeting chromatin-modifying proteins at specified genomic locations; (3) regulating the binding of specific transcription factors; and (4) regulating alternative splicing or in the cytoplasm via (5) acting as a sponge or a decoy for miRNAs; (6) encoding the functional micro-peptides; (7) binding and suppressing the translation of mRNAs; and (8) binding specific mRNAs to enhance stability. The figure was created using Biorender (https://biorender.com/)
As previously noted, most lncRNAs preferentially localize in the nucleus and chromatin, and growing evidence suggests that some nuclear lncRNAs epigenetically control gene expression by modifying chromatin structure [17]. LncRNAs mediate changes in chromatin and gene expression in two ways. First, they interact directly with chromatin-modifying enzymes. They act as guides in cis or trans, attracting chromatin modifiers to specific genomic loci to promote DNA methylation or histone modification. These modifications regulate chromatin states and effects on gene expression [18]. In the second mechanism; lncRNAs act as adaptors, connecting specific chromatin loci to ATP-dependent chromatin-remodelling complexes and serving as guides for the regulation of nucleosome remodelling and gene expression [18].
Furthermore, lncRNAs have been discovered to be important regulators of epigenetic processes, including X-chromosome inactivation, genomic imprinting, cellular differentiation, and maintenance of cell identity [19]. LncRNAs regulate gene expression at the transcriptional level by different mechanisms as (A) Gene transcription regulation; they facilitate mobilization of the transcription factors to the promoter region of target genes, interacting with RNA polymerase II, affecting protein localization, and acting as transcriptional activators or repressors. (B) Repression of target gene transcription; they form lncRNA-DNA hybrids with the target gene and promote DNA methylation or acetylation [20]. LncRNAs mediate mRNA stability, translation, degradation, and pre-mRNA alternative splicing genes. Thus they affect gene expression and regulate biological processes at the post-transcriptional level.
LncRNAs function as competing for endogenous RNAs (ceRNAs) or sponges, decoys, signals, guides and scaffolds according to their mode of action [21]. LncRNAs act as ceRNAs or endogenous microRNAs (miRNAs) sponges through two pathways; (A) they can sequester miRNAs, preventing them from attaching to target mRNAs. (B) In the case in which lncRNAs and miRNAs have common binding sites in the target mRNA, lncRNAs can interact directly with target mRNA transcripts, blocking miRNA binding sites in mRNA molecules. Interactions between miRNAs and ceRNAs are important for regulating many basic biological functions [22].
Decoy lncRNAs act as modulators of transcription by trapping or decreasing the availability of different regulatory factors, including catalytic proteins, miRNAs, transcription factors and sub-units of larger chromatin-modifying complexes [23]. Scaffold lncRNAs have a vital structural role, they serve as platforms for the assembly of multiple-component complexes, and other regulatory co-factors such as ribonucleoprotein (RNP) complexes. RNP complexes can regulate gene expression by targeting certain genomic regions [24]. Guide lncRNAs can regulate transcription by binding with different factors like regulatory or enzymatically active proteins and organize their localization on specific sites on the genome either cis (adjacent) or trans (distant) from their transcriptional locus [25]. Signal lncRNAs function as molecular signals to control transcription in response to different stimuli. As a result, the presence and production of these components act as an indicator for transcription activity and also under specific conditions, they can modulate specific signaling pathways [26].
In summary, lncRNAs are widely expressed and play important roles in gene regulation. They can modulate chromatin function, regulate the assembly and function of membraneless nuclear bodies, modify the stability and translation of cytoplasmic mRNAs, and interfere with signaling pathways depending on their localization and specific interactions with DNA, RNA, and proteins.
The effect of lncRNAs on the efficacy of cancer immunotherapy
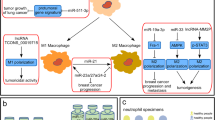
Rapid advances in transcriptomic profiling have led to the identification of immune-related lncRNAs as regulators of immune cell-specific gene expression that mediates both immune stimulation and suppression, implying that lncRNAs could improve the efficacy of immunotherapy against tumors. Tables 1 and 2 summarize the regulatory role of lncRNAs in cancer progression or cancer suppression via different immune responses. Some lncRNAs act as immunomodulators, influencing inflammatory cytokine production, altering innate immune cells, or creating resistance to immunotherapy [27]. Others can affect antigen presentation, control the production of PD-L1, and regulate the actions of immunosuppressive cells to modify the T-cell-mediated immune response that leads to immunosuppression [28]. In this review, we focus on lncRNAs associated with the immune response as a target for improving immunotherapy resistance in different cancer types (Fig. 3). For Example, in acute lymphoblastic leukemia (ALL), lnc- Insulin receptor (INSR) has a role in tumor growth and progression via overexpression of T regulatory cells (Treg) and decreases the percentage of cytotoxic T lymphocytes (CTLs) [29]. In addition, lnc INSR directly attaches and blocks INSR ubiquitination, leading to activation of INSR and PI3K/AKT-signaling pathway, which results in induction of suppressive immune microenvironment (IME).
PI3K is an abundant protein in white blood cells and abnormality in its signaling pathway inhibits T cell activation. Also, the PI3K/AKT pathway enables T cell receptor signaling to control the expression of forkhead box P3 (Foxp3), which binds to certain areas of DNA and helps to regulate the activity of genes involved in immune system regulation [30]. LncRNA HOTAIR increases the immunologic rejection in mice leukaemia cells via the activation of the Wnt/-catenin pathway, in which mice given HOTAIR mimics as well as small interfering RNA HOTAIR. The results revealed the up regulation of lncRNA HOTAIR, which resulted in an increase in the percentage of leukocytes and decreasing the percentage of PLTs and haemoglobin concentration. Furthermore, increased HOTAIR suppressed Ig synthesis, natural killer (NK) cell activity, and the ratio of CD4/CD8 T cell subsets in B-lymphocytes, and lowered the levels of transforming growth factor, interferon, interleukin-10, and tumor necrosis factor in PB and T-lymphocyte proliferation. Additionally, over-expressed HOTAIR boosted the expression of cyclinD1, GSK-3, and c-Myc in the bone marrow of mice, resulting in better survival and proliferation [31].
In breast cancer, lnc- T-cell leukemia/lymphoma 6 (TCL6) correlated with immune infiltration and is therefore considered a useful prognostic molecular marker. Low TCL6 expression is linked to a poor prognosis, particularly in progesterone receptor-negative (PR) and luminal B breast cancer patients. Additionally, the lncRNA TCL6 may influence immune-related pathways, including JAK/STAT. TCL6 regulates tumor-associated B cells, CD8 + T cells, CD4 + T cells, neutrophils, and Dendritic cells (DCs). Furthermore, TCL6 is associated with tumor-infiltrating lymphocytes (TILs) and immunological checkpoint markers, including PD-1, PD-L1, PD-L2, and CTLA-4 [32].
LncRNA p21 plays an important role as a regulator of tumor-associated macrophages (TAMs) in breast cancer. TAMs show different phenotypes in cancer development, where they have a pro-inflammatory phenotype, generate an anti-tumor type-1 inflammatory response, and suppress tumor cell proliferation by generating TNF, reactive oxygen species (ROS), or phagocytosis during the primary stage of tumor development. In contrast, tumor microenvironment-educated macrophages release IL-10 and TGF and inhibit the activation of CTLs and NK cells at advanced stages of tumor development. Downregulation of lncRNA p21 facilitates interaction between p53 and Mouse double minute 2 homolog (MDM2), resulting in activation of NF-κB and STAT3 signaling pathways, reversed TAMs phenotype and generating TNF-α to destroy cancer cells [33].
LncRNA Cyclooxygenase-2 (Cox-2) is located about 50 kb downstream of the mouse Cox-2 gene. LncRNA Cox-2 is a broad-acting regulatory component of the inflammatory response regulation circuit, where the repression or activation of various immune genes can mediate by it. In hepatocellular carcinoma (HCC), lncRNA Cox-2 has an important role in the suppression of the tumor immune evasion and development by enhancing the polarization of M1 macrophages and suppressing the polarization of M2 macrophages [34]. In contrast, the lnc-epidermal growth factor receptor (EGFR) enhances HCC progression and development. LncRNA EGFR enhances differentiation of Treg and suppresses CTLs by binding with EGFR and blocking its interaction with c-Casitas B-lineage Lymphoma (c-CBL). The latter is an E3 ubiquitin-protein ligase that helps in cell signaling and protein ubiquitination. Additionally, lncRNA EGFR has a positive feedback loop in Treg, where it increases the expression of EGFR resulting in activation of the expression of its downstream extracellular signal-regulated kinases 1/2 (ERK1/2), Activator protein 1 (AP-1) and Foxp3 expression, which induce immunosuppression in HCC patients [35].
In a recent in silico study, Xia P. and colleagues used a similar strategy with The Cancer Genome Atlas (TCGA) and Chinese Glioma Genome Atlas (CGGA) patients. A total number of 812 immune-related lncRNAs founded to be linked directly to glioma. The authors developed a risk score formula using Cox regression and the least absolute shrinkage and selection operator (LASSO) analysis to investigate the differences in overall survivals (OSs) between high- and low-risk groups. The risk score (RS) method contained eleven immune-related lncRNAs that were linked with survival. There is a marked relation between high-risk score cases that had poor survival in both the TCGA and CGGA groups related to Glioma patients. The RS formula exhibited a good prediction accuracy in the CGGA dataset equals (5-year Area under Curve (AUC) = 0.730) and could accurately forecast the prognosis of glioma patients that equals (5-year AUC = 0.749). These results from the model’s robust prediction function, give specific guiding values for glioblastoma etiology and clinical treatment analysis, allowing users to find prospective therapeutic targets for glioma treatment [36].
In another study, the immune-related lncRNA signature (IRLS) in bladder cancer was determined using the LASSO Cox regression model. Using LASSO Cox regression analysis, the authors chose five prognostic lncRNAs to create IRLS. They discovered that IRLS positively linked with immune cell infiltration and expression of important immunological checkpoints in the tumor microenvironment (TME), implying that immunosuppressive TME may play a role in poor prognosis [37]. LncRNA NKX2-1-AS1 was originally detected in lung carcinoma cells. Overexpression of lncRNA NKX2-1-AS1 can reduce cell migration and immune system evasion by decreasing the expression of CD274, the gene encoding PD-L1 [38]. In contrast, lncRNA plasmacytoma variant translocation 1 (Pvt1) induces lung cancer progression and development, where HIF-1α regulates the expression of Pvt1 under hypoxia. Pvt1 activates Myeloid-derived suppressor cells (MDSCs) which have a role in blocking T-cell-induced antitumor responses resulting in induction of immunosuppression activity in lung carcinoma [39].
In summary, lncRNAs are expressed in a variety of immune cells and have a role in both innate and adaptive immunity. The discovery of lncRNAs has provided a unique perspective for investigating the regulation of TME. LncRNAs promote the formation of an immunosuppressive microenvironment through related different pathways, like the PI3K/AKT pathway, Wnt/-catenin pathway, JAK/STAT pathway and TNF-α/ NF-κB pathway, thereby controlling the escape of tumors from immune surveillance and promoting the development of metastasis and drug resistance in different cancer types.
The role of lncRNAs in cancer stem cell resistance to immunotherapy
The cancer immune-editing theory and the hierarchy model of cancer initially explained the ability of cancer stem cells (CSCs) to resist immunological destruction. Pluripotent and long-living cancer cells are subpopulations of CSCs which are responsible for drug resistance, recurrence, and metastasis and tumor initiation [40]. CSCs can hide from the immune-mediated destruction and interact with different immune cells by releasing several intracellular, soluble and membrane-bound factors that allow them to survive and inhibit innate and adaptive anti-tumor immune responses in their niche [41]. The CSCs can dysregulate the expression of various anti-apoptotic proteins, including survivin, Bcl-xL, and Bcl-2 in different tumor types. Also, CSCs can evade immune-mediated apoptosis promoted by NK and effector T cells, or immunotherapeutic antibodies by secreting many cytokines and other molecules, such as IL-10, IL-6, IL-4, FasL, and prostaglandin E2 (PGE2). In addition, CSCs maintain the immunosuppressive TME and enhance tumor infiltration by releasing diverse immunosuppressive cytokines and chemotactic factors such as TGF-β, IL-10 and IL-2. These factors otherwise suppress the activities of effector immune cells such as NK cells, CTLs and T cells or/and enhance immunosuppressive immune cell subpopulations, including MDSCs, Tregs and TAMs [42]. Also, other studies on multifaceted interactions between CSCs and the immune system showed that CSCs can recruit and enhance the properties of macrophages into TAMs. TAMs usually express M2 phenotypes which have pro-tumor and immunosuppressive properties and are related to cancer progression and recurrence [43]. TAMs are required for CSCs self-renewal and maintenance in different cancer types via activation of the STAT3/NF-κB pathway (Sainz et al., 2016). DCs are APCs that generate innate or adaptive immune responses. CSCs can recruit immunosuppressive properties of DCs by producing immunosuppressive cytokines, including IL-13, IL-10, IL-4, and co-inhibitory molecules like B7-H3, IDO1 and PD-L1 which induce tumor properties of DCs [44].
The MDSCs play a crucial role in enhancing an immunosuppressive environment in TME and are employed as a prognostic biomarker for patients` response to immunotherapy and their rate of survival. The MSDCs can be categorized into two subtypes based on their different nuclear morphologies: granulocytic-MDSC (gMDSC) and monocytic-MDSC (mMDSC). They have two different immunosuppressive mechanisms. Cross linking between CSCs and MDSCs at the tumor site can enhance tumorigenesis and metastasis in different cancer types. For example; CSCs activate IL-6/STAT3 signaling, which enhances the differentiation of monocytes to MDSCs in breast cancer (Welte et al., 2016). Meanwhile, in ovarian cancer, MDSCs promote the stem-like features of CSCs by increasing the expression of (PGE2) and PD-L1 [45].
In HCC, MDSCs move to the tumor site via ENTPD2/CD39 L1 signaling under hypoxic conditions where liver CSCs are abundant. These MDSCs promote HCC development and decrease the efficiency of PD1 treatment (Chiu et al., 2017). Tregs are CD4 + T cells that promote tumor growth and are usually identified by the Foxp3 + CD25 + CD4 + T cell subset. They suppress effector T cells and other immune cells by secreting immunosuppressive cytokines such as IL-35, IL-10, and TGF- β. The interplay between CSCs and Treg in TME, suggests their roles in maintaining immunosuppression and promoting tumor infiltration of different types of cancers. Mainly, Treg infiltration is induced by CSCs in the TME via co-stimulatory molecules and STAT3 signaling, whereas Tregs regulate CSCs proliferation and expansion either directly through IL-17 and PGE2 production or indirectly through TGF-mediated angiogenesis and epithelial-mesenchymal transition (EMT) [44].
NK cells represent a subpopulation of cytotoxic lymphocytes, known as large granular lymphocytes (LGL) which account for 5–20% of all circulating lymphocytes in humans. NKs are critical for innate immune response and are considered one of the cell types which has the most potential for targeting and killing the tumor cells. CSCs can evade NK by increasing the expression of MHC-I on their surface, where most immune responses are triggered by NK in tumor cells through down-regulation of MHC-I. Also, EMT-derived NK cell is one of the cellular pathways which NK cells promote in CSCs according to the type of cancer. For example; in melanoma, NK cells secret IFN-γ and TNF-α and enhance CSCs to undergo EMT, inducing them to invasive phenotypes and increasing the expression of stemness markers [46].
In contrast, in colorectal cancer, EMT promotes anti-tumor immune response by increasing the expression of NK group 2, member D (NKG2D) receptor which has an important role in cancer immunosurveillance [47]. LncRNAs have a potential impact on immune modulation and regulation of CSCs. Several studies showed that TME and CSCs niches have a crucial role in the maintenance of lncRNA-regulated therapy resistance in CSCs. For example, in HCC and breast cancer, Hypoxia-associated lncRNA (HAL) and lncRNAs runt-related transcription factor 1 Intronic Transcript 1 (RUNX1-IT1) enhance cancer stemness and increase resistance to apoptosis, under the regulation of Hypoxia which represents one of the key features of TME [48, 49]. As mentioned above, crosslinking between CSCs and two pathways- TNF-α/ NF-κB signaling and IL-6/STAT3- regulates the differentiation of many immune cells in TME, promoting immunosurveillance and cancer progression.
LncRNA down-regulated in liver cancer stem cells (lnc-DILC) has an inhibitory role in liver cancer stem cells’ self-renewal and suppresses its expansion by down-regulation of IL-6 transcription and STAT3 activation. In addition, lnc-DILC modulates the crosstalk between TNF-α/ NF-κB signalling and IL-6/STAT3, resulting in a decreased hepatic inflammation and liver cancer stem cell proliferation [50]. Another study showed that crosslinking between lncRNA H19 and TAMs in EMT induced stemness and accelerated HCC invasion of HCC cells in vitro through modulation and activation of the miR-193b/MAPK1 axis, suggesting that lncRNA H19 could be a promising anti-cancer therapy strategy for reducing HCC aggression and improving clinical outcomes [51].
In glioblastoma, stem-like subpopulation cells which are exited in multiform and have a role in tumor progression, resistance to apoptosis and recurrence are called glioma stem cells (GSCs). Crosstalk of the miR-146b-5p/HuR/lincRNA-p21 axis inhibits expression and activity of β-catenin, resulting in increased apoptosis and radio sensitivity, decreased cell viability, neurosphere formation capacity and stem cell marker expression, and induced differentiation in GSCs [52]. LncRNA HOTAIR regulates proliferation, invasion, colony formation, and self-renewal capacity through the modulation of the miR-34a/ Sox2/ p53 /p21 axis in breast cancer CSCs [53].
In summary, CSCs release a variety of immunosuppressive cytokines and chemotactic factors such as; PGE2, IL-10, TGF-β, IL-4, IL-13 and IL-35 that affect different immune cells in the TME and promote tumor invasion and progression. Also, the interaction between lncRNAs, CSCs and immune cells in TME, has a key role in immune escape and resistance to immunotherapies through different cellular pathways including; EMT, IL-6/STAT3 signaling pathway, STAT3/NF-κB pathway and TNF-α/ NF-κB pathway.
The effect of lncRNAs on the miRNAs involved in immunotherapy responses
The miRNAs are a subset of noncoding RNA with a length of ~ 22 nucleotides that have a functional role in the posttranscriptional regulation of protein-coding genes by mRNA cleavage, direct translational suppression, and/or mRNA instability. LncRNAs affect miRNAs through a variety of pathways, as mentioned above, which control a variety of biological processes. The interaction between miRNAs and lncRNAs can influence tumor growth, invasion, and metastasis by enhancing the activation of oncogenic pathways and reducing the expression of tumor suppressors or vice versa [54].
The relationship between lncRNAs and miRNAs was shown to alter cancer immunotherapy resistance in several earlier investigations (Tables 3 and 4). For Example, in breast cancer, overexpression of lncRNA small nucleolar RNA host gene 1 (SNHG1) could decrease immune escape and tumor progression by enhancing the expression of miRNA- 448 and repression of Indoleamine 2,3-dioxygenase (IDO) level, resulting in inhibition of Treg differentiation. The IDO is a vital component of the immune system that aids in natural defense against a variety of infections. Also, it has a role in the suppression of T cell immunity and promoting the maturation and differentiation of Treg cells [55]. In contrast, another study by Liang and his colleagues showed that knockdown of lncRNA breast cancer-related transcript 1 (BCRT1) could decrease breast cancer progression in vitro and in vivo. LncRNA BCRT1 acts as a ceRNA for miRNA-1303 which targets Poly pyrimidine tract-binding protein 3 (PTBP3), thus protecting PTBP3 from degradation and promoting breast cancer progression. PTBP3 is an essential RNA-binding protein that regulates gene expression and influences the biological behaviour of many malignancies. It also plays a key role in RNA alternative splicing. In addition, lncRNA BCRT1 can transfer to macrophages by exosomes, accelerating M2 polarization and promoting its impact on tumor progression [56].
In HCC, overexpression of lncRNA fetal-lethal non-coding developmental regulatory RNA (FENDRR) reduces Treg differentiation, and cell proliferation and induces apoptosis in vitro and in vivo by sponging miRNA-423-5p [57]. In contrast, LINC00662 enhanced HCC tumor progression and metastasis in vivo by competitively binding and inhibiting miRNA-15a, miRNA‐16, and miRNA‐107, up regulation of WNTA3 and activation of Wnt/β‐catenin signaling and M2 macrophage polarization [58].
Diffuse large B cell lymphoma (DLBCL) is the most common disorder derived from the B-lymphocytes. LncRNA small nucleolar RNA host gene 14 (SNHG14) induced immune evasion and DLBCL progression by sponging miRNA-5590-3p, increasing the expression of Zinc finger E-box binding homeobox 1 (ZEB1). ZEB1 is a transcriptional factor that functions as an oncogene regulating migration, invasion, EMT, and development of various types of cancer [59].
Meanwhile, lncRNA SMAD5 antisense RNA 1 (SMAD5-AS1) suppressed immune escape and DLBCL progression by directly sponging miRNA-135b-5p, increasing adenomatous polyposis coli gene expression, and suppressing Wnt/β-catenin pathway. This result showed that SMAD5-AS1 could be used as a DLBCL biomarker and treatment target [60].
Interestingly, lncRNA HOTAIR enhances immune escape and tumor progression of both gastric cancer and cervical cancer by promoting the expression of human leukocyte antigen (HLA)-G. HLA-G is a member of the non-classical MHC family and plays a key role in tumor cell escape from host immune surveillance by inhibiting immune cell activities. In gastric cancer, HOTAIR acts as a ceRNA by sponging miRNA-152, resulting in promoting HLA-G expression [61]. Meanwhile, in cervical cancer, HOTAIR triggers the expression of HLA-G by directly sponging and inhibiting miRNA-148a [62].
In summary, both miRNAs and lncRNAs can influence tumor growth, invasion, and metastasis by enhancing the activation of oncogenic pathways and reducing the expression of tumor suppressors, hence regulating tumor growth, invasion, and metastasis. Some examples of lncRNAs-miRNAs interactions which contributed to enhancement of immune escape and tumor progression of different cancer types, include lncRNA SNHG1/ miRNA- 448, lncRNA BCRT1/ miRNA-1303, lncRNA FENDRR/ miRNA-423-5p, lncRNA SNHG14/ miRNA-5590-3p, lncRNA SMAD5-AS1/ miRNA-135b-5p and lncRNA HOTAIR. Also, lncRNA HOTAIR promotes the expression of HLA-G in both gastric and cervical cancer by sponging two different miRNAs; miRNA-152 and miRNA-152, respectively.
Studying the clinical trials of lncRNAs to find a potential prognostic factor for efficient immunotherapy
The relevance of lncRNA as a predictive factor for cancer cell immunotherapy responsiveness was previously demonstrated in a clinical trial. The data for this study came from TCGA of immunotherapy patients, which included 419 cancer patients. Patients were divided into two groups: (a) 348 patients with bladder cancer from the IMvigor 210 trial phase 2 who were treated with the PD-L1 inhibitor “Atezolizumab,“ and (b) 71 patients with melanoma who were treated with anti-PD-1, anti-cytotoxic T-lymphocyte–associated protein 4 (CTLA4), and cytokine tumor vaccine. Other cancer patients who participated in this study were 493 patients with lung squamous cell carcinoma, 1082 patients with breast cancer, 406 patients with bladder cancer, and 457 patients with melanoma. LASSO was included in the study since since the patients were naive to immunotherapy medications. To expect an appropriate immunotherapeutic response, researchers employed a prediction analysis method that produces models with excellent prediction accuracy. Four of the most significant lncRNAs, AC002116-2, AP000251-1, TMEM147-AS1, and NKILA, have functional immune predictions, according to the findings. Patients with lower levels of these lncRNAs had a higher OS, overall response, and full response length [63].
Another clinical investigation found that lncRNAs have a function in the organization of immunity and the TME in non-small cell lung cancer (NSCLC), which might be useful for prognosis. To differentiate a lncRNA Signature (TILSig) as an indication of immune cell infiltration in patients with NSCLC, researchers developed a unique computational technique . This method was developed using information gathered from an integrated investigation of immune and clinical lncRNA profiles in 115 immune cell lines, 187 NSCLC cell lines, and 1533 NSCLC patients. TILSig divided the patients into two categories: (A) Patients in the high-risk group are immune-cold and have less immune cell infiltration; (B) patients in the low-risk group are immunologically hot and have more immune cell infiltration. Seven TILncRNAs (HCG26, PSMB8-AS1, TNRC6C-AS1, CARD8-AS1, HCP5, LOC286437, and LINC02256) were shown to be associated with immune infiltration in NSCLS patients. Furthermore, patients in the immunological-hot group had a considerably higher survival rate and immune cell infiltration than those in the immune-cold group [64].
Another clinical trial intended to create a lncRNA-based risk signature and nomogram that might predict OS in gastric cancer (GC) patients. The primary cohort consisted of 341 patients with clinical and lncRNA expression data in The Cancer Genome Atlas stomach adenocarcinoma (TCGA STAD), the internal validation cohort consisted of 172 randomly assigned patients, and the external validation cohort consisted of 300 patients from the GSE62254 dataset. Gene set enrichment analysis (GSEA) was also used to investigate the pathway enrichment for the risk signature [65]. The expression patterns of various lncRNAs were also investigated in clinical samples from ten GC patients. The findings indicated a 14-lncRNA signature that was strongly associated with GC patients’ OS and performed well on C-index, the area under the curve, and calibration curve assessments. In univariate and multivariate Cox regression analysis, the lncRNA signature was demonstrated to be an independent predictor of GC patients. Therefore, a nomogram combining the lncRNA signature and clinical factors was developed to predict OS in patients with GC in the original cohort, and it demonstrated high predictive values for survival in the TCGA cohort and the other two validation cohorts. GSEA also observed that the newly identified lncRNAs may affect carcinogenesis and prognosis in GC patients via influencing the autophagy pathway. According to experimental validation, the expression of lncRNAs in clinical samples and the STAD dataset followed the same pattern. Both the risk signature and the nomogram are excellent prognostic markers for GC patients, according to these studies [65].
Current challenges and future perspectives
This review mainly focuses on the crosslinking between lncRNAs and TME which affects the response of cancer patients to immunotherapy. Immunotherapy has become one of the most well-established treatment options for a wide range of cancers. The most recent forms of immunotherapies with excellent outcomes, notably in hematologic malignancies, are CPIs and CAR-T-cells [66]. Many changes and mutations in cancer cells as well as other stromal cells in the tumor microenvironment can lead to immune escape and resistance. Immune-related lncRNAs play a key role in the regulation of immunological cell-specific gene expression, which modifies immune processes by controlling the environment and activities of immune cells as well as anti-inflammatory substances [1]. LncRNAs have the features of tissue-specific expression. They are characterized by relative stability in circulating body fluids, which make lncRNAs useful as cancer biomarkers and facilitate non-invasive detection. Immunotherapy based on lncRNAs has numerous advantages. LncRNAs can regulate a number of downstream target genes by participating in a variety of cell signalling pathways, which can help to control cancer treatment. Furthermore, many regulatory sites of lncRNAs can interact and interfere with other molecules, facilitating the development of new structure-based anticancer medicines [67].
More importantly, lncRNAs can also influence the interaction between CSCs and immune cells in the tumor microenvironment, potentially resulting in CSC resistance to immunotherapy and increased tumor growth and progression. Immunological-related lncRNAs might influence immune responses either directly by influencing adjacent protein-coding genes or indirectly by sponging miRNAs through several pathways [68]. Finally, we sought to outline and emphasize the role and levels of expression that have been linked to immune cell formation, differentiation, and activation, which may influence immunotherapeutic responses for a variety of malignancies and other disorders. Future research may be required to elucidate the many processes and pathways of immune-related lncRNAs in the tumor microenvironment, which might influence cancer development and progression.
Conclusion
LncRNAs play a key function in modifying the TME and controlling tumor cell immune escape. As a result, lncRNA-based targeted cancer immunotherapy has a bright future ahead of it. Despite the ongoing issues with lncRNA-based therapy, as research advances and becomes more refined, the use of lncRNA as a therapeutic target will contribute to the development of novel cancer therapeutic techniques.
Abbreviations
- LncRNA:
-
long noncoding RNAs.
- CI:
-
Cancer immunotherapy.
- APC:
-
antigen –presenting cells.
- mAbs:
-
monoclonal antibodies.
- CAR-T cell:
-
antigen receptor-T cell.
- ICIs:
-
immune checkpoint inhibitors.
- m7G:
-
7-methyl guanosine.
- ORFs:
-
open reading frames.
- TLR:
-
Toll-like receptor.
- ceRNAs:
-
competing endogenous RNAs.
- miRNAs:
-
microRNAs.
- RNP:
-
ribonucleoprotein.
- ALL:
-
Acute lymphoblastic leukemia.
- INSR:
-
Insulin receptor.
- Treg:
-
T regulatory cells.
- CTLs:
-
cytotoxic T lymphocytes.
- IME:
-
immune microenvironment.
- Foxp3:
-
forkhead box P3.
- NK:
-
natural killer.
- TCL6:
-
T-cell leukemia/lymphoma 6.
- PR:
-
Progesterone receptor.
- DCs:
-
Dendritic cells.
- TILs:
-
tumor-infiltrating lymphocytes.
- TAMs:
-
tumor-associated macrophages.
- ROS:
-
reactive oxygen species.
- MDM2:
-
Mouse double minute 2 homolog.
- Cox-2:
-
Cyclooxygenase-2.
- HCC:
-
hepatocellular carcinoma.
- EGFR:
-
epidermal growth factor receptor.
- c-CBL:
-
c-Casitas B-lineage Lymphoma.
- ERK1/2:
-
extracellular signal-regulated kinases 1/2.
- AP-1:
-
Activator protein 1.
- TCGA:
-
The Cancer Genome Atlas.
- CGGA:
-
Chinese Glioma Genome Atlas.
- OSs:
-
overall survivals.
- RS:
-
risk score.
- IRLS:
-
immune-related lncRNA signature.
- TME:
-
tumor microenviroment.
- Pvt1:
-
plasmacytoma variant translocation 1.
- MDSCs:
-
Myeloid-derived suppressor cells.
- CSCs:
-
cancer stem cells.
- PGE2:
-
prostaglandin E2.
- EMT:
-
epithelial–mesenchymal transition.
- LGL:
-
large granular lymphocytes.
- HAL:
-
Hypoxia-associated lncRNA.
- RUNX1-IT1:
-
runt-related transcription factor 1 Intronic Transcript 1.
- lnc-DILC:
-
LncRNA down-regulated in liver cancer stem cells.
- GSCs:
-
glioma stem cells.
- SNHG1:
-
small nucleolar RNA host gene 1.
- IDO:
-
Indoleamine 2,3-dioxygenase.
- BCRT1:
-
breast cancer-related transcript 1.
- PTBP3:
-
Poly pyrimidine tract-binding protein 3.
- FENDRR:
-
fetal-lethal non-coding developmental regulatory RNA.
- DLBCL:
-
Diffuse large B cell lymphoma.
- ZEB1:
-
Zinc finger E-box binding homeobox 1.
- SMAD5-AS1:
-
lncRNA SMAD5 antisense RNA 1.
- HLA:
-
human leukocyte antigen.
- NSCLC:
-
non- lung small cell cancer.
- GSEA:
-
Gene set enrichment analysis.
References
Hu QS, Ye YQ, Chan LC, Li YJ, Liang K, Lin AF et al (2019) Oncogenic lncRNA Downregulates Cancer Cell Antigen Presentation and Intrinsic Tumor Suppression. Nat Immunol 7:835–851
Emens LA, Paolo A, Ascierto PK, Darcy S, Demaria, Alexander MM, Eggermont WL, Redmond B, Seliger, Francesco M, Marincola (2017) Cancer Immunotherapy: Opportunities And Challenges In The Rapidly Evolving Clinical Landscape. Eur J Cancer 81:116–129
Ahmed W, Mofed D, Zekri AR, El-Sayed N, Rahouma M, Sabet S (2018) Antioxidant activity and apoptotic induction as mechanisms of action of Withania somnifera (Ashwagandha) against a hepatocellular carcinoma cell line. J Int Med Res 4:1358–1369
Kaufman HL, Atkins MB, Subedi P, Wu J, Chambers J, Joseph Mattingly T et al (2019) The promise of Immuno-oncology: implications for defining the value of cancer treatment. J Immunother Cancer 7:129
Stanculeanu DL, Daniela Z, Lazescu A, Bunghez R, Anghel R (2016) Development of new immunotherapy treatments in different cancer types. J Med Life 9:240–248
Galluzzi L, Vacchelli E, Bravo-San Pedro JM, Buqué A, Senovilla L, Baracco EE et al (2014) Classification of current anticancer immunotherapies. Oncotarget 5:12472–12508
Rossi M, Young JW (2005) Human dendritic cells: potent antigen-presenting cells at the crossroads of innate and adaptive immunity. J Immunol 3:1373–1381
Kakimi K, Karasaki T, Matsushita H, Sugie T (2017) Advances in personalized cancer immunotherapy. Breast Cancer 1:16–24
McLane LM, Abdel-Hakeem MS, Wherry EJ (2019) CD8 T Cell Exhaustion During Chronic Viral Infection and Cancer. Annu Rev Immunol 37:457–495
Statello L, Guo CJ, Chen LL, Huarte M (2021) Gene regulation by long non-coding RNAs and it biological functions. Nat Rev Mol Cell Biol 22:96–118
Choi S, Kim HW, Nam JW (2019) The small peptide world in long noncoding RNAs. Brief Bioinform 20:1853–1864
Winkle M, El-Daly S, Fabbri M, Calin G (2021) Noncoding RNA Therapeutics-Challenges and Potential Solutions. Nat Rev Drug Discov 8:629–651
Atianand MK, Caffrey DR, Fitzgerald KA (2017) Immunobiology of Long Noncoding RNAs. Annu Rev Immunol 35:177–198
Grote P, Wittler L, Hendrix D, Koch F, Währisch S, Beisaw A et al (2013) The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev Cell 24:206–214
Fang Y, Fullwood MJ (2016) Roles, Functions, and Mechanisms of Long Non-coding RNAs in Cancer. Genom Proteom Bioinform 14:42–54
Zhang X, Wang W, Zhu W, Dong J, Cheng Y, Yin Z, Shen F (2019) Mechanisms and Functions of Long Non-Coding RNAs at Multiple Regulatory Levels. Int J Mol Sci 20:5573
Cogill SB, Wang L (2014) Co-expression Network Analysis of Human lncRNAs and Cancer Genes. Cancer Inf 13:49–59
Böhmdorfer G, Wierzbicki AT (2015) Control of Chromatin Structure by Long Noncoding RNA. Trends Cell Biol 10:623–632
Furlan G, Rougeulle C (2016) Function and evolution of the long noncoding RNA circuitry orchestrating X-chromosome inactivation in mammals. Wiley Interdiscip Rev RNA 7:702–722
Yoon JH, Abdelmohsen K, Gorospe M (2013) Posttranscriptional gene regulation by long noncoding RNA. J Mol Biol 19:3723–3730
Chowdhury IH, Narra HP, Sahni A, Khanipov K, Schroeder CLC, Patel J, Fofanov Y, Sahni SK (2017) Expression Profiling of Long Noncoding RNA Splice Variants in Human Microvascular Endothelial Cells: Lipopolysaccharide Effects In vitro.Mediat. Inflamm;3427461
Sebastian-delaCruz M, Gonzalez-Moro I, Olazagoitia-Garmendia A, Castellanos-RubioA (2021) ; 7: p.3
Kallen AN, Zhou XB, Xu J, Qiao C, Ma J, Yan L et al (2013) The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol Cell 52:101–112
Yang L, Froberg J, Lee J (2014) Long noncoding RNAs: fresh perspectives into the RNA world. Trends Biochem Sci 1:35–43
Ma L, Bajic V, Zhang Z (2013) On the classification of long non-coding RNAs. RNA Biol 10:924–933
Gao N, Li Y, Li J, Gao Z, Yang Z, Li Y et al (2020) Long Non-Coding RNAs: The Regulatory Mechanisms, Research Strategies, and Future Directions in Cancers.Frontiers In Oncology; 10
Zhou Y, Zhu Y, Xie Y, Ma X (2019) The Role of Long Non-coding RNAs in Immunotherapy Resistance. Front Oncol 9:1292
Jiang W, Pan S, Chen X, Wang Z, Zhu X (2021) The role of lncRNAs and circRNAs in the PD-1/PD-L1 pathway in cancer immunotherapy. Mol Cancer 20:1
Wang Y, Yang X, Sun X, Rong L, Kang M, Wu P et al (2018) Bone marrow infiltrated Lnc-INSR induced suppressive immune microenvironment in pediatric acute lymphoblastic leukemia.Cell Death Dis;9
Wang J, Cardoso R, Marreros N, Müller N, Lundström-Stadelmann B, Siffert M, Vuitton DA, Boué F, Lin R, Wen H, Gottstein B (2018) Foxp3+ T Regulatory Cells as a Potential Target for Immunotherapy against Primary Infection with Echinococcus multilocularis Eggs. Infect Immun 10:e00542–e00518
Li GJ, Ding H, Miao D (2019) Long-noncoding RNA HOTAIR inhibits immunologic rejection of mouse leukemia cells through activating the Wnt/β-catenin signaling pathway in a mouse model of leukemia. J Cell Physiol 234:10386–10396
Zhang Y, Li Z, Chen M, Chen H, Zhong Q, Liang L, Li B (2020) lncRNA TCL6 correlates with immune cell infiltration and indicates worse survival in breast cancer. Breast Cancer 27:573–585
Zhou L, Tian Y, Guo F, Yu B, Li J, Xu H, Su Z (2020) LincRNA-p21 knockdown reversed tumor-associated macrophages function by promoting MDM2 to antagonize* p53 activation and alleviate breast cancer development. Cancer Immunol Immunother 69:835–846
Ye Y, Xu Y, Lai Y, He W, Li Y, Wang R (2017) Long non-coding RNA cox-2 prevents immune evasion and metastasis of hepatocellular carcinoma by altering M1/M2 macrophage polarization. J Cell Biochem 119:2951–2963
Jiang R, Tang J, Chen Y, Deng L, Ji J, Xie Y, Jia W, Chu W, Sun WM (2017) B. ells differentiation thus promoting hepatocellular carcinoma immune evasion. Nat Commun; 8
Xia P, Li Q, Wu G, Huang Y (2021) An Immune-Related lncRNA Signature to Predict Survival In Glioma Patients. Cell Mol Neurobiol 41:365–375
Cao R, Yuan L, Ma B, Wang G, Tian Y (2020) Immune-related long non-coding RNA signature identified prognosis and immunotherapeutic efficiency in bladder cancer (BLCA). Cancer Cell Int 20:1–18
Kathuria H, Millien G, McNally L, Gower AC, Tagne JB, Cao Y, Ramirez MI (2018) NKX2-1-AS1 negatively regulates CD274/PD-L1, cell-cell interaction genes, and limits human lung carcinoma cell migration.Sci Rep;8
Zheng Y, Tian X, Wang T, Xia X, Cao F, Tian J, Xu P, Ma J, Xu H, Wang S (2019) Long noncoding RNA Pvt1 regulates the immunosuppression activity of granulocytic myeloid-derived suppressor cells in tumor-bearing mice. Mol Cancer 18:1–12
Bruttel V, Wischhusen J Cancer Stem Cell Immunology: Key to Understanding Tumorigenesis and Tumor Immune Escape?. Frontiers in Immunology.2014;5
López de Andrés J, Griñán-Lisón C, Jiménez G, Marchal JA (2020) Cancer stem cell secretome in the tumor microenvironment: a key point for an effective personalized cancer treatment. J Hematol Oncol 13:136
Eptaminitaki GC, Wolff N, Stellas D, Sifakis K, Baritaki S (2021) Long Non-Coding RNAs (lncRNAs) in Response and Resistance to Cancer Immunosurveillance and Immunotherapy. Cells 10:3313
Malfitano AM, Pisanti S, Napolitano F, Di Somma S, Martinelli R (2020) Portella G.Tumor-associated macrophage status in cancer treatment. Cancers (Basel).; 12:1987
Lei MML, Lee TKW (2021) Cancer Stem Cells: Emerging Key Players in Immune Evasion of Cancers. Front Cell Dev Biol 9:692940
Komura N, Mabuchi S, Shimura K, Yokoi E, Kozasa K, Kuroda H et al (2020) The role of myeloid-derived suppressor cells in increasing cancer stem-like cells and promoting PD-L1 expression in epithelial ovarian cancer. Cancer Immunol Immunother 69:2477–2499
Huergo-Zapico L, Parodi M, Cantoni C, Lavarello C, Fernández-Martínez JL, Petretto A, DeAndrés-Galiana EJ, Balsamo M, López-Soto A, Pietra G, Bugatti M, Munari E, Marconi M, Mingari MC, Vermi W, Moretta L, González S, Vitale M (2018) NK-cell Editing Mediates Epithelial-to-Mesenchymal Transition via Phenotypic and Proteomic Changes in Melanoma Cell Lines. Cancer Res 15:3913–3925
Lopez-Soto A, Huergo-Zapico L, Galvan JA, Rodrigo L, de Herreros AG, Astudillo A et al (2013) Epithelial-mesenchymal transition induces an antitumor immune response mediated by NKG2D receptor. J Immunol 190:4408–4419
García-Venzor A, Mandujano-Tinoco EA, Lizarraga F, Zampedri C, Krötzsch E, Salgado RM, Dávila-Borja VM, Encarnación-Guevara S, Melendez-Zajgla J (2019) Microenvironment-regulated lncRNA-HAL is able to promote stemness in breast cancer cells. Biochim Biophys Acta Mol Cell Res 1866:118523
Sun L, Wang L, Chen T, Shi Y, Yao B, Liu Z, Wang Y, Li Q, Liu R, Niu Y (2020) LncRNA RUNX1-IT1 which is downregulated by hypoxia-driven histone deacetylase 3 represses proliferation and cancer stem-like properties in hepatocellular carcinoma cells. Cell Death &Disease 11:95
Wang X, Sun W, Shen W, Xia M, Chen C, Xiang D et al (2016) Long non-coding RNA DILC regulates liver cancer stem cells via IL-6/STAT3 axis. J Hepatol 64:1283–1294
Yang W, Yu H, Shen Y, Liu Y, Yang Z, Sun T (2016) MiR-146b-5p overexpression attenuates stemness and radioresistance of glioma stem cells by targeting HuR/lincRNA-p21/β-catenin pathway. Oncotarget 7:41505–41526
Ye Y, Guo J, Xiao P, Ning J, Zhang R, Liu P, Yu W, Xu L, Zhao Y, Yu J (2020) Macrophages-induced long noncoding RNA H19 up-regulation triggers and activates the miR-193b/MAPK1 axis and promotes cell aggressiveness in hepatocellular carcinoma. Cancer Lett 469:310–322
Deng J, Yang M, Jiang R, An N, Wang X, Liu B (2017) Long non-coding RNA HOTAIR regulates the proliferation, self-renewal capacity, tumor formation and migration of the cancer stem- like cell (CSC) subpopulation enriched from breast cancer cells.PLoS One;12
Ratti M, Lampis A, Ghidini M, Salati M, Mirchev MB, Valeri N, Hahne JC (2020) MicroRNAs (miRNAs) and Long Non-Coding RNAs (lncRNAs) as New Tools for Cancer Therapy: First Steps from Bench to Bedside. Target Oncol 3:261–278
Pei X, Wang X, Li H (2018) LncRNA SNHG1 regulates the differentiation of Treg cells and affects the immune escape of breast cancer via regulating miR-448/IDO. Int J Biol Macromol 118:24–30
Liang Y, Song X, Li Y, Chen B, Zhao W, Wang L, Zhang H, Liu Y, Han D, Zhang N, Ma T, Wang Y, Ye F, Luo D, Li X, Yang Q (2020) LncRNA BCRT1 promotes breast cancer progression by targeting miR-1303/PTBP3 axis. Mol Cancer 8:85
Yu Z, Zhao H, Feng X, Li H, Qiu C, Yi X, Tang H, Zhang J (2019) Long Non-coding RNA FENDRR Acts as a miR-423-5p Sponge to Suppress the Treg-Mediated Immune Escape of Hepatocellular Carcinoma Cells. Molecular therapy. Nucleic acids 17:516–529
Tian X, Wu Y, Yang Y, Wang J, Niu M, Gao S, Qin T, Bao D (2020) Long noncoding RNA LINC00662 promotes M2 macrophage polarization and hepatocellular carcinoma progression via activating Wnt/β-catenin signaling. Mol Oncol 14:462–483
Zhao L, Liu Y, Zhang J, Liu Y, Qi Q (2019) LncRNA SNHG14/miR-5590-3p/ZEB1 positive feedback loop promoted diffuse large B cell lymphoma progression and immune evasion through regulating PD-1/PD-L1 checkpoint. Cell Death Dis 10:1–15
Zhao CC, Jiao Y, Zhang YY, Ning J, Zhang YR, Xu J, Wei W, Kang-Sheng G (2019) Lnc SMAD5-AS1 as ceRNA inhibit proliferation of diffuse large B cell lymphoma via Wnt/β-catenin pathway by sponging miR-135b-5p to elevate expression of APC. Cell Death Dis 10:1–15
Song B, Guan Z, Liu F, Sun D, Wang K, Qu H (2015) Long non-coding RNA HOTAIR promotes HLA-G expression via inhibiting miR-152 in gastric cancer cells. Biochem Biophys Res Commun 3:807–813
Sun J, Chu H, Ji J, Huo G, Song Q, Zhang X (2016) Long non-coding RNA HOTAIR modulates HLA-G expression by absorbing miR-148a in human cervical cancer. Int J Oncol 49:943–952
Yu Y, Zhang W, Li A, Chen Y, Ou Q, He Z, Zhang Y, Liu R, Yao H, Song E (2020) Association of Long Noncoding RNA Biomarkers With Clinical Immune Subtype and Prediction of Immunotherapy Response in Patients With Cancer. JAMA Netw open 3:e202149
Sun J, Zhang Z, Bao S, Yan C, Hou P, Wu N, Su J, Xu L, Zhou M (2020) Identification of tumor immune infiltration-associated lncRNAs for improving prognosis and immunotherapy response of patients with non-small cell lung cancer. J Immunother Cancer 1:e000110
Kechao N, Zhitong D, Zhihua Z, Yi W, Jinglin P, Xiaotao J (2020) Identification of a 14-lncRNA Signature and Construction of a Prognostic Nomogram Predicting Overall Survival of Gastric Cancer. DNA Cell Biol 3:1532–1544
Shah NN, Fry TJ (2019) Mechanisms of resistance to CAR T cell therapy. Nat Rev Clin Oncol 16:372–385
Zhang Y, Liu Q, Liao Q (2020) Long noncoding RNA: a dazzling dancer in tumor immune microenvironment. J Exp Clin Cancer Res 39:1–25
Sun B, Liu C, Li H, Zhang L, Luo G, Liang S, Lü M (2020) Research progress on the interactions between long non-coding RNAs and microRNAs in human cancer (Review). Oncol Lett 19:595–605
Zhang X, Lian Z, Padden C, Gerstein MB, Rozowsky J, Snyder M, Gingeras TR, Kapranov P, Weissman SM, Newburger PE (2009) A myelopoiesis-associated regulatory intergenic noncoding RNA transcript within the human HOXA cluster. Blood 113:2526–2534
Huang D, Chen J, Yang L, Ouyang Q, Li J, Lao L et al (2018) NKILA lncRNA promotes tumor immune evasion by sensitizing T cells to activation-induced cell death. Nat Immunol 19:1112–1125
Sang L, Ju H, Liu G, Tian T, Ma G, Lu Y (2018) LncRNA CamK-A Regulates Ca2+-Signaling-Mediated Tumor Microenvironment Remodeling. Mol Cell 72:601
Huang J, Ma L, Song W, Lu B, Huang Y, Dong H, Ma X, Zhu Z, Zhou R (2017) LncRNA-MALAT1 Promotes Angiogenesis of Thyroid Cancer by Modulating Tumor‐Associated Macrophage FGF2 Protein Secretion. J Cell Biochem 12:4821–4830
Wu K, Zhao Z, Liu K, Zhang J, Li G, Wang L (2017) Long noncoding RNA lnc-sox5 modulates CRC tumorigenesis by unbalancing tumor microenvironment. Cell Cycle 16:1295–1301
Liang Z, Liu H, Wang F, Xiong L, Zhou C, Hu T et al (2019) LncRNA RPPH1 promotes colorectal cancer metastasis by interacting with TUBB3 and by promoting exosomes-mediated macrophage M2 polarization. Cell Death Dis 10:1–17
Ji J, Yin Y, Ju H, Xu X, Liu W, Fu Q, Hu J, Zhang X, Sun B (2018) Long non-coding RNA Lnc-Tim3 exacerbates CD8 T cell exhaustion via binding to Tim-3 and inducing nuclear translocation of Bat3 in HCC. Cell Death Dis 9:1–11
Xie C, Guo Y, Lou S (2020) LncRNA ANCR Promotes Invasion and Migration of Gastric Cancer by Regulating FoxO1 Expression to Inhibit Macrophage M1 Polarization. Dig Dis Sci 65:2863–2872
Xiong G, Yang L, Chen Y, Fan Z (2015) Linc-POU3F3 promotes cell proliferation in gastric cancer via increasing T-reg distribution. Am J Transl Res 7:2262–2269
Gao Y, Wang T, Li Y, Zhang Y, Yang R (2018) Lnc-chop Promotes Immunosuppressive Function of Myeloid-Derived Suppressor Cells in Tumor and Inflammatory Environments. J Immunol 200:2603–2614
Tian X, Ma J, Wang T, Tian J, Zheng Y, Peng R et al (2018) Long non-coding RNA RUNXOR accelerates MDSC-mediated immunosuppression in lung cancer. BMC Cancer 18:1–10
Ma F, Lei YY, Ding MG, Luo LH, Xie YC, Liu XL (2020) LncRNA NEAT1 Interacted With DNMT1 to Regulate Malignant Phenotype of Cancer Cell and Cytotoxic T Cell Infiltration via Epigenetic Inhibition of p53, cGAS, and STING in Lung Cancer.Front Genet;11
Sun Y, Xu J (2019) TCF-4 regulated lncrna-xist promotes m2 polarization of macrophages and is associated with lung cancer. Onco Targets Ther 12:8055–8062
Shang A, Wang W, Gu C, Chen C, Zeng B, Yang Y et al (2019) Long non-coding RNA HOTTIP enhances IL-6 expression to potentiate immune escape of ovarian cancer cells by upregulating the expression of PD-L1 in neutrophils.J Exp Clin Cancer Res;38
Lin A, Li C, Xing Z (2016) The LINK-A lncRNA activates normoxic HIF1α signalling in triple-negative breast cancer. Nat Cell Biol 18:213–224
Gao Y, Sun W, Shang W, Li Y, Zhang D, Wang T, Zhang X, Zhang S, Zhang Y, Yang R (2018) Lnc-C/EBPβ negatively regulates the suppressive function of myeloid-derived suppressor cells. Cancer Immunol Res 6:1352–1363
Zhou Q, Tang X, Tian X, Tian J, Zhang Y, Ma J, Xu H, Wang S (2018) LncRNA MALAT1 negatively regulates MDSCs in patients with lung cancer. J Cancer 9:2436–2442
Charpentier M, Croyal M, Carbonnelle D, Fortun A, Florenceau L, Rabu C, Krempf M, Labarrière N, Lang F (2016) IRES-dependent translation of the long non-coding RNA meloe in melanoma cells produces the most immunogenic MELOE antigens. Oncotarget 7:59704–59713
Wang QM, Lian GY, Song Y, Huang YF, Gong Y (2019) LncRNA MALAT1 promotes tumorigenesis and immune escape of diffuse large B cell lymphoma by sponging miR-195.Life Sci;231
Qian CS, Li LJ, Huang HW, Yang HF, Wu DP (2020) MYC-regulated lncRNA NEAT1 promotes B cell proliferation and lymphomagenesis via the miR-34b-5p-GLI1 pathway in diffuse large B-cell lymphoma. Cancer Cell Int 20:1–13
Ni C, Fang QQ, Chen WZ, Jiang JX, Jiang Z, Ye J et al (2020) Breast cancer-derived exosomes transmit lncRNA SNHG16 to induce CD73 + γδ1 Treg cells. Signal Transduct Target Ther 5:1–14
Liu SQ, Zhou ZY, Dong X, Guo L, Zhang KJ (2020) LncRNA GNAS-AS1 facilitates ER + breast cancer cells progression by promoting M2 macrophage polarization via regulating miR-433-3p/GATA3 axis.Biosci Rep;40
Yin Z, Zhou Y, Ma T, Chen S, Shi N, Zou Y, Hou B, Zhang C (2020) Down-regulated lncRNA SBF2-AS1 in M2 macrophage-derived exosomes elevates miR-122-5p to restrict XIAP, thereby limiting pancreatic cancer development. J Cell Mol Med 24:5028–5038
Zhou WY, Zhang MM, Liu C, Kang Y, Wang JO, Yang XH (2019) Long noncoding RNA LINC00473 drives the progression of pancreatic cancer via upregulating programmed death-ligand 1 by sponging microRNA-195-5p. J Cell Physiol 234:23176–23189
Hou ZH, Xu XW, Fu XY, Zhou L, Du, Liu SP, Tan DM (2020) Long non-coding RNA MALAT1 promotes angiogenesis and immunosuppressive properties of HCC cells by sponging miR-140. Am J Physiol - Cell Physiol 318:649–663
Li Z, Feng C, Guo J, Hu X, Xie D (2019) GNAS-AS1/miR-4319/NECAB3 axis promotes migration and invasion of non-small cell lung cancer cells by altering macrophage polarization. Funct Integr Genom 1:17–28
Shang W, Gao Y, Tang Z, Zhang Y, Yang R (2019) The pseudogene OLFR29-PS1 promotes the suppressive function and differentiation of monocytic MDSCs. Cancer Immunol Res 7:813–827
Zhang Y, Feng J, Fu H, Liu C, Yu Z, Sun Y et al (2020) Corrigendum: Coagulation Factor X Regulated by CASC2c Recruited Macrophages and Induced M2 Polarization in Glioblastoma Multiforme (Frontiers in Immunology. Front Immunol 11:934
Xing F, Liu Y, Wu SY, Wu K, Sharma S, Mo YY et al (2018) Loss of XIST in breast cancer activates MSN-c-Met and reprograms microglia via exosomal miRNA to promote brain metastasis. Cancer Res 78:4316–4330
Wei M, CU Z, Zheng L, zhao M, Wang X (2020) Long non-coding RNA GAS5 promotes natural killer cell cytotoxicity against gastric cancer by regulating miR-18a. Neoplasma 67:1085–1093
Zhou Y, Zhao W, Mao L, wei, Wang Y, li, Xia L, Cao M, Shen J, Chen J (2018) Long non-coding RNA NIFK-AS1 inhibits M2 polarization of macrophages in endometrial cancer through targeting miR-146a. Int J Biochem Cell Biol 104:25–33
Funding
This work is supported by the internal Zewail City of Science and Technology fund ZC 016-2019 and the open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
All authors contributed to the paper’s information gathering, conception and design. The data collection, data contribution or analysis tools, analyzation, interpretation, conceived, and designed the study, wrote the paper by Dina Mofeed. The data analyzation, interpretation, designed the graphs, and paper wrote by Jihad I. Omran. The study supervision, conceived, designed, and data interpretation by Salwa Sabet. Study supervised, conceived, designed, and data interpreted by Ahmed A. Baiomy. Study supervised, revised, and final version approval of the manuscript by Marwan Emara. The study supervised, conceived, designed, data interpretation, revised, and final version approval of the manuscript by Tamer Z. Salem.
Corresponding author
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mofed, D., Omran, J.I., Sabet, S. et al. The regulatory role of long non- coding RNAs as a novel controller of immune response against cancer cells. Mol Biol Rep 49, 11775–11793 (2022). https://doi.org/10.1007/s11033-022-07947-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07947-4