Abstract
Introduction
The unbound brain extracelullar fluid (brainECF) to plasma steady state partition coefficient, Kp,uu,BBB, values provide steady-state information on the extent of blood-brain barrier (BBB) transport equilibration, but not on pharmacokinetic (PK) profiles seen by the brain targets. Mouse models are frequently used to study brain PK, but this information cannot directly be used to inform on human brain PK, given the different CNS physiology of mouse and human. Physiologically based PK (PBPK) models are useful to translate PK information across species.
Aim
Use the LeiCNS-PK3.0 PBPK model, to predict brain extracellular fluid PK in mice.
Methods
Information on mouse brain physiology was collected from literature. All available connected data on unbound plasma, brainECF PK of 10 drugs (cyclophosphamide, quinidine, erlotonib, phenobarbital, colchicine, ribociclib, topotecan, cefradroxil, prexasertib, and methotrexate) from different mouse strains were used. Dosing regimen dependent plasma PK was modelled, and Kpuu,BBB values were estimated, and provided as input into the LeiCNS-PK3.0 model to result in prediction of PK profiles in brainECF.
Results
Overall, the model gave an adequate prediction of the brainECF PK profile for 7 out of the 10 drugs. For 7 drugs, the predicted versus observed brainECF data was within two-fold error limit and the other 2 drugs were within five-fold error limit.
Conclusion
The current version of the mouse LeiCNS-PK3.0 model seems to reasonably predict available information on brainECF from healthy mice for most drugs. This brings the translation between mouse and human brain PK one step further.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mouse studies recapitulating central nervous system (CNS) diseases have long been used to study human diseases and drug treatment [1, 2], including those related to the CNS [3]. However, such information is not directly translatable to the human situation. Therefore, it is important to seek for translational approaches.
As unbound drug concentration-time profiles (PK) at CNS target sites drive the CNS effect [4, 5] these are most important. Microdialysis is the most adequate technique to assess the unbound drug concentration-time profiles in the brain extracellular fluid (brainECF) and in the different cerebrospinal fluid (CSF) compartments in preclinical species [6,7,8,9,10,11,12], but assessment of unbound brain PK in human by microdialysis is highly restricted for ethical reasons. The question is therefore on how to bridge the translational gap between preclinical information to be used in drug development and the clinical setting.
In preclinical species, the unbound brainECF to plasma steady state partition coefficient, the Kp,uu,BBB, is a very important value and is often obtained to provide information on the extent of blood-brain barrier (BBB) transport equilibration but is does not provide information on the brain PK profiles, as seen by the brain target sites. It is a ratio, and does not inform on PK profiles, as seen by receptors and other targets.
As PK profiles are driven by the combination of drug properties and the body (system) physiology, the physiological differences between species prevents a direct translation. However, physiologically based pharmacokinetic (PBPK) models explicitly take systems physiology into account, and when combined with drug properties allows prediction of PK profiles, and therefore should be able to bridge the findings between different species, such as rat or mouse and human.
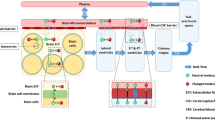
Earlier, we developed a comprehensive CNS PBPK model in rat and human, the LeiCNS-PK3.0 [13, 14]. This model includes the BBB and blood-CSF barrier (BCSFB) characteristics and surfaces, the brainECF, brain intracellular fluid (brainICF), cerebrospinal fluid (CSF) in lateral ventricles, third and fourth ventricles, cisterna magna, and subarachnoid space, and their volumes and flows, and the brain cell membranes surfaces, brain cell volumes, lysosome volumes, and pH values in all the compartments. With that it allows prediction of a drug’s blood to brain and intra-brain transport processes, including nonspecific binding, when the drug properties were used as input together with the plasma PK after the mode of drug administration of choice. The predicted observed unbound drug PK in brainECF and different CSF compartments in rats and humans were within less than two-fold error, demonstrating rat-to-human translatability of CNS PK profiles [13].
A LeiCNS-PK3.0 mouse version could help to translate mouse CNS PK to that of human, and thereby bridge a lot of mouse data for human interpretation. In this study, we searched for healthy mouse CNS physiological parameters from literature, and connected unbound plasma PK, and associated brainECF PK, as obtained by microdialysis. Such data was available for 10 drugs, from different mouse strains/types, with different physicochemical properties. Using the plasma data, plasma PK models were developed, to inform the LeiCNS-PK3.0 model, together with the detailed mouse CNS physiological parameters and calculation of Kp,uu,BBB. Here we report and discuss the results on the performance of this model to predict brainECF data in mice.
Data and Methods
The LeiCNS-PK3.0 model structure [13] was informed on mouse CNS physiological information and mouse drug unbound plasma and associated brainECF PK as far as available from literature. For 10 drugs such PK information was available, and plasma PK model were available for 3 drugs and for 7 drugs a plasma PK model was developed. Furthermore, the physicochemical and biological properties of these drugs were obtained/ calculated. All is explained below.
Drugs
The following 10 drugs were used to evaluate the predictive performance of the mouse version of the LeiCNS-PK3.0 model: cyclophosphamide, quinidine, erlotonib, phenobarbital, colchicine, ribociclib, topotecan, cefradroxil, prexasertib, and methotrexate.
Drug physicochemical properties
The physicochemical properties of the drugs were extracted from DrugBank release version 5.1.9 [15] and are presented in Table I. Lipophilicity (as logP) was estimated using the ALOGPS [16], while the acid/base ionization constants, polar surface area, and hydrogen bond donor/acceptor values were provided by the Chemaxon method [17].
In Vivo Data
Plasma and associated brainECF concentration-time profiles (Table II) of part of the drugs were kindly provided by the Stewart lab (St. Jude Children's Research Hospital, Memphis, Tennessee, USA) or otherwise extracted from literature with WebPlotDigitizer version 4.5 (https://automeris.io/WebPlotDigitizer/).
Mouse CNS Physiological Parameters
Mouse parameter values for CNS physiology were obtained from literature. Values for BBB and blood-CSF barrier (BCSFB) characteristics and surfaces; brainECF, brainICF, CSF in lateral ventricles, third and fourth ventricles, cisterna magma and subarachnoid space, volumes and flows; brain cell membranes surfaces, brain cell volumes, lysosome volumes; and pH values in all the compartments were collected. In case multiple values were found, the mean value was calculated. The surface area of the BBB was calculated with two approaches and the mean value was computed. The first, using the microvessels average radius, length density, and brain volume, while the second using the brain vessel surface area to brain volume ratio and total brain volume.
Plasma PK Modeling
The plasma PK models of cyclophosphamide, ribociclib, and prexasertib were available from literature [7, 9, 22]. The plasma PK parameters of cefadroxil were estimated using Monolix version 2021R2 (Lixoft, Orsay, France). Plasma PK model parameters of the other 6 drugs were estimated using NONMEM version 7.4.3 (ICON, Dublin, Ireland) [23]. Population plasma PK models were developed and used as input to the CNS PBPK model. In brief, one-, two-, three- compartment models were fitted to total plasma concentrations, accounting for the associated interindividual variabilities (where possible) using an exponential function, and for the residual unexplained variability using proportional or combined proportional and additive error models. The final model was selected based on likelihood ratio test with p<0.05 corresponding to an objective function value decline of 3.84, visual predictive check (VPC) plots, precision of the parameter estimates (%RSE), and the basic goodness of fit plots.
Drug Biological Properties–Calculation of Kp,uu,BBB Values
Kp,uu,BBB values, defined as the ratio of the unbound drug in brainECF to that of plasma at steady state, reflect the extent of drug transport across a barrier (i.e. BBB or BCSFB). These values may differ from 1 due to transporters at these barriers [24]. Kp,uu,BBB values were calculated by the ratio of influx and efflux clearances across the BBB, respectively, or by the ratio of the AUC 0-∞ at the brainECF to that of plasma, respectively [24]. Where unavailable, the influx and efflux clearances of the unbound drug across the BBB were estimated by combining the respective population plasma PK model and a one-compartment model representing the whole brain. Then, these Kp,uu,BBB values were used to calculate the asymmetry factors at the BBB that reflect the net active transport across these barriers in the LeiCNS-PK3.0 model.
Mouse LeiCNS-PK3.0 Model Evaluation and Data Analysis
As indicated, the LeiCNS-PK 3.0 mouse model was developed using the previously published model structure of the rat and human LeiCNS-PK3.0 model versions. The mouse CNS physiological parameters were given as input, together with the Kp,uu,BBB values, and the plasma PK parameters. The LeiCNS-PK3.0 mouse model predictions of brainECF were evaluated by comparison with the observed CNS PK data, using visual predictive checks (VPC). In addition, using the prediction errors, the percentage average fold error (%AFE) and percentage absolute average fold error (%AAFE) were computed as described previously [13] and were used to evaluate the bias and the accuracy of the model predictions, respectively. Data analysis and visualization were performed in R (version 4.1.2) [25]. The LeiCNS-PK3.0 model simulations were also performed in R, using the package RxODE (version 1.1.4) and the LSODA (Livermore Solver for Ordinary Differential Equations) Fortran package [26].
Results
Mouse CNS Physiological Parameters
Mouse parameter values for CNS physiology were obtained from literature. When more values of a certain parameter were found, the mean value was used in the LeiCNSPK3.0 mouse model. Table III lists all parameter values and relevant assumptions.
Plasma PK Modelling
For a given dose regimen, for the LeiCNS-PK3.0 model the associated plasma PK model parameters were used as input and forcing function as an input and forcing function to reduce errors due to the potential imprecise plasma PK predictions of a whole body PBPK model. The plasma PK models parameters are reported in Table IV and the model prediction of brainECF data against the observed brainECF data and associated errors are depicted in Figs. 1 and 2, respectively. Generally, the models were estimated within two-fold error and the in vivo plasma PK data were accurately described.
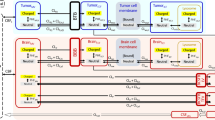
Visual predictive check plots evaluating the predictive accuracy of the mouse version of the LeiCNS-PK3.0 model. Ten drugs with different physicochemical properties and affinities to active transporters were used to evaluate the model predictions. The solid lines and colored band represent the median and 95% prediction interval, respectively, of the model’s prediction of the unbound pharmacokinetic profile at the plasma (red), brain extracellular fluid (yellow). The black dots represent the unbound drug concentrations measured in mice. Drugs were simulated with various routes of administrations: cyclophosphamide, quinidine and phenobarbital were intraperitoneal; erlotinib and ribociclib were orally; prexasertib was subcutaneous; colchicine, topotecan, cefadroxil and methotrexate were intravenously administered. Please note the different axes scales
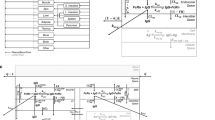
Box plot of the relative accuracy errors to evaluate the prediction accuracy of the current mouse version of the LeiCNS-PK3.0 model. The predictions of the ten drugs in plasma (red) and brain extracellular fluid (yellow) were evaluated using the relative accuracy errors. The green and yellow vertical lines represent two- and five-fold error limit, respectively. The predictions of methotrexate and prexasertib were beyond the two-fold errors but were within the five-fold error
Drug Biological Properties – Calculation Of Kp,uu,BBB Values
As input for the LeiCNS-PK3.0 model values of the asymmetry factors, the Kp,uu,BBB values for the different drugs were calculated as clearance in over clearance out of the brainECF. Results are shown in Table V.
Mouse LeiCNS-PK3.0 Model Evaluation And Data Analysis
Model validation was performed by comparing the data-independent LeiCNS-PK3.0 model predictions at brainECF to drug concentrations measured in vivo with microdialysis (Fig. 1). Overall, the LeiCNS-PK3.0 model predictions were good for 7 out of 10 drugs. For phenobarbital, the prediction of brainECF was slower (lower Cmax, slower elimination rate) than the actual data. For prexasertib, the model predictions of brainECF were faster (higher and earlier Cmax, higher elimination rate), while for methotrexate the model prediction of underestimated the elimination phase of brainECF data.
The LeiCNS-PK3.0 model bias was assessed using relative accuracy errors (%AFE), which was 99.6% and 76.5%, for plasma and brainECF, respectively. The model’s predictivity of the typical CNS PK profile was evaluated using the %AAFE, which was 105 % and 152 %, for plasma and brainECF, respectively. Figure 2 displays the visual predictive checks. It shows that the LeiCNS-PK3.0 mouse model could adequately predict the brainECF drug concentrations, within the two-fold error limit for 8 out of the 10 drugs.
Discussion
Translation between mouse and human CNS PK data would be an important step forward in CNS drug development, as animal data can be used in a better way. Lots of total plasma and total brain concentrations in mice are available, however, the link to the human situation cannot be directly made. The Kp,uu,BBB (or Kp,uu,ECF) value can be obtained using steady-state ratios of unbound brain over unbound plasma concentrations, but this ratio can have the same value for multiple combinations of plasma and brainECF PK. Brain targets, mostly extracellular, however, see the PK profiles, and therefore we need insights into the relationship between plasma PK and brainECF PK profiles. In this study we explored the use of the LeiCNS-PK3.0 model to predict brainECF PK profiles, based on unbound plasma PK profiles. If such a model would be adequate, it may be used to be further extended to other CNS compartments, and ultimately may also use mouse Kp,uu,brain values to predict brainECF and other CNS location PK profiles for translation to the human situation.
In this study, we validated the LeiCNS-PK3.0 model for its use to predict mouse brainECF data. Earlier versions of the LeiCNS-PK3.0 model have shown to adequately predict rat and human CNS unbound PK profiles in multiple CNS physiological compartments [13]. Here we used all available data on unbound plasma PK and associated brainECF PK profiles, as well as literature information on details of the mouse CNS physiology, to explore the ability of this mouse version of the LeiCNS-PK3.0 model to predict brainECF data.
Many published studies have reported the development of whole-body mouse PBPK models, accounting also for the brain [10, 32, 58,59,60,61,62]. These models were used to predict mouse PK profiles in multiple organs and to translate the PK profiles to humans. However, these models do not distinguish the brain cells, brainICF and brainECF [32, 58], while also do not account for the presence of lysosomes and non-specific binding [10, 32, 58]. Explicit distinction between all CNS physiological compartments, particularly the main target sites: brainECF and brainICF, is very relevant for more accurate assessment of the concentration-effect relationship [63]. Our LeiCNS-PK3.0 model inputs are physiological parameters, drug physicochemical properties, and Kp,uu values, which can be obtained from in vivo, in vitro [64], or in silico [65] studies. None of the model parameters was estimated and, therefore, the model is translatable to other species, including humans, and to predict the CNS PK of small molecule drugs.
The current mouse LeiCNS-PK3.0 model is the first mouse CNS PBPK model of small molecule drugs, to the best of the authors’ knowledge, that accounts mechanistically for the mouse CNS physiology, including the different compartments and drug transport modes, bulk fluid flow, pH differences, and non-specific binding. The first step was to see if the model could adequately predict brainECF, of available mouse data sets with associated plasma PK profiles, with fu,plasma information, and brainECF. 10 drugs were found (cyclophosphamide, quinidine, erlotonib, phenobarbital, colchicine, ribociclib, topotecan, cefradroxil, prexasertib, and methotrexate), with data from different mouse strains/types. Overall, the LeiCNS-PK3.0 model brainECF predictions were good for 7 out of 10 drugs. For phenobarbital, the prediction of brainECF was slower (lower Cmax, slower elimination rate) than the actual data. For prexasertib, the model predictions of brainECF were faster (higher and earlier Cmax, higher elimination rate), while for methotrexate the model prediction might overestimated the Cmax and underestimated Tmax (not enough early time data to know), while it underestimated the elimination phase of brainECF data. This could not be due to the plasma PK input, as all models rather precisely described the plasma PK profiles. Analytical assays might also be a source of some deviation, but not detailed enough information on the analytical assays for high and low concentration CV% were provided to assess this possibility. Potential differences between the CNS physiology of the mouse types/strains could contribute, but the number of drugs studied with the unbound plasma and brainECF data is too little to further analyze such a possibility. More data should be produced to further evaluate the mouse LeiCNS-PK3.0 model, while its performance is already quite a step forward.
Another aspect could be the mouse BBB surface area (SA). It is reported in literature to be 240 cm2/g brain equivalent to 86.4 cm2 for a 360-μg mouse brain [66]. This value when used in the LeiCNS-PK3.0 model resulted in poor prediction of brainECF PK profile (results not shown). In comparison, BBB SA in rats and humans were 82 and 120 cm2/g brain, respectively [13], implying that 240 cm2/g brain could be an overestimation of mouse BBB surface area. Hence, we calculated a mean mouse BBB SA of 19.8 cm2 using two techniques: a value of 18.8 cm2 using the surface area per unit volume of different brain regions [42], weighted by the regional volume [35] and corrected for the total brain volume and another value of 20.7 cm2 using the average microvessels diameter and length density, corrected for total brain volume [41, 42]. The new value resulted in better prediction of brainECF PK profiles of cyclophosphamide and topotecan, while that of other drugs in our dataset remained the same. This approach is what we call the “handshake approach” [67], as in our opinion, we can especially learn back from in vivo data, and therefore (CNS) PBPK models should not only be informed by in vitro or in silico information, to improve physiological parameter values in the PBPK models.
Besides methodological and/or physiological aspects, drugs physicochemical properties could play a role in passive BBB transport. We considered the polar surface area (PSA) to play a role [68], being relatively high for methotrexate and prexasertib (206 and 135 Å2, respectively). However, cefadroxil also has a high PSA value (133 Å2), but could be predicted within two-fold error, while that of phenobarbital is much lower (75 Å2) and not within two-fold error. Another consideration was to compare the number of hydrogen bond acceptors (HBA) and/or donors (HBD). For methotrexate HBA/HBD was 12/6, and for prexasertib it was 8/3. However, for phenobarbital this was 3/2. So, no clear pattern for HBA and HBD neither. Then, the comparison of the mouse versus the rat CNS LeiCNS-PK3.0 model performance could only be done for methotrexate, as the only drug for which appropriate data was observed in mouse and rat. In the rat model, methotrexate brainECF data were within 251 %AAFE versus 433 %AAFE for that in mice. Altogether, this indicates the need for more in-depth analysis of the combination of multiple physicochemical properties, as well as exploring potential physiological aspects that may vary between different mouse strains and/or methodologies used to measure the physiology. Although our goal is to reuse as much animal data as possible, and to save animal lives, It might even be necessary to have additional microdialysis data on CNS drug distribution in mice produced, in which also other CNS locations and end-of-experiment total brain concentrations can be obtained in conjunction (connected data, [69].
This model will be further improved in depth analysis of the influence of drug physicochemical properties. Furthermore, using the “handshake” approach [67], the impact of physiological values used in the model will be studied, and improved, assuming that in vivo data “tell the truth”. Next steps will be to make use of ex-vivo plasma, and plasma binding, as well as brain homogenate and brain binding, to calculate Kp,uu,brain values [70], by which the model can predict full pharmacokinetic profiles in the different compartments”.
Altogether, the current mouse LeiCNS-PK3.0 model shows adequate predictions of observed brainECF data for 7 out of the 10 drugs for which the unbound plasma PK and associated brainECF data were available. While some deviating predictions were also observed, the mouse LeiCNS-PK3.0 holds promise for further development to be useful as a translational tool to predict the healthy, and ultimate diseased human CNS PK profiles, also from using PK data obtained from mice.
Data Availability
Data sharing is not applicable to this article as no new data was generated in this study.
References
Garattini S, Grignaschi G. Animal testing is still the best way to find new treatments for patients. Eur J Intern Med. 2017;39:32–5. https://doi.org/10.1016/j.ejim.2016.11.013.
Murillo-Cuesta S, Artuch R, Asensio F, de la Villa P, Dierssen M, Enríquez JA, Fillat C, Fourcade S, Ibáñez B, Montoliu L, Oliver E, Pujol A, Salido E, Vallejo M, Varela-Nieto I. The value of mouse models of rare diseases: A spanish experience. Front Genet. 2020; 11: 583932. https://doi.org/10.3389/fgene.2020.583932.
Hall AM, Roberson ED. Mouse models of Alzheimer’s disease. Brain Res Bull. 2012;88:3–12. https://doi.org/10.1016/j.brainresbull.2011.11.017.
de Lange ECM, Hesselink MB, Danhof M, de Boer AG, Breimer DD. The use of intracerebral microdialysis to determine changes in blood-brain barrier transport characteristics. Pharm Res. 1995;12:129–33. https://doi.org/10.1023/a:1016207208406.
Summerfield SG, Yates JWT, Fairman DA. Free drug theory – no longer just a hypothesis? Pharm Res. 2022;39:213–22. https://doi.org/10.1007/s11095-022-03172-7.
de Lange ECM, Hesselink MB, Danhof M, de Boer AG, Breimer DD. The use of intracerebral microdialysis to determine changes in blood-brain barrier transport characteristics 1995b.
Campagne O, Davis A, Zhong B, Nair S, Haberman V, Patel YT, Janke L, Roussel MF, Stewart CF. Cns penetration of cyclophosphamide and metabolites in mice bearing group 3 medulloblastoma and non-tumor bearing mice. J Pharm Pharm Sci. 2019;22:612–29. https://doi.org/10.18433/JPPS30608.
Hammarlund-Udenaes M. The use of microdialysis in CNS drug delivery studies: Pharmacokinetic perspectives and results with analgesics and antiepileptics. Adv Drug Deliv Rev. 2000;45:283–94. https://doi.org/10.1016/S0169-409X(00)00109-5.
Patel YT, Davis A, Baker SJ, Campagne O, Stewart CF. CNS penetration of the CDK4/6 inhibitor ribociclib in non-tumor bearing mice and mice bearing pediatric brain tumors. Cancer Chemother Pharmacol. 2019;84:447–52. https://doi.org/10.1007/s00280-019-03864-9.
Perkins RS, Davis A, Campagne O, Owens TS, Stewart CF. CNS penetration of methotrexate and its metabolite 7-hydroxymethotrexate in mice bearing orthotopic Group 3 medulloblastoma tumors and model-based simulations for children. Drug Metab Pharmacokinet 2022; 100471. https://doi.org/10.1016/j.dmpk.2022.100471.
Westerhout J, Van Den Berg DJ, Hartman R, Danhof M, De Lange ECM. Prediction of methotrexate CNS distribution in different species - Influence of disease conditions. Eur J Pharm Sci. 2014;57:11–24. https://doi.org/10.1016/j.ejps.2013.12.020.
Yamamoto Y, Välitalo PA, van den Berg DJ, Hartman R, van den Brink W, Wong YC, Huntjens DR, Proost JH, Vermeulen A, Krauwinkel W, Bakshi S, Aranzana-Climent V, Marchand S, Dahyot-Fizelier C, Couet W, Danhof M, van Hasselt JGC, de Lange ECM. A Generic Multi-Compartmental CNS Distribution Model Structure for 9 Drugs Allows Prediction of Human Brain Target Site Concentrations. Pharm Res. 2017;34:333–51. https://doi.org/10.1007/s11095-016-2065-3.
Saleh MAA, Loo CF, Elassaiss-Schaap J, De Lange ECM. Lumbar cerebrospinal fluid-to-brain extracellular fluid surrogacy is context-specific: insights from LeiCNS-PK3.0 simulations. J Pharmacokinet Pharmacodyn. 2021;48:725–41. https://doi.org/10.1007/s10928-021-09768-7.
Yamamoto Y, Välitalo PA, Wong YC, Huntjens DR, Proost JH, Vermeulen A, Krauwinkel W, Beukers MW, Kokki H, Kokki M, Danhof M, van Hasselt JGC, de Lange ECM. Prediction of human CNS pharmacokinetics using a physiologically-based pharmacokinetic modeling approach. Eur J Pharm Sci. 2018;112:168–79. https://doi.org/10.1016/j.ejps.2017.11.011.
Wishart DS, Feunang YD, Guo AC, Lo EJ, Marcu A, Grant JR, Sajed T, Johnson D, Li C, Sayeeda Z, Iynkkaran NAI, Liu Y, Maciejewski A, Gale N, Wilson A, Chin L, Cummings R, Le D, Pon A, Knox C, Wilson M. DrugBank 5.0: a major update to the DrugBank database for 2018. Nucl Acids Res. 2017;46:D1074–82.
Mannhold R, Poda GI, Ostermann C, Tetko IV. Calculation of molecular lipophilicity: state-of-the-art and comparison of logp methods on more than 96,000 compounds. J Pharm Sci. 2009;98:861–93. https://doi.org/10.1002/jps.
Manchester J, Walkup G, Rivin O, You Z. Evaluation of pka estimation methods on 211 druglike compounds. J Chem Inf Model. 2010;50:565–71. https://doi.org/10.1021/ci100019p.
Sziráki I, Erdo F, Trampus P, Sike M, Molnár PM, Rajnai Z, Molnár J, Wilhelm I, Fazakas C, Kis E, Krizbai I, Krajcsi P. The use of microdialysis techniques in mice to study P-gp function at the blood-brain barrier. J Biomol Screen. 2013;18:430–40.
Jones DR, Hall SD, Jackson EK, Branch RA, Wilkinson GR. Brain uptake of benzodiazepines: Effects of lipophilicity and plasma protein binding. J Pharmacol Exp Ther. 1988;245:816–22.
Pierre A. Evrard, Corinne Ragusi, Gabrielle Boschi, Roger K. Verbeeck, Jean-Michel Scherrmann (1998) Simultaneous microdialysis in brain and blood of the mouse: extracellular and intracellular brain colchicine disposition. Brain Res. 786(1-2):122–7. https://doi.org/10.1016/S0006-8993(97)01454-6.
Chen X, Keep RF, Liang Y, Zhu H-J, Hammarlund-Udenaes M, Hu Y, Smith DE. Influence of peptide transporter 2 (PEPT2) on the distribution of cefadroxil in mouse brain: A microdialysis study. Biochem Pharmacol. 2017:13189–97. https://doi.org/10.1016/j.bcp.2017.02.005.
Campagne O, Davis A, Maharaj AR, Zhong B, Stripay J, Farmer D, Roussel MF, Stewart CF. CNS penetration and pharmacodynamics of the CHK1 inhibitor prexasertib in a mouse Group 3 medulloblastoma model. Eur J Pharm Sci. 2020;142:105106. https://doi.org/10.1016/j.ejps.2019.105106.
Bauer RJ NONMEM Users guide: Introduction to NONMEM 7.4.3, In: ICON Plc 2019. https://doi.org/10.1017/CBO9781107415324.004.
Hammarlund-Udenaes M, Fridén M, Syvänen S, Gupta A. On the rate and extent of drug delivery to the brain. Pharm Res. 2008;25:1737–50. https://doi.org/10.1007/s11095-007-9502-2.
R Core Team. R: A language and environment for statistical computing. Vienna: R Foundation for Statistical Computing; 2019.
Fidler M, Hallow M, Wilkins J, Wang W. RxODE: Facilities for simulating from ODE-based models 2019.
Kovačević N, Henderson JT, Chan E, Lifshitz N, Bishop J, Evans AC, Henkelman RM, Chen XJ. A threedimensional mri atlas of the mouse brain with estimates of the average and variability. Cereb Cortex. 2004;15(5):639–45. https://doi.org/10.1093/cercor/bhh165.
Singh Badhan RK, Chenel M, Penny JI. Development of a physiologically-based pharmacokinetic model of the rat central nervous system. Pharmaceutics. 2014;6:97–136. https://doi.org/10.3390/pharmaceutics6010097.
Fenneteau F, Turgeon J, Couture L, Michaud V, Li J, Nekka F. Assessing drug distribution in tissues expressing P-glycoprotein through physiologically based pharmacokinetic modeling: model structure and parameters determination Abstract. Theor Biol Medical Model. 2009;6(1). https://doi.org/10.1186/1742-4682-6-2.
Mizoguchi T, Minakuchi H, Ishisaka M, Tsuruma K, Shimazawa M, Hara H. Behavioral abnormalities with disruption of brain structure in mice overexpressing VGF Abstract. Sci Rep. 2017;7(1). https://doi.org/10.1038/s41598-017-04132-7.
Baker KL, Daniels SB, Lennington JB, Lardaro T, Czap A, Notti RQ, Cooper O, Isacson O, Frasca S, Conover JC. Neuroblast protuberances in the subventricular zone of the regenerative MRL/MpJ mouse. J Comp Neurol. 2006;498(6):747–61. https://doi.org/10.1002/cne.21090.
Hu ZY, Lu J, Zhao Y. A physiologically based pharmacokinetic model of alvespimycin in mice and extrapolation to rats and humans. Br J Pharmacol. 2014;171:2778–89. https://doi.org/10.1111/bph.12609.
Brown RP, Delp MD, Lindstedt SL, Rhomberg LR, Beliles RP. Physiological parameter values for physiologically based pharmacokinetic models. Toxicol Ind Health. 1997;13(4):407–84. https://doi.org/10.1177/074823379701300401.
Pardridge WM. CSF blood-brain barrier and brain drug delivery. Expert Opin Drug Deliv. 2016;13(7):963–75. https://doi.org/10.1517/17425247.2016.1171315.
Badea A, Ali-Sharief AA, Johnson GA. Morphometric analysis of the C57BL/6J mouse brain. Neuroimage. 2007;37:683–93. https://doi.org/10.1016/j.neuroimage.2007.05.046.
Dorr AE, Lerch JP, Spring S, Kabani N, Henkelman RM. High resolution three-dimensional brain atlas using an average magnetic resonance image of 40 adult C57Bl/6J mice. NeuroImage. 2008;42(1):60–9. https://doi.org/10.1016/j.neuroimage.2008.03.037.
Hino K, Otsuka S, Ichii O, Hashimoto Y, Kon Y. Strain differences of cerebral ventricles in mice: Can the MRL/MpJ mouse be a model for hydrocephalus? Jpn J Vet Res. 2009;57:3–11. https://doi.org/10.14943/jjvr.57.1.3.
Ray LA, Heys JJ. Fluid flow and mass transport in brain tissue. Fluids. 2019;4(4):196. https://doi.org/10.3390/fluids4040196.
Rudick RA, Zirretta DK, Herndon RM. Clearance of albumin from mouse subarachnoid space: a measure of CSF bulk flow. J Neurosci Methods. 1982;6(3):253–9. https://doi.org/10.1016/0165-0270(82)90088-7.
Matthew J., Simon Jeffrey J., Iliff (2016) Regulation of cerebrospinal fluid (CSF) flow in neurodegenerative neurovascular and neuroinflammatory disease BBA-Mol Basis Dis 1862(3) 442-451. https://doi.org/10.1016/j.bbadis.2015.10.014.
Smith AF, Doyeux V, Berg M, Peyrounette M, Haft-Javaherian M, Larue AE, Slater JH, Lauwers F, Blinder P, Tsai P, Kleinfeld D, Schaffer CB, Nishimura N, Davit Y, Lorthois S. Brain capillary networks across species: A few simple organizational requirements are sufficient to reproduce both structure and function. Front Physiol. 2019;10:1–22. https://doi.org/10.3389/fphys.2019.00233.
Boero JA, Ascher J, Arregui A, Rovainen C, Woolsey TA. Increased brain capillaries in chronic hypoxia. J Appl Physiol. 1999;86:1211–9. https://doi.org/10.1152/jappl.1999.86.4.1211.
Thomsen MS, Humle N, Hede E, Moos T, Burkhart A, Thomsen LB. The bloodbrain barrier studied in vitro across species. PLOS One. 2021;16(3):e0236770. https://doi.org/10.1371/journal.pone.0236770.
Janiurek MM, Soylu-Kucharz R, Christoffersen C, Kucharz K, Lauritzen M. Apolipoprotein M-bound sphingosine-1-phosphate regulates blood–brain barrier paracellular permeability and transcytosis. eLife. 2019;8 https://doi.org/10.7554/eLife.49405.
Poulin P, Theil FP. A priori prediction of tissue: Plasma partition coefficients of drugs to facilitate the use of physiologically-based pharmacokinetic models in drug discovery. J Pharm Sci. 2000;89:16–35. https://doi.org/10.1002/(SICI)1520-6017(200001)89:13.0.CO;2-E.
Iversen NK, Malte H, Baatrup E, Wang T. The normal acid-base status of mice. Respir Physiol Neurobiol. 2012;180:252–7. https://doi.org/10.1016/j.resp.2011.11.015.
Tyrtyshnaia AA, Lysenko LV, Madamba F, Manzhulo IV, Khotimchenko MY, Kleschevnikov AM. Acute neuroinflammation provokes intracellular acidification in mouse hippocampus. J Neuroinflammation. 2016;13(1). https://doi.org/10.1186/s12974-016-0747-8.
Kasper D, Planells-Cases R, Fuhrmann JC, Scheel O, Zeitz O, Ruether K, Schmitt A, Poët M, Steinfeld R, Schweizer M, Kornak U, Jentsch TJ. Loss of the chloride channel ClC-7 leads to lysosomal storage disease and neurodegeneration. The EMBO Journal. 2005;24(5):1079–91. https://doi.org/10.1038/sj.emboj.7600576.
Majumdar A, Cruz D, Asamoah N, Buxbaum A, Sohar I, Lobel P, Maxfield FR. Activation of microglia acidifies lysosomes and leads to degradation of alzheimer amyloid fibrils. Mol Biol Cell. 2007;18:1490–6. https://doi.org/10.1091/mbc.E06.
Pla A, Pascual M, Consuelo G. Autophagy constitutes a protective mechanism against ethanol toxicity in mouse astrocytes and neurons. PLOS One 2016;11(4) e0153097. https://doi.org/10.1371/journal.pone.0153097.
Henry AG, Aghamohammadzadeh S, Samaroo H, Chen Y, Mou K, Needle E, Hirst WD. Pathogenic LRRK2 mutations through increased kinase activity produce enlarged lysosomes with reduced degradative capacity and increase ATP13A2 expression. Hum Mol Genet. 2015;24(21):6013–28. https://doi.org/10.1093/hmg/ddv314.
Bae M, Patel N, Xu H, Lee M, Tominaga-Yamanaka K, Nath A, Geiger J, Gorospe M, Mattson MP, Haughey NJ. Activation of TRPML1 clears intraneuronal Aβ in preclinical models of HIV infection. J Neurosci. 2014;34(34):11485–503. https://doi.org/10.1523/JNEUROSCI.0210-14.2014.
Christensen HL, Barbuskaite D, Rojek A, Malte H, Christensen IB, Füchtbauer AC, Füchtbauer E-M, Wang T, Praetorius J, Damkier HH. The choroid plexus sodium-bicarbonate cotransporter NBCe2 regulates mouse cerebrospinal fluid pH. J Physiol. 2018;596(19):4709–28. https://doi.org/10.1113/JP275489.
Herculano-Houzel S, Mota B, Lent R. Cellular scaling rules for rodent brains. Proc Natl Acad Sci. 2006;103(32):12138–43. https://doi.org/10.1073/pnas.0604911103.
Keep RF, Jones HC. A morphometric study on the development of the lateral ventricle choroid plexus choroid plexus capillaries and ventricular ependyma in the rat Developmental. Brain Res. 1990;56(1):47–53. https://doi.org/10.1016/0165-3806(90)90163-S.
Chan GNY, Saldivia V, Yang Y, Pang H, de Lannoy I, Bendayan R. In vivo induction of P-glycoprotein expression at the mouse blood-brain barrier: an intracerebral microdialysis study. J Neurochem. 2013;127(3):342–52. https://doi.org/10.1111/jnc.12344.
Zhuang Y, Fraga CH, Hubbard KE, Hagedorn N, Panetta JC, Waters CM, Stewart CF. Topotecan central nervous system penetration is altered by a tyrosine kinase inhibitor. Cancer Res. 2006;66:11305–13. https://doi.org/10.1158/0008-5472.CAN-06-0929.
Hudachek SF, Gustafson DL. Physiologically based pharmacokinetic model of lapatinib developed in mice and scaled to humans. J Pharmacokinet Pharmacodyn. 2013;40:157–76. https://doi.org/10.1007/s10928-012-9295-8.
Hughes JH, Upton RN, Reuter SE, Rozewski DM, Phelps MA, Foster DJR. Development of a physiologically based pharmacokinetic model for intravenous lenalidomide in mice. Cancer Chemother Pharmacol. 2019;84:1073–87. https://doi.org/10.1007/s00280-019-03941-z.
Methaneethorn J, Naosang K, Kaewworasut P, Poomsaidorn C, Lohitnavy M. Development of a Physiologically-based pharmacokinetic model of δ9-tetrahydrocannabinol in mice, rats, and pigs. Eur J Drug Metab Pharmacokinet. 2020;45:487–94. https://doi.org/10.1007/s13318-020-00616-6.
Pawaskar DK, Straubinger RM, Fetterly GJ, Hylander BH, Repasky EA, Ma WW, Jusko WJ. Physiologically based pharmacokinetic models for everolimus and sorafenib in mice. Cancer Chemother Pharmacol. 2013;71:1219–29. https://doi.org/10.1007/s00280-013-2116-y.
Zake DM, Kurlovics J, Zaharenko L, Komasilovs V, Klovins J, Stalidzans E. Physiologically based metformin pharmacokinetics model of mice and scale-up to humans for the estimation of concentrations in various tissues. PLoS One 2021; 16:. https://doi.org/10.1371/journal.pone.0249594.
Saleh MAA, Bloemberg JS, Elassaiss-Schaap J, de Lange ECM. Drug Distribution in Brain and Cerebrospinal Fluids in Relation to IC50 Values in Aging and Alzheimer’s Disease, Using the Physiologically Based LeiCNS-PK3.0 Model. Pharm Res. 2022;39:1303–19. https://doi.org/10.1007/s11095-022-03281-3.
Yamamoto Y, Välitalo PA, Huntjens DR, Proost JH, Vermeulen A, Krauwinkel W, Beukers MW, Van Den Berg DJ, Hartman R, Wong YC, Danhof M, Van Hasselt JGC, De Lange ECM. Predicting drug concentration-time profiles in multiple CNS compartments using a comprehensive physiologically-based pharmacokinetic model. CPT Pharmacomet Syst Pharmacol. 2017;6:765–77. https://doi.org/10.1002/psp4.12250.
Gupta M, Bogdanowicz T, Reed MA, Barden CJ, Weaver DF. The Brain exposure efficiency (BEE) score. ACS Chem Neurosci. 2020;11:205–24. https://doi.org/10.1021/acschemneuro.9b00650.
Garberg P, Ball M, Borg N, Cecchelli R, Fenart L, Hurst RD, Lindmark T, Mabondzo A, Nilsson JE, Raub TJ, Stanimirovic D, Terasaki T, Öberg JO, Österberg T. In vitro models for the blood-brain barrier. Toxicol Vitr. 2005;19:299–334. https://doi.org/10.1016/j.tiv.2004.06.011.
Hirasawa M, Saleh MAA, de Lange ECM. The Extension of the LeiCNS-PK3.0 model in combination with the “Handshake” approach to understand brain tumor pathophysiology. Pharm Res. 2022;39:1343–61. https://doi.org/10.1007/s11095-021-03154-1.
Pye CR, Hewitt WM, Schwochert J, Haddad TD, Townsend CE, Etienne L, Lao Y, Limberakis C, Furukawa A, Mathiowetz AM, Price DA, Liras S, Lokey RS. Nonclassical Size Dependence of Permeation Defines Bounds for Passive Adsorption of Large Drug Molecules. J Med Chem. 2017;60:1665–72. https://doi.org/10.1021/acs.jmedchem.6b01483.
de Lange EC. The mastermind approach to CNS drug therapy: translational prediction of human brain distribution, target site kinetics, and therapeutic effects. Fluids Barriers CNS. 2013;10:12. https://doi.org/10.1186/2045-8118-10-12.
de Lange ECM, Hammarlund Udenaes M. Understanding the blood-brain barrier and beyond: challenges and opportunities for novel cns therapeutics. Clin Pharmacol Ther. 2022;111:758–73. https://doi.org/10.1002/cpt.2545.
Funding
This project was funded by Leiden Academic Center for Drug Research (LACDR), Leiden University, Leiden, The Netherlands and has received funding from the European Union’s Horizon 2020 research and innovation program under grant agreement number 848068.
Author information
Authors and Affiliations
Contributions
Mohammed AA Saleh: Conceptualization, Data collection, Data analysis, Model simulations, writing original draft, review and editing. Berfin Gülave: Conceptualization, Data collection, Data analysis, Model simulations, writing original draft, review and editing. Olivia Campagne: Data supply. Clinton F Stewart: Data Supply. Jeroen Elassaiss-Schaap: Conceptualization, writing original draft, review and editing. Elizabeth CM de Lange: Conceptualization, writing original draft, review and editing.
Corresponding author
Ethics declarations
Conflict of Interest
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Saleh, M.A.A., Gülave, B., Campagne, O. et al. Using the LeiCNS-PK3.0 Physiologically-Based Pharmacokinetic Model to Predict Brain Extracellular Fluid Pharmacokinetics in Mice. Pharm Res 40, 2555–2566 (2023). https://doi.org/10.1007/s11095-023-03554-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11095-023-03554-5