Abstract
Strawberry (Fragraria x ananassa) is a crop affected by various soil-borne fungal pathogens with mostly non-specific foliar symptoms and often requiring laboratory isolation for correct diagnosis. Moreover, these nonspecific foliar symptoms, appreciated by the human eye, appear after some time following infection by the pathogen. Early detection of plant diseases is one of the primary objectives in agriculture because it may contribute to identifying more tolerant cultivars in breeding programs and optimise pesticide use in agricultural production with earlier applications in emerging disease foci. New technologies, such as remote sensing and machine learning (ML) algorithms, have arisen as potential tools to improve the ability to detect and classify different crop diseases. The combined use of hyperspectral imagery and ML algorithms were investigated to detect and classify the physiological stress caused by early infections of Fusarium wilt in strawberry plants. Six ML models, namely artificial neural network, decision tree, K-nearest neighbour, support vector machine, multinomial logistic regression and Naïve Bayes were developed to estimate physiological stress associated with Fusarium wilt disease. The results showed that stomatal conductance (gs) and photosynthesis (A) declined even without visual symptoms of the disease. Among the six ML models evaluated, the artificial neural network model showed the highest classification performance with an overall accuracy of 81%, regardless of the physiological parameter utilized for model training. Moreover, the artificial neural network accurately predicted the absolute values of both physiological parameters (gs and A) based on the complete spectral signature from visually healthy foliar tissue, achieving coefficients of determination of 84% and 81%, respectively. Consequently, ML models utilizing physiological response data and hyperspectral imaging exhibited remarkable robustness, facilitating the estimation of Fusarium wilt severity in strawberry plants even without visual symptoms.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Strawberries are the most produced and commercialized berries in the world. Since 2016, strawberry production has increased by 8% to reach 396,000 ha worldwide, producing 8,893 Mt (FAO, 2019). In Spain, strawberries are grown mainly in Huelva, a province covering most red fruit production. In 2021, this province’s strawberry-cultivated area was around 6,700 ha, representing 93% of the national total. Fusarium oxysporum f sp. fragrariae (Winks and Y.N. Williams) (Fof) is one of the most dangerous soilborne pathogens affecting strawberries worldwide (Henry et al., 2017). Fusarium oxysporum Schltdl. is an asexual ascomycete that causes severe vascular wilt disease in many crops (Booth et al., 1971). This saprophytic/necrotrophic fungus may survive in soil organic matter as chlamydospores for several years. The formae speciales of F. oxysporum usually have a narrow host range, often restricted to a single plant species (Kistler et al., 1998), and there are non-pathogenic strains morphologically indistinguishable from pathogenic strains (Suga et al., 2012). In strawberry crops, Fof inoculum generally includes contaminated soil and plants with latent infection (Katoh et al., 2021).
Since the prohibition of methyl bromide, there has been a notable increase in soilborne fungal diseases affecting strawberries, with unclear alternative control strategies (Holmes et al., 2020). Host plant resistance against these diseases has become paramount in strawberry breeding programs. Extensive research by Pincot et al. (2018, 2022) has identified five dominant resistance genes (Fw1-5) against F. oxysporum in strawberries. Several studies, including those by Borrero et al. (2017) and Fang et al. (2012), have focused on identifying resistant strawberry cultivars. It is essential to associate these cultivars with the corresponding pathotypes of F. oxysporum, as defined by Henry et al. (2021) and Pincot et al. (2022).
Besides breeding for disease resistance, the early detection and timely control of diseases are also critical factors in preventing yield losses (Garrido et al., 2011). Standard methods of early disease detection involve processes such as culturing, isolation, and molecular techniques following visual inspection of the plants (Debode et al., 2009; Koike & Gordon, 2015). Polimerase Chain Reaction primers allowed to detect Fof propagules in field soil, but they may miss specific Fof pathotypes (Koike & Gordon, 2015). Burkhardt et al. (2019) also introduced primers for Fof detection, while Katoh et al. (2021) devised a rapid LAMP assay requiring minimal equipment. Despite their accuracy, these methods are labor-intensive for large-scale scouting (Tapia-Tussell et al., 2008; Lu et al., 2017), and they cannot detect the Spanish pathotype of Fof.
For these reasons, non-destructive methods for detecting plant diseases have generated considerable interest among researchers and farmers. The integration of advanced sensors for proximal and remote sensing with accessible machine-learning algorithms holds promise for monitoring strawberry growth effectively. Such tools can aid breeding programs and disease control in strawberry fields by identifying infected plants before visible symptom occurs. This capability is especially critical for accurately assessing disease tolerance levels among resistant cultivars and determining whether non-pathogenic F. oxysporum strains impact crop physiology, potentially threatening productivity despite the absence of visible disease symptoms.
Various studies have explored non-destructive techniques for plant disease identification, mainly using RGB, multispectral, and hyperspectral images (Zheng et al., 2021). Hyperspectral systems, covering a wide spectral range with high resolution, showed promising results in the early detection of asymptomatic pathogens (Zhang et al., 2020). Siedliska et al. (2018) achieved over 97% accuracy in detecting fungal infections in strawberries using VNIR/SWIR hyperspectral analysis and a back propagation artificial neural network (ANN) model. Similarly, using hyperspectral data and ML algorithms, Lu et al. (2017) successfully detected Anthracnose crown rot in field-grown strawberries. Yeh et al. (2013, 2016) demonstrated early detection of foliar anthracnose in strawberries, achieving around 80% accuracy using different hyperspectral image analysis techniques. These studies underscore the potential of hyperspectral imaging for disease detection before visible symptoms emerge. Additionally, Pujari et al. (2016) compared ML models for classifying crop diseases based on plant symptom characteristics, with support vector machines (SVM) outperforming ANN with 92% accuracy.
The detection of F. oxysporum in other agricultural crop species has undergone significant advancements owing to the integration of diverse innovative techniques. Dang et al. (2020) pioneered a creative approach to detecting radish wilt using RGB and NIR images from UAVs. Studies such as those conducted by Navarro et al. (2022) and Shih et al. (2023) have elucidated distinct spectral features associated with Fusarium wilt, enabling early detection even before visible symptoms manifest in wild rocket and moth orchids, respectively. Through methods like spectral angle mapper (SAM) and constrained energy minimization (CEM), coupled with ANN, SVM, and Random Forest Classifiers (RFC), researchers have effectively identified disease-specific signals while suppressing background noise in hyperspectral spectra. Additionally, integrating thermography, as highlighted by Rispail and Rubiales (2015), showed promising results in assessing the resistance of pea plants to Fusarium wilt and other soil-borne pathogens by detecting subtle temperature or transpiration rate changes, providing valuable insights into plant physiological responses to disease stressors.
These findings highlight the capability of proximal and remote sensors, combined with ML algorithms, to develop practical, non-destructive disease detection methods with high accuracy and efficiency. Building upon these arguments, the objectives of this work are: (1) to assess the visual symptomatology and physiological impairment, particularly leaf gas exchange, induced by six F. oxysporum isolates obtained from strawberry plants; (2) to evaluate the effectiveness of hyperspectral images obtained from ‘non-symptomatic’ leaf tissue, combined with machine learning models, in distinguishing between healthy and Fusarium wilted strawberry plants grown under controlled conditions, and determining the severity of physiological impairment caused by F. oxysporum f sp. fragrariae.
These objectives are designed to address the uncertainties surrounding the physiological impacts of ‘non-pathogenic’ F. oxysporum isolates on apparently healthy strawberry plants, and to explore the potential of remote sensing and machine learning for early detection and severity assessment of Fusarium wilt.
Materials and methods
Plant material and experimental design
This work was carried out at the Higher Technical School of Agricultural Engineering (ETSIA) facilities of the University of Seville (Seville, 37º 21’ N, 5º 56’ W). Strawberry plants cv. Palmeritas (Masiá Ciscar, Lepe, Spain) were grown in pots of 1 L volume using a completely randomized block experimental design with four blocks and seven treatments. One treatment served as a control treatment (non-inoculated). By contrast, the other six treatments involved different Fusarium oxysporum isolates obtained from strawberry plants (inoculated plants), with eight and four replications for control and different isolates, respectively. The plants were inoculated with the following isolates: F74, F111, F115, F134, F138 and F141. The isolate F74 was characterized as Fof causing strawberry Fusarium wilt in different trials (Borrero et al., 2017; Henry et al., 2021). This Spanish isolate has a distinct genotype and phenotype compared to other isolates from other countries (Henry et al., 2021). The other isolates in this study were isolated in Spain and considered non-pathogenic in previous trials (data not shown). Subsequently, the pots were randomly placed in a growth chamber under controlled environmental conditions with temperatures set at 28ºC during the day and 24ºC at night, maintaining a twelve-hour photoperiod. The plants were manually irrigated twice weekly with two distinct nutrient solutions: one comprised of Peter’s foliar feed (27 + 15 + 12; N + P2O5 + K2O) (Scotts, Heerlen, The Netherlands) at 0.5 g L-1, and the other consisting of CaCl2 · 2H2O, with a dosage of 0.7 g L-1 (Panreac Química SLU, Barcelona, Spain).
Inoculation conditions
The fungal isolates used belong to the Plant Pathology Laboratory of ETSIA. All these F. oxysporum isolates were stored at -80 °C before being used. The inoculum of all the Fusaria was grown for seven days in AMAP culture medium (Borrero et al., 2009) containing 10 g of agar, 10 g of malt extract, 2.0 g of L-asparagine, 0.5 g of Peter’s foliar feed and micronutrients per L of distilled water. To obtain a conidial suspension, sterile water (5 mL) was added to each culture plate before scraping the culture surface with a sterile bent metallic rod. The concentration of conidia was determined with a hemocytometer and adjusted to 2 × 106 conidia mL− 1. The plants were inoculated by root-dipping in the abovementioned suspension for 30 min after root pruning. The inoculation was performed one week before further measurements.
Hyperspectral imaging system and data acquisition
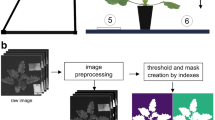
A Headwall Nano-Hyperspec hyperspectral pushbroom camera (Headwall Photonics, Bolton, UK) with a spectral range from 397 to 1003 nm and a spectral resolution of 272 bands was used to perform the measurements. The camera sensor is a line-scan, meaning the number of lines scanned (y-axis) depends on the scanning time. By contrast, in the ground projection, 640 constant lines are measured (x-axis), with a spectral resolution of 272 bands per projection line (z-axis). The sensor exposure time was 15 ms, and the frame rate was 15 ms. The camera was fixed to an aluminium frame at a height of 0.5 m with a scanning angle of 0º (nadir view). The pots were placed on a conveyor belt powered by a 12 V motor (electric motor output of 6.1 V and 1 A) under the aluminium structure, moving at a constant speed of 0.06 m s− 1 (Fig. 1). Additional artificial light was provided by two 400 W tungsten halogen lamps (Maurer Ferritalia, Padova, Italy) fixed to both sides of the structure. The hyperspectral camera was connected via an Ethernet cable to a computer to record the images and subsequent data transfer. Measurements were made weekly (six to eight days) for four weeks, described as M1, M2, M3 and M4, respectively.
Data processing
Hyperspectral data underwent radiometric calibration to obtain spectral reflectance data, for which dark and white reference measurements were performed. Covering the camera shutter, a dark reference value was obtained for each sampling date. For white reference, a diffuse reflectance target (Labsphere, USA, Spectralon® diffuse Reflectance target, 127 × 127 mm, 99% reflectance, calibrated) was placed aside each plant. The pixels corresponding to the white reference panel were identified using the 728 nm wavelength, as this band exhibited the highest signal intensity among the panel pixels. To determine the spectral signature of the white reference, only the pixels with the top 10% highest radiation values in the 728 nm band were selected and then averaged. Hyperspectral data processing and analysis were performed using the open-source programming language Python (Python Software Foundation, https://www.python.org/). Firstly, the hyperspectral image was corrected as follows:
where raw is the uncorrected image file and dark and white denote the dark and white reference values, respectively.
The hyperspectral images were then subjected to Savitzky-Golay smoothing. Then, the first 35 and the last 40 wavebands were removed to reduce possible noise due to high and random variation of spectral reflectance in these bands, resulting in images with a total of 197 spectral bands. The dataset was then normalized between 0 and 1. Finally, a manual pixel selection process was carried out within regions of interest (ROIs) to obtain an average spectral signature per plant of leaf tissues without any visual symptoms of the disease. Subsequently, a ‘csv’ file is composed as a dataset in which the rows represent individual plants, and the columns represent the spectral bands. This dataset consists of the selected ROIs’ average spectral signature, representing visually healthy leaf tissue.
Disease severity and physiological measurements
On the same days that hyperspectral images were taken, measurements of the severity and leaf gas exchange were also made. Disease severity was assessed, considering leaf wilting to be the main symptom of Fusarium wilt. Weekly observations were taken for disease evolution. Plants were scored with the relative number of symptomatic leaves per plant (number of symptomatic leaves / total number of leaves) (Hernández-Muñiz et al., 2023). The area-under-the-disease-progress-curve-standardized (AUDPCs) per pot was calculated from the disease severity values by the trapezoidal integration method between the onset of symptoms and the ending time of the assay and by dividing the total duration (days) of the epidemic the assay (Campbell & Madden, 1990). In addition, leaf gas exchange measurements were taken on one fully expanded leaf per plant without any visual symptoms of the disease. Stomatal conductance (gs) and leaf photosynthesis rate (A) were measured using a differential portable system (CIRAS-3 Model, PP-Systems, Amesbury, U.S.A.), in which photosynthetically active radiation (PAR) and air CO2 concentration were set to 1200 µmol m− 2 s− 1 and 400 µmol mol− 1, respectively.
Machine learning models description
Six different classifier models, artificial neural network (ANN), decision tree (DT), K-nearest neighbour (K-NN), support vector machine (SVM), multinomial logistic regression (MLR), and Naïve Bayes (NB), were developed using the complete spectral signatures of both inoculated and non-inoculated. The dataset was divided into training and validation sets, with 80% and 20% of individuals, respectively. Due to the imbalanced nature of the dataset, stratified cross-validation (Szeghalmy & Fazekas, 2023) was conducted to train and validate the models to ensure that the proportion of examples from each class is maintained in the validation set. Early stopping and regularization were used to avoid model overfitting. The algorithms used were tested with different settings. The model configurations finally used in the study are described in what follows. A Multi-Layer Perceptron (MLP) framework was adopted for the ANN model architecture, comprising input, hidden, and output layers. The optimal number of neurons was determined as the number of input data, followed by 150, 100, and 50 hidden layers, and the output corresponding to the number of classes described. Learning rate (0.001), training epochs (500), rectified linear unit (ReLU) activation function after hidden layers, and Adam as the solver function were set. The ANN model was used with two configurations: as a classifier (the output layer is the class to which the plant belongs according to the level of disease severity) and as a regression (the output layer is the value of the physiological indicator used in each case, gs or A). For this purpose, a ‘softmax’ output layer was defined for the ANN classifier version and a ‘linear’ output layer for the regression one.
For the DT algorithm, a decision tree was constructed using the Gini Index criterion, and the ‘best’ attribute was selected to optimize the splitting of the provided dataset. The KNN algorithm was set to minimize the influence of outliers or incorrect classes, with K = 3 (considering the three nearest neighbours) and ‘distance’ as the weight parameter. This choice was made because neighbours closer to a query point will have a greater influence than neighbours further away. The SVM algorithm was set with a linear-based kernel to find an optimal hyperplane for effectively separating data classes. The MLR algorithm, an extension of binary logistic regression for situations with more than two categories, was configured with the ‘newton-cg’ solver and the ‘multinomial’ parameter. The NB algorithm was set to use a Gaussian distribution model due to the dataset size. An approach based on confusion matrices and classification reports was employed to evaluate the model’s performance. Matrices provided a visual representation of each class classified by the corresponding model. The Y-axis of each matrix represented the actual values (true labels), and the X-axis represented the values that the model predicted (estimated labels). Each matrix showed the classification results in terms of true positives (TP), false negatives (FN), false positives (FP) and true negatives (TN). This allowed for the evaluation of the model’s performance in identifying each class.
Statistical analysis
The severity, stomatal conductance, and leaf photosynthesis rate of each F. oxysporum isolate were analysed using one-way ANOVA to determine the effects of the isolates during the experiment. Heterogeneous groups were separated using Tukey’s test (HSD) (P < 0.05) to discriminate differences between isolates. Severity data were transformed for normalization following the Box-Cox methodology, which employs power transformations to correct unequal variances and non-linearity in the relationship. The machine learning models used Linear regressions to compare the estimated and measured values of stomatal conductance and leaf photosynthesis rate. The root means square error (RMSE) and the mean absolute error (MAE) values were calculated using the same algorithms of the chosen models to analyse predictive performance.
The data presented for models were unbalanced since the number of data included in each class was not equal, so the metric used is represented as ‘F1-score’ since it follows a weighting between the ‘Precision’ and ‘Recall’ metrics of the classification report. Precision measures the quality of the classification and is calculated as VP / (VP + FP), while the Recall informs about the identification capacity and is calculated as TP / (TP + FN). The F1-score combines these two metrics in a single measure as the harmonic mean between Precision and Recall values. Thus, F1-score balances the classification quality and identification ability in unbalanced cases.
Results
Fusarium wilt severity
Figure 2 shows the visual disease severity caused by the different isolates of F. oxysporum on the last sampling date (M4), the only one in which significant differences between control and inoculated plants were observed. Only two isolates (F74, F141) showed a noticeable increase in visual disease symptoms on leaf tissues during the disease. However, due to the high variability of visible symptoms observed in F141, only F74 showed significant differences with the control treatment.
Visual disease severity on leaf tissues caused by each isolate of Fusarium oxysporum on the last sampling date. Boxplots were represented for each treatment as control (non-inoculated) plants (n = 8) and six Fusarium oxysporum isolates (n = 4). The star indicates significant differences between each F. oxysporum isolate and control plants (P < 0.05)
Leaf gas exchange responses to F. Oxysporum isolates
Figure 3 shows the values of gs measured during the experiment. The control treatment (non-inoculated plants) showed relatively constant gs values in the 50–60 mmol m− 2 s1 range. All F. oxysporum isolates showed significantly lower gs values throughout the four sampling dates than the control treatment. However, F111 and F115 did not show significant differences until the second and third sampling dates. The isolate F74 caused the most severe reduction in gs, with a 35% reduction in gs relative to the control treatment in M1 that dropped to 90% in M3 and M4. The rest of the isolates showed a gs decrease with respect to the control treatment, which was less severe than that observed in F74, being significant in all cases from M3 onwards and with percentage reductions in the 15–30% range.
Stomatal conductance of control and inoculated plants for each sampling day was described as M1 (a), M2 (b), M3 (c) and M4 (d). Boxplots were represented for each treatment as control (non-inoculated) plants (n = 8) and six Fusarium oxysporum isolates (n = 4). The star indicates significant differences between each F. oxysporum isolate and control plants (P < 0.05)
Figure 4 shows A measured in control and inoculated plants during the experiment. The control plants showed relatively constant A values throughout the experiment, with average values around 6 µmol m− 2 s− 1. As observed with gs, all F. oxysporum isolates caused a significant reduction in A relative to the control treatment. However, F111 and F115 did not do so until the second and third sampling dates, respectively. F74 was again the isolate that caused the greatest severity in A reduction, with up to 90% reductions on the last two sampling dates. The remaining isolates showed a progressive decrease of A, reaching values close to 30% on the previous sampling date.
Leaf photosynthesis rate (A) of control and inoculated plants for each sampling day was described as M1 (a), M2 (b), M3 (c) and M4 (d). Boxplots were represented for each treatment as control (non-inoculated) plants (n = 8) and six Fusarium oxysporum isolates (n = 4). The star indicates significant differences between each F. oxysporum isolate and control plants (P < 0.05)
Classification and estimation of Fusarium wilt severity by machine learning algorithms
Plants inoculated with the six F. oxysporum isolates were used for model training since they significantly reduced physiological parameters compared to control plants. To train the classification response, the plants were categorized into four categories based on the severity of physiological damage. The established categories were a healthy plant, mild, moderate, and severe impairment. To assess the suitability of using either stomatal conductance or leaf photosynthesis rate as indicators of physiological damage, the plants were categorized twice, once using gs and the other using A as an indicator of biotic stress. In both cases, the variables gs and A were normalized by dividing their values by the maximum value observed for both variables throughout the experiment. Once the variables gs and A were normalized, the normalized ranges of gs and A used in the categorization of plants were as follows: healthy plant (1-0.85), mild impairment (0.85 − 0.75), moderate impairment (0.75 − 0.65), and severe impairment (< 0.65).
shows the average spectral signatures obtained throughout the entire study period for each class for both categorization criteria, i.e. based on gs and A. It can be observed that the spectral region with the most significant variability in reflectance between classes was from the green region (550 nm) to the red region (670 nm), while differences are less noticeable in the NIR region (700–900 nm)
Figure 5 Average spectral signatures and standard deviations for each class were distinguished as follows: class 1 (healthy plant), class 2 (mild impairment), class 3 (moderate impairment), and class 4 (severe impairment). The plants were categorized using stomatal conductance-based (a) and leaf photosynthesis rate-based (b) physiological impairment. For the gs-based and A-based categorizations, the number of individuals belonging to each class was 35 (class 1), 37 (class 2), 30 (class 3), 26 (class 4) and 23 (class 1), 29 (class 2), 34 (class 3), 42 (class 4), respectively.
Figure 6 displays the confusion matrices for the gs-based physiological impairment classes categorized by all studied models. Additionally, Table 1 presents the results of these models in the classification report. The ANN model (Fig. 6a) best classified healthy plants and those with varying degrees of physiological damage (Table 1). Analysing the F1-score metric, this model demonstrated an overall accuracy of 85%, followed by the DT model (77%). The other evaluated models (KNN, SVM, MLR, and NB) exhibited a low classification performance, with overall F1-score values below 60%.
Analysing the performance of the top-performing models (ANN and DT), it can be observed in Fig. 6 that both models correctly classified 100% of healthy individuals. The classification of individuals with some kind of physiological impairment based on gs was slightly better in ANN, which correctly classified 15 out of 19 individuals. In comparison, 13 individuals were correctly classified by DT. Another critical aspect of the ANN model compared to DT is that it misclassified only one individual with biotic stress as a healthy plant. By contrast, the DT model misclassified three individuals as healthy plants. The remaining evaluated models encountered significant difficulties in correctly classifying healthy and diseased individuals, with SVM’s performance being notably poor.
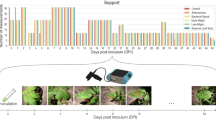
Confusion matrices were obtained for artificial neural network (a), decision tree (b), K-nearest neighbour (c), support vector machine (d), multinomial logistic regression (e) and Naïve Bayes (f) classifiers using gs-based physiological impairment classes. Four classes were established: healthy plants, mild, moderate, and severe impairment. ‘True label’ corresponds to the actual class, while ‘Predicted label’ represents the class predicted by the model. The colour scale indicates the number of randomly selected plants or samples (support) used for prediction
Figure 7 shows the confusion matrices obtained with the six classifiers for classes established using the A as a physiological indicator of biotic stress severity. As it has been observed when plants were classified using gs as a physiological indicator, the ANN algorithm (Fig. 7a) also demonstrated the highest classification performance, correctly classifying all healthy plants and 19 out of 21 diseased plants. However, the performance of the ANN model to correctly classify the severity level of diseased plants was somewhat poorer than that observed using gs-based classes, with 15 out of 21 plants classified correctly for 15 out of 19 correctly classified in the case of gs-based classes. Analysing the F1-score metrics, the ANN model showed an overall accuracy of 77%, followed by the DT model with 65% (Table 2). In both cases, weaker classification performance was observed compared to that shown by the same models when classes were defined based on gs as a physiological indicator. The rest of the evaluated models (KNN, SVM, MLR, and NB) showed low classification potential, with overall F1-score values in the 30–50% range.
Confusion matrices obtained for artificial neural network (a), decision tree (b), K-nearest neighbour (c), support vector machine (d), multinomial logistic regression (e) and Naïve Bayes (f) classifiers using A-based physiological impairment classes. Four classes were established: healthy plants, mild, moderate, and severe impairment. ‘True label’ corresponds to the actual class, while ‘Predicted label’ represents the class predicted by the model. The colour scale indicates the number of randomly selected plants or samples (support) used for prediction
The best classification model (ANN) was also trained as a regression model to assess its reliability in predicting actual gs and A values in non-inoculated and inoculated strawberry plants. Figure 8 shows the relationships between measured and predicted gs (Fig. 8a) and A (Fig. 8b) values. Stomatal conductance presented a coefficient of determination (R2) of 0.84, an RMSE value of 5.03 mmol m− 2 s− 1 and a MAE value of 3.94 mmol m− 2 s− 1. The R2, RMSE and MAE values for A were 0.81, 0.64 µmol m− 2 s− 1 and 0.41 µmol m− 2 s− 1, respectively. The slopes of the regression lines were 0.87 (gs) and 0.99 (A), whereas the intercepts were 3.23 (gs) and − 0.13 (A).
Discussion
The disease caused by pathogenic F. oxysporum isolates is characterized by the plant’s difficulty in absorbing water and nutrients from the soil (Yadeta & Thomma, 2013), as the fungus affects the plant’s vascular system. In horticultural species such as tomato or cucumber, the disease caused by F. oxysporum has resulted in severe reductions in stomatal conductance (gs) and net photosynthesis rate (A) (Carmona et al., 2021; Lorenzini et al., 1997; Wang et al., 2015), with the latter being down-regulated not only by diffusional limitations (gs) but also by biochemical impairment in the leaf photosynthetic apparatus (Nogués et al., 2002; Susič et al., 2018).
Our findings show that strawberry plants inoculated with the pathogenic Fof isolate also exhibited the characteristic pattern of reduced gs and A as described for other species, with reductions of up to 90% in gs and A compared to control plants (non-inoculated) around 15 days after inoculation (Fig. 3). However, it has also been shown that F. oxysporum isolates considered as non-pathogenic may also cause moderate gs and A impairment, with reductions of up to 20–30% of these variables without any visible symptoms of the disease. This can have consequences at the production level, and its detection requires the development of simple and early detection methods due to the absence of any visual symptoms.
The results obtained in this study reveal that among the six machine learning models evaluated, the ANN model exhibited the highest classification performance, both when disease severity was based on gs and when it was based on A. The overall accuracy values evaluated through the F1-score metrics were 0.85 (gs-based) and 0.77 (A-based). To the best of our knowledge, this is the first study in which machine learning models have been developed for the early detection of F. oxysporum f sp. fragrariae in strawberries based on hyperspectral images. Therefore, it cannot directly compare the performance of the ANN model with other F. oxysporum detection studies in strawberries. However, for detecting powdery mildew in strawberry plants using RGB images, the ANN model also showed the highest classification performance (93.8%) (Chang et al., 2019). In strawberry fruits affected by Botrytis and Anthracnose, the ANN model was also highly accurate (> 95%) for disease detection using hyperspectral images (Siedliska et al., 2018). Ye et al. (2020) also evaluated three models (i.e., SVM, RF, and ANN) for detecting areas affected by Fusarium wilt in banana plantations using multispectral images taken from drones. In this case, all models showed similar accuracies (around 90%) in detecting visually infected areas. Although they did not evaluate the ANN model, Yeh et al. (2016) assessed the reliability of three models for classifying healthy and inoculated strawberry plants with visible symptoms of Anthracnose disease. In all cases, they obtained an accuracy close to 80%, similar to those obtained in this study for classifying F. oxysporum-infected strawberry plants without visual symptoms.
The development of machine learning models trained on hyperspectral images of strawberry plant´s apparently healthy (non-symptomatic) plant tissue represents, therefore, a valuable tool for early detection of physiological impairments caused by F. oxysporum isolates. Moreover, establishing disease severity classes based on stress indicators such as gs and A, commonly used for characterizing crop responses to environmental stresses, marks a departure from traditional methodologies that often rely on visual symptom severity for class determination. However, since most biotic and abiotic stresses decrease these physiological parameters (Abdulridha et al., 2019; Pantazi et al., 2017; Susič et al., 2018), ML models calibrated on the interactions between spectral signatures and variations in gs and A would require further testing under field conditions to determine their accuracy in detecting F. oxysporum infections in real-world scenarios with multiple stressors.
However, there is evidence in the literature that the combination of hyperspectral images and ML algorithms allows for precise discrimination between biotic and abiotic stresses in plants, as there is a certain degree of specificity in the spectral bands altered by each factor. In this sense, Susič et al. (2018) successfully differentiated nematode infestation from water deficiency in tomato plants using shortwave infrared spectral regions associated with O-H and C–H stretches. Pantazi et al. (2017) classified reflectance data from winter wheat crops with over 95% accuracy, identifying health status, nitrogen stress, and disease presence using SOM-based models. As demonstrated by Abdulridha et al. (2019), optimal band selection significantly improves disease detection accuracy, achieving up to 98% accuracy in identifying laurel wilt disease in avocados against nutrient deficiency. These advances underscore the efficacy of combining hyperspectral imaging and machine learning for precisely detecting and classifying plant stresses. Ortiz et al. (2019) found that spectral bands associated with F. oxysporum infection were identified in the visible range, while bands related to water stress were found in the near-infrared range, suggesting high specificity and sensitivity in detecting and dis.
These findings suggest that the models trained in this study can offer a high degree of specificity in detecting F. oxysporum f sp. fragariae (Fof) and the severity of its physiological damage. As future research directions, it is recommended to evaluate the performance of the models in detecting F. oxysporum f sp. fragariae under uncontrolled conditions and multiple stress factors, as well as across different cultivars and with a larger number of pathogenic F. oxysporum isolates.
Conclusions
Pathogenic Fof isolates caused rapid wilting of strawberry plants with severe drops in leaf gas exchange variables. The non-pathogenic F. oxysporum isolates also caused a reduction in leaf gas exchange variables in infected strawberry plants, although much less severe. The combined use of machine learning models and hyperspectral images obtained from healthy-appearing leaf tissue (without visual disease symptomology) allows the classification of healthy and diseased strawberry plants with different levels of physiological impairment. The ANN-based model provided the best classification performance, with an overall accuracy of more than 80% when plant classes were defined based on their stomatal conductance values. Further improvement and refinement of the model with new datasets are still needed to enhance the discrimination between Fof severity classes and to ensure high detection accuracies under multi-stressor environments (e.g., those potentially found under field conditions) and multi-cultivar environments (e.g., those found in breeding programs).
Data availability
The data will be available as requested and/or in a public repository once accepted.
Code Availability
The code will be available as requested and/or in a public repository once accepted.
References
Abdulridha, J., Ehsani, R., Abd-Elrahman, A., & Ampatzidis, Y. (2019). A remote sensing technique for detecting laurel wilt disease in avocado in presence of other biotic and abiotic stresses. Computers and Electronics in Agriculture, 156, 549–557. https://doi.org/10.1016/j.compag.2018.12.018.
Booth, C. (Ed.). (1971). Fungal culture media. Pages 49–94 in: Methods in Microbiology (Vol. 4). Academic.
Borrero, C., Trillas, I., & Avilés, M. (2009). Carnation Fusarium wilt suppression in four composts. European Journal of Plant Pathology, 123, 425–433. https://doi.org/10.1007/s10658-008-9380-4.
Borrero, C., Bascón, J., Gallardo, M. Á., Orta, M. S., & Avilés, M. (2017). New foci of strawberry Fusarium wilt in Huelva (Spain) and susceptibility of the most commonly used cultivars. Scientia Horticulturae, 226, 85–90. https://doi.org/10.1016/j.scienta.2017.08.034.
Burkhardt, A., Henry, P. M., Koike, S. T., Gordon, T. R., & Martin, F. (2019). Detection of Fusarium oxysporum f. sp. fragariae from infected Strawberry plants. Plant Disease, 103(5), 1006–1013. https://doi.org/10.1094/PDIS-08-18-1315-RE.
Campbell, C., & Madden, L. (1990). Introduction to plant disease epidemiology. Wiley.
Carmona, S. L., del Pilar Villarreal-Navarrete, A., Burbano-David, D., Gómez-Marroquín, M., Torres-Rojas, E., & Soto-Suárez, M. (2021). Protection of tomato plants against Fusarium oxysporum f. sp. lycopersici induced by chitosan. Revista Colombiana De Ciencias Hortícolas, 15(3), 2422–3719.
Chang, M., Shin, Nguyen-Quang, P., & Prithiviraj (2019). Comparison of image texture based supervised learning classifiers for strawberry powdery mildew detection. AgriEngineering, 1(3), 434–452. https://doi.org/10.3390/agriengineering1030032.
Dang, L. M., Wang, H., Li, Y., Min, K., Kwak, J. T., Lee, O. N., & Moon, H. (2020). Fusarium wilt of radish detection using RGB and near infrared images from Unmanned Aerial vehicles. Remote Sensing, 12(17), 2863. https://doi.org/10.3390/rs12172863.
Debode, J., Van Hemelrijck, W., Baeyen, S., Creemers, P., Heungens, K., & Maes, M. (2009). Quantitative detection and monitoring of Colletotrichum acutatum in strawberry leaves using real-time PCR. Plant Pathology, 58(3), 504–514. https://doi.org/10.1111/j.1365-3059.2008.01987.x.
Fang, X., Phillips, D., Verheyen, G., Li, H., Sivasithemparam, K., & Barbetti, M. J. (2012). Yields and resistance of strawberry cultivars to crown and root diseases in the field, and cultivar responses to pathogens under controlled environment conditions. Phytopathologia Mediterranea, 51(1), 69–84. http://www.jstor.org/stable/43872357.
FAO (Food and Agriculture Organization) (2019). FAOSTAT. http://faostat.fao.org. (last accessed 10th of August 2022).
Garrido, C., Carbú, M., Fernández-Acero, F., González-Rodríguez, V., & Cantoral, J. (2011). New insights in the study of strawberry fungal pathogens. Genes Genomes Genomics, 5(1), 24–39.
Henry, P. M., Kirkpatrick, S. C., Islas, C. M., Pastrana, A. M., Yoshisato, J. A., Koike, S. T., Daugovish, O., & Gordon, T. R. (2017). The population of Fusarium oxysporum f. sp. fragariae, cause of Fusarium wilt of strawberry, in California. Plant Disease, 101(4), 550–556. https://doi.org/10.1094/PDIS-07-16-1058-RE.
Henry, P. M., Pincot, D. D. A., Jenner, B. N., Borrero, C., Aviles, M., Nam, M. H., Epstein, L., Knapp, S. J., & Gordon, T. R. (2021). Horizontal chromosome transfer and independent evolution drive diversification in Fusarium oxysporum f. sp. fragariae. New Phytologist, 230(1), 327–340. https://doi.org/10.1111/nph.17141.
Hernández-Muñiz, P., Borrero, C., Ordóñez-Martín, J., Pastrana, A. M., & Avilés, M. (2023). Optimization of the use of industrial wastes in anaerobic soil disinfestation for the control of Fusarium wilt in strawberry. Plants, 12(18), 3185. https://doi.org/10.3390/plants12183185.
Holmes, G. J., Mansouripour, S. M., & Hewavitharana, S. S. (2020). Strawberries at the crossroads: Management of soilborne diseases in California without methyl bromide. Phytopathology, 110(5), 956–968. https://doi.org/10.1094/PHYTO-11-19-0406-IA.
Katoh, H., Yamazaki, S., Fukuda, T., Sonoda, S., Nishigawa, H., & Natsuaki, T. (2021). Detection of Fusarium oxysporum f. sp. fragariae by using loop-mediated isothermal amplification. Plant Disease, 105(4), 1072–1079. https://doi.org/10.1094/PDIS-03-20-0590-RE.
Kistler, H. C., Alabouvette, C., Baayen, R. P., Bentley, S., Brayford, D., Coddington, A., Correll, J., Daboussi, M. J., Elias, K., Fernández, D., Gordon, T. R., Katan, T., Kim, H. G., Leslie, J. F., Martyn, R., Migueli, D., Moore, Q., O’Donnell, N. Y., Ploetz, K., Rutherford, R. C., Summerell, M. A., Waalwijk, B., C., & Woo, S. (1998). Systematic numbering of vegetative compatibility groups in the plant pathogenic fungus fusarium oxysporum. Phytopathology, 88, 30–32.
Koike, S. T., & Gordon, T. R. (2015). Management of Fusarium wilt of strawberry. Crop Protection, 73, 67–72. https://doi.org/10.1016/j.cropro.2015.02.003.
Lorenzini, G., Guidi, L., Nali, C., Ciompi, S., & Soldatini, G. F. (1997). Photosynthetic response of tomato plants to vascular wilt diseases. Plant Science, 124(2), 143–152. https://doi.org/10.1016/S0168-9452(97)04600-1.
Lu, J., Ehsani, R., Shi, Y., Abdulridha, J., de Castro, A. I., & Xu, Y. (2017). Field detection of anthracnose crown rot in strawberry using spectroscopy technology. Computers and Electronics in Agriculture, 135(C), 289–299. https://doi.org/10.1016/j.compag.2017.01.017.
Navarro, A., Nicastro, N., Costa, C., Pentangelo, A., Cardarelli, M., Ortenzi, L., & Pane, C. (2022). Sorting biotic and abiotic stresses on wild rocket by leaf-image hyperspectral data mining with an artificial intelligence model. Plant Methods, 18(1), 45. https://doi.org/10.1186/s13007-022-00880-4.
Nogués, S., Cotxarrera, L., Alegre, L., & Trillas, M. I. (2002). Limitations to photosynthesis in tomato leaves induced by Fusarium wilt. New Phytologist, 154(2), 461–470. https://doi.org/10.1046/j.1469-8137.2002.00379.x.
Ortiz, J. C. M., Carvajal, L. M. H., & Fernandez, V. B. (2019). Detection of significant wavelengths for identifying and classifying Fusarium oxysporum during the incubation period and water stress in Solanum lycopersicum plants using reflectance spectroscopy. Journal of Plant Protection Research, 59(2), 244–254. https://doi.org/10.24425/jppr.2019.129290.
Pantazi, X. E., Moshou, D., Oberti, R., West, J., Mouazen, A. M., & Bochtis, D. (2017). Detection of biotic and abiotic stresses in crops by using hierarchical self organizing classifiers. Precision Agriculture, 18, 383–393. https://doi.org/10.1007/s11119-017-9507-8.
Pincot, D. D. A., Poorten, T. J., Hardigan, M. A., Harshman, J. M., Acharya, C. B., Cole, G. S., Gordon, T. R., Stueven, M., Edger, P. P., & Knapp, S. J. (2018). Genome-wide Association Mapping uncovers Fw1, a Dominant Gene conferring resistance to Fusarium Wilt in Strawberry. G3: Genes Genomes Genetics, 8(5), 1817–1828. https://doi.org/10.1534/g3.118.200129.
Pincot, D. D. A., ·Feldmann, M. J., Hardigan, M. A., Vachev, M. V., Henry, P. M., Gordon, T. R., Bjornson, M., Rodriguez, A., Cobo, N., ·Famula, R. A., Cole, G. S., Coaker, G. L., & Knapp, S. J. (2022). Novel Fusarium wilt resistance genes uncovered in natural and cultivated strawberry populations are found on three non–homoeologous chromosomes. Theoretical and Applied Genetics, 135, 2121–2145.
Pujari, D., Yakkundimath, R., & Byadgi, A. S. (2016). SVM and ANN based classification of plant diseases using feature reduction technique. International Journal of Interactive Multimedia and Artificial Intelligence, 3(7), 6–14. https://doi.org/10.9781/ijimai.2016.371.
Rispail, N., & Rubiales, D. (2015). Rapid and efficient estimation of pea resistance to the soil-borne pathogen Fusarium oxysporum by infrared imaging. Sensors (Basel, Switzerland), 15(2), 3988–4000. https://doi.org/10.3390/s150203988.
Shih, M. S., Chang, K. C., Chou, S. A., Liu, T. S., & Ouyang, Y. C. (2023). The automated detection of Fusarium wilt on Phalaenopsis using VIS-NIR and SWIR hyperspectral imaging. Remote Sensing, 15(17), 4174. https://doi.org/10.3390/rs15174174.
Siedliska, A., Baranowski, P., Zubik, M., Mazurek, W., & Sosnowska, B. (2018). Detection of fungal infections in strawberry fruit by VNIR/SWIR hyperspectral imaging. Postharvest Biology and Technology, 139, 115–126. https://doi.org/10.1016/j.postharvbio.2018.01.018.
Suga, H., Hirayama, Y., Morishima, M., Suzuki, T., Kageyama, K., & Hyakumach, M. (2012). Development of PCR primers to identify Fusarium oxysporum f. sp. fragariae. Plant Disease, 97(5), 619–625. https://doi.org/10.1094/PDIS-07-12-0663-RE.
Susič, N., Žibrat, U., Širca, S., Strajnar, P., Razinger, J., Knapič, M., & Stare, B. G. (2018). Discrimination between abiotic and biotic drought stress in tomatoes using hyperspectral imaging. Sensors and Actuators B: Chemical, 273, 842–852. https://doi.org/10.1016/j.snb.2018.06.121.
Szeghalmy, S., & Fazekas, A. (2023). A comparative study of the use of stratified cross-validation and distribution-balanced stratified cross-validation in Imbalanced Learning. Sensors (Basel, Switzerland), 23(4), 2333.
Tapia-Tussell, R., Quijano-Ramayo, A., Cortes-Velazquez, A., Lappe, P., Larque-Saavedra, A., & Perez-Brito, D. (2008). PCR-based detection and characterization of the fungal pathogens Colletotrichum gloeosporioides and Colletotrichum Capsici causing anthracnose in papaya (Carica papaya l.) in the Yucatan Peninsula. Molecular Biotechnology, 40(3), 293–298. https://doi.org/10.1007/s12033-008-9093-0.
Wang, M., Sun, Y., Sun, G., Liu, X., Zhai, L., Shen, Q., & Guo, S. (2015). Water balance altered in cucumber plants infected with Fusarium oxysporum f. sp. cucumerinum. Scientific Reports, 5(1), 7722. https://doi.org/10.1038/srep07722.
Yadeta, K. A., & Thomma, B. P. H. (2013). The xylem as battleground for plant hosts and vascular wilt pathogens. Frontiers in Plant Science, 4(97). https://doi.org/10.3389/fpls.2013.00097.
Ye, H., Huang, W., Huang, S., Cui, B., Dong, Y., Guo, A., Ren, Y., & Jin, Y. (2020). Identification of banana fusarium wilt using supervised classification algorithms with UAV-based multi-spectral imagery. International Journal of Agricultural and Biological Engineering, 13(3), 136–142. https://doi.org/10.25165/j.ijabe.20201303.5524.
Yeh, Y. H. F., Chung, W. C., Liao, J. Y., Chung, C. L., Kuo, Y. F., & Lin, T. T. (2013). A comparison of machine learning methods on hyperspectral plant disease assessments. IFAC Proceedings Volumes, 46(4), 361–365. https://doi.org/10.3182/20130327-3-JP-3017.00081.
Yeh, Y. H., Chung, W. C., Liao, J. Y., Chung, C. L., Kuo, Y. F., & Lin, T. T. (2016). Strawberry foliar anthracnose assessment by hyperspectral imaging. Computers and Electronics in Agriculture, 122, 1–9. https://doi.org/10.1016/j.compag.2016.01.012.
Zhang, N., Yang, G., Pan, Y., Yang, X., Chen, L., & Zhao, C. (2020). A review of advanced technologies and development for hyperspectral-based plant disease detection in the past three decades. Remote Sensing, 12(19), 3188. https://doi.org/10.3390/rs12193188.
Zheng, C., Abd-elrahman, A., & Whitaker, V. (2021). Remote sensing and machine learning in crop phenotyping and management, with an emphasis on applications in strawberry farming. Remote Sensing, 13(3), 1–29. https://doi.org/10.3390/rs13030531.
Acknowledgements
The Proyecto RTI2018 provided support for this study–094537-B-I00 (MCIU/AEI/FEDER, UE), Proyecto US-1380608 (US/JUNTA/FEDER, UE) and Contrato PRJ202104102 (Masiá Ciscar, S.A.).
Funding
Funding for open access publishing: Universidad de Sevilla/CBUA
Author information
Authors and Affiliations
Contributions
P. Castro-Valdecantos: Conceptualization, Methodology, Software, Validation, Formal Analysis and Investigation, Writing – Original Draft, Visualization. G. Egea: Conceptualization, Methodology, Formal Analysis and Investigation, Writing – Review and Editing, Supervision, Funding acquisition. C. Borrero: Conceptualization, Methodology, Writing – Review and Editing. M. Pérez-Ruiz: Conceptualization, Writing – Review and Editing, Supervision, Funding acquisition. M. Avilés: Conceptualization, Writing – Review and Editing, Supervision, Funding acquisition.
Corresponding author
Ethics declarations
Ethical approval
The Research Ethics Act guides the work with ethics at NIBIO. The research and activities in this study do not conflict with NIBIO’s ethical guidelines, which comprise personal and business conduct, research ethics, and good scientific and publication practice.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Castro-Valdecantos, P., Egea, G., Borrero, C. et al. Detection of fusarium wilt-induced physiological impairment in strawberry plants using hyperspectral imaging and machine learning. Precision Agric (2024). https://doi.org/10.1007/s11119-024-10173-6
Accepted:
Published:
DOI: https://doi.org/10.1007/s11119-024-10173-6