Abstract
This article relates epilithic dry- and wet-seasonal bacterial biofilm composition to water quality along Río de la Sabana near Acapulco, Mexico. Samples were taken from various locations including nearly pristine upland locations, adjacent to residential floodplain developments, and immediately upstream from an estuarine lagoon. Bacterial composition was identified through sequential DNA analysis at the phylum, class, order, and family levels, with most of these categorized as heterotrophs, autotrophs, denitrifiers, nitrogen fixers, pathogens, and/or potential bioremediators based on generalized literature-sourced assignments. The results were interpreted in terms of location by extent of effluent pollution, and by dry versus wet seasonal changes pertaining to biofilm composition, related bacterial functions, and the following water quality parameters: temperature, pH, dissolved oxygen, biological and chemical oxygen demand, fecal and total bacteria counts, methylene blue active substances, electrical conductivity, and nitrite, nitrate, ammonium, sulfate, and phosphate concentrations. It was found that epilithic bacterial biofilm diversity was richest during the wet season, was more varied in abundance along the upland locations, and was dominated by Proteobacteria and Bacteroidetes with bioremediation and pathogen functions along effluent-receiving river locations. Low-abundance families associated with anaerobic and denitrifying functions were more prevalent during the wet season, while low-abundance families associated with aerobic, N2-fixing and pH-elevating functions were more prevalent during the dry season.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
This article reports on epilithic bacterial biofilm composition, functions, and changes along Río de la Sabana as it flows north to south east of Acapulco, Mexico. This river and its floodplain provide high biodiversity habitats, but these are now affected by rapidly evolving urban encroachments (Arriaga-Cabrera et al. 2008). As previously reported, these encroachments have not only accelerated environmental degradation and water pollution along the river and its coastal lagoon (CONAGUA 2012; Pineda-Mora et al. 2018), but also increased flood-induced health vulnerabilities in low-lying settlements across and adjacent to the Río de la Sabana Valley (CONAGUA 2012; Rodríguez-Herrera et al. 2013).
In general, biofilms are densely populated and self-stabilizing microbiomes (Besemer 2015; Peipoch et al. 2015; Findlay and Battin 2016; Nicholls and Crompton 2017) involving algae, diatoms, fungi, bacteria, and protozoa, all enveloped in an extracellular polymeric substance matrix (EPS) on biotic and/or abiotic surfaces (Nadell et al. 2009; Villeneuve et al. 2013; Nadell et al. 2016). As such, biofilms undoubtedly affect a variety of processes and functions by way of organic matter production and decomposition, nutrient uptake and cycling, and contaminant absorption and processing (Findlay and Battin 2016; Shahsavari et al. 2017; Guasch et al. 2017; Banerjee et al. 2018). In addition, several studies have stressed that microbial biofilm compositions are subject to physical, chemical, and biological changes in water flow and quality (Pesce et al. 2010; Montuelle et al. 2010; Tlili et al. 2011; Baek et al. 2013; Langenheder et al. 2016; Bouchez et al. 2016; Hu et al. 2017). These changes can now be revealed in terms of operational taxonomic units (OTUs) using next generation sequencing (NGS) DNA technologies (Ghiglione et al. 2014; Tan et al. 2015; Pawlowski et al. 2018). With this background, this article reports on changes in bacterial biofilm composition, diversity, and functions as affected by the extent of residential effluent pollution and related seasonal influences along Río de la Sabana. The emphasis is on epilithic biofilms because their bacterial composition would directly be affected by locational changes in water quality, varying flow rates, and desiccating exposure incidences.
2 Materials and Methods
2.1 Study Area and Field Sampling
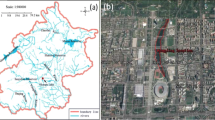
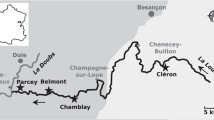
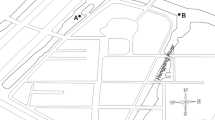
Río de la Sabana begins in the north-east portion of the Acapulco municipality, with its headwaters reaching 2000 m above sea level. Its main flow channel is 30.7 km long and widens to about 60 m at its confluence with the coastal Tres Palos lagoon (CONAGUA 2012). Its 466 km2 basin area supports permanent flow channels with a cumulative length of 727 km, and a channel density of 1.55 km−1 (Villegas-Romero et al. 2009). The steep and mostly pristine terrain of the upper basin feeds the main flow channels with short-duration runoff. In contrast, the lower part of the terrain widens into the broad and densely populated La Sabana Valley floodplain (Pineda-Mora et al. 2018).
This study was carried out in 2017 during the dry (January) and the rainy (July) season, with samples taken from the basin headwaters to the floodplain estuary (Fig. 1, Table 1) as per the following sampling zonation (Dorigo et al. 2008):
-
1.
Reference Zone (Zone 1; Locations S1, S2): located on the upper part of the sub-basin, meeting national and international standards for the protection of aquatic life, representing oligotrophic water quality conditions
-
2.
Transition Zone (Zone 2; Locations S3, S4): a peri-urban zone located between the upper and mid-low parts of the basin.
-
3.
Pollution Zone (Zone 3; Locations S5, S6): representing a densely populated area and eutrophic water quality conditions.
-
4.
Recovery Zone (Zone 4; Location S7): a confluence area that combines water from densely populated area west of the river, with water also received from the low population area in the east.
In total, 14 composite biofilm samples were gathered from submerged rock surfaces (Fig. 2): 7 per dry season, and 7 per rainy season using the point contact method (Lyautey et al. 2010). At each location, three rocks of similar type and surface area (about 15 cm in diameter) were selected from a transverse profile approximately 0.3, 0.5, and 0.7 m apart. At least 30 g were collected per sample through scraping the rocks with sterile disposable brushes and using sterile Falcon tubes for collection (50 mL). These samples were stored on ice until ready for centrifugation at 6,000 rpm for 10 min to eliminate excess water. Thereafter, the samples were stored at − 20 °C until processed for DNA extraction.
Examples of biofilm-covered rocks lifted from Location 2 representing oligotrophic upland water conditions (top), and from Location 5 representing eutrophic water conditions near the residential floodplain developments (bottom). Biofilms were found to be thicker, less translucent, and greener along Locations S4 to S7
2.2 DNA Extraction, Sequencing, and Phylogenetic Analysis
Deoxyribonucleic acid (DNA) was extracted from the biofilms according to the methodology proposed by Ceja-Navarro et al. (2010) and Navarro-Noya et al. ( 2013). Sample extractions were done in triplicate. Doing so yielded 42 DNA samples for RNA analysis at Applied Biological Materials Inc. This procedure involved two-stage PCR amplification with gen 16S rDNA V3–V4 specific primers. The genomic library was subsequently generated with an Agilent 2100 Bioanalyzer, followed by qPCR quantification. DNA sequencing was accomplished using a MiSeq 600 cycle flow cell. The resulting Bcl archives were converted into FastQC data. Sequencing quality control was performed with the FastQC tool. The resulting data were processed using the QUIIME v. 1.8.0. program, with OTUs determined using pick_de_novo_otus.py software to a similarity of > 97% against the Greengenes database (Caporaso et al. 2010). Taxonomic assignments (phylum, order, class, family) were subsequently done by means of an rRNA Bayesian classifier of the Ribosomal Data Project, up to a confidence threshold of 80% (Supplementary Data I). The relative abundance of DNA sequences (i.e., paired V3 -V4 16s rRNA sequences) identified as taxonomic units at ≥ 1% are listed in Supplementary Data II. Where possible, these units were assigned as bioremediators, pathogens, aerobes, anaerobes (including facultative anaerobes), N2-fixers, and/or denitrifiers at the family or order level through literature search (Supplementary Data III). The raw nucleotide sequence reads of this study have been deposited in NCBI under BioProject ID PRJNA643497, and can be accessed using the NCBI SRA accession id entries from SAMN15468712 to SAMN15468753 (42 entries: 3 epilithic biofilm samples taken from each of the 7 locations during the dry and rainy season).
2.3 Water Quality Sampling and Analysis
As already reported by Pineda-Mora et al. (2018), water quality was sampled at the same locations during the dry season (January) and rainy season (July) in 2017 (Table 1). Fourteen physicochemical and bacteriological water quality parameters (Table 2, Supplementary Data IV) were measured following established protocols in accordance with Mexican references (NMX) and standardized alternative methods. Temperature, pH, and electrical conductivity were measured in situ. The other parameters were analyzed in a laboratory authorized through Mexican Organization of Accreditation (AC). Temperature, pH, electrical conductivity, ammonium nitrogen, nitrates, nitrites, methylene blue active substances (anionic surfactants), sulfates, and phosphates parameters were measured in triplicate. Dissolved oxygen, biochemical oxygen demand, chemical oxygen demand, total coliforms, and fecal coliforms were determined by way of single measurements per sample.
2.4 Statistical Analyses
The bacterial relative abundance data, identified by order, class, family, and unclassified, were compiled into spreadsheets. The BiodiversityR package (Kindt and Kindt 2019) in R studio was used to determine microbial diversity indices by location and season: diversity by type (Richness, Kindt and Kindt 2019), by type and abundance (Simpson 1949), and by diversity of dominants (Berger and Parker 1970).
The sums of the relative taxonomic abundance frequencies contributing to bacterial composition, functions, biodiversity indices, and water quality parameters were all compiled by location and season. Locations S1, S2. S3, S4, S5, S6, and S7 where coded 1, 2, 3, 4, 7, 6, and 5, respectively, to reflect the overall extent of residential effluent pollution. Season was coded 0 when dry and 1 when wet. The resulting variable-to-variable association patterns of these sums were (i) factor analyzed (Principal components, 2 factors, Varimax transformation) and (ii) were subjected to multivariate regression analyses to quantify the extent to which bacterial relative OTU abundances, functions, and water quality parameters relate to each other by location and season. The resulting two-factor analyses plots were sectorized by extent of pollution and season so that Sectors I and II refer to least-polluted (S1, S2, S3) and Sectors III and IV refer to most polluted (S4, S5, S6, S7) locations during the wet and dry seasons, respectively.
3 Results
3.1 Bacterial Biofilm Abundances, Phylum Level
The > 1% relative abundances of the DNA-recognized bacterial phylum (p), class (c), order (o), or family (f) in the Río de la Sabana biofilms are presented in Supplementary Data I. Altogether, 11 entries were identified at the phylum level, 27 at the class level, and 68 entries at the family/order level. Of the ≥ 1% entries, 27 were present during the dry and wet seasons, 20 occurred during the dry season only, and 24 occurred during the wet season only.
The bar-charted phylum abundance display in Fig. 3 (i) across all locations and seasons combined, (ii) across all locations by season, and (iii) across the least to most polluted locations by season revealed the following differences: increased pollution promoted the relative abundance on Proteobacteria, Bacteroidetes, Chloroflexi, at the S4-S7 locations but decreased the relative abundances of Actinobacteria, Firmicutes, Cyanobacteria, and Thermi. During the wet season, Proteobacteria and Bacteroidetes further increased their relative dominance at the S4-S7 locations at the expense of Firmicutes and Chloroflexi.
These compositional phylum changes are further summarized in Fig. 4 by the 2-factor analysis plot that differentiates the relative abundance sums of each phylum as they fall into Sector I (Locations S1-S3; least polluted, wet season), II (Locations S1-S3; least polluted, dry season), III (Locations S4-S7; wet, most polluted), and IV (Locations S4–S7; dry, most polluted). Proteobacteria, Chloroflexi, and Synergistetes clustered along the polluted Sector III to IV transitions. Acidobacteria, Actinobacteria, and Planctomycetes prevailed more in Sector I, while Thermi, Firmicutes, and Cyanobacteria prevailed more in Sector II.
3.2 Bacterial Biofilm Abundances, Class/Family Levels
The relative abundance numbers in Tables 3 and 4, and the bar charts compiled as Supplementary Data I show how the bacterial orders and families were distributed by Sectors I to IV from the least to most polluted locations by season. Among these families, Comamonadaceae (motile aerobic Betaproteobacteria), Rhodobacteraceae (chemo-organo and phototrophic Alphaproteobacteria), Sphingomonadaceae (aerobic chemo-heterotrophic Alphaproteobacteria), and Moraxellaceae (saprophytic Gammaproteobacteria) can be seen to be the most dominant (Fig. 5). However, only Comamonadaceae and Rhodobacteraceae were present at ≥ 1% at all seven locations and both seasons (Table 3). DNA sequences associated with Moraxellaceae ≥ 1% were absent at S2 during the wet season, and absent at S1, S2, S3 during the dry season.
Most of the other families occurred at < 10% abundances, but their relative abundances were strongly affected by pollution and season, and somewhat related to their sector I to IV phylum assignments as apparent from Fig.4 and Tables 3 and 4. For example, classes and families associated with Acidobacteria, Cyanobacteria, and Planctomycetes occurred for the most part in Sector I and II (least polluted during the wet or dry seasons, respectively). In detail, the phototrophic Acaryochloridaceae, Chamasiphomnaceae, Chroococcales, Phormidiaceae, Pseudanabaenales, Stramenopiles, and Xenococcaceae members of the Cyanobacteria phylum occurred with relative abundances ≥ 1% only at the upland S1 to S4 locations (Sectors I and II), and more so during the dry than the wet season (Table 3, Fig. 5).
Factor analyzing the Tables 3 and 4 entries by extent of pollution and season revealed three clusters in the resulting 2-factor, 4-Sector plot in Fig. 6: one cluster in Sector I (least polluted, wet season), one in Sector II (least polluted, dry season), and one straddling Sectors III and IV (most polluted, with entries somewhat influenced by dry and wet season). In this, residential pollution or lack thereof also impacted the relative abundances of the bacterial OTUs. In detail, Alpha- and Betaproteobacteria occurred in all four sectors; Gammaproteobacteria were exclusively found in Sectors III and IV, while Deltaproteobacteria were considerably less abundant (Fig. 6; Supplementary Data I). While Proteobacteria dominated in Sectors III and to IV, some were also be associated with Sectors I and II, especially soil-associated Rhizobiales, Hyphomicrobiaceae, Phyllobacteriaceae, Bradyrhizobiaceae, Oxalobacteraceae, Acetobacteraceae, Phyllobacteriaceae, and Syntrophobacteraceae.
Families listed under Thermi (Trueperaceae, Deinococcaceae, Thermaceae, Erysipelotrichaceae) where present at ≥ 1% relative abundance spread across Sectors I, II, and III, while Firmicutes classes and families (i.e., Mogibacteriaceae, Clostridiales, Bacillaceae, Christensenellaceae, Peptostreptococcaceae, Ruminococcaceae, Exiguobacteraceae) spread across Sectors I to IV (Fig. 6). In contrast, Bacteriodetes families as represented by Cytophagaceae, Chitinophagaceae, Flavobacteriaceae, and Weeksellaceae were present at ≥ 1% during the wet season only. Sphingomonadaceae were present at ≥ 1% at most locations except at S5 and S6 during the dry season: hence more abundant at the non-polluted locations during the wet season (Table 3). Rhizobiales (N2-fixing microsymbiotic Alphaproteobacteria) were also present during the dry and wet season, but with a strong preference for the less polluted upper Río de la Sabana locations during the dry season.
3.3 Bacterial Biofilm Diversity
The bacterial diversity assessment results based on the Table 3 entries are listed in Table 5, by location and season as coded. Factor analyzing these results identified pollution and season as predictive factors for bacterial richness by type (Richness index, higher during the wet than the dry season), diversity abundance (Simpson index, increasing with decreasing pollution level), and dominance (Berger index, increasing with increasing pollution level). Expressed in regression terms, the resulting Factor 1 scores were related to pollution extent, and the Factor 2 scores related to season, as follows:
3.4 Bacterial Biofilm Functions
The results of summing the relative abundances of bacterial families, classes, orders, or phylum levels according to their presumed bioremediation, pathogen, aerobic, anaerobic, N2-fixing, and denitrifying functions (see Supplementary Data III) are presented Fig. 7, by season and location. These plots reveal higher relative abundances of bioremediators, pathogens, anaerobes, and denitrifiers at Locations S4 to S6. In contrast, aerobes and N2-fixers were nearly evenly distributed across the S1 to S7 locations, and registered low relative abundances during the dry and wet seasons.
3.5 Water Quality Related to Microbial Biofilm Diversity and Functions
Factor analyzing the water quality measures as listed in Supplementary Data IV in combination with the bacterial diversity results in Table 5 and the relative abundances by bacterial functions as plotted in Fig. 7 led to the following observations by way of Fig. 8:
-
1.
High fecal coliform counts, high electrical conductivities, and elevated temperatures were—for the most part—associated with the polluted locations.
-
2.
These locations were also associated with high pathogen and bioremediator occurrences, which—in turn—registered elevated biological and chemical oxygen demands (BOD, COD), elevated NH3, NO2−, NO3−, SO42− and PO43− concentrations, and low DO concentrations.
-
3.
There were also seasonal differences: pH as well as NO2−, NO3−, NH3 and SO42− concentrations scored higher during the dry season.
-
4.
The bacterial Richness index fell on Sector I, meaning that the fresh-water locations had the highest variety of OTUs during the wet season. In comparison, OTU relative abundances as per the Simpson index fell on Sector II, meaning that the relative OTU abundance numbers varied the most within the freshwater locations during the dry season. In contrast, the diversity index for dominance (Berger) fell into Sector III in association with increased effluent pollution and increased abundances of biofilm-resident denitrifiers, anaerobes, aerobes, bioremediators, and pathogens. These location- and season-specific diversity changes can also be discerned through inspecting the Supplementary Data I OTU bar charts, and most clearly so by focusing on the OTU class compilation.
-
5.
Aerobes prevailed more consistently during the dry than the wet season (Sector IV), anaerobes and denitrifiers dominated during the wet season (Sector III), while N2-fixers prevailed in Sector II.
Factor analysis plot combining the general association patterns between (i) relative bacterial biofilm taxa abundances pertaining to bioremediators, pathogens, aerobes, anaerobes, denitrifiers and N2-fixers (white white squares), (ii) water-quality parameters (Table 2; colored dots), and (iii) the bacterial diversity indices (black circles), by Sectors I to IV, with each signifying one of the four dry and wet season combinations pertaining to low and high pollution extent, as in Fig. 4
By way of regression analysis, BOD and COD varied by location and season as follows:
where locations and seasons were coded again as per Table 5. Correspondingly, dissolved oxygen levels decreased with increased biological and chemical oxygen demand, and this decrease was more pronounced during the wet season, i.e.,
Note that the relative abundance values for the aerobic and anaerobic biofilm families did not enter this formulation in a significant way. This likely related to the general expectation that aerobes and anaerobes function best when and where DO remained steady at high or low levels, respectively.
In reference to seasonal water quality changes, anaerobes including denitrifiers were prevalent under neutral pH condition. In contrast, N2-fixers and organic-matter consuming aerobes were associated with increasing pH levels likely due to (i) converting N2 to NH3 and (ii) heterotrophic release of CO2 from organic acid groups. Relating the reported pH values to season, location, and bacterial biofilm functions generated the following equation:
where N2-fixers and bioremediators were entered by their sum of relative abundance per location and season.
In terms of electrical conductivity, which indexes overall effluent concentrations, the regression analysis produced the following result:
Hence, EC levels were not only lower during the wet season (likely due to higher flow rate and subsequent dilution) but were also lower where DO levels remained high, and where pollution loads were low to absent, i.e., at Locations S1 to S3.
The best-fitted scatterplots for Eqs. 3, 5, 6, and 7 are presented in Fig. 9 to further illustrate how water quality was affected by location and season. In particular, the changes on BOD (and therefore COD as well) were categorically relatable to season, and linearly relatable to location from least to most polluted. In contrast, the changes in DO were both categorically related to location and season, while the changes in pH and EC were—for the most part—categorical by season, and clustered by location.
4 Discussion
4.1 Bacterial Changes at the Phylum Level
The overall numerical ranking sequence of phylum abundance across locations and seasons was as follows:
-
Proteobacteria > Bacterioidetes > Firmicutes > Actinobacteria > Chloroflexi > Cyanobacteria > Thermi > Acidobacteria > Planctonmycetes > Verrumicrobia.
Other stream and riverine biofilm studies also report Proteobacteria, Bacterioides, and Firmicutes to be dominant, followed in abundance by Cyanobacteria, Actinobacteria, Chloroflexi, and Verrumicrobia (e.g., Peipoch et al. 2015; Fang et al. 2017; Lin et al. 2019). Remarkably, increasing pollution levels rendered Proteobacteria, Bacterioides, Firmicutes, and Chloroflexi to dominate all the other biofilm bacteria at > 98% relative abundance during the dry and the wet season. In addition, residential effluents rendered Proteobacteria and Bacterioides to become even more dominant during the wet season through diminishing the relative abundances of Firmicutes and Chloroflexi. Among the Proteobacteria, there was a further pollution-induced dry-to-wet seasonal shift towards greater Betaproteobacteria than Gammaproteobacteria dominance. This observation parallels the relative abundance of Proteobacteria in active sludge as reported by Jiang et al. (2008):
Betaproteobacteria 39.8% > Gammaproteobacteria 20.15% > Alphaproteobacteria 6.79% > Deltaproteobacteria 4.85%.
Hence, Alphaproteobacteria would dominate biofilms in low nutrient waters such as those found at the S1, S2, and S3 locations, and heterotrophic Betaproteobacteria and Gammaproteobacteria would dominate and co-dominate polluted waters such as those found at the S4, S5, S6, and S7 locations.
At the less polluted S1 to S3 locations, Actinobacteria and Planctomycetes became more prevalent from the dry to the wet season, while the opposite occurred with Fermicutes and Thermi. These changes were likely due to (i) higher detrital availabilities that would increase Actinobacteria and Planctomycetes activities during the wet season while (ii) Fermicutes and heat-resistant Thermi would be more adapted to persist on rocky surfaces during the dry season when exposed to periods of intermittent desiccation and heat stress (Marxsen et al. 2010).
The low presence of Cyanobacteria at Locations S4 to S7 was likely caused by pollution-enhanced and light-blocking green algae growth, and this would especially be so under continuous flow conditions, as reported by Sabater et al. (2016). In addition, lower Cyanobacteria abundance at the polluted locations may also be related to high molar N:P ratios, as reported by Peipoch et al. (2019). Determining whether this also applies at the Río de la Sabana sampling locations led to the following regression result, at least in a contributing sense:
with Pollution extent and Season coded as in Table 5. Hence, Cyanobacteria abundance was lower during the wet season and at the polluted locations and was also somewhat reduced by increasing molar N:P ratios. These ratios varied from 48 to 420, were significantly lower during the wet season (p = 0.01), but were not significantly affected by location (p = 0.17).
Relating the sum of bacterial abundances at the family, order, or phylum level as listed in Tables 3 and 4 to the water quality measures by location and season did not produce significant regression results, except for Bacteriodetes and Proteobacteria, as follows:
where Location is coded as in Table 5. These equations and their best-fitted scatterplots in Fig. 10 indicate that the epilithic biofilm presence of Bacteriodetes decreases with increasing pH and increasing DO levels, while the presence of Proteobacteria increases with increasing effluents of residential pollutants.
4.2 Bacterial Changes at the Order/Family Level
Of the 68 taxonomic units identified, bioremediators belonging to the Comamonodaceae, Rhodobacteriae, and Moraxellaceae families dominated at locations S5 and S6, likely due to increased effluent/pollution levels. In contrast, biofilm-forming Sphingomonadaceae (Glaeser and Kämpfer 2014) and biofilm-supporting Rhizobiaceae were more abundant along the upland streams (Figs. 2 and 3). Despite high coliform counts at Locations S5 and S6 (Supplementary Data IV), relative DNA sequences symptomatic of Enterobacteriaceae generally remained low at < 1% at all biofilm sampling locations. This would in part be due to low survival and growth of Escherichia coli biofilms on non-living surfaces (Beloin et al. 2008; Abberton et al. 2016), where Comamonodaceae, Rhodobacteriae, Moraxellaceae, and Sphingomonadaceae generally dominate (Widder et al. 2014).
Many of the bacterial families that were present but in low abundance tended to be (i) aerobic at Locations S1 to S3 (e.g., Ellin6075, Ellin6529, Acidimicrobiales, Actinomycetales, Gaiellaceae) and (ii) anaerobic at Locations S5 to S7 (e.g., Bacteroidales, Flavobacteriaceae, Desulfobulbaceae, Chromatiaceae). These prevalence differences would be in line with the corresponding stream-to-river water aeration and pollution-related changes as discussed above in reference to Sectors I to IV. Nevertheless, aerobic families such as Comamonadaceae, Rhodobacteraceae, Rhodocyclaceae, Weeksellaceae, and Xanthomonadaceae would function well at Locations S4 to S7 as aerobic sludge and waste-water treatment agents. Also of note are the contrasting occurrences of Exiguobacteriales (i.e., Bacilli members of the Firmicutes phylum: highly temperature-adaptive heterotrophs, see White et al. ( 2019) and Weeksellaceae (i.e., Flavobacteriales members of the Bacteroidetes phylum). Both are known to be aerobic to facultative intestinal chemo-organotrophs (Wikipedia 2020): Exiguobacteriales were only present at ≥ 1% during the dry season (Table 4), but Weeksellaceae—similar to Moraxellaceae—were mostly present at the S4, S5, and S6 locations during the wet season (Table 2).
The simultaneous biofilm presence of aerobes and anaerobes attest to spatial biofilm self-arrangements: aerobes on the outside, anaerobes on the inside. Furthermore, the biofilm dominance of the aerobic families would likely have contributed—at least in part—to the dry seasonal increases on water pH due to progressive alkalinity-revealing consumption of organic matter (Kayombo et al. 2002; Bernal et al. 2008; Abdullahi et al. 2008). The quantitative extent to which the epilithic biofilms affected water quality from Locations S1 to S7, however, remains unknown since that would depend on, e.g., incoming effluent loads, flow rates, and combined biofilm- and plankton-based organic matter consumption and mineralization (Romaní et al. 2004). A recent study reported an overall 14% biofilm uptake of dissolved organic matter from an agricultural stream, versus 4% from a forest stream (Graeber et al. 2019).
While there are significant correlations between bacterial biofilm diversities, functions and relative phylum, class, order, family abundances by location and/or season (Figs. 4, 6, 8, 9, 10), it is difficult to quantitatively relate specific water quality measures to bacterial biofilm functions. In part, this refers to the fact that biofilms function synergistically as self-stabilizing microbiomes with their own overall input, output, and life-supporting requirements (Peipoch et al. 2015; Toyofuku et al. 2016; Zeglin 2015). To illustrate, one would have expected that the measured changes in water pH and DO would have influenced bacterial abundances at the individual phylum, order, and family levels, but only Bacteriodetes were so correlated. In part, Bacteriodetes may be more sensitive to increasing pH and DO levels (Eq. 9) because of their anaerobic requirements and inability to produce spores. In general, however, biofilm-internal pH and DO values likely did not correlate with biofilm-external pH and DO measurements since the former would be the result of balanced microbial biofilm coexistences. For example, biofilm internal DO levels would result from combined O2 producing and reducing activities, and biofilm internal pH levels would depend on (i) the organic acidity of biofilm structures and cellular exudates, and (ii) the mix of cellular nutrient cation and anion uptake, respectively.
To discern which effects on water quality were brought about by specific family functions by location and season would be difficult due to wide phenotypic variabilities per bacterial family. In this regard, Comamonadaceae represent 29 genera within an overall 16S rRNA gene sequence similarity of 93–97%. Yet, these genera differ by function as aerobic organotrophs, anaerobic denitrifiers, Fe3+-reducers, hydrogen oxidizers, photoautotrophs, photoheterotrophs, and fermenters (Willems 2014a). Rhodobacteraceae are even more diverse, containing 100 genera involving aerobic photo- and chemo-heterotrophs, with some performing photosynthesis under anaerobic conditions. Characteristically, members of this family facilitate sulfur and carbon biogeochemical cycling, often in symbiosis with aquatic micro- and macro-organisms (Pujalte et al. 2014). Moraxellaceae include 5 genera with non-fermentative saprophytic functions, with some species recognized as pathogens (Teixeira and Merquior 2014). Sphingomonadaceae, known for their important roles as bioremediators, involve 13 genera with facultative to anaerobic fermentation functions in soils, freshwater systems, sludge, rhizospheres, and phyllospheres (Glaeser and Kämpfer 2014). Rhizobiales are primarily involved with N2-fixation, with some forming root nodules while others dominate in tap-water biofilms, with some being pathogenic (Potter 2013; Wang et al. 2018), and some being pathogen controlling. Pathogen-controlling species are associated with Actinobacteria (Wink et al. 2017), with Acidimicrobiales, Actinomycetales, and Nocardioidaceae occurring with ≥ 1% abundances at the S1, S2, and S3 locations of this study.
To reveal further details about positive and negative bacterial interactions in addition to those surmised above along Río de la Sabana locations, it would be necessary to conduct multiple location- and season-specific DNA sampling studies. In addition, such an effort should be expanded to include other biofilm constituents, e.g., green algae and diatoms (Kouzuma and Watanabe 2015; Ramanan et al. 2016). The focus on epilithic biofilms could also be expanded to biofilms residing on other mineral surfaces (e.g., sediments) and submerged macrophytes. Also, valuable would be to analyze how dissolved and total elemental concentrations in biofilms relate to corresponding changes in water quality.
5 Conclusions
The above results show how bacterial biofilm composition, functions, and water quality measures varied along Río de la Sabana from the least to most polluted locations and by season. The main effect of residential effluent discharge expressed itself through biofilm thickening caused by green algae growth and a bacterial dominance shift towards heterotrophic Proteobacteria (Comamonadaceae, Moraxellaceae, Rhodobacteraceae) and Bacteroides (Weeksellaceae). This dominance shift towards common wastewater and sludge OTUs was most pronounced during the wet season and occurred on the expense of the ≥1% relative abundance of most other families. By bacterial biofilm function and water quality associations, Locations S5 and S6 registered the highest levels of bioremediators and pathogens. Water-sampled fecal coliform counts, biological and chemical oxygen demands, electrical conductivity, and NH3, NO3−, NO2−, PO43− and SO42−, concentrations were also generally highest at these locations but more so during the dry than the wet season. Dissolved oxygen levels registered highest at the least pollution locations (S1 and S2) during the wet and dry season. In contrast, pH remained near neutral during the wet season but increased to alkaline levels during the dry season. Apart from general factor-analyzed location and seasonal association patterns between bacterial composition, diversity, functions and water quality measures, correlational changes in relative abundance for any particular OTU could only be discerned for Bacteriodetes with changing pH and DO levels in sampled stream and river water.
References
Abberton, C. L., Bereschenko, L., van der Wielen, P. W. J. J., & Smith, C. J. (2016). Survival, biofilm formation, and growth potential of environmental and enteric Escherichia coli strains in drinking water microcosms. Applied and Environment Microbiology, 82, 5320–5331.
Abdullahi, Y. A., Akunna, J. C., White, N. A., Hallett, P. D., & Wheatley, R. (2008). Investigating the effects of anaerobic and aerobic post-treatment on quality and stability of organic fraction of municipal solid waste as soil amendment. Bioresource Technology, 99, 8631–8636.
Arriaga-Cabrera, L., Aguilar-Sierra, J., Alcocer-Durand, R., Jiménez-Rosenberg, E., Muñoz-López, E., & Vázquez-Domínguez, E. (2008). Regiones hidrológicas prioritarias. Comisión Nacional para el Conocimiento y Uso de la Biodiversidad, CONABIO.
Baek, S., Son, M., & Shim, W. (2013). Effects of chemically enhanced water-accommodated fraction of Iranian heavy crude oil on periphytic microbial communities in microcosm experiment. Bulletin of Environmental Contamination & Toxicology, 5(90), 605–610.
Banerjee, S., Schlaeppi, K., & Heijden, M. (2018). Keystone taxa as drivers of microbiome structure and functioning. Nature Reviews Microbiology, 16(9), 567–557.
Beloin, C., Roux, A., & Ghigo, J.-M. (2008). Escherichia coli biofilms. Current Topics in Microbiology and Immunology, 322, 249–289.
Berger, W. H., & Parker, F. L. (1970). Diversity of planktonic foraminifera in deep-sea sediments. Science, 3937(168), 1345–1347.
Bernal, C. B., Vázquez, G., Quintal, I. B., & Bussy, A. L. (2008). Microalgal dynamics in batch reactors for municipal wastewater treatment containing dairy sewage water. Water, Air, and Soil Pollution, 190, 259–279.
Besemer, K. (2015). Biodiversity, community structure and function of biofilms in stream ecosystems. Research in Microbiolgy, 166(10), 774–781.
Bouchez, T., Blieux, A., Dequiedt, S., Domaizon, I., Dufresne, A., Ferreira, S., et al. (2016). Molecular microbiology methods for environmental diagnosis. Environmental Chemistry Letters, 4(14), 423–441.
Caporaso, G., Kuczynski, J., Stombaugt, J., Bittinger, K., Bushman, F., Costello, E., et al. (2010). QIIME allows analysis of high through put community sequencing data. Nature Methods, 7(5), 335–336.
Ceja-Navarro, J. A., Rivera-Orduña, F. N., Patiño-Zúñiga, L., Vila-Sanjurjo, A., Crossa, J., Govaerts, B., et al. (2010). Phylogenetic and multivariate analyses to determine the effects of different tillage and residue management practices on soil bacteria communities. Applied and Environmental Microbiology, 76(11), 3685–3691.
CONAGUA (2012) Plan operativo general: Proyecto de suministro de agua potable y saneamiento de las zonas marginadas del valle de la Sabana en el estado de Guerrero. Acapulco, México. Comisión de Agua Potable, Alcantarillado y Saneamiento del Estado de Guerrero.
Dorigo, U., Lefranc, M., Leboulanger, C. H., Montuelle, B., & Humbert, J. (2008). Spatial heterogeneity of periphytic microbial communities in a small pesticide-polluted river. FEMS Microbiology Ecology, 67(3), 491–501.
Fang, H., Chen, Y., Huang, L., & Guojian, H. (2017). Analysis of biofilm bacterial communities under different shear stresses using size-fractionated sediment. Science Reports, 7, 1299. https://doi.org/10.1038/s41598-017-01446-4.
Findlay, R., & Battin, T. (2016). The microbial ecology of benthic environments. In M. Yates, C. Nakatsu, R. Miller, & S. Pillai (Eds.), Manual of Environmental Microbiology (pp. 421.1–421.20). ASM Press.
Ghiglione, J., Martin-Laurent, F., Stachowski-Haberkorn, S., Pesce, S., & Vuilleumier, S. (2014). The coming of age of microbial ecotoxicology: Report on the first two meetings in France. Environmental Science and Pollution Research, 24(21), 14241–14245.
Glaeser, S. P., & Kämpfer, P. (2014). The Family Sphingomonadaceae. In E. Rosenberg, E. F. DeLong, S. Lory, E. Stackebrandt, & F. Thompson (Eds.), The Prokaryotes (pp. 641–707). Berlin: Springer.
Graeber, D., Gücker, B., Wild, R., Anlanger, C., Kamjunke, N., Norf, C., et al. (2019). Biofilm-specific uptake does not explain differences in whole-stream DOC tracer uptake between a forest and an agricultural stream. Biogeochemistry, 144, 85–101.
Guasch, H., Bonet, B., Bonnineau, C., & Barral, L. (2017). Microbial biomarkers. In C. Cravo-Laureau, C. Cagnon, B. Lauga, & R. Duran (Eds.), Microbial Ecotoxicology (pp. 251–274). Springer International Publishing AG.
Hu, A., Ju, F., Hou, F., Li, J., Yang, X., Wang, H., Mulla, S., et al. (2017). Strong impact of anthropogenic contamination on the co-occurrence patterns of a riverine microbial community. Environmental Microbiology, 12(19), 4993–5009.
Jiang, X., Ma, M., Li, J., Lu, A., & Zhong, Z. (2008). Bacterial diversity of activesludge in wastewater treatment plant. Earth Science Frontiers, 15, 163–168.
Kayombo, S., Mbwette, T. S. A., Mayo, A. W., Katima, J. H. Y., & Jørgensen, S. E. (2002). Diurnal cycles of variation of physical–chemical parameters in waste stabilization ponds. Ecological Engineering, 18, 287–291.
Kindt, R., & Kindt, M. R. (2019). Package for community ecology and suitability analysis.
Kouzuma, A., & Watanabe, K. (2015). Exploring the potential of algae/bacteria interactions. Current Opinion in Biotechnology, 33, 125–129.
Krieg, R. N., Ludwing, W., Euzéby, J., & Whitman, W. B. (2010). Order Bacteroidales. In N. R. Krieg, J. T. Staley, D. R. Brown, B. P. Hedlund, B. J. Paster, & N. L. Ward (Eds.), Bergey’s manual of systematic bacteriology (pp. 25–26). Springer.
Kuever, J. (2014). The Family Desulfobulbaceae. In E. Rosenberg, E. F. DeLong, S. Lory, E. Stackebrandt, & F. Thompson (Eds.), The Prokaryotes (pp. 75–86). Berlin: Springer.
Langenheder, S., Wang, J., Karjalainen, S., Laamanen, T., Tolonen, K., Vilmi, A., & Heino, J. (2016). Bacterial metacommunity organization a highly–connected aquatic system. FEMS Microbiology Ecology, 4(93), 1–30.
Lin, Q., Sekar, R., Marrs, R., & Zhang, Y. (2019). Effect of river ecological restoration on biofilm microbial community composition. Water, 11(6), 1244.
Lyautey, E., Bouletreau, S., Madigou, E., & Garabetian, F. (2010). Bacterial community succession in natural river biofilm assemblages. Microbial Ecology, 5(4), 589–601.
Marxsen, J., Zoppini, A., & Wilczek, S. (2010). Microbial communities in streambed sediments recovering from desiccation. FEMS Microbiology Ecology, 71, 374–386.
Montuelle, B., Dorigo, U., Bérard, A., Volat, B., Bouchez, A., Tlili, A., Gouy, V., et al. (2010). The periphyton as a multimetric bioindicator for assessing the impact of land use on rivers: an overview of the Ardieres – Morcille experimental watershed (France). In R. J. Stevenson & S. Sabater (Eds.), Global change and river ecosystems – implications for structure, function and ecosystem services (pp. 123–141). Dordrecht: Springer.
Nadell, C., Xavier, J., & Foster, K. (2009). The sociobiology of biofilms. FEMS Microbiology Reviews, 1(33), 206–224.
Nadell, C., Drescher, K., & Foster, K. (2016). Spatial structure, cooperation and competition in biofilms. Nature Reviews Microbiology, 9(14), 589–600.
Navarro-Noya, Y. E., Suárez-Arriaga, M. C., Rojas-Valdes, A., Montoya-Ciriaco, N. M., Gómez-Acata, S., Fernández-Luqueño, F., & Dendooven, L. (2013). Pyrosequencing analysis of the bacterial community in drinking water wells. Microbial ecology, 66(1), 19-29.
Nicholls, S., & Crompton, L. (2017). The effect of rivers, streams, and canals on property values. River Research and Applications, 9(33), 1–10.
Pawlowski, J., Kelly-Quinn, M., Altermatt, F., Apothéloz-Perret-Gentil, L., Beja, P., Boggero, A., Borja, A., et al. (2018). The future of biotic indices in the ecogenomic era: integrating (e) DNA metabarcoding in biological assessment of aquatic ecosystems. Science of the Total Environment, 1(637–638), 1295–1310.
Peipoch, M., Jones, R., & Valett, H. M. (2015). Spatial patterns in biofilm diversity across hierarchical levels of river-floodplain landscapes. PLoS One, 10(12), 1–14.
Peipoch, M., Miller, S. R., Antao, T. R., & Valett, H. M. (2019). Niche partitioning of microbial communities in riverine floodplains. Scientific Reports, 9, 16384. https://doi.org/10.1038/s41598-019-52865-4.
Pesce, S., Margoum, C., & Montuelle, B. (2010). In situ relationships between spatio-temporal variations in diuron concentrations and phototrophic biofilm tolerance in a contaminated river. Water Research, 44, 1941–1949.
Pineda-Mora, D., Toribio-Jiménez, J., Leal-Ascencio, T., Juárez-López, A., González-González, J., Ruvalcaba-Ledezma, J., et al. (2018). Emerging water quality issues along Rio de la Sabana, Mexico. Journal of Water Resource and Protection, 1, 621–636.
Potter, M. E. (2013). Brucellosis. In J. G. Morris & M. E. Potter (Eds.), Foodborne infections and intoxications (pp. 239–250). Elsevier.
Pujalte, M. J., Lucena, T., Ruvira, M. A., Arahal, D. R., & Macián, M. C. (2014). The Family Rhodobacteraceae. In E. Rosenberg, E. F. DeLong, S. Lory, E. Stackebrandt, & F. Thompson (Eds.), The Prokaryotes (pp. 439–512). Berlin: Springer.
Ramanan, R., Kim, B.-H., Cho, D.-H., Oh, H. M., & Kim, H. S. (2016). Algae–bacteria interactions: evolution, ecology and emerging applications. Biotechnology Advances, 34, 14–29.
Rodríguez-Herrera, A. L., Olivier-Salomé, B., López-Velasco, R., Barragán-Mendoza, M., Cañedo-Villareal, R., & Valera-Pérez, M. A. (2013). La contaminación y riesgo sanitario en zonas urbanas de la subcuenca del río de la Sabana, ciudad de Acapulco, México. Revista Gestión y Ambiente, 1(16), 85–96.
Romaní, A. M., Guasch, H., Muñoz, I., Ruana, J., Vilalt, E., et al. (2004). Biofilm structure and function and possible implications for riverine DOC dynamics. Microbial Ecology, 47, 316–328.
Sabater, S., Timoner, X., Borrego, C., & Acuña, V. (2016) Stream biofilm responses to flow intermittency: from cells to ecosystems. Frontiers in Environmental Science 4 1-14 https://www.frontiersin.org/article/10.3389/fenvs.2016.00014.
Sakamoto, M. (2014). The Family Porphyromonadaceae. In E. Rosenberg, E. F. DeLong, S. Lory, E. Stackebrandt, & F. Thompson (Eds.), The Prokaryotes (pp. 811–824). Berlin: Springer.
Shahsavari, E., Aburto-Medina, A., Khudur, L., Taha, M., & Ball, A. (2017). From microbial ecology to microbial ecotoxicology. In C. Cravo-Laureau, C. Cagnon, B. Lauga, & R. Duran (Eds.), Microbial Ecotoxicology (pp. 17–31). Cham: Springer.
Simpson, E. H. (1949). Measurement of diversity. Nature, 163(4148), 688.
Stackebrandt, E. (2014). The family Acidimicrobiaceae. In E. Rosenberg, E. F. DeLong, S. Lory, E. Stackebrandt, & F. Thompson (Eds.), The Prokaryotes (pp. 5–12). Berlin: Springer.
Tan, B., Ng, C., Nshimyimana, J. P., Loh, L. L., Gin, K., & Thompson, J. R. (2015). Next-generation sequencing (NGS) for assessment of microbial water quality: current progress, challenges, and future opportunities. Frontiers in microbiology, 6, 1027.
Teixeira, L. M., & Merquior, V. L. C. (2014). The family Moraxellaceae. In E. Rosenberg, E. F. DeLong, S. Lory, E. Stackebrandt, & F. Thompson (Eds.), The Prokaryotes (pp. 443–476). Berlin: Springer.
Tlili, A., Marechal, M., Montuelle, B., Volat, B., Dorigo, U., & Bérard, A. (2011). Use of the MicroResp™ method to assess pollution-induced community tolerance to metals for lotic biofilms. Environmental Pollution, 1(159), 18–24.
Toyofuku, M., Inaba, T., Kiyokawa, T., Obana, N., Yawata, Y., & Nomura, N. (2016). Environmental factors that shape biofilm formation. Bioscience, Biotechnology, and Biochemistry, 80, 7–12.
Villegas-Romero, I., Oropesa-Mota, J. L., Martínez-Ménes, M., & Mejía-Sáenz, E. (2009). Trayectoria y relación lluvia-escurrimiento causados por el huracán paulina en la cuenca del Río La Sabana, Guerrero, México. Agrociencia, 43, 345–356.
Villeneuve, A., Montuelle, B., Pesce, S., & Bouchez, A. (2013). Environmental river biofilms as biological indicators of the impact of chemical contaminants. In J. Férard & C. Blaise (Eds.), Encyclopedia of Aquatic Ecotoxicology (pp. 443–455). Dordrecht: Springer.
Wang, F., Li, W., Li, Y., Zhang, J., Chen, J., Zhang, W., & Wu, X. (2018). Molecular analysis of bacterial community in the tap water with different water ages of a drinking water distribution system. Frontiers of Environmental Science & Engineering, 3(12), 1–10.
Wei, Z., Hu, X., Li, X., Zhang, Y., Jiang, L., Li, J., et al. (2017). The rhizospheric microbial community structure and diversity of deciduous and evergreen forests in Taihu Lake area, China. PLoS One, 12, e0174411. https://doi.org/10.1371/journal.pone.0174411.
White III, R. A., Soles, S. A., Gavelis, G., Gosselin, E., Slater, G. G., Lim, D. S. S., et al. (2019). The complete genome and physiological analysis of the eurythermal Firmicute Exiguobacterium chiriqhucha strain RW2 isolated from a freshwater microbialite, widely adaptable to broad thermal, pH, and salinity ranges. Frontiers in Microbiology, 9, 34189.
Widder, S., Besemer, K., Singer, G.A., Ceola, S., Bertuzzo, E., Quince, C., et al. 2014. Fluvial network organization imprints on microbial co-occurrence networks. Proceedings of the National Academy of Sciences of the United States of America (PNAS) 111:12799–12804.
Wikipedia (2020). Bacteroidetes, https://en.wikipedia.org/wiki/Bacteroidetes. Retrieved 15:10, February 17, 2020.
Willems, A. (2014a). The Family Comamonadaceae. In E. Rosenberg, E. F. DeLong, S. Lory, E. Stackebrandt, & F. Thompson (Eds.), The Prokaryotes (pp. 777–851). Berlin: Springer.
Wink, J., Mohammadipanah, F., & Hamedi, J. (2017). Biology and biotechnology of actinobacteria. Springer. 395 pp. https://doi.org/10.1007/978-3-319-60339-1.
Zeglin, L. H. (2015). Stream microbial diversity in response to environmental changes: review and synthesis of existing research. Frontiers in Microbiology, 18. https://doi.org/10.3389/fmicb.2015.00454.
Acknowledgements
We thank Perla E. Hernández, Augusto Rojas, Enrique J. Flores, Manuel Ramírez, Luis A. Cornejo, Jair J. Pineda, Miguel A. Santos, Noemi Cortez and the Applied Biological Materials Inc. team, especially Jeffrey Chu, director of the NGS Department, for sample processing and the DNA data analyses.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 223 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pineda-Mora, D., Juárez-López, A.L., Toribio-Jiménez, J. et al. Diversity and Functions of Epilithic Riverine Biofilms. Water Air Soil Pollut 231, 391 (2020). https://doi.org/10.1007/s11270-020-04692-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-020-04692-x